Abstract
Acetic acid is a common inhibitor present in lignocellulose hydrolysate, which inhibits the ethanol production by yeast strains. Therefore, the cellulosic ethanol industry requires yeast strains that can tolerate acetic acid stress. Here we demonstrate that overexpressing a yeast native arginase-encoding gene, CAR1, renders Saccharomyces cerevisiae acetic acid tolerance. Specifically, ethanol yield increased by 27.3% in the CAR1-overexpressing strain compared to the control strain under 5.0 g/L acetic acid stress. The global intracellular amino acid level and compositions were further analyzed, and we found that CAR1 overexpression reduced the total amino acid content in response to acetic acid stress. Moreover, the CAR1 overexpressing strain showed increased ATP level and improved cell membrane integrity. Notably, we demonstrated that the effect of CAR1 overexpression was independent of the spermidine and proline metabolism, which indicates novel mechanisms for enhancing yeast stress tolerance. Our studies also suggest that CAR1 is a novel genetic element to be used in synthetic biology of yeast for efficient production of fuel ethanol.
Keywords: Saccharomyces cerevisiae, Arginase, CAR1, Acetic acid tolerance, Amino acid metabolism, Ethanol production
1. Introduction
Increasing greenhouse emissions coupled with rising reliance on fossil energy have compelled policy makers to promote the production of biofuels, such as bioethanol, from lignocellulose biomass using budding yeast Saccharomyces cerevisiae [[1], [2], [3]]. However, growth and metabolism repression by inhibitors present in lignocellulose hydrolysate is one of the limiting factors for economic cellulosic ethanol production [[4], [5], [6], [7], [8]]. Acetic acid is a common inhibitor generated at large quantity during the pretreatment of lignocellulose biomass. It is toxic to the yeast cell and inhibit cellulosic ethanol production due to at least the following three reasons: Firstly, the protons dissociated from acetic acid in the cytosol lead to cytoplasmic acidification, thereby inhibiting important metabolic processes [9,10]; secondly, acetic acid can severely inhibit the activities of transporters and metabolic enzymes [[9], [10], [11], [12]]; and thirdly, acetic acid can induce oxidative stress [13]. Due to the fact that multiple genes are involved in acetic acid tolerance [14], It is necessary to reveal new genes conferring to improved acetic acid tolerance to develop efficient yeast cell factories for bio-conversion of lignocellulosic biomass.
Great efforts have been made to render yeast tolerant to acetic acid by systematic manipulating key genes involved in stress tolerance. For example, enhancement of acetic acid tolerance in S. cerevisiae was achieved by the overexpression of transcriptional factors such as Znf1p [15], Ppr1p [13,16] and Haa1p [17], the overexpression of protein kinases such as Hog1 [18] and Kic1p [19], the overexpression of a histone modification enzyme Set5p [13], elevated pression of the eukaryotic translation factor eIF5A [20], as well as the deletion of acetic acid importer Ady2p [21,22]. On the other hand, protective agents or metal ions were known to exert beneficial effects to yeast cells under stress conditions [23,24]. Among the protective amino acids, it was found that the presence of proline improved growth of S. cerevisiae in the presence of acetic acid stress [[25], [26], [27], [28]]. In our previous studies, we demonstrated that the presence of zinc sulfate exerted protection for S. cerevisiae towards acetic acid stress [29], and using a zinc-finger-based artificial transcription factor can improve the acetic acid tolerance of S. cerevisiae [30]. Furthermore, we also found by integrative transcriptomic and proteomic analysis that elevated expression of arginase (Car1p), which catabolizes arginine to ornithine and urea [31], at both mRNA (by 2.34 folds) and protein (by 1.23 folds) levels associated with the zinc sulfate supplementation during continuous ethanol fermentation. Considering that arginine can be converted to proline, we assume that overexpression of CAR1 may exert protective effects towards S. cerevisiae by improving stress resistance.
In this study, we overexpressed CAR1 in S. cerevisiae and evaluated its effect on acetic acid tolerance. The engineered strain exhibited improved growth and ethanol fermentation performance under acetic acid stress compared to the control strain. We also examined the effects of CAR1 overexpression on transcription of amino acid biosynthesis genes and the intracellular amino acid content. Our work suggested novel roles and mechanisms of yeast CAR1 gene, and also provide a viable strategy to enhance cellulosic ethanol production efficiency by overexpressing arginase.
2. Material and methods
2.1. Strains and culture medium
Escherichia coli DH5α was used for plasmid propagation. The laboratory strains S. cerevisiae S288c and the industrial strain S. cerevisiae 4126 were used for this work. All strains were listed in Table 1.
Table 1.
Yeast strains used in this study.
| Strain | Description | Source |
|---|---|---|
| S. cerevisiae S288c | Laboratory yeast strain for research | Donated by Prof. Jens Nielsen at Chalmers University, Sweden |
| S. cerevisiae 4126 | An industrial yeast strain for ethanol fermentation | Donated by Mrs Jana Otrubo at University of Waterloo, Canada |
| S. cerevisiae 4126-CAR1 | S. cerevisiae 4126 carrying HO::KanMX-PPGK1-CAR1-TCYC1 | This study |
| S. cerevisiae 4126-HO | S. cerevisiae 4126 carrying HO::KanMX | This study |
| S. cerevisiae S288c-CAR1 | S. cerevisiae S288c carrying HO::KanMX-PPGK1-CAR1-TADH1 | This study |
| S. cerevisiae S288c-HO | S. cerevisiae S288c carrying HO::KanMX | This study |
Luria-Bertani medium (NaCl 10 g/L, yeast extract 5.0 g/L and tryptone 10 g/L) was used for E. coli culture, in which 100 μg/mL Ampicilin was supplemented for transformant selection and plasmid propagation. S. cerevisiae strains were cultured in the YPD medium composed of glucose 20 g/L, peptone 20 g/L and yeast extract 10 g/L. For solid agar plates, 20 g/L Bacto agar was added into the YPD medium, and G418 was supplemented at 300 μg/mL for screening the transformant of S. cerevisiae. Fermentation medium composed of 4 g/L yeast extract, 3 g/L peptone and 100 g/L glucose was used for ethanol fermentation by S. cerevisae strains. Acetic acid tolerance was exerted on by adding 5 g/L acetic acid into the YPD agar medium and liquid media, and the initial pH of the media was measured at ∼3.5.
2.2. Construction of recombinant yeast strains
All primers used in this study were listed in Table S1. The open reading frame (ORF) of CAR1 was amplified by polymerase chain reaction (PCR) using the genomic DNA extracted from S. cerevisiae S288c as the template. Then the ORF of CAR1 was cloned into the pHO vector [32] at the XmaI and PacI restriction sites, yielding the CAR1 overexpression plasmid with the PGK1 promoter and the CYC1 terminator. The recombinant plasmid was named pHO-CAR1, and the sequence was confirmed by DNA sequencing at Sangon Biotech, Shanghai, China. Finally, both the empty vector pHO and the CAR1-overexpressing vector pHO-CAR1 were linearized by NotI and transformed into S. cerevisiae 4126 and S. cerevisiae S288c by electroporation as previously described [33]. After screening and verification of the gene integration using diagnostic PCR, the recombinant yeast strain with CAR1 overexpression based on S. cerevisiae 4126 was designated as 4126-CAR1, while the strain transformed with the empty vector was named as 4126-HO.
2.3. Yeast culture and ethanol fermentation
To compare the fermentation performance of S. cerevisiae strains, batch fermentation was carried out in 250 mL flasks with 100 mL fermentation medium, and the initial optical density at 600 nm (OD600) was adjusted to about 0.2 for inoculation with all experiments. All flasks were sealed with rubber plugs for CO2 to be released. After inoculation, cells were incubated in a rotatory shaker operating at 30 °C and 150 rpm. To test acetic acid tolerance for yeast strains during the fermentation, 5.0 g/L acetic acid was added into the fermentation medium before inoculation. Fermentation medium without acetic acid addition served as the control. To test its effect on the fermentation, ornithine was added into the fermentation medium at 0.1, 1.0 and 10.0 mM, respectively. All fermentation experiments described above were conducted without pH control.
2.4. Assessment for yeast growth, cell viability and acetic stress tolerance
Yeast growth was determined by detecting OD600 for the culture with a spectrophotometer (Multiskan GO, ThermoFisher Scientific, Finland). Yeast cell viability was determined by counting the colony forming units (CFUs), which was normalized to that detected for the control: yeast cells cultured at their growth phase without stressful conditions. Both the sample and its control were diluted serially to same cell densities (OD600) suitable for counting CFU, and CFU for the control was designated as 100%. Acetic acid stress tolerance of yeast cells was evaluated on agar plates by spot assay according to reference [13].
2.5. Analysis for ethanol fermentation
The fermentation broth of 1 mL was centrifuged at 10,000×g at room temperature for 2 min to remove yeast cells. The supernatant was further filtrated with 0.22 μm membrane and diluted properly for analysis by high performance liquid chromatography (HPLC) with 20 μL sample uploaded. Glucose, glycerol, acetic acid, ethanol and furfural in the fermentation both were analyzed with high performance liquid chromatography (HPLC, Waters e2695, Waters, MA, USA) equipped with Aminex HPX-87H column (300 mm × 7.8 mm, Bio-Rad, Hercules, CA), the Refractive Index Detector (RID, Waters 2414, Waters, MA, USA) and a UV/Vis Detector (Waters 2498, Waters, MA, USA). The operating temperatures for the column and RI were 65 °C and 50 °C, respectively, and 4 mmol/L H2SO4 was used as the mobile phase.
2.6. Real-time quantitative PCR analysis
Reverse transcription quantitative PCR (RT-qPCR) was performed to determine the relative expression of mRNAs. S. cerevisiae cells cultured in the fermentation medium were grown to their exponential phase, and then harvested by centrifuging at 10,000×g for 5 min at 4 °C. Total RNA was purified from the yeast cells using the Spin Column Yeast total RNA Purification Kit (Sangon, Shanghai, China) following the manufacturer's instruction, which was then reversely transcribed into cDNA using the PrimeScript™ RT Reagent Kit with gDNA Eraser (Takara, Dalian, China) for the PCR reaction using iQ™ SYBR® Green Supermix and CFX Connect™ Real-Time PCR Detection System (Bio-Rad, Shanghai, China). Relative gene expression levels were determined by the 2−ΔΔCt method [34] using ALG9 as the internal control. All the primers for RT-qPCR were listed in Supplementary Table S1.
2.7. Detection of reactive oxygen species accumulation
Yeast cells were collected at the log phase (∼OD600 1.0) by centrifugation and washed twice with 0.1 M PBS (pH 7.0), followed by re-suspending in the PBS containing 10 μM 2′, 7′-dichlorofluorescein diacetate (DCFH-DA). The reactive oxygen species (ROS) accumulation in yeast cells was measured and calculated following the methods described in the previous work [29].
2.8. Determination of intracellular ATP concentration
The intracellular ATP concentration was determined via luminescence method using the ATP Assay Kit (Beyotime, Nanjing, Jiangsu, China) following the manufacturers’ instructions. Briefly, yeast cells were harvest by centrifugation at 4 °C and 10,000×g for 2 min. Then cell pellets were washed twice and immediately lysed using the lysis buffer under low temperature provided by ice. After removing cell debris by centrifugation again at 4 °C, crude cell extract was collected and protected with ice for determining intracellular ATP concentration immediately. The luminescence was detected with the Multi-Mode Detector (Enspire 2300, PerkinElmer, USA). The whole process was completed within 1 h, and the final ATP concentration was normalized with the protein concentration of the crude cell extract determined by the BCA protein assay kit (Beyotime, Nanjing, Jiangsu, China).
2.9. Plasma membrane integrity assay
Yeast cell membrane integrity was determined as described previously [35], with minor modifications. Briefly, 1 mL of yeast cells cultured to exponential-phase were harvested, and washed twice with 0.1 M PBS (pH 7.0). Then, the cells suspended in 1 mL PBS were treated with 15 μM propidium iodide (PI) for 20 min in dark. After washing twice with PBS, cell pellets were re-suspended in 1 mL PBS. The PI fluorescence intensity (excitation, 485 nm; emission, 635 nm) of 200 μL cell suspension was measured by a Multi-Mode Detection (Enspire 2300, PerkinElmer, USA). The final fluorescence intensity of each sample was normalized with cell concentration (OD600).
2.10. Determination of intracellular amino acid compositions
The intracellular amino acid compositions in S. cerevisiae cells were determined by High Speed Amino Acid Analyzer (AAA) (Hitachi L-8900, Tokyo, Japan). Yeast cells growing at their exponential phase were harvested from fermentation broth by centrifuging at 4 °C, 3000×g for 5 min. After being washed twice in sterilized distilled water, cell pellets were suspended in 1 mL 0.1 M HCl. Then 500 μL glass beads with diameter of 0.5 mm were added into the suspension, which was vortexed vigorously for 10 min in a metal bath pre-cooled at −20 °C to disrupt the cells. After being centrifuged at 12,000×g and 4 °C for 5 min, 800 μL supernatant was well mixed with 400 μL 16.7% (w/v) sulfosalicylic acid and incubated at 4 °C for 1 h. The mixture was centrifuged at 15,000×g and 4 °C for 30 min, and the supernatant was collected and neutralized with 1 M NaOH. The supernatant was filtered by 0.22 μm filter before being analyzed. The final amino acid content was normalized by dry cell weight (DCW). The DCW of each sample was determined with twice washed cells incubated at 45 °C for over 48 h.
2.11. Determination of the concentration of spermidine
Dansyl chloride derivatization based HPLC method was adopt for the detection of polyamines, which was modified from a method described previously [36]. In brief, for derivatization, 0.125 mL of saturated NaHCO3 solution and 0.25 mL of dansyl chloride solution (5 mg/mL in acetone) were added to 0.25 mL of sample (dilute samples whenever needed with minimal medium). The samples for determination of the concentration of spermidine was prepared with the powder dried under vacuum from the crude cell extract was prepared with the same method as that for the determination of intracellular amino acid compositions. For each sample, 500 μL of the crude cell extract solution was dried under vacuum, and then re-suspended and diluted with minimal medium whenever needed. After being incubated at 40 °C for 1 h in dark with occasional shaking, the derivatization reaction was stopped by adding 0.275 mL of methanol into the reaction mixture. Then samples filtered through a 25-mm syringe filter (0.45 μm Nylon) were used for HPLC detection. The following chromatographic condition are used: C18 (100 mm × 4.6 mm; i. d., 2.6 μm; Phenomenex Kinetex), excitation wavelength 340 nm, emission wavelength 515 nm, sample injecting 1.5 μL, column temperature 40 °C, detector sensitivity was set at level 7, acquisition starts at 4.0 min. The mobile phase is water and methanol with the speed of 1 mL/min. The elution program is as follows: 50%–65% methanol (0–5 min), 65%–75% methanol (5–7.5 min), 75%–87.5% methanol (7.5–9.5 min), 87.5%–100% methanol (9.5–10.5 min), 100% methanol (10.5–11.5 min), 100%–50% methanol (11.5–13.5 min), 50% methanol (13.5–16 min).
2.12. Statistical analysis
All experiments were performed in triplicates. The results of RT-qPCR, antioxidant enzyme activities, ATP content, and amino acid compositions were expressed as mean values and standard deviations. Statistical analysis was performed using the t-test at the significance: p < 0.05 (*), p < 0.01 (**) and p < 0.001 (***).
3. Results
3.1. CAR1 overexpression enhances acetic acid tolerance of S. cerevisiae
Previous studies showed that CAR1 is regulated by its native promoter comprising multiple regulatory elements that can response to environmental signals [37]. To enable constitutive expression of CAR1, the PGK1 promoter was used to drive its overexpression in this study. After recombinant strains were obtained and verified, their acetic acid tolerance was evaluated and compared with the control strain (Fig. 1).
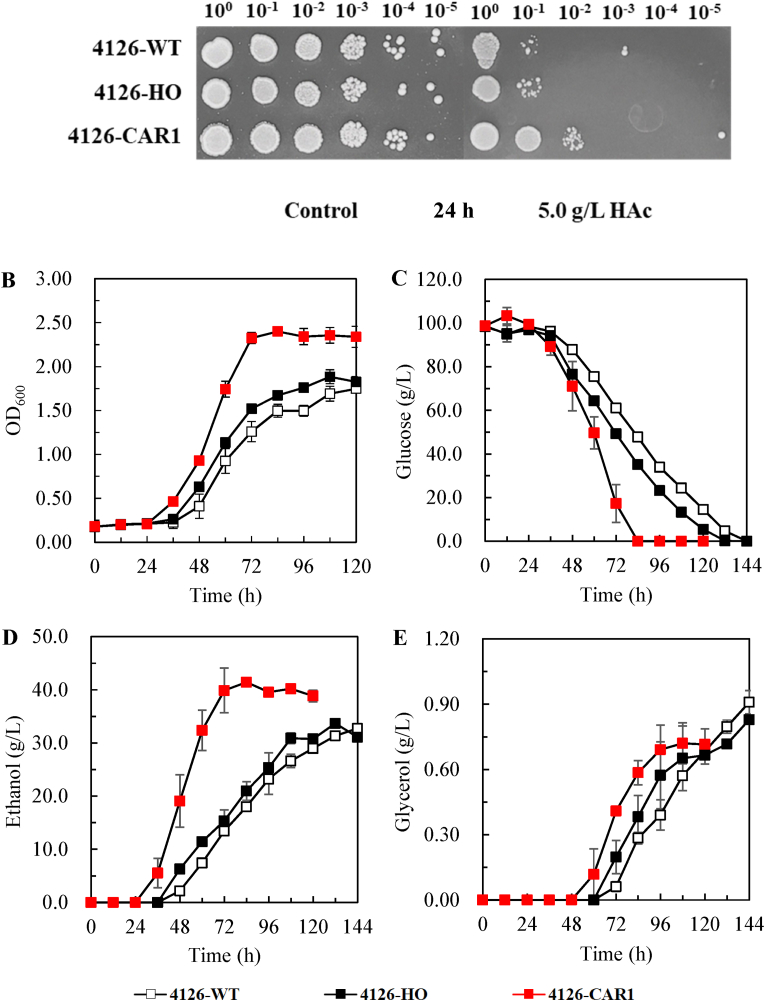
Fig. 1.
Impact of CAR1 overexpression on the growth of S. cerevisiae 4126 strains on YPD agar plates supplemented with 5.0 g/L acetic acid to test their tolerance to the inhibitor and without acetic acid supplementation as the control (A), and ethanol fermentation by S. cerevisiae 4126 engineered with CAR1 overexpression using YPD medium composed of 4 g/L yeast extract, 3 g/L peptone, 100 g/L glucose and 5 g/L acetic acid (B–E). The data for ethanol fermentation were expressed as the means for triplicate with standard deviations. 4126-WT, S. cerevisiae 4126; 4126-HO, S. cerevisiae 4126 engineered with the empty plasmis; 4126-CAR1, S. cerevisiae 4126 engineered with CAR1 overexpression, respectively.
Spot assay showed improved acetic acid tolerance in the CAR1-overexpressing strain S. cerevisiae 4126-CAR1 (4126-CAR1) when comparing with the wild type strain S. cerevisiae 4126 (4126-WT) and the strain carrying the empty vector S. cerevisiae 4126-HO (4126-HO). Without acetic acid stress, no significant difference was observed in the growth of these strains. Similar results were observed when CAR1 overexpression was engineered in S. cerevisiae S288c (Fig. S1A).
Ethanol fermentation was quantitatively evaluated for these strains in the presence of 5.0 g/L acetic acid, and 4126-CAR1 showed improved growth with over 30% higher cell density (Fig. 1B−E). As a result, both glucose consumption and ethanol production of the CAR1 overexpressing strain were facilitated with less glycerol produced, and consequently ethanol productivity was doubled, with ethanol yield increased by 23.5% and 27.3%, respectively compared with 4126-HO and 4126-WT (Table 2). Similar results were observed when CAR1 overexpression was engineered in S. cerevisiae S288c (Figs. S1B–E). Since no significant difference was detected between 4126-WT and 4126-HO, we selected 4126-HO as the only control strain for the following studies. Similar results were observed for ethanol fermentation from the hydrolysate of corn stover pretreated with dilute acid (Fig. S2). The acetic acid concentrations of 4126-WT, 4126-HO, 4126-CAR1 decreased as the fermentation was completed.
Table 2.
Ethanol fermentation performance of S. cerevisiae 4126-WT, 4126-HO and 4126- CAR1 in the presence of 5.0 g/L acetic acid.
| Yeast strain | Fermentation time (h) | Ethanol production (g/L) | Ethanol yield (g/g) | Ethanol productivity (g/L/h) |
|---|---|---|---|---|
| 4126-WT | 144 | 32.73 ± 0.15 | 0.33 ± 0.002 | 0.23 ± 0.002 |
| 4126-HO | 132 | 33.68 ± 0.02 | 0.34 ± 0.000 | 0.26 ± 0.003 |
| 4126-CAR1 | 84 | 40.86 ± 0.58 | 0.42 ± 0.006 | 0.49 ± 0.021 |
3.2. CAR1 overexpression affects intracellular amino acid composition
We are interested in whether such a CAR1 overexpression led to the changes of amino acid metabolism in S. cerevisiae under acetic acid stress, and therefore examined the intracellular amino acid compositions for those strains under conditions with and without such a stress. Totally 26 amino acids and their derivatives were analyzed (Table S2), and the statistical analysis indicated that 24 of them were significantly affected by the CAR1 overexpression (Fig. 2E). While the content of isoleucine, serine, histidine, asparagine and methionine was substantially decreased in S. cerevisiae 4126-CAR1 compared to that detected for S. cerevisiae 4126-HO under the acetic acid stress condition, no such an impact was observed for the CAR1 overexpression strain when no acetic acid stress was applied, suggesting a differential regulation on amino acid metabolism under the stressful condition. We also compared the global amino acid profile of the control strain with or without acetic acid stress, and found that it was different from the case observed in the CAR1-overexpressing strain. Therefore, we speculate that CAR1 overexpression might affect the overall amino acid metabolism of S. cerevisiae under acetic acid stress conditions.
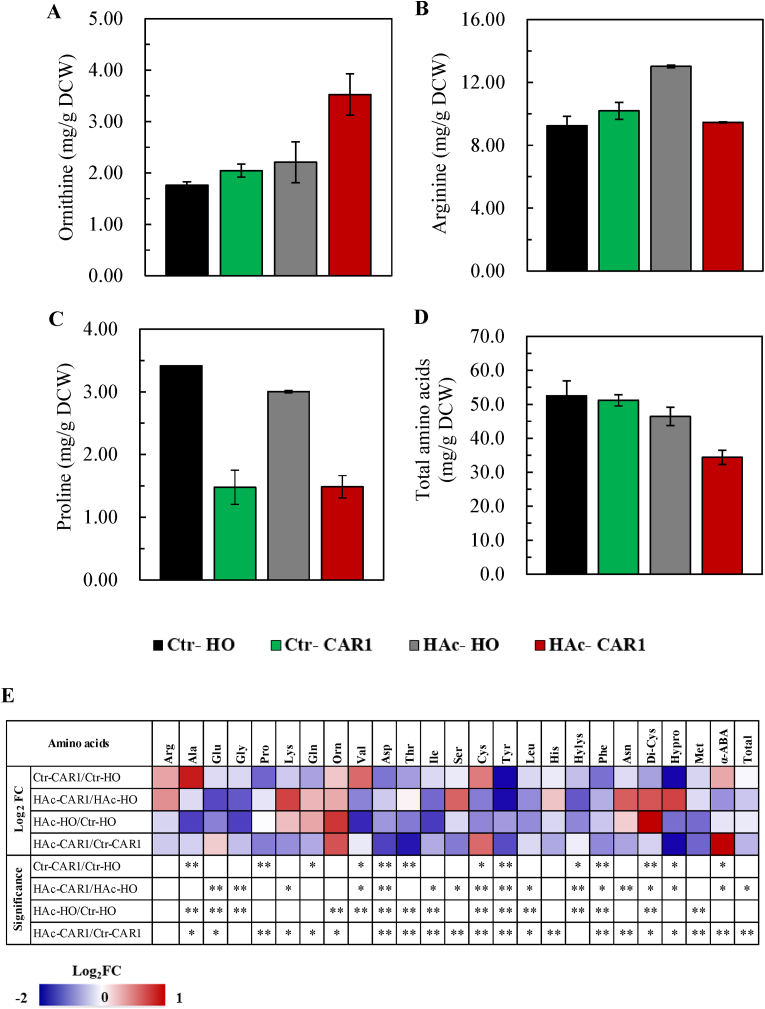
Fig. 2.
An overview on the profile of amino acid metabolism (A, B, C and D) and the heat map (E) highlighting the fold changes of intracellular amino acids. Ctr-CAR1 and Ctr-HO represent S. cerevisiae 4126 engineered with CAR1 overexpression and the empty plasmid cultured in the YPD medium without acetic acid supplementation, and HAc-CAR1 and HAc-HO represent those two recombinat strains cultivated Detailed data for amino acids are available in Table S2. Abbreviations: Ala, alanine; Arg, arginine; Asn, asparaginate; Asp, asparate; Cys, cysteine; Di-Cys, dicyseine; Gln, glutamine; Glu, glutamate; Gly, glycine; His, histidine; Hylys, hydroxylysine; Hypro, hydroxyproline; Ile, isoleusine; Lys, lysine; Leu, leucine; Met, methionine; Orn, ornithine; Phe, phenylalanine; Pro, proline; Ser, serine; Thr, thronine; Tyr, tyrosine; Val, valine; α-ABA, α-aminobutyric acid. Yeast cells were harvested at their exponential growth phase for the analysis with triplicate under the statistic significance *p < 0.05 and **p < 0.01.
Analysis of intracellular amino acids revealed that ornithine was increased in S. cerevisiae 4126-CAR1 under the acetic acid stress condition (HAc-CAR1) compared to that detected with the control strain (HAc-HO), but the accumulation of arginine and proline was decreased by 27.3% and 50.6%, respectively, making the total amino acids reduced as high as 25.9% (34.38 mg/gDCW v.s. 46.42 mg/gDCW (Fig. 2A−D and Table S2). On the other hand, no significant difference was observed in ornithine, arginine and total amino acids between S. cerevisiae 4126-CAR1 (Ctr-CAR1) and S. cerevisiae 4126-HO (Ctr-HO) when no acetic acid stress was applied, and the content of proline was decreased by the overexpression of CAR1.
The results highlighted in Fig. 2 were detected for yeast cells cultured in the rich medium supplemented with yeast extract and peptone for laboratory research, and amino acids were available for potential uptake by the yeast cells, which might affect the profile of their intracellular amino acids. We therefore performed culture using the minimal medium without amino acids to rule out such a concern (Fig. S3 and Table S3). The same profiles were obtained, confirming that the effect of CAR1 overexpression on yeast acetic acid tolerance was independent on the uptake of amino acids.
Notably, under the acetic acid stress condition, ornithine content in S. cerevisiae 4126-CAR1 (HAc-CAR1) increased by about 60.0% when compared with that in the control strain (HAc-HO). However, unexpectedly, the contribution of increased ornithine content to the growth of S. cerevisiae was negligible (Fig. S4A, B and E), despite the fact that the supplementation of ornithine at 10 mM increased the intracellular ornithine content by 442% (Fig. S4C), respectively. The cell viability was not significantly affected by supplementation of ornithine at 10 mM as well (Fig. S4F). We therefore conclude that CAR1 overexpression in S. cerevisiae improves acetic acid tolerance through mechanisms other than affecting ornithine content. We did not find significant changes in acetic acid concentration when comparing the CAR1 overexpressing strain with that of the control strain. Therefore, the improved growth is not related to elevated acetyl-CoA by acetic acid utilization.
3.3. Transcriptional profiling of genes involved in ornithine, arginine and proline metabolism influenced by CAR1 overexpression
Since CAR1 encodes arginase for arginine degradation, we first quantified the relative transcription of genes related to arginine catabolism. As can be seen in Fig. 3A and S5A, no matter whether or not acetic acid stress was applied, the transcription of CAR1 was enhanced in S. cerevisiae 4126-CAR1, which perturbed the expression of genes involved in arginine catabolism. Since arginine degradation is the major upstream pathway for ornithine synthesis, the CAR1 overexpression enhanced ornithine anabolism, and consequently intracellular accumulation of ornithine was observed, no matter whether the acetic acid stress was applied (Table S2).
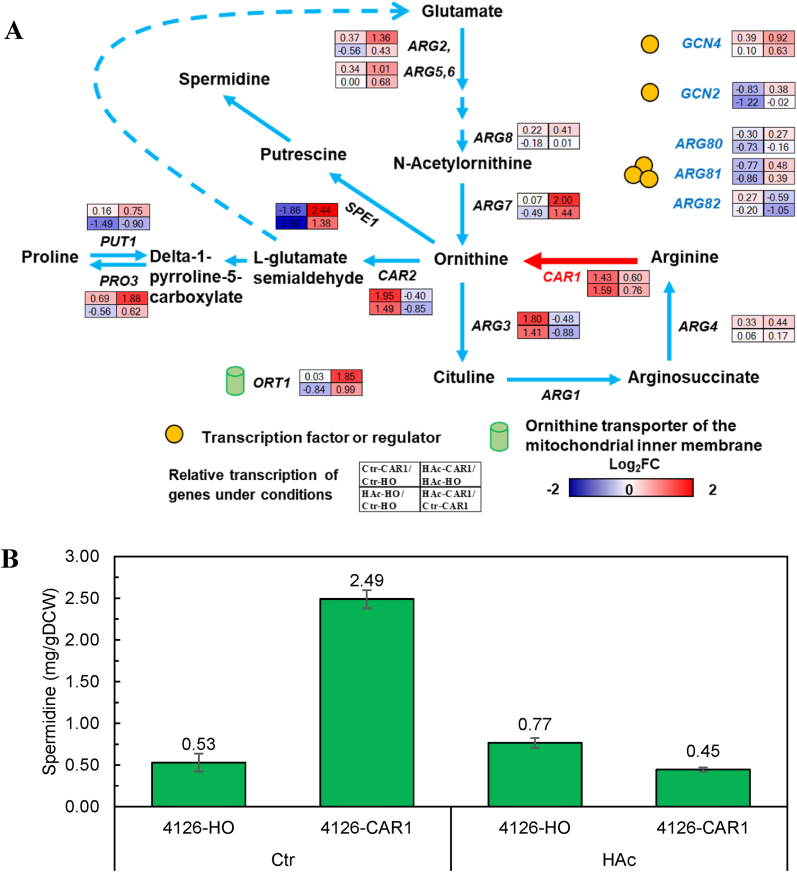
Fig. 3.
Transcription of genes related to the arginine metabolism (A) and intracellular spermidine contents of the recombinant strains (B). Relative gene expression was determined by the 2−ΔΔCt method using the expression of ALG9 as the internal control, which is highlighted by the fold change (Log2FC). Pathways for arginine metabolism was constructed based on information from the KEGG Pathway (http://www.genome.jp/kegg/). Intracellular spermidine contents of the recombinat strains were determined dansyl chloride derivatization based HPLC method as described in the Material and Method section. Yeast cells grew to exponential-phase in fermentation medium were harvest and used for the determination of transcription and intracellular spermidine contents.
When acetic acid stress was applied, the overexpression of CAR1 in 4126-CAR1 was compromised, which consequently affected transcription of other genes. Comparing 4126-CAR1 with 4126-HO, the transcription of ornithine decarboxylase gene SPE1 and mitochondrial ornithine acetyltransferase encoding gene ARG7 was significantly enhanced, while the expression of ornithine transaminase gene CAR2 and ornithine carbamoyltransferase (OTCase) encoding gene AGR3 were down-regulated. Although the transcription of P5C reductase encoding gene PRO3 was enhanced, it wouldn't contribute substantially to ornithine catabolism to proline due to the down-regulation of CAR2. As a result, the accumulation of proline was suppressed since about 50% lower proline was detected in 4126-CAR1 than that in 4126-HO.
3.4. Acetic acid tolerance rendered by CAR1 overexpression is independent on the spermidine metabolism
Previous studies demonstrated that the elevation of polyamine spermidine level in S. cerevisiae could improve tolerance to various lignocellulose-derived inhibitors [36,38]. Moreover, another study revealed that yeast cells could cope with stress by activating polyamine metabolism [39,40]. In this study, the expression level of SPE1, the enzyme responsible for the initial reaction in polyamine synthesis, was quantified in this study. The results showed that SPE1 transcription was upregulated by 4.43-fold in 4126-CAR1 compared to 4126-HO. (Fig. 3A). As mentioned earlier, ornithine is the essential substrate for polyamine biosynthesis. In this study, we measured the ornithine content in S. cerevisiae 4126-CAR1 (HAc-CAR1) and found that it was approximately 60.0% higher than that in the control strain (HAc-HO). Based on this observation, we hypothesized that the increased production of polyamines, particularly spermidine, could be the underlying mechanism for acetic acid tolerance. We quantified the spermidine content of 4126-CAR1 and 4126-HO under acetic acid stress. Contrary to our expectation, spermidine content in 4126-CAR1 (0.45 mg/gDCW) was marginally lower than that in 4126-HO (0.77 mg/gDCW) (Fig. 3B). These results indicated that the enhanced acetic acid tolerance by CAR1-overexpression was not related to the elevated spermidine level. Therefore, we proposed that a novel mechanism, distinct from the previously reported spermidine-independent pathway [36,38], was involved in acetic acid resistance upon CAR1 overexpression.
3.5. CAR1 overexpression increase ATP content and enhance cell membrane integrity
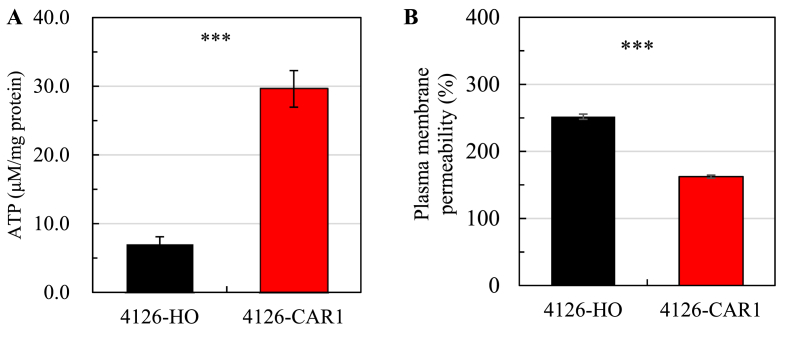
In response to acetic acid stress, cytoplasmic membrane proton-pumping ATPases and vacuolar proton-pumping ATPase are employed to pump protons out at the expense of ATP consumption [41]. Hence, it is of great significance to maintain relatively high ATP content to resist acetic acid stress. The intracellular ATP content in S. cerevisiae 4126-CAR1 was 3.23-fold higher than that detected in S. cerevisiae 4126-HO under the acetic acid stress condition (Fig. 4A), indicating that the CAR1 overexpression might be conducive to maintain the high level of intracellular ATP in S. cerevisiae.
Fig. 4.
Intracellular ATP (A) and plasma membrane permeability (B) for S. cerevisiae 4126-CAR1 and S. cerevisiae 4126-HO. The yeast cells were harvested at their exponential growth phase at 36 h and 48 h, respectively, with their OD600 of 0.46 and 0.63 for the analysis with triplicate under the statistical significance ***p < 0.001.
Cell membranes are important barrier against environmental stresses. It was reported that good membrane integrity contributed to maintaining cell activity under acetic acid conditions [22,42]. Plasma membrane permeability characterized by the propidium iodide uptake significantly decreased in S. cerevisiae 4126-CAR1 compared to that detected with S. cerevisiae 4126-HO under the acetic acid stress condition (Fig. 4B), suggesting a less damage in its cell membranes. The transcription of genes involved in ergosterol biosynthesis was quantified to explore molecular mechanism underlying this phenomenon, and the expression of ERG3, ERG4, ERG6 and ERG28, was significantly up-regulated in S. cerevisiae 4126-CAR1 compared to that observed in S. cerevisiae 4126-HO regardless of the presence of acetic acid (Fig. S5C). Therefore, the CAR1 overexpression in S. cerevisiae enhanced ergosterol biosynthesis either directly or indirectly, which consequently improved its cell membrane integrity [43].
4. Discussion
In this study, we observed remarkable improved acetic acid tolerance by CAR1 overexpression in S. cerevisiae, which provides alternative strategy for yeast strain development for efficient cellulosic biorefinery. Previous studies mostly focused on the effect of individual amino acids on the stress tolerance of S. cerevisiae [25,[44], [45], [46]]. However, we observed significant decrease of total amino acid content by CAR1 overexpression under acetic acid stress, which was not observed in the non-stressed condition. Our report is the first to reveal decreased total amino acid content under stress condition by overexpression a single gene CAR1. It was reported that single gene deletion influences the global amino acid metabolome of yeast cells [47], and our experimental results further revealed different changes of amino acid composition under stress condition comparing with stress free condition.
Our current studies also revealed that overexpression of CAR1 regulates acetic acid tolerance through mechanisms different from reported ones. Both arginine and proline are known protectants for yeast cells to tolerate various stresses. It was reported that improved intracellular arginine increased ethanol tolerance of S. cerevisiae YS58 [46], and increased proline improved tolerance to multiple lignocellulose-derived inhibitors including furfural, acetic acid and phenol [45]. Decreased ornithine as well as proline content was observed when metabolomics analysis was performed to study the adaptation of an industrial yeast strain in a mixture of inhibitors containing acetic acid, furfural and phenol [48]. Although ornithine metabolism is closely related to arginine and proline, significantly decreased arginine and proline contents comparing to that detected in the control strain were observed in S. cerevisiae 4126-CAR1 under the acetic acid stress condition in this work, suggesting different mechanisms from that of proline in regulating acetic acid tolerance by CAR1 overexpression.
It was suggested that the accumulation of spermidine in S. cerevisiae enhanced tolerance to multiple lignocellulose-derived inhibitors [36,38]. However, spermidine content in CAR1 overexpression strain decreased by 58.4% when compared with the control strain in the presence, indicating a mechanism significantly different from the previous studies [36,38]. Intracellular accumulation of amino acids is a dynamic balance between their uptake from medium and metabolism through synthesis and degradation, particularly when rich medium with various amino acids is used. In order to address such a concern, both S. cerevisiae 4126-CAR1 and S. cerevisiae 4126-HO were cultured with the minimal medium without amino acids supplemented (Table S3). Although data with individual amino acids are different, we confirmed that the CAR1 overexpression still showed beneficial effects on growth of S. cerevisiae under acetic acid stress. These results suggest that the effects of CAR1 overexpression is not dependent on uptake of amino acids in the culture medium, at least under the conditions employed in this study.
Consistent with the changes in amino acid contents, changes in transcription of genes involved in amino acid translocation as well as arginine metabolism also showed significant differences, suggesting regulation of amino acid metabolism gene expression by CAR1. Expression of genes for arginine metabolism is regulated by the multimeric complex composed of ArgR encoded by ARG80, ARG81 and ARG82 with their expression regulated by the transcription factor encoded by GCN4 that can be activated by a protein kinase encoded by GCN2 [44]. The expression of these genes was also regulated in S. cerevisiae 4126-CAR1 under the acetic acid stress condition to affect its amino acid metabolism. In addition, amino acid metabolism regulated at post-translational level also contributed to the equivalence of intracellular amino acid content [44]. More efforts are needed to elucidate the direct mechanism relating to modulation of amino acid by CAR1 overexpression for improved acetic acid tolerance in the future.
Compared to the control, a higher intracellular ATP content was observed in S. cerevisiae 4126-CAR1. Under acid stress, yeast cells pump both the proton and the anion out of the cell at the expense of ATP, and studies have found that increasing acid stress resulted in higher cellular ATP levels [49], suggesting that higher ATP levels may increase yeast tolerance to acid stress. Our results are consistent with this hypothesis. Since biosynthesis of arginine, proline is energy-consuming, we thus propose that at least the partial mechanism underlying improved acetic acid tolerance in the CAR1 overexpressing strain is related to repressing biosynthesis of certain amino acids to elevate its intracellular ATP level, which benefits tolerance to acetic acid as reported [13,49]. In addition, we also observed improved integrity for cell membranes by CAR1 overexpression, which added evidence in the effect of CAR1 in modulating membrane lipid metabolism. Although it is difficult to distinguish direct impacts from indirect ones, our results reveal possible link between overexpression of CAR1 and metabolism of total amino acid, ATP and lipid to maintain high viability of S. cerevisiae under acetic acid stress, and therefore improve its ethanol fermentation performance.
In conclusion, we confirmed that CAR1 overexpression in S. cerevisiae improve its tolerance to acetic acid and thereby boosts ethanol production. Our results also suggest that the underlining mechanism are different from the known ones related to proline and spermidine, and might related to decreased total amino acid metabolism for the engineered strain, particularly the regulation on its biosynthesis of amino acids for reducing their intracellular accumulation properly to save energy in the form of ATP. Nevertheless, follow-up studies warrant elucidating the direct mechanism for the improved stress tolerance. Our studies also suggest that CAR1 is a novel genetic element to be used in synthetic biology of yeast for efficient production of fuel ethanol.
Funding information
This work was supported financially by National Key Research and Development Program (No. 2022YFE0108500) and National Natural Science Foundation of China (No. 21978168) to X.-Q.Z. JFQ appreciates the grant from Sichuan Natural Science Foundation (No. 2023NSFSC0132).
CRediT authorship contribution statement
Liang Xiong: carried out the major part of the experiments and drafted the manuscript, designed the experiments. Ya-Ting Wang: revised the manuscript. Ming-Hai Zhou: included some additional experiments and refined the manuscript. Hiroshi Takagi: were involved in the data analysis and discussion, revised the manuscript. Jiu-Fu Qin: designed the experiments, were involved in the data analysis and discussion, revised the manuscript. Xin-Qing Zhao: conceived the study, designed the experiments, were involved in the data analysis and discussion, revised the manuscript.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgement
The authors appreciate assistance from the former graduate Keyu Zhang, Xiaowen Wei, Dr. Qian Li and Dr. Mingming Zhang in the experimental work.
Footnotes
Peer review under responsibility of KeAi Communications Co., Ltd.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.synbio.2024.05.013.
Contributor Information
Jiufu Qin, Email: jiufuq@scu.edu.cn.
Xin-Qing Zhao, Email: xqzhao@sjtu.edu.cn.
Appendix A. Supplementary data
The following is the Supplementary data to this article.
References
- 1.Nielsen J. Yeast cell factories on the horizon. Science. 2015;349(6252):1050–1051. doi: 10.1126/science.aad2081. [DOI] [PubMed] [Google Scholar]
- 2.Antar M., Lyu D.M., Nazari M., Shah A.T., Zhou X.M., Smith D.L. Biomass for a sustainable bioeconomy: an overview of world biomass production and utilization. Renew Sustain Energy Rev. 2021;139(18) doi: 10.1016/j.rser.2020.110691. [DOI] [Google Scholar]
- 3.Srivastava R.K., Shetti N.P., Reddy K.R., Kwon E.E., Nadagouda M.N., Aminabhavi T.M. Biomass utilization and production of biofuels from carbon neutral materials. Environ Pollut. 2021;276(25) doi: 10.1016/j.envpol.2021.116731. [DOI] [PubMed] [Google Scholar]
- 4.Jonsson L.J., Alriksson B., Nilvebrant N.O. Bioconversion of lignocellulose: inhibitors and detoxification. Biotechnol Biofuels. 2013;6(10):16. doi: 10.1186/1754-6834-6-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhao X.Q., Xiong L., Zhang M.M., Bai F.W. Towards efficient bioethanol production from agricultural and forestry residues: exploration of unique natural microorganisms in combination with advanced strain engineering. Bioresour Technol. 2016;215:84–91. doi: 10.1016/j.biortech.2016.03.158. [DOI] [PubMed] [Google Scholar]
- 6.Bhatia S.K., Jagtap S.S., Bedekar A.A., et al. Recent developments in pretreatment technologies on lignocellulosic biomass: effect of key parameters, technological improvements, and challenges. Bioresour Technol. 2020;300(13) doi: 10.1016/j.biortech.2019.122724. [DOI] [PubMed] [Google Scholar]
- 7.Wang S.Z., Sun X.X., Yuan Q.P. Strategies for enhancing microbial tolerance to inhibitors for biofuel production: a review. Bioresour Technol. 2018;258:302–309. doi: 10.1016/j.biortech.2018.03.064. [DOI] [PubMed] [Google Scholar]
- 8.Yuan B., Wang W.B., Wang Y.T., Zhao X.Q. Regulatory mechanisms underlying yeast chemical stress response and development of robust strains for bioproduction. Curr Opin Biotechnol. 2024;86 doi: 10.1016/j.copbio.2024.103072. [DOI] [PubMed] [Google Scholar]
- 9.Giannattasio S., Guaragnella N., Zdralevic M., Marra E. Molecular mechanisms of Saccharomyces cereyisiae stress adaptation and programmed cell death in response to acetic acid. Front Microbiol. 2013;4(7):33. doi: 10.3389/fmicb.2013.00033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Guaragnella N., Bettiga M. Acetic acid stress in budding yeast: from molecular mechanisms to applications. Yeast. 2021;38(7):391–400. doi: 10.1002/yea.3651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ding J., Bierma J., Smith M.R., et al. vol. 97. 2013. p. 7416. (Acetic acid inhibits nutrient uptake in Saccharomyces cerevisiae: auxotrophy confounds the use of yeast deletion libraries for strain improvement). 16. [DOI] [PubMed] [Google Scholar]
- 12.Chaves S.R., Rego A., Martins V.M., Santos-Pereira C., Sousa M.J., Corte-Real M. Regulation of cell death induced by acetic acid in yeasts. Front Cell Dev Biol. 2021;9(20) doi: 10.3389/fcell.2021.642375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhang M.M., Zhao X.Q., Cheng C., Bai F.W. Improved growth and ethanol fermentation of Saccharomyces cerevisiae in the presence of acetic acid by overexpression of SET5 and PPR1. Biotechnol J. 2015;10(12):1903–1911. doi: 10.1002/biot.201500508. [DOI] [PubMed] [Google Scholar]
- 14.Camara E., Olsson L., Zrimec J., Zelezniak A., Geijer C., Nygard Y. Data mining of Saccharomyces cerevisiae mutants engineered for increased tolerance towards inhibitors in lignocellulosic hydrolysates. Biotechnol Adv. 2022;57(14) doi: 10.1016/j.biotechadv.2022.107947. [DOI] [PubMed] [Google Scholar]
- 15.Songdech P., Ruchala J., Semkiv M.V., et al. Overexpression of transcription factor ZNF1 of glycolysis improves bioethanol productivity under high glucose concentration and enhances acetic acid tolerance of Saccharomyces cerevisiae. Biotechnol J. 2020;15(7):10. doi: 10.1002/biot.201900492. [DOI] [PubMed] [Google Scholar]
- 16.Li B., Wang L., Wu Y.J., Xia Z.Y., Yang B.X., Tang Y.Q. Improving acetic acid and furfural resistance of xylose-fermenting Saccharomyces cerevisiae strains by regulating novel transcription factors revealed via comparative transcriptomic analysis. Appl Environ Microbiol. 2021;87(10) doi: 10.1128/aem.00158-21. 15. 21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tanaka K., Ishii Y., Ogawa J., Shima J. Enhancement of acetic acid tolerance in Saccharomyces cerevisiae by overexpression of the HAA1 gene, encoding a transcriptional activator. Appl Environ Microbiol. 2012;78(22):8161–8163. doi: 10.1128/aem.02356-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ye P.L., Wang X.Q., Yuan B., Liu C.G., Zhao X.Q. Manipulating cell flocculation-associated protein kinases in Saccharomyces cerevisiae enables improved stress tolerance and efficient cellulosic ethanol production. Bioresour Technol. 2022;348(11) doi: 10.1016/j.biortech.2022.126758. [DOI] [PubMed] [Google Scholar]
- 19.Chen H.Q., Xing Q., Cheng C., et al. Identification of Kic1p and Cdc42p as novel targets to engineer yeast acetic acid stress tolerance. Front Bioeng Biotechnol. 2022;10(12) doi: 10.3389/fbioe.2022.837813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cheng Y.F., Zhu H., Du Z.D., et al. Eukaryotic translation factor eIF5A contributes to acetic acid tolerance in Saccharomyces cerevisiae via transcriptional factor Ume6p. Biotechnol Biofuels. 2021;14(1) doi: 10.1186/s13068-021-01885-2. 13. 38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.de Kok S., Nijkamp J.F., Oud B., et al. Laboratory evolution of new lactate transporter genes in a jen1Δ mutant of Saccharomyces cerevisiae and their identification as ADY2 alleles by whole-genome resequencing and transcriptome analysis. FEMS Yeast Res. 2012;12(3):359–374. doi: 10.1111/j.1567-1364.2011.00787.x. [DOI] [PubMed] [Google Scholar]
- 22.Zhang M.M., Zhang K.Y., Mehmood M.A., Zhao Z.K., Bai F.W., Zhao X.Q. Deletion of acetate transporter gene ADY2 improved tolerance of Saccharomyces cerevisiae against multiple stresses and enhanced ethanol production in the presence of acetic acid. Bioresour Technol. 2017;245:1461–1468. doi: 10.1016/j.biortech.2017.05.191. [DOI] [PubMed] [Google Scholar]
- 23.Ismail K.S.K., Sakamoto T., Hasunuma T., Zhao X.Q., Kondo A. Zinc, magnesium, and calcium ion supplementation confers tolerance to acetic acid stress in industrial Saccharomyces cerevisiae utilizing xylose. Biotechnol J. 2014;9(12):1519–1525. doi: 10.1002/biot.201300553. [DOI] [PubMed] [Google Scholar]
- 24.Ko J.K., Um Y., Lee S.M. Effect of manganese ions on ethanol fermentation by xylose isomerase expressing Saccharomyces cerevisiae under acetic acid stress. Bioresour Technol. 2016;222:422–430. doi: 10.1016/j.biortech.2016.09.130. [DOI] [PubMed] [Google Scholar]
- 25.Greetham D., Takagi H., Phister T.P. Presence of proline has a protective effect on weak acid stressed Saccharomyces cerevisiae. Antonie Leeuwenhoek. 2014;105(4):641–652. doi: 10.1007/s10482-014-0118-3. [DOI] [PubMed] [Google Scholar]
- 26.Takagi H. Metabolic regulatory mechanisms and physiological roles of functional amino acids and their applications in yeast. Biosci Biotechnol Biochem. 2019;83(8):1449–1462. doi: 10.1080/09168451.2019.1576500. [DOI] [PubMed] [Google Scholar]
- 27.Mukai Y., Kamei Y., Liu X., et al. Proline metabolism regulates replicative lifespan in the yeast Saccharomyces cerevisiae. Microb Cell. 2019;6(10):482–490. doi: 10.15698/mic2019.10.694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chu C., Zong H., Zhuge B., Lu X.Y. Effect and application of proline metabolism-related gene CgMCUR1 on stress tolerance of Candida glycerinogenes and Saccharomyces cerevisiae. J Appl Microbiol. 2023;134(6) doi: 10.1093/jambio/lxad108. 10. [DOI] [PubMed] [Google Scholar]
- 29.Wan C., Zhang M.M., Fang Q., et al. The impact of zinc sulfate addition on the dynamic metabolic profiling of Saccharomyces cerevisiae subjected to long term acetic acid stress treatment and identification of key metabolites involved in the antioxidant effect of zinc. Metallomics. 2015;7(2):322–332. doi: 10.1039/c4mt00275j. [DOI] [PubMed] [Google Scholar]
- 30.Ma C., Wei X.W., Sun C.H., et al. Improvement of acetic acid tolerance of Saccharomyces cerevisiae using a zinc-finger-based artificial transcription factor and identification of novel genes involved in acetic acid tolerance. Appl Microbiol Biotechnol. 2015;99:2441–2449. doi: 10.1007/s00253-014-6343-x. [DOI] [PubMed] [Google Scholar]
- 31.Qin J.F., Zhou Y.J.J., Krivoruchko A., et al. Modular pathway rewiring of Saccharomyces cerevisiae enables high-level production of L-ornithine. Nat Commun. 2015;6(11):8224. doi: 10.1038/ncomms9224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.He L.Y., Zhao X.Q., Bai F.W. Engineering industrial Saccharomyces cerevisiae strain with the FLO1-derivative gene isolated from the flocculating yeast SPSC01 for constitutive flocculation and fuel ethanol production. Appl Energy. 2012;100:33–40. doi: 10.1016/j.apenergy.2012.03.052. [DOI] [Google Scholar]
- 33.Manivasakam P., Schiestl R.H. High efficiency transformation of Saccharomyces cerevisiae by electroporation. Nucleic Acids Res. 1993;21(18):4414–4415. doi: 10.1093/nar/21.18.4414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Teste M.A., Duquenne M., Francois J.M., Parrou J.L. Validation of reference genes for quantitative expression analysis by real-time RT-PCR in Saccharomyces cerevisiae. BMC Mol Biol. 2009;10(15):99. doi: 10.1186/1471-2199-10-99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Fang Z.J., Chen Z.X., Wang S., et al. Overexpression of OLE1 enhances cytoplasmic membrane stability and confers resistance to cadmium in Saccharomyces cerevisiae. Appl Environ Microbiol. 2017;83(1) doi: 10.1128/aem.02319-16. 11. 16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kim S.K., Jin Y.S., Choi I.G., Park Y.C., Seo J.H. Enhanced tolerance of Saccharomyces cerevisiae to multiple lignocellulose-derived inhibitors through modulation of spermidine contents. Metab Eng. 2015;29:46–55. doi: 10.1016/j.ymben.2015.02.004. [DOI] [PubMed] [Google Scholar]
- 37.Smart W.C., Coffman J.A., Cooper T.G. Combinatorial regulation of the Saccharomyces cerevisiae CAR1 (arginase) promoter in response to multiple environmental signals. Mol Cell Biol. 1996;16(10):5876–5887. doi: 10.1128/MCB.16.10.5876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kim S.K., Jo J.H., Jin Y.S., Seo J.H. Enhanced ethanol fermentation by engineered Saccharomyces cerevisiae strains with high spermidine contents. Bioproc Biosyst Eng. 2017;40(5):683–691. doi: 10.1007/s00449-016-1733-3. [DOI] [PubMed] [Google Scholar]
- 39.Kim S.K., Auh J.H. Evaluating the engineered Saccharomyces cerevisiae with high spermidine contents for increased tolerance to lactic, succinic, and malic acids and increased xylose fermentation. Biotechnol Bioproc Eng. 2021;26(1):47–54. doi: 10.1007/s12257-020-0020-y. [DOI] [Google Scholar]
- 40.Olin-Sandoval V., Yu J.S.L., Miller-Fleming L., et al. Lysine harvesting is an antioxidant strategy and triggers underground polyamine metabolism. Nature. 2019;572(7768):249. doi: 10.1038/s41586-019-1442-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Martinez-Munoz G.A., Kane P. Vacuolar and plasma membrane proton pumps collaborate to achieve cytosolic pH homeostasis in yeast. J Biol Chem. 2008;283(29):20309–20319. doi: 10.1074/jbc.M710470200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Zheng D.Q., Wu X.C., Wang P.M., et al. Drug resistance marker-aided genome shuffling to improve acetic acid tolerance in Saccharomyces cerevisiae. J Ind Microbiol Biotechnol. 2011;38(3):415–422. doi: 10.1007/s10295-010-0784-8. [DOI] [PubMed] [Google Scholar]
- 43.Ferraz L., Vorauer-Uhl K., Sauer M., Sousa M.J., Branduardi P. Impact of ergosterol content on acetic and lactic acids toxicity to Saccharomyces cerevisiae. Yeast. 2023;40(3–4):152–165. doi: 10.1002/yea.3828. [DOI] [PubMed] [Google Scholar]
- 44.Ljungdahl P.O., Daignan-Fornier B. Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics. 2012;190(3):885–929. doi: 10.1534/genetics.111.133306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wang X., Bai X., Chen D.F., Chen F.Z., Li B.Z., Yuan Y.J. Increasing proline and myo-inositol improves tolerance of Saccharomyces cerevisiae to the mixture of multiple lignocellulose-derived inhibitors. Biotechnol Biofuels. 2015;8(13):142. doi: 10.1186/s13068-015-0329-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Cheng Y.F., Du Z.L., Zhu H., Guo X.N., He X.P. Protective effects of arginine on Saccharomyces cerevisiae against ethanol stress. Sci Rep. 2016;6(12) doi: 10.1038/srep31311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mulleder M., Calvani E., Alam M.T., et al. Functional metabolomics describes the yeast biosynthetic regulome. Cell. 2016;167(2):553. doi: 10.1016/j.cell.2016.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Wang X., Li B.Z., Ding M.Z., Zhang W.W., Yuan Y.J. Metabolomic analysis reveals key metabolites related to the rapid adaptation of Saccharomyce cerevisiae to multiple inhibitors of furfural, acetic acid, and phenol. OMICS. 2013;17(3):150–159. doi: 10.1089/omi.2012.0093. [DOI] [PubMed] [Google Scholar]
- 49.Guo Z.P., Olsson L. Physiological response of Saccharomyces cerevisiae to weak acids present in lignocellulosic hydrolysate. FEMS Yeast Res. 2014;14(8):1234–1248. doi: 10.1111/1567-1364.12221. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.