Summary
The efficacy of chimeric antigen receptor (CAR)-engineered T cell therapy is suboptimal in most cancers, necessitating further improvement in their therapeutic actions. However, enhancing antitumor T cell response inevitably confers an increased risk of cytokine release syndrome associated with monocyte-derived interleukin-6 (IL-6). Thus, an approach to simultaneously enhance therapeutic efficacy and safety is warranted. Here, we develop a chimeric cytokine receptor composed of the extracellular domains of GP130 and IL6RA linked to the transmembrane and cytoplasmic domain of IL-7R mutant that constitutively activates the JAK-STAT pathway (G6/7R or G6/7R-M452L). CAR-T cells with G6/7R efficiently absorb and degrade monocyte-derived IL-6 in vitro. The G6/7R-expressing CAR-T cells show superior expansion and persistence in vivo, resulting in durable antitumor response in both liquid and solid tumor mouse models. Our strategy can be widely applicable to CAR-T cell therapy to enhance its efficacy and safety, irrespective of the target antigen.
Keywords: adoptive immunotherapy, chimeric antigen receptor, CAR-T, cytokine release syndrome, neurotoxicity, IL-6, IL-7, JAK-STAT signaling, hematological malignancy, solid tumor
Graphical abstract

Highlights
-
•
A chimeric cytokine receptor G6/7R captures IL-6 via extracellular IL-6 receptors
-
•
G6/7R CAR-T cells show constitutively active IL-7 signaling
-
•
G6/7R promotes CAR-T cell proliferation and effector functions
-
•
G6/7R enhances antitumor efficacy of CAR-T cells in vivo
Yoshikawa et al. develop a chimeric cytokine receptor (G6/7R) composed of extracellular IL-6 receptors and intracellular IL-7 receptor with a constitutively active mutation. G6/7R-expressing CAR-T cells significantly reduce the proinflammatory cytokine IL-6 in vitro and demonstrate durable antitumor efficacy in multiple tumor models.
Introduction
Adoptive immunotherapy using chimeric antigen receptor (CAR)-engineered T cells has shown unprecedented efficacy against relapsed or refractory B cell malignancies and multiple myeloma.1,2,3,4,5 However, its long-term efficacy is not yet optimal, and objective clinical responses have not been observed in most cases following CAR-T cell therapy against solid tumors. Analysis of the clinical data indicates that improving the peak expansion and/or long-term persistence of the infused CAR-T cells can enhance the long-term efficacy of therapy.1,6,7,8 To this end, multiple strategies have been developed to augment the proliferative and survival capacity of CAR-T cells. For example, the incorporation of cytokine signaling significantly improves durable therapeutic efficacy by increasing both effector functions and the persistence of infused CAR-T cells.9,10,11,12,13,14,15
In addition to augmenting therapeutic efficacy, safety issues associated with CAR-T cell infusion need to be addressed. Cytokine release syndrome (CRS) and immune effector cell-associated neurotoxicity syndrome (ICANS) are two of the most frequent and sometimes serious side effects that occur irrespective of the target antigen.16 Recent studies have demonstrated that interleukin-6 (IL-6) and IL-1 derived from monocyte-lineage cells, including macrophages, are centrally associated with the development of CRS and ICANS, respectively.17,18 The use of tocilizumab, a monoclonal antibody against the IL-6 receptor, has been established as a treatment for CRS in clinical settings.19 However, the median time from treatment initiation to response is around 4 days, during which patients require intensive care in case any adverse symptoms manifest.20 In addition to pretreatment tumor burden, efficient expansion of infused CAR-T cells is associated with high-grade CRS development, suggesting that the enhancement of CAR-T cell functions could inevitably confer an increased risk for severe CRS.21,22,23,24,25
Preemptive or prophylactic use of tocilizumab before CAR-T cell infusion showed a promising result in ameliorating the severity of CRS.26,27 Moreover, the efficacy of anakinra (an IL-1R antagonist) in mitigating ICANS has been tested.28,29 However, these therapeutics cannot completely abrogate high-grade adverse events. Incorporating mechanisms that can suppress the activity of IL-6 and IL-1 in CAR-T cells will provide a substantial advantage in ameliorating the risk of severe CRS. This study aimed to simultaneously address the aforementioned challenges by developing a chimeric cytokine receptor with dual functionalities.
Results
Development of a chimeric receptor that sequesters IL-6 and induces constitutive IL-7 signaling
To simultaneously address the risk of CRS and loss of durable therapeutic efficacy in CAR-T cell therapy, we designed a chimeric fusion receptor with dual functionality (Figure 1A; Table S1). We used the extracellular domain of IL6 receptor alpha chain (IL6RA) or the fused domains of GP130 (also known as IL6R beta chain) and IL6RA in order to capture IL-6 with a higher affinity.30 Among the several cytokines associated with T cell functions, activation of IL-7 signaling provides CAR-T cells with a durable antitumor efficacy.9,13,31 To ensure stable IL-7 signaling, we linked the transmembrane and intracellular domains of the IL-7 receptor alpha chain (IL7RA) that contains a constitutively active mutant (insertion of the amino acids PPCL) to the extracellular domain of IL6R (6/7R or G6/7R).9,32 Both chimeric receptors were successfully expressed on the cell surface at comparable levels upon retroviral transduction (Figure S1A). As shown in Figure 1B, the G6/7R-expressing anti-CD19 CAR-T cells captured recombinant IL-6 more efficiently than the 6/7R CAR-T cells. Cell surface expression of the G6/7R quickly downregulated following the addition of IL-6 and then recovered, likely due to the internalization of IL-6 through the chimeric receptor (Figures S1B and S1C). To elucidate the dynamic changes in IL-6 localization, we measured extracellular and intracellular levels of IL-6 at different time points (Figure 1C). Recombinant IL-6 supplemented in the culture medium started to decrease as early as 1 h after addition (Figure 1D). In marked contrast, intracellular IL-6 levels were elevated until 3 h and subsequently decreased, suggesting the degradation of IL-6 absorbed into anti-CD19 CAR-T cells (Figure 1E). We also analyzed the absorption of IL-6 through the chimeric receptor in vivo. NOD-SCID IL2rgnull (NSG) mice engrafted with the CD19+ B cell leukemia cell line NALM-6 were treated with anti-CD19 CAR or anti-CD19 CAR+G6/7R T cells and infused with recombinant IL-6 (Figure 1F). Although plasma IL-6 concentration progressively decreased due to its short half-life,33 the mice infused with G6/7R anti-CD19 CAR-T cells showed reduced IL-6 levels at each time point, suggesting that G6/7R CAR-T cells effectively capture circulating IL-6 in vivo (Figure 1G).
Figure 1.
Construction of a chimeric receptor that captures IL-6 into T cells
(A) A schematic diagram of the chimeric receptor developed in this study. Extracellular IL6R or fusion domains of GP130-IL6R were combined with constitutively active IL7R containing PPCL residues to form 6/7R or G6/7R, respectively (SP: signal peptide).
(B) T cells transduced with anti-CD19 CAR alone or anti-CD19 CAR and 6/7R or G6/7R were incubated in the presence of recombinant IL-6 (3,000 pg/mL). The concentration of IL-6 within the culture supernatants was analyzed 48 h later (n = 3 cultures, one-way ANOVA with multiple comparison test). Representative data of two independent experiments.
(C–E) Anti-CD19 CAR-T cells with or without ectopic expression of G6/7R were treated with 40 ng of recombinant IL-6. Extracellular (D) and intracellular (E) concentrations of IL-6 at the indicated time points were measured by ELISA (n = 3 cultures, unpaired two-tailed Student’s t test; ND, not detected).
(F) Experimental design demonstrating the infusion of T cells engineered with anti-CD19 CAR alone or with anti-CD19 CAR and G6/7R into NALM6-bearing NSG mice followed by intravenous administration of recombinant IL-6.
(G) Plasma IL-6 levels were quantified at the indicated time points (n = 7 mice, unpaired two-tailed Student’s t test of the log-transformed values).
In (B), (D), (E), and (G), horizontal lines denote mean values. See also Figure S1 and Table S1.
G6/7R contributes to constitutive JAK-STAT signaling in CD19-targeting CAR-T cells
We next analyzed how the chimeric cytokine receptor affected anti-CD19 CAR-T cell functions. Phosphorylation of STAT5, the main downstream signaling molecule of IL-7, was significantly elevated in the G6/7R CAR-T cells under cytokine-free conditions irrespective of IL-6 exposure (Figures 2A and 2B). STAT3 phosphorylation was also elevated in the resting condition and slightly increased upon IL-6 treatment, suggesting that IL-6 engagement activated downstream signaling likely through the interaction of the chimeric receptor with endogenous GP130. To verify transcriptional changes induced by IL-7 signaling in G6/7R CAR-T cells, we compared gene expression profiles of anti-CD19 CAR alone or anti-CD19 CAR+G6/7R-expressing T cells with or without treatment with IL-6 through RNA-seq analysis. Overall, 1,956 genes were differentially expressed among the four groups (false discovery rate [FDR] < 0.01). Principal component analysis clearly segregated standard CAR-T cells and G6/7R CAR-T cells, while IL-6 treatment had a minor impact on gene expression signature (Figures 2C and 2D). Genes upregulated in G6/7R CAR-T cells included canonical STAT3/5 target genes (CISH and SOCS3), those essential for effector functions (GZMB, FOSL1, and FOLS2), and antiapoptotic genes (BCL2 and BCL3), suggesting the association of the transcriptional changes with potent and constitutive activation of the JAK-STAT pathway (Figure S2A). This was also corroborated by gene set enrichment analysis (GSEA), which demonstrated the significant enrichment of the gene sets upregulated upon IL-7 treatment in G6/7R anti-CD19 CAR-T cells (Figure 2E).
Figure 2.
Constitutive IL-7 signaling confers a proliferative advantage to anti-CD19 CAR-T cells
(A and B) T cells transduced with anti-CD19 CAR alone or in combination with 6/7R or G6/7R were rested overnight in cytokine-free media and left untreated or treated with 10 ng/mL IL-6 for 30 min. Representative fluorescence-activated cell sorting plots (A) and mean fluorescence intensity (B) of phosphorylated STAT5 and STAT3 in the CD3+ T cell population (n = 4 cultures, unpaired two-tailed Student’s t test for comparison between untreated and IL6-treated samples; one-way ANOVA with multiple comparison test for comparison among different CAR-T cells. NS, not significant.
(C–E) Anti-CD19 CAR or anti-CD19 CAR+G6/7R T cells were rested overnight in cytokine-free media and left untreated or treated with IL-6 for 30 min, and their gene expression profiles were compared by RNA-seq (n = 3 different donor samples for each). (C and D) Principal component analysis (C) and unsupervised hierarchical clustering (D) of differentially expressed genes among the four groups (false discovery rate [FDR] < 0.01). (E) Gene set enrichment analysis (GSEA) between control and G6/7R-expressing rested CAR-T cells. Genes induced by IL-7 were used as gene sets (n = 3 samples, p values were calculated by an empirical phenotype-based permutation test).
(F) Fold expansion of anti-CD19 CAR alone or anti-CD19 CAR and G6/7R-engineered T cells 1 week after stimulation with NALM-6. CAR-T cells were cultured without cytokine supplementation (n = 4 cultures, unpaired two-tailed Student’s t test). Representative data of three independent experiments.
(G) Fold expansion of anti-CD19 CAR or anti-CD19 CAR and G6/7R-transduced T cells restimulated with NALM-6 once and cultured in the presence of IL-2 without additional antigen stimulation (n = 4 cultures, unpaired two-tailed Student’s t test at each time point, ∗∗p < 0.01).
(H) The proliferation of anti-CD19 CAR and anti-CD19 CAR+G6/7R T cells incubated with or without 20 ng/mL IL-6 for 3 days without antigen stimulation (n = 4 cultures, one-way ANOVA with multiple comparison test).
(I) Cytokine production of anti-CD19 CAR-T cells with or without G6/7R upon restimulation with NALM-6 was analyzed by intracellular flow cytometry (n = 4 different donor samples, paired two-tailed Student’s t test).
(J) Cytotoxic activity of anti-CD19 CAR or anti-CD19 CAR+G6/7R T cells against the indicated target cells as evaluated by flow cytometry (n = 3 cultures, unpaired two-tailed Student’s t test).
In (B), (F), (H), and (J), horizontal lines denote mean values. See also Figure S2.
We then examined the functional properties of G6/7R-engineered anti-CD19 CAR-T cells. The G6/7R CAR-T cells displayed significantly better proliferation than T cells engineered with CAR alone upon antigen stimulation (Figure 2F). The G6/7R anti-CD19 CAR-T cells continued proliferation significantly longer than standard CAR-T cells in the absence of antigen restimulation; however, they eventually lost viability, suggesting that constitutive IL7R signaling does not result in malignant transformation of CAR-T cells (Figure 2G). Although IL-6-mediated STAT3 activation was modest, it promoted T cell survival in G6/7R anti-CD19 CAR-T cells incubated in a cytokine-free medium (Figure 2H). Production of IFN-γ and cytolytic activity of anti-CD19 CAR-T cells upon coculture with target tumor cells were slightly improved by the expression of G6/7R (Figures 2I, 2J, and S2B), which is consistent with RNA-seq results indicating enhanced expression of genes associated with effector functions.
JAK-STAT pathway activation plays a central role in the improved function of G6/7R anti-CD19 CAR-T cells
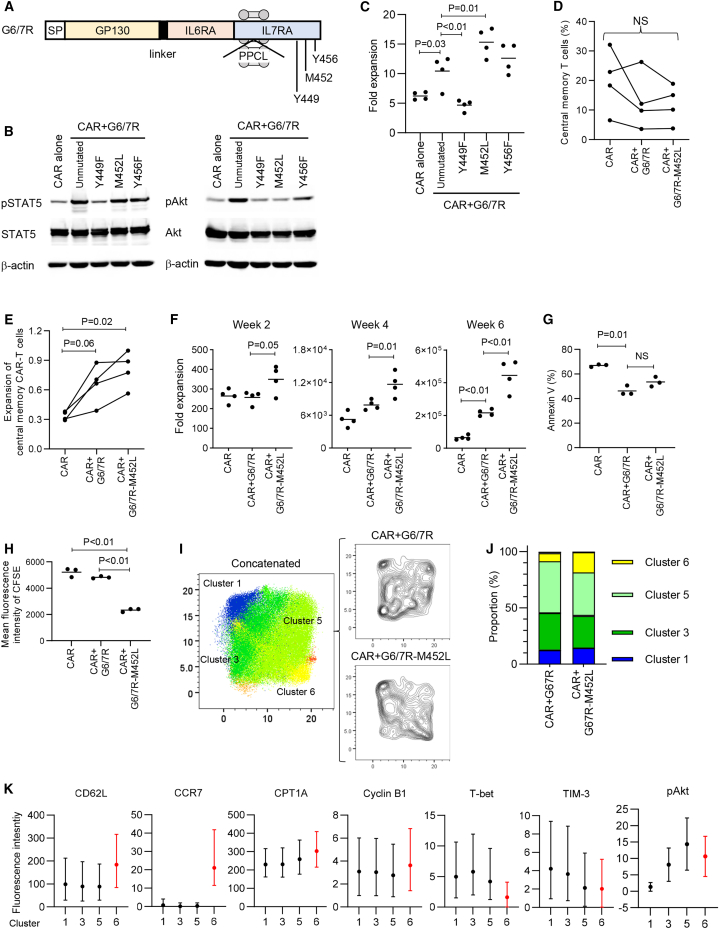
IL-7 signaling induces activation of the Akt pathway in addition to the JAK-STAT pathway, both of which play a vital role in T cell functions.34,35 To investigate which signaling pathway predominantly contributes to the superior CAR-T cell proliferation, we generated G6/7R with point mutations at C-terminal amino acid residues (Figure 3A). The tyrosine Y449 is required for both STAT5 and Akt activation.36 Although the M452 residue is essential for recruiting the p85 subunit of phosphatidylinositol 3-kinase (PI3K), it can be replaced by leucine (YXXL motif) for STAT5 recruitment.37 Consistent with these previous findings, the Y449F mutation abrogated phosphorylation of both STAT5 and Akt, while the M452L mutation selectively suppressed Akt phosphorylation without affecting STAT5 activation in G6/7R-transduced anti-CD19 CAR-T cells (Figure 3B). The Y456F mutation did not affect either STAT5 or Akt signaling.
Figure 3.
Attenuated Akt signaling in G6/7R-M452L supports long-term proliferation of anti-CD19 CAR-T cells
(A) A schematic diagram of the individual mutants in G6/7R.
(B) T cells transduced with anti-CD19 CAR alone or with anti-CD19 CAR and G6/7R with the indicated mutations were rested overnight in cytokine-free media. Immunoblotting evaluated the phosphorylated and total levels of STAT3, STAT5, and Akt. Representative data of two experiments.
(C) Anti-CD19 CAR or anti-CD19 CAR+G6/7R T cells with the indicated mutations were stimulated with NALM-6 and cultured without exogenous cytokines. The data shown are fold expansion of the CAR-T cells on day 7 following stimulation (n = 4 cultures, one-way ANOVA with multiple comparison test). Representative data of three independent experiments.
(D–H) Anti-CD19 CAR-T cells with or without ectopic expression of G6/7R or G6/7R-M452L were restimulated with NALM-6. The data shown are the frequency (D) and fold expansion (E) of CD8+ central memory T cells on day 7 (n = 4 different donor samples, repeated measures one-way ANOVA with multiple comparison test), cumulative fold expansion of CAR-T cells upon weekly stimulation until week 6 (F, n = 4 cultures, one-way ANOVA with multiple comparison test), the frequency of Annexin V+ cells on day 7 (G, n = 3 cultures, one-way ANOVA with multiple-comparison test), and mean fluorescence intensity of CFSE on day 7 (H, n = 3 cultures, one-way ANOVA with multiple-comparison test). In (H), T cells were labeled with CFSE before stimulation. NS, not significant.
(I–K) Mass cytometry analysis was performed in G6/7R and G6/7R-M452L anti-CD19 CAR-T cells 7 days after restimulation. (I) An EmbedSOM-based plot of concatenated samples to visualize the distribution of G6/7R and G6/7R-M452L CD8+ CAR-T cells. Individual cells are colored according to the clusters obtained by FlowSOM. Each sample consists of a pool of three cultures from one donor. (J) Proportions of each cluster in G6/7R and G6/7R-M452L CAR-T cells. Color annotations are provided for the four major populations (clusters 1, 3, 5, and 6). (K) Distribution of fluorescence intensity of the indicated molecules in each cluster. Median values with interquartile range are shown.
In (C) and (F–H), horizontal lines denote mean values. See also Figure S3 and Table S2.
Among the generated mutants, the G6/7R-M452L promoted the proliferation of anti-CD19 CAR-T cells significantly better than the unmutated G6/7R (Figure 3C). On day 7 after restimulation, anti-CD19 CAR-T cells acquired central memory (CD45RA−CCR7+CD62L+CD28+CD27+) and effector memory (CD45RA−CCR7−) phenotypes (Figure S3A). Although both G6/7R and G6/7R-M452L CAR-T cells showed a tendency of accelerated differentiation compared with standard CAR-T cells due to their enhanced proliferation (Figure 3D), G6/7R-M452L CAR-T cells accomplished significantly better expansion of cells with a central memory phenotype than standard CAR-T cells (Figure 3E). The proliferative advantage of G6/7R-M452L CAR-T cells became more prominent upon repeated stimulations (Figure 3F). The enhanced proliferation was due to a higher rate of cell division without significant changes in the frequency of apoptosis after antigen stimulation (Figures 3G and 3H). The G6/7R-M452L-expressing CAR-T cells showed cytokine production and cytolytic activity comparable to G6/7R CAR-T cells (Figures S3B and S3C).
We took a closer look at the molecular profiles of anti-CD19 CAR-T cells with or without G6/7R or G6/7R-M452L. RNA sequence analysis of the antigen-restimulated anti-CD19 CAR-T cells showed that the gene expression profiles of the G6/7R and G6/7R-M452L CAR-T cells were very similar. While conventional CAR-T cells formed an independent cluster by unsupervised clustering, G6/7R and G6/7R-M452L CAR-T cells were grouped in the same cluster, which was subdivided according to the donor (Figure S3D). There were 4,383 differentially expressed genes (p < 0.01) between conventional and G6/7R CAR-T cells, of which 3,428 genes were also differentially expressed in G6/7R-M452L versus conventional CAR-T cells (Figure S3E). Only 187 genes showed differential expression between G6/7R and G6/7R-M452L CAR-T cells. These results suggest that, at least in the bulk sample data, the M452L mutation does not cause significant changes in gene expression.
We then performed mass cytometry analysis to compare the profiles of three types of CD8+ anti-CD19 CAR-T cells at the single-cell level (Table S2). Conventional CAR-T cells again formed a distinct cluster when visualized with the EmbedSOM algorithm based on the profiles of 16 markers (Figure S3F).38 As expected, the area enriched with conventional CAR-T cells was characterized by low levels of phospho-STAT5. To explore differential profiles between G6/7R and G6/7R-M452L CAR-T cells, we next projected these two groups alone with EmbedSOM (Figure 3I). Clustering with flowSOM identified 4 predominant subsets, of which cluster #6 was enriched in G6/7R-M452L CAR-T cells (Figure 3J). Cells within this population were characterized by increased expression of the memory markers CD62L and CCR7, CPT1A (related to fatty acid oxidation), and cyclin B1 (related to cell cycle) and decreased levels of the effector transcription factor T-bet and the exhaustion marker TIM-3 (Figures 3K and S3G). Akt phosphorylation levels were most upregulated in the cluster 5, which was enriched for G6/7R CAR-T cells. These results suggest that less differentiated and exhausted T cells with superior proliferative capacity tended to be maintained in G6/7R-M452L CAR-T cells compared to G6/7R CAR-T cells.
G6/7R and G6/7R-M452L CAR-T cells provide durable antitumor efficacy in multiple tumor models
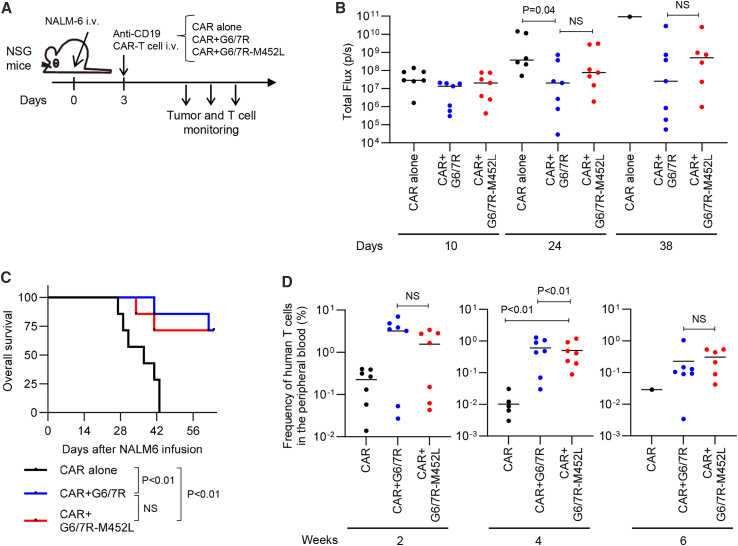
We then analyzed the antitumor efficacy of G6/7R and G6/7R-M452L CAR-T cells in vivo. When infused into NALM-6-bearing NSG mice, both G6/7R and G6/7R-M452L-engineered anti-CD19 CAR-T cells showed a more durable antitumor response compared with T cells transduced with anti-CD19 CAR alone (control), which resulted in superior survival of the treated mice (Figures 4A–4C, S4A, and S4B). Importantly, both G6/7R and G6/7R-M452L CAR-T cells showed better persistence compared with the control CAR-T cells (Figure 4D). There was no significant difference in therapeutic activity between G6/7R and G6/7R-M452L CAR-T cells. These results highlight a therapeutic advantage of the chimeric receptor-driven JAK-STAT activation in CD19-targeting CAR-T cells.
Figure 4.
Superior antitumor effects of G6/7R-M452L anti-CD19 CAR-T cells in a leukemia model
(A) Experimental design showing the transplantation of NALM6-GL-bearing NSG mice with 5 × 105 anti-CD19 conventional CAR-T cells or G6/7R- or G6/7R-M452L-expressing CAR-T cells on day 3.
(B) The total flux of luciferase activity was quantified by in vivo bioluminescent imaging at the indicated time points following NALM6-GL infusion (n = 7 mice for each group, one-way ANOVA with multiple comparison test of the log-transformed values).
(C) The Kaplan-Meier curve for progression-free survival of the treated mice (n = 7 mice, log rank test). The data shown are a composite of three independent experiments.
(D) The frequency of human T cells in the peripheral blood at the indicated time points (n = 7 mice, one-way ANOVA with multiple comparison test of the log-transformed values).
In (B) and (D), horizontal lines denote mean values. See also Figure S4.
Next, we investigated whether G6/7R enhances CAR-T cell efficacy against solid tumors. Similar to anti-CD19 CAR-T cells, expression of G6/7R or G6/7R-M452L in T cells engineered with CAR genes encoding different antibody fragments or costimulatory domains (CD28 or 41BB) significantly augmented their proliferative capacity upon antigen stimulation (Figure 5A). A marginal proliferative advantage of G6/7R-M452L over G6/7R was observed in anti-mesothelin 28z and anti-GD2 BBz CAR-T cells but not in anti-GD2 28z CAR-T cells.
Figure 5.
G6/7R-M452L enhances the antitumor efficacy of anti-mesothelin CAR-T cells against solid tumors
(A) Fold expansion of mesothelin or GD2-targeting CAR-T cells transduced with G6/7R or G6/7R-M452L after restimulation with the target antigen (n = 4 cultures, one-way ANOVA with multiple comparison test).
(B) NOG-ΔMHC mice were subcutaneously inoculated with the pancreatic cancer cell line AsPC-1 and treated with anti-mesothelin CAR-T cells with or without different cytokine receptors. CAR-T cells were also transduced with the luciferase gene.
(C) Tumor volume was monitored longitudinally (n = 6 or 7 mice for each).
(D) Human T cell frequencies were analyzed in peripheral blood at the indicated time points (n = 6 or 7 mice for each, one-way ANOVA with multiple comparison test).
(E) In vivo bioluminescence imaging was used to quantify the total flux of luciferase activity (n = 6 or 7 mice for each, one-way ANOVA with multiple comparison test for the log-transformed values).
(F) Body weight of the treated mice relative to the weight on day 0 (n = 6 or 7 mice for each). ∗p < 0.05, ∗∗p < 0.01.
In (A), (D), and (E), horizontal lines denote mean values. See also Figure S5.
To assess the therapeutic efficacy of G6/7R-M452L CAR-T cells in a solid tumor model, NOG-ΔMHC mice were subcutaneously inoculated with the mesothelin-expressing pancreatic cancer cell line AsPC-1 and then infused with anti-mesothelin CAR-T cells with or without chimeric cytokine receptors (G6/7R, G6/7R-M452L, or CD34/7R; Figure 5B). The CD34/7R is a previously reported cytokine receptor consisting of the extracellular domain of CD34 and the transmembrane and cytoplasmic domain of the constitutively active IL7R mutant to transduce constitutive IL-7 signaling (Table S1).9 We also expressed the luciferase gene in CAR-T cells to track their distribution longitudinally. As shown in Figure 5C, all cytokine receptors similarly enhanced the antitumor efficacy of anti-mesothelin CAR-T cells. However, peripheral blood analysis showed superior expansion of G6/7R-M452L CAR-T cells compared to G6/7R or CD34/7R CAR-T cells (Figure 5D). Consistent with this, in vivo imaging analysis also showed that G6/7R-M452L CAR-T cells expanded and persisted significantly better than the other CAR-T cells (Figures 5E and S5). None of the mice experienced lethal weight loss during the follow-up period (Figure 5F). These results suggest that G6/7R-M452L anti-mesothelin CAR-T cells can expand efficiently and induce durable therapeutic responses in vivo without causing toxicity.
We further evaluated the properties of G6/7R-M452L anti-mesothelin CAR-T cells in a high tumor burden model. NSG mice were inoculated with AsPC-1 and 13 days later infused with mesothelin-targeting conventional or G6/7R-M452L CAR-T cells expressing the luciferase gene (Figure 6A). As seen in the previous experiments, G6/7R-M452L-expressing CAR-T cells suppressed tumor progression more significantly than conventional CAR-T cells (Figures 6B and 6C). No apparent weight loss was observed in treated mice (Figure S6A). The G6/7R-M452L CAR-T cells were detected at a higher level in the peripheral blood than the control CAR-T cells (Figure 6D). While both CAR-T cells were effectively localized within the tumor site, G6/7R-M452L CAR-T cells showed significantly better accumulation at each time point, which likely resulted in optimal control of tumor progression (Figures 6E and S6B). When analyzed at early time points, anti-mesothelin CAR-T cells migrated to the lungs first and then gradually moved to the tumor site within a week (Figures S6C and S6D).
Figure 6.
Functional and phenotypic properties of G6/7R-M452L anti-mesothelin CAR-T cells in a solid tumor model
(A) NSG mice subcutaneously inoculated with AsPC1 were infused with T cells engineered with anti-mesothelin CAR alone or anti-mesothelin CAR and G6/7R-M452L. CAR-T cells were cotransduced with the luciferase gene.
(B) Monitoring of the subcutaneous tumor volume (n = 7 mice for each).
(C) Progression-free survival of the treated mice (n = 7 mice, log rank test).
(D) Frequency of human T cells in the peripheral blood at the indicated time points (n = 7 mice, unpaired two-tailed Student’s t test).
(E) The total flux of the luciferase activity was analyzed by in vivo bioluminescent imaging (n = 7 mice, unpaired two-tailed Student’s t test of the log-transformed values). In (A)–(E), the data are a composite of two independent experiments. ∗p < 0.05, ∗∗p < 0.01.
(F and G) The subcutaneous tumor was harvested 6–9 weeks following anti-mesothelin CAR-T cell infusion in the AsPC-1 tumor model shown in (A). (F) Percentage of CD8+ CAR-T cells expressing exhaustion markers (PD-1, LAG-3, and TIM-3) (n = 6 mice for the CAR alone group and n = 8 mice for the CAR+G6/7R-M452L group). Error bars denote SD. (G) Phospho-STAT3 and STAT5 levels in the CD8+ CAR-T cell population were quantified by flow cytometry (n = 8, unpaired two-tailed Student’s t test).
(H) AsPC-1-bearing NSG mice were infused with G6/7R or G6/7R-M452L anti-mesothelin CAR-T cells. CAR-T cells persisting in the spleen were analyzed for Ki-67 expression (n = 8 mice, unpaired two-tailed Student’s t test).
(I) Anti-mesothelin G6/7R-M452L CAR-T cells persisting after regression of AsPC-1 were collected from spleen and transplanted into tumor-free NSG mice (n = 9 mice).
(J) The frequency of human T cells in the peripheral blood of mice with secondary transplantation was analyzed by flow cytometry (n = 9 mice).
(K) Total flux of the luciferase expressed in CAR-T cells was serially monitored by in vivo imaging. For comparison, luciferase-transduced Jurkat cells were transplanted into NSG mice and monitored using the same protocol (n = 9 mice for the CAR-T cell group, and n = 8 mice for the Jurkat group).
In (D), (E), (G), (H), (J), and (K), horizontal lines denote mean values. See also Figure S6.
We also performed phenotypic analysis of tumor-infiltrating CAR-T cells targeting mesothelin. Both conventional and G6/7R-M452L CAR-T cells highly upregulated immunoinhibitory receptors at the tumor site (Figures 6F and S6E). At the tumor site, G6/7R-M452L CAR-T cells still maintained elevated levels of pSTAT3 and pSTAT5 (Figure 6G). IL-2 and its downstream target STAT5 are essential for regulatory T cell (Treg) functions.39 Although a proportion of CD4+ CAR-T cells had Treg-like phenotypes (CD25+CD127−; Figure S6F), the frequency was not significantly different regardless of G6/7R-M452L expression (Figure S6G). We also compared the proliferative capacity of G6/7R and G6/7R-M452L anti-mesothelin CAR-T cells in vivo. As observed in in vitro assays, G6/7R-M452L CAR-T cells exhibited elevated levels of Ki-67 compared to G6/7R CAR-T cells, confirming the superior proliferation provided by this mutant cytokine receptor (Figure 6H).
Constitutive JAK-STAT activation is frequently detected in T cell malignancies.40 To assess whether G6/7R-M452L CAR-T cells show evidence of malignant transformation, we collected G6/7R-M452L anti-mesothelin CAR-T cells from the spleen after regression of subcutaneous AsPC-1 and secondarily transplanted them into irradiated NSG mice (Figures 6I, S6H, and S6I). Human T cells were barely detectable in the peripheral blood (Figure 6J) or by in vivo imaging analysis (Figure 6K) during the 2-month follow-up period. We also transplanted the T cell lymphoma cell line Jurkat cells transduced with luciferin into NSG mice and confirmed that they rapidly spread throughout the body and induced lethal weight loss (Figure S6J). These results suggest that although G6/7R-M452L CAR-T cells can expand potently in vivo, they do not undergo malignant transformation, at least during the time period monitored.
CAR-T cells expressing G6/7R-M452L and IL1R2 effectively sequester monocyte-derived IL-6 and IL-1β
CAR-T cells produce multiple effector cytokines upon antigen encounter and activate monocyte-lineage cells, which then trigger the development of CRS through the secretion of proinflammatory cytokines. In addition to IL-6, multiple studies have revealed that the upregulated expression of IL-1β is related to a neurological disorder.17,18,41,42 To simultaneously sequester IL-6 and IL-1β by CAR-T cells, we linked G6/7R-M452L, anti-CD19 CAR, and IL1R2, which functions as a high-affinity decoy receptor that prevents IL-1β from binding to the signaling receptor IL1R1 (Figure 7A).43 When transduced in T cells, IL1R2 was hardly expressed on the cell surface but was abundantly detected in the culture supernatant, suggesting that its extracellular domain is secreted through shedding as previously reported (Figures S7A and S7B).44 CAR-T cells expressing G6/7R-M452L and IL1R2 simultaneously reduced IL-6 and IL-1β concentrations within the supernatant (Figures 7B and 7C).
Figure 7.
G6/7R-M452L anti-CD19 CAR-T cells can absorb monocyte-derived IL-6
(A) A schematic diagram of the all-in-one vector expressing G6/7R-M452L, anti-CD19 CAR, and IL1R2.
(B and C) CAR-T cells coexpressing G6/7R-M452L and IL1R2 were incubated for the indicated periods in the presence of recombinant IL-6 and IL-1β. Concentrations of IL-6 and IL-1β in the culture supernatant were quantified by ELISA (n = 3 cultures; ∗∗p < 0.01, unpaired two-tailed Student’s t test).
(D and E) Anti-CD19 CAR-T cells with or without G6/7R and IL1R2 were incubated with NALM-6 and macrophages differentiated from peripheral blood monocytes. Culture supernatant was collected 24 or 48 h after coculture and analyzed for the indicated cytokine concentration by ELISA (E, n = 4 cultures, unpaired two-tailed Student’s t test). Representative data of two experiments.
(F) NOG-EXL mice reconstituted with human cord blood cells were infused with CD19-targeting CAR-T cells with or without the indicated cytokine receptors (n = 7–10 mice for each group). Tocilizumab was administered on day 1 when indicated.
(G and H) The frequency of CAR-T cells in the peripheral blood (G) and plasma IL-6 levels (H) were analyzed on days 1, 3, 7, and 10 after CAR-T cell infusion (n = 7–10 mice per group, one-way ANOVA with multiple comparison test of the log-transformed values). ∗p < 0.05, ∗∗p < 0.01; NS, not significant.
In (E), (G), and (H), horizontal lines denote mean values. See also Figure S7.
We also tested whether G6/7R-M452L and IL1R2 could effectively suppress monocyte-derived cytokines. We stimulated anti-CD19 CAR-T cells expressing G6/7R-M452L and IL1R2 with NALM-6 and then added it to macrophages differentiated from peripheral blood monocytes (Figure 7D). As expected, CAR-T cells expressing G6/7R-M452L and IL1R2 effectively reduced both IL-6 and IL-1β levels in the culture supernatant (Figure 7E).
We then investigated whether the G6/7R-M452L can reduce monocyte-derived IL-6 levels in vivo using a humanized mouse model. NOG-EXL mice expressing human GM-CSF and human IL-3 were transplanted with CD34+ cord blood cells to reconstitute human hematopoiesis (Figure 7F).45 3 weeks after cord blood infusion, reconstitution of human B cells (CD19+) and monocytes (CD14+CD33+) was successfully demonstrated (Figure S7C). The humanized mice were then infused with CD19-targeting CAR-T cells with or without a chimeric cytokine receptor. In this model, CAR-T cells are activated upon recognition of normal B cells and then induce monocytes to secrete inflammatory cytokines including IL-6. We can evaluate the consumption of IL-6 by CAR-T cells if all other conditions (antigen load, CAR-T cell proliferation, other cell sources expressing the IL-6 receptor, etc.) are comparable. We also tested the chimeric receptor CD34/7R-M452L, which activates the JAK-STAT pathway similarly to G6/7R-M452L without IL-6 sequestration. CAR-T cells with CD34/7R-M452L or G6/7R-M452L expanded significantly better than conventional CAR-T cells, consistent with the data shown in multiple tumor models (Figure 7G). Along with superior T cell expansion, CD34/7R-M452L CAR-T cells induced higher levels of IL-6 compared with conventional CAR-T cells (Figure 7H). Despite potent proliferation, G6/7R-M452L CAR-T cells induced less IL-6 than CD34-M452L CAR-T cells. The mice infused with conventional or G6/7R-M452L CAR-T cells had comparable levels of IL-6. These results suggest that the extracellular IL-6 receptors likely contribute to the consumption of IL-6 produced from monocytes in vivo. Most of the mice infused with CAR-T cells showed progressive weight loss irrespective of IL-6 reduction (Figure S7D). Tocilizumab administration did not prevent weight loss, suggesting that the observed weight loss in this model is induced independently of IL-6.
Discussion
In the present study, we developed a chimeric cytokine receptor G6/7R that enhances the therapeutic efficacy of CAR-T cell therapy. While G6/7R CAR-T cells efficiently absorbed extracellular IL-6 in vitro, their infusion did not significantly reduce plasma IL-6 levels compared to control CAR-T cells in a mouse model. Considering that CRS is induced by regionally activated macrophages at the tumor-T cell interaction site,17 it is a reasonable strategy to equip CAR-T cells with mechanisms to eliminate CRS mediators. CAR-T cells efficiently internalized IL-6 through chimeric receptor-mediated endocytosis. Although a specific internalization motif has not been identified in IL7Rα, its intracellular domain seems to be equipped with clathrin-mediated endocytosis of the bound cytokine.46 In real clinical settings, both the number of CAR-T cells and IL-6 levels change dynamically during the antitumor response. In addition, it should be noted that absorption kinetics differ between in vitro and in vivo. Careful evaluation of the relationship between CAR-T cell dose, expansion, and serum IL-6 levels in vivo would be required in future studies. One recent study demonstrated that an anti-IL6 single-chain variable fragment (scFv) secreted by CAR-T cells effectively neutralized serum IL-6 in the treated patients.47 Although the mode of action is different from ours, these results provide encouraging evidence that CAR-T cell-derived “IL-6 inhibitors” can indeed suppress the development of severe CRS.
In addition to the direct inhibition of inflammatory cytokines, repressing mediators associated with macrophage activation is another strategy to prevent the development of CRS. Pharmacologic or genetic inhibition of GM-CSF production by CAR-T cells has been shown to reduce the risk of CRS and neurotoxicity.41 A recent study showed that knockout of IFN-γ in CAR-T cells also suppressed macrophage activation, thereby resulting in attenuating CRS-related cytokine production including IL-6.42
Although we showed that the increase in plasma IL-6 levels was significantly repressed by ectopic expression of G6/7R-M452L in CAR-T cells in the humanized mouse model, there are several other confounding factors that affect plasma IL-6 levels. First, efficient clearance of target B cells by G6/7R-M452L CAR-T cells may also have contributed to the reduced IL-6 levels. Second, human IL-6 can also bind to the mouse IL-6 receptor, the expression of which is dynamically altered by proinflammatory stimuli.48 In addition, we could not detect clinical symptoms specifically related to IL-6. Most of the treated mice lost body weight progressively. This toxicity is unlikely to be related to elevated levels of IL-6 considering that the weight loss was also observed in mice treated with tocilizumab, a well-established treatment for CRS. Similarly, we did not observe neurological symptoms following CAR-T cell infusion. The establishment of a model that truly mimics the development of CRS will be required in future studies. Nevertheless, our data demonstrated that chimeric receptor-engineered CAR-T cells can efficiently sequester both IL-6 and IL-1 derived from monocytes in vitro, which simulates cytokine secretion profiles in the clinical settings.42
The effect of IL-7 signaling in enhancing antitumor efficacy has already been established.9,13,31 In particular, constitutive STAT5 signaling contributes to the enhancement of both effector functions and long-term survival of T cells as memory T cells.49,50,51 Unexpectedly, we found that attenuated Akt signaling by the M452L mutation conferred a slight proliferative advantage to G6/7R CAR-T cells. While Akt signaling is required for potent effector T cell functions, previous studies have shown that its chronic activation negatively affects T cell proliferation due to reduced mitochondrial biogenesis.52,53 These studies also showed that pharmacological inhibition of Akt restored mitochondrial activity, suggesting that chronic Akt signaling-mediated T cell dysfunction may be prevented or reversed by intermittent silencing of the signal. In another study, Weber et al. also showed that transient inhibition of tonic CAR signaling reinvigorated the effector function of exhausted CAR-T cells.54 Tonic signaling is induced by some of the CAR constructs with the CD28 signaling domain and includes activation of Akt signaling. Our mass cytometry data showing a relative increase in the less differentiated and exhausted population in G6/7R-M452L CAR-T cells compared to G6/7R CAR-T cells are consistent with these findings. Another approach to prevent the potential negative effects of chronic Akt signaling would be to develop a receptor that induces IL-7 signaling in an IL-6-dependent manner. However, the in vitro functional differences associated with the M452L mutation did not necessarily translate into a difference in therapeutic efficacy in vivo. Both G6/7R and G6/7R-M452L CAR-T cells showed similarly better antitumor function than control CAR-T cells in multiple tumor models. These results suggest that the effect of Akt signaling activity, if present, is marginal to the overall therapeutic activity of G6/7R CAR-T cells.
In addition to constitutive IL-7 signaling, engagement of IL-6 in G6/7R augmented phosphorylation of STAT3, which is likely induced by the recruitment of endogenous GP130 to the IL6RA-IL6 complex. STAT3 plays an essential role in memory T cell formation, and its activation contributes to the increased proliferation of CAR-T cells.6,10,55,56 Although we confirmed that IL-6 treatment enhances G6/7R-expressing CAR-T cell proliferation in vitro, it would also be interesting to evaluate how exposure to IL-6 affects CAR-T cell properties in vivo.
One of the concerns related to constitutive IL-7 signaling is the malignant transformation of CAR-T cells. Lymphoma development was recently reported in piggyBac transposon-mediated CAR-T cells, which has never been observed in CAR-T cells generated by a virus vector.57 Although we did not observe aberrant expansion of G6/7R-M452L CAR-T cells after tumor eradication, careful evaluation of long-term T cell survival would be required in future clinical trials. Incorporating a suicide gene that can eliminate CAR-T cells is one approach to ensure safety. Another strategy is to construct a receptor that induces cytokine signaling in a ligand-dependent manner. For example, Lange et al. recently reported a chimeric GM-CSF receptor with the cytoplasmic domains of IL-18R, in which CAR-T cell-derived GM-CSF is converted to IL-18 signaling and augments their effector functions.58
In summary, our chimeric receptor can improve efficacy of CAR-T cell therapy and potentially contribute to the suppression of CRS. This technology can be applicable to a broad range of gene-engineered T cell therapy, irrespective of the target antigen, including CAR-T cells and TCR-T cells.
Limitations of the study
Although we found superior proliferation and in vivo persistence of G6/7R-M452L CAR-T cells compared to G6/7R CAR-T cells, gene expression and mass cytometry analysis did not provide sufficient mechanistic insight to explain these functional differences. Another limitation is that the present study did not establish an in vivo mouse model that truly recapitulates the development of CRS. In addition, we did not demonstrate efficient sequestration of plasma IL-6 by G6/7R CAR-T cells compared to control CAR-T cells in vivo.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| APC anti-human CD8 | BioLegend | Cat#301049; RRID: AB_2562054 |
| FITC anti-human CD8 | BioLegend | Cat#301050; RRID: AB_2562055 |
| PerCP/Cyanine5.5 anti-human CD8 | BioLegend | Cat#344710; RRID: AB_2044010 |
| Brilliant Violet 421 anti-human CD8 | BioLegend | Cat#301036; RRID: AB_10960142 |
| PE anti-human CD4 | BioLegend | Cat#300539; RRID: AB_2562053 |
| Alexa Fluor 647 anti-human CD45 | BioLegend | Cat#304018; RRID: AB_389336 |
| FITC anti-human CD45 | BioLegend | Cat#304038; RRID: AB_2562050 |
| PerCP/Cyanine5.5 anti-human CD271 | BioLegend | Cat#345112; RRID: AB_11204075 |
| PerCP/Cyanine5.5 anti-human CD126 (IL-6Rα) | BioLegend | Cat#352812; RRID: AB_2562719 |
| PE anti-human CD126 (IL-6Rα) | BioLegend | Cat#352804; RRID: AB_10900066 |
| APC anti-human CD130 (gp130) | BioLegend | Cat#362006; RRID: AB_2563404 |
| FITC anti-human CD45RA | BD Pharmingen | Cat#555488; RRID: AB_395879 |
| Pacific Blue anti-human CD197 (CCR7) | BioLegend | Cat#353210; RRID: AB_10918984 |
| APC/Cyanine7 anti-human CD62L | BioLegend | Cat#304814; RRID: AB_493582 |
| Brilliant Violet 510 anti-human CD28 | BioLegend | Cat#302936; RRID: AB_2562030 |
| PE/Cyanine7 anti-human CD27 | BioLegend | Cat#356412; RRID: AB_2562258 |
| FITC anti-human IL-2 | BioLegend | Cat#500304; RRID: AB_315091 |
| PE/Cyanine7 anti-human IFN-γ | BioLegend | Cat#502528; RRID: AB_2123323 |
| Brilliant Violet 421 anti-human TNF-α | BioLegend | Cat#502932; RRID: AB_10960738 |
| FITC anti-human CD3 | BioLegend | Cat#300440; RRID: AB_2562046 |
| Pacific Blue anti-STAT3 (pY705) | BD Phosflow | Cat#560312; RRID: AB_1645327 |
| Alexa Fluor 647 anti-STAT3 (pTyr705) | BioLegend | Cat#651008; RRID: AB_1645327 |
| Alexa Fluor 647 anti-STAT5 (pY694) | BD Phosflow | Cat#612599; RRID: AB_2572086 |
| APC anti-human IL-1 RII | R&D Systems | Cat#FAB663A; RRID: AB_10569839 |
| Brilliant Violet 421 anti-human CD127 | BioLegend | Cat#351310; RRID: AB_10960140 |
| PE-Cy7 anti-human CD25 | BioLegend | Cat#302612; RRID:AB_314282 |
| PE-Cy7 anti-human CD279 (PD-1) | BD Pharmingen | Cat#561272; RRID: AB_10611585 |
| PE anti-human CD223 (LAG-3) | BioLegend | Cat#369306; RRID: AB_2629592 |
| Brilliant Violet 421 anti-human CD366 (Tim-3) | BioLegend | Cat#345008; RRID: AB_11218598 |
| Pacific Blue anti-human CD14 | Biolegend | Cat#325616; RRID: AB_830689 |
| PE anti-human CD19 | BioLegend | Cat#302208; RRID: AB_314238 |
| PE-Cy7 anti-human CD33 | Biolegend | Cat#366618; RRID: AB_2566420 |
| PE Anti-Ki-67 | BD Pharmingen | Cat#567719; RRID: AB_2916708 |
| Anti-STAT5 rabbit mAb | Cell Signaling Technology | Cat#94205; RRID: AB_2737403 |
| Anti-Phospho-Stat5 (Tyr694) | Cell Signaling Technology | Cat#4322; RRID: AB_10544692 |
| Anti-Akt | Cell Signaling Technology | Cat#9272; RRID: AB_329827 |
| Anti-Phospho-Akt (Ser473) | Cell Signaling Technology | Cat#4060; RRID: AB_2315049 |
| Anti-Mouse IgG (H + L), HRP Conjugated | Promega | Cat#W4021; RRID: AB_430834 |
| Anti-Rabbit IgG (H + L), HRP Conjugated | Promega | Cat#W4011; RRID: AB_430833 |
| Anti-β-Actin | Santa Cruz Biotechnology | Cat#sc-47778; RRID:AB_626632 |
| Anti-human CD3 (OKT3) | Tonbo Biosciences | Cat#40-0037-U500; RRID: AB_2621438 |
| Anti-Human CD8a-112Cd | Standard BioTools | Cat#92J035112 |
| Anti-Human pStat5 [Y694]-147Sm | Standard BioTools | Cat#3147012C; RRID: AB_2661819 |
| Anti-Human CD271-149Sm | Standard BioTools | Cat#3149017C |
| Anti-Human CD223/LAG-3 -150ND | Standard BioTools | Cat#3150030C |
| Purified anti-human CD28 (Maxpar® Ready) Antibody | Biolegend | Cat#302937; RRID: AB_2563737 |
| Anti-pAkt [S473]-152Sm | Standard BioTools | Cat#3152005C; RRID: AB_2811246 |
| Anti-Human CD62L-153Eu | Standard BioTools | Cat#3153004C; RRID: AB_2810245 |
| Anti-Glucose Transporter GLUT1 | Abcam | Cat#ab196357; RRID: AB_2832207 |
| Anti-CPT1A | Abcam | Cat#ab128568; RRID: AB_11141632 |
| Anti-Human CD197/CCR7-159Tb | Standard BioTools | Cat#3159003C; RRID: AB_2938859 |
| Anti-Human CD130-160Gd | Standard BioTools | Cat#3160017C |
| Anti-Human T-bet/TBX21-161Dy | Standard BioTools | Cat#3161033C |
| Anti-CyclinB1-164Dy | Standard BioTools | Cat#3164010C; RRID: AB_3086684 |
| Anti-pCREB [S133]-165Ho | Standard BioTools | Cat#3165009C; RRID: AB_2661832 |
| Anti-pRb [S807/S811]-166Er | Standard BioTools | Cat#3166011C; RRID: AB_2924785 |
| Anti-Human Ki-67-168Er | Standard BioTools | Cat#3168001C; RRID: AB_2810856 |
| Anti-Human CD366/TIM-3-169Tm | Standard BioTools | Cat#3169028C; RRID: AB_2905650 |
| Anti-pS6 [S235/S236]-172Yb | Standard BioTools | Cat#3172008C; RRID: AB_2938622 |
| Anti-pHistone H3 [S28]-175Lu | Standard BioTools | Cat#3175012C; RRID: AB_2847869 |
| Bacterial and virus strains | ||
| E.coli DH5α competent cells | Takara Bio | Cat#9057 |
| Biological samples | ||
| Human peripheral blood mononuclear cells (PBMCs) | Cellular Technology Limited | CTL-UP1 |
| Human cord blood mononuclear cells | STEMCELL Technologies | ST-70007 |
| Chemicals, peptides, and recombinant proteins | ||
| Recombinant human IL-2 | Novartis | Cat#87-890 |
| Recombinant human IL-15 | PeproTech | Cat#200-15 |
| Recombinant human IL-6 | PeproTech | Cat#200-06 |
| Recombinant human IL-1β | PeproTech | Cat#200-01B |
| Recombinant human GM-CSF | PeproTech | Cat#300-23 |
| Recombinant human M-CSF | PeproTech | Cat#300-25 |
| Brefeldin A Solution (1,000X) | BioLegend | Cat#420601 |
| CD14 MicroBeads, human | Miltenyi Biotec | 130-050-201 |
| CD34 MicroBeads, human | Miltenyi Biotec | 130-046-702 |
| Tocilizumab (anti-IL-6R) | Selleck | A2012 |
| Tumor Dissociation kit, human | Miltenyi Biotec | 130-095-929 |
| RetroNectin | Takara Bio | T100B |
| RIPA Buffer | Nacalai Tesque | 08714–04 |
| Matrigel Matrix | Corning | 354234 |
| D-Luciferin Potassium Salt | FUJIFILM Wako | 126–05116 |
| Cell-ID Cisplatin-198Pt | FLUIDIGM | 201198 |
| Human TruStain FcX | Biolegend | 422302 |
| Cell-ID Intercalator-Ir | FLUIDIGM | 201192A |
| Maxper Cell Acquisition Solution | FLUIDIGM | 201240 |
| Critical commercial assays | ||
| Human IL-1 beta/IL-1F2 DuoSet ELISA | R&D Systems | DY201 |
| ELISA MAX Deluxe Set Human IL-6 | BioLegend | 430504 |
| Human IL-1 RII Quantikine ELISA Kit | R&D Systems | DR1B00 |
| Human IL-1β Enhanced Sensitivity Flex Set | BD Biosciences | 561509 |
| Human IL-6 Enhanced Sensitivity Flex Set | BD Biosciences | 561512 |
| CellTrace CFSE Cell Proliferation Kit | Thermo Fisher Scientific | C34554 |
| LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit | Thermo Fisher Scientific | L34976 |
| Cyto-Fast Fix/Perm Buffer Set | BioLegend | 426803 |
| PE-Cy7 Annexin V | BioLegend | 640951 |
| Maxpar X8 Antibody labeling kit, 151Eu- 4Rxn | Standard BioTools | 201151A |
| Maxpar® X8 Antibody Labeling Kit, 155Gd—4 Rxn | Standard BioTools | 201155A |
| Maxpar® X8 Antibody Labeling Kit, 158Gd—4 Rxn | Standard BioTools | 201158A |
| Deposited data | ||
| RNA-seq of CAR-T cells (Figure 2) | This paper | GSE216734 |
| Mass cytometry analysis of CAR-T cells | This paper | FlowRepository (http://flowrepository.org): FR-FCM-Z77Y |
| RNA-seq of CAR-T cells (Figure S3) | This paper | GSE255145 |
| Experimental models: Cell lines | ||
| K562 | Japanese Collection of Research Bioresources (JCRB) | JCRB0019 |
| Raji | Japanese Collection of Research Bioresources (JCRB) | JCRB9012 |
| NALM-6 | Cell Resource Center for Biomedical Research | TKG 0413 |
| PG13 | American Type Culture Collection (ATCC) | CRL-3597 |
| AsPC-1 | American Type Culture Collection (ATCC) | CRL-1682 |
| Jurkat, clone E6-1 | American Type Culture Collection (ATCC) | TIB-152 |
| Plat-A | Toshio Kitamura | N/A |
| Plat-E | Toshio Kitamura | N/A |
| Lenti-X 293T | Takara Bio | Cat#632180 |
| K562-OKT3/CD80 | Yoshikawa et al.59 | N/A |
| K562-CD19 | Yoshikawa et al.59 | N/A |
| K562-mesothelin | Yoshikawa et al.59 | N/A |
| NALM6-GL | Yoshikawa et al.59 | N/A |
| NALM6-GD2 | Yoshikawa et al.59 | N/A |
| Experimental models: Organisms/strains | ||
| NOD-SCID IL2rgnull (NSG) | Jackson Laboratory | Strain #:005557 |
| NOG-Iab KO (NOG-ΔMHC) | In-Vivo Science | N/A |
| NOG-hGM-CSF/hIL-3 Tg (NOG-EXL) | In-Vivo Science | N/A |
| Recombinant DNA | ||
| pMX | Toshio Kitamura | N/A |
| Anti-CD19 scFv (clone FMC63) | Nicholson et al.60 | N/A |
| Anti-mesothelin scFv (clone ss1) | Li et al.61 | N/A |
| Anti-GD2 scFv (clone 14g2a) with E101K mutation | Rossig et al.62 Horwacik et al.63 |
N/A |
| 28z CAR | Yoshikawa et al.59 | N/A |
| BBz CAR | Yoshikawa et al.59 | N/A |
| pLVpro-MND | Takara Bio | N/A |
| LVpro Packaging Mix | Takara Bio | 6195 |
| 6/7R | Table S1 | N/A |
| G6/7R | Table S1 | N/A |
| G6/7R-M452L | Table S1 | N/A |
| IL1R2 | This manuscript | N/A |
| CD34/7R | This manuscript; originally designed by Shum et al.9 | N/A |
| CD34/7R-M452L | Table S1 | N/A |
| EGFP-P2A-luciferase | Yoshikawa et al.59 | N/A |
| Software and algorithms | ||
| Living Image v4.7.4 | PerkinElmer | https://www.perkinelmer.com/lab-products-and-services/resources/in-vivo-imaging-software-downloads.html |
| Flowjov10.9.0 | BD Biosciences | https://www.flowjo.com/ |
| GraphPad Prism v10.0.0 | GraphPad software | https://www.graphpad.com/ |
| Kuant v2.0 | Vilber Bio Imaging | https://www.vilber.com/ |
| GSEA v4.0.3 | Broad Institute | https://www.gsea-msigdb.org/gsea/index.jsp |
| R v4.0.5 | R project | https://www.r-project.org/ |
| fastp v0.23.2 | Chen et al.64 | https://github.com/OpenGene/fastp |
| Hisat2 v2.1.0 | Kim et al.65 | https://daehwankimlab.github.io/hisat2/ |
| HTseq v0.11.2 | Anders et al.66 | https://github.com/htseq/htseq |
| edgeR v3.28.1 | Robinson et al.67 | https://bioconductor.org/packages/release/bioc/html/edgeR.html |
| Subread | Liao et al.68 | https://subread.sourceforge.net/ |
| EmbedSOM | Kratochvíl et al.38 | https://www.flowjo.com/exchange/#/plugin/profile?id=19 |
| flowSOM | Van Gassen et al.69 | https://www.flowjo.com/exchange/#/plugin/profile?id=7 |
Resource availability
Lead contact
Requests for further information and reagents should be directed to and will be fulfilled by the lead contact, Yuki Kagoya (ykagoya@keio.jp).
Materials availability
Plasmids and cell lines generated in this study will be made available for academic research purpose upon reasonable request.
Data and code availability
The RNA-seq datasets presented in Figures 2 and S3 have been deposited in the NCBI Gene Expression Omnibus under the accession number GSE216734 and GSE255145, respectively. Mass cytometry data were deposited in FlowRepository (FR-FCM-Z77Y). No original code is included in this paper. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Experimental model and subject details
Cell lines
The erythroleukemia cell line K562 and B-cell lymphoma cell line Raji were obtained from the Japanese Collection of Research Bioresources (JCRB) cell bank (Osaka, Japan). The CD19+ B-cell leukemia cell line NALM-6 was obtained from the Cell Resource Center for Biomedical Research, Tohoku University (Sendai, Japan). The PG13 retroviral packaging cell line, pancreatic cancer cell line AsPC-1, and T cell lymphoma cell line Jurkat were purchased from the American Type Culture Collection (ATCC) (Manassas, VA). The Plat-A and Plat-E packaging cell lines were kindly provided by Dr. T. Kitamura. The K562-OKT3/CD80, which expresses single-chain variable fragment (scFv) from the anti-CD3 mAb OKT3 and CD80 on the cell surface has been previously described.59 NALM6-GL was created by retroviral transduction of NALM-6 with the EGFP-P2A-luciferase gene. GD2-expressing NALM6-GL (NALM6-GD2) was generated by transducing NALM6-GL with GM2/GD2 synthase and GD3 synthase.59 K562, NALM-6, Raji, AsPC1, and their derivatives were cultured in RPMI-1640 (Nacalai Tesque, Kyoto, Japan) with 10% FBS (Nichirei Biosciences, Tokyo, Japan). Plat-A, Plat-E, and PG13 cells were cultured in DMEM (Nacalai Tesque) with 10% FBS.
Primary human cells
Healthy donor-derived peripheral blood mononuclear cells (PBMCs) were purchased from Cellular Technology Limited (Cleveland, OH). Human cord blood cells were provided by the RIKEN BioResource Research Center (Ibaraki, Japan) or purchased from STEMCELL Technologies (Vancouver, Canada). This study was performed in accordance with the tenets of the Declaration of Helsinki and approved by the Research Ethics Board of the Aichi Cancer Center, Nagoya, Japan (approval number: 2021-0-183 and 2020-2-14) and the Research Ethics Board of Keio University School of Medicine (approval number: 20231098).
Mice
For the mouse experiments, 4- to 10-week-old male NSG mice (The Jackson Laboratory, Bar Harbor, ME) and 6- to 8-week-old male NOG-ΔMHC and NOG-EXL mice (In-Vivo Science Inc., Tokyo, Japan) were used. All animal experiments were approved by the Animal Care and Use Committee of the Aichi Cancer Center Research Institute, Nagoya, Japan (approval number: R2-8(3)-A) and the Animal Care and Use Committee of Keio University (approval number: A2023-013).
Method details
Construction of CAR and chimeric cytokine receptor genes
Amino acid sequences of the chimeric receptors generated in this study are provided in Table S1. We used scFv derived from the clones FMC63,60 ss1,61 and 14g2a with E101K mutation62,63 for the CAR genes targeting CD19, mesothelin, and GD2, respectively. The FMC63, ss1, and 14g2a scFv sequences were followed by the CD28 and CD3z signaling domains (28z). We also used the CD8α hinge/transmembrane domain and the 4-1BB and CD3z signaling domains for the 14g2a CAR (BBz).10 A truncated form of the NGFR gene (tNGFR) was linked to the CAR genes using a Furin-SGSG-P2A sequence (tNGFR-P2A-CAR) to discriminate CAR-transduced T cells using flow cytometry as previously described.59 Chimeric receptor genes were also connected to a CAR gene via a Furin-SGSG-P2A sequence or expressed from a separate plasmid. All the cDNA genes were inserted into the pMX retroviral plasmid kindly provided by Dr. T. Kitamura (Foundation for Biomedical Research and Innovation at Kobe, Kobe, Japan). For the experiments using humanized mice to evaluate IL-6 secretion, we designed lentivirus plasmids using the pLVpro lentivirus vector as a backbone (Takara Bio, Shiga, Japan). The chimeric cytokine receptor, CD19-targeting 28z CAR, and IL1R2 sequences were linked by a P2A sequence. The MND promoter was used for transgene expression.
Retroviral transduction and T cell culture
Healthy donor-derived PBMCs were stimulated with K562-OKT3/CD80 at an effector to target (E:T) ratio of 7:1 and cultured in the presence of recombinant IL-2 (100 IU/mL, Nipro, Osaka, Japan) and IL-15 (10 ng/mL, PeproTech, Waltham, MA) unless otherwise indicated. Retroviral transduction in T cells was performed on day 2 after the initial stimulation using RetroNectin (Takara Bio). We prepared stable PG13 packaging cell line by treating them with Plat-E-derived retroviruses. These PG13-derived retroviruses were used for T cell infection. The CD19-targeting CAR-T cells were restimulated by coculturing them with NALM-6 at an E:T ratio of 1:1. Anti-mesothelin and anti-GD2 CAR-T cells were restimulated by K562-mesothelin at 5:1 and NALM6-GD2 at 1:1, respectively. For serial stimulation, 2.5×105 CAR-T cells were stimulated with NALM-6 and expanded in the presence of 100 IU/mL IL-2. The CAR-T cells were harvested one week later and restimulated at 2.5×105 cells in fresh medium with 100 IU/mL of IL-2.
Lentiviral transduction and T cell culture
For lentiviral transduction, PBMCs were stimulated on a plate coated with anti-CD3 mAb (clone OKT3, Tonbo Biosciences, San Diego, CA) and RetroNectin (Takara Bio) at 5 μg/ml and 25 μg/mL, respectively. The stimulated cells were added to a lentiviral supernatant and centrifuged at 1,000g for half an hour. Lentivirus was produced by transient transfection of Lenti-X 293T cells (Takara Bio) with a pLVpro lentivirus plasmid and LVpro Packaging Mix (Takara Bio).
Flow cytometry analysis
Flow cytometry analysis was performed with BD LSRFortessa (BD Biosciences, Franklin Lakes, NJ) or CytoFLEX S Flow Cytometer (Beckman, Brea, CA). The antibodies used for analysis are listed in the key resources table. Cytokine production by CAR-T cells was analyzed by coculturing CAR-T cells with target cells for 6 h. After the incubation, Brefeldin A (BioLegend, San Diego, CA) was added to the cultures for 2 h. The cells were fixed and permeabilized using a Cyto-Fast Fix/Perm kit (BioLegend) and subjected to intracellular staining. For intracellular flow cytometry analysis of phosphorylated STAT proteins, T cells were fixed in 4% paraformaldehyde and then permeabilized with ice-cold methanol. Ki-67 staining was performed after T cells were fixed and permeabilized with a Cyto-Fast Fix/Perm kit (BioLegend). All the data analysis was performed using FlowJo software (version 10.8.1; BD Biosciences). To estimate the cytotoxic activity of CAR-T cells, 1×105 CAR-T cells were cultured with the EGFP+ or carboxyfluorescein succinimidyl ester (CFSE)-labeled target tumor cells. We recorded the absolute cell counts of viable tumor cells by flow cytometry. Dead cells were excluded using the LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit (Thermo Fisher Scientific). The frequency of live tumor cells was calculated as the ratio of the cell counts that were incubated without CAR-T cell coculture.
Quantification of cytokine concentration
Cytokine concentrations in the culture supernatants were quantified by ELISA according to the manufacturer’s instructions. The kits used for analysis are summarized in the key resources table. To measure intracellular IL-6 concentrations by ELISA, cells were lysed with RIPA buffer. Extracted lysates and standard IL-6 were denatured at 95°C for 5 min. Recombinant cytokines (human IL-6 and IL-1β) were purchased from Peprotech. To obtain M0 macrophage, we isolated CD14+ monocytes from PBMC using anti-CD14 MicroBeads (Miltenyi Biotec, Bergisch Gladbach, Germany) and cultured them for 6 days in the presence of recombinant M-CSF (10 ng/mL, Peprotech). The obtained macrophage was then mixed with CAR-T cells and NALM-6 and cultured them for 24–48 h to obtain the supernatants. Plasma IL-6 levels were quantified by flow cytometry using the BD CBA Human IL-6 Flex Set (BD Biosciences) according to the manufacturer’s instructions.
Immunoblotting analysis
To analyze phosphoprotein levels, CAR-T cells were incubated in the absence of cytokines. Total cell lysates were collected on the next day. T cells were treated with the Akt inhibitor Akti-1/2 (Selleck, Houston, TX) overnight when indicated. Defined amounts of protein were separated on 8% gels by SDS–PAGE. The gels were transferred to Transblot Turbo PVDF membranes (Bio-Rad, Hercules, CA). The membranes were stained with the primary antibodies at 4°C overnight, and then with horseradish peroxidase (HRP)-conjugated goat anti-rabbit-IgG (H + L) or anti-mouse-IgG (H + L) secondary antibodies (Promega, Madison, WI). The following antibodies were used for analysis: anti-STAT5 (clone D2O6Y; Cell Signaling Technology; 1:1,000 dilution), anti-phospho-STAT5 (Tyr 694) (clone D47E7; Cell Signaling Technology; 1:1,000 dilution), anti-Akt (Cell Signaling Technology; 1:1,000 dilution), anti-phospho-Akt (Ser473) (clone D9E; Cell Signaling Technology; 1:1,000 dilution), anti-β-actin (clone C4; Santa Cruz Biotechnology; 1:1000 dilution), HRP-conjugated anti-mouse-IgG(H + L) (Promega; 1:1,000 dilution), and HRP-conjugated anti-rabbit-IgG(H + L) (Promega; 1:1,000 dilution). All images were acquired with FUSION SOLO.7S.EDGE (Vilber Lourmat, Collégien, France) or ImageQuant LAS 4000 (GE Healthcare, Chicago, IL).
In vivo experiments
To analyze the absorption of recombinant IL-6 in vivo, NSG mice were intravenously transplanted with 2.5×106 NALM-6 cells per mouse and then treated with 3×106 CAR-T cells per mouse. The mice were infused with 2 mg of recombinant IL-6 on day 7 after CAR-T cell infusion. Plasma was collected at 1, 2, and 4 h after IL-6 infusion to measure the IL-6 concentration. In the NALM-6 leukemia model, mice were intravenously infused with 2.5×106 NALM6-GL cells and then treated with 0.5×106 CAR-T cells on day 3 following infusion. Tumor burden was monitored using IVIS Lumina II (PerkinElmer, Waltham, MA) or Newton 7.0 (Vilber, Collégien, France).
For the AsPC-1 solid tumor model comparing the efficacy of CAR-T cells with different cytokine receptors, NOG-ΔMHC mice were inoculated subcutaneously on the abdomen with 1×106 AsPC-1 cells. Mesothelin-targeting CAR-T cells (2 × 106 T cells) were intravenously infused 4 days later. In the high tumor-burden model, NSG mice were inoculated subcutaneously on the abdomen with 1.5×106 AsPC1 cells in Matrigel (Corning Inc, Corning, NY). We infused 0.5×106 mesothelin-targeting CAR-T cells on day 13. Progression-free survival was defined as the time when the tumor volume exceeds 200 mm3 at two consecutive time points or 220 mm3. For tumor-infiltrating CAR-T cell analysis, subcutaneous tumors were harvested and dissociated into single cell suspensions using the Tumor Dissociation Kit (Miltenyi Biotec) and gentleMACS Dissociator (Miltenyi Biotec).
To evaluate the malignant transformation of infused CAR-T cells, splenocytes were harvested from the AsPC-1-inoculated mice after tumor shrinkage (days 40–47), and the frequency of human T cells were analyzed by flow cytometry. NSG mice were irradiated with 1 Gy and infused with the splenocytes containing 1×106 human T cells. For comparison, NSG mice were infused with luciferase-transduced Jurkat cells using the same protocol. In all the solid tumor models, T cells were co-transduced with the luciferase gene to analyze their localization by in vivo imaging.
To determine IL-6 secretion from monocytic cells in vivo, NOG-EXL mice were transplanted with 5×104 CD34+ hematopoietic stem/progenitor cells (HSPCs) derived from umbilical cord blood samples after irradiation. CD34+ HSPCs were isolated using the CD34 MicroBeads Kit (Miltenyi Biotec). After confirmation of the reconstitution of human hematopoiesis, mice were infused with the indicated number of CAR-T cells on day 21 and serially analyzed for body weight, plasma IL-6 levels, and human T cell expansion. When indicated, mice were intravenously infused with tocilizumab (10 mg/kg, Selleck) the day after CAR-T cell infusion.
In all the experiments, mice were monitored for their overall health status and body weight at least twice a week and euthanized upon exhibiting one of the following symptoms: >20% loss of initial body weight, pronounced lethargy, severe diarrhea, hunched posture, and severe dermatitis.
RNA-sequencing
Control or G6/7R-expressing CAR-T cells derived from three different donors were incubated overnight in cytokine-free media. On the next day, CD8+ CAR-T cells were isolated by FACSAria III (BD Biosciences) for RNA purification before or after treatment with IL-6 (50 ng/mL). The purified total RNA was subjected to RNA-sequencing analysis (RIKEN Genesis, Tokyo, Japan). The FASTQ files were quality-trimmed using the Fastp tool (version 0.20.0).64 The trimmed reads were mapped to the GRCh38 human reference genome with Hisat2 (version 2.1.0)65 and read counts for individual genes were quantified by HTseq (version 0.11.2).66 Differentially expressed genes between groups were identified using edgeR (version 3.28.1).67 Unsupervised hierarchical clustering and principal component analysis of differentially expressed genes between the four groups (FDR <0.01) were performed using the heatmap.2 function and the ggplot function, respectively. We used log-transformed counts-per-million reads as gene expression levels. All the above analysis was carried out using the R software environment (version 4.0.5). Gene set enrichment analysis (GSEA) was performed with the GSEA v2 software (Broad Institute) using the genes upregulated upon IL-7 treatment, which were extracted from the publicly available data: GSE58262 (p < 0.05 and fold-change >2) and GSE180020 (p < 0.05 and the log fold-change >2).
To compare gene expression profiles among control, G6/7R, and G6/7R-M452L CAR-T cells, CAR-T cells derived from three different donor PBMCs were individually restimulated on day 7 with NALM-6 at an E:T ratio of 1:1. CD8+CAR+ (for the control group) or CD8+CAR+GP130+ (for the G6/7R and G6/7R-M452L groups) T cell populations were isolated for RNA collection using an MA900 cell sorter (Sony, Tokoy, Japan). RNA sequencing was performed by Azenta Japan (Tokyo, Japan). After quality trimming and mapping to the human genome as described above, read counts for individual genes were calculated using featureCounts of the Subread package (version 2.0.6).68 Differentially expressed genes between the three groups were identified using edgeR (version 3.40.2). Unsupervised hierarchical clustering and principal component analysis was performed as described above. A Venn diagram was generated using the VennDiagram package (version 1.7.3).
Mass cytometry analysis
For mass cytometry analysis, CAR-T cells were stimulated with NALM-6 and expanded in cytokine-free medium for 7 days. The collected cells were stained with Cell-ID Cisplatin-198Pt (Standard Biotools, South San Francisco, CA) to distinguish dead cells. After Fc receptor blocking with Human TruStain FcX (BioLegend), cells were labeled with antibodies for cell-surface markers, fixed with 4% paraformaldehyde, and then permeabilized with ice-cold methanol. The permeabilized cells were further labeled with antibodies for the intracellular targets, fixed with 1.6% formaldehyde, and stained with Cell-ID Intercalator-Ir. All antibodies used are listed in Table S2. Antibodies to CD28, CPT1A and GLUT1 were conjugated to metal isotopes using the Maxpar X8 Antibody Labeling Kit (Standard BioTools) according to the manufacturer’s protocol.
Data were collected on a Helios mass cytometer (Standard BioTools) by St. Luke’s SRL Advanced Clinical Research Center, Inc (Tokyo, Japan). For clustering analysis, equal numbers of CD8+CAR+ conventional CAR-T cells, CD8+CAR+GP130+ G6/7R and G6/7R-M452L CAR-T cell populations were concatenated and analyzed using the EmbedSOM algorithm (v2.2.0).38 Eight metaclusters were formed on concatenated samples of G6/7R and G6/7R-M452L CAR-T cells using flowSOM (v4.0.0) and overlaid on the EmbedSOM map.69 For both EmbedSOM and flowSOM analyses, we used the markers listed in Table S2 (16 in total, excluding CD8, NGFR, and CD130). All analysis was performed using Flowjo software (BD Biosciences).
Quantification and statistical analysis
Statistical analysis
Statistical significance between the two groups was estimated using a two-tailed paired or unpaired Student’s t test. Comparisons among three or more groups were performed using repeated measures or ordinary one-way ANOVA with multiple-comparison test. When indicated, the log-transformed values were used for comparison. The overall survival of the mice treated with CAR-T cells was depicted by the Kaplan–Meier curve. The survival differences were compared with the log rank test. Differences were considered statistically significant at a p-value <0.05. All statistical analyses were performed using GraphPad Prism 10 software. No statistical method was used to predetermine the sample size.
Acknowledgments
This work was supported by AMED under grant numbers JP22bm0704066 (Y.K.), JP22ym0126072 (Y.K.), JP22ama221303 (Y.K.), and JP23zf0127009 (Y.K.); JST FOREST Program (grant number JPMJFR2060, Japan, Y.K.); JSPS KAKENHI grant numbers 20H03543 and 21K19422 (Y.K.); Takara Bio, Inc (Y.K.); Aichi Cancer Center Joint Research Project on Priority Areas (Y.K.); the Princess Takamatsunomiya Cancer Research Foundation (Y.K.); Takeda Science Foundation (Y.K.); The Cell Science Research Foundation (Y.K.); Uehara Memorial Foundation (Y.K.); Japanese Society of Hematology Research Grant (Y.K.); Astellas Foundation for Research on Metabolic Disorders (Y.K.); SGH Foundation (Y.K.); Multiple Myeloma Research Grant from Myeloma Patients and Families (Y.K.); Kobayashi Foundation for Cancer Research (Y.K.); Project Mirai Cancer Research Grants (Y.K.); KAKENHI grant numbers 22K15575 (Y.I.) and 22H02910 (S.I.); Japanese Government (Monbukagakusho: MEXT) Scholarship (Y.L.); and Otsuka Toshimi Scholarship Foundation (H.Z.).
Author contributions
Y.K. designed the project. T.Y., Y.I., Z.W., S.I., H.K., T.N., H.Z., Y.L., T.M., and Y.K. performed the experiments. S.O. and Y.A. designed lentivirus vectors and contributed to the writing of the manuscript. T.Y. and Y.K. analyzed the results and wrote the manuscript.
Declaration of interests
This study was partly supported by a commercial research grant from Takara Bio, Inc. S.O. and Y.A. are employees of Takara Bio Inc. Aichi Prefecture has filed a patent application related to this study on which Y.K. and T.Y. are named as inventors.
Declaration of generative AI and AI-assisted technologies in the writing process
During the preparation of the present work, the authors used DeepL Write (https://www.deepl.com/write) in order to correct spelling and grammatical mistakes. The authors thoroughly reviewed and edited the texts after using this tool. The authors take full responsibility for the final content of the paper.
Published: April 25, 2024
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xcrm.2024.101526.
Supplemental information
References
- 1.Maude S.L., Laetsch T.W., Buechner J., Rives S., Boyer M., Bittencourt H., Bader P., Verneris M.R., Stefanski H.E., Myers G.D., et al. Tisagenlecleucel in Children and Young Adults with B-Cell Lymphoblastic Leukemia. N. Engl. J. Med. 2018;378:439–448. doi: 10.1056/NEJMoa1709866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Locke F.L., Ghobadi A., Jacobson C.A., Miklos D.B., Lekakis L.J., Oluwole O.O., Lin Y., Braunschweig I., Hill B.T., Timmerman J.M., et al. Long-term safety and activity of axicabtagene ciloleucel in refractory large B-cell lymphoma (ZUMA-1): a single-arm, multicentre, phase 1-2 trial. Lancet Oncol. 2019;20:31–42. doi: 10.1016/s1470-2045(18)30864-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Abramson J.S., Palomba M.L., Gordon L.I., Lunning M.A., Wang M., Arnason J., Mehta A., Purev E., Maloney D.G., Andreadis C., et al. Lisocabtagene maraleucel for patients with relapsed or refractory large B-cell lymphomas (TRANSCEND NHL 001): a multicentre seamless design study. Lancet. 2020;396:839–852. doi: 10.1016/S0140-6736(20)31366-0. [DOI] [PubMed] [Google Scholar]
- 4.Wang M., Munoz J., Goy A., Locke F.L., Jacobson C.A., Hill B.T., Timmerman J.M., Holmes H., Jaglowski S., Flinn I.W., et al. KTE-X19 CAR T-Cell Therapy in Relapsed or Refractory Mantle-Cell Lymphoma. N. Engl. J. Med. 2020;382:1331–1342. doi: 10.1056/NEJMoa1914347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Munshi N.C., Anderson L.D., Jr., Shah N., Madduri D., Berdeja J., Lonial S., Raje N., Lin Y., Siegel D., Oriol A., et al. Idecabtagene Vicleucel in Relapsed and Refractory Multiple Myeloma. N. Engl. J. Med. 2021;384:705–716. doi: 10.1056/NEJMoa2024850. [DOI] [PubMed] [Google Scholar]
- 6.Fraietta J.A., Lacey S.F., Orlando E.J., Pruteanu-Malinici I., Gohil M., Lundh S., Boesteanu A.C., Wang Y., O'Connor R.S., Hwang W.T., et al. Determinants of response and resistance to CD19 chimeric antigen receptor (CAR) T cell therapy of chronic lymphocytic leukemia. Nat. Med. 2018;24:563–571. doi: 10.1038/s41591-018-0010-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Raje N., Berdeja J., Lin Y., Siegel D., Jagannath S., Madduri D., Liedtke M., Rosenblatt J., Maus M.V., Turka A., et al. Anti-BCMA CAR T-Cell Therapy bb2121 in Relapsed or Refractory Multiple Myeloma. N. Engl. J. Med. 2019;380:1726–1737. doi: 10.1056/NEJMoa1817226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gardner R.A., Finney O., Annesley C., Brakke H., Summers C., Leger K., Bleakley M., Brown C., Mgebroff S., Kelly-Spratt K.S., et al. Intent-to-treat leukemia remission by CD19 CAR T cells of defined formulation and dose in children and young adults. Blood. 2017;129:3322–3331. doi: 10.1182/blood-2017-02-769208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Shum T., Omer B., Tashiro H., Kruse R.L., Wagner D.L., Parikh K., Yi Z., Sauer T., Liu D., Parihar R., et al. Constitutive Signaling from an Engineered IL7 Receptor Promotes Durable Tumor Elimination by Tumor-Redirected T Cells. Cancer Discov. 2017;7:1238–1247. doi: 10.1158/2159-8290.CD-17-0538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kagoya Y., Tanaka S., Guo T., Anczurowski M., Wang C.H., Saso K., Butler M.O., Minden M.D., Hirano N. A novel chimeric antigen receptor containing a JAK-STAT signaling domain mediates superior antitumor effects. Nat. Med. 2018;24:352–359. doi: 10.1038/nm.4478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chmielewski M., Abken H. CAR T Cells Releasing IL-18 Convert to T-Bet(high) FoxO1(low) Effectors that Exhibit Augmented Activity against Advanced Solid Tumors. Cell Rep. 2017;21:3205–3219. doi: 10.1016/j.celrep.2017.11.063. [DOI] [PubMed] [Google Scholar]
- 12.Ma X., Shou P., Smith C., Chen Y., Du H., Sun C., Porterfield Kren N., Michaud D., Ahn S., Vincent B., et al. Interleukin-23 engineering improves CAR T cell function in solid tumors. Nat. Biotechnol. 2020;38:448–459. doi: 10.1038/s41587-019-0398-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Adachi K., Kano Y., Nagai T., Okuyama N., Sakoda Y., Tamada K. IL-7 and CCL19 expression in CAR-T cells improves immune cell infiltration and CAR-T cell survival in the tumor. Nat. Biotechnol. 2018;36:346–351. doi: 10.1038/nbt.4086. [DOI] [PubMed] [Google Scholar]
- 14.Yeku O.O., Purdon T.J., Koneru M., Spriggs D., Brentjens R.J. Armored CAR T cells enhance antitumor efficacy and overcome the tumor microenvironment. Sci. Rep. 2017;7 doi: 10.1038/s41598-017-10940-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hurton L.V., Singh H., Najjar A.M., Switzer K.C., Mi T., Maiti S., Olivares S., Rabinovich B., Huls H., Forget M.A., et al. Tethered IL-15 augments antitumor activity and promotes a stem-cell memory subset in tumor-specific T cells. Proc. Natl. Acad. Sci. USA. 2016;113:E7788–E7797. doi: 10.1073/pnas.1610544113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Morris E.C., Neelapu S.S., Giavridis T., Sadelain M. Cytokine release syndrome and associated neurotoxicity in cancer immunotherapy. Nat. Rev. Immunol. 2022;22:85–96. doi: 10.1038/s41577-021-00547-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Giavridis T., van der Stegen S.J.C., Eyquem J., Hamieh M., Piersigilli A., Sadelain M. CAR T cell-induced cytokine release syndrome is mediated by macrophages and abated by IL-1 blockade. Nat. Med. 2018;24:731–738. doi: 10.1038/s41591-018-0041-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Norelli M., Camisa B., Barbiera G., Falcone L., Purevdorj A., Genua M., Sanvito F., Ponzoni M., Doglioni C., Cristofori P., et al. Monocyte-derived IL-1 and IL-6 are differentially required for cytokine-release syndrome and neurotoxicity due to CAR T cells. Nat. Med. 2018;24:739–748. doi: 10.1038/s41591-018-0036-4. [DOI] [PubMed] [Google Scholar]
- 19.Kotch C., Barrett D., Teachey D.T. Tocilizumab for the treatment of chimeric antigen receptor T cell-induced cytokine release syndrome. Expert Rev. Clin. Immunol. 2019;15:813–822. doi: 10.1080/1744666x.2019.1629904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Le R.Q., Li L., Yuan W., Shord S.S., Nie L., Habtemariam B.A., Przepiorka D., Farrell A.T., Pazdur R. FDA Approval Summary: Tocilizumab for Treatment of Chimeric Antigen Receptor T Cell-Induced Severe or Life-Threatening Cytokine Release Syndrome. Oncol. 2018;23:943–947. doi: 10.1634/theoncologist.2018-0028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Schuster S.J., Bishop M.R., Tam C.S., Waller E.K., Borchmann P., McGuirk J.P., Jäger U., Jaglowski S., Andreadis C., Westin J.R., et al. Tisagenlecleucel in Adult Relapsed or Refractory Diffuse Large B-Cell Lymphoma. N. Engl. J. Med. 2019;380:45–56. doi: 10.1056/NEJMoa1804980. [DOI] [PubMed] [Google Scholar]
- 22.Teachey D.T., Lacey S.F., Shaw P.A., Melenhorst J.J., Maude S.L., Frey N., Pequignot E., Gonzalez V.E., Chen F., Finklestein J., et al. Identification of Predictive Biomarkers for Cytokine Release Syndrome after Chimeric Antigen Receptor T-cell Therapy for Acute Lymphoblastic Leukemia. Cancer Discov. 2016;6:664–679. doi: 10.1158/2159-8290.Cd-16-0040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Brentjens R.J., Davila M.L., Riviere I., Park J., Wang X., Cowell L.G., Bartido S., Stefanski J., Taylor C., Olszewska M., et al. CD19-targeted T cells rapidly induce molecular remissions in adults with chemotherapy-refractory acute lymphoblastic leukemia. Sci. Transl. Med. 2013;5 doi: 10.1126/scitranslmed.3005930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hay K.A., Hanafi L.A., Li D., Gust J., Liles W.C., Wurfel M.M., López J.A., Chen J., Chung D., Harju-Baker S., et al. Kinetics and biomarkers of severe cytokine release syndrome after CD19 chimeric antigen receptor-modified T-cell therapy. Blood. 2017;130:2295–2306. doi: 10.1182/blood-2017-06-793141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Santomasso B.D., Park J.H., Salloum D., Riviere I., Flynn J., Mead E., Halton E., Wang X., Senechal B., Purdon T., et al. Clinical and Biological Correlates of Neurotoxicity Associated with CAR T-cell Therapy in Patients with B-cell Acute Lymphoblastic Leukemia. Cancer Discov. 2018;8:958–971. doi: 10.1158/2159-8290.Cd-17-1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kadauke S., Myers R.M., Li Y., Aplenc R., Baniewicz D., Barrett D.M., Barz Leahy A., Callahan C., Dolan J.G., Fitzgerald J.C., et al. Risk-Adapted Preemptive Tocilizumab to Prevent Severe Cytokine Release Syndrome After CTL019 for Pediatric B-Cell Acute Lymphoblastic Leukemia: A Prospective Clinical Trial. J. Clin. Oncol. 2021;39:920–930. doi: 10.1200/jco.20.02477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Caimi P.F., Pacheco Sanchez G., Sharma A., Otegbeye F., Ahmed N., Rojas P., Patel S., Kleinsorge Block S., Schiavone J., Zamborsky K., et al. Prophylactic Tocilizumab Prior to Anti-CD19 CAR-T Cell Therapy for Non-Hodgkin Lymphoma. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.745320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wehrli M., Gallagher K., Chen Y.-B., Leick M.B., McAfee S.L., El-Jawahri A.R., DeFilipp Z., Horick N., O'Donnell P., Spitzer T., et al. Single-center experience using anakinra for steroid-refractory immune effector cell-associated neurotoxicity syndrome (ICANS) J. Immunother. Cancer. 2022;10 doi: 10.1136/jitc-2021-003847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Strati P., Ahmed S., Kebriaei P., Nastoupil L.J., Claussen C.M., Watson G., Horowitz S.B., Brown A.R.T., Do B., Rodriguez M.A., et al. Clinical efficacy of anakinra to mitigate CAR T-cell therapy-associated toxicity in large B-cell lymphoma. Blood Adv. 2020;4:3123–3127. doi: 10.1182/bloodadvances.2020002328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ancey C., Küster A., Haan S., Herrmann A., Heinrich P.C., Müller-Newen G. A fusion protein of the gp130 and interleukin-6Ralpha ligand-binding domains acts as a potent interleukin-6 inhibitor. J. Biol. Chem. 2003;278:16968–16972. doi: 10.1074/jbc.C300081200. [DOI] [PubMed] [Google Scholar]
- 31.Markley J.C., Sadelain M. IL-7 and IL-21 are superior to IL-2 and IL-15 in promoting human T cell-mediated rejection of systemic lymphoma in immunodeficient mice. Blood. 2010;115:3508–3519. doi: 10.1182/blood-2009-09-241398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Shochat C., Tal N., Bandapalli O.R., Palmi C., Ganmore I., te Kronnie G., Cario G., Cazzaniga G., Kulozik A.E., Stanulla M., et al. Gain-of-function mutations in interleukin-7 receptor-α (IL7R) in childhood acute lymphoblastic leukemias. J. Exp. Med. 2011;208:901–908. doi: 10.1084/jem.20110580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Castell J.V., Geiger T., Gross V., Andus T., Walter E., Hirano T., Kishimoto T., Heinrich P.C. Plasma clearance, organ distribution and target cells of interleukin-6/hepatocyte-stimulating factor in the rat. Eur. J. Biochem. 1988;177:357–361. doi: 10.1111/j.1432-1033.1988.tb14384.x. [DOI] [PubMed] [Google Scholar]
- 34.Foxwell B.M., Beadling C., Guschin D., Kerr I., Cantrell D. Interleukin-7 can induce the activation of Jak 1, Jak 3 and STAT 5 proteins in murine T cells. Eur. J. Immunol. 1995;25:3041–3046. doi: 10.1002/eji.1830251109. [DOI] [PubMed] [Google Scholar]
- 35.Swainson L., Kinet S., Mongellaz C., Sourisseau M., Henriques T., Taylor N. IL-7–induced proliferation of recent thymic emigrants requires activation of the PI3K pathway. Blood. 2007;109:1034–1042. doi: 10.1182/blood-2006-06-027912. [DOI] [PubMed] [Google Scholar]
- 36.Osborne L.C., Dhanji S., Snow J.W., Priatel J.J., Ma M.C., Miners M.J., Teh H.-S., Goldsmith M.A., Abraham N. Impaired CD8 T cell memory and CD4 T cell primary responses in IL-7Rα mutant mice. J. Exp. Med. 2007;204:619–631. doi: 10.1084/jem.20061871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Cui G., Shimba A., Ma G., Takahara K., Tani-ichi S., Zhu Y., Asahi T., Abe A., Miyachi H., Kitano S., et al. IL-7R–Dependent Phosphatidylinositol 3-Kinase Competes with the STAT5 Signal to Modulate T Cell Development and Homeostasis. J. Immunol. 2020;204:844–857. doi: 10.4049/jimmunol.1900456. [DOI] [PubMed] [Google Scholar]
- 38.Kratochvíl M., Koladiya A., Vondrášek J. Generalized EmbedSOM on quadtree-structured self-organizing maps. F1000Res. 2019;8:2120. doi: 10.12688/f1000research.21642.2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Fontenot J.D., Rasmussen J.P., Gavin M.A., Rudensky A.Y. A function for interleukin 2 in Foxp3-expressing regulatory T cells. Nat. Immunol. 2005;6:1142–1151. doi: 10.1038/ni1263. [DOI] [PubMed] [Google Scholar]
- 40.Waldmann T.A., Chen J. Disorders of the JAK/STAT Pathway in T Cell Lymphoma Pathogenesis: Implications for Immunotherapy. Annu. Rev. Immunol. 2017;35:533–550. doi: 10.1146/annurev-immunol-110416-120628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sterner R.M., Sakemura R., Cox M.J., Yang N., Khadka R.H., Forsman C.L., Hansen M.J., Jin F., Ayasoufi K., Hefazi M., et al. GM-CSF inhibition reduces cytokine release syndrome and neuroinflammation but enhances CAR-T cell function in xenografts. Blood. 2019;133:697–709. doi: 10.1182/blood-2018-10-881722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bailey S.R., Vatsa S., Larson R.C., Bouffard A.A., Scarfò I., Kann M.C., Berger T.R., Leick M.B., Wehrli M., Schmidts A., et al. Blockade or Deletion of IFNγ Reduces Macrophage Activation without Compromising CAR T-cell Function in Hematologic Malignancies. Blood Cancer Discov. 2022;3:136–153. doi: 10.1158/2643-3230.Bcd-21-0181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.McMahan C.J., Slack J.L., Mosley B., Cosman D., Lupton S.D., Brunton L.L., Grubin C.E., Wignall J.M., Jenkins N.A., Brannan C.I. A novel IL-1 receptor, cloned from B cells by mammalian expression, is expressed in many cell types. EMBO J. 1991;10:2821–2832. doi: 10.1002/j.1460-2075.1991.tb07831.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Orlando S., Matteucci C., Fadlon E.J., Buurman W.A., Bardella M.T., Colotta F., Introna M., Mantovani A. TNF-alpha, unlike other pro- and anti-inflammatory cytokines, induces rapid release of the IL-1 type II decoy receptor in human myelomonocytic cells. J. Immunol. 1997;158:3861–3868. [PubMed] [Google Scholar]
- 45.Vendramin R., Katopodi V., Cinque S., Konnova A., Knezevic Z., Adnane S., Verheyden Y., Karras P., Demesmaeker E., Bosisio F.M., et al. Activation of the integrated stress response confers vulnerability to mitoribosome-targeting antibiotics in melanoma. J. Exp. Med. 2021;218 doi: 10.1084/jem.20210571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Henriques C.M., Rino J., Nibbs R.J., Graham G.J., Barata J.T. IL-7 induces rapid clathrin-mediated internalization and JAK3-dependent degradation of IL-7Ralpha in T cells. Blood. 2010;115:3269–3277. doi: 10.1182/blood-2009-10-246876. [DOI] [PubMed] [Google Scholar]
- 47.Xue L., Yi Y., Xu Q., Wang L., Yang X., Zhang Y., Hua X., Chai X., Yang J., Chen Y., et al. Chimeric antigen receptor T cells self-neutralizing IL6 storm in patients with hematologic malignancy. Cell Discov. 2021;7:84. doi: 10.1038/s41421-021-00299-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.van Dam M., Müllberg J., Schooltink H., Stoyan T., Brakenhoff J.P., Graeve L., Heinrich P.C., Rose-John S. Structure-function analysis of interleukin-6 utilizing human/murine chimeric molecules. Involvement of two separate domains in receptor binding. J. Biol. Chem. 1993;268:15285–15290. [PubMed] [Google Scholar]
- 49.Hand T.W., Cui W., Jung Y.W., Sefik E., Joshi N.S., Chandele A., Liu Y., Kaech S.M. Differential effects of STAT5 and PI3K/AKT signaling on effector and memory CD8 T-cell survival. Proc. Natl. Acad. Sci. USA. 2010;107:16601–16606. doi: 10.1073/pnas.1003457107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Previte D.M., Martins C.P., O’Connor E.C., Marre M.L., Coudriet G.M., Beck N.W., Menk A.V., Wright R.H., Tse H.M., Delgoffe G.M., Piganelli J.D. Lymphocyte Activation Gene-3 Maintains Mitochondrial and Metabolic Quiescence in Naive CD4+ T Cells. Cell Rep. 2019;27:129–141.e4. doi: 10.1016/j.celrep.2019.03.004. [DOI] [PubMed] [Google Scholar]
- 51.Quinn K.M., Hussain T., Kraus F., Formosa L.E., Lam W.K., Dagley M.J., Saunders E.C., Assmus L.M., Wynne-Jones E., Loh L., et al. Metabolic characteristics of CD8+ T cell subsets in young and aged individuals are not predictive of functionality. Nat. Commun. 2020;11:2857. doi: 10.1038/s41467-020-16633-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Scharping N.E., Menk A.V., Moreci R.S., Whetstone R.D., Dadey R.E., Watkins S.C., Ferris R.L., Delgoffe G.M. The tumor microenvironment represses T cell mitochondrial biogenesis to drive intratumoral T cell metabolic insufficiency and dysfunction. Immunity. 2016;45:701–703. doi: 10.1016/j.immuni.2016.08.009. [DOI] [PubMed] [Google Scholar]
- 53.Banerjee A., Li D., Guo Y., Mei Z., Lau C., Chen K., Westwick J., Klauda J.B., Schrum A., Lazear E.R., Krupnick A.S. A reengineered common chain cytokine augments CD8+ T cell–dependent immunotherapy. JCI Insight. 2022;7 doi: 10.1172/jci.insight.158889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Weber E.W., Parker K.R., Sotillo E., Lynn R.C., Anbunathan H., Lattin J., Good Z., Belk J.A., Daniel B., Klysz D., et al. Transient rest restores functionality in exhausted CAR-T cells through epigenetic remodeling. Science. 2021;372 doi: 10.1126/science.aba1786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Cui W., Liu Y., Weinstein J.S., Craft J., Kaech S.M. An interleukin-21-interleukin-10-STAT3 pathway is critical for functional maturation of memory CD8+ T cells. Immunity. 2011;35:792–805. doi: 10.1016/j.immuni.2011.09.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Siegel A.M., Heimall J., Freeman A.F., Hsu A.P., Brittain E., Brenchley J.M., Douek D.C., Fahle G.H., Cohen J.I., Holland S.M., Milner J.D. A Critical Role for STAT3 Transcription Factor Signaling in the Development and Maintenance of Human T Cell Memory. Immunity. 2011;35:806–818. doi: 10.1016/j.immuni.2011.09.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Micklethwaite K.P., Gowrishankar K., Gloss B.S., Li Z., Street J.A., Moezzi L., Mach M.A., Sutrave G., Clancy L.E., Bishop D.C., et al. Investigation of product-derived lymphoma following infusion of piggyBac-modified CD19 chimeric antigen receptor T cells. Blood. 2021;138:1391–1405. doi: 10.1182/blood.2021010858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Lange S., Sand L.G.L., Bell M., Patil S.L., Langfitt D., Gottschalk S. A Chimeric GM-CSF/IL18 Receptor to Sustain CAR T-cell Function. Cancer Discov. 2021;11:1661–1671. doi: 10.1158/2159-8290.Cd-20-0896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yoshikawa T., Wu Z., Inoue S., Kasuya H., Matsushita H., Takahashi Y., Kuroda H., Hosoda W., Suzuki S., Kagoya Y. Genetic ablation of PRDM1 in antitumor T cells enhances therapeutic efficacy of adoptive immunotherapy. Blood. 2022;139:2156–2172. doi: 10.1182/blood.2021012714. [DOI] [PubMed] [Google Scholar]
- 60.Nicholson I.C., Lenton K.A., Little D.J., Decorso T., Lee F.T., Scott A.M., Zola H., Hohmann A.W. Construction and characterisation of a functional CD19 specific single chain Fv fragment for immunotherapy of B lineage leukaemia and lymphoma. Mol. Immunol. 1997;34:1157–1165. doi: 10.1016/s0161-5890(97)00144-2. [DOI] [PubMed] [Google Scholar]
- 61.Li Q., Verschraegen C.F., Mendoza J., Hassan R. Cytotoxic activity of the recombinant anti-mesothelin immunotoxin, SS1(dsFv)PE38, towards tumor cell lines established from ascites of patients with peritoneal mesotheliomas. Anticancer Res. 2004;24:1327–1335. [PubMed] [Google Scholar]
- 62.Rossig C., Nuchtern J.G., Brenner M.K. Selection of human antitumor single-chain Fv antibodies from the B-cell repertoire of patients immunized against autologous neuroblastoma. Med. Pediatr. Oncol. 2000;35:692–695. doi: 10.1002/1096-911x(20001201)35:6<692::aid-mpo45>3.0.co;2-7. [DOI] [PubMed] [Google Scholar]
- 63.Horwacik I., Golik P., Grudnik P., Kolinski M., Zdzalik M., Rokita H., Dubin G. Structural Basis of GD2 Ganglioside and Mimetic Peptide Recognition by 14G2a Antibody. Mol. Cell. Proteomics. 2015;14:2577–2590. doi: 10.1074/mcp.M115.052720. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Chen S., Zhou Y., Chen Y., Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–i890. doi: 10.1093/bioinformatics/bty560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kim D., Paggi J.M., Park C., Bennett C., Salzberg S.L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37:907–915. doi: 10.1038/s41587-019-0201-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Anders S., Pyl P.T., Huber W. HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–169. doi: 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Robinson M.D., McCarthy D.J., Smyth G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–140. doi: 10.1093/bioinformatics/btp616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Liao Y., Smyth G.K., Shi W. The R. package Rsubread is easier, faster, cheaper and better for alignment and quantification of RNA sequencing reads. Nucleic Acids Res. 2019;47:e47. doi: 10.1093/nar/gkz114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Van Gassen S., Callebaut B., Van Helden M.J., Lambrecht B.N., Demeester P., Dhaene T., Saeys Y. FlowSOM: Using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A. 2015;87:636–645. doi: 10.1002/cyto.a.22625. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The RNA-seq datasets presented in Figures 2 and S3 have been deposited in the NCBI Gene Expression Omnibus under the accession number GSE216734 and GSE255145, respectively. Mass cytometry data were deposited in FlowRepository (FR-FCM-Z77Y). No original code is included in this paper. Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.