Abstract
The identification of novel tumor‐specific proteins or antigens is of great importance for diagnostic and therapeutic applications in pancreatic cancer. Using oligonucleotide microarrays, we identified a broad spectrum of differentially expressed pancreatic cancer‐related genes. Of these, we selected an overexpressed expressed sequence taq and cloned a 721‐bp full‐length cDNA with an open reading frame of 196 amino acids. This novel gene was localized on the Homo sapiens 16p13.3 chromosomal locus, and its nucleotide sequence matched the Homo sapiens similar to common salivary protein 1 (LOC124220). We named the gene pancreatic adenocarcinoma up‐regulated factor. The pancreatic adenocarcinoma up‐regulated factor was secreted into the culture medium of pancreatic adenocarcinoma up‐regulated factor‐overexpressing Chinese hamster ovary cells, had an apparent molecular mass of ~25 kDa, and was N‐glycosylated. The induction of pancreatic adenocarcinoma up‐regulated factor in Chinese hamster ovary cells increased cell proliferation, migration, and invasion ability in vitro. Subcutaneous injection of mice with Chinese hamster ovary/pancreatic adenocarcinoma up‐regulated factor cells resulted in 3.8‐fold greater tumor sizes compared to Chinese hamster ovary/mock cells. Reverse transcription–polymerase chain reaction and western blotting with antirecombinant human pancreatic adenocarcinoma up‐regulated factor antibodies confirmed that pancreatic adenocarcinoma up‐regulated factor was highly expressed in six of eight pancreatic cancer cell lines. Immunohistochemical staining of human pancreatic cancer tissues also showed pancreatic adenocarcinoma up‐regulated factor overexpression in the cytoplasm of cancer cells. Transfection with pancreatic adenocarcinoma up‐regulated factor‐specific small‐interfering RNA reduced cancer cell migration and invasion in vitro. Treatment with antirecombinant human pancreatic adenocarcinoma up‐regulated factor in vitro and in vivo reduced proliferation, migration, invasion, and tumorigenic ability. Collectively, our results suggest that pancreatic adenocarcinoma up‐regulated factor is a novel secretory protein involved in pancreatic cancer progression and might be a potential target for the treatment of pancreatic cancer. (Cancer Sci 2009; 100: 828–836)
Pancreatic cancer has an extremely poor prognosis with a 5‐year survival rate of less than 5%.( 1 ) The identification of novel tumor‐specific antigens and signal pathways could improve prognosis and help overcome obstacles such as late diagnoses, aggressive tumors, and resistance to existing therapeutic regimens.( 2 , 3 )
Over the past decade, several pancreatic cancer‐related genes have been identified. These genes include tumor suppressor genes (cvclin‐dependent kinase 2A [p16/CDKN2A], Tumor protein p53 [Tp53], and Mothers against decapentaplegic homolog 4 [SMAD4]), oncogenes (Kirsten rat sarcoma viral oncogene 2 [KRAS2]), and genome‐maintenance genes (Breast cancer susceptibility protein 2 [BRCA2]),( 4 , 5 , 6 , 7 , 8 ) and in a recent study using oligonucleotide microarrays, we found that a novel tumor‐associated gene, Cancer‐Up regulated Gene 2 (CUG2), was up‐regulated.( 9 ) The identification of these cancer‐related genes has increased our understanding of pancreatic cancer development and led to the discovery of several genes that may be useful as diagnostic biomarkers or as therapeutic molecular targets.
Studies of gene expression patterns using serial analyses, cDNA arrays, and oligonucleotide arrays have provided useful information concerning the molecular characteristics of pancreatic cancers that can be used to distinguish closely related cancer subtypes.( 10 , 11 , 12 , 13 ) Using oligonucleotide microarrays, we identified a novel gene, named pancreatic adenocarcinoma up‐regulated factor (PAUF), which was highly expressed in human pancreatic cancer. Results of a database search indicated that PAUF encoded a hypothetical protein that matched the Homo sapiens similar to common salivary protein 1 (LOC124220; synonyms: HRPE773, PRO1567) gene. This homology is consistent with the functional nature of both the pancreas and salivary glands; both are secretory organs and consist of morphologically and functionally similar secretory units.( 14 ) Moreover, a recent sequence alignment study predicted that HPRE773 was a putative human ortholog of the mouse Demilune cell and parotid protein (Dcpp) gene and the rat Common salivary protein 1 (CSP1) gene.( 15 )
In this study, we further characterized the hypothetical protein PAUF. The effects of PAUF in vitro and in vivo were determined by increasing or decreasing its expression. Our findings suggest that PAUF plays an important role in cancer progression and may have the potential to become a novel diagnostic and therapeutic target in pancreatic cancer.
Materials and Methods
Microarray expression analysis. Global gene expression profiles from normal pancreas (n = 24) and from pancreatic adenocarcinomas (n = 32) were examined using a HG‐U133 GeneChip set (Affymetrix, Santa Clara, CA, USA) in a collaboration with Yonsei University College of Medicine (Seoul, Korea), Gene Logic Inc. (Gaithersburg, MD, USA), and LG Life Sciences Ltd. (Daejeon, Korea). Expression values of all gene fragments were normalized and analyzed using Microarray Suite 5.0 (Affymetrix). Outliers were detected by principal component analyses using the MatLab program (The MathWorks Inc., Natick, MA, USA) and were excluded from the clustering analysis. Confidence limits were set at a P‐value < 0.05 for each differentially expressed gene fragment. Unigene (National Center for Biological Information, Bethesda, MD, USA) was used to identify expressed sequence taq (EST) clusters that matched the selected Affymetrix fragments.
The Ethical Committee and Institutional Review Board of Yonsei University College of Medicine approved the protocol of tissue acquisition from surgical specimens, and written informed consent was obtained from each patient.
Cell lines. Eight pancreatic cancer cell lines (AsPC‐1, BxPC‐3, Capan‐1, Capan‐2, CFPAC‐1, HPAC, MIAPaCa‐2, and PANC‐1), Chinese hamster ovary (CHO) and CHO/dihydrofolate reductase (dhFr)‐cells, and NIH/3T3 cells were purchased from American Type Culture Collection (Manassas, VA, USA). Peripheral blood mononuclear cells (PBMC), phytohemagglutinin (PHA)‐stimulated PBMC, and bone marrow stromal cells (BMSC) were kindly provided by Dr H.S. Kim (Department of Laboratory Medicine and Cell Therapy Center, Yonsei University College of Medicine). All cells were grown in conditioned medium and maintained in an atmosphere of 5% CO2 at 37°C.
Quantitative real‐time and reverse transcription (RT) polymerase chain reaction (PCR) and northern blot analysis. Quantitative real‐time RT‐PCR was performed on the HG‐U133 GeneChip set using total RNA from each sample and Taqman EZ RT‐PCR Core Reagents (Applied Biosystems, Foster City, CA, USA) according to the manufacturer's protocol. PAUF primers, designed from the sequence information file of Affymetrix fragments, were used in the assay. C‐terminal binding protein 1 (CTBP; GenBank accession no. AF091555) was used as a reference gene.( 9 ) For northern blot analyses, Human Multiple Tissue Northern blots (Clontech, Palo Alto, CA, USA) were hybridized with a radioactively labeled probe. The EST sequence was radioactively labeled by the random primer method and used as the probe for hybridization. The expression of PAUF in pancreatic cancer cells was examined using RT‐PCR. The PCR conditions were 95°C for 30 s, 55°C for 30 s, and 72°C for 30 s, for a total of 35 cycles. Beta‐actin (ACTB) was used as a reference gene. Primer sequences are reported in Supporting Table S1.
Cloning and sequencing of cDNA. Full‐length cDNA was obtained using the GeneTrapper cDNA Positive Selection System (Invitrogen, Carlsbad, CA, USA). A gene‐specific oligonucleotide was selected based on the Affymetrix fragment sequence. After biotinylation, the oligonucleotide was incubated with a single‐stranded human placenta library (Origene Technologies Inc., Rockville, MD, USA). The cDNA of the positive clone was eluted and subjected to DNA sequencing. The nucleotide sequence was defined as PAUF (GenBank accession no. EF067313).
Construction of the expression vector. For mammalian expression, the predicted coding region of PAUF was inserted into pcDNA3.1(+)‐Myc/His (Invitrogen) using the BamHI and XbaI sites, producing a pcDNA3.1(+)‐PAUF‐Myc/His plasmid. For production of recombinant human PAUF (rhPAUF), pcDNA3.1(+)‐PAUF‐Fc was constructed by inserting the coding region of PAUF, a thrombin cleavage sequence, and an constant region of immunoglobulin (Fc) sequence along with the gene for dhFr into the plasmid pcDNA3.1(+) (Invitrogen).
Establishment of stable cell lines. To obtain transformants that would stably express PAUF, the pcDNA3.1(+)‐PAUF‐Myc/His plasmid was transfected into CHO cells using LipofectaminePLUS reagent (Invitrogen). Stably transfected cells were selected by G418. For production of rhPAUF, pcDNA3.1(+)‐PAUF‐Fc was transfected into CHO/dhFr‐cells. Clones that expressed rhPAUF were selected with G418 and were adapted further by the stepwise increase of methotrexate (Sigma‐Aldrich, St. Louis, MO, USA).
Preparation and purification of antirecombinant human PAUF (antirhPAUF) polyclonal antibodies (pAb). The rhPAUF was purified from a medium of PAUF‐Fc‐transfected CHO/dhFr‐cells. The medium was purified through a 10‐mL protein A‐agarose column (Pierce, Rockford, IL, USA) and further purified with size‐exclusion chromatography using sepharose 200 HR resin (Amersham Biosciences, Buckinghamshire, UK). The C‐terminal Fc region was cleaved from PAUF‐Fc using the Thrombin Cleavage Capture Kit (Novagen, Madison, WI, USA) and was removed using ImmunoPure Immobilized Protein A (Pierce). Two New Zealand white female rabbits were immunized with purified rhPAUF. Freund's complete adjuvant (Sigma‐Aldrich) was used for the primary immunization. The antiserum was subsequently purified using an ImmunoPure (A Plus) IgG Purification Kit (Pierce). A second purification using immobilized recombinant human Fc (AminoLink Plus Immobilization Kit; Pierce) was performed to remove residual anti‐Fc activity from the antirhPAUF pAb. We confirmed that this antibody could specifically recognize the purified rhPAUF protein and exogenously introduced his‐tagged PAUF protein. This antibody was used for western blotting, immunohistochemical staining, and in vivo study is described below.
Down‐regulation of PAUF by transient transfection of small‐interfering RNA (siRNA) targeting for the PAUF gene. The three sets of 25‐nucleotide stealth RNAi targeting LOC124220 (si#1, si#2, and si#3) were customarily synthesized by Invitrogen, and stealth RNAi duplexes whose guanine‐cytosine (GC) content was similar to that of each duplex siRNA from Invitrogen were used as negative control. Sequences of three synthesized oligonucleotides are reported in Supporting Table S1. Stealth RNAi were transfected into CFPAC‐1 cells using Lipectamine™ RNAi Max transfection agent (Invitrogen) according to the manufacturer's protocol. At 48 h post‐transfecion, cells were harvested and subjected to total RNA extraction and a migration and invasion assay. Secreted protein was prepared from culture medium at 48 h post‐transfection.
Protein extraction and western blot analysis. Cell lysates were extracted in lysis buffer containing 50 mM Tris, 150 mM NaCl, 25 mM β‐glycerophosphate, 25 mM NaF, 0.5 M ethylenediaminetetraacetic acid (EDTA), 1% NP‐40, 0.1 mM phenylmethylsulfonyl fluoride (PMSF), and 1% protease inhibitor cocktail (Sigma‐Aldrich). For secretary protein preparation, the culture medium was centrifuged, and cellular components were discarded. The culture medium was concentrated using microcon centrifugal filters (10 kDa cutoff; Amicon, Houston, TX, USA). Cell lysates and media were separated by sodium dodecylsulfate–polyacrylamide gel electrophoresis (SDS‐PAGE) and transferred to polyvinylidene fluoride (PVDF) membrane (Millipore, Billerica, MA, USA). The membranes were blocked in 5% (w/v) non‐fat dry milk and probed with either the anti‐His antibody (1:1000; Santa Cruz Biotechnology, Santa Cruz, CA, USA) or antirhPAUF antibodies. The immunoreactive proteins were visualized using the West Pico Chemiluminescent Substrate (Pierce).
Immunohistochemical staining. Six cancer and adjacent non‐cancerous tissues were obtained by surgical resection. Paraformaldehyde‐fixed, paraffin‐embedded tissue sections were deparaffinized in xylene and rehydrated in a graded ethanol series. Endogenous peroxidase activity was blocked in 0.3% (v/v) hydrogen peroxide. Microwave antigen retrieval was performed in citrate buffer (0.01 M, pH 6.0). To prevent non‐specific staining, the sections were preincubated with 10% (v/v) normal donkey serum. The blocked sections were then incubated with antirhPAUF pAb (1:500), followed by incubation with biotinylated Link Universal and streptavidin–horseradish peroxidase (HRP) conjugate (DakoCytomation, Ft. Collins, CO, USA). The sections were stained with chromogen and counterstained with Harris hematoxylin solution (Sigma‐Aldrich).
In vitro glycosidase treatment. For protein deglycosylation, the protein was denatured and reduced by boiling for 10 min in 0.5% (w/v) SDS/1% (v/v) β‐mercaptoethanol. The secretion product was adjusted to a final concentration of 0.1 mg/mL in 50 mM sodium phosphate (pH 7.5), 1% (w/v) NP‐40, 0.3% (w/v) SDS, and 0.6% (v/v) β‐mercaptoethanol and was digested for 1 h at 37°C with 400 units of Peptide‐N‐glycosidase F (PNGase F) (New England Biolabs, Ipswich, MA, USA). Controls were treated identically as experimental samples but without the addition of enzymes. For deglycosylation of endogenous PAUF, CFPAC‐1 cells were seeded at 100‐mm dish and cultured for 24 h, and then medium was changed with fresh medium containing 2 µg/mL of tunicamycin (Sigma‐Aldrich) and cultured for additional 24 h. Controls were treated with the same volume of dimethyl sulfoxide. Proteins were then precipitated by acetone treatment, resolubilized in lysis buffer, and analyzed by western blotting.
N‐terminal sequencing. Purified protein from the culture medium was resolved by SDS‐PAGE and transferred to PVDF membrane (Millipore). The membrane was stained with Ponceau‐S and dried. The stained band was then excised from the membrane for N‐terminal sequencing, which was carried out at the Korea Basic Science Institute on a Procise 492 cLC protein sequencer (Applied Biosystems).
Measurement of cell growth. Cells were seeded at 2 × 103 cells per well in 24‐well plates. Every 24 h, the cells were removed with trypsin/EDTA digestion, and the number of cells was counted. Each experiment was performed in triplicate at each time‐point.
Migration and invasion assay. Migration and invasion assays were performed in Transwell plates (CoStar, Bethesda, MD, USA) and a Boyden chamber (Neuro Probe, Gaithersburg, MD, USA), respectively. For the invasion assay, detached cells (1 × 105) were seeded in the upper chamber coated with Matrigel. The lower wells of the chamber were filled with standard media. After incubation at 37°C, the cells on the upper surface of the membrane were completely removed with a moist cotton swab. For both assays, the cells were fixed with methanol and stained. They were counted and photographed by microscopy at ×100 magnification. All assays were performed in triplicate.
In vivo tumorigenesis and inhibition of tumor growth by antirhPAUF pAb. The CHO/PAUF and CHO/Mock cells (5 × 106/150 µL of serum‐free culture medium) were injected subcutaneously into 6‐week‐old BALB/c nude female mice (five mice per cell type). Tumor formation was monitored twice a week by measuring the width and the length of the mass. To evaluate the PAUF inhibitory effect, CFPAC‐1 (5 × 106/150 µL of serum‐free culture medium) was injected into 6‐week‐old BALB/c nude female mice, and when the tumor reached a size of 125–150 mm3, the mice were randomized into two groups: the antirhPAUF pAb‐treated group and the control group (six mice in each group). The antirhPAUF pAb‐treated group was injected with 500 µg of antirhPAUF pAb via the tail vein twice a week for 3 weeks. The control group was injected with 500 µg of preimmune serum rabbit IgG. Serial tumor volumes were measured twice a week and calculated by the formula V (mm3) = A × B2, with A as the largest dimension and B as the perpendicular diameter.
Results
Identification of differentially expressed PAUF mRNA in pancreatic ductal adenocarcinomas. We compared the gene expression of pancreatic ductal adenocarcinoma with the normal pancreas using the Affymetrix HG‐U133 GeneChip set. The microarray data showed 1681 known and unknown genes that were differentially expressed in pancreatic cancer and normal tissue samples (fold change > 2, P < 0.05). From the unknown genes, we selected an up‐regulated Affymetrix fragment (11.13‐fold, P < 0.001) in pancreatic cancer that did not increase in normal tissues (Fig. 1a). After sequencing (see below), this gene was named PAUF. Quantitative real‐time RT‐PCR confirmed the up‐regulation of PAUF mRNA in pancreatic ductal adenocarcinomas (Fig. 1b). Similar to data generated by the oligonucleotide microarray (Fig. 1a), the expression level of PAUF increased 11.33‐fold (P = 0.013) in pancreatic cancer tissue samples. Matched sample comparisons in other tissue subsets also confirmed up‐regulated PAUF in cancerous tissue compared to adjacent non‐cancerous tissue (Supporting Fig. S1). These results confirmed that the transcription of PAUF increased in pancreatic cancer.
Figure 1.
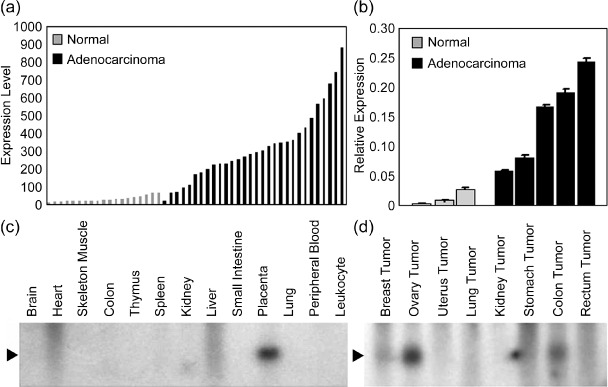
Expression of pancreatic adenocarcinoma up‐regulated factor (PAUF) in human tumor and non‐cancerous tissues. (a) Expression levels of PAUF in normal pancreas and pancreatic adenocarcinomas were compared with data acquired from Affymetrix HG‐U133 chip set analysis. PAUF was significantly up‐regulated in pancreatic adenocarcinomas compared to normal pancreatic tissue. (b) The differential expression of PAUF was validated by quantitative real‐time polymerase chain reaction in pancreatic cancer compared to adjacent normal pancreas. The relative expression value of PAUF was measured by threshold value relative to C‐terminal binding protein (CTBP). (c) The size of PAUF mRNA transcripts was measured using Human 12‐lane MTN blot containing various human normal tissues; only the placenta showed strong PAUF expression. (d) Northern blot analysis of PAUF from tumor tissues using MTN Human Tumor Blot. High expression of PAUF was found in colon, ovary, and stomach tumors.
Combined mining of the above data by GeneExpress Oncology Datasuite (Gene Logic, Inc., Columbia, MD, USA) and expression levels of PAUF in various malignant neoplasms compared to control tissues are shown in the Supporting Table S2.
Sequencing and cloning of full‐length PAUF cDNA. The size of mRNA transcripts and expression patterns of PAUF were determined by northern blot analysis. A northern blot containing total RNAs from various human tissues and tumors was used. The Northern blot showed a single transcript for this gene, which was approximately 0.8 kb in size (Fig. 1c,d). High expression of PAUF was noted in colon and ovarian tumors, consistent with results obtained by GeneExpress Oncology Datasuite analysis (Fig. 1d and Supporting Table S2). Only the placenta showed high expression of PAUF in control tissues (Fig. 1c).
To obtain full‐length cDNA, we screened the placental library and obtained a clone containing a 721‐bp insert with a putative open reading frame of 196 amino acids (Fig. 2a). Comparison with genome databases showed that it was located on chromosome 16p13.3 with four exons. The gene was named PAUF and submitted to GenBank (accession no. EF067313). A homology search using the BLAST database indicated that the sequence of PAUF matched the sequence of the Homo sapiens similar to common salivary protein 1 mRNA (LOC124220; HRPE773; GenBank accession no. NM_145252), but showed a single base difference in two regions of the 3′‐untranslated region. Although the N‐terminus of the PAUF transcript was shorter than the similar to common salivary protein 1 transcript (the mRNA sequence was recently updated to 846 bp), both the PAUF protein and the similar to common salivary protein 1 were predicted in silico to express the same putative secretory protein (data not shown). The PAUF sequence was compared using the SMART and EMBL protein databases. The sequence had a putative hydrophobic 40‐amino acid signal peptide and a putative jacalin‐related lectin domain but did not share homology with any known protein. The probable signal cleavage predicted by the CBS prediction site (http://www.cbs.dtu.dk/services/) would result in the production of a 17‐kDa secreted protein with an N‐terminal glycine (residue 41 of the intact protein). Analysis of the predicted protein using the PredictProtein server (http://cubic.bioc.columbia.edu/predictprotein/) suggested that PAUF was a monomer with one N‐glycosylation site, two protein kinase C phosphorylation sites, two casein kinase II phosphorylation sites, and three N‐myristoylation sites.
Figure 2.

Nucleotide and amino acid sequences of pancreatic adenocarcinoma up‐regulated factor (PAUF) cDNA. (a) PAUF contains 721 bp. An inframe termination codon is located at nucleotide positions 600–602 ( ), and the canonical 3′‐polyadenylation signal AATAAA is located at nucleotide positions 698–703 (_). PAUF encodes a 196‐amino acid protein, including a 40‐amino acid signal peptide (bold type; cleavage site, bold arrow) with putative post‐translational modification sites. Specifically, one N‐glycosylation site at amino acids 185–189 (
), and the canonical 3′‐polyadenylation signal AATAAA is located at nucleotide positions 698–703 (_). PAUF encodes a 196‐amino acid protein, including a 40‐amino acid signal peptide (bold type; cleavage site, bold arrow) with putative post‐translational modification sites. Specifically, one N‐glycosylation site at amino acids 185–189 ( ); three phosphorylation sites at amino acids 7–9, 53–56, and 125–128 (
); three phosphorylation sites at amino acids 7–9, 53–56, and 125–128 ( ); and three N‐myristorylation sites at amino acids 64–69, 90–95, and 137–143 (
); and three N‐myristorylation sites at amino acids 64–69, 90–95, and 137–143 ( ) were predicted in silico. (b) The amino acid sequences of human PAUF, chimpanzee XP_523271 (similar to HRPE773), rhesus monkey XP_001086870 (similar to zymogen granule membrane protein 16 isoform 2), and mouse NP_064294 (demilone cell and parotid protein [Dcpp1]) were aligned by CLUSTAL W (1.83) multiple sequence alignment. Amino acid identities in all sequences are indicated by an asterisk while conserved and semiconserved substitutions are shown by a colon and a dot, respectively.
) were predicted in silico. (b) The amino acid sequences of human PAUF, chimpanzee XP_523271 (similar to HRPE773), rhesus monkey XP_001086870 (similar to zymogen granule membrane protein 16 isoform 2), and mouse NP_064294 (demilone cell and parotid protein [Dcpp1]) were aligned by CLUSTAL W (1.83) multiple sequence alignment. Amino acid identities in all sequences are indicated by an asterisk while conserved and semiconserved substitutions are shown by a colon and a dot, respectively.
The PAUF sequence was compared by sequence alignment with the GenBank and EMBL protein databases. Protein sequence similarity was particularly high in the putative chimpanzee protein (XP_523271) and the putative rhesus monkey protein (XP_001086870; Fig. 2b). The predicted chimpanzee and rhesus monkey homologs displayed 98% and 92% amino acid identity and 100% and 95% amino acid similarity, respectively, to the PAUF amino acid sequence. Conversely, when aligned with the above sequences, the mouse Dcpp protein (NP_064294) showed relatively low identity (31%) and similarity (45%).
Establishment of PAUF overexpressing CHO cells and analysis of PAUF post‐translational modification. To define whether PAUF could be secreted into the extracellular medium, the recombinant plasmid pcDNA3.1(+)‐PAUF‐Myc/His (CHO/PAUF) or the pcDNA3.1(+)‐Myc/His vector (CHO/Mock) were stably transfected into CHO cells and selected by G418. Selected clones of transformed cells had the same manners of PAUF expression and phenotypes. One clone of CHO/Mock and CHO/PAUF were used for further studies. Western blot analysis in CHO/PAUF cells revealed the presence of His‐tagged PAUF in the culture medium, indicating that PAUF was secreted (Fig. 3a). However, the PAUF fusion protein in the culture medium was detected as larger than PAUF from cell lysates, and differed from the predicted size (Fig. 3a). Two‐dimensional (2DE) gel analysis of the PAUF fusion protein in the culture medium revealed multiple spots (data not shown), suggesting post‐translational modification of the PAUF. To investigate glycosylation as a potential post‐translational modification, PAUF was purified from the culture medium of CHO/PAUF cells and treated with PNGase F; this enzyme treatment resulted in complete deglycosylation of PAUF as demonstrated by a decreased apparent molecular mass of PAUF (Fig. 3b). The secretion of N‐glycosylated PAUF in human pancreatic cancer cells was confirmed by tunicamycin. By tunicamycin treatment, CFPAC‐1 cells secreted the deglycosylated form of PAUF suggesting the glycosylation of exo‐ and endogenous secreted PAUF proteins (Fig. 3c). The deglycosylated PAUF bands revealed as double bands, and this may suggest the presence of additional post‐translational modification. N‐terminal peptide sequencing using excised PAUF bands from a SDS‐PAGE gel demonstrated that the first five amino acids were Gly–Lys–Met–Tyr–Gly–Pro (residue 41–46), and consistent with in silico predictions, the cleavage site was positioned between amino acids 40 and 41 (data not shown).
Figure 3.

Subcellular localization and post‐translational modification of pancreatic adenocarcinoma up‐regulated factor (PAUF). The pcDNA3.1(+)‐PAUF‐Myc/His (CHO/PAUF) or pcDNA3.1(+)‐Myc/His vectors (CHO/Mock) were transfected stably into Chinese hamster ovary (CHO) cells. (a) Western blot analysis of the parental and transfected CHO cell lines: PAUF expression was detected with anti‐His antibody in cell lysates (CL) and culture medium (CM) from CHO/PAUF cells. (b) Secreted PAUF protein was purified from culture medium of CHO/PAUF and digested with Peptide‐N‐glycosidase F (PNGase F) for 1 h. The samples were electrophoresed and immunoblotted with anti‐His antibody. Treatment with PNGase F resulted in complete de‐glycosylation of PAUF as demonstrated by immunoreactivity of anti‐His antibodies with protein bands of lower molecular mass. (c) Secreted PAUF protein was purified from medium of 2‐µg/mL tunicamycin (TM) treated CFPAC‐1 cells. Treatment with tunicamycin resulted in complete de‐glycosylation of PAUF as demonstrated by immunoreactivity of antirhPAUF antibodies with protein bands of lower molecular mass.
PAUF changes phenotypes of CHO cells in vitro. The biological function of PAUF was determined by comparing cell growth rate, migration, and invasion ability between CHO/Mock and CHO/PAUF cells. Induction of PAUF increased the proliferation rate of CHO cells by 1.2‐fold (Fig. 4a). The doubling time of CHO/PAUF cells (18.6 h) was relatively shorter than that of CHO/Mock cells (21.6 h). No accompanying morphological changes due to the overexpression of PAUF were observed. Overexpression of PAUF also resulted in increased migration and invasion potential of CHO cells. In CHO/PAUF cells, cell migration increased 2.1‐fold compared to CHO/Mock cells: 74.23 ± 2.83 (mean ± SEM) cells/microscopic field were observed for CHO/Mock cells and 145.8 ± 11.67 cells/microscopic field for CHO/PAUF cells (Fig. 4b). Cell invasion also increased 3.2‐fold in cells expressing PAUF (37.33 ± 5.81 in CHO/Mock versus 101.7 ± 9.39 cells/microscopic field in CHO/PAUF) (Fig. 4c). Other clones of PAUF‐stable cells showed similar PAUF‐dependent enhancement of growth, cell migration, and invasion ability (Supporting Fig. S2). These results demonstrated that PAUF expression was associated with increased motility and invasiveness of cells in vitro.
Figure 4.

Effects of pancreatic adenocarcinoma up‐regulated factor (PAUF) expression on cell proliferation, invasion, and tumorigenicity. (a) The transfected cells (2 × 103 cells per well) were counted every 24 h using a hemocytometer. The Chinese hamster ovary (CHO)/PAUF ( ) cells showed a higher proliferation rate compared to CHO/Mock cells (
) cells showed a higher proliferation rate compared to CHO/Mock cells ( ). (b) Cell migration ability was assessed based on the number of cells per microscopic field (×100) that migrated through the pores of a Transwell chamber to the under surface of the membrane after an incubation time of 24 h. In CHO/PAUF cells, cell migration increased compared to CHO/Mock cells. (c) Cell invasion ability was assessed by counting the number of cells per microscopic field (×100) that migrated through Matrigel invasion chambers after 48 h. Cell invasion ability increased in CHO/PAUF cells compared to CHO/Mock cells. (d) The CHO/PAUF cells were injected into the flank of 6‐week‐old female Balb/c nude mice. The CHO/Mock cells were used as a control. Mice were monitored for 32 days after injection (n = 5 for each). Data are shown as the mean ± SEM. Representative images of tumor xenografts after 32 days shows that grafting with CHO/PAUF cells produced larger tumors than CHO/Mock treatments.
). (b) Cell migration ability was assessed based on the number of cells per microscopic field (×100) that migrated through the pores of a Transwell chamber to the under surface of the membrane after an incubation time of 24 h. In CHO/PAUF cells, cell migration increased compared to CHO/Mock cells. (c) Cell invasion ability was assessed by counting the number of cells per microscopic field (×100) that migrated through Matrigel invasion chambers after 48 h. Cell invasion ability increased in CHO/PAUF cells compared to CHO/Mock cells. (d) The CHO/PAUF cells were injected into the flank of 6‐week‐old female Balb/c nude mice. The CHO/Mock cells were used as a control. Mice were monitored for 32 days after injection (n = 5 for each). Data are shown as the mean ± SEM. Representative images of tumor xenografts after 32 days shows that grafting with CHO/PAUF cells produced larger tumors than CHO/Mock treatments.
PAUF promotes tumor formation in vivo. To investigate the effect of PAUF on tumor formation in vivo, both CHO/PAUF and CHO/Mock cells were injected subcutaneously into nude mice, and the tumor volume was monitored. The CHO/PAUF cells promoted dramatic increases in tumor size compared to CHO/Mock cells (Fig. 4d). After 32 days, PAUF‐derived xenografts showed a 3.8‐fold increase in tumor volume compared to mock xenografts (Fig. 4d); the average tumor size (± SEM) was 1491 ± 280.7 mm3 in PAUF‐derived xenografts compared to 391.9 ± 302.2 mm3 in control mice. Two of the five CHO/Mock cell injections did not result in tumor formation.
Expression of PAUF in pancreatic cancer cells and human pancreatic cancer tissues. mRNA and protein expression profiles of PAUF in various pancreatic cancer cells were analyzed. The CHO/PAUF cells were used as a positive control. The PAUF mRNA was variably expressed in six of eight pancreatic cancer cell lines. In contrast, PAUF was not detected in normal or transformed cell lines: PBMC, PHA‐stimulated PBMC, BMSC, NIH/3T3, and CHO cells had no detectable PAUF mRNA (Fig. 5A).
Figure 5.

Expression of pancreatic adenocarcinoma up‐regulated factor (PAUF) mRNA and protein in pancreatic cells and tissue. (A) Reverse transcription–polymerase chain reaction analysis of PAUF mRNA expression in various cell lines. Upper, the expression of PAUF mRNA in the normal cell lines: peripheral blood mononuclear cells (PBMC), PBMC_st, NIH/3T3, and bone marrow stromal (BMSC) cells. The CHO/PAUF cells served as a positive control for PAUF. Lower, the expression of PAUF mRNA pancreatic cancer cell lines and in parental, vector‐ (CHO/Mock), and CHO/PAUF cells. Beta‐actin (ACTB) served as an internal control. AsPC‐1; BxPC‐3; CFPAC‐1; HPAC, human pancreatic cancer cell line; MIAPaCa‐2; PANC‐1. (B) Western blot analysis of PAUF in pancreatic cancer cell lines; 20 µg of protein from culture media (CM) and cell lysates (CL) were probed with antirecombinant human PAUF (antirhPAUF) antibodies. For CHO/PAUF cells, which served as a positive control, 0.2 µg of protein from the culture medium was loaded. Molecular mass markers (kDa) are shown on the left. (C) Expression of PAUF in pancreatic tissue. Immunohistochemical staining with antirhPAUF antibodies revealed strong cytoplasmic overexpression of PAUF in cancer and islets cells. No significant staining was observed in normal ductal and acinar cells. (a) normal pancreas (×400); (b) pancreatic adenocarcinoma (×100); (c) pancreatic adenocarcinoma (×400); (d) metaplastic duct in chronic pancreatitis (×1000); (e) pancreatic adenocarcinoma (×1000); and (f) pancreatic adenocarcinoma (×1000).
We next generated antirhPAUF antibodies in rabbits as described in ‘Materials and Methods’. Anti‐rhPAUF antibodies were immunoreactive with PAUF in cell lysates and in the culture medium from six of the eight pancreatic cancer cell lines (Fig. 5B), confirming PAUF expression and secretion.
Expression of PAUF was also confirmed in human pancreatic cancer tissues (Fig. 5C). Immunohistochemical staining revealed strong cytoplasmic overexpression of PAUF in cancer cells and preneoplastic lesions such as metaplastic and dysplastic ducts (Fig. 5Cb–f). Strong immunoreactivity in apicolateral membrane in metaplastic ductal cells was clearly noticed (Fig. 5Cd). The islet cells also showed strong immunoreactivity (Fig. 5 Ca); however, normal pancreatic ducts, acinar cells, interstitial cells, and blood vessels did not react with the antibody.
Knockdown effect of PAUF in pancreatic cancer cells by siRNA. For knockdown of PAUF, CFPAC‐1 cells were transiently transfected with three sets of stealth RNAi targeting LOC124220 (si#1, si#2, and si#3). At 48 h post‐transfection, mRNA and secreted PAUF expression levels were down regulated in all three sets of siRNA‐transfected cells compared to control siRNA‐transfected cells (Fig. 6a). Down‐regulation of PAUF also resulted in decreased migration and invasion potential of CFPAC‐1 cells. In PAUF down‐regulated CFPAC‐1 cells, cell migratory potential decreased compared to negative control siRNA‐transfected cells: 338.5 ± 61.5 (mean ± SEM) cells/microscopic field were observed for negative control cells, 30.5 ± 0.5 cells/microscopic field for si#1 cells, 60.5 ± 5.5 cells/microscopic field for si#2 cells, and 122.0 ± 7.0 cells/microscopic field for si#3 cells (Fig. 6b). Cell invasion also decreased in PAUF down‐regulated CFPAC‐1 cells: 492.5 ± 23.5 cells/microscopic field were observed for control cells, 239.5 ± 1.5 cells/microscopic field for si#1 cells, 318.0 ± 2.0 cells/microscopic field for si#2 cells, and 368.5 ± 1.5 cells/microscopic field for si#3 cells (Fig. 6c). These findings demonstrated that down‐regulation of PAUF was associated with the suppression of cancer cell migration and invasion in vitro.
Figure 6.

Effects of inhibiting pancreatic adenocarcinoma up‐regulated factor (PAUF) in pancreatic cancer cells. Knockdown effect with PAUF small‐interfering RNA (siRNA)‐transfected CFPAC‐1 cells (a–c). PAUF siRNA were transiently transfected into CFPAC‐1 cells for 48 h. (a) At 48 h post‐transfection, cells were harvested and subjected to total RNA extraction, and reverse transcription–polymerase chain reaction was performed. Secreted protein was prepared from culture medium at 48 h post‐transfection. mRNA and protein levels of PAUF of siRNA‐transfected CFPAC‐1 cells decreased. ACTB, beta‐actin. (b) Cell migration ability was assessed based on the number of cells per microscopic field (×200) that migrated through the pores of a Transwell chamber to the under surface of the membrane after an incubation time of 24 h. In PAUF siRNA‐tranfected CFPAC‐1 cells, cell migration ability decreased compared to negative control cells. (c) Cell invasion ability was assessed by counting the number of cells per microscopic field (×200) that migrated through Matrigel invasion chambers after 48 h. Cell invasion ability was decreased in PAUF siRNA‐tranfected CFPAC‐1 cells compared to negative control cells. Effects of inhibiting PAUF with antirecombinant human PAUF (antirhPAUF) antibodies (d–f). The graphs (d,e) show the mean ± SD (% of antibody‐treated cells/non‐treated cells per microscopic field) from triplicate samples. (d) CFPAC‐1 cells were plated on a Boyden chamber and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies ( ) or normal rabbit IgG (
) or normal rabbit IgG ( ). AntirhPAUF antibodies effectively inhibited the migration of CFPAC‐1 in a dose‐dependent manner. (e) CFPAC‐1 cells were plated on Matrigel‐coated filters and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies (
). AntirhPAUF antibodies effectively inhibited the migration of CFPAC‐1 in a dose‐dependent manner. (e) CFPAC‐1 cells were plated on Matrigel‐coated filters and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies ( ) or normal rabbit IgG (
) or normal rabbit IgG ( ). Invasion through Matrigel‐coated membranes was dose‐dependently inhibited by antirhPAUF antibodies. (f) Tumor‐bearing mice generated by the injection of CFPAC‐1 were treated with antirhPAUF antibodies (
). Invasion through Matrigel‐coated membranes was dose‐dependently inhibited by antirhPAUF antibodies. (f) Tumor‐bearing mice generated by the injection of CFPAC‐1 were treated with antirhPAUF antibodies ( ) or normal rabbit IgG (
) or normal rabbit IgG ( ) via the tail vein twice a week (n = 6 for each) and tumor mass was monitored (left) for 32 days. Data are shown as the mean ± SEM. The starting day of antibody injection is represented by the arrow. The representative image shows the results of tumor xenografts at the end of the experiments.
) via the tail vein twice a week (n = 6 for each) and tumor mass was monitored (left) for 32 days. Data are shown as the mean ± SEM. The starting day of antibody injection is represented by the arrow. The representative image shows the results of tumor xenografts at the end of the experiments.
Inhibition of cancer cell migration and invasion by antirhPAUF pAb. The effect of exogenous antirhPAUF antibodies on the motility and invasiveness of the human pancreatic cancer cell CFPAC‐1 was evaluated. As shown in Figure 6(d,e), antirhPAUF antibodies effectively inhibited the migration and invasion of CFPAC‐1 in a dose‐dependent manner. Similar effects were observed in other pancreatic cancer cell lines such as AsPC‐1, BxPC‐3, MIAPaCa‐2, and PANC‐1 (Supporting Fig. S3).
Inhibition of tumor growth in vivo by antirhPAUF pAb. To investigate the consequences of PAUF inhibition on tumor growth in vivo, the antibodies were administered via the tail vein to tumor‐bearing mice generated by injection of CFPAC‐1. Treatment with antirhPAUF antibodies inhibited the tumor growth (Fig. 6f), and compared to non‐specific rabbit IgG‐treated mice, treatment with antirhPAUF antibodies resulted in significantly smaller tumors (Fig. 6f). The average tumor size (± SEM) in antibody‐treated mice was 950.7 ± 202.5 mm3 compared to 2732 ± 932.2 mm3 in controls. Body weight did not change in either group.
Discussion
We identified a novel EST fragment that was highly expressed in pancreatic cancer. This secretory protein, PAUF, contained a signal peptide at its NH2‐terminus, and PAUF mRNA and protein were expressed abundantly in most of the tested pancreatic cancer cell lines and in human pancreatic cancer tissues. Based on the following in vitro and in vivo observations, we can propose that PAUF functions as a growth factor and plays a role in tumor progression: overexpression of PAUF enhanced cell growth, migration, and invasion; the migration and invasion‐promoting effects of PAUF were down‐regulated and neutralized in vitro by siRNA and antirhPAUF antibodies in a dose‐dependent manner; xenografts of PAUF‐expressing CHO cells not only promoted tumor growth but also resulted in metastatic nodules at the peritoneum and omentum of mice in vivo (data not shown); and antirhPAUF antibodies reduced tumor growth in vivo.
The nucleotide sequence of PAUF matched the mRNA sequence of the Homo sapiens similar to common salivary protein 1 (LOC124220), a hypothetical protein with an unknown function. According to Clark et al. the LOC124220 protein is a secretory protein.( 16 ) The LOC124220 protein was discovered by human gastric cancer transcriptome analysis,( 17 ) and consistent with its putative function. PAUF was expressed in gastric cancer as detected by our Genechip analysis. According to Mullins et al. the HRPE773 (GenBank AAQ89380.1) protein was suggested to be the putative human CSP‐1/Dcpp.( 15 ) Note that Dcpp genes on mouse chromosome 17 and the human ortholog on chromosome 16 are flanked by the mouse and human orthologs of several genes including a family of serine proteases.
Many growth factors have been shown to participate in normal and abnormal pancreatic growth,( 18 , 19 , 20 , 21 , 22 , 23 , 24 ) and PAUF might be one of these factors. Recent studies suggest that various growth factors of solid tumors are cancer treatment targets. Accordingly, our results suggest that PAUF is a potential target for the treatment of pancreatic cancer. Inhibiting PAUF with specific antibodies resulted in dramatic decreases in tumor growth, migration, and invasion, which implies that down‐regulation of PAUF might be a novel therapy for pancreatic cancer.
The increased expression of PAUF in pancreatic cancer tissues, as shown by GeneChip analysis and quantitative real‐time PCR, suggest that PAUF may be a reliable biomarker for pancreatic adenocarcinoma. Many clinical markers and therapeutic targets in cancer are glycoproteins, such as cancer antigen (carbohydrate antigen Sialyl Lewisa[CA])‐19‐9 in pancreatic cancer, CA125 in ovarian cancer, human epidermal growth factor receptor 2 (receptor tyrosine‐protein kinase erbB‐2 [ERBB2/Her2/neu]) in breast cancer, and prostate‐specific antigen in prostate cancer. Protein glycosylation is a common post‐translational modification and plays a fundamental role in many biological processes,( 25 , 26 ) according to our computational and experimental analysis, PAUF is also glycosylated. Furthermore, glycoproteins secreted into the bloodstream comprise a major part of the serum proteome.( 27 ) Our analyses also showed that PAUF has a putative jacalin‐related lectin domain. Jacalin is from the seed of the Artocarpus heterophyllus (jackfruit), which is specific for galactose and can bind to the tumor‐associated T‐antigen.( 28 ) Moreover, proteins containing lectins may be involved in cell–cell communications, host–pathogen interactions, cancer metastasis, embryogenesis, and tissue development.( 29 )
A BLAST search for potential mammalian PAUF protein orthologs using GenBank protein databases revealed significant homology only to primates such as chimpanzees and rhesus monkeys. Mullins et al. reported that human HRPE773 is the human ortholog of the mouse Dcpp gene and the rat CSP1 gene, but the amino acid identity between these two genes is only 31%.( 15 ) The degree of conservation of coding regions between humans and mice has been reported to be nearly 85%.( 30 ) Therefore, a low conservation rate of PAUF in primates suggests that PAUF might be a rapidly evolving primate gene. Similarly, genes responsible for higher cognitive functions such as G72 and disrupted‐in schizophrenia‐1 (DISC1) have low sequence identity between rodents and primates, indicating that they have developed recently on the evolutionary scale.( 31 , 32 , 33 )
In conclusion, we report a novel secretory protein, PAUF, which is highly expressed in pancreatic cancer and might be involved in cancer progression. Further studies for the identification of its receptor, its downstream signaling pathways, other post‐translational modification, and its molecular mechanisms of tumor progression are required. These future studies may enable us to develop novel diagnostic tests for pancreatic cancer and could lead to new therapeutic approaches to treat pancreatic cancer.
Supporting information
Fig. S1. Expression of pancreatic adenocarcinoma up‐regulated factor (PAUF) in cancerous tissues and adjacent non‐cancerous tissues from matched samples. Seven cases were compared using quantitative real‐time polymerase chain reaction, which showed up‐regulated PAUF in cancerous tissues.
Fig. S2. The effects of pancreatic adenocarcinoma up‐regulated factor (PAUF) on cell proliferation, migration, and invasion in transfected Chinese hamster ovary (CHO) clones. (a) The transfected cells (1.25 × 104 cells per well) were counted every 24 h using a hemocytometer. The CHO/PAUF clones showed a higher proliferation rate compared to CHO/Mock clones. (b) Cell migration ability was assessed based on the number of cells per microscopic field (×200) that migrated through the pores of a Transwell chamber to the under surface of the membrane after an incubation time of 24 h. In CHO/PAUF clones, cell migration increased compared to CHO/Mock clones. (c) Cell invasion ability was assessed by counting the number of cells per microscopic field (×200) that migrated through Matrigel invasion chambers after 48 h. Cell invasion ability increased in CHO/PAUF clones compared to CHO/Mock clones.
Fig. S3. Effects of inhibiting pancreatic adenocarcinoma up‐regulated factor (PAUF) with antirecombinant human PAUF (antirhPAUF) antibodies in pancreatic cancer cell lines. (a) Four pancreatic cancer cell lines were plated on a Boyden chamber and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies () or normal rabbit IgG (). AntirhPAUF antibodies effectively inhibited the migration of cancer cells in a dose‐dependent manner. (b) Four pancreatic cancer cell lines were plated on Matrigel‐coated filters and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies () or normal rabbit IgG (). Invasion through Matrigel‐coated membranes was dose‐dependently inhibited by antirhPAUF antibodies.
Table S1. Sequences of primer set for reverse transcription–polymerase chain reaction and polymerase chain reaction and synthesized oligonucleotide sequences for siRNA.
Table S2. Expression levels of the pancreatic adenocarcinoma up‐regulated factor (PAUF) gene in various malignant neoplasms. PAUF expression levels were compared to normal control tissues from data from Affymetrix HG‐U133 chip set analysis using GeneExpress Oncology Datasuite. Mucinous adenocarcinomas of colon cancer, mucinous cystadenocarcinomas of ovarian cancer, and signet ring cell carcinomas of stomach cancer exhibited significant up‐regulation of PAUF compared to its expression in normal tissues.
Please note: Wiley‐Blackwell are not responsible for the content or functionality of any supporting materials supplied by the authors. Any queries (other than missing material) should be directed to the corresponding author for the article.
Supporting info item
Acknowledgments
This study was supported by grant FG03‐32‐02, FG07‐32‐02 and FG08‐32‐02 of the 21C Frontier Functional Human Genome Project from the Ministry of Science and Technology of Korea, and in part by a grant from the Brain Korea 21 Project for Medical Sciences of Yonsei University College of Medicine.
References
- 1. Jemal A, Murray T, Ward E et al . Cancer Statistics, 2005. CA: A Cancer J Clin 2005; 55: 10–30. [DOI] [PubMed] [Google Scholar]
- 2. Yeo CJMDC, John L, Sohn TA. Six hundred fifty consecutive pancreaticoduodenectomies in the 1990s: pathology, complications, and outcomes. Ann Surg 1997; 226: 248–60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Hawes RH, Xiong Q, Waxman I, Chang KJ, Evans DB, Abbruzzese JL. A multispecialty approach to the diagnosis and management of pancreatic cancer. Am J Gastroenterol 2000; 95: 17–31. [DOI] [PubMed] [Google Scholar]
- 4. Hu YX, Watanabe H, Ohtsubo K et al . Frequent loss of p16 expression and its correlation with clinicopathological parameters in pancreatic carcinoma. Clin Cancer Res 1997; 3: 1473–7. [PubMed] [Google Scholar]
- 5. Dong M, Nio Y, Tamura K et al . Ki‐ras point mutation and p53 expression in human pancreatic cancer: a comparative study among Chinese, Japanese, and Western patients. Cancer Epidemiol Biomarkers Prev 2000; 9: 279–84. [PubMed] [Google Scholar]
- 6. Wilentz RE, Su GH, Dai JL et al . Immunohistochemical labeling for Dpc4 mirrors genetic status in pancreatic adenocarcinomas: a new marker of DPC4 inactivation. Am J Pathol 2000; 156: 37–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Liu Q, Yan YX, McClure M, Nakagawa H, Fujimura F, Rustgi AK. MTS‐1 (CDKN2) tumor suppressor gene deletions are a frequent event in esophagus squamous cancer and pancreatic adenocarcinoma cell lines. Oncogene 1995; 10: 619–22. [PubMed] [Google Scholar]
- 8. Bruce AR, Lingyi H, Moria W. et al. Amplification and overexpression of the AKT2 oncogene in a subset of human pancreatic ductal adenocarcinomas. Mol Carcinogenesis 1998; 21: 81–6. [PubMed] [Google Scholar]
- 9. Lee S, Gang J, Jeon SB et al . Molecular cloning and functional analysis of a novel oncogene, cancer‐upregulated gene 2 (CUG2). Biochem Biophys Res Comms 2007; 360: 633–9. [DOI] [PubMed] [Google Scholar]
- 10. Iacobuzio‐Donahue CA, Ashfaq R, Maitra A et al . Highly expressed genes in pancreatic ductal adenocarcinomas: a comprehensive characterization and comparison of the transcription profiles obtained from three major technologies. Cancer Res 2003; 63: 8614–22. [PubMed] [Google Scholar]
- 11. Iacobuzio‐Donahue CA, Maitra A, Olsen M et al . Exploration of global gene expression patterns in pancreatic adenocarcinoma using cDNA microarrays. Am J Pathol 2003; 162: 1151–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Iacobuzio‐Donahue CA, Maitra A, Shen‐Ong GL et al . Discovery of novel tumor markers of pancreatic cancer using global gene expression technology. Am J Pathol 2002; 160: 1239–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Iacobuzio‐Donahue CAH, Ralph H. Gene expression in neoplasms of the pancreas: applications to diagnostic pathology. Adv Anat Pathol 2003; 10: 125–34. [DOI] [PubMed] [Google Scholar]
- 14. McManaman J, Reyland M, Thrower E. Secretion and fluid transport mechanisms in the mammary gland: comparisons with the exocrine pancreas and the salivary gland. J Mammary Gland Biol Neoplasia 2006; 11: 249–68. [DOI] [PubMed] [Google Scholar]
- 15. Mullins JJ, Mullins LJ, Dunbar DR, Brammar WJ, Gross KW, Morley SD. Identification of a human ortholog of the mouse Dcpp gene locus, encoding a novel member of the CSP‐1/Dcpp salivary protein family. Physiol Genomics 2006; 28: 129–40. [DOI] [PubMed] [Google Scholar]
- 16. Clark HF, Gurney AL, Abaya E et al . The secreted protein discovery initiative (SPDI), a large‐scale effort to identify novel human secreted and transmembrane proteins: a bioinformatics assessment. Genome Research 2003: 2265–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Jung‐Hwa O, Jin Ok Y, Yoonsoo H et al . Transcriptome analysis of human gastric cancer. Mammalian Genome 2005; V16: 942–54. [DOI] [PubMed] [Google Scholar]
- 18. Ho JJL, Farrelly ER, Kim YS. Phorbol ester reduces phosphorylation of epidermal growth factor receptor in pancreatic cancer cells by activation of a tyrosine phosphatase. Biochem Biophys Res Comms 1999; 265: 728–33. [DOI] [PubMed] [Google Scholar]
- 19. I EI‐Hariry MPaNRL . FGF‐1 and FGF‐2 modulate the E‐cadherin/catenin system in pancreatic adenocarcinoma cell lines. Br J Cancer 2001; 84: 1656–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Wagner MLM, Cahn M, Korc M. Suppression of fibroblast growth factor receptor signaling inhibits pancreatic cancer growth in vitro and in vivo . Gastroenterology 1998; 114: 798–807. [DOI] [PubMed] [Google Scholar]
- 21. Flossmann‐Kast BBM, Jehle PM, Hoeflich A, Adler G, Lutz MP. Src stimulates insulin‐like growth factor I (IGF‐I)‐dependent cell proliferation by increasing IGF‐I receptor number in human pancreatic carcinoma cells. Cancer Res 1998; 58: 3551–4. [PubMed] [Google Scholar]
- 22. Datto MB, Li Y, Panus JF, Howe DJ, Xiong Y, Wang X. Transforming growth factor {beta} induces the cyclin‐dependent kinase inhibitor p21 through a p53‐independent mechanism. PNAS 1995; 92: 5545–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. James WF, Cynthia AM, William ES. Increased tumorigenicity in the human pancreatic cell line MIA PaCa‐2 Is associated with an aberrant regulation of an IGF‐1 autocrine loop and lack of expression of the TGF‐β type RII receptor. J Cellular Physiol 1995; 165: 155–63. [DOI] [PubMed] [Google Scholar]
- 24. Marko K, Pam T, Murray K. TGF‐β‐1 up‐regulates cyclin D1 expression in colo‐357 cells, whereas suppression of cyclin d1 levels is associated with down‐regulation of the type I TGF‐β receptor. Int J Cancer 1999; 83: 247–54. [DOI] [PubMed] [Google Scholar]
- 25. Bertozzi CR, Kiessling LL. Chemical glycobiology. Science 2001; 291: 2357–64. [DOI] [PubMed] [Google Scholar]
- 26. Rudd PM, Elliott T, Cresswell P, Wilson IA, Dwek RA. Glycosylation and the immune system. Science 2001; 291: 2370–6. [DOI] [PubMed] [Google Scholar]
- 27. Leigh N, Anderson NGA. Proteome and proteomics: new technologies, new concepts, and new words. Electrophoresis 1998; 19: 1853–61. [DOI] [PubMed] [Google Scholar]
- 28. Jeyaprakash AA, Geetha Rani P, Banuprakash Reddy G et al . Crystal structure of the Jacalin‐T‐antigen complex and a comparative study of Lectin‐T‐antigen complexes. J Mol Biol 2002; 321: 637–45. [DOI] [PubMed] [Google Scholar]
- 29. Sharon N, Lis H. Lectins as cell recognition molecules. Science 1989; 246: 227–34. [DOI] [PubMed] [Google Scholar]
- 30. Waterston RHL‐TK, Birney E, Rogers J, Abril JF, Lander ES. Initial sequencing and comparative analysis of the mouse genome. Nature 2002; 420: 520–62. [DOI] [PubMed] [Google Scholar]
- 31. Bord L, Wheeler J, Paek M et al . Primate disrupted‐in‐schizophrenia‐1 (DISC1): high divergence of a gene for major mental illnesses in recent evolutionary history. Neuroscience Res 2006; 56: 286–93. [DOI] [PubMed] [Google Scholar]
- 32. Chumakov I, Blumenfeld M, Guerassimenko O et al . Genetic and physiological data implicating the new human gene G72 and the gene for d‐amino acid oxidase in schizophrenia. Proc Natl Acad Sci 2002; 99: 13675–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Clark AG, Glanowski S, Nielsen R et al . Inferring nonneutral evolution from human‐chimp‐mouse orthologous gene trios. Science 2003; 302: 1960–3. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Fig. S1. Expression of pancreatic adenocarcinoma up‐regulated factor (PAUF) in cancerous tissues and adjacent non‐cancerous tissues from matched samples. Seven cases were compared using quantitative real‐time polymerase chain reaction, which showed up‐regulated PAUF in cancerous tissues.
Fig. S2. The effects of pancreatic adenocarcinoma up‐regulated factor (PAUF) on cell proliferation, migration, and invasion in transfected Chinese hamster ovary (CHO) clones. (a) The transfected cells (1.25 × 104 cells per well) were counted every 24 h using a hemocytometer. The CHO/PAUF clones showed a higher proliferation rate compared to CHO/Mock clones. (b) Cell migration ability was assessed based on the number of cells per microscopic field (×200) that migrated through the pores of a Transwell chamber to the under surface of the membrane after an incubation time of 24 h. In CHO/PAUF clones, cell migration increased compared to CHO/Mock clones. (c) Cell invasion ability was assessed by counting the number of cells per microscopic field (×200) that migrated through Matrigel invasion chambers after 48 h. Cell invasion ability increased in CHO/PAUF clones compared to CHO/Mock clones.
Fig. S3. Effects of inhibiting pancreatic adenocarcinoma up‐regulated factor (PAUF) with antirecombinant human PAUF (antirhPAUF) antibodies in pancreatic cancer cell lines. (a) Four pancreatic cancer cell lines were plated on a Boyden chamber and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies () or normal rabbit IgG (). AntirhPAUF antibodies effectively inhibited the migration of cancer cells in a dose‐dependent manner. (b) Four pancreatic cancer cell lines were plated on Matrigel‐coated filters and incubated for 24 h in the presence of the indicated concentration of antirhPAUF antibodies () or normal rabbit IgG (). Invasion through Matrigel‐coated membranes was dose‐dependently inhibited by antirhPAUF antibodies.
Table S1. Sequences of primer set for reverse transcription–polymerase chain reaction and polymerase chain reaction and synthesized oligonucleotide sequences for siRNA.
Table S2. Expression levels of the pancreatic adenocarcinoma up‐regulated factor (PAUF) gene in various malignant neoplasms. PAUF expression levels were compared to normal control tissues from data from Affymetrix HG‐U133 chip set analysis using GeneExpress Oncology Datasuite. Mucinous adenocarcinomas of colon cancer, mucinous cystadenocarcinomas of ovarian cancer, and signet ring cell carcinomas of stomach cancer exhibited significant up‐regulation of PAUF compared to its expression in normal tissues.
Please note: Wiley‐Blackwell are not responsible for the content or functionality of any supporting materials supplied by the authors. Any queries (other than missing material) should be directed to the corresponding author for the article.
Supporting info item


