Abstract
Ion channels are critical in enabling ion movement into and within cells and are important targets for pharmacological interventions in different human diseases. In addition to their ion transport abilities, ion channels interact with signalling and scaffolding proteins, which affects their function, cellular positioning, and links to intracellular signalling pathways. The study of “channelosomes” within cells has the potential to uncover their involvement in human diseases, although this field of research is still emerging. LRRC8A is the gene that encodes a crucial protein involved in the formation of volume-regulated anion channels (VRACs). Some studies suggest that LRRC8A could be a valuable prognostic tool in different types of cancer, serving as a biomarker for predicting patients’ outcomes. LRRC8A expression levels might be linked to tumour progression, metastasis, and treatment response, although its implications in different cancer types can be varied. Here, publicly accessible databases of cancer patients were systematically analysed to determine if a correlation between VRAC channel expression and survival rate exists across distinct cancer types. Moreover, we re-evaluated the impact of LRRC8A on cellular proliferation and migration in colon cancer via HCT116 LRRC8A-KO cells, which is a current topic of debate in the literature. In addition, to investigate the role of LRRC8A in cellular signalling, we conducted biotin proximity-dependent identification (BioID) analysis, revealing a correlation between VRAC channels and cell-cell junctions, mechanisms that govern cellular calcium homeostasis, kinases, and GTPase signalling. Overall, this dataset improves our understanding of LRRC8A/VRAC and explores new research avenues while identifying promising therapeutic targets and promoting inventive methods for disease treatment.
Subject terms: Chloride channels, Protein-protein interaction networks
Introduction
Ion channels mediate the movement of ions across cellular membranes and play important roles in the development and progression of several human diseases [1–4]. Channel proteins are complex structures consisting of a central ion-selective pore and various interacting proteins. Channel-interacting proteins (CIPs) can play a critical role in regulating biophysical properties, such as permeability and gating, or act as signalling and scaffolding proteins that influence the interaction of ion channels with upstream and downstream cellular signalling pathways [1–3, 5, 6].
The increasing knowledge of ion channels as macromolecular signalling complexes is transforming research methods in this field. Although high-throughput gene profiling techniques are commonly employed, studying the ion channel interactome, or channelosome, can reveal interconnected networks that affect not only ion channel activity but also complex signalling pathways. This approach is crucial in identifying groups of molecules that could act as targets for interventions aiming to manage or ameliorate pathological conditions. Additionally, it is essential to acknowledge that network-based approaches provide a more nuanced comprehension of the complex pathways involved in disease.
The identification of protein-protein interactions (PPIs) within a cellular context presents significant challenges, as conventional methods often prove ineffective in a natural cellular environment. Classical approaches may fail to ensure the detection of interactions involving weak or transient interactors, as well as those subject to spatio-temporal regulation. These limitations result in a substantial loss of valuable information, stemming from an incomplete understanding of the dynamic nature of these interactions. Initially, research in CIPs field mainly focused on cation channels, particularly those selectively permeable to calcium, potassium, and sodium ions. Nevertheless, in recent years, there has been a growing interest in anion channels, i.e., chloride (Cl) channels. These channels play a vital role in various cellular processes and their abnormal expression and/or function is associated with various human diseases, such as cystic fibrosis, myotonia, epilepsy, hyperekplexia, lysosomal storage diseases, deafness, renal salt wasting, kidney stones, osteopetrosis and numerous tumour development of numerous types of tumours [7–9].
Among anion channels, the volume-regulated anion channel (VRAC) is emerging as a promising pharmacological target in human pathology and oncology. VRAC has a broad permeability, enabling the transfer of Cl- and other anions, organic compounds, neurotransmitters, taurine, and signalling molecules [10–15]. VRAC plays a critical role in regulating cell volume by reducing it through a process called regulatory volume decrease (RVD) and maintaining cell volume homeostasis. Due to its role in RVD, VRAC has been proposed to be involved in cell volume changes during different aspects of cancer cell behaviour and response to therapies. The activity of VRAC has been linked to cancer cell proliferation, metastasis, and multidrug resistance [11, 16]. Nonetheless, the biological function and prognostic value of the gene encoding the pore-forming subunit of VRAC, LRRC8, require better delineation, as contrasting results have been reported.
In this work, we examined the publicly accessible TCGA database to comprehensively analyse the relationship between LRRC8 expression in cancer and patient survival. We focused specifically on the colon cancer context.
Despite the functional characterization of VRAC has been known for several decades, its molecular identity has been unveiled only recently [17, 18]. These investigations shed light on VRAC as a heteromeric assembly that consists of subunits belonging to the LRRC8 gene family, comprising of five distinct members (LRRC8A-E) [17, 18]. Evolutionary, LRRC8 proteins are formed by combining a pannexin-like transmembrane protein and an intracellular leucine-rich repeat domain (LRRD) [19]. LRRC8 proteins are made up of four transmembrane segments and a C-terminal leucine-rich repeat domain. The N-terminal domain was recently found to fold back into the pore from the cytoplasm, taking part in determining ion selectivity and possibly gating [20]. The precise proportions and arrangement of subunits needed to create functional LRRC8 heteromers have not yet been fully discovered. To operate efficiently, VRAC necessitates the presence of LRRC8A and at least one subunit among the LRRC8B-E isoforms [17, 18]. Recently, in cryoEM studies of heteromeric LRRC8A/LRRC8C complexes, different stoichiometries have been reported: Rutz et al. found 4 LRRC8A subunits and 2 LRRC8C subunits in heteromers [21], while Kern et al. reported a 5 LRRC8A/1 LRRC8C architecture [22]. In general, most cells express more than two different LRRC8 genes and LRRC8A-E assemble in various configurations, leading to the formation of VRACs with differing functional characteristics. At present, we have limited knowledge of the identities of the subunits that compose the VRAC pore, as well as how the channel is activated by cellular swelling.
Over the past three decades, intensive research endeavours have unveiled an extensive network of potential PPIs that exert a pivotal role in modulating the activity of VRAC. Experimental evidence indicates that the activation of VRAC currents can be achieved through purinergic signalling [23], a process involving calcium (Ca2+) signalling and protein phosphorylation events [24], as well as by the activation of bradykinin receptor signalling, which is intricately regulated by reactive oxygen species (ROS) and Ca2+ nanodomains [23, 25]. Furthermore, ROS have been suggested to influence the activation of VRAC by EGF [26]. Additionally, VRAC can be induced isovolumetrically by intracellular GTPγS [27, 28]. Volume-independent activation of VRAC is also induced by sphingosine-1-phosphate, which is generated by bacterial lipopolysaccharide-activated S-kinase, PDGF, TNFα, thrombin, IgE-bound antigen, and especially ATP [29].
Despite considerable effort, there is an insufficient amount of information available on its signalling networks. In 2016, Syeda et al. generated a cell line expressing LRRC8A with a FLAG-tag to biochemically investigate the protein and its associated partners [30]. The detection of an 800-kDa complex in native gels implies potential interactions between LRRC8 subunits and other proteins. The authors used mass spectrometry (MS) to identify the proteins associated with LRRC8A. Only peptides from the four LRRC8 family members were detected, and no other binding partners were found. Probably, the lack of data on VRAC interaction factors in the literature owes to the channel’s hydrophobicity and the technical challenges associated with biochemical manipulation. Indeed, in the authors themselves, suggest that the use of detergents and the tag affinity purification process may have resulted in the loss of probable associations [30].
In the present investigation, we have undertaken an in-depth exploration of the PPI network associated with LRRC8A. This endeavour was accomplished by the application of the state-of-the art BioID technique, with a specific emphasis on the central subunit LRRC8A. We exploited the BioID methodology to construct an exhaustive compendium of proteins engaged in interactions with LRRC8A/VRAC. These interactions encompass a spectrum of strengths, temporal dynamics, and indirect connections. The discerned proteins are presumed to establish close or direct functional affiliations with the LRRC8A subunit, thereby harbouring the potential to bestow valuable insights into the pathophysiological mechanisms and underlying functional roles.
Results
LRRC8s alterations and expression in human cancers: impact on patient survival
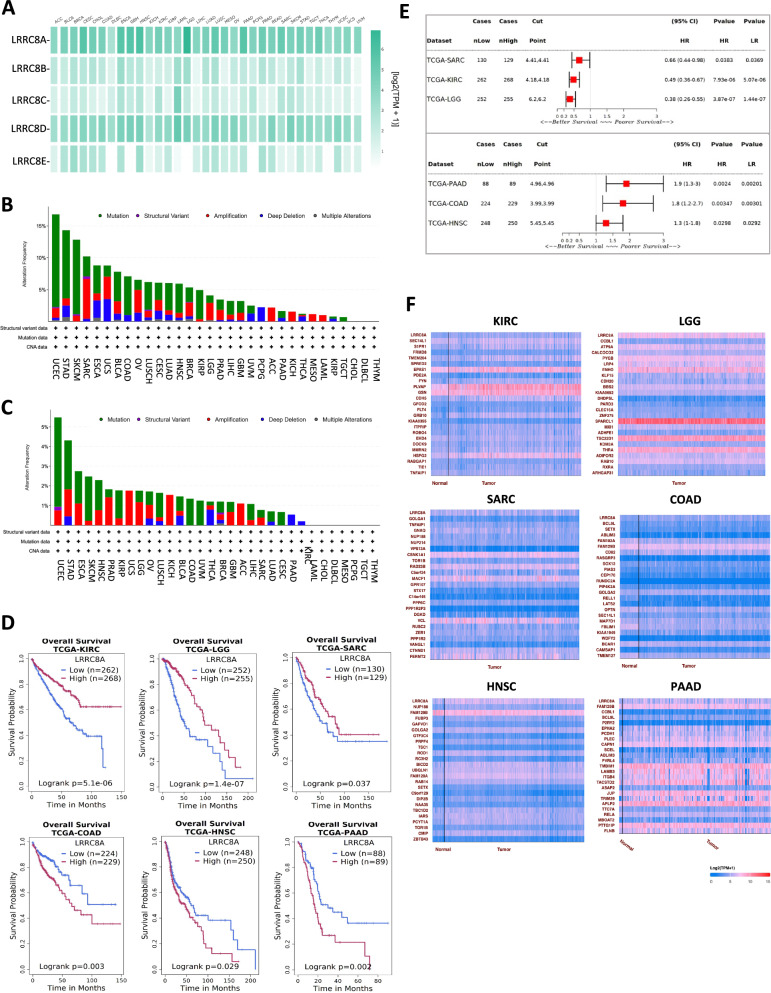
The GEPIA database [31] was used to evaluate the expression profile of LRRC8 genes. A comparative analysis of multiple genes revealed that LRRC8A and LRRC8D exhibit higher expression in tumours when compared to other LRRC8 genes, providing an overall characterization (Fig. 1A). Exploring the publicly accessible TCGA database enabled us to systematically investigate the correlation between VRAC channels in specific cancer types and patient survival. To assess comprehensive alterations, we analysed all LRRC8 genes (LRRC8A-E) using the TCGA Pan-Cancer Atlas dataset, combining data from 32 human cancers, encompassing a total of 10,953 patients. Genetic alterations within the cBioPortal database were categorized as mutations, deep deletions, gene amplifications, structural variants, and multiple alterations [32, 33]. The graphical representation depicting cancer types illustrates a discernible pattern of genomic alterations in LRRC8 genes across the TCGA PanCancer cohorts (Fig. 1B). The results indicate that the 10 types of cancer with the highest frequency of alterations were uterine corpus endometrial carcinoma (UCEC), stomach adenocarcinoma (STAD), skin cutaneous melanoma (SKCM), sarcoma (SARC) and oesophageal adenocarcinoma (ESCA), uterine carcinosarcoma (UCS), bladder urothelial carcinoma (BLCA), colorectal adenocarcinoma (COAD), ovarian serous cystadenocarcinoma (OV), and lung squamous cell carcinoma (LUSC). The cumulative mutation frequency ranged from 16.82% for UCEC to 6.16% for LUSC. Truncating, in-frame, or missense mutations were prevalent in LRRC8 genes, while amplifications, multiple alterations, or deep deletions, especially homozygous deletions in non-aneuploidy cases, were less frequently observed. Figure 1C delineates the specific genetic alterations of the LRRC8A gene.
Fig. 1. LRRC8s family in human cancers.
A Differential expression of LRRC8s family in human cancers. LRRC8s multiple gene comparison was performed using TCGA and GTEx datasets and using the GEPIA database. Data were normalized as transcripts per kilobase million (TPM) values. TPM values were converted to log2-normalized transcripts per million [log2(TPM + 1)]. B Mutation frequencies of LRRC8s in 32 cancer studies were retrieved from cBioPortal (TCGA Pan-Cancer Atlas dataset). C Mutation frequencies of LRRC8A in 32 cancer studies were retrieved from cBioPortal (TCGA Pan-Cancer Atlas dataset). D Survival plots based on LRRC8A expression level in represented tumours were obtained through Kaplan–Meier analysis by sorting samples for high and low LRRC8A expression groups according to Survival Genie software. E The forest plot illustrates hazard ratio (HR) analyses. The results of the Wald test (HR P-value) and log-rank test (LR P-value) are also displayed. F LRRC8A correlated differentially expressed genes and related pathways. The top 25 positively LRRC8A co-expressed genes were mapped using the TCGA KIRC, LGG, SARC, COAD, HNSC, and PAAD datasets in the ULCAN database.
To highlight the prognostic impact of LRRC8A expression in patients with cancer, we evaluated the Kaplan-Meier analysis to classify samples into groups with high and low LRRC8A expression. Our findings indicate a significant association between LRRC8A expression levels and patient prognosis across six different types of cancer. Patients with high LRRC8A levels had a significantly worse prognosis in cases of COAD, Head-Neck Squamous Cell Carcinoma (HNSC), and Pancreatic adenocarcinoma (PAAD). Conversely, Kidney renal clear cell carcinoma (KIRC), Low-grade glioma (LGG), and SARC demonstrated a favourable outcome (Fig. 1D). However, statistically significant results were not obtained for other types of tumours. The forest plot (Fig. 1E) illustrating hazard ratio (HR) analyses, was generated using the Survival Genie software, a web-based platform designed for conducting survival analyses in both paediatric and adult cancer populations [16]. This graph illustrates the HR and corresponding 95% confidence intervals for two distinct groups analysed univariably. The results of the Wald test (HR P-value) and log-rank test (LR P-value) are also displayed. Important details, such as the cut-off point used to classify patients as having high or low expression, and the sample sizes for each group, are provided. In the figure, the hazard ratio is represented by a central box (significant correlations are highlighted in red), and the lower and upper bounds of the 95% confidence interval are indicated by horizontal lines. For example, for COAD, the patients were divided into two groups, namely high (n = 229 samples) and low (n = 224 samples). It has been established those patients in the high group, with a median cut-off point of 3.99, exhibit an unfortunate prognosis (HR = 1.8; 95% CI 1.2-2.7; P = 0.00347 by Wald-test, and P = 0.0030 by log-rank test) (Fig. 1E).
Fascinated by the dual role of LRRC8A/VRAC in tumours, which is linked to both favourable and unfavourable prognoses (as illustrated in Fig. 1D and E), our aim was to recognize differentially expressed genes (DEGs) related to LRRC8A to illuminate its functional roles (Fig. 1F). To achieve this objective, a thorough examination of genes with a positive correlation to LRRC8A in KIRC, LGG, SARC, COAD, HNSC, and PAAD tumour types was conducted in their respective TCGA datasets. This was accomplished by utilizing the UALCAN database [34, 35]. Over 100 differentially DEGs were found to be shared between tumour types associated with both a positive prognosis (KIRC, LGG, and SARC) and those linked to a negative prognosis (COAD, HNSC and PAAD,) within the TCGA tumours datasets. This suggests that these common genes may play a critical role in tumour progression and outcome. The levels of LRRC8A transcripts discovered in different tumours and the difference in LRRC8A expression between normal and tumours tissue lack justification for the impact of genes on survival, suggesting that aspects beyond gene expression need to be studied.
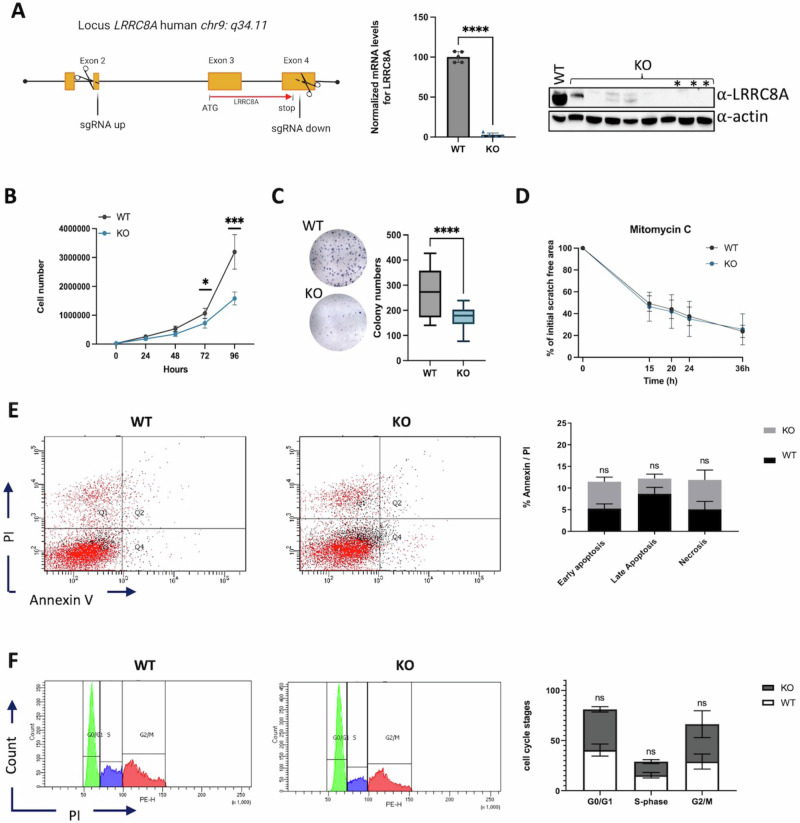
Our in-depth analysis, as illustrated in Fig. 2 supports the above findings and specifically examines the effects of LRRC8A deletion on HCT116 cell behaviour. The increased expression of LRRC8A seems to be linked with reduced survival among CRC patients with positive lymph nodes, indicating LRRC8A proteins’ possible involvement in CRC metastasis by aiding cell migration [36]. However, these findings contradict those reported by Liu et al. [37], who showed that the deletion of LRRC8A or all members of LRRC8 did not reduce the migration of HCT116 cells. To gain new insights, we employed CRISPR/Cas9 technology to generate a new LRRC8A-deficient HCT116 knockout cell line. Our model completely removed the LRRC8A gene, and we isolated two independent monoclonal cell lines for further analysis. Target site-specific PCR was utilized to evaluate any modifications in the genomic DNA sequence. The lack of LRRC8A expression was verified by Sanger sequencing, Western blot, and RT-qPCR (Fig. 2A). To investigate the potential effects of LRRC8A, cell proliferation and wound healing assays were performed on HCT116 cells that either expressed or lacked LRRC8A (Fig. 2B–D). Cell counting over a period of up to 96 hours demonstrated a significant decrease in proliferation rate for the KO clones in comparison to the WT controls at both 72 hours (p = 0.0130) and 96 hours (p < 0.0001), suggesting an involvement of LRRC8A in cell proliferation (Fig. 2B). To further explore the influence of LRRC8A on cell proliferation further, a colony formation assay was conducted. The data demonstrated a substantial reduction (p < 0.0001) in colony count for the KO clones relative to the controls, consistent with the outcomes of the cell proliferation assay (Fig. 2C). Finally, we explored the function of LRRC8A in cell migration via a wound healing assay. Supplementary treatment of cells with mitomycin C validated that the formation of a confluent cell monolayer post-scratching was attributed to migration proficiency rather than cellular proliferation. No significant differences were found in the migration rate between the controls and KO clones during both experimental conditions (Fig. 2D).
Fig. 2. Characterization of LRRC8A knockout HCT116 cells.
A Strategy of CRISPR/Cas9 editing applied for LRRC8A and characterization of LRRC8A KO HCT116 cells. KO-LRRC8A monoclonal HCTT16 cell lines were isolated and evaluated for the expression of their LRRC8A gene using real-time qPCR. HCT116 WT cells were used as a control, and the housekeeping genes ACTB and TBP were employed. Gene expression data were normalised to the control and presented as a percentage variation (n = 5, one-way ANOVA, ****p < 0.0001). In addition, the α-LRRC8A antibody was used to assess the protein expression of LRRC8A, with HCT116 WT cells as the control. Protein extracts (50 μg) were loaded for each sample, with actin serving as the loading control. Single clones indicated by a red asterisk were selected for further analysis. B Growth curves of KO- and WT-LRRC8A HCT116 cells. At the 96-hour, it was found that KO clones exhibited a lower proliferation rate compared to HCT116 WT control (n = 4, two-way ANOVA, ***p-value < 0.001). C Colony formation assay. LRRC8A-KO cells formed significantly fewer colonies compared to HCT116 WT control (n = 5, one-way ANOVA, *p-value < 0.5). D The Wound Healing Assay examined migratory capacity by measuring the percentage of the initial scratch area free at various time points (from t = 0 h to t = 36 h) using ImageJ software. In both experimental settings (mitomycin 5 μg/mL), HCT116 WT control and LRRC8A-KO clones displayed no significant differences in scratch closure speed (n = 4, two-way ANOVA). Illustrative histograms showing HCT116 WT and KO clone. Statistical analysis of independent triplicate experiments showed non-significant differences between WT and the KO cells in terms of apoptosis (E) and cell cycle (F). Data were compared by Nonparametric T- test with a significance level of p < 0.05.
Numerous studies have suggested that LRRC8A supports cell survival under hypotonic conditions and facilitates tumorigenesis by suppressing apoptosis both in vitro and in vivo [38, 39]. Consequently, we investigated the influence of LRRC8A on apoptosis in HCT116 WT and KO cells using an Annexin V/propidium iodide assay. However, our results revealed no discernible impact on apoptosis events with the deletion of LRRC8A (Fig. 2E). Additionally, the analysis of the cell cycle did not reveal any significant differences in the G0/G1, S, and G2/M phases (Fig. 2F).
Identification and Analysis of the LRRC8A Differentially Expressed Genes (DEGs) in colon Cancer
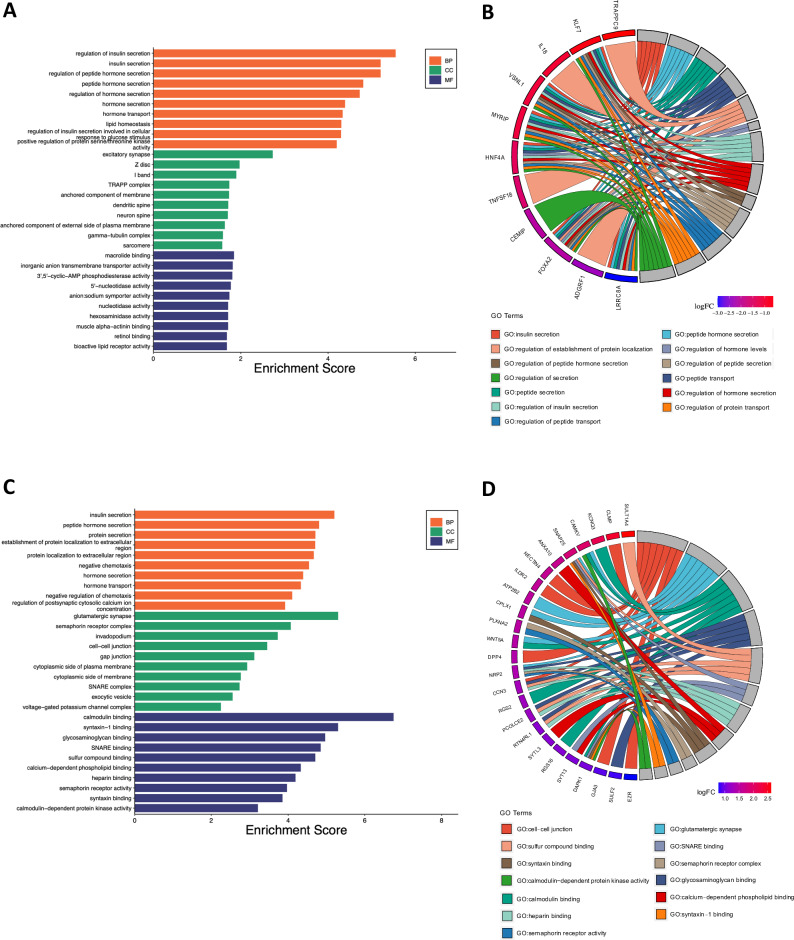
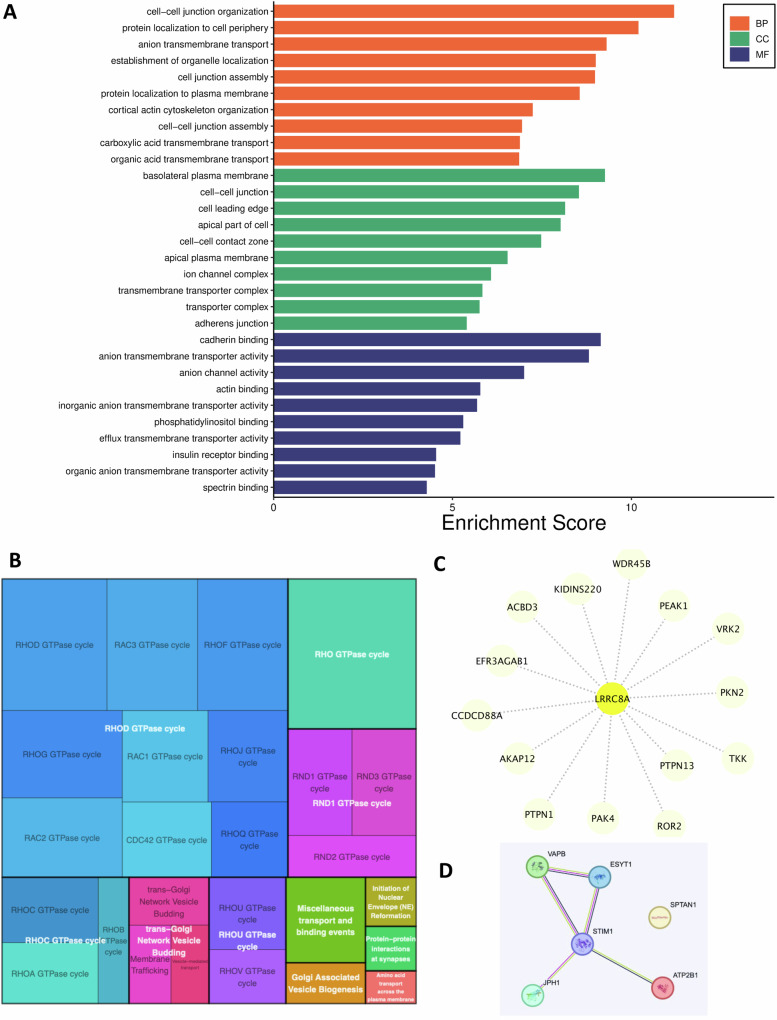
An RNA-Seq analysis was conducted to assess the impact of the LRRC8A deletion on HCT116 gene expression. A total of 125 genes displayed differential expression in LRRC8A KO versus WT cells: 56 genes being down-regulated and 69 up-regulated in LRRC8A KO cells (Table 1). Figure 3 shows the Gene Ontology (GO) enrichment scores for genes that were identified as either down-regulated or up-regulated. Panels 3A and 3C illustrate the GO enrichment for down-regulated and up-regulated genes, respectively. The enrichment bars are categorized by Biological Processes (BP) in orange, Cellular Components (CC) in green, and Molecular Functions (MF) in blue. Greater enrichment significance is indicated by longer bars, suggesting a more substantial association with the set of genes being analyzed. For example, in Fig. 3A, processes such as ‘regulation of insulin secretion’ show notable enrichment, indicating significant down-regulation. Figure 3C shows that the up-regulated genes are significantly enriched in processes such as ‘insulin secretion’ and components such as ‘cell junctions’. Functions such as ‘calmodulin binding’ are particularly prominent. Figures 3B and D display chord diagrams that visually represent the relationships between genes and their corresponding GO terms. Figure 3B shows down-regulated genes, while Fig. 3D shows up-regulated genes. The outer circle of each diagram lists genes, which are connected by coloured chords to specific GO terms that describe BP, CC, and MF. These chords display the network of associations between genes and their functions or locations within the cell. The chords are colour-matched with the corresponding GO categories they represent. On the gene side, a colour gradient illustrates the log-fold change (logFC) in gene expression. Warmer colours indicate a higher degree of down- or up-regulation. These diagrams summarise the complex interactions and functional implications of the observed changes in gene expression in the study.
Table 1.
DEGs in LRRC8A KO CRC cells.
| Differentially Expressed Genes (DEG) | ||
|---|---|---|
| ADGRF1 | ENSG00000153292 | DOWN |
| ALDH8A1 | ENSG00000118514 | UP |
| ANKRD22 | ENSG00000152766 | DOWN |
| ANXA10 | ENSG00000109511 | UP |
| APOBEC3G | ENSG00000239713 | UP |
| ARHGAP29 | ENSG00000137962 | DOWN |
| ARHGEF10L | ENSG00000074964 | DOWN |
| ARRDC4 | ENSG00000140450 | UP |
| ATP2B2 | ENSG00000157087 | UP |
| B3GALT5 | ENSG00000183778 | UP |
| CACNB4 | ENSG00000182389 | UP |
| CAMKV | ENSG00000164076 | UP |
| CCN3 | ENSG00000136999 | UP |
| CD24 | ENSG00000272398 | DOWN |
| CEMIP | ENSG00000103888 | DOWN |
| CLMP | ENSG00000166250 | UP |
| CPLX1 | ENSG00000168993 | UP |
| CRYBG1 | ENSG00000112297 | DOWN |
| CRYZL2P | ENSG00000242193 | DOWN |
| DAPK1 | ENSG00000196730 | UP |
| DPP4 | ENSG00000197635 | UP |
| DYNLT1 | ENSG00000146425 | UP |
| EZR | ENSG00000092820 | UP |
| FAM222A-AS1 | ENSG00000255650 | UP |
| FOXA2 | ENSG00000125798 | DOWN |
| FRMD4B | ENSG00000114541 | DOWN |
| FRMPD3 | ENSG00000147234 | UP |
| GARIN5A | ENSG00000142530 | DOWN |
| GGT5 | ENSG00000099998 | DOWN |
| GJA3 | ENSG00000121743 | UP |
| GPAT3 | ENSG00000138678 | UP |
| GPD1 | ENSG00000167588 | UP |
| GPR55 | ENSG00000135898 | UP |
| GREB1L | ENSG00000141449 | UP |
| GTF2H5 | ENSG00000272047 | UP |
| HKDC1 | ENSG00000156510 | UP |
| HNF4A | ENSG00000101076 | DOWN |
| IKZF2 | ENSG00000030419 | DOWN |
| IL18 | ENSG00000150782 | DOWN |
| ILDR2 | ENSG00000143195 | UP |
| INAVA | ENSG00000163362 | DOWN |
| KCNQ3 | ENSG00000184156 | UP |
| KLF7 | ENSG00000118263 | DOWN |
| KLRK1-AS1 | ENSG00000245648 | DOWN |
| LCN2 | ENSG00000148346 | DOWN |
| LGR4 | ENSG00000205213 | UP |
| LINC00698 | ENSG00000244342 | DOWN |
| LINC02593 | ENSG00000223764 | UP |
| LNCOC1 | ENSG00000253741 | DOWN |
| LOC102723553 | ENSG00000273590 | DOWN |
| LPAR1 | ENSG00000198121 | DOWN |
| LRRC8A | ENSG00000136802 | DOWN |
| LYPD3 | ENSG00000124466 | UP |
| MECOM + B58:B124 | ENSG00000085276 | UP |
| MEF2C | ENSG00000081189 | UP |
| MGLL | ENSG00000074416 | UP |
| MNS1 | ENSG00000138587 | DOWN |
| MTUS1-DT | ENSG00000253671 | DOWN |
| MYRIP | ENSG00000170011 | DOWN |
| NAPRT | ENSG00000147813 | DOWN |
| NAV2 | ENSG00000166833 | DOWN |
| NAV3 | ENSG00000067798 | DOWN |
| NECTIN4 | ENSG00000143217 | UP |
| NEMP2-DT | ENSG00000233654 | DOWN |
| NFIB | ENSG00000147862 | DOWN |
| NR2F1 | ENSG00000175745 | DOWN |
| NR2F1-AS1 | ENSG00000237187 | DOWN |
| NRIP1 | ENSG00000180530 | DOWN |
| NRP2 | ENSG00000118257 | UP |
| NT5E | ENSG00000135318 | DOWN |
| PALLD | ENSG00000129116 | DOWN |
| PCAT2 | ENSG00000254166 | DOWN |
| PCDH19 | ENSG00000165194 | UP |
| PCOLCE2 | ENSG00000163710 | UP |
| PDE4B | ENSG00000184588 | DOWN |
| PHYHD1 | ENSG00000175287 | UP |
| PKIB | ENSG00000135549 | DOWN |
| PLXNA2 | ENSG00000076356 | UP |
| PURPL | ENSG00000250337 | UP |
| PXDN | ENSG00000130508 | UP |
| RBP1 | ENSG00000114115 | DOWN |
| RBPMS | ENSG00000157110 | UP |
| RGS16 | ENSG00000143333 | UP |
| RGS2 | ENSG00000116741 | UP |
| RNF43 | ENSG00000108375 | DOWN |
| RTN4RL1 | ENSG00000185924 | UP |
| RUNX2 | ENSG00000124813 | UP |
| SAMD11 | ENSG00000187634 | UP |
| SERPING1 | ENSG00000149131 | UP |
| SLC16A6 | ENSG00000108932 | UP |
| SLC17A9 | ENSG00000101194 | UP |
| SLC5A5 | ENSG00000105641 | DOWN |
| SLCO3A1 | ENSG00000176463 | UP |
| SNAP25 | ENSG00000132639 | UP |
| SRGAP1 | ENSG00000196935 | DOWN |
| STK39 | ENSG00000198648 | DOWN |
| STYK1 | ENSG00000060140 | DOWN |
| SULF2 | ENSG00000196562 | UP |
| SULT1A4 | ENSG00000213648 | UP |
| SYT13 | ENSG00000019505 | UP |
| SYTL3 | ENSG00000164674 | UP |
| TACSTD2 | ENSG00000184292 | DOWN |
| TLL1 | ENSG00000038295 | UP |
| TMEM154 | ENSG00000170006 | UP |
| TMEM181 | ENSG00000146433 | UP |
| TMEM200A | ENSG00000164484 | DOWN |
| TNFRSF10C | ENSG00000173535 | UP |
| TNFSF18 | ENSG00000120337 | DOWN |
| TRAPPC9 | ENSG00000167632 | DOWN |
| TRHDE | ENSG00000072657 | UP |
| TRPV2 | ENSG00000187688 | UP |
| TULP4 | ENSG00000130338 | UP |
| VGLL3 | ENSG00000206538 | UP |
| VSNL1 | ENSG00000163032 | DOWN |
| WNT5A | ENSG00000114251 | UP |
| ZNF585B | ENSG00000245680 | DOWN |
| ZNF595 | ENSG00000272602 | DOWN |
| ZNF704 | ENSG00000164684 | DOWN |
A total of 125 genes displayed differential expression in LRRC8A KO versus WT cells. 56 genes being down-regulated and 69 up-regulated in LRRC8A KO cells.
Fig. 3. Gene Ontology (GO) enrichment analysis of down- and up-regulated genes across three principal ontologies.
A The graph provides the enrichment scores from a Gene Ontology analysis focusing on down-regulated genes, categorized into the domains of Biological Processes (BP), Cellular Components (CC), and Molecular Functions (MF). The bars, color-coded as orange for BP, green for CC, and blue for MF, illustrate the degree of each GO term within the studied dataset. Notably, BP terms such as regulation of insulin secretion’ and ‘lipid homeostasis’ show the highest enrichment scores, indicating their significant down-regulation in the genomic profile. B The figure delineates a series of genes with their corresponding log fold changes (logFC), indicating a down-regulation in expression levels. The color-coded segments represent individual genes, while the connecting ribbons illustrate their binding to specific GO terms associated. The GO terms lists include cell-cell junctions, synaptic activity, and binding activities (such as calmodulin and glycosaminoglycan). C The chart depicts the enrichment scores derived from a GO analysis for up-regulated genes. Notably, “calmodulin-dependent protein kinase activity” in the MF category and “insulin secretion” in the BP category exhibit the highest enrichment scores, suggesting significant up-regulation in these functional areas. D The figure draws the genes with their corresponding logFC. GO terms associated with these genes are listed, including those related to cell-cell junctions, synaptic activity, and binding activities (such as calmodulin and glycosaminoglycan).
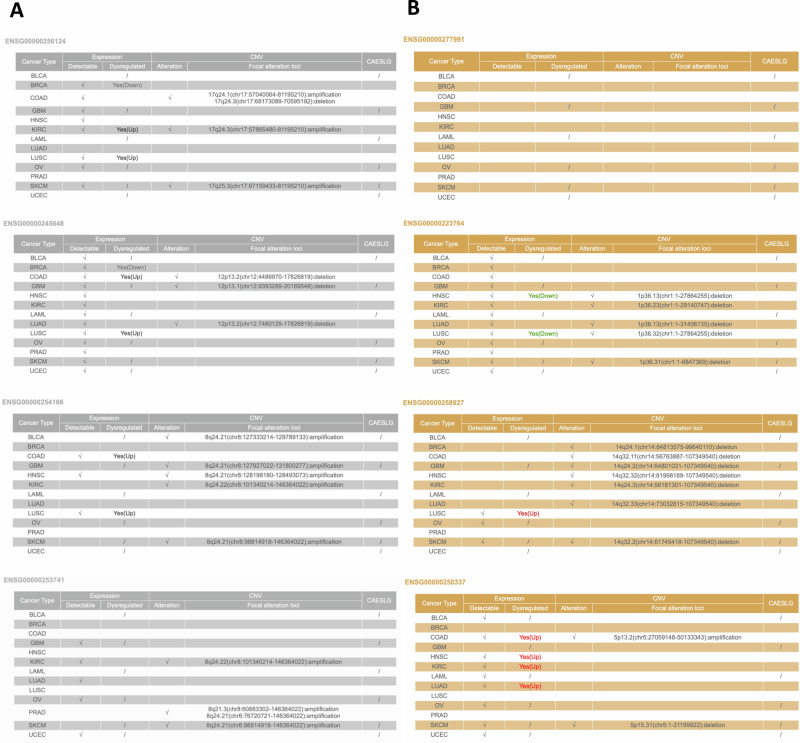
Furthermore, our attention was directed towards long non-coding RNAs (lncRNAs). Recent research has shown a significant correlation between aberrant expression patterns of lncRNAs and various complex human diseases, particularly cancer. The increasing repertoire of lncRNAs has led to their characterization as either ‘oncogenes’ or ‘tumor suppressors’. Dysregulation of lncRNAs has been linked to the initiation, progression, and metastasis of cancer [40, 41]. The Cancer LncRNome Atlas was used to investigate alterations in individual lncRNAs at transcriptional, genomic, and epigenetic levels in human cancers to identify relevant information (Table 2 and Fig. 4). The observed lncRNAs exhibited varied patterns of regulation across different cancer types, with some showing upregulation and others downregulation (Table 2 and Fig. 4). This differential expression, coupled with their location on the chromosomes as noted in the data, hints at the complex genetic architecture that underlies cancer.
Table 2.
List of lncRNAs.
| Regulation | Ensembl ID | log2 Fold Change | Adj.Pval | Symbol | Chr |
|---|---|---|---|---|---|
| Down | ENSG00000256124 | −6.55320993004657 | 2.72e−03 | LINC01152 | 17q24.3 |
| Down | ENSG00000245648 | −1.4700236550104 | 1.85e−05 | KLRK1-AS1 | 12p13.2 |
| Down | ENSG00000254166 | −1.4339799975236 | 1.03e−05 | CASC19 | 8q24.21 |
| Down | ENSG00000253741 | −1.07168286754586 | 2.20e−03 | LNCOC1 | 8q24.3 |
| Up | ENSG00000277991 | 4.24840538102438 | 9.66e−05 | 21p11.2 | |
| Up | ENSG00000258927 | 3.31905187015385 | 4.01e−03 | 14q32.13 | |
| Up | ENSG00000223764 | 1.49820395651212 | 3.96e−03 | LINC02593 | 1p36.33 |
| Up | ENSG00000250337 | 1.44062288967738 | 1.34e−03 | PURPL | 5p14.1 |
The expression is evaluated considering LRRC8A-KO vs WT in HCT116.
Fig. 4. Expression Profiles of lncRNAs Across Various Cancer Types.
The tables systematically compare the expression profiles of long non-coding RNAs (lncRNAs) across different cancer types. A and B illustrate, respectively, the downregulated and upregulated lncRNAs in various types of cancers. Each row represents a specific lncRNA, with columns indicating detectability, expression dysregulation, alterations, and the localization of focal alterations within the genome, as identified in the CAESLG database.
Defining the LRRC8A interactome in living cells
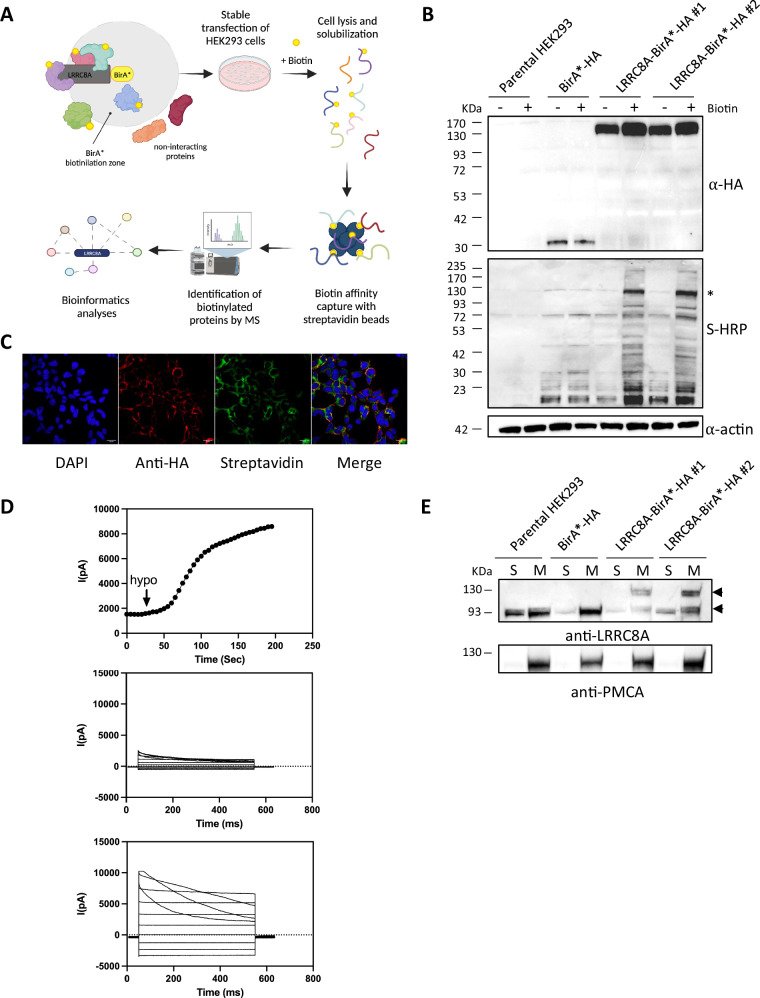
To identify regulators of LRRC8A, we employed the BioID technique (Fig. 5A). WT human LRRC8A was fused with BirA*-HA and HEK293 cell lines stably expressing the LRRC8A-BirA*-HA fusion protein were generated. Stable expression was employed to maximize the formation of VRAC heteromers. The parental cell line and cells expressing solely the BirA* enzyme served as negative and positive controls, respectively. To validate the proper production of the BirA* protein fusion, we performed western blot experiments using Streptavidin-HRP to assess the degree of biotinylation in the presence and absence of exogenous biotin (Fig. 5B). As expected, distinct bands of varying sizes were observed in LRRC8A-BirA* cells in the presence of biotin. Conversely, minimal levels of biotinylation were detected in control lysates, both in the presence and absence of biotin (Fig. 5B). Subsequently, to verify that the genetic fusion of LRRC8A with BirA*-HA did not alter intracellular localization and channel function, we conducted confocal microscopy and patch-clamp experiments (Fig. 5C, D). The presence of BirA*-HA did not impact channel activity (Fig. 5D), consistent with observations for LRRC8A with a C-terminal GFP tag—a protein with a similar molecular weight and steric hindrance to the BirA* enzyme [18]. Fluorescence microscopy confirmed that the fusion protein localized to the plasma membrane similarly to endogenous channels (Fig. 5C) Biotinylation was further confirmed using Alexa Fluor 488 nm-conjugated streptavidin (Fig. 5C). Additionally, western blots confirmed the localization of the fusion protein in membranes (Fig. 5E).
Fig. 5. Characterization of cells stably expressing LRRC8A-BioID fusion protein.
A Schematic representation of the BioID technique, a method for exploring protein complexes in live cells [114, 120]. Within the BioID methodology, the target protein is expressed in cells as a fusion with a specialized tagging enzyme, BirA R118G (a promiscuous mutant biotin ligase, hereinafter referred to as BirA*). This enzyme utilizes exogenous biotin to catalyse the formation of biotinoyl-5’-AMP, a highly reactive molecule that biotinylates primary amines, such as the lysine side chain, within a proximity of approximately 10 nm [121]. Subsequently, cells are lysed, and the labeled proteins are subjected to affinity purification, followed by detection through MS. The identification of pertinent biotinylated proteins is then accomplished through quantitative and statistical methodologies B Verification of BirA* activity in cells expressing LRRC8A-BirA*-HA by Western blot. The expression of the LRRC8A-BirA*-HA fusion protein was assessed using an anti-HA antibody in the absence and presence of biotin. BirA* activity in cells expressing LRRC8A-BirA*-HA was assessed by WB using streptavidin-HRP for the detection of biotinylated proteins. Activity was assessed in the presence and absence of biotin (50 μM), using HEK293 cells and cells transfected with pcDNA3.1 MCS-BirA(R118G)-HA (in the blot indicated as BirA*) as controls. Actin was used as a loading control. For each sample, 50 μg of total protein extract was loaded. C Validation of the subcellular localization of the LRRC8A-BirA*-HA protein by immunofluorescence. Nuclei were visualized by DAPI staining; α-HA antibody and Alexa Fluor 586-conjugated red-emitting secondary antibody were used to visualize LRRC8A-BirA*-HA; streptavidin-HRP was used to visualize biotinylated proteins. D Validation of LRRC8A-BirA*-HA protein channel activity by patch clamp: Representative time course of current activation upon perfusion with a hypotonic solution of cells co-expressing the 8A-BirA*-8E heteromers. E Validation of the subcellular localization of the LRRC8A-BirA*-HA protein by Western Blotting. The localization of the LRRC8A-BirA*-HA protein was assessed in the two clones and in HEK293 cells and cells transfected with pcDNA3.1 MCS-BirA(R118G)-HA using the α-LRRC8A antibody. For each sample, 50 μg of protein extract enriched with the cytoplasmic (S) or membrane (M) protein fraction was loaded. The α-PMCA antibody was used to exclude contamination of the S-fraction by proteins from the M-fraction.
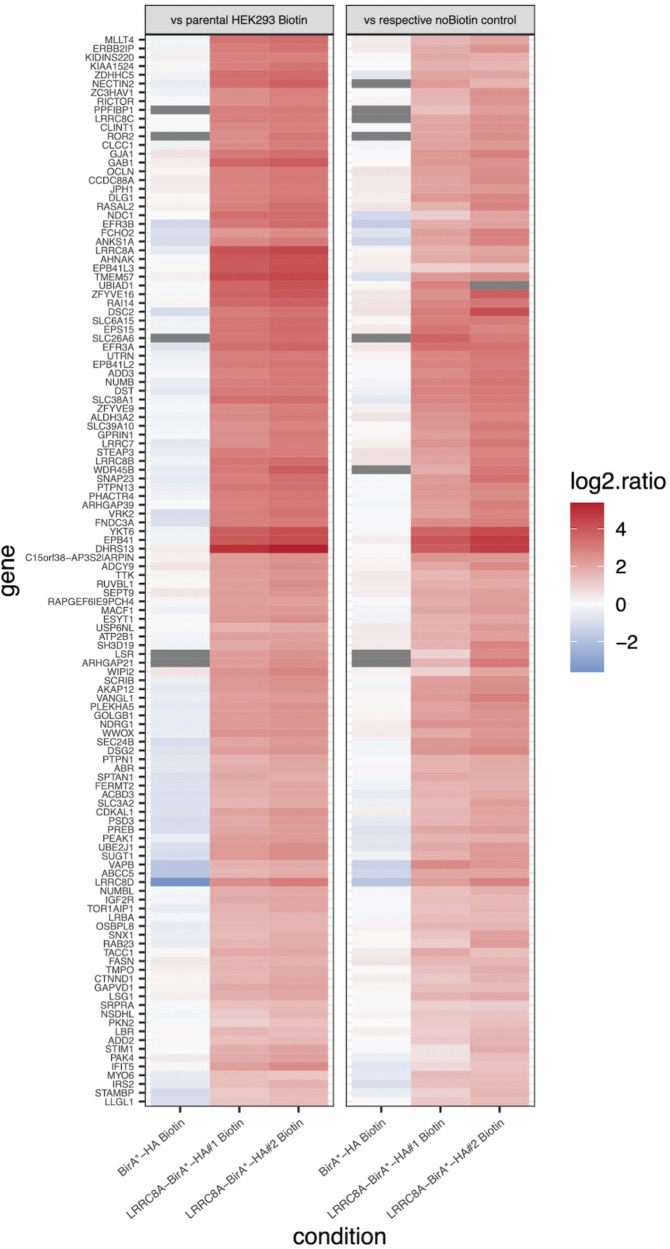
Following the validation of the correct cellular localization and function of LRRC8A-BioID, purified biotinylated proteins were analysed with quantitative mass spectrometry (MS) using tandem mass tag labelling (TMT). The tagged proteins were isolated through streptavidin affinity purification, TMT labelled and analysed by LC-MS/MS. Raw files were analysed using IsobarQuant and Mascot (v2.2.07) (see overview of normalized TMT reporter ion intensities in Figure S1A for an overview of samples). Quantification of individual protein enrichment was determined by comparing their relative abundance in each LRRC8A-BirA* sample with the experimentally paired Venus-BirA* control (no Biotin control, see Volcano Plots in Figure S1B). To obtain a comprehensive understanding of the proximal protein environment surrounding the VRAC complex, a stringent cut-off was applied to the dataset. Only proteins quantified with two unique peptide matches were retained for analysis. Proteins were further tested for differential abundance using a moderated t-test by applying the limma R package. As a significance cut-off we used a false discovery rate (FDR) of less than 0.05 and a fold change (FC) of at least 2-fold. Out of 1227 proteins, we selected 122 enriched hits (around 10% of the total) with a significant up-biotinylation in the comparisons ‘LRRC8A-BirA*-HA#1 Biotin vs LRRC8A-BirA*-HA#1 noBiotin’ or ‘LRRC8A-BirA*-HA#2 Biotin vs LRRC8A-BirA*-HA#2 noBiotin’ (see Volcano Plots in Figure S1B, Fig. 6, Table 3).
Fig. 6. MS results: identification of hits proteins.
Heatmap reporting the log2 fold changes as indicated in the heading (left column ratio against the parental HEK293 Biotin control and right column against the corresponding noBiotin control) for the 122 identified enriched proteins (hits).
Table 3.
List and annotations of hit proteins from BioID analyses.
| Protein | stringID | Annotation |
|---|---|---|
| ABCC5 | 9606.ENSP00000333926 | Multidrug resistance-associated protein 5; Acts as a multispecific organic anion pump which can transport nucleotide analogs; Belongs to the ABC transporter superfamily. ABCC family. Conjugate transporter (TC 3.A.1.208) subfamily |
| ABR | 9606.ENSP00000303909 | Active breakpoint cluster region-related protein; GTPase-activating protein for RAC and CDC42. Promotes the exchange of RAC or CDC42-bound GDP by GTP, thereby activating them; C2 domain containing |
| ACBD3 | 9606.ENSP00000355777 | Acyl-coa binding domain containing 3; Golgi resident protein GCP60; Involved in the maintenance of Golgi structure by interacting with giantin, affecting protein transport between the endoplasmic reticulum and Golgi. Involved in hormone-induced steroid biosynthesis in testicular Leydig cells (By similarity). Recruits PI4KB to the Golgi apparatus membrane; enhances the enzyme activity of PI4KB activity via its membrane recruitment thereby increasing the local concentration of the substrate in the vicinity of the kinase; A-kinase anchoring proteins |
| ADD2 | 9606.ENSP00000264436 | Beta-adducin; Membrane-cytoskeleton-associated protein that promotes the assembly of the spectrin-actin network. Binds to the erythrocyte membrane receptor SLC2A1/GLUT1 and may therefore provide a link between the spectrin cytoskeleton to the plasma membrane. Binds to calmodulin. Calmodulin binds preferentially to the beta subunit; Belongs to the aldolase class II family. |
| ADD3 | 9606.ENSP00000348381 | Gamma-adducin; Membrane-cytoskeleton-associated protein that promotes the assembly of the spectrin-actin network. Plays a role in actin filament capping. Binds to calmodulin; Belongs to the aldolase class II family. Adducin subfamily |
| AHNAK | 9606.ENSP00000367263 | Neuroblast differentiation-associated protein AHNAK; May be required for neuronal cell differentiation; PDZ domain containing |
| AKAP12 | 9606.ENSP00000384537 | A-kinase anchor protein 12; Anchoring protein that mediates the subcellular compartmentation of protein kinase A (PKA) and protein kinase C (PKC); A-kinase anchoring proteins |
| ALDH3A2 | 9606.ENSP00000345774 | Aldehyde dehydrogenase 3 family member a2; Fatty aldehyde dehydrogenase; Catalyzes the oxidation of long-chain aliphatic aldehydes to fatty acids. Active on a variety of saturated and unsaturated aliphatic aldehydes between 6 and 24 carbons in length. Responsible for conversion of the sphingosine 1-phosphate (S1P) degradation product hexadecenal to hexadecenoic acid |
| ANKS1A | 9606.ENSP00000353518 | Ankyrin repeat and SAM domain-containing protein 1A; Regulator of different signaling pathways. Regulates EPHA8 receptor tyrosine kinase signaling to control cell migration and neurite retraction (By similarity); Ankyrin repeat domain containing |
| ARHGAP21 | 9606.ENSP00000379709 | Rho GTPase-activating protein 21; Functions as a GTPase-activating protein (GAP) for RHOA and CDC42. Downstream partner of ARF1 which may control Golgi apparatus structure and function. Also required for CTNNA1 recruitment to adherens junctions; PDZ domain containing |
| ARHGAP39 | 9606.ENSP00000366522 | Rho gtpase-activating protein 39; Rho GTPase activating protein 39 |
| ATP2B1 | 9606.ENSP00000392043 | Plasma membrane calcium-transporting ATPase 1; This magnesium-dependent enzyme catalyzes the hydrolysis of ATP coupled with the transport of calcium out of the cell; ATPases Ca2+ transporting |
| CCDC88A | 9606.ENSP00000338728 | Coiled-coil domain containing 88a; Girdin; Plays a role as a key modulator of the AKT-mTOR signaling pathway controlling the tempo of the process of newborn neurons integration during adult neurogenesis, including correct neuron positioning, dendritic development and synapse formation (By similarity). Enhances phosphoinositide 3-kinase (PI3K)- dependent phosphorylation and kinase activity of AKT1/PKB, but does not possess kinase activity itself (By similarity). Phosphorylation of AKT1/PKB thereby induces the phosphorylation of downstream effectors GSK3 and FOXO1/FKHR |
| CDKAL1 | 9606.ENSP00000274695 | Threonylcarbamoyladenosine tRNA methylthiotransferase; Catalyzes the methylthiolation of N6- threonylcarbamoyladenosine (t(6)A), leading to the formation of 2- methylthio-N6-threonylcarbamoyladenosine (ms(2)t(6)A) at position 37 in tRNAs that read codons beginning with adenine; Belongs to the methylthiotransferase family. CDKAL1 subfamily |
| CLCC1 | 9606.ENSP00000349456 | Chloride channel CLIC-like protein 1; Seems to act as a chloride ion channel; Tetraspan junctional complex superfamily |
| CLINT1 | 9606.ENSP00000429824 | Clathrin interactor 1; Binds to membranes enriched in phosphatidylinositol 4,5- bisphosphate (PtdIns(4,5)P2). May have a role in transport via clathrin-coated vesicles from the trans-Golgi network to endosomes. Stimulates clathrin assembly |
| CTNND1 | 9606.ENSP00000382004 | Catenin delta-1; Binds to and inhibits the transcriptional repressor ZBTB33, which may lead to activation of target genes of the Wnt signaling pathway (By similarity). Associates with and regulates the cell adhesion properties of both C-, E- and N-cadherins, being critical for their surface stability. Implicated both in cell transformation by SRC and in ligand-induced receptor signaling through the EGF, PDGF, CSF-1 and ERBB2 receptors. Promotes GLIS2 C-terminal cleavage; Belongs to the beta-catenin family |
| DHRS13 | 9606.ENSP00000368173 | Dehydrogenase/reductase SDR family member 13; Putative oxidoreductase |
| DLG1 | 9606.ENSP00000345731 | Disks large homolog 1; Essential multidomain scaffolding protein required for normal development (By similarity). Recruits channels, receptors and signaling molecules to discrete plasma membrane domains in polarized cells. May play a role in adherens junction assembly, signal transduction, cell proliferation, synaptogenesis and lymphocyte activation. Regulates the excitability of cardiac myocytes by modulating the functional expression of Kv4 channels. Functional regulator of Kv1.5 channel; Belongs to the MAGUK family |
| DSC2 | 9606.ENSP00000280904 | Desmocollin-2; Component of intercellular desmosome junctions. Involved in the interaction of plaque proteins and intermediate filaments mediating cell-cell adhesion. May contribute to epidermal cell positioning by mediating differential adhesiveness between cells that express different isoforms |
| DSG2 | 9606.ENSP00000261590 | Desmoglein-2; Component of intercellular desmosome junctions. Involved in the interaction of plaque proteins and intermediate filaments mediating cell-cell adhesion |
| DST | 9606.ENSP00000307959 | Dystonin; Cytoskeletal linker protein. Acts as an integrator of intermediate filaments, actin and microtubule cytoskeleton networks. Required for anchoring either intermediate filaments to the actin cytoskeleton in neural and muscle cells or keratin- containing intermediate filaments to hemidesmosomes in epithelial cells. The proteins may self-aggregate to form filaments or a two- dimensional mesh. Regulates the organization and stability of the microtubule network of sensory neurons to allow axonal transport. Mediates docking of the dynein/dynactin motor complex to vesicle cargos |
| EFR3A | 9606.ENSP00000254624 | Protein EFR3 homolog A; Component of a complex required to localize phosphatidylinositol 4-kinase (PI4K) to the plasma membrane. The complex acts as a regulator of phosphatidylinositol 4-phosphate (PtdIns(4)P) synthesis (Probable). In the complex, EFR3A probably acts as the membrane-anchoring component. Also involved in responsiveness to G-protein-coupled receptors; it is however unclear whether this role is direct or indirect |
| EFR3B | 9606.ENSP00000384081 | Protein EFR3 homolog B; Component of a complex required to localize phosphatidylinositol 4-kinase (PI4K) to the plasma membrane. The complex acts as a regulator of phosphatidylinositol 4-phosphate (PtdIns(4)P) synthesis (Probable). In the complex, EFR3B probably acts as the membrane-anchoring component. Also involved in responsiveness to G-protein-coupled receptors; it is however unclear whether this role is direct or indirect; Armadillo-like helical domain containing |
| EPB41 | 9606.ENSP00000345259 | Protein 4.1; Protein 4.1 is a major structural element of the erythrocyte membrane skeleton. It plays a key role in regulating membrane physical properties of mechanical stability and deformability by stabilizing spectrin-actin interaction. Recruits DLG1 to membranes. Required for dynein-dynactin complex and NUMA1 recruitment at the mitotic cell cortex during anaphase |
| EPB41L2 | 9606.ENSP00000338481 | Band 4.1-like protein 2; Required for dynein-dynactin complex and NUMA1 recruitment at the mitotic cell cortex during anaphase; Erythrocyte membrane protein band 4.1 |
| EPB41L3 | 9606.ENSP00000343158 | Band 4.1-like protein 3; Tumor suppressor that inhibits cell proliferation and promotes apoptosis. Modulates the activity of protein arginine N- methyltransferases, including PRMT3 and PRMT5; Erythrocyte membrane protein band 4.1 |
| EPS15 | 9606.ENSP00000360798 | Epidermal growth factor receptor substrate 15; Involved in cell growth regulation. May be involved in the regulation of mitogenic signals and control of cell proliferation. Involved in the internalization of ligand-inducible receptors of the receptor tyrosine kinase (RTK) type, in particular EGFR. Plays a role in the assembly of clathrin-coated pits (CCPs). Acts as a clathrin adapter required for post-Golgi trafficking. Seems to be involved in CCPs maturation including invagination or budding. Involved in endocytosis of integrin beta- 1 (ITGB1) and transferrin receptor (TFR) |
| ERBB2IP | 9606.ENSP00000426632 | Erbin; Acts as an adapter for the receptor ERBB2, in epithelia. By binding the unphosphorylated ‘Tyr-1248’ of receptor ERBB2, it may contribute to stabilize this unphosphorylated state. Inhibits NOD2-dependent NF-kappa-B signaling and proinflammatory cytokine secretion; Belongs to the LAP (LRR and PDZ) protein family |
| ESYT1 | 9606.ENSP00000267113 | Extended synaptotagmin-1; Binds glycerophospholipids in a barrel-like domain and may play a role in cellular lipid transport (By similarity). Binds calcium (via the C2 domains) and translocates to sites of contact between the endoplasmic reticulum and the cell membrane in response to increased cytosolic calcium levels. Helps tether the endoplasmic reticulum to the cell membrane and promotes the formation of appositions between the endoplasmic reticulum and the cell membrane; Belongs to the extended synaptotagmin family |
| FASN | 9606.ENSP00000304592 | Fatty acid synthase; Fatty acid synthetase catalyzes the formation of long- chain fatty acids from acetyl-CoA, malonyl-CoA and NADPH. This multifunctional protein has 7 catalytic activities and an acyl carrier protein; Seven-beta-strand methyltransferase motif containing |
| FCHO2 | 9606.ENSP00000393776 | F-BAR domain only protein 2; Functions in an early step of clathrin-mediated endocytosis. Has both a membrane binding/bending activity and the ability to recruit proteins essential to the formation of functional clathrin-coated pits. Has a lipid-binding activity with a preference for membranes enriched in phosphatidylserine and phosphoinositides (Pi(4,5) biphosphate) like the plasma membrane. Its membrane-bending activity might be important for the subsequent action of clathrin and adaptors in the formation of clathrin-coated vesicles |
| FERMT2 | 9606.ENSP00000342858 | Fermitin family homolog 2; Scaffolding protein that enhances integrin activation mediated by TLN1 and/or TLN2, but activates integrins only weakly by itself. Binds to membranes enriched in phosphoinositides. Enhances integrin-mediated cell adhesion onto the extracellular matrix and cell spreading; this requires both its ability to interact with integrins and with phospholipid membranes. Required for the assembly of focal adhesions. Participates in the connection between extracellular matrix adhesion sites and the actin cytoskeleton |
| FNDC3A | 9606.ENSP00000441831 | Fibronectin type-III domain-containing protein 3A; Mediates spermatid-Sertoli adhesion during spermatogenesis; Belongs to the FNDC3 family |
| GAB1 | 9606.ENSP00000262995 | GRB2-associated-binding protein 1; Adapter protein that plays a role in intracellular signaling cascades triggered by activated receptor-type kinases. Plays a role in FGFR1 signaling. Probably involved in signaling by the epidermal growth factor receptor (EGFR) and the insulin receptor (INSR) |
| GAPVD1 | 9606.ENSP00000377664 | GTPase-activating protein and VPS9 domain-containing protein 1; Acts both as a GTPase-activating protein (GAP) and a guanine nucleotide exchange factor (GEF), and participates in various processes such as endocytosis, insulin receptor internalization or LC2A4/GLUT4 trafficking. Acts as a GEF for the Ras-related protein RAB31 by exchanging bound GDP for free GTP, leading to regulate LC2A4/GLUT4 trafficking. In the absence of insulin, it maintains RAB31 in an active state and promotes a futile cycle between LC2A4/GLUT4 storage vesicles and early endosomes |
| GJA1 | 9606.ENSP00000282561 | Gap junction alpha-1 protein; Gap junction protein that acts as a regulator of bladder capacity. A gap junction consists of a cluster of closely packed pairs of transmembrane channels, the connexons, through which materials of low MW diffuse from one cell to a neighboring cell. May play a critical role in the physiology of hearing by participating in the recycling of potassium to the cochlear endolymph. Negative regulator of bladder functional capacity: acts by enhancing intercellular electrical and chemical transmission |
| GOLGB1 | 9606.ENSP00000377275 | Golgin subfamily B member 1; May participate in forming intercisternal cross-bridges of the Golgi complex |
| GPRIN1 | 9606.ENSP00000305839 | G protein-regulated inducer of neurite outgrowth 1; May be involved in neurite outgrowth |
| IFIT5 | 9606.ENSP00000360860 | Interferon-induced protein with tetratricopeptide repeats 5; Interferon-induced RNA-binding protein involved in the human innate immune response. Has a broad and adaptable RNA structure recognition important for RNA recognition specificity in antiviral defense. Binds precursor and processed tRNAs as well as poly-U-tailed tRNA fragments. Specifically binds single-stranded RNA bearing a 5’-triphosphate group (PPP-RNA), thereby acting as a sensor of viral single-stranded RNAs. |
| IGF2R | 9606.ENSP00000349437 | Cation-independent mannose-6-phosphate receptor; Transport of phosphorylated lysosomal enzymes from the Golgi complex and the cell surface to lysosomes. Lysosomal enzymes bearing phosphomannosyl residues bind specifically to mannose-6- phosphate receptors in the Golgi apparatus and the resulting receptor-ligand complex is transported to an acidic prelyosomal compartment where the low pH mediates the dissociation of the complex. This receptor also binds IGF2. Acts as a positive regulator of T-cell coactivation, by binding DPP4; CD molecules |
| IRS2 | 9606.ENSP00000365016 | Insulin receptor substrate 2; May mediate the control of various cellular processes by insulin; Pleckstrin homology domain containing |
| JPH1 | 9606.ENSP00000344488 | Junctophilin-1; Junctophilins contribute to the formation of junctional membrane complexes (JMCs) which link the plasma membrane with the endoplasmic or sarcoplasmic reticulum in excitable cells. Provides a structural foundation for functional cross-talk between the cell surface and intracellular calcium release channels. JPH1 contributes to the construction of the skeletal muscle triad by linking the t-tubule (transverse-tubule) and SR (sarcoplasmic reticulum) membranes |
| KIAA1524 | 9606.ENSP00000295746 | Cellular inhibitor of pp2a; Protein CIP2A; Oncoprotein that inhibits PP2A and stabilizes MYC in human malignancies. Promotes anchorage-independent cell growth and tumor formation |
| KIDINS220 | 9606.ENSP00000256707 | Ankyrin repeat-rich membrane spanning protein; Kinase D-interacting substrate of 220 kDa; Promotes a prolonged MAP-kinase signaling by neurotrophins through activation of a Rap1-dependent mechanism. Provides a docking site for the CRKL-C3G complex, resulting in Rap1-dependent sustained ERK activation. May play an important role in regulating postsynaptic signal transduction through the syntrophin-mediated localization of receptor tyrosine kinases such as EPHA4. In cooperation with SNTA1 can enhance EPHA4-induced JAK/STAT activation. |
| LBR | 9606.ENSP00000339883 | Delta14-sterol reductase (lamin-B receptor); Lamin-B receptor; Anchors the lamina and the heterochromatin to the inner nuclear membrane; Tudor domain containing |
| LLGL1 | 9606.ENSP00000321537 | LLGL1, scribble cell polarity complex component; Lethal(2) giant larvae protein homolog 1; Cortical cytoskeleton protein found in a complex involved in maintaining cell polarity and epithelial integrity. Involved in the regulation of mitotic spindle orientation, proliferation, differentiation and tissue organization of neuroepithelial cells. Involved in axonogenesis through RAB10 activation thereby regulating vesicular membrane trafficking toward the axonal plasma membrane |
| LRBA | 9606.ENSP00000349629 | Lipopolysaccharide-responsive and beige-like anchor protein; May be involved in coupling signal transduction and vesicle trafficking to enable polarized secretion and/or membrane deposition of immune effector molecules; Armadillo-like helical domain containing |
| LRRC7 | 9606.ENSP00000035383 | Leucine-rich repeat-containing protein 7; Required for normal synaptic spine architecture and function. Necessary for DISC1 and GRM5 localization to postsynaptic density complexes and for both N-methyl D-aspartate receptor-dependent and metabotropic glutamate receptor-dependent long term depression; Belongs to the LAP (LRR and PDZ) protein family |
| LRRC8B | 9606.ENSP00000332674 | Volume-regulated anion channel subunit LRRC8B; Non-essential component of the volume-regulated anion channel (VRAC, also named VSOAC channel), an anion channel required to maintain a constant cell volume in response to extracellular or intracellular osmotic changes. The VRAC channel conducts iodide better than chloride and may also conduct organic osmolytes like taurine. Channel activity requires LRRC8A plus at least one other family member (LRRC8B, LRRC8C, LRRC8D or LRRC8E); channel characteristics depend on the precise subunit composition |
| LRRC8C | 9606.ENSP00000359483 | Volume-regulated anion channel subunit LRRC8C; Non-essential component of the volume-regulated anion channel (VRAC, also named VSOAC channel), an anion channel required to maintain a constant cell volume in response to extracellular or intracellular osmotic changes. The VRAC channel conducts iodide better than chloride and may also conduct organic osmolytes like taurine. Channel activity requires LRRC8A plus at least one other family member (LRRC8B, LRRC8C, LRRC8D or LRRC8E); channel characteristics depend on the precise subunit composition |
| LRRC8D | 9606.ENSP00000338887 | Volume-regulated anion channel subunit LRRC8D; Non-essential component of the volume-regulated anion channel (VRAC, also named VSOAC channel), an anion channel required to maintain a constant cell volume in response to extracellular or intracellular osmotic changes. The VRAC channel conducts iodide better than chloride and may also conduct organic osmolytes like taurine. Channel activity requires LRRC8A plus at least one other family member (LRRC8B, LRRC8C, LRRC8D or LRRC8E); channel characteristics depend on the precise subunit composition |
| LSG1 | 9606.ENSP00000265245 | Large subunit GTPase 1 homolog; GTPase required for the XPO1/CRM1-mediated nuclear export of the 60 S ribosomal subunit. Probably acts by mediating the release of NMD3 from the 60 S ribosomal subunit after export into the cytoplasm |
| LSR | 9606.ENSP00000480821 | Lipolysis-stimulated lipoprotein receptor; Probable role in the clearance of triglyceride-rich lipoprotein from blood. Binds chylomicrons, LDL and VLDL in presence of free fatty acids and allows their subsequent uptake in the cells (By similarity); Belongs to the immunoglobulin superfamily. LISCH7 family |
| MACF1 | 9606.ENSP00000354573 | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5; Isoform 2: F-actin-binding protein which plays a role in cross-linking actin to other cytoskeletal proteins and also binds to microtubules. Plays an important role in ERBB2-dependent stabilization of microtubules at the cell cortex. Acts as a positive regulator of Wnt receptor signaling pathway and is involved in the translocation of AXIN1 and its associated complex (composed of APC, CTNNB1 and GSK3B) from the cytoplasm to the cell membrane (By similarity). |
| MLLT4 | 9606.ENSP00000375960 | Afadin; Belongs to an adhesion system, probably together with the E-cadherin-catenin system, which plays a role in the organization of homotypic, interneuronal and heterotypic cell-cell adherens junctions (AJs). Nectin- and actin-filament-binding protein that connects nectin to the actin cytoskeleton |
| MYO6 | 9606.ENSP00000358994 | Unconventional myosin-VI; Myosins are actin-based motor molecules with ATPase activity. Unconventional myosins serve in intracellular movements. Myosin 6 is a reverse-direction motor protein that moves towards the minus-end of actin filaments. Has slow rate of actin-activated ADP release due to weak ATP binding. Functions in a variety of intracellular processes such as vesicular membrane trafficking and cell migration. Required for the structural integrity of the Golgi apparatus via the p53-dependent pro-survival pathway. |
| NDC1 | 9606.ENSP00000360483 | NDC1 transmembrane nucleoporin; Nucleoporin NDC1; Component of the nuclear pore complex (NPC), which plays a key role in de novo assembly and insertion of NPC in the nuclear envelope. Required for NPC and nuclear envelope assembly, possibly by forming a link between the nuclear envelope membrane and soluble nucleoporins, thereby anchoring the NPC in the membrane |
| NDRG1 | 9606.ENSP00000404854 | N-myc downstream regulated 1; Protein NDRG1; Stress-responsive protein involved in hormone responses, cell growth, and differentiation. Acts as a tumor suppressor in many cell types. Necessary but not sufficient for p53/TP53- mediated caspase activation and apoptosis. Has a role in cell trafficking, notably of the Schwann cell, and is necessary for the maintenance and development of the peripheral nerve myelin sheath. Required for vesicular recycling of CDH1 and TF. May also function in lipid trafficking. Protects cells from spindle disruption damage. |
| NECTIN2 | 9606.ENSP00000252483 | Nectin-2; Modulator of T-cell signaling. Can be either a costimulator of T-cell function, or a coinhibitor, depending on the receptor it binds to. Upon binding to CD226, stimulates T-cell proliferation and cytokine production, including that of IL2, IL5, IL10, IL13, and IFNG. Upon interaction with PVRIG, inhibits T-cell proliferation. These interactions are competitive. Probable cell adhesion protein; Belongs to the nectin family |
| NSDHL | 9606.ENSP00000359297 | Sterol-4-alpha-carboxylate 3-dehydrogenase, decarboxylating; Involved in the sequential removal of two C-4 methyl groups in post-squalene cholesterol biosynthesis; Short chain dehydrogenase/reductase superfamily |
| NUMB | 9606.ENSP00000451300 | Numb, endocytic adaptor protein; Protein numb homolog; Plays a role in the process of neurogenesis. Required throughout embryonic neurogenesis to maintain neural progenitor cells, also called radial glial cells (RGCs), by allowing their daughter cells to choose progenitor over neuronal cell fate. Not required for the proliferation of neural progenitor cells before the onset of neurogenesis. Also involved postnatally in the subventricular zone (SVZ) neurogenesis by regulating SVZ neuroblasts survival and ependymal wall integrity. May also mediate local repair of brain ventricular wall damage |
| NUMBL | 9606.ENSP00000252891 | Numb-like protein; Plays a role in the process of neurogenesis. Required throughout embryonic neurogenesis to maintain neural progenitor cells, also called radial glial cells (RGCs), by allowing their daughter cells to choose progenitor over neuronal cell fate. Not required for the proliferation of neural progenitor cells before the onset of embryonic neurogenesis. Also required postnatally in the subventricular zone (SVZ) neurogenesis by regulating SVZ neuroblasts survival and ependymal wall integrity. Negative regulator of NF-kappa-B signaling pathway. The inhibition of NF- kappa-B a […] |
| OCLN | 9606.ENSP00000347379 | Occludin; May play a role in the formation and regulation of the tight junction (TJ) paracellular permeability barrier. It is able to induce adhesion when expressed in cells lacking tight junctions; Protein phosphatase 1 regulatory subunits |
| OSBPL8 | 9606.ENSP00000261183 | Oxysterol-binding protein-related protein 8; Lipid transporter involved in lipid countertransport between the endoplasmic reticulum and the plasma membrane: specifically exchanges phosphatidylserine with phosphatidylinositol 4-phosphate (PI4P), delivering phosphatidylserine to the plasma membrane in exchange for PI4P, which is degraded by the SAC1/SACM1L phosphatase in the endoplasmic reticulum. Binds phosphatidylserine and PI4P in a mutually exclusive manner. Binds oxysterol, 25- hydroxycholesterol and cholesterol; Belongs to the OSBP family |
| PAK4 | 9606.ENSP00000469413 | Serine/threonine-protein kinase PAK 4; Serine/threonine protein kinase that plays a role in a variety of different signaling pathways including cytoskeleton regulation, cell migration, growth, proliferation or cell survival. Activation by various effectors including growth factor receptors or active CDC42 and RAC1 results in a conformational change and a subsequent autophosphorylation on several serine and/or threonine residues. Phosphorylates and inactivates the protein phosphatase SSH1, leading to increased inhibitory phosphorylation of the actin binding/depolymerizing factor cofilin |
| PEAK1 | 9606.ENSP00000452796 | Pseudopodium-enriched atypical kinase 1; Tyrosine kinase that may play a role in cell spreading and migration on fibronectin. May directly or indirectly affect phosphorylation levels of cytoskeleton-associated proteins MAPK1/ERK and PXN |
| PHACTR4 | 9606.ENSP00000362942 | Phosphatase and actin regulator 4; Regulator of protein phosphatase 1 (PP1) required for neural tube and optic fissure closure, and enteric neural crest cell (ENCCs) migration during development. Acts as an activator of PP1 by interacting with PPP1CA and preventing phosphorylation of PPP1CA at ‘Thr-320’. During neural tube closure, localizes to the ventral neural tube and activates PP1, leading to down-regulate cell proliferation within cranial neural tissue and the neural retina. Also acts as a regulator of migration of enteric neural crest cells (ENCCs) by activating PP1 |
| PKN2 | 9606.ENSP00000359552 | Serine/threonine-protein kinase N2; PKC-related serine/threonine-protein kinase and Rho/Rac effector protein that participates in specific signal transduction responses in the cell. Plays a role in the regulation of cell cycle progression, actin cytoskeleton assembly, cell migration, cell adhesion, tumor cell invasion and transcription activation signaling processes. Phosphorylates CTTN in hyaluronan-induced astrocytes and hence decreases CTTN ability to associate with filamentous actin. Phosphorylates HDAC5, therefore lead to impair HDAC5 import. |
| PLEKHA5 | 9606.ENSP00000404296 | Pleckstrin homology domain-containing family a member 5; Pleckstrin homology domain containing A5 |
| PPFIBP1 | 9606.ENSP00000314724 | Ppfia binding protein 1; Liprin-beta-1; May regulate the disassembly of focal adhesions. Did not bind receptor-like tyrosine phosphatases type 2 A; Sterile alpha motif domain containing |
| PREB | 9606.ENSP00000260643 | Prolactin regulatory element-binding protein; Guanine nucleotide exchange factor that specifically activates the small GTPase SAR1B. Mediates the recruitement of SAR1B and other COPII coat components to endoplasmic reticulum membranes and is therefore required for the formation of COPII transport vesicles from the ER; WD repeat domain containing |
| PSD3 | 9606.ENSP00000324127 | PH and SEC7 domain-containing protein 3; Guanine nucleotide exchange factor for ARF6; Pleckstrin homology domain containing |
| PTPN1 | 9606.ENSP00000360683 | Tyrosine-protein phosphatase non-receptor type 1; Tyrosine-protein phosphatase which acts as a regulator of endoplasmic reticulum unfolded protein response. Mediates dephosphorylation of EIF2AK3/PERK; inactivating the protein kinase activity of EIF2AK3/PERK. May play an important role in CKII- and p60c-src-induced signal transduction cascades. May regulate the EFNA5-EPHA3 signaling pathway which modulates cell reorganization and cell-cell repulsion. May also regulate the hepatocyte growth factor receptor signaling pathway through dephosphorylation of MET |
| PTPN13 | 9606.ENSP00000394794 | Tyrosine-protein phosphatase non-receptor type 13; Tyrosine phosphatase which regulates negatively FAS- induced apoptosis and NGFR-mediated pro-apoptotic signaling. May regulate phosphoinositide 3-kinase (PI3K) signaling through dephosphorylation of PIK3R2; FERM domain containing |
| RAB23 | 9606.ENSP00000417610 | RAB23, member RAS oncogene family; Ras-related protein Rab-23; The small GTPases Rab are key regulators of intracellular membrane trafficking, from the formation of transport vesicles to their fusion with membranes. Rabs cycle between an inactive GDP-bound form and an active GTP-bound form that is able to recruit to membranes different set of downstream effectors directly responsible for vesicle formation, movement, tethering and fusion. Together with SUFU, prevents nuclear import of GLI1, and thereby inhibits GLI1 transcription factor activity. |
| RAI14 | 9606.ENSP00000427123 | Retinoic acid induced 14; Ankycorbin; Plays a role in actin regulation at the ectoplasmic specialization, a type of cell junction specific to testis. Important for establishment of sperm polarity and normal spermatid adhesion. May also promote integrity of Sertoli cell tight junctions at the blood-testis barrier; Ankyrin repeat domain containing |
| RAPGEF6 | 9606.ENSP00000296859 | Rap guanine nucleotide exchange factor 6; Guanine nucleotide exchange factor (GEF) for Rap1A, Rap2A and M-Ras GTPases. Does not interact with cAMP; PDZ domain containing |
| RASAL2 | 9606.ENSP00000356621 | Ras GTPase-activating protein nGAP; Inhibitory regulator of the Ras-cyclic AMP pathway; C2 and RasGAP domain containing |
| RICTOR | 9606.ENSP00000296782 | Rapamycin-insensitive companion of mTOR; Subunit of mTORC2, which regulates cell growth and survival in response to hormonal signals. mTORC2 is activated by growth factors, but, in contrast to mTORC1, seems to be nutrient- insensitive. mTORC2 seems to function upstream of Rho GTPases to regulate the actin cytoskeleton, probably by activating one or more Rho-type guanine nucleotide exchange factors. mTORC2 promotes the serum-induced formation of stress-fibers or F-actin. mTORC2 plays a critical role in AKT1 ‘ |
| ROR2 | 9606.ENSP00000364860 | Tyrosine-protein kinase transmembrane receptor ROR2; Tyrosine-protein kinase receptor which may be involved in the early formation of the chondrocytes. It seems to be required for cartilage and growth plate development (By similarity). Phosphorylates YWHAB, leading to induction of osteogenesis and bone formation. In contrast, has also been shown to have very little tyrosine kinase activity in vitro. May act as a receptor for wnt ligand WNT5A which may result in the inhibition of WNT3A-mediated signaling; I-set domain containing |
| RUVBL1 | 9606.ENSP00000318297 | RuvB-like 1; May be able to bind plasminogen at cell surface and enhance plasminogen activation; AAA ATPases |
| SCRIB | 9606.ENSP00000349486 | Protein scribble homolog; Scaffold protein involved in different aspects of polarized cells differentiation regulating epithelial and neuronal morphogenesis. Most probably functions in the establishment of apico-basal cell polarity. May function in cell proliferation regulating progression from G1 to S phase and as a positive regulator of apoptosis for instance during acinar morphogenesis of the mammary epithelium. May also function in cell migration and adhesion and hence regulate cell invasion through MAPK signaling. May play a role in exocytosis and in the targeting synaptic vesicle |
| SEC24B | 9606.ENSP00000428564 | Protein transport protein Sec24B; Component of the coat protein complex II (COPII) which promotes the formation of transport vesicles from the endoplasmic reticulum (ER). The coat has two main functions, the physical deformation of the endoplasmic reticulum membrane into vesicles and the selection of cargo molecules for their transport to the Golgi complex. Plays a central role in cargo selection within the COPII complex and together with SEC24A may have a different specificity compared to SEC24C and SEC24D. May package preferentially cargos with cytoplasmic DxE or LxxLE motifs |
| SEPT9 | 9606.ENSP00000391249 | Septin-9; Filament-forming cytoskeletal GTPase (By similarity). May play a role in cytokinesis (Potential). May play a role in the internalization of 2 intracellular microbial pathogens. Belongs to the TRAFAC class TrmE-Era-EngA-EngB-Septin- like GTPase superfamily. Septin GTPase family |
| SH3D19 | 9606.ENSP00000302913 | SH3 domain-containing protein 19; May play a role in regulating A disintegrin and metalloproteases (ADAMs) in the signaling of EGFR-ligand shedding. May be involved in suppression of Ras-induced cellular transformation and Ras-mediated activation of ELK1. Plays a role in the regulation of cell morphology and cytoskeletal organization |
| SLC26A6 | 9606.ENSP00000378920 | Solute carrier family 26 member 6; Apical membrane anion-exchanger with wide epithelial distribution that plays a role as a component of the pH buffering system for maintaining acid-base homeostasis. Acts as a versatile DIDS-sensitive inorganic and organic anion transporter that mediates the uptake of monovalent anions like chloride, bicarbonate, formate and hydroxyl ion and divalent anions like sulfate and oxalate. Function in multiple exchange modes involving pairs of these anions |
| SLC38A1 | 9606.ENSP00000449756 | Sodium-coupled neutral amino acid transporter 1; Functions as a sodium-dependent amino acid transporter. Mediates the saturable, pH-sensitive and electrogenic cotransport of glutamine and sodium ions with a stoichiometry of 1:1. May also transport small zwitterionic and aliphatic amino acids with a lower affinity. May supply glutamatergic and GABAergic neurons with glutamine which is required for the synthesis of the neurotransmitters glutamate and GABA; Solute carriers |
| SLC39A10 | 9606.ENSP00000386766 | Solute carrier family 39 (zinc transporter), member 10; Zinc transporter ZIP10; May act as a zinc-influx transporter; Belongs to the ZIP transporter (TC 2.A.5) family |
| SLC3A2 | 9606.ENSP00000367123 | 4F2 cell-surface antigen heavy chain; Required for the function of light chain amino-acid transporters. Involved in sodium-independent, high-affinity transport of large neutral amino acids such as phenylalanine, tyrosine, leucine, arginine and tryptophan. Involved in guiding and targeting of LAT1 and LAT2 to the plasma membrane. When associated with SLC7A6 or SLC7A7 acts as an arginine/glutamine exchanger, following an antiport mechanism for amino acid transport, influencing arginine release in exchange for extracellular amino acids. |
| SLC6A15 | 9606.ENSP00000266682 | Sodium-dependent neutral amino acid transporter B(0)AT2; Functions as a sodium-dependent neutral amino acid transporter. Exhibits preference for the branched-chain amino acids, particularly leucine, valine and isoleucine and methionine. Mediates the saturable, pH-sensitive and electrogenic cotransport of proline and sodium ions with a stoichiometry of 1:1. May have a role as transporter for neurotransmitter precursors into neurons. In contrast to other members of the neurotransmitter transporter family, does not appear to be chloride-dependent; Solute carriers |
| SNAP23 | 9606.ENSP00000249647 | Synaptosomal-associated protein 23; Essential component of the high affinity receptor for the general membrane fusion machinery and an important regulator of transport vesicle docking and fusion; Belongs to the SNAP-25 family |
| SNX1 | 9606.ENSP00000261889 | Sorting nexin-1; Involved in several stages of intracellular trafficking. Interacts with membranes containing phosphatidylinositol 3- phosphate (PtdIns(3 P)) or phosphatidylinositol 3,5-bisphosphate (PtdIns(3,5)P2). Acts in part as component of the retromer membrane-deforming SNX-BAR subcomplex. The SNX-BAR retromer mediates retrograde transport of cargo proteins from endosomes to the trans-Golgi network (TGN) and is involved in endosome-to-plasma membrane transport for cargo protein recycling. The SNX-BAR subcomplex functions to deform the donor membrane into a tubular profile |
| SPTAN1 | 9606.ENSP00000361824 | Spectrin alpha chain, non-erythrocytic 1; Fodrin, which seems to be involved in secretion, interacts with calmodulin in a calcium-dependent manner and is thus candidate for the calcium-dependent movement of the cytoskeleton at the membrane; EF-hand domain containing |
| SRPRA | 9606.ENSP00000328023 | Signal recognition particle receptor subunit alpha; Component of the SRP (signal recognition particle) receptor. Ensures, in conjunction with the signal recognition particle, the correct targeting of the nascent secretory proteins to the endoplasmic reticulum membrane system |
| STAMBP | 9606.ENSP00000377633 | STAM-binding protein; Zinc metalloprotease that specifically cleaves ‘Lys-63’- linked polyubiquitin chains. Does not cleave ‘Lys-48’-linked polyubiquitin chains (By similarity). Plays a role in signal transduction for cell growth and MYC induction mediated by IL-2 and GM-CSF. Potentiates BMP (bone morphogenetic protein) signaling by antagonizing the inhibitory action of SMAD6 and SMAD7. Has a key role in regulation of cell surface receptor-mediated endocytosis and ubiquitin-dependent sorting of receptors to lysosomes. Endosomal localization of STAMBP is required for efficient EGFR degradation |
| STEAP3 | 9606.ENSP00000376822 | Metalloreductase STEAP3; Endosomal ferrireductase required for efficient transferrin-dependent iron uptake in erythroid cells. Participates in erythroid iron homeostasis by reducing Fe(3 + ) to Fe(2 + ). Can also reduce of Cu(2 + ) to Cu(1 + ), suggesting that it participates in copper homeostasis. Uses NADP(+) as acceptor. May play a role downstream of p53/TP53 to interface apoptosis and cell cycle progression. Indirectly involved in exosome secretion by facilitating the secretion of proteins such as TCTP; STEAP family |
| STIM1 | 9606.ENSP00000478059 | Stromal interaction molecule 1; Plays a role in mediating store-operated Ca(2 + ) entry (SOCE), a Ca(2 + ) influx following depletion of intracellular Ca(2 + ) stores. Acts as Ca(2 + ) sensor in the endoplasmic reticulum via its EF-hand domain. Upon Ca(2 + ) depletion, translocates from the endoplasmic reticulum to the plasma membrane where it activates the Ca(2 + ) release-activated Ca(2 + ) (CRAC) channel subunit ORAI1. Involved in enamel formation. Activated following interaction with STIMATE, leading to promote STIM1 conformational switch; Sterile alpha motif domain containing |
| SUGT1 | 9606.ENSP00000367208 | SGT1 homolog, MIS12 kinetochore complex assembly cochaperone; Protein SGT1 homolog; May play a role in ubiquitination and subsequent proteasomal degradation of target proteins |
| TACC1 | 9606.ENSP00000321703 | Transforming acidic coiled-coil-containing protein 1; Likely involved in the processes that promote cell division prior to the formation of differentiated tissues |
| TMEM57 | 9606.ENSP00000363463 | Macoilin 1; Plays a role in the regulation of neuronal activity |
| TMPO | 9606.ENSP00000266732 | Lamina-associated polypeptide 2, isoform alpha; May be involved in the structural organization of the nucleus and in the post-mitotic nuclear assembly. Plays an important role, together with LMNA, in the nuclear anchorage of RB1; Belongs to the LEM family |
| TOR1AIP1 | 9606.ENSP00000435365 | Torsin-1A-interacting protein 1; Required for nuclear membrane integrity. Induces TOR1A and TOR1B ATPase activity and is required for their location on the nuclear membrane. Binds to A- and B-type lamins. Possible role in membrane attachment and assembly of the nuclear lamina |
| TTK | 9606.ENSP00000358813 | Serine/threonine-protein kinase ttk/mps1; Dual specificity protein kinase TTK; Phosphorylates proteins on serine, threonine, and tyrosine. Probably associated with cell proliferation. Essential for chromosome alignment by enhancing AURKB activity (via direct CDCA8 phosphorylation) at the centromere, and for the mitotic checkpoint |
| UBE2J1 | 9606.ENSP00000451261 | Ubiquitin-conjugating enzyme E2 J1; Catalyzes the covalent attachment of ubiquitin to other proteins. Functions in the selective degradation of misfolded membrane proteins from the endoplasmic reticulum (ERAD); Belongs to the ubiquitin-conjugating enzyme family |
| UBIAD1 | 9606.ENSP00000366006 | UbiA prenyltransferase domain-containing protein 1; Prenyltransferase that mediates the formation of menaquinone-4 (MK-4) and coenzyme Q10. MK-4 is a vitamin K2 isoform present at high concentrations in the brain, kidney and pancreas, and is required for endothelial cell development. Mediates the conversion of phylloquinone (PK) into MK-4, probably by cleaving the side chain of phylloquinone (PK) to release 2- methyl-1,4-naphthoquinone (menadione; K3) and then prenylating it with geranylgeranyl pyrophosphate (GGPP) to form MK-4. |
| USP6NL | 9606.ENSP00000277575 | USP6 N-terminal-like protein; Acts as a GTPase-activating protein for RAB5A and RAB43. Involved in receptor trafficking. In complex with EPS8 inhibits internalization of EGFR. Involved in retrograde transport from the endocytic pathway to the Golgi apparatus. Involved in the transport of Shiga toxin from early and recycling endosomes to the trans-Golgi network. Required for structural integrity of the Golgi complex |
| UTRN | 9606.ENSP00000356515 | Utrophin; May play a role in anchoring the cytoskeleton to the plasma membrane; Zinc fingers ZZ-type |
| VANGL1 | 9606.ENSP00000347672 | VANGL planar cell polarity protein 1 |
| VAPB | 9606.ENSP00000417175 | Vesicle-associated membrane protein-associated protein B/C; Participates in the endoplasmic reticulum unfolded protein response (UPR) by inducing ERN1/IRE1 activity. Involved in cellular calcium homeostasis regulation |
| VRK2 | 9606.ENSP00000408002 | Serine/threonine-protein kinase VRK2; Serine/threonine kinase that regulates several signal transduction pathways. Isoform 1 modulates the stress response to hypoxia and cytokines, such as interleukin-1 beta (IL1B) and this is dependent on its interaction with MAPK8IP1, which assembles mitogen-activated protein kinase (MAPK) complexes. Inhibition of signal transmission mediated by the assembly of MAPK8IP1-MAPK complexes reduces JNK phosphorylation and JUN-dependent transcription. Phosphorylates ‘Thr-18’ of p53/TP53, histone H3, and may also phosphorylate MAPK8IP1 |
| WDR45B | 9606.ENSP00000376139 | Wd repeat domain phosphoinositide-interacting protein 3; Component of the autophagy machinery that controls the major intracellular degradation process by which cytoplasmic materials are packaged into autophagosomes and delivered to lysosomes for degradation. Binds phosphatidylinositol 3-phosphate (PtdIns3P) forming on membranes of the endoplasmic reticulum upon activation of the upstream ULK1 and PI3 kinases and is recruited at phagophore assembly sites where it regulates the elongation of nascent phagophores downstream of WIPI2 |
| WIPI2 | 9606.ENSP00000288828 | WD repeat domain phosphoinositide-interacting protein 2; Early component of the autophagy machinery being involved in formation of preautophagosomal structures and their maturation into mature phagosomes in response to phosphatidylinositol 3-phosphate (PtdIns3P). May bind PtdIns3P |
| WWOX | 9606.ENSP00000457230 | WW domain-containing oxidoreductase; Putative oxidoreductase. Acts as a tumor suppressor and plays a role in apoptosis. Required for normal bone development (By similarity). May function synergistically with p53/TP53 to control genotoxic stress-induced cell death. Plays a role in TGFB1 signaling and TGFB1-mediated cell death. May also play a role in tumor necrosis factor (TNF)-mediated cell death. Inhibits Wnt signaling, probably by sequestering DVL2 in the cytoplasm; Short chain dehydrogenase/reductase superfamily |
| YKT6 | 9606.ENSP00000223369 | Synaptobrevin homolog YKT6; Vesicular soluble NSF attachment protein receptor (v- SNARE) mediating vesicle docking and fusion to a specific acceptor cellular compartment. Functions in endoplasmic reticulum to Golgi transport; as part of a SNARE complex composed of GOSR1, GOSR2 and STX5. Functions in early/recycling endosome to TGN transport; as part of a SNARE complex composed of BET1L, GOSR1 and STX5. Has a S-palmitoyl transferase activity; SNAREs |
| ZC3HAV1 | 9606.ENSP00000242351 | Zinc finger CCCH-type antiviral protein 1; Antiviral protein which inhibits the replication of viruses by recruiting the cellular RNA degradation machineries to degrade the viral mRNAs. Binds to a ZAP-responsive element (ZRE) present in the target viral mRNA, recruits cellular poly(A)- specific ribonuclease PARN to remove the poly(A) tail, and the 3’- 5’ exoribonuclease complex exosome to degrade the RNA body from the 3’-end. It also recruits the decapping complex DCP1-DCP2 through RNA helicase p72 (DDX17) to remove the cap structure of the viral mRNA to initiate its degradation |
| ZDHHC5 | 9606.ENSP00000287169 | Palmitoyltransferase ZDHHC5; Palmitoyl acyltransferase for the G-protein coupled receptor SSTR5. Also palmitoylates FLOT2 (By similarity); Zinc fingers DHHC-type |
| ZFYVE16 | 9606.ENSP00000337159 | Mad, mothers against decapentaplegic interacting protein; Zinc finger FYVE domain-containing protein 16; May be involved in regulating membrane trafficking in the endosomal pathway. Overexpression induces endosome aggregation. Required to target TOM1 to endosomes; Protein phosphatase 1 regulatory subunits |
| ZFYVE9 | 9606.ENSP00000287727 | Mad, mothers against decapentaplegic interacting protein; Zinc finger FYVE domain-containing protein 9; Early endosomal protein that functions to recruit SMAD2/SMAD3 to intracellular membranes and to the TGF-beta receptor. Plays a significant role in TGF-mediated signaling by regulating the subcellular location of SMAD2 and SMAD3 and modulating the transcriptional activity of the SMAD3/SMAD4 complex. Possibly associated with TGF-beta receptor internalization; Zinc fingers FYVE-type |
Prepared using https://string-db.org/.
None of the 122 proteins displayed enrichment in the biotin conditions of the BioID-only samples or the non-transfected HEK293 cells (see heatmap in Fig. 6). This indicates that these proteins underwent biotinylation due to their proximity to LRRC8A when fused with BirA*. Notably, the LRRC8A-associated proteins comprise almost all the known elements of the VRAC complex (LRRC8B/C/D). The lack of the LRRC8E subunit poses no issue, as only LRRC8A is necessary for channel function and must bind with at least one other isoform [10, 13, 17, 18, 42]. Additionally, certain results match with details labelled in GeneMANIA, a versatile and user-friendly online platform for hypothesizing about gene function, scrutinizing gene catalogues, and prioritizing genes for functional assays. LRRC8A has been found to be functionally associated with USP6NL, LBR, SH3D19, and TMPO, as reported in [43], where the authors combine publicly available hybrid radiation datasets from different species to provide a highly reliable map of mammalian genetic interactions. Additionally, we discovered several unforeseen protein interactors, including STIM1, PTPN1, PTPN13, and RICTOR.
To examine the findings of BioID, we implemented gene set analysis in accordance with the designated methodology from the Methods section. Subsequently, we conducted pathway enrichment analysis by utilizing GO and Pathway Reactome (PR) on the compilation of results from BioID. The figures exhibiting the ten most considerably enriched terms (p < 0.05) in each grouping are illustrated in Fig. 7A. The biological processes highlighted here include the cell-cell junctions organization, protein localization to cell periphery, anion transmembrane transport, establishment of organelle localization and cell junction assembly. Notably, the molecular functions identified in the GO terms, in addition to anion transport, comprise of phosphatidylinositol and insulin receptor binding. The cellular components with the highest enrichment were cell-cell junction, leading edge of the cell, actin cytoskeleton, cell-substrate junction, focal adhesion, cell cortex and basal plasma membrane. The list of cell-cell junction organisations includes CTNND1, DLG1, DSG2, EPB41L3, FERMT2, GJA1, LSR, NUMB, NUMBL, OCLN and PKN2 as shown in Fig. 7C. Through our analysis of PR enrichment (FDR < 0.05), we also observed physical proximity of LRRC8A to proteins associated with Rho subfamilies (Fig. 7B). Moreover, our BioID investigation indicates a prospective correlation between the VRAC channel and the management of cellular calcium. This involves proteins such as ATP2B1, ESYT1, JPH1, STIM1, SPTAN1, and VAPB (Fig. 7D).
Fig. 7. Pathway analysis of the interacting proteins.
A Analysis of GO-term enrichment, showcasing the top 10 significant biological processes, molecular functions, and cellular components, respectively, derived from the 120 hit proteins. B Reactome pathway enrichment FDR < 0.05. C Cell-cell junctions exhibited the greatest enrichment in cellular components. The chart explores the range of cell-cell junction structures that contributed to this increased enrichment. D Schematic representation of calcium-related proteins identified using BioID. The figure was created using https://string-db.org.
Discussion
Numerous investigations have examined the correlation between ion channels and cancer, suggesting their prospective application as targets for oncological therapies [4]. This paper presents a comprehensive outlook of publicly accessible databases addressing VRAC to diverse cancer types. Our meta-analysis infers a conceivable association between LRRC8A - the essential VRAC subunit- expression, and the survival of patients with cancer. Gene modifications of LRRC8A were examined across various tumour types (Fig. 1A–F). The LRRC8A mutation frequency is highest in UCEC, STAD, SKCM, SARC, and ESCA. However, the Kaplan-Meier analysis indicates a correlation between high LRRC8A expression and reduced survival rates only in COAD, HNSC, and PAAD patients (Fig. 1D). In HNSC, there is considerable evidence to suggest that VRAC function plays a critical role in mediating cisplatin sensitivity. Additionally, a molecular network analysis of LRRC8A has identified interactions with oncological factors, including HDAC4, NOTCH, and SNAI1 [44–48]. In PAAD, LRRC8A is associated with cell proliferation, migration, drug resistance, and immune infiltration [9]. Moreover, investigation of the LRRC8A gene’s co-expression and its network in PAAD patients has revealed prominent enrichment of genes situated close to LRRC8A in the PI3K-AKT and focal adhesion pathways [9]. On the contrary, detailed functions and interactions of VRAC in colon cancer are not yet clear and subjected of discussion. Some findings suggest that LRRC8A could serve as a novel biomarker for predicting the survival of colon cancer patients and that is a central mediator in mediating multiple signaling pathways to promote metastasis and targeting LRRC8A. LRRC8A proteins were found highly expressed in hematogenous metastasis from human colorectal cancer samples. The oxaliplatin-resistant HCT116 cells highly expressed LRRC8A, which was related to impaired proliferation and enhanced migration. The over-expressed LRRC8A slowed proliferation and increased migration ex vivo and in vivo [36]. Very recently, LRRC8A proteins were found highly expressed in hematogenous metastasis from human colorectal cancer samples. The elevated LRRC8A upregulated the focal adhesion, MAPK, AMPK, and chemokine signaling pathways via phosphorylation and dephosphorylation. Inhibition of LRRC8A impeded the TNF-α signaling cascade and TNF-α-induced migration. LRRC8A binding to PIP5K1B regulated the PIP2 formation, providing a platform for LRRC8A to mediate cell signaling transduction. Importantly, LRRC8A self- regulated its transcription via NF-κB1 and NF-κB2 pathways and the upregulation of NIK/NF-κB2/LRRC8A transcriptional axis was unfavorable for colon cancer patients [49]. However, in 2019, Liu et al. indicated that VRAC is not essential for the proliferation and migration of human colon cancer cells [37]. In this manuscript, we focused on a human CRC line derived from an adult male (HCT116 cells). Our independent clones of HCT116 LRRC8A-KO exhibited decreased proliferation rates and colony formation. However, the role in cell migration appears to be insignificant (Fig. 2A–C)
For further information in CRC context, we carried out an RNA-Seq analysis to evaluate the effects of LRRC8A deletion on the behaviour of HCT116 cells (Table 1). In the list of down-regulated genes, we found, amongst others, VSNL1, HNF4α, FOXA2, and KLF7. VSNL1 is a member of the neuronal calcium sensor protein family that controls calcium-dependent cell signalling and signal transduction by modulating adenylyl cyclase expression in a cAMP-dependent manner [50]. The precise functions of VSNL-1 have not been elucidated, but it appears to have several roles in tumour invasion and metastasis. Akagi et al. [51], using mRNA microarrays, evaluated the expression of VSNL-1 in >100 colorectal cancer patients. They observed that, compared to low VSNL-1 expression, high VSNL-1 expression was significantly associated with a high rate of lymph node metastases and poor prognosis for patients. It has been reported that down-regulation of VSNL1 inhibits the proliferation, migration, and invasion of colorectal cancer cells. Co-IP experiments indicated that VSNL1 can bind to COL10A1, which, when up-regulated, can promote colorectal cell proliferation, migration and invasion and reverse the effect of sh-VSNL1 on colorectal cancer cells [52]. Similarly, KLFs have been associated with the development of some cancers. Overexpression of KLF7 has been associated with worse prognosis in gastric and lung cancers, according to recent studies [53, 54]. However, mRNA expression levels of KLF7 have also been reported to be significantly elevated in CRC tissues [55]. In the list of down-regulated genes, we also found IL-18. It is known that the exit of Cl- ions from immune cells activates the NLRP3 inflammasome. Green et al. [56] found that, in macrophages, VRACs are the only Cl- channels involved in NLRP3 inflammasome activation when the cell volume changes.
In the list of up-regulated genes, we found: CCN3, CLMP, ILDR2, NECTIN4, DPP4, GJA3, and EZR. In particular, CCN3, a scaffolding protein that controls and balances the interconnection between individual signalling pathways, is involved in numerous biological processes that promote cancer development. CCN3 has antitumour effects in many tumours including CRC [57–61].
Although proteins are single entities, they hardly carry out their biological functions independently. Instead, they combine and create complex and dynamic molecular machines. A global map of PPIs within cellular systems offers fundamental understandings into the functioning of a specific protein. One of the main objectives of biological research has been the analysis and comprehension of molecular interactions within cells. Constructing maps that work out not only direct, binary protein-protein interactions but also incorporate information on indirect interactions is a significant challenge; these interactions include those between proteins that do not directly interact with each other but are part of the same multiprotein complex and proximal protein networks. To identify interactomes, methods must also consider the dynamic nature of various cellular processes and overcome the technical challenges presented by weak protein-protein interactions. The present study employs the innovative BioID proximal-labeling technique to disclose the extensive interactome of LRRC8A in human cells. The reliable cellular localization of biotinylated proteins was validated using LRRC8A-BioID in HEK293 cells before performing MS analysis (Fig. 5). By conducting three independent MS runs, approximately 120 proteins were identified, which represent around 10% of all proteins. To determine the significance of the BioID results, enrichment analyses were conducted using GO and PR (Figs. 6 and 7).
Although BioID and RNA-Seq are distinct in their focus, with BioID focusing on immediate protein interactions and RNA-Seq on gene expression dynamics, the combined results of these methods enrich our understanding of LRRC8A’s intricate role. LRRC8A has been identified as a key regulatory factor in crucial cellular pathways, as illustrated in Figs. 3 and 5. These findings are relevant in the context of colorectal carcinoma, where LRRC8A may regulate mechanisms impacting intercellular communication, metabolic signaling, and cellular stress responses. Disrupted calcium homeostasis could facilitate tumor progression. Recently, it has been revealed that LRRC8A is one of the components of the exosomes released from colon cancer HCT116 cells. Exosomes play a crucial role in intercellular communications within the microscopic tumor environment, facilitating the progression of colon cancer. Recently, it has been revealed that LRRC8A is one of the components of the exosomes released from colon cancer HCT116 cells. Exosomes play a crucial role in intercellular communications within the microscopic tumor environment, facilitating the progression of colon cancer [62]. Additionally, compromised cellular junctions might enhance metastatic potential, leading to a more aggressive and invasive tumor phenotype. Our research also examines hyperinsulinemia, which is prevalent in colorectal cancer. This condition could worsen tumor growth by promoting cellular proliferation and inhibiting apoptosis. Additionally, compromised cellular junctions might enhance metastatic potential, leading to a more aggressive and invasive tumor phenotype. Our research also touches on hyperinsulinemia, prevalent in colorectal cancer, which could exacerbate tumor growth by promoting cellular proliferation and inhibiting apoptosis. The calcium pathway, including IP3R, is implicated in resistance to cell destruction, including evasion of NK cell-mediated killing, positioning it as a possible therapeutic target.
EZR, a member of the ERM protein family, is a common component in RNAseq and BioID. Our data shows that EZR interacts with LRRC8A, and its expression is upregulated in KO cells (Table 3 and Fig. 3D). EZR acts as a linker connecting the actin cytoskeleton to the plasma membrane. The interaction is essential for maintaining cell shape, polarity, and surface structure integrity. EZR participates in several signaling pathways that regulate cell survival, proliferation, and motility [63]. It is involved in the Rho signaling pathway, which impacts actin filament assembly and disassembly, essential for cell movement and structural integrity. EZR also interacts with proteins such as PI3K (phosphoinositide 3-kinase), influencing pathways that control cell growth and survival. EZR interacts with various other cellular proteins, including CD44, a cell-surface glycoprotein involved in cell-cell interactions, cell adhesion, and migration. It interacts with membrane proteins such as ICAM-1 and VCAM-1, which are involved in leukocyte trafficking and immune responses. [64–69]. According to data from The Cancer Genome Atlas (TCGA), EZR expression is significantly decreased in colon cancer, with downregulation observed in 78% of colon adenocarcinoma (COAD) cases and 25% of rectal adenocarcinoma (READ) cases. In our models with LRRC8A KO cells, we observed an upregulation of EZR. This increase may indicate a compensatory mechanism in response to the downregulation of EZR observed in COAD. This could reflect adaptive changes in the cellular environment, or the activation of specific pathways triggered by the absence of LRRC8A.
Additionally, we have conducted an in-depth analysis of long non-coding RNAs (lncRNAs), which are increasingly recognized for their critical roles in cancer biology. LncRNAs, which do not produce proteins, play a crucial role in modulating gene expression and influencing key cellular processes such as growth, survival, and metastasis. They can function as either oncogenes or tumor suppressors. KLRL1-AS1 (ENSG00000245648) and CASC19 (ENSG00000254166), both down-regulated, exhibit atypical expression patterns in colon cancer, indicating significant roles in its pathology (Table 2 and Fig. 4A). KLRL1-AS1 is typically upregulated in colorectal cancer (CRC) cells. It affects cell proliferation by influencing the cell cycle and DNA damage response under TP53 regulation. Koldo Garcia-Etxebarria et al. identified a critical SNP within KLRL1-AS1 (rs10845123) that correlates with CRC prognosis and impacts five-year patient survival rates. [70]. Conversely, Wang et al. suggest that CASC19 acts as an oncogene in CRC progression, potentially serving as a valuable biomarker for CRC diagnosis and treatment [71]. This aligns with our findings where CASC19’s down-regulation in LRRC8A-KO cells is associated with decreased tumor growth. Additionally, the lncRNA ENSG00000250337, known as PURPL (a p53 level regulator increased by p53), exhibits up-regulation in LRRC8A-KO cells (Table 2 and Fig. 4B). This finding is particularly compelling as it contradicts studies by Li et al., who showed that depleting PURPL in colorectal cancer cells increases basal p53 levels and hinders growth, indicating that in its absence, MYBBP1A more effectively stabilizes p53 [72]. This complex interaction may not play a significant role in the reduced tumor growth observed in LRRC8A-KO cells [73, 74]. Also, two novel lncRNAs, ENSG00000277991 and ENSG00000258927, were found to be up-regulated (Table 2 and Fig. 4B). It is worth noting that ENSG00000258927 has only been studied in the context of ovarian cancer and is a newly discovered, uncatalogued lncRNA with limited information on its expression [73, 74].
Considering the limited information available on non-coding RNAs, these findings are significant and could be a valuable focus for further bioinformatic investigations using TCGA databases or other RNA sequencing analyses in the field of colorectal cancer. Additionally, these two lncRNAs may have the potential to serve as markers for identification, prognosis, or even therapeutic targeting in colorectal cancer.
Interestingly, among the LRRC8A interacting proteins, more than 40 have previously been found to interact with the LRRC8A ancestor, PANX1 [75]. Among these, ACBD3, CTNND1, DLG1, FERMT2, NUMB and NUMBL are common proteins found in the cell-cell junction that have been identified. DLG1 is a MAGUK scaffolding protein that regulates the localization and function of multiple ion channels [76, 77]. We also identified three proteins belonging to the 4.1 family: EPB41, EPB41L2 and EPB41L3. These proteins have essential functions in the assembly and maintenance of specific transmembrane protein complexes in the plasma membrane [78, 79]. 4.1 R performs a pivotal function in preserving the regularity of cell structure by bridging the spectrin-actin cytoskeleton to plasma membrane proteins and attracting DLG1 to the membranes. While hitherto, no research has linked VRAC to DLG1 or 4.1 R, it has been shown to associate with other channels such as NaV1.5, a sodium channel, and TRPC4, a non-selective cation channel that allows the transfer of calcium [80, 81].
Another group of significant interactors includes cadherin-binding proteins, such as AHNAK. The interaction between AHNAK and PANX1 was recently demonstrated using BioID techniques. The finding was further confirmed by co-immunoprecipitation, in combination with mass spectrometry [75]. The correlation between AHNAK and PANX1 led to initial insights into the mechanisms by which PANX1 suppresses malignant properties in rhabdomyosarcoma [75].
The dynamic mechanism of post-translational modification regulates several phases of the ion channel life cycle, such as maturation, trafficking, signalling and regulation [82–86]. Phosphorylation is thought to alter the probability of channel opening, gating, voltage dependence, desensitisation, and permeability. In the case of VRAC, post-translational modifications may focus on modifying the intracellular loop connecting the transmembrane pore to the leucine-rich repeat domain. This region contains numerous phosphorylation sites for amino acid residues such as serine, threonine, and tyrosine. Several studies suggest that PKC (protein kinase C) is a main regulator of VRAC activation [87–92]. Furthermore, a recent publication suggests that hypotonicity can significantly influence VRAC activation through the involvement of PKD [93]. Interestingly, AKAP12 and PKN2, two proteins related to PKC and PKA were identified in the list of interactors. AKAP12 controls the subcellular localisation of PKA and PKC through tethering. It can interact with these protein kinases and phosphatases to facilitate signalling pathways. PKN2 plays a critical role in the organisation of the actin cytoskeleton by activating Rho GTPase in coordination with PKC. We also found other kinase-related proteins, such as ACBD3, CCDCD88A, EFR3A, GAB1, PAK4, PEAK1, PTPN1, PTPN13, ROR2, KIDINS220, TKK, VRK2 and WDR45B (see Fig. 7D).
Analysing the Reactome pathways (FDR < 0.05), the physical proximity of LRRC8A to proteins linked to Rho subfamilies was found (Fig. 5E). Our BioID analysis results align with previous findings that suggest a potential association between intracellular GTPγS and VRAC, implying that the Rho pathway may regulate VRAC activation, as reported in [27, 94, 95]. Rho proteins are thought to shift between their active form (Rho-GTP) on the cell membrane to an inactive form (Rho-GDP) in the cytoplasm. Rho GTPases regulate various cellular processes that demand dynamic reorganization of the cytoskeleton, such as cell migration, adhesion, division, polarity establishment, and intracellular transport [96, 97]. Previous research has shown that intracellular application of GTPγS, a non-hydrolysable analogue of GTP, can activate VRAC currents, even in isotonic conditions [27, 28, 98]. In contrast, GDPβS, a non-hydrolysable analogue, results in the time-dependent inhibition of VRAC [99]. Although the role of GTPγS has been subject to extensive scrutiny, the current understanding of how GTPγS modulates VRAC signalling remains limited.
As mentioned previously, VRAC is an important pathway for anion transport during cell volume regulation. It is typically activated in response to cell swelling, but how the channel senses swelling remains unclear. Lemonnier and co-workers provided evidence for colocalization of VRAC with store-operated Ca2+ channels and showed that activation of VRAC is strongly dependent on Ca2+ release through IP3R [100]. They concluded that VRAC is regulated within Ca2+ microdomains. Similarly, Akita and collaborators found that activation is regulated by high concentration regions of intracellular Ca2+ in the immediate vicinity of open Ca2+-permeable channels, called Ca2+ nanodomains [101, 102]. However, Liu et al. showed that intracellular Ca2+ is necessary but not sufficient to activate LRRC8A-mediated currents [103]. Our BioID analysis suggests a potential link between the VRAC channel and the regulation of cellular calcium. The list includes ATP2B1, ESYT1, JPH1, STIM1, SPTAN1 and VAPB (Fig. 7F). Of particular interest is the correlation with STIM1. STIM1 is functionally related to the CRAC channel, allowing the influx of Ca2+ ions from the extracellular space into the cytosol upon depletion of stored Ca2+ ions in the ER [104]. After Ca2+ depletion, STIM1 forms oligomers and migrates to ER-PM junctions. The subsequent interaction of STIM1 with ORAI1 and ORAI2 causes the opening of the CRAC channels [105]. ATP2B1 is a plasma membrane ATP dependent calcium pump. It regulates insulin sensitivity through Ca2+/calmodulin signalling pathway by regulating AKT1 activation and NOS3 activation in endothelial cells [106]. ESYT1 binds Ca2+ and translocates to sites of contact between the endoplasmic reticulum and the plasma membrane in response to elevated cytosolic Ca2+ levels. It assists in the tethering of the ER to the plasma membrane [107]. Furthermore, JPH1 provides a structural foundation for functional crosstalk between the cell surface and intracellular Ca2+ release channels [108]. SPTAN1 is a protein with an EF-hand domain that appears to be involved in secretion, and it interacts with calmodulin in a Ca2+-dependent manner. This interaction allows it to move the cytoskeleton across the membrane [109].
Conclusion
This study investigates the correlation between the expression levels of VRAC genes and patient survival across different cancer types. The research uncovers a noteworthy correlation between LRRC8A expression and prognosis in specific oncological contexts. Furthermore, RNA-Seq analysis was employed to examine DEGs in CRC cells that lack LRRC8A, providing novel insights into the genetic alterations caused by the absence of this ion channel. This analysis explores the LRRC8A interactome in living mammalian cells, providing insight into the molecular dynamics of VRAC.
The study of PPIs and gene regulatory systems is crucial for comprehending the genetic and molecular basis of diseases and advancing therapeutic developments. BioID is an effective method for identifying interactions, regardless of the solubility properties of the proteins involved. This technique is particularly useful for analysing proteins that are challenging to purify or have variable solubility, such as ion channels. However, it is essential to recognise the inherent limitations of BioID, like any experimental technique. Future validations will be necessary, but these initial findings are crucial as they represent some of the first focused explorations of the VRAC channel members. This data establishes the foundation for more detailed studies on the functions and interactions of VRAC channels. It highlights the importance of these early insights in the fields of cellular biology and physio/pathology.
Materials and Methods
Plasmid constructs
The wild-type human LRRC8A gene was subcloned from the “8a-pcdna3-hektor” plasmid provided by Dr. Raul Estevez into pcDNA3.1-MCS-BirA(R118G)-HA (Addgene #36047(Roux, Kim et al. 2012)) using primers that modified the stop codon and appended restriction enzyme sequences to the 5’ overhangs. The reaction made use of the following PCR primers: The reaction utilized the NheI forward (5′-aaagctagcaccatgattccggtgacagagctccgctac-3’) and EcoRI reverse (5’- tgcgaattctgcggccttcagccctccacag-3’) primers. The amplification was conducted using the Phusion enzyme (NEB England). Confirmation of successful LRRC8A-BirA* fusion protein-containing clones was assessed via sequencing (Eurofins Genomics).
Cell cultures
The study used HEK293 cells and HCT116 cells (received as a gift from Prof. Szabò at the University of Padua). The cells were cultured in Dulbecco’s modified Eagle medium (DMEM) with 10% fetal bovine serum (FBS), 10 mM HEPES, 100 U/ml penicillin, and 100 U/ml streptomycin along with 1X non-essential amino acids (NAA) from Gibco. The cells were grown at 37 °C under a humidified atmosphere with 5% CO2. T25 flasks were used to culture the cells. Upon reaching 70 to 80% confluency, the cells were trypsinized and seeded in culture flasks or Petri dishes with a 10 to 20% density. Coverslips were included in the Petri dishes used for the experiments. For biotinylation assays, the cells were incubated for 24 hours at 37 °C in media with 50 μM of biotin, under 5% CO2.
Survival analysis using Kaplan–Meier Plotter and GEPIA
The correlation between LRRC8A expression and clinical outcomes in cancer was evaluated through the analysis of Kaplan-Meier plots and survival analysis module available on SurvivalGenie (https://bbisr.shinyapps.winship.emory.edu/SurvivalGenie/, accessed on 10 November 2023). The PP-network of LRRC8A-E was analysed using the Search Tool for Retrieval of Interacting Genes/Proteins (https://string-db.org/, accessed on 28 August 2023) and GeneMANIA (http://www.genemania.org, accessed on 10 November 2023).
Generation of HCT116 LRRC8A knockout cell lines using CRISPR/Cas9 technology
The CRISPR/CAS9 method employed the pSpCas9n(BB)-2A-Puro (PX459) plasmid (Addgene Plasmid #48139 Zhang Lab), according to the instructions provided by Ran et al. (2013). The sgRNA guides were designed specifically to remove exon 3. The following sgRNA sequences were utilized: HsLRR8A_u11 FOR CACCGGCTATCTGCGCGTCGGCTGT, HsLRR8A_u11 REV AAACACAGCCGACGCGCAGATAGCC, HsLRR8A_u12 FOR CACCGTGGCTCTGCTATCTGCGCGT, and HsLRR8A_u12 REV AAACACGCGCAGATAGCAGAGCCAC were used as sgRNA sequences. The following sgRNA sequences were utilized: HsLRR8A_d13 FOR CACCGCCTGGGGCCGCTTGTGAGTC, HsLRR8A_d13 REV AAACGACTCACAAGCGGCCCCAGGC, HsLRR8A_d14 FOR CACCGCCTGGCTGTCCGGGAGTTCT, and HsLRR8A_d14 REV AAACAGAACTCCCGGACAGCCAGGC were used as sgRNA sequences. The annealed and phosphorylated guides were connected to the vector at the BbsI (NEB) restriction site beneath the U6 promoter and authenticated through sequencing. A total of one hundred thousand HCT116 cells were cultured in a six-well plate. The next day, several pairs of single guides were employed to deliver 2.5 micrograms of plasmid DNA (1.25 μg per guide) to the HCT116 cells via the TransIT®-LT1 transfection reagent (Mirus). 48 hours post-transfection, the cell medium had puromycin at a concentration of 1.5 μg/ml (Gibco), which persisted for 96 hours. After selection, the cells were stepwise diluted to 0.5 cells per well in a 96-well plate to limit the presence of multiple cells in a single well. Actively growing cells were gathered and centrifuged at 600 g for 5 minutes. The medium was discarded, and the DNA was extracted from the pellet using a MyTaq Extract-PCR kit (Meridian Bioscience) in accordance with the manufacturer’s guidelines. The quantity was calculated using a Thermo Scientific ND2000 spectrophotometer.
Evaluation of cell proliferation
25,000 HCT116 cells were seeded in 12-well plates. The cells were detached from the plates at 24-, 36-, 48- and 72-hours intervals using 100 μL Gibco trypsin and then diluted in 100 μL of medium. Cell proliferation was assessed daily by using the Logos Biosystem’s LUNA™II Automated Cell Counter.
Evaluation of cell migration
The potential for cell migration was measured using wound-scratch experiments. HCT116 cells were seeded in 24-well culture plates and allowed to grow until they reached 80-90% confluence. A treatment of 5 μg/mL mitomycin C (Sigma Aldrich) was administered one hour before the scratch. A plastic 200 μl pipette tip was used to scratch the monolayer of cells. Following a wash in PBS, the medium was substituted with phenol red-free medium containing 5% FBS. The same fields were photographed at 0, 15, 20, and 24 hours after performing the scratch with a Leica DMI4000 inverted microscope. The area of the scratch was measured using the ImageJ software (NIH). The migration rate was quantified as a percentage of the initial scraped area.
Colony formation assay
A total of six hundred HCT116 cells were cultured in a standard culture medium for 6 days using a 6-well plate. The medium was then discarded, and the cells were washed with PBS twice and then fixed with 3.8% paraformaldehyde for 30 minutes. After three more washes with PBS, the cells were stained with 0.1% crystal violet at room temperature for 15 minutes. The PBS was used to remove the staining until the colonies were cleared. At a magnification of 1 × 0.5, the Leica stereo microscope MZ16F was used to capture the images. The software used to identify the number and dimensions of the colonies was ImageJ (NIH).
Quantification of apoptotic events
HCT116 cells were cultivated in DMEM medium supplemented with 10% FBS and seeded in a six-well plate at a density of 5 × 10^5 cells. After 24 hours, the cells were collected, washed with PBS, and centrifuged at 300 g. The cells were subsequently stained with 2.5 µl of annexin V and propidium iodide (PI) in binding buffer (Thermo Fisher) for 15 minutes in the dark at 4 °C to determine apoptotic events. The BD LSR Fortessa X-20 flow cytometer was employed to evaluate cell populations for annexin V-positive or double-positive cells. Histograms for APC-A, PE-A, SSC-A, and FSC-A were obtained.
Cell Cycle Analysis
HCT116 cells were cultivated using DMEM medium supplemented with 10% FBS, and then cultured in a 6-well plate with a density of 5 × 10^5 cells. After 24 hours, the cells underwent cold PBS washing twice and were treated with 70% ethanol-based cell permeabilization at ice-cold temperatures for an hour. Following incubation at 37 °C with a solution containing 50 µg/ml propidium iodide (Sigma-Aldrich) and 10 ng/ml RNase A (Qiagen) for one hour, the cells underwent centrifugation at 1000 g/min, were washed with PBS, and analysed using BD LSR Fortessa X-20 flow cytometry. Following incubation at 37 °C with a solution containing 50 µg/ml PI (Sigma-Aldrich) and 10 ng/ml RNase A (Qiagen) for one hour, the cells underwent centrifugation at 1000 g/min, were washed with PBS, and analysed using BD LSR Fortessa X-20 flow cytometry. Cell cycle stages were determined, and quantification of PE-A, FSC-AA, and SSC-A histograms was performed using BD FACS Diva 9.0.
RNA-seq analyses: alignment, pre-processing, and differential expression
HCT116 cells were seeded in a 6-well plate with a density of 0.4 × 10^6 cells for WT and 0.5 × 10^6 cells for KO. RNA was extracted using RNeasy Mini Kit (Qiagen) according to the recommended protocol. Reads were aligned to the reference genome with STAR (v 2.7.10a) [110] and quantified with RSEM (v1.3.1). The indexed genome was built with RSEM starting from Ensembl’s Homo Sapiens DNA primary assembly (release 106) [111]. To identify the differentially expressed genes we used the edgeR R package [112]. We provided as input the filtered raw counts with the design matrix defined by the dichotomous variables for the different clones. The RLE normalization was applied to the samples. False Discovery Rate (FDR) less than 0.01 was used to significantly select DEG.
The LncRNome analyses was performed using the Cancer LncRNome Atlas is a comprehensive database or resource that catalogs and characterizes long non-coding RNAs (lncRNAs) associated with various types of cancer (http://fcgportal.org/TCLA/search.php).
Transfections and the generation of stable cell lines
The TransIT®-LT1 transfection reagent (Mirus) was used to transfect cells with LRRC8A-BirA* following the manufacturer’s recommended protocols for BioID. After transfection, HEK293 cells were selected with 750 μg/mL Geneticin (G418) to create stable cell lines. Monoclonal cells were isolated and grown following colony formation, and their stable expression was confirmed using Western blotting. Stable HEK293 cell lines were maintained using G418.
Patch-clamp analyses
HEK-5X-KO lrrc8-/- cells used for patch clamp recordings were knock-out for all five genes encoding lrrc8 subunits [14] and were kindly provided by Thomas Jentsch (Berlin). Cells were cultured in DMEM (Pan Biotech) supplemented with 10% FBS, 1% penicillin/streptomycin and 1% glutamine and maintained at 37 °C in a 5% CO2, 100% humidity atmosphere. Cells were grown on plastic tissue culture dishes and splitted every 3–4 days. Cells were co-transfected using the effectene reagent (Qiagen) with the BirA-tagged LRRC8A plasmid and LRRC8E in pCDNA3.1 as in [92]. For the identification of positively transfected cells, a plasmid encoding the CD8 antigen was co-transfected. The transfected cells were identified by microbeads coated with anti-CD8 antibodies (Dynabeads M-450 CD 8; ThermoFisher) as described in [113].
Currents were recorded 24–36 h after transfection. The standard current-voltage protocol (IV) for stimulation consisted of 500 ms-long voltage steps ranging from -80 to 120 mV in 20 mV increments. Current response to the various stimuli were monitored using the “time course protocol”, which consisted of successive steps of 50 ms pulses to −75, −25, 0, 25, and 75 mV every 5 s.
The standard extracellular isotonic solution contained in mM: 145 NaCl, 6 KCl, 1.5 CaCl2, 1 MgCl2, 10 HEPES, 10 glucose (pH 7.4, 310 mOsm). Hypotonic solution contained in mM: 105 NaCl, 6 CsCl, 1.5 CaCl2, 1 MgCl2, 10 HEPES, 10 glucose (pH7.4, 230 mOsm). The standard pipette solution to monitor VRAC activation upon hypotonic perfusion contained (in mM) 100 K-Gluconate, 40 CsCl, 2 MgCl2, 1.9 CaCl2, 5 EGTA-NMDG,1 Na2ATP, and 10 HEPES-NMDG, pH 7.3 (290 mOsm).
Immunofluorescence
The cells that were transfected with BioID constructs were fixed using 3.8% paraformaldehyde and permeabilized in 0.1% Triton-X100 in phosphate buffer saline (PBS) for five minutes. The coverslips were incubated with primary antibody, anti-HA (Abcam, 1:500), overnight. After washing in PBS, the coverslips were then incubated with streptavidin coupled with Alexa Fluor 488 (Invitrogen, 1:1000), and goat anti-rabbit 568 (Invitrogen, 1:500) secondary antibodies. The coverslips were washed three more times using PBS, then mounted on glass slides using ProLong Gold containing 4′,6′-diamidino-2-phenylindole (DAPI; Invitrogen). Images of the cells were captured using a 63x oil objective with a Leica SP5 confocal microscope (Leica Microsystem, Wetzlar, Germany).
Membrane extraction
The ProteoExtract kit (Merck/Sigma-Aldrich) was used according to the recommended protocol to separate and collect soluble and membrane protein fractions.
Immunoblotting
Whole-cell extracts were prepared using a lysis buffer composed of 25 mM Tris-HCl pH 7.8, 2.5 mM EDTA, 10% (v/v) glycerol, and 1% (v/v) NP40, supplemented with a Protease Inhibitor Cocktail (Sigma) and 2 mM DTT. The lysates were resolved by SDS-PAGE using 4-12% ExpressPlus® PAGE gels (GenScript) and then transferred to a membrane (polyvinylidene fluoride, PVDF, Amersham™Hybond™ P 0.45μm). The membranes were checked for equal loading using a Ponceau solution, followed by washing and blocking for one hour in 5% skim milk in Tris-buffered saline (TBS: 10 mM Tris, 150 mM NaCl, pH 7.4). The blotted proteins were detected by incubation overnight at 4 °C with the following antibodies after washing three times with TBS-T (TBS plus 0.05% Tween). All Western blots were developed using the Clarity Western ECL substrate (Bio-Rad) and then imaged on the ChemiDoc Imager (Bio-Rad). We used the following primary antibodies: Streptavidin-HPR (1:40.000, Thermofisher, #21130), anti-PMCA (1:2000, Invitrogen #MA3-914), anti-actin (1:2000, Millipore #MAB1501), anti-HA (1:1000, Abcam #AB9110), anti-LRRC8A (1:1000, Bethyl #A304-175A).
The detection of biotinylated proteins was conducted with specific adaptations tailored for nitrocellulose membranes. The detection of biotinylated proteins was carried out with the following modifications. After transfer, membranes (nitrocellulose, Amersham™Hybond™ P 0.45μm) were blocked for 30 minutes in 1% BSA in phosphate-buffered saline (PBS) containing 0.2% Triton X-100 and then incubated in the same buffer with HRP-conjugated streptavidin (Thermo Fisher Scientific, 21130, 1:40000) for 40 minutes.
Streptavidin pull-down of biotinylated proteins
For MS analysis, large-scale pull-downs were conducted by seeding 1 × 106 cells expressing BioID-LRRC8A fusion proteins on four 10-cm plates. The cells were grown to 70–80% confluence and then incubated in complete media with 50 μM biotin for 24 hours. Cells were washed with PBS and lysed in RIPA lysis buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1% (v/v) NP-40, 0.1% (w/v) SDS, 0.5% (w/v) sodium deoxycholate, 1 mM DTT, and protease inhibitors) at room temperature. The lysate was then incubated on ice for 15 minutes, as described in a previous study [114]. The cell lysate samples were centrifuged at 13000 × g for 10 minutes at 4 °C. The supernatant underwent Streptavidin-based pull-down using MyOneDynabeads Streptavidin C1 (Thermo Fisher Scientific), following the method outlined in [115].
To test for protein biotinylation, we reserved around 10% of the whole sample specifically for Western blot analysis. The proteins were extracted from the beads by boiling them with SDS for 5 minutes at 95 °C. The remaining sample was used for MS analysis.
Mass spectrometry
The raw IsobarQuant output files (protein.txt – files) were processed using the R programming language (www.r-project.org). Only proteins that were quantified with at least two unique peptides were considered for analysis. Raw TMT reporter ion intensities (‘signal_sum’ columns) were first cleaned for batch effects using limma [116] and further normalized using vsn (variance stabilization normalization [117]). The differential expression of the proteins was tested using the limma package. The replicate information was added as a factor in the design matrix given as an argument for the limma ‘lmFit’ function. 122 potentially interacting proteins were defined as proteins with significant change (fdr <= 0.05 and fold-change >= 2) in comparisons ‘LRRC8A-BirA*-HA#1 Biotin vs LRRC8A-BirA*-HA#1 noBiotin’ or ‘LRRC8A-BirA*-HA#2 Biotin vs LRRC8A-BirA*-HA#2 noBiotin’.
LC-MS/MS acquisition
An UltiMate 3000 RSLC nano LC system (Dionex) was used, which had a trapping cartridge (µ-Precolumn C18 PepMap 100, 5 µm, 300 µm i.d. x 5 mm, 100 Å) and an analytical column (nanoEase™ M/Z HSS T3 column 75 µm x 250 mm C18, 1.8 µm, 100 Å, Waters). The trapping was performed for 6 minutes using a constant trapping solution flow of 0.05% trifluoroacetic acid in water at a rate of 30 µL/min on the trapping column. The peptides were then eluted using solvent A (0.1% formic acid in water, 3% DMSO) with a constant flow rate of 0.3 µL/min through the analytical column. The proportion of solvent B (0.1% formic acid in acetonitrile, 3% DMSO) was gradually increased during this process. The outlet of the analytical column was directly connected to an Orbitrap Fusion™ Lumos™ Tribrid™ Mass Spectrometer (Thermo Fisher) utilizing the Nanospray Flex™ ion source in the positive ion mode.
The peptides were received by the Fusion Lumos using a Pico-Tip Emitter with a 10 µm tip a diameter of 360 µm and an inner diameter of 20 µm (New Objective) while applying a spray voltage of 2.4 kV. The capillary temperature was maintained at 275 °C. The whole mass scan was obtained in the orbitrap with a resolution of 120,000 in profile mode. Mass ranges from 375 to 1500 m/z were employed. The filling time was restricted to a maximum of 50 ms and no more than 4 × 105 ions. Data-dependent acquisition (DDA) with a fill time of 94 ms and a limitation of 1 × 105 ions was carried out. Orbitrap’s resolution was set at 30,000. We used a collision energy of 38 in normalized units. We acquired MS2 data in profile mode.
MS data analysis – Isobarquant
IsobarQuant and Mascot (v2.2.07) were used to process the acquired data, which were searched against a Uniprot Homo sapiens proteome database (UP000005640) containing common contaminants and reversed sequences. The following modifications were included in the search parameters Carbamidomethyl (C) and TMT11 (K) (fixed modification), Acetyl (protein N-terminal), Oxidation (M) and TMT11 (N-terminal) (variable modifications). A mass error tolerance of 10 ppm was set for the full scan (MS1) and 0.02 Da for the MS/MS spectra (MS2). Other parameters were: trypsin as protease with a maximum of two missed cleavages allowed; a minimum peptide length of seven amino acids; at least two unique peptides were required for protein identification. The false discovery rate at the peptide and protein level was set at 0.01.
Mass spectrometry data analysis
The raw IsobarQuant output files (protein.txt – files) were processed using the R programming language (www.r-project.org). Only proteins that were quantified with at least two unique peptides and identified in all mass spec runs were considered for analysis. Raw reporter ion intensities (signal_sum columns) were first cleaned for batch effects using limma [116] and further normalized using vsn (variance stabilization normalization [117]. Missing values were imputed with the ‘knn’ method using the Msnbase package [118]. The differential expression of the proteins was tested using the limma package. The replicate information was added as a factor in the design matrix given as an argument for the limma lmFit function. Furthermore, the imputed values were given a weight of 0.05 in the ‘lmFit’ function. A protein was annotated as a hit with a false discovery rate (FDR) smaller than 5% and a fold change of at least 100% and as a candidate with an FDR below 20% and a fold-change (FC) of at least 50%.
Bioinformatic analysis
We used the list of selected proteins to identify significantly enriched functional categories. We conducted enrichment analyses using the clusterprofiler R package (Wu, Hu et al., 2021) on the Gene Ontology (GO) categories of biological process (BP), molecular function (MF), and cellular component (CC), as well as the Reactome and KEGG pathway databases [119]. We employed the False Discovery Rate (FDR) to control for multiple testing. We identified significantly enriched GO terms and Reactome pathways using a false discovery rate (FDR) threshold of 0.05. To reduce the redundancy of significant GO terms, we employed semantic similarity distance, which is implemented in the R package rrvo (https://ssayols.github.io/rrvgo). The findings were graphically summarized using dot, scatter, and tree plots. The maps that displayed significantly enriched KEGG pathways were color-coded according to the logarithmic fold change (logFC) of the proteins.
Statistical analysis
The data is presented as means ± standard error of the mean. Data analysis was performed using the GraphPad Prism 8 software. Two-way ANOVA with Bonferroni’s test was used for comparing the data due to two variables. We compared two groups using the unpaired Student’s t-test. Additional statistical information is available in the figure legends.
Supplementary information
Acknowledgements
We thank Dr. Mandy Rettel at the Proteomics Core Facility, EMBL Heidelberg, Germany, for the Mass Spec support. Furthermore, V.Ch. acknowledges and appreciates Prof. I. Szabò for insightful discussions and support in establishing her independent laboratory. V.Ch. expresses gratitude for the financial support received from various sources, including DOR (Dotazione Ordinaria della Ricerca dipartimentale) projects — Calls 2021-2022-2023 entitled “Ion channel proteins and their role in cell signalling”, the local SEED-PRID project 2021 entitled “Setting up in vivo interactome for cancer research”, the Integrated Budget for Interdepartmental Research 2023 titled “The protein-protein interactions network of the Volume-Regulated Anion Channel (VRAC): from uncertainty to molecular details” (BIRD-PRID, grant number BIRD239198/23), the Projects of National Interest Research 2022 (PRIN, grant number 2022ZY7ATN) and the research project “Mitochondrial ATP-sensitive potassium channels in health and disease” sponsored by Fondazione Cassa di Risparmio di Padova e Rovigo — Scientific Excellence Research Grant 2021, M.P. was partially funded by the European Union - NextGenerationEU and by the Ministry of University and Research (MUR), National Recovery and Resilience Plan (NRRP), Mission 4, Component 2, Investment 1.5, project “RAISE - Robotics and AI for Socio-economic Empowerment” (ECS00000035). MALAS expresses gratitude for the fellowship provided by WWCR (grant number 22-0348).
Author contributions
Conceptualization VC, MF, EP VCh; Methodology, VC, MF, EP, MB, SS, SB, FS, AV, MALAS, CR, MP, VCh; Investigation, VC, MF, EP, MB, SS, SB, FS, AV, MALAS, CR, MP, VCh; Writing – Original Draft, VCh, VC, MF, EP, SS; Editing- MB, MP. Funding, VCh; MP; Supervision, VCh. All authors read, edited, and approved the final manuscript.
Data availability
The datasets used in the current study are available from the corresponding authors on reasonable request.
Competing interests
The authors declare no competing interests.
Consent for publication
All authors gave consent for publication of the manuscript in Cell Death Discovery.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Veronica Carpanese, Margherita Festa, Elena Prosdocimi.
Change history
8/29/2024
A Correction to this paper has been published: 10.1038/s41420-024-02123-y
Supplementary information
The online version contains supplementary material available at 10.1038/s41420-024-02032-0.
References
- 1.Al-Sabi A, Abd El-Aziz TM, Yu P, Rowe AH, Wulff H. Editorial: Ion channels in health and disease. Front Physiol. 2022;13:1093210. 10.3389/fphys.2022.1093210 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Banderali U, Leanza L, Eskandari N, Gentile S. Potassium and chloride ion channels in cancer: a novel paradigm for cancer therapeutics. Rev Physiol Biochem Pharm. 2022;183:135–55. 10.1007/112_2021_62 [DOI] [PubMed] [Google Scholar]
- 3.Bachmann M, Pontarin G, Szabo I. The contribution of mitochondrial ion channels to cancer development and progression. Cell Physiol Biochem. 2019;53:63–78. 10.33594/000000198 [DOI] [PubMed] [Google Scholar]
- 4.Checchetto V, Leanza L, De Stefani D, Rizzuto R, Gulbins E, Szabo I, Mitochondrial K. Mitochondrial K+ channels and their implications for disease mechanisms.Pharmacol Ther. 2021;227:107874. 10.1016/j.pharmthera.2021.107874 [DOI] [PubMed] [Google Scholar]
- 5.Stoilova-McPhie S, Ali S, Laezza F. Protein-protein interactions as new targets for ion channel drug discovery. Austin J Pharmacol Ther. 2013;1:5. [PMC free article] [PubMed] [Google Scholar]
- 6.Lee A, Fakler B, Kaczmarek LK, Isom LL. More than a pore: ion channel signaling complexes. J Neurosci. 2014;34:15159–69. 10.1523/JNEUROSCI.3275-14.2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gururaja Rao S, Patel NJ, Singh H. Intracellular chloride channels: novel biomarkers in diseases. Front Physiol. 2020;11:96. 10.3389/fphys.2020.00096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Martinez AH, Mohiuddin SS. Biochemistry, Chloride Channels. 2023 Jul 17. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2024. [PubMed]
- 9.Xu R, Hu Y, Xie Q, Zhang C, Zhao Y, Zhang H. et al. LRRC8A is a promising prognostic biomarker and therapeutic target for pancreatic adenocarcinoma. Cancers. 2022;14:5526. 10.3390/cancers14225526 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Planells-Cases R, Lutter D, Guyader C, Gerhards NM, Ullrich F, Elger DA, et al. Subunit composition of VRAC channels determines substrate specificity and cellular resistance to Pt-based anti-cancer drugs. EMBO J. 2015;34:2993–3008. 10.15252/embj.201592409 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Pedersen SF, Klausen TK, Nilius B. The identification of a volume-regulated anion channel: an amazing Odyssey. Acta Physiol. 2015;213:868–81. 10.1111/apha.12450 [DOI] [PubMed] [Google Scholar]
- 12.Pedersen SF, Okada Y, Nilius B. Biophysics and physiology of the Volume-Regulated Anion Channel (VRAC)/Volume-Sensitive Outwardly Rectifying Anion Channel (VSOR). Pflug Arch. 2016;468:371–83. 10.1007/s00424-015-1781-6 [DOI] [PubMed] [Google Scholar]
- 13.Jentsch TJ, Lutter D, Planells-Cases R, Ullrich F, Voss FK. VRAC: molecular identification as LRRC8 heteromers with differential functions. Pflug Arch. 2016;468:385–93. 10.1007/s00424-015-1766-5 [DOI] [PubMed] [Google Scholar]
- 14.Lutter D, Ullrich F, Lueck JC, Kempa S, Jentsch TJ. Selective transport of neurotransmitters and modulators by distinct volume-regulated LRRC8 anion channels. J Cell Sci. 2017;130:1122–33. 10.1242/jcs.196253 [DOI] [PubMed] [Google Scholar]
- 15.Osei-Owusu J, Yang J, Vitery MDC, Qiu Z. Molecular biology and physiology of Volume-Regulated Anion Channel (VRAC). Curr Top Membr. 2018;81:177–203. 10.1016/bs.ctm.2018.07.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Dwivedi B, Mumme H, Satpathy S, Bhasin SS, Bhasin M. Survival Genie, a web platform for survival analysis across pediatric and adult cancers. Sci Rep. 2022;12:3069. 10.1038/s41598-022-06841-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Qiu Z, Dubin AE, Mathur J, Tu B, Reddy K, Miraglia LJ, et al. SWELL1, a plasma membrane protein, is an essential component of volume-regulated anion channel. Cell. 2014;157:447–58. 10.1016/j.cell.2014.03.024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Voss FK, Ullrich F, Münch J, Lazarow K, Lutter D, Mah N, et al. Identification of LRRC8 heteromers as an essential component of the volume-regulated anion channel VRAC. Science. 2014;344:634–8. 10.1126/science.1252826 [DOI] [PubMed] [Google Scholar]
- 19.Abascal F, Zardoya R. LRRC8 proteins share a common ancestor with pannexins, and may form hexameric channels involved in cell-cell communication. Bioessays. 2012;34:551–60. 10.1002/bies.201100173 [DOI] [PubMed] [Google Scholar]
- 20.Liu H, Polovitskaya MM, Yang L, Li M, Li H, Han Z, et al. Structural insights into anion selectivity and activation mechanism of LRRC8 volume-regulated anion channels. Cell Rep. 2023;42:112926. 10.1016/j.celrep.2023.112926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rutz S, Deneka D, Dittmann A, Sawicka M, Dutzler R. Structure of a volume-regulated heteromeric LRRC8A/C channel. Nat Struct Mol Biol. 2023;30:52–61. 10.1038/s41594-022-00899-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kern DM, Bleier J, Mukherjee S, Hill JM, Kossiakoff AA, Isacoff EY, et al. Structural basis for assembly and lipid-mediated gating of LRRC8A:C volume-regulated anion channels. Nat Struct Mol Biol. 2023;30:841–52. 10.1038/s41594-023-00944-6 [DOI] [PubMed] [Google Scholar]
- 23.Akita T, Okada Y. Characteristics and roles of the volume-sensitive outwardly rectifying (VSOR) anion channel in the central nervous system. Neuroscience. 2014;275:211–31. 10.1016/j.neuroscience.2014.06.015 [DOI] [PubMed] [Google Scholar]
- 24.Mongin AA, Kimelberg HK. ATP potently modulates anion channel-mediated excitatory amino acid release from cultured astrocytes. Am J Physiol Cell Physiol. 2002;283:C569–78. 10.1152/ajpcell.00438.2001 [DOI] [PubMed] [Google Scholar]
- 25.Liu HT, Akita T, Shimizu T, Sabirov RZ, Okada Y. Bradykinin-induced astrocyte-neuron signalling: glutamate release is mediated by ROS-activated volume-sensitive outwardly rectifying anion channels. J Physiol. 2009;587:2197–209. 10.1113/jphysiol.2008.165084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Varela D, Simon F, Riveros A, Jørgensen F, Stutzin A. NAD(P)H oxidase-derived H(2)O(2) signals chloride channel activation in cell volume regulation and cell proliferation. J Biol Chem. 2004;279:13301–4. 10.1074/jbc.C400020200 [DOI] [PubMed] [Google Scholar]
- 27.Doroshenko P, Neher E. Volume-sensitive chloride conductance in bovine chromaffin cell membrane. J Physiol. 1992;449:197–218. 10.1113/jphysiol.1992.sp019082 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Voets T, Manolopoulos V, Eggermont J, Ellory C, Droogmans G, Nilius B. Regulation of a swelling-activated chloride current in bovine endothelium by protein tyrosine phosphorylation and G proteins. J Physiol. 1998;506:341–52. 10.1111/j.1469-7793.1998.341bw.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Burow P, Klapperstück M, Markwardt F. Activation of ATP secretion via volume-regulated anion channels by sphingosine-1-phosphate in RAW macrophages. Pflug Arch. 2015;467:1215–26. 10.1007/s00424-014-1561-8 [DOI] [PubMed] [Google Scholar]
- 30.Syeda R, Qiu Z, Dubin AE, Murthy SE, Florendo MN, Mason DE, et al. LRRC8 proteins form volume-regulated anion channels that sense ionic strength. Cell. 2016;164:499–511. 10.1016/j.cell.2015.12.031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Tang Z, Li C, Kang B, Gao G, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–W102. 10.1093/nar/gkx247 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Noy NF, Shah NH, Whetzel PL, Dai B, Dorf M, Griffith N, et al. BioPortal: ontologies and integrated data resources at the click of a mouse. Nucleic Acids Res. 2009;37:W170–3. 10.1093/nar/gkp440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Whetzel PL, Noy NF, Shah NH, Alexander PR, Nyulas C, Tudorache T, et al. BioPortal: enhanced functionality via new Web services from the National Center for Biomedical Ontology to access and use ontologies in software applications. Nucleic Acids Res. 2011;39:W541–5. 10.1093/nar/gkr469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chandrashekar DS, Karthikeyan SK, Korla PK, Patel H, Shovon AR, Athar M, et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18–27. 10.1016/j.neo.2022.01.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi BVSK, et al. UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19:649–58. 10.1016/j.neo.2017.05.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zhang H, Deng Z, Zhang D, Li H, Zhang L, Niu J, et al. High expression of leucine‑rich repeat‑containing 8A is indicative of a worse outcome of colon cancer patients by enhancing cancer cell growth and metastasis. Oncol Rep. 2018;40:1275–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Liu T, Stauber T. The Volume-Regulated Anion Channel LRRC8/VRAC is dispensable for cell proliferation and migration. Int J Mol Sci. 2019;20:2663. 10.3390/ijms20112663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Konishi T, Shiozaki A, Kosuga T, Kudou M, Shoda K, Arita T, et al. LRRC8A expression influences growth of esophageal squamous cell carcinoma. Am J Pathol. 2019;189:1973–85. 10.1016/j.ajpath.2019.06.006 [DOI] [PubMed] [Google Scholar]
- 39.Kurashima K, Shiozaki A, Kudou M, Shimizu H, Arita T, Kosuga T. et al. LRRC8A influences the growth of gastric cancer cells via the p53 signaling pathway. Gastric Cancer. 2021;24:1063–75. 10.1007/s10120-021-01187-4 [DOI] [PubMed] [Google Scholar]
- 40.Hauptman N, Glavač D. Long non-coding RNA in cancer. Int J Mol Sci. 2013;14:4655–69. 10.3390/ijms14034655 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Fatica A, Bozzoni I. Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet. 2014;15:7–21. 10.1038/nrg3606 [DOI] [PubMed] [Google Scholar]
- 42.Schober AL, Wilson CS, Mongin AA. Molecular composition and heterogeneity of the LRRC8-containing swelling-activated osmolyte channels in primary rat astrocytes. J Physiol. 2017;595:6939–51. 10.1113/JP275053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lin A, Wang RT, Ahn S, Park CC, Smith DJ. A genome-wide map of human genetic interactions inferred from radiation hybrid genotypes. Genome Res. 2010;20:1122–32. 10.1101/gr.104216.109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zeng LS, Yang XZ, Wen YF, Mail SJ, Wang MH, Zhang MY, et al. Overexpressed HDAC4 is associated with poor survival and promotes tumor progression in esophageal carcinoma. Aging. 2016;8:1236–49. 10.18632/aging.100980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Cai JY, Xu TT, Wang Y, Chang JJ, Li J, Chen XY, et al. Histone deacetylase HDAC4 promotes the proliferation and invasion of glioma cells. Int J Oncol. 2018;53:2758–68. [DOI] [PubMed] [Google Scholar]
- 46.Aster JC, Pear WS, Blacklow SC. The varied roles of notch in cancer. Annu Rev Pathol. 2017;12:245–75. 10.1146/annurev-pathol-052016-100127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Yuan X, Wu H, Xu H, Xiong H, Chu Q, Yu S, et al. Notch signaling: an emerging therapeutic target for cancer treatment. Cancer Lett. 2015;369:20–7. 10.1016/j.canlet.2015.07.048 [DOI] [PubMed] [Google Scholar]
- 48.Siemer S, Fauth T, Scholz P, Al-Zamel Y, Khamis A, Gül D. Profiling Cisplatin resistance in head and neck cancer: a critical role of the VRAC ion channel for chemoresistance. Cancers. 2021;13:4831 10.3390/cancers13194831 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Zhang H, Liu R, Jing Z, Li C, Fan W, Li H, et al. LRRC8A as a central mediator promotes colon cancer metastasis by regulating PIP5K1B/PIP2 pathway. Biochim Biophys Acta Mol Basis Dis. 2024;1870:167066. 10.1016/j.bbadis.2024.167066 [DOI] [PubMed] [Google Scholar]
- 50.Braunewell KH, Spilker C, Behnisch T, Gundelfinger ED. The neuronal calcium-sensor protein VILIP modulates cyclic AMP accumulation in stably transfected C6 glioma cells: amino-terminal myristoylation determines functional activity. J Neurochem. 1997;68:2129–39. 10.1046/j.1471-4159.1997.68052129.x [DOI] [PubMed] [Google Scholar]
- 51.Akagi T, Hijiya N, Inomata M, Shiraishi N, Moriyama M, Kitano S. Visinin-like protein-1 overexpression is an indicator of lymph node metastasis and poor prognosis in colorectal cancer patients. Int J Cancer. 2012;131:1307–17. 10.1002/ijc.27341 [DOI] [PubMed] [Google Scholar]
- 52.He C, Liu W, Xiong Y, Wang Y, Pan L, Luo L, et al. VSNL1 promotes cell proliferation, migration, and invasion in colorectal cancer by binding with COL10A1. Ann Clin Lab Sci. 2022;52:60–72. [PubMed] [Google Scholar]
- 53.Jiang Z, Yu T, Fan Z, Yang H, Lin X. Krüppel-like Factor 7 is a marker of aggressive gastric cancer and poor prognosis. Cell Physiol Biochem. 2017;43:1090–9. 10.1159/000481748 [DOI] [PubMed] [Google Scholar]
- 54.Niu R, Tang Y, Xi Y, Jiang D. High expression of Krüppel-like Factor 7 indicates unfavorable clinical outcomes in patients with lung Adenocarcinoma. J Surg Res. 2020;250:216–23. 10.1016/j.jss.2019.12.053 [DOI] [PubMed] [Google Scholar]
- 55.Huang Z, He H, Qiu F, Qian H. Expression and prognosis value of the KLF family members in colorectal cancer. J Oncol. 2022;2022:6571272. 10.1155/2022/6571272 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Green. JP, Swanton. T, Morris. LV, El-Sharkawy. LY, Cook. J, Yu. S, et al. LRRC8A is essential for hypotonicity-, but not for DAMP-induced NLRP3 inflammasome activation. Elife. 2020;9:e59704. [DOI] [PMC free article] [PubMed]
- 57.Gupta N, Wang H, McLeod TL, Naus CC, Kyurkchiev S, Advani S, et al. Inhibition of glioma cell growth and tumorigenic potential by CCN3 (NOV). Mol Pathol. 2001;54:293–9. 10.1136/mp.54.5.293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.McCallum L, Lu W, Price S, Lazar N, Perbal B, Irvine AE. CCN3 suppresses mitogenic signalling and reinstates growth control mechanisms in Chronic Myeloid Leukaemia. J Cell Commun Signal. 2012;6:27–35. 10.1007/s12079-011-0142-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Liu S, Liu Z, Bi D, Yuan X, Liu X, Ding S, et al. CCN3 (NOV) regulates proliferation, adhesion, migration and invasion in clear cell renal cell carcinoma. Oncol Lett. 2012;3:1099–104. 10.3892/ol.2012.607 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Dobson JR, Taipaleenmäki H, Hu YJ, Hong D, van Wijnen AJ, Stein JL, et al. hsa-mir-30c promotes the invasive phenotype of metastatic breast cancer cells by targeting NOV/CCN3. Cancer Cell Int. 2014;14:73. 10.1186/s12935-014-0073-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Li J, Ye L, Sun PH, Zheng F, Ruge F, Satherley LK, et al. Reduced NOV expression correlates with disease progression in colorectal cancer and is associated with survival, invasion and chemoresistance of cancer cells. Oncotarget. 2017;8:26231–44. 10.18632/oncotarget.15439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhang H, Cui S, Jing Z, Fu G, Liu R, Zhao W, et al. LRRC8A is responsible for exosome biogenesis and volume regulation in colon cancer cells. Biochem J. 2023;480:701–13. 10.1042/BCJ20220614 [DOI] [PubMed] [Google Scholar]
- 63.Song Y, Ma X, Zhang M, Wang M, Wang G, Ye Y, et al. Ezrin Mediates Invasion and Metastasis in Tumorigenesis: A Review. Front Cell Dev Biol. 2020;8:588801. 10.3389/fcell.2020.588801 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Fehon RG, McClatchey AI, Bretscher A. Organizing the cell cortex: the role of ERM proteins. Nat Rev Mol Cell Biol. 2010;11:276–87. 10.1038/nrm2866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Bretscher A, Edwards K, Fehon RG. ERM proteins and merlin: integrators at the cell cortex. Nat Rev Mol Cell Biol. 2002;3:586–99. 10.1038/nrm882 [DOI] [PubMed] [Google Scholar]
- 66.Mackay DJ, Esch F, Furthmayr H, Hall A. Rho- and rac-dependent assembly of focal adhesion complexes and actin filaments in permeabilized fibroblasts: an essential role for ezrin/radixin/moesin proteins. J Cell Biol. 1997;138:927–38. 10.1083/jcb.138.4.927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Yonemura S, Hirao M, Doi Y, Takahashi N, Kondo T, Tsukita S. Ezrin/radixin/moesin (ERM) proteins bind to a positively charged amino acid cluster in the juxta-membrane cytoplasmic domain of CD44, CD43, and ICAM-2. J Cell Biol. 1998;140:885–95. 10.1083/jcb.140.4.885 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Gautreau A, Poullet P, Louvard D, Arpin M. Ezrin, a plasma membrane-microfilament linker, signals cell survival through the phosphatidylinositol 3-kinase/Akt pathway. Proc Natl Acad Sci USA. 1999;96:7300–5. 10.1073/pnas.96.13.7300 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Heiska L, Alfthan K, Grönholm M, Vilja P, Vaheri A, Carpén O. Association of ezrin with intercellular adhesion molecule-1 and -2 (ICAM-1 and ICAM-2). Regulation by phosphatidylinositol 4, 5-bisphosphate. J Biol Chem. 1998;273:21893–900. 10.1074/jbc.273.34.21893 [DOI] [PubMed] [Google Scholar]
- 70.Garcia-Etxebarria K, Etxart A, Barrero M, Nafria B, Segues Merino NM, Romero-Garmendia I. et al. Genetic variants as predictors of the success of colorectal cancer treatments. Cancers. 2023;15:4688. 10.3390/cancers15194688 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Wang XD, Lu J, Lin YS, Gao C, Qi F. Functional role of long non-coding RNA CASC19/miR-140-5p/CEMIP axis in colorectal cancer progression. World J Gastroenterol. 2019;25:1697–714. 10.3748/wjg.v25.i14.1697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Li XL, Subramanian M, Jones MF, Chaudhary R, Singh DK, Zong X, et al. Long noncoding RNA PURPL suppresses Basal p53 levels and promotes tumorigenicity in colorectal cancer. Cell Rep. 2017;20:2408–23. 10.1016/j.celrep.2017.08.041 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Li N, Li J, Mi Q, Xie Y, Li P, Wang L, et al. Long non-coding RNA ADAMTS9-AS1 suppresses colorectal cancer by inhibiting the Wnt/β-catenin signalling pathway and is a potential diagnostic biomarker. J Cell Mol Med. 2020;24:11318–29. 10.1111/jcmm.15713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wang Y, Fu J, Yang L, Liang Z. Long non-coding RNA SNHG20 promotes colorectal cancer cell proliferation, migration and invasion via miR-495/STAT3 axis. Mol Med Rep. 2021;23:31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Xiang X, Langlois S, St-Pierre ME, Blinder A, Charron P, Graber TE, et al. Identification of pannexin 1-regulated genes, interactome, and pathways in rhabdomyosarcoma and its tumor inhibitory interaction with AHNAK. Oncogene. 2021;40:1868–83. 10.1038/s41388-020-01623-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Fourie C, Li D, Montgomery JM. The anchoring protein SAP97 influences the trafficking and localisation of multiple membrane channels. Biochim Biophys Acta. 2014;1838:589–94. 10.1016/j.bbamem.2013.03.015 [DOI] [PubMed] [Google Scholar]
- 77.Marziali F, Dizanzo MP, Cavatorta AL, Gardiol D. Differential expression of DLG1 as a common trait in different human diseases: an encouraging issue in molecular pathology. Biol Chem. 2019;400:699–710. 10.1515/hsz-2018-0350 [DOI] [PubMed] [Google Scholar]
- 78.Walensky LD, Gascard P, Fields ME, Blackshaw S, Conboy JG, Mohandas N, et al. The 13-kD FK506 binding protein, FKBP13, interacts with a novel homologue of the erythrocyte membrane cytoskeletal protein 4.1. J Cell Biol. 1998;141:143–53. 10.1083/jcb.141.1.143 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Parra M, Gascard P, Walensky LD, Snyder SH, Mohandas N, Conboy JG. Cloning and characterization of 4.1G (EPB41L2), a new member of the skeletal protein 4.1 (EPB41) gene family. Genomics. 1998;49:298–306. 10.1006/geno.1998.5265 [DOI] [PubMed] [Google Scholar]
- 80.Baines AJ, Bennett PM, Carter EW, Terracciano C. Protein 4.1 and the control of ion channels. Blood Cells Mol Dis. 2009;42:211–5. 10.1016/j.bcmd.2009.01.016 [DOI] [PubMed] [Google Scholar]
- 81.Cioffi DL, Wu S, Alexeyev M, Goodman SR, Zhu MX, Stevens T. Activation of the endothelial store-operated ISOC Ca2+ channel requires interaction of protein 4.1 with TRPC4. Circ Res. 2005;97:1164–72. 10.1161/01.RES.0000193597.65217.00 [DOI] [PubMed] [Google Scholar]
- 82.Shipston MJ. Ion channel regulation by protein palmitoylation. J Biol Chem. 2011;286:8709–16. 10.1074/jbc.R110.210005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Howie J, Reilly L, Fraser NJ, Vlachaki Walker JM, Wypijewski KJ, Ashford ML, et al. Substrate recognition by the cell surface palmitoyl transferase DHHC5. Proc Natl Acad Sci USA. 2014;111:17534–9. 10.1073/pnas.1413627111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Yang HQ, Martinez-Ortiz W, Hwang J, Fan X, Cardozo TJ, Coetzee WA. Palmitoylation of the K. Proc Natl Acad Sci USA. 2020;117:10593–602. 10.1073/pnas.1918088117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Gök C, Main A, Gao X, Kerekes Z, Plain F, Kuo CW, et al. Insights into the molecular basis of the palmitoylation and depalmitoylation of NCX1. Cell Calcium. 2021;97:102408. 10.1016/j.ceca.2021.102408 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Gao X, Kuo CW, Main A, Brown E, Rios FJ, Camargo LL, et al. Palmitoylation regulates cellular distribution of and transmembrane Ca flux through TrpM7. Cell Calcium. 2022;106:102639. 10.1016/j.ceca.2022.102639 [DOI] [PubMed] [Google Scholar]
- 87.Meyer K, Korbmacher C. Cell swelling activates ATP-dependent voltage-gated chloride channels in M-1 mouse cortical collecting duct cells. J Gen Physiol. 1996;108:177–93. 10.1085/jgp.108.3.177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Chou CY, Shen MR, Hsu KS, Huang HY, Lin HC. Involvement of PKC-alpha in regulatory volume decrease responses and activation of volume-sensitive chloride channels in human cervical cancer HT-3 cells. J Physiol. 1998;512:435–48. 10.1111/j.1469-7793.1998.435be.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Hermoso M, Olivero P, Torres R, Riveros A, Quest AF, Stutzin A. Cell volume regulation in response to hypotonicity is impaired in HeLa cells expressing a protein kinase Calpha mutant lacking kinase activity. J Biol Chem. 2004;279:17681–9. 10.1074/jbc.M304506200 [DOI] [PubMed] [Google Scholar]
- 90.Gong W, Xu H, Shimizu T, Morishima S, Tanabe S, Tachibe T, et al. ClC-3-independent, PKC-dependent activity of volume-sensitive Cl channel in mouse ventricular cardiomyocytes. Cell Physiol Biochem. 2004;14:213–24. 10.1159/000080330 [DOI] [PubMed] [Google Scholar]
- 91.Bertelli S, Remigante A, Zuccolini P, Barbieri R, Ferrera L, Picco C, et al. Mechanisms of activation of LRRC8 volume regulated anion channels. Cell Physiol Biochem. 2021;55:41–56. 10.33594/000000329 [DOI] [PubMed] [Google Scholar]
- 92.Bertelli S, Zuccolini P, Gavazzo P, Pusch M. Molecular determinants underlying volume-regulated anion channel subunit-dependent oxidation sensitivity. J Physiol. 2022;600:3965–82. 10.1113/JP283321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.König, B, Hao, Y, Schwartz, S, Plested, AJ, Stauber, TA. FRET sensor of C-terminal movement reveals VRAC activation by plasma membrane DAG signaling rather than ionic strength. Elife. 2019;8. [DOI] [PMC free article] [PubMed]
- 94.Doroshenko P. Second messengers mediating activation of chloride current by intracellular GTP gamma S in bovine chromaffin cells. J Physiol. 1991;436:725–38. 10.1113/jphysiol.1991.sp018576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Nilius B, Eggermont J, Voets T, Buyse G, Manolopoulos V, Droogmans G. Properties of volume-regulated anion channels in mammalian cells. Prog Biophys Mol Biol. 1997;68:69–119. 10.1016/S0079-6107(97)00021-7 [DOI] [PubMed] [Google Scholar]
- 96.Jaffe AB, Hall A. Rho GTPases: biochemistry and biology. Annu Rev Cell Dev Biol. 2005;21:247–69. 10.1146/annurev.cellbio.21.020604.150721 [DOI] [PubMed] [Google Scholar]
- 97.Vega FM, Ridley AJ. Rho GTPases in cancer cell biology. FEBS Lett. 2008;582:2093–101. 10.1016/j.febslet.2008.04.039 [DOI] [PubMed] [Google Scholar]
- 98.Estevez AY, Bond T, Strange K. Regulation of I(Cl,swell) in neuroblastoma cells by G protein signaling pathways. Am J Physiol Cell Physiol. 2001;281:C89–98. 10.1152/ajpcell.2001.281.1.C89 [DOI] [PubMed] [Google Scholar]
- 99.Botchkin LM, Matthews G. Chloride current activated by swelling in retinal pigment epithelium cells. Am J Physiol. 1993;265:C1037–45. 10.1152/ajpcell.1993.265.4.C1037 [DOI] [PubMed] [Google Scholar]
- 100.Lemonnier L, Prevarskaya N, Shuba Y, Vanden Abeele F, Nilius B, Mazurier J, et al. Ca2+ modulation of volume-regulated anion channels: evidence for colocalization with store-operated channels. FASEB J. 2002;16:222–4. 10.1096/fj.01-0383fje [DOI] [PubMed] [Google Scholar]
- 101.Akita T, Okada Y. Regulation of bradykinin-induced activation of volume-sensitive outwardly rectifying anion channels by Ca2+ nanodomains in mouse astrocytes. J Physiol. 2011;589:3909–27. 10.1113/jphysiol.2011.208173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Akita T, Fedorovich SV, Okada Y. Ca2+ nanodomain-mediated component of swelling-induced volume-sensitive outwardly rectifying anion current triggered by autocrine action of ATP in mouse astrocytes. Cell Physiol Biochem. 2011;28:1181–90. 10.1159/000335867 [DOI] [PubMed] [Google Scholar]
- 103.Liu Y, Zhang H, Men H, Du Y, Xiao Z, Zhang F, et al. Volume-regulated Cl. Am J Physiol Cell Physiol. 2019;317:C466–C80. 10.1152/ajpcell.00507.2018 [DOI] [PubMed] [Google Scholar]
- 104.Putney JW. Origins of the concept of store-operated calcium entry. Front Biosci (Sch Ed). 2011;3:980–4. 10.2741/202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Sallinger M, Grabmayr H, Humer C, Bonhenry D, Romanin C, Schindl R. et al. Activation mechanisms and structural dynamics of STIM proteins. J Physiol. 2023;602:1475–507. 10.1113/JP283828 [DOI] [PubMed] [Google Scholar]
- 106.Long Y, Xia JY, Chen SW, Gao CL, Liang GN, He XM, et al. ATP2B1 gene silencing increases insulin sensitivity through facilitating Akt activation via the Ca. Int J Biol Sci. 2017;13:1203–12. 10.7150/ijbs.19666 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Giordano F, Saheki Y, Idevall-Hagren O, Colombo SF, Pirruccello M, Milosevic I, et al. PI(4,5)P(2)-dependent and Ca(2+)-regulated ER-PM interactions mediated by the extended synaptotagmins. Cell. 2013;153:1494–509. 10.1016/j.cell.2013.05.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Lehnart SE, Wehrens XHT. The role of junctophilin proteins in cellular function. Physiol Rev. 2022;102:1211–61. 10.1152/physrev.00024.2021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Sreeja JS, John R, Dharmapal D, Nellikka RK, Sengupta S. A fresh look at the structure, regulation, and functions of Fodrin. Mol Cell Biol. 2020;40:e00133–20. 10.1128/MCB.00133-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinforma. 2013;29:15–21. 10.1093/bioinformatics/bts635 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinforma. 2011;12:323. 10.1186/1471-2105-12-323 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinforma. 2010;26:139–40. 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Jurman ME, Boland LM, Liu Y, Yellen G. Visual identification of individual transfected cells for electrophysiology using antibody-coated beads. Biotechniques. 1994;17:876–81. [PubMed] [Google Scholar]
- 114.Le Sage V, Cinti A, Mouland AJ. Proximity-dependent biotinylation for identification of interacting proteins. Curr Protoc Cell Biol. 2016;73:17.9.1–9.2. [DOI] [PubMed] [Google Scholar]
- 115.Roux KJ, Kim DI, Burke B. BioID: a screen for protein-protein interactions. Curr Protoc Protein Sci. 2013;74:19.23.1–19.23.14. 10.1002/0471140864.ps1923s74 [DOI] [PubMed] [Google Scholar]
- 116.Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic acids Res. 2015;43:e47. 10.1093/nar/gkv007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Huber W, von Heydebreck A, Sültmann H, Poustka A, Vingron M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinforma. 2002;18:S96–104. 10.1093/bioinformatics/18.suppl_1.S96 [DOI] [PubMed] [Google Scholar]
- 118.Gatto L, Lilley KS. MSnbase-an R/Bioconductor package for isobaric tagged mass spectrometry data visualization, processing and quantitation. Bioinforma. 2012;28:288–9. 10.1093/bioinformatics/btr645 [DOI] [PubMed] [Google Scholar]
- 119.Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30. 10.1093/nar/28.1.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Roux KJ, Kim DI, Burke B, May DG. BioID: A screen for protein-protein interactions. Curr Protoc Protein Sci. 2018;91:19.23.1–19.23.15. 10.1002/cpps.51 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Kim DI, Birendra KC, Zhu W, Motamedchaboki K, Doye V, Roux KJ. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc Natl Acad Sci USA. 2014;111:E2453–61. 10.1073/pnas.1406459111 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets used in the current study are available from the corresponding authors on reasonable request.