Abstract
The presence of veterinary drug residues in aquatic products represents a significant challenge to food safety. The current detection methods, limited in both scope and sensitivity, underscore the urgent need for more advanced techniques. This research introduces a swift and potent screening technique using high-performance liquid chromatography-high-resolution mass spectrometry (HPLC-HRMS) and a refined QuEChERS protocol, allowing simultaneous qualitative and semi-quantitative analysis of 192 residues. A comprehensive database, employing full scan mode and data-dependent secondary mass spectroscopy, enhances screening accuracy. The method involves efficient extraction using 90% acetonitrile, dehydration with Na2SO4, and acetic acid, followed by cleanup using dispersive solid-phase extract sorbent primary secondary amine. It is suitable for samples with varying fat content, offering detection limits ranging from 0.5 to 10 μg/kg, high recovery rates (60–120%), and low relative standard deviations (<20%). Practical application has validated its effectiveness for multi-residue screening, marking a significant advancement in food safety evaluation.
Keywords: High resolution mass spectrometry, QuEChERS, Veterinary drug, Aquatic products, Screening
Chemical compounds studied in this article: Amantadine (PubChem CID: 2130), Tetracyclines (PubChem CID: 54675776), Chloramphenicol (PubChem CID: 5959), Malachite green (PubChem CID: 11294), Metronidazole (PubChem CID: 4173), Norfloxacin (PubChem CID: 4539), Enrofloxacin (PubChem CID: 71188), Sulfadiazine (PubChem CID: 5215), Sulfamethoxazole (PubChem CID: 5329), Diazepam (PubChem CID: 3016)
Highlights
-
•
Simultaneous detection of 15 classes of drug residue in aquatic products by HPLC-HRMS.
-
•
An extensive database involving 192 veterinary drugs was constructed.
-
•
A rapid and simple sample preparation method was developed.
-
•
The method is suitable for samples with variable fat content.
1. Introduction
Since the last century, the significance of aquaculture as a source of nutrition has greatly increased, drawing attention to the misuse and residues of veterinary drugs in aquaculture products from both consumers and governments (Kong, Wang, Huang, & Yu, 2018; Park, Choi, Kang, Kwon, & j., 2022; Uchida et al., 2016; Zhang, Bleeker, & Liu, 2015). Consequently, there is an urgent need for advanced technologies to monitor these residues to guarantee the safety and health of consumers (Castilla-Fernández et al., 2021; Wang et al., 2021; Yu et al., 2023). Several techniques have been established for detecting veterinary drugs in aquatic animal foods, among which enzyme-linked immunosorbent assay (ELISA), gas chromatography–mass spectrometry (GC–MS), and high performance liquid chromatography-mass spectrometry (HPLC-MS) are the most widely used (Han et al., 2022; Jung et al., 2021; Shahsavari et al., 2022; Zhu et al., 2016). Despite their effectiveness, these methods have limitations such as the narrow range of detectable compounds in a single test, the requirement for extensive mass spectrometry optimization for each compound, sensitivity to matrix interference, and lower resolution, potentially leading to false positives. However, multi-residue veterinary drug detection techniques can enhance efficiency and lower environmental impact by reducing reliance on organic solvents, and provid an essential advancement for aquatic product safety and consumer health safeguarding. High-performance liquid chromatography-high-resolution mass spectrometry (HPLC-HRMS) has emerged as a superior technique, known for its accuracy in the quick detection of drug residues (Casado, Brigden, Santillo, & Johnston, 2019; Magalhaes, Freitas, Sofia Vila Pouca, Barbosa, & Ramos, 2020; Mehl, Schmidt, Schmidt, & Morlock, 2021; Zhang et al., 2016).
The “Quick, Easy, Cheap, Effective, Rugged, and Safe” (QuEChERS) extraction method, initially designed for pesticide analysis (Anastassiades, Lehotay, Štajnbaher, & Schenck, 2019), has shown to be more favorable compared to traditional methods like liquid-liquid extraction (LLE)(Lopes, Reyes, Romero-Gonzalez, Vidal, & Frenich, 2012; Shin et al., 2018; Zhang, Guo, Yan, Sun, & Zhang, 2015) and solid-phase extraction (SPE)(Melekhin et al., 2022; Wang et al., 2022; Zhao et al., 2017). Its simplicity and high extraction efficiency make it suitable for a variety of applications, including extracting veterinary drug residues from complex matrices like aquatic products (Gonzalez-Curbelo, Varela-Martinez, & Riano-Herrera, 2022; Lazartigues et al., 2011; Monteiro et al., 2022; Syrgabek & Alimzhanova, 2022). The adaptability of QuEChERS to the complex aquatic matrices results in more efficient extraction for trace residues detection. Its effectiveness in large-scale residue screening is well documented, along with its ability to be applied in various matrices without the need for salting out or intricate cleanup processes (Wang et al., 2023; Wang, Tian, Ai, & Liang, 2023; Zhao et al., 2017).
Despite these advancements, current meths still have limitations that need to be addressed, such as the limited number of screened drugs and unsatisfactory recovery rates. The comprehensive validation of sample preparation methods across different matrices are also insufficient, highlighting the need for a more adaptable screening range. Additionally, there is a pressing need to expand the drug screening database.
This study developed a simple, rapid, sensitive, and accurate screening strategy for multiple veterinary drug residues in aquatic products. A database was established including retention time, accurate mass-to-charge ratio of precursors and fragments, and screening protocols for 192 drugs across 15 categories. After optimizing and validating the extraction conditions, including the addition of sodium sulfate, acidic mediums, adsorbents, redissolution solutions, and filtration for products with varying fat content exemplified by grass carp and crab, the method was applied to thirty locally sourced samples. This approach enables the detection of a wide range of veterinary drugs, meeting the stringent residue limits set by the European Union and China. Covering 15 categories and 192 veterinary drugs, it plays an essential role in the prompt and precise evaluation of drug residues in aquatic products, thereby minimizing associated safety incidents.
2. Materials and methods
2.1. Materials
All 192 veterinary drug standards, as detailed in Table S1, were obtained at high purity from First Standard (Tianjin, China). These standards encompass fifteen distinct categories: twenty-four β-agonists, three antiviral drugs, eleven macrolides, four avermectins, twenty-seven sulfonamides, nineteen fluoroquinolones, four tetracyclines, four amide alcohols, six nitrofurans, twenty-seven nitroimidazoles, benzimidazoles and their metabolites, thirty-two steroid hormones, nineteen sedatives, four triphenylmethane compounds, and eight other types of pharmaceuticals. HPLC-grade acetonitrile (ACN) and methanol (MeOH) were purchased from ANPEL Laboratory Technologies (Shanghai) Inc. Dichloromethane (DCM), ethyl acetate, acetic acid, and n-hexane were supplied by JT Baker (USA). Formic acid (FA) was procured from Fluka (Germany). Neutral aluminum oxide (NA), primary secondary amine (PSA), octadecylsilyl (C18), and polar-enhanced polymer (PEP) sorbents were provided by Agela Technologies. Citric acid monohydrate, disodium hydrogen phosphate dodecahydrate, tetramethyl-p-phenylenediamine dihydrochloride (TMPD), and anhydrous sodium sulfate were purchased from Sinopharm Chemical Reagent Co., Ltd. The experiment utilized deionized water from the Milli-Q® direct water purification system.
2.2. Instrumentation
The instrumental analysis was conducted using an HPLC system coupled to a Q-Exactive™ benchtop mass spectrometer with a heated electrospray ionization source (Thermo Fisher Scientific, San Francisco, USA). The HPLC system utilized was a Dionex Ultimate 3000, equipped with an autosampler, a binary pump, and a solvent degasser, performing chromatographic separation on a ZORBAX Eclipse Plus C18 RRHD column. It was maintained at 40 °C in the column oven. The mobile phase consisted of ultrapure water with 5 mM/L ammonium acetate (A) and MeOH (B), each containing 0.2% FA, at a flow rate of 0.4 mL/min following a specific gradient elution profile detailed in the protocol. The gradient elution protocol was executed as follows: 0 min, 5% B; 1 min, 5% B; 3 min, 20% B; 8 min, 50% B; 19 min, 95% B; 23 min, 95% B; 23.1 min, 5% B; 27 min, 5% B. The injection volume remained constant at 4 μL throughout the analysis.
The mass spectrometer was configured with an electrospray voltage of ±4.0 kV, sheath gas flow at 50 psi, sweep gas at 0 psi, and auxiliary gas at 13 psi. The capillary temperature was maintained at 320 °C with an auxiliary gas heater set to 40 °C. The S-lens RF level was adjusted to 70. Full-scan and data-dependent MS2 scans were carried out with a loop count of 3 and an isolation window of 1.5 m/z. The detection range covered 70 to 1050 m/z, with resolutions of 70,000 for full MS analysis and 17,500 for dd-MS2 scans.
2.3. Database establishment and identification of compounds
To facilitate qualitative screening, the database was established through ultra-HPLC coupled to an Orbitrap HRMS, utilizing TraceFinder EFS3.0 software for data management. It comprises comprehensive chemical and analytical profiles for all compounds, including names, CAS numbers, molecular formulas, weights, structures, types, chromatographic retention times, and mass spectrometric details (high-resolution spectra and secondary fragments). Calibration standards were introduced under finely tuned conditions to enrich the database with accurate mass measurements and fragmentation patterns, thereby enabling the confident identification in further analyses. The identification of target compounds within the samples was achieved by correlating retention times (within 0.1 min), precursor ions (with a mass deviation <10 ppm), and a single fragment ion (with a mass deviation <20 ppm) against the database. This comparison was executed using TraceFinder EFS 3.0 software.
2.4. Sample preparation and pretreatment
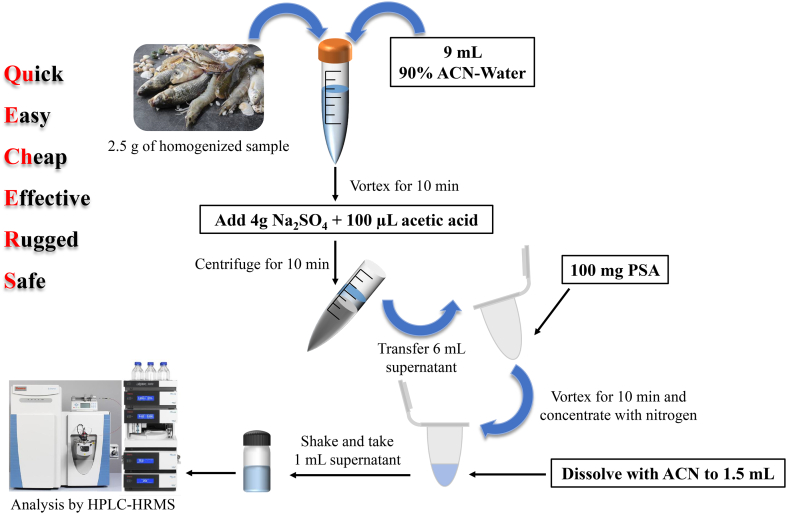
The schematic diagram of procedures for the Rapid Screening of 192 Veterinary Drug Residues in Aquatic Products using HPLC-HRMS Coupled with QuEChERS is depicted in Fig. 1. Local market samples of grass carp and crab were homogenized and kept at −18 °C for analysis. Blank samples were utilized to prepare standards that matched the matrix and assess the recovery rates of analytes via spiking experiments. Prior to the spiking experiment and preparation of matrix-matched standards, the samples were tested to confirm the absence of analytes.
Fig. 1.
Schematic diagram of procedures for Rapid Screening of 192 Veterinary Drug Residues in Aquatic Products using HPLC-HRMS Coupled with QuEChERS.
Prior to processing, the samples were thawed to room temperature. A 2.50 g ± 0.05 g sample was carefully weighed and transferred into a 50 mL centrifuge tube. Subsequently, 9 mL of water-ACN (1:9 v/v) solution was added, followed by thorough vortexing, and the addition of 4 g of Na2SO4 and 100 μL of acetic acid. The mixture was then centrifuged at 10,000 rpm for 10 min, after which 6 mL of the supernatant was mixed with 100 mg PSA, vortexed for 10 min, and then 3 mL supernatant was reduced under nitrogen to 0.5 mL at 35 °C. The concentrated solution was then filtered through a 0.22 μm H-PTFE filter before HPLC-HRMS analysis.
2.5. Method validation
2.5.1. Matrix effect (ME)
The ME in various aquatic products after pretreatment were calculated by evaluating the peak area ratio of a 20 ng/mL matrix-matched standard solution to that of a solvent standard solution, utilizing the formula ME (%) = (1 - Ab/As) × 100%, where Ab and As denote the peak areas of the matrix matching standard solution and the solvent standard solution, respectively.
2.5.2. Screening detection limit (SDL)
Based on the pretreatment method in Section 2.4, samples were spiked with a mixed standard solution to achieve target concentrations of 0.5, 1, 2, 5, and 10 μg/kg in various aquatic products. This process was conducted with six replicates for each concentration level. SDLs for each compound were determined in this study by introducing specific concentrations into aquatic samples and observing the consistency of detection across all six replicates. The validation process was meticulously aligned with the SANTE/2019/12682 guidelines, setting the SDL at the lowest spiked concentration where consistent detection was achievable. This approach ensures the stability of the screening method in reliably identifying trace-level residues, thereby adhering to stringent regulatory standards (De Paepe et al., 2019; Wang, Tian, et al., 2023; Wang, Yang, et al., 2023).
2.5.3. Accuracy and precision
Accuracy and precision were verified across different matrices at the designated spiking concentrations for various compounds. The validation process included six replicate analyses per concentration level. Quantification was performed by comparing the ratios between the samples and those of the single-point matrix-matched standard solutions for each compound, which were then multiplied by the concentration of standard solutions to get the concentration in the samples. Recovery was calculated using the formula: Recovery (%) = (A0/Ab) × 100%, where A0 represents the peak area of the spiked matrix-matched solution, and Ab denotes the peak area of the matrix-matching standard solution.
2.6. Statistical analysis
Statistical analysis of the experimental data was conducted using Microsoft Excel. The values in the research were reported as averages based on either three or six replicates. The significance difference test between groups was performed at a threshold of p < 0.05 using the SPSS 16.0 software package (SPSS Inc., Chicago, IL., USA). Analysis was conducted via variance (ANOVA) followed by Duncan's test.
3. Results and discussion
3.1. Mass database and data analysis
Detecting the presence of target compounds in samples is crucial and entails comparing the compound list to a multi-residue drug database. The process begins with automatically determining the molecular formulas, ionization modes, and adduct ions of protonated compounds using mass calculation software. Data for 192 candidate compounds, including retention time, m/z of precursors, and fragment ions, were systematically collected via HPLC-HRMS analysis using a standard solution at a concentration of 100 μg/L. Finally, this data was synthesized to construct a comprehensive multi-residue drug database, including detailed information such as compound names, structures, accurate molecular weights, retention times (RTs), and fragment ions (Table S1).
3.2. Optimization of sample pretreatment
The effectiveness of extraction procedures in isolating veterinary drug residues from complex matrices is crucial, particularly due to their typically low levels of concentration. Evaluation of extraction efficiency was conducted considering various factors such as the choice of extraction solvent, incorporation of sodium sulfate, acidification, adsorption, and filtration (Masia, Suarez-Varela, Llopis-Gonzalez, & Pico, 2016).
3.2.1. Establishment of extract method
The solubility range and chemical inertness make ACN, MeOH, and ethyl acetate popular choices for extracting a broad spectrum of polarity in veterinary drug residues. Specifically, ACN is preferred due to its minimal fat solubility, efficient deproteinization, and miscibility with water, facilitating the extraction of polar analytes. The Na2-EDTA-McIlvaine buffer solution has been observed to enhance the extraction of tetracyclines and quinolones. The addition of methylene chloride optimizes the extraction by adjusting the solvent polarity. Furthermore, for alkaline compounds, an acidic medium may potentially improve their extraction by forming neutral molecules.
In this study, grass carp and crab were chosen as matrices to represent low-fat and high-fat aquatic products, respectively. Seven extraction methods with diverse solvent mixtures were evaluated, including buffer solutions, acetic acid, and methylene chloride. The detailed extraction procedures and efficiencies are shown in Table 1 and Fig. 2.
Table 1.
Seven methods for extracting multi-residue drugs from aquatic products.
| No. | Extraction procedures |
|---|---|
| A | 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-water-ACN (1:9, V/V) |
| B | 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-acetic acid-water-ACN (0.2:0.8:9, V/V/V) -2 mg/mL TMDP solution |
| C | Water-ACN (2:8; V/V) |
| D | Acetic acid - water-ACN (0.2:1.8:8, V/V/V) |
| E | 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-DCM-ACN (2:8; V/V) |
| F | 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-water-ACN (1:9, V/V)‑sodium sulfate (5 g)-acetic acid (100 μL) |
| G | 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-water-ACN (1:9, V/V)‑sodium sulfate (5 g)-acetic acid (100 μL) the residue is extracted by ethyl acetate after 0.1 mol/L Na2 EDTA-Mcllvaine buffer solution-ACN‑sodium sulfate-acetic acid extraction. |
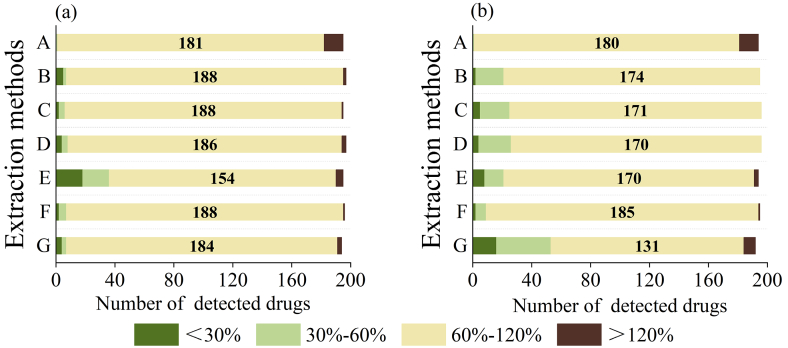
Fig. 2.
The distribution of veterinary drugs with varying recovery levels in the grass carp matrix (a) and the crab matrix (b). Extract procedure: A: B: C: D: E: F: G, corresponding to that described in Table 1.
The optimized extraction methods resulted in recoveries ranging from 60% to 120%, with method E being an exception. Six of the methods allowed for the detection of over 180 compounds in grass carp, showcasing a broad extraction efficiency. For high-fat crab samples, methods A and F exhibited a slight decrease in detectable compounds to 180 and 185, respectively. Furthermore, the recoveries obtained from methods A and F remained above 60%. These results suggest the excellent extraction efficiency of methods A and F. While method A offered slightly higher recovery rates, method F was able to detect more compounds, making it the preferable choice for multi-residue screening. As a result, method F, utilizing a 0.1 mol/L Na2-EDTA-McIlvaine buffer solution with water, ACN, sodium sulfate, and acetic acid (1:9 v/v), was selected for further optimization.
3.2.2. Optimization of Na2-EDTA buffer solution
The role of complexing agents like EDTA in enhancing the recovery of specific antibiotic residues, especially tetracyclines and quinolones, has been extensively documented (Kaufmann, Butcher, Maden, & Widmer, 2008). Preliminary experiments mirrored these observations, revealing low recovery rates for tetracycline analogs at a spiked concentration of 10 ng/g. This resulted in reduced sensitivity, thereby limiting detection capabilities. To address this issue, an EDTA buffer solution was integrated into the extraction protocol to assess its impact on recovery rates. Contrary to expectations, as illustrated in Fig. S1., the results demonstrated that the presence of the EDTA buffer did not improve recovery rates. In fact, the recovery of compounds was higher without the EDTA buffer, particularly within the crab matrix. Based on these findings, the use of an EDTA buffer was excluded from the sample pretreatment process in favor of conditions that yield higher recoveries for a wider range of compounds.
3.2.3. Optimization of sodium sulfate
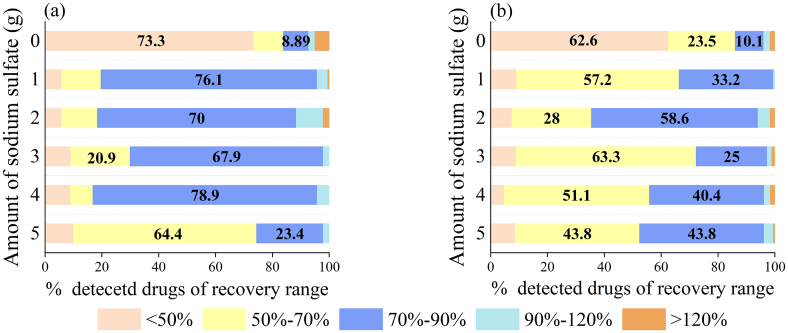
This study evaluated the impact of different amounts of sodium sulfate (Na2SO4) on the removal of extracted water and the enhancement of extraction efficiency. Sodium sulfate concentrations were varied from 0 to 5 g and added to spiked grass carp and crab samples at a constant concentration of 25 μg/kg, as shown in Fig. 3. In the grass carp matrix, the incremental addition of Na2SO4 showed no marked impact on the compound detection across a recovery range of 30–120%. The optimal separation was achieved with 4 g of Na2SO4, where the recovery rates were most consistent, falling within the 70–90% bracket. For crab matrix, which have a higher fat content, utilizing a smaller quantity of 2 g Na2SO4 proved to yield better recovery rates within the established initial range. Due to the high fat content, over-dehydrating the sample may lessen the extraction efficiency. However, extending the evaluation to a broader recovery range, the use of 4 g of Na2SO4 exhibited improved performance compared to 2 g, suggesting that a higher salt concentration may compensate for the reduced efficiency caused by the dehydration of the extract. Taking both matrices into account, the study found that 4 g of Na2SO4 struck an optimal balance between efficient extraction and phase separation. This amount had been confirmed as effective for both low-fat and high-fat aquatic product matrices.
Fig. 3.
Profile of veterinary drug quantities across different recovery ranges in the grass carp matrix(a) and the crab matrix(b).
3.2.4. Optimization of acidic medium
The impact of acetic acid on extraction efficiency was examined by comparing recovery rates with and without its presence during the sample preparation, as detailed in Fig. S2. In grass carp samples, the inclusion of acetic acid led to a roughly 10% increase in the recovery of various compounds, including zilpaterol, ractopamine, amantadine, chlordimeform, malachite green, and danofloxacin. This improvement is believed to stem from acetic acid aiding in the more effective analyte desorption from the sample matrix. On the other hand, the crab matrix displayed 169 compounds with recovery rates falling within the 50–90% range when acetic acid was included. Based on these findings, the addition of 100 μL of acetic acid into the extraction solvent proved to be optimal, boosting overall recovery rates for both low-fat and high-fat aquatic matrices.
3.2.5. Optimization of adsorbents
The cleanup step in complex aquatic matrix analysis can reduce interference and maintain the integrity of the mass spectrometer. This study compared the different efficacy of seven adsorbents. Through the utilization of 100 mg of each dispersed solid-phase extraction material, such as d-C18, d-PSA, d-Neutral Alumina, d-PEP, and two solid-phase extraction cartridges — PRiME HLB and EMR Lipid — alongside a commercial purification cartridge, Cleanert, their recovery rates and matrix effects were evaluated as depicted in Fig. 4. For the grass carp matrix, the detection rates of compounds across the tested adsorbents ranged between 185 and 191. In the context of minimizing matrix effects, d-C18, d-PSA, and EMR Lipid performed well, with over 85% of compounds exhibiting matrix effect values within the desirable range of −20% to +20%. Notably, the recovery rates for 92% of compounds utilizing d-C18 and d-PSA sorbents fell within 30–120%, slightly outperforming EMR Lipid.
Fig. 4.
The proportion profiles of veterinary drugs across varying ranges of recoveries (a) (c) and matrix effects (b) (d) under seven different cleanup treatments for both the grass carp and crab matrices.
In the crab matrix, the extract efficacy after sorbents cleanup followed the order of d-PSA > d-C18 = Cleanert > d-Neutral Alumina > d-PEP > PRiME HLB > EMR Lipid. However, when considering matrix effects, the ranking shifted to d-Neutral Alumina > d-PSA > d-C18 > d-PEP > Cleanert > EMR Lipid > PRiME HLB. Notably, d-PSA stood out as the top performer due to its dual advantages of efficient lipid removal and minimal impact on recovery rates, resulting in the least matrix effect among the tested sorbents.
Subsequent exploration delved into determining the optimal amount of d-PSA by assessing dosages ranging from 0 to 200 mg. The maximum number of detectable compounds was observed with 100 mg of d-PSA, where both the grass carp and crab matrices exhibited over 92% of compounds with recoveries between 50 and 120%, as illustrated in Fig. S3. Therefore, 100 mg of d-PSA was established as the optimal adsorbent choice for subsequent analyses based on the balance between purity and recovery efficiency.
3.2.6. Optimization of re-dissolved solution
Fine-tuning the re-dissolution step plays a pivotal role in augmenting the sensitivity of the analytical method. In this study, the supernatant obtained post d-PSA purification was concentrated under a gentle nitrogen stream to achieve a volume of 0.5 mL. Subsequently, it was diluted to a final volume of 1.5 mL using a carefully chosen reconstitution solution. The selection was predicated on its capacity to thoroughly dissolve the target analytes while ensuring that both the baseline and peak shapes remained unaffected during sample injection (Wang, Tian, et al., 2023).
The evaluation encompassed two prospective reconstitution solutions: ACN and ACN with 1% FA. Comparative analysis, as illustrated in Fig. S4., revealed a notable reduction in recovery when FA was incorporated into the reconstitution solution for both grass carp and crab matrices. Based on these findings, ACN emerged as the superior reconstitution solution due to its ability to dissolve analytes effectively while preserving chromatographic performance.
3.2.7. Optimization of filter
Filtering the reconstituted sample solution is essential for ensuring clarity and preventing instrument contamination prior to analysis. The study observed the occasional presence of flocculent suspensions after re-dissolving samples in ACN, likely originating from non-soluble extracts in the aquatic product matrix. While d-PSA is effective at impurity removal, filtration is required to eliminate any remaining particulates. The effect of four different types of 0.22 μm disposable microporous filters — polyethersulfone (PES), Nylon, hydrophilic polytetrafluoroethylene (H-PTFE), and polytetrafluoroethylene (PTFE) — were examined on the recovery of veterinary drug residues.
The findings indicated that the H-PTFE filter exhibited the most reliable performance, successfully mitigating analyte adsorption and achieving the highest recovery rates within the desirable 60–120% range for both grass carp and crab matrices (Fig. S5). In comparison, the other filters demonstrated slightly inferior performance in maintaining compound recovery rates within the same range. Based on these findings, the H-PTFE filter was determined to be the preferred option for filtering reconstituted sample solutions prior to instrumental analysis.
3.3. Method validation
3.3.1. Matrix effect
The matrix effect played a significant role in method accuracy and was assessed in two representative aquatic products with differing fat contents: grass carp and crab (Hoff et al., 2015). Post-optimization, standard preparations were diluted to 10 ng/mL using the optimized blank solutions, and the resulting matrix-matched standard signals were compared to the solvent standards. As outlined in Table S3, >90% of the compounds in grass carp exhibited matrix inhibitory effects between −30% and + 30%. Crabs, being a high-fat matrix, showed that 85% of compounds exhibited similar inhibitory effects, with approximately 3% of compounds displaying more pronounced inhibitory effects (>50%). This suppression could be attributed to ionization competition between the high-fat content and the target analytes in the ionization source. Despite some compounds exhibiting significant matrix effects, the method proved effective for simultaneous detection and semi-quantitative analysis across fish products with varying fat contents.
3.3.2. SDL
SDL, a critical parameter for method assessment, reflects the sensitivity and applicability across various matrices. As shown in Table S2 and Fig. S6, the method effectively identified a wide range of compounds across six diverse aquatic matrices: grass carp, crab, bullfrog, scallop, shrimp, and yellow croaker. Starting from the lowest spiked level of 0.5 μg/kg, the method screened 142 to 153 compounds across these matrices. The detection rate increased with increasing spiked concentration: at 1 μg/kg, 128 and 167 compounds were detected; at 2 μg/kg, 151 to 175 compounds were identified; at 5 μg/kg, 172 to 176 compounds were captured; and at 10 μg/kg, 177 to 181 compounds were detected across the matrices. The detection scope expanded further at 50 μg/kg, with 192 compounds detectable in the matrices, demonstrating the method's broad applicability and robust performance for screening diverse aquatic products. The compounds posing challenges for detection with satisfactory recoveries and SDL in some matrices (with obvious recoveries decrease or not detectable in some matrices at SDL) belongs to β-agonists class, steroid hormones class, and macrolides class (Fig. S7).
3.3.3. Accuracy and precision
There are numerous potential factors influencing the differences in detection rates among different matrices, such as the concentration, solubility in extract solvent, and polarity of the compounds to be detected, as well as the fat and protein content in different matrices. Hence, the accuracy and precision of this method were validated by assessing spiked recovery rates and the relative standard deviation (RSD) across a range of concentrations. Utilizing a refined sample pretreatment technique, standard solutions were introduced into blank aquatic samples with varying levels of fat content (low and high) at concentrations ranging from 1 to 10 μg/kg. The recovery and precision data for the 192 target compounds are systematically presented across three concentration levels in Tables S3 and S4.
In the grass carp samples, the recovery rates for 115, 134, and 164 compounds were found to be within the acceptable range of 60–120% at spiked levels of 1, 5, and 10 μg/kg, respectively, as depicted in Table S3. However, the crab samples exhibited lower recovery consistency, with only 53, 118, and 146 compounds falling within a 50–120% recovery range at 1, 5, and 10 μg/kg, as detailed in Table S4. This variability in recovery rates for crab samples is likely due to the complex extraction challenges posed by the high-fat content in these samples.
Precision, denoted by RSD, was calculated from six replicates for each target compound at the specified concentrations. Within the grass carp matrix, >80% of compounds exhibited RSD values ranging from 0.15% to 19.59%. Conversely, according to Table S4, the results from the crab matrix indicated that over 74.4% of compounds had RSD values ranging from 0.2% to 19.84%, demonstrating the stability of the method.
These findings confirm the method suitability for the semi-quantitative analysis of the majority of compounds selected for this study. The method demonstrates a high level of reliability, even at lower concentration levels, making it a valuable tool for routine screening in diverse aquatic matrices.
Additionally, an intriguing phenomenon was observed. As depicted in Fig. S8, at the spiked concentration of 10 μg/kg, the recovery distribution of six categories including tetracyclines, amide alcohols, nitrofurans, triphenylmethane compounds, sulfonamides, sedatives, and other pharmaceuticals stably remained at 70% or above in aquatic product matrices with varying fat contents (grass carp and crab). However, when applying this procedure to screen for the following categories of multi-residue drugs in the high-fat content matrices such as β-agonists, antiviral drugs, steroid hormones, nitroimidazoles, benzimidazoles and their metabolites, fluoroquinolone, macrolides, avermectins, and other pharmaceuticals, the recovery rates tended to fall within a range of 70% to 50%, or even lower compared to low-fat matrices.
3.4. Application to real samples
To evaluate the method practicality for routine surveillance of 192 veterinary drugs, 30 samples of diverse aquatic products from local markets, including blackfish, grouper, white shrimp, Chinese mitten crab, and scallops, were obtained. The analysis was conducted using TraceFinder EFS3.0 software, enabling a semi-quantitative assessment of the detected analytes. Quantitation of the identified compounds was performed by comparing the peak areas of the samples with those of corresponding spiked standards. The comprehensive screening data is compiled in Table S5.
The results revealed the widespread presence of veterinary drugs, particularly hormone-related substances such as testosterone propionate, megestrol, clobetasol 17-propionate, methylprednisolone, trenbolone, and nandrolone phenylpropionate, predominantly in Chinese mitten crab and scallops. Additionally, amantadine was confirmed in 29 samples, reflecting its extensive use as an antiviral drug in aquaculture (Zhu, Miao, Zhang, Wang, & Wang, 2021). However, to address potential resistance issues associated with these compounds in human treatments, the use of amantadine in animal husbandry has been prohibited since 2005, including in countries such as the United States and China (Tsuruoka et al., 2017).
Interestingly, diazepam, which is prohibited in China for use in animal-derived products, was detected in 23 samples, indicating potential unauthorized usage or environmental accumulation within the aquatic food chain (Corcoran, Winter, & Tyler, 2010). Additionally, antibiotics, specifically sulfonamides and fluoroquinolones, were detected with sulfamethoxazole in 24 samples and enrofloxacin in 14 samples. In one instance, a blackfish sample exhibited enrofloxacin levels at 114.77 μg/kg, surpassing the maximum residue limits (MRLs) set by Chinese food safety standards, underscoring the necessity for stringent monitoring and regulation enforcement.
4. Conclusions
This study established a method for screening and semi-quantifying 192 veterinary drug residues in aquatic products, primarily utilizing QuEChERS pretreatment coupled with HPLC-HRMS. This method accurately measured parent and fragment ions in MS ddMS2 full scan mode, fulfilling qualitative screening requirements and enabling the detection of multiple drugs used in livestock and poultry, showcasing its broad applicability in veterinary pharmaceuticals. The key advantages included simple pretreatment, effective purification, high sensitivity, and excellent stability, enhancing its high detection efficacy and precision. Its practicality was affirmed through successful analysis of real samples, uncovering the presence of drug residues ranging from hormone-related substances to antibiotics, some of which exceeded legal limits. In addition, the expandable database offer flexibility and allow inclusion of accurate mass date for newly identified compounds, facilitating rapid adaptation to evolving screening demands. This approach serves as a vital step towards comprehensive monitoring of multi-residue veterinary drugs in aquaculture, ensuring both food safety and public health.
CRediT authorship contribution statement
Guangxin Yang: Writing – review & editing, Methodology, Funding acquisition, Data curation, Conceptualization. Junyu Zhang: Writing – original draft, Methodology, Investigation. Yunyu Tang: Validation, Formal analysis. Cong Kong: Writing – review & editing, Supervision, Resources, Funding acquisition, Conceptualization. Siman Li: Validation. Shouying Wang: Validation, Investigation. Shuhai Ding: Validation, Conceptualization. Lin Gu: Writing – review & editing, Supervision. Xiaosheng Shen: Validation, Resources. Aka Alla Martin: Visualization, Formal analysis. Hai Chi: Writing – review & editing, Resources, Funding acquisition.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
The authors would like to thank the financial support from the Science and Technology Commission of Shanghai Municipality (22142201400), National Natural Science Foundation of China (31701698), Central Public-interest Scientific Institution Basal Research Fund, CAFS (2020TD72), the Special Research Fund for the National Non-profit Institutes (East China Sea Fisheries Research Institute, Chinese Academy of Fishery Sciences, China) (2016T09).
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.fochx.2024.101504.
Contributor Information
Cong Kong, Email: kongcong@gmail.com, kongc@ecsf.ac.cn.
Hai Chi, Email: chih@ecsf.ac.cn.
Appendix A. Supplementary data
Fig. S1-S8, Table S1-S5 about the optimization of sample preparation, the screenig result of aquatic products, the database, the verified SDL, matrix effect, and recoveries of each target in different matrices, are shown in the supplementary data.
Data availability
Data will be made available on request.
References
- Anastassiades M., Lehotay S.J., Štajnbaher D., Schenck F.J. Fast and easy multiresidue method employing acetonitrile extraction/partitioning and “dispersive solid-phase extraction” for the determination of pesticide residues in produce. Journal of AOAC International. 2019;86(2):412–431. [PubMed] [Google Scholar]
- Casado J., Brigden K., Santillo D., Johnston P. Screening of pesticides and veterinary drugs in small streams in the European Union by liquid chromatography high resolution mass spectrometry. Science of the Total Environment. 2019;670:1204–1225. doi: 10.1016/j.scitotenv.2019.03.207. [DOI] [PubMed] [Google Scholar]
- Castilla-Fernández D., Moreno-González D., Bouza M., Saez-Gómez A., Ballesteros E., García-Reyes J.F., et al. Assessment of a specific sample cleanup for the multiresidue determination of veterinary drugs and pesticides in salmon using liquid chromatography/tandem mass spectrometry. Food Control. 2021;130 [Google Scholar]
- Corcoran J., Winter M.J., Tyler C.R. Pharmaceuticals in the aquatic environment: A critical review of the evidence for health effects in fish. Critical Reviews in Toxicology. 2010;40(4):287–304. doi: 10.3109/10408440903373590. [DOI] [PubMed] [Google Scholar]
- De Paepe E., Wauters J., Van Der Borght M., Claes J., Huysman S., Croubels S., et al. Ultra-high-performance liquid chromatography coupled to quadrupole orbitrap high-resolution mass spectrometry for multi-residue screening of pesticides, (veterinary) drugs and mycotoxins in edible insects. Food Chemistry. 2019;293:187–196. doi: 10.1016/j.foodchem.2019.04.082. [DOI] [PubMed] [Google Scholar]
- Gonzalez-Curbelo M.A., Varela-Martinez D.A., Riano-Herrera D.A. Pesticide-residue analysis in soils by the QuEChERS method: A review. Molecules. 2022;27(13) doi: 10.3390/molecules27134323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han F., Chen Q.-P., Yang G., Zhai W., Kouadio Fodjo E., Tian L.-L., et al. Determination of 18 β-agonists in aquatic products using high-performance liquid chromatography tandem mass spectrometry. ACS Food Science & Technology. 2022;2(2):337–346. [Google Scholar]
- Hoff R.B., Rubensam G., Jank L., Barreto F., Peralba Mdo C., Pizzolato T.M., et al. Analytical quality assurance in veterinary drug residue analysis methods: Matrix effects determination and monitoring for sulfonamides analysis. Talanta. 2015;132:443–450. doi: 10.1016/j.talanta.2014.08.046. [DOI] [PubMed] [Google Scholar]
- Jung H.N., Park D.H., Yoo K.H., Cho H.J., Shim J.H., Shin H.C., et al. Simultaneous quantification of 12 veterinary drug residues in fishery products using liquid chromatography-tandem mass spectrometry. Food Chemistry. 2021;348 doi: 10.1016/j.foodchem.2021.129105. [DOI] [PubMed] [Google Scholar]
- Kaufmann A., Butcher P., Maden K., Widmer M. Quantitative multiresidue method for about 100 veterinary drugs in different meat matrices by sub 2-microm particulate high-performance liquid chromatography coupled to time of flight mass spectrometry. Journal of Chromatography. A. 2008;1194(1):66–79. doi: 10.1016/j.chroma.2008.03.089. [DOI] [PubMed] [Google Scholar]
- Kong C., Wang Y., Huang Y., Yu H. Multiclass screening of >200 pharmaceutical and other residues in aquatic foods by ultrahigh-performance liquid chromatography-quadrupole-Orbitrap mass spectrometry. Analytical and Bioanalytical Chemistry. 2018;410(22):5545–5553. doi: 10.1007/s00216-018-1124-5. [DOI] [PubMed] [Google Scholar]
- Lazartigues A., Wiest L., Baudot R., Thomas M., Feidt C., Cren-Olive C. Multiresidue method to quantify pesticides in fish muscle by QuEChERS-based extraction and LC-MS/MS. Analytical and Bioanalytical Chemistry. 2011;400(7):2185–2193. doi: 10.1007/s00216-011-4945-z. [DOI] [PubMed] [Google Scholar]
- Lopes R.P., Reyes R.C., Romero-Gonzalez R., Vidal J.L., Frenich A.G. Multiresidue determination of veterinary drugs in aquaculture fish samples by ultra high performance liquid chromatography coupled to tandem mass spectrometry. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences. 2012;895-896:39–47. doi: 10.1016/j.jchromb.2012.03.011. [DOI] [PubMed] [Google Scholar]
- Magalhaes D., Freitas A., Sofia Vila Pouca A., Barbosa J., Ramos F. The use of ultra-high-pressure-liquid-chromatography tandem time-of-flight mass spectrometry as a confirmatory method in drug residue analysis: Application to the determination of antibiotics in piglet liver. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences. 2020;1153 doi: 10.1016/j.jchromb.2020.122264. [DOI] [PubMed] [Google Scholar]
- Masia A., Suarez-Varela M.M., Llopis-Gonzalez A., Pico Y. Determination of pesticides and veterinary drug residues in food by liquid chromatography-mass spectrometry: A review. Analytica Chimica Acta. 2016;936:40–61. doi: 10.1016/j.aca.2016.07.023. [DOI] [PubMed] [Google Scholar]
- Mehl A., Schmidt L.J., Schmidt L., Morlock G.E. High-throughput planar solid-phase extraction coupled to orbitrap high-resolution mass spectrometry via the autoTLC-MS interface for screening of 66 multi-class antibiotic residues in food of animal origin. Food Chemistry. 2021;351 doi: 10.1016/j.foodchem.2021.129211. [DOI] [PubMed] [Google Scholar]
- Melekhin A.O., Tolmacheva V.V., Goncharov N.O., Apyari V.V., Dmitrienko S.G., Shubina E.G., et al. Multi-class, multi-residue determination of 132 veterinary drugs in milk by magnetic solid-phase extraction based on magnetic hypercrosslinked polystyrene prior to their determination by high-performance liquid chromatography-tandem mass spectrometry. Food Chemistry. 2022;387 doi: 10.1016/j.foodchem.2022.132866. [DOI] [PubMed] [Google Scholar]
- Monteiro S.H., Lehotay S.J., Sapozhnikova Y., Ninga E., Moura Andrade G.C.R., Lightfield A.R. Validation of the QuEChERSER mega-method for the analysis of pesticides, veterinary drugs, and environmental contaminants in tilapia (Oreochromis Niloticus) Food Additives & Contaminants. Part A, Chemistry, Analysis, Control, Exposure & Risk Assessment. 2022;39(4):699–709. doi: 10.1080/19440049.2021.2020911. [DOI] [PubMed] [Google Scholar]
- Park H., Choi S.Y., Kang H.-S., Kwon N., j. Multi residue determination of 96 veterinary drug residues in domestic livestock and fishery products in South Korea. Aquaculture. 2022;553 [Google Scholar]
- Shahsavari S., Shariatifar N., Arabameri M., Mahmoudi R., Moazzen M., Ghajarbeygi P. Analysis of polychlorinated biphenyls in cream and ice cream using modified QuEChERS extraction and GC-QqQ-MS/MS method: A risk assessment study. International Journal of Dairy Technology. 2022;75(2):448–459. [Google Scholar]
- Shin D., Kang H.-S., Jeong J., Kim J., Choe W.J., Lee K.S., et al. Multi-residue determination of veterinary drugs in fishery products using liquid chromatography-tandem mass spectrometry. Food Analytical Methods. 2018;11(6):1815–1831. [Google Scholar]
- Syrgabek Y., Alimzhanova M. Modern analytical methods for the analysis of pesticides in grapes: A review. Foods. 2022;11(11) doi: 10.3390/foods11111623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuruoka Y., Nakajima T., Kanda M., Hayashi H., Matsushima Y., Yoshikawa S., et al. Simultaneous determination of amantadine, rimantadine, and memantine in processed products, chicken tissues, and eggs by liquid chromatography with tandem mass spectrometry. Journal of Chromatography B. 2017;1044-1045:142–148. doi: 10.1016/j.jchromb.2017.01.014. [DOI] [PubMed] [Google Scholar]
- Uchida K., Konishi Y., Harada K., Okihashi M., Yamaguchi T., Do M.H., et al. Monitoring of antibiotic residues in aquatic products in urban and rural areas of Vietnam. Journal of Agricultural and Food Chemistry. 2016;64(31):6133–6138. doi: 10.1021/acs.jafc.6b00091. [DOI] [PubMed] [Google Scholar]
- Wang C., Chen M., Hu Q., Bai H., Wang C., Ma Q. Non-lethal microsampling and rapid identification of multi-residue veterinary drugs in aquacultured fish by direct analysis in real time coupled with quadrupole-Orbitrap high-resolution mass spectrometry. Microchemical Journal. 2021;160 [Google Scholar]
- Wang H., Tian H., Ai L.F., Liang S.X. Screening and quantification of 146 veterinary drug residues in beef and chicken using QuEChERS combined with high performance liquid chromatography-quadrupole orbitrap mass spectrometry. Food Chemistry. 2023;408 doi: 10.1016/j.foodchem.2022.135207. [DOI] [PubMed] [Google Scholar]
- Wang J., Zhao W., Guo W., Li Y., Jiang R., Li H., et al. Simultaneous screening and analysis of 155 veterinary drugs in livestock foods using ultra-high performance liquid chromatography tandem quadrupole linear-ion-trap mass spectrometry. Food Chemistry. 2022;393 doi: 10.1016/j.foodchem.2022.133260. [DOI] [PubMed] [Google Scholar]
- Wang S., Yang G., Tang Y., Wang Y., Shen X., Si W., et al. Multi-residue screening of pesticides in aquatic products using high-performance liquid chromatography-tandem high-resolution mass spectrometry. Foods. 2023;12(6) doi: 10.3390/foods12061131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu X., Xu F., Zhang R., Liu H., Sun A., Zhang L., et al. Simultaneous determination and dietary intake risk assessment of 60 herbicide residues in aquatic products. Science of the Total Environment. 2023;883 doi: 10.1016/j.scitotenv.2023.163633. [DOI] [PubMed] [Google Scholar]
- Zhang S., Guo Y., Yan Z., Sun X., Zhang X. A selective biomarker for confirming nitrofurazone residues in crab and shrimp using ultra-performance liquid chromatography-tandem mass spectrometry. Analytical and Bioanalytical Chemistry. 2015;407(30):8971–8977. doi: 10.1007/s00216-015-9058-7. [DOI] [PubMed] [Google Scholar]
- Zhang Y., Bleeker A., Liu J. Nutrient discharge from China’s aquaculture industry and associated environmental impacts. Environmental Research Letters. 2015;10(4) [Google Scholar]
- Zhang Y., Liu X., Li X., Zhang J., Cao Y., Su M., et al. Rapid screening and quantification of multi-class multi-residue veterinary drugs in royal jelly by ultra performance liquid chromatography coupled to quadrupole time-of-flight mass spectrometry. Food Control. 2016;60:667–676. [Google Scholar]
- Zhao F., Gao X., Tang Z., Luo X., Wu M., Xu J., et al. Development of a simple multi-residue determination method of 80 veterinary drugs in Oplegnathus punctatus by liquid chromatography coupled to quadrupole Orbitrap mass spectrometry. Journal of Chromatography. B, Analytical Technologies in the Biomedical and Life Sciences. 2017;1065-1066:20–28. doi: 10.1016/j.jchromb.2017.09.013. [DOI] [PubMed] [Google Scholar]
- Zhu D., Mu Y., Li Q., Pang X., Wang X., Liu Y., et al. A novel method for artificial antigen synthesis and preparation of a polyclonal antibody for the sensitive determination of leucomalachite green in fish samples by enzyme-linked immunoassay. Analytical Methods. 2016;8(32):6236–6243. [Google Scholar]
- Zhu S., Miao B., Zhang Y.Z., Wang D.S., Wang G.X. Amantadine, a promising therapeutic agent against viral encephalopathy and retinopathy. Journal of Fish Diseases. 2021;45(3):451–459. doi: 10.1111/jfd.13574. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Fig. S1-S8, Table S1-S5 about the optimization of sample preparation, the screenig result of aquatic products, the database, the verified SDL, matrix effect, and recoveries of each target in different matrices, are shown in the supplementary data.
Data Availability Statement
Data will be made available on request.