Summary
Background
In order to prevent the emergence and spread of future variants of concern of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), developing vaccines capable of stopping transmission is crucial. The SARS-CoV-2 vaccine NDV-HXP-S can be administered live intranasally (IN) and thus induce protective immunity in the upper respiratory tract. The vaccine is based on Newcastle disease virus (NDV) expressing a stabilised SARS-CoV-2 spike protein. NDV-HXP-S can be produced as influenza virus vaccine at low cost in embryonated chicken eggs.
Methods
The NDV-HXP-S vaccine was genetically engineered to match the Omicron variants of concern (VOC) BA.1 and BA.5 and tested as an IN two or three dose vaccination regimen in female mice. Furthermore, female mice intramuscularly (IM) vaccinated with mRNA-lipid nanoparticles (LNPs) were IN boosted with NDV-HXP-S. Systemic humoral immunity, memory T cell responses in the lungs and spleens as well as immunoglobulin A (IgA) responses in distinct mucosal tissues were characterised.
Findings
NDV-HXP-S Omicron variant vaccines elicited high mucosal IgA and serum IgG titers against respective SARS-CoV-2 VOC in female mice following IN administration and protected against challenge from matched variants. Additionally, antigen-specific memory B cells and local T cell responses in the lungs were induced. Host immunity against the NDV vector did not interfere with boosting. Intramuscular vaccination with mRNA-LNPs was enhanced by IN NDV-HXP-S boosting resulting in improvement of serum neutralization titers and induction of mucosal immunity.
Interpretation
We demonstrate that NDV-HXP-S Omicron variant vaccines utilised for primary immunizations or boosting efficiently elicit humoral and cellular immunity. The described induction of systemic and mucosal immunity has the potential to reduce infection and transmission.
Funding
This work was partially funded by the NIAIDCenters of Excellence for Influenza Research and Response (CEIRR) and by the NIAID Collaborative Vaccine Innovation Centers and by institutional funding from the Icahn School of Medicine at Mount Sinai. See under Acknowledgements for details.
Keywords: COVID-19, Mucosal immune response, Low cost vaccine platform, Variant vaccine, NDV vector, mRNA vaccine, Prime-pull vaccination
Research in context.
Evidence before this study
In an unprecedented effort, many vaccines for SARS-CoV-2 have been developed with remarkable speed as a response to the coronavirus disease 2019 (COVID-19) pandemic. The viral vector vaccine NDV-HXP-S was designed as a low-cost vaccine allowing self-sufficient production of SARS-CoV-2 vaccines in low- and middle-income countries. This is possible because the NDV-HXP-S vaccine can be grown in embryonated chicken eggs, which allows manufacturing in existing influenza vaccine production facilities. The vaccine is based on the avirulent LaSota strain of Newcastle disease virus (NDV) that has been evaluated as a vaccine vector for many animal pathogens as well as an oncolytic agent in humans. Since NDV is known to tolerate large insertions into its genome, a modified version of the spike protein was inserted and is expressed on the virion’s surface. It has been shown that NDV-HXP-S can be used as an inactivated intramuscular (IM) or live intranasal (IN) vaccine, conferring protection in mice and hamsters. Furthermore, in pre-clinical studies in mice, next-generation NDV-HXP-S variant vaccines (Beta, Gamma and Delta) used in multivalent formulations protected against phylogenetically distant SARS-CoV-2 VOC. Good safety and immunogenicity was displayed in humans in clinical phase I and II trials.
Added value of this study
This study demonstrates that NDV-HXP-S administered IN efficiently induces mucosal humoral and cellular immunity. The NDV-HXP-S vaccine construct can easily be genetically engineered to match variants of concern (VOC). The BA.1 and BA.5 variant vaccines induce high binding and neutralizing serum IgG titers as well as IgA responses in several different mucosal tissues against the respective Omicron variants. Furthermore, the NDV-HXP-S variant vaccines protect against SARS-CoV-2 BA.1 virus challenge in mice. As a proof of concept, IN boosting with NDV-HXP-S after IM mRNA-lipid nanoparticles (LNPs) vaccination elicits good mucosal immunity and improves systemic immunity. Finally, the impact of vector immunity was addressed and no impediment of immune responses against the SARS-CoV-2 spike was found.
Implications of all the available evidence
This data supports the usage of an intranasal booster after mRNA vaccination in order to induce mucosal immunity.
Introduction
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has caused a pandemic with tremendous human suffering worldwide. Many of the newly emerging variants of concern (VOC) have shown an increase in transmissibility and immune evasion.1 This made it difficult for the first generation vaccines based on the ancestral strain to protect against asymptomatic to mild infection and transmission.2, 3, 4 Matched strain boosters might be able to restore a higher level of protection.5, 6, 7
While intramuscular (IM) administered SARS-CoV-2 vaccines induce high levels of serum antibodies as well as memory B cells and circulating effector CD4+ and CD8+ T cells, IM vaccines appear to fail at efficiently inducing mucosal immunity.8, 9, 10, 11, 12 Intranasal (IN) vaccinations could fill that gap.13 Like natural SARS-CoV-2 infection, IN vaccination has been shown to induce potent antiviral immune memory at sites of administration in animal models.14, 15, 16, 17, 18, 19, 20, 21, 22, 23 Several virus based vaccine platforms like adenovirus, measles, influenza, parainfluenza virus 5, and Newcastle disease virus have had success when administered IN in preclinical models.14, 15, 16, 17, 18, 19, 20, 21 Furthermore, boosting IN with unadjuvanted recombinant spike protein after IM priming elicits mucosal immune memory within the respiratory tract.22 These increases in mucosal IgA and tissue-resident memory cells lead to significantly reduced viral loads in the upper and lower airways. The induced mucosal immunity can prevent disease and potentially block the transmission of the virus. However, developing and validating a protective and safe intranasal vaccine (against any pathogen) has proven difficult with only limited success in the past. The only intranasal vaccine approved by the FDA is FluMist.23,24 Additionally, an inhaled aerosolised Ad5-nCoV vaccine (CanSino), a live-attenuated influenza virus vector-based vaccine delivered via intranasal spray (dNS1-RBD), and an intranasal ChAd vaccine (iNCOVACC) were recently licensed in China (CanSino, dNS1-RBD) and India (iNCOVACC).25, 26, 27, 28, 29
We have developed a vaccine candidate (NDV-HXP-S) based on the avirulent LaSota strain of Newcastle disease virus (NDV) that expresses a modified version of the SARS-CoV-2 spike protein on its surface.30, 31, 32 NDV has been shown to be a very immunogenic vaccine when administered intranasally in mice and hamster models.21,30,32 Compared to the work of others, like Warner and colleagues, we have modified the spike with stabilizing and membrane anchoring adaptations.21 Most importantly, their work focusses on the ancestral spike.21 Conversely, we created next-generation variant vaccines (Beta, Gamma and Delta) and showed that a trivalent IM formulation protects against phylogenetically distant SARS-CoV-2 VOC in mice.33 The NDV-HXP-S vaccine can be produced at low cost in embryonated chicken eggs. This allows manufacturing of the NDV-HXP-S vaccine in existing influenza vaccine production facilities located around the globe, enabling generation of self-sufficient supplies of coronavirus disease 2019 (COVID-19) vaccine in many low- and middle-income countries.32 Additionally, results from clinical phase I/II trials have demonstrated a good safety and immunogenicity profile (Thailand, inactivated IM, phase I and II, NCT04764422; Vietnam, inactivated IM, phase I and II, NCT04830800), with additional trials in the United States (live IN, phase I; NCT05181709), Mexico (live IN/IM, phase II/III; NCT05205746), and Brazil (inactivated IM, phase II/III; NCT05354024) on the way.34, 35, 36, 37
We believe that a mucosal (intranasal) vaccine platform easily adjustable to new variants will aid in the struggle against the rapidly evolving SARS-CoV-2. In the present study, mice were vaccinated IN with NDV-HXP-S BA.1 and BA.5 variant vaccines and the cellular and humoral immune responses induced at different mucosal sites were assessed. In addition, protection against SARS-CoV-2 BA.1 infection was shown. Furthermore, we determined the mucosal and systemic immunity after an IM mRNA lipid nanoparticles (LNP) vaccination followed by IN boosting with NDV-HXP-S in mice.
Methods
Cells
Baby hamster kidney cells stably expressing human angiotensin converting enzyme 2 (BHK-ACE2) and BHK-derived cell line stably expressing T7 RNA polymerase (BSRT7) were used in this study.38,39 Douglas Foster-1 (DF-1) cells were purchased from American Type Culture Collection (ATCC, CRL-12203, RRID: CVCL_0570). Cercopithecus aethiops kidney epithelial cells expressing transmembrane protease, serine 2 and human angiotensin-converting enzyme 2 (Vero E6-TMPRSS2-T2A-ACE2) were purchased from ATCC (NR-54970, RRID:CVCL_C7NK). BHK-ACE2, BSRT7 and DF-1 cells were maintained in Dulbecco’s Modified Eagle’s Medium (DMEM; Gibco, REF 11995073) containing 10% (vol/vol) heat inactivated fetal bovine serum (FBS), 100 units/mL of penicillin and 100 ug/mL of streptomycin (P/S; Gibco, REF 15140122) with BSRT7 and BHK-ACE2 cells kept at 37 °C and DF-1 cells at 39 °C with 5% CO2. Vero E6-TMPRSS2-T2A-ACE2 were maintained in DMEM containing 10% (vol/vol) heat inactivated FBS, 10 mM N-2-hydroxyethylpiperazine-N-2-ethane sulfonic acid buffering agent (HEPES; Gibco, REF 15630080) and 10 mg/mL puromycin (Gibco, REF A1113803) at 37 °C with 5% CO2. Cell lines were confirmed to be mycoplasma free upon arrival and tested at monthly intervals.
Plasmids
Stabilizing Hexa-Pro (HXP) mutations (F817P, A892P, A899P, A942P, K986P, V987P) were introduced into the S–F chimera by PCR and the S–F sequence was subsequently inserted into the pNDV_LS/L289A rescue plasmid between the P and M genes using the In-Fusion HD Cloning Kit by Takara Bio (Cat#639650) as described.30, 31, 32,40 The HXP spike was adapted to new variants by introducing the appropriate mutations into the HXP-S sequence in silico and obtained as gBlock gene fragments from Integrated DNA Technologies.33 The recombinant products were transformed into MAX Efficiency Stbl2 chemically competent cells (Thermo Fisher Scientific, REF 10268-019) to generate the pNDV-HXP-S rescue plasmid. The plasmids were purified using NucleoBond Xtra Maxi Plus kit (Macherey–Nagel, REF 740416.50).
Rescue of NDV-HXP-S
As described in our previous studies,31 BSRT7 cells stably expressing the T7 polymerase were seeded onto 6-well plates at 5 × 105 cells per well in duplicate. The next day, cells were transfected with 2 μg of pNDV-HXP-S, 1 μg of pTM1-NP, 0.5 μg of pTM1-P, 0.5 μg of pTM1-L, and 1 μg of pCI-T7opt and were re-suspended in 250 μL of Opti-Minimum Essential Medium (Opti-MEM, Gibco, REF 31985-070). The plasmid cocktail was then gently mixed with 15 μL of TransIT LT1 transfection reagent (Mirus, Prod. No. MIR 2305). To increase rescue efficiency, BSRT7-DF-1 co-culture was established the next day as described previously.41 Specifically, transfected BSRT7 and DF-1 cells were washed with warm phosphate-buffered saline (PBS, pH 7.4, Gibco, REF 10010-023) and trypsinised (using 3 mL of 0.05% Trypsin EDTA 1x, Gibco, REF 25300). Trypsinised cells were neutralised with 6 mL of growth media. BSRT7 cells were mixed with DF-1 cells (around 1:2.5) in a 10-cm dish. The co-culture was incubated at 37 °C overnight. The next day, the medium was removed, and cells were gently washed with warm PBS twice, Opti-MEM supplemented with 1% P/S, and 0.1 μg/mL of tosyl phenylalanyl chloromethyl ketone (TPCK) - treated trypsin was added. The co-cultures were incubated for 2 or 3 days before inoculation into 8- or 9-day-old specific pathogen-free (SPF) embryonated chicken eggs (Charles River Laboratories). To inoculate eggs, cells and supernatants were harvested and homogenised by several syringe strokes. Two hundred μL of the mixture were injected into each egg. Eggs were incubated at 37 °C for 3 days and cooled at 4 °C overnight. Allantoic fluid was harvested from cooled eggs, and the rescue of the viruses was determined by hemagglutination (HA) assays. The genetic stability of the recombinant viruses was evaluated across multiple passages in 10-day-old SPF embryonated chicken eggs.33 Furthermore, RNA of the rescued viruses were extracted, RT-PCR was performed to amplify the cDNA segments of the viral genome and then sequenced by Sanger sequencing (Psomagen).
Preparation of concentrated virus
Allantoic fluid was clarified by centrifugation to remove cell debris at 3441×g at 4 °C for 30 min using a Sorvall Legend RT Plus Refrigerated Benchtop Centrifuge (Thermo Fisher Scientific) as described previously.31 The allantoic fluid was carefully laid on top of a 20% sucrose cushion in NTE buffer (100 mM NaCl, 10 mM Tris–HCl, 1 mM ethylenediamine tetraacetic acid (EDTA), pH 7.4). The live virus was pelleted through a sucrose cushion, removing soluble egg proteins, via ultra-centrifugation in a Beckman L7-65 ultracentrifuge at 112,499×g for 2 h at 4 °C using a Beckman SW28 rotor (Beckman Coulter). After aspirating off the supernatants, the virus pellets were re-suspended in PBS. The total protein content was determined using the bicinchoninic acid (BCA) assay (Thermo Fisher Scientific, REF 23225).
Virus titration by EID50
As described in our previous studies, the fifty percent of egg embryo infectious dose (EID50) assay was performed in 9- to 11-day old chicken embryonated eggs.33 The virus in allantoic fluid was 10-fold serially diluted in PBS, resulting in 10−5 to 10−10 dilutions of the virus. Of each dilution, 100 μl were injected into each egg for a total of 5–10 eggs per dilution. The eggs were incubated at 37 °C for 3–4 days and then cooled at 4 °C overnight. The allantoic fluid was subsequently collected and analysed by HA assay. The EID50 titer of the NDV, determined by the number of HA-positive and HA-negative eggs in each dilution, was calculated using the Reed and Muench method.
Sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE)
The concentrated NDV-HXP-S was mixed with 2x Laemmli sample buffer (BIO-RAD, Cat. #1610737), 2-mercaptoethanol (1:40, Sigma–Aldrich, M3148) and PBS at appropriate amounts to reach a total protein content of 15 μg in 50 μl per gel slot. The mixture was heated at 90 °C for 5 min. The samples were mixed by pipetting and loaded to a 4–20% 10-well Mini-Protean TGX precast gel (Bio-RAD, Cat. #4561094). Ten μl of the Novex Sharp Pre-Stained Protein standard (Thermo Fisher Scientific, REF 57318) was used as the ladder. The electrophoresis was run in Tris/glycine SDS/Buffer (25 mM Tris, 192 mM glycine, 0.1% SDS, pH8.3). For Coomassie blue staining, the gel was washed with distilled water at room temperature (RT) several times until the dye front in the gel was no longer visible. The gel was stained with 20 mL of SimplyBlue SafeStain (Thermo Fisher Scientific, LC6065) overnight. The SimplyBlue SafeStain was decanted and the gel was washed with distilled water several times until the background was clear. Gels were imaged using the Bio-Rad Universal Hood II Molecular imager (Bio-Rad) and processed by Image Lab Software (Bio-Rad).
Ethics
All the animal experiments were performed in accordance with protocols (PROTO202000098) approved by the Icahn School of Medicine at Mount Sinai Institutional Animal Care and Use Committee (IACUC).
Animal experiments
All the animals were housed in a temperature/humidity-controlled facility with 12 h light/dark cycle. All experiments with live SARS-CoV-2 were performed in the Centers for Disease Control and Prevention/US Department of Agriculture-approved biosafety level 3 (BSL-3) biocontainment facility of the Global Health and Emerging Pathogens Institute at the Icahn School of Medicine at Mount Sinai, in accordance with institutional biosafety requirements. Healthy female mice that are 6–8 week-old were used for this study due to their behavioural stability as compared to male mice. Male mice tend to be difficult to manage (especially in the BSL-3 facility) because of their aggressive behaviour, often resulting in inhumane consequences to their cage mates and may compromise statistical power of the study due to loss of animals.
Mouse immunization
Six-to eight-week-old female BALB/c mice (RRID: IMSR_JAX:000651) were purchased from The Jackson Laboratory (JAX). For IN vaccination, mice were anesthetised with a ketamine/xylazine cocktail before vaccine administration of 10 μl of total volume split between each nostril of the anesthetised mouse. The live concentrated NDV-HXP-S and its variant adapted versions were used at EID50 of 105, 106 or 107 in PBS. The animals were boosted with the same preparation 21 or 28 days after receiving the initial dose and for some experiments received a third IN vaccination approximately 6 months later. Live purified WT NDV and PBS were utilised as controls.
Six-to eight-week-old female B6.cg-Tg (K18-ACE2)2Prlmn/J mice (RRID: IMSR_JAX:034860) were purchased from The Jackson Laboratory (JAX) and were vaccinated with Pfizer BNT162b2 mRNA vaccine at 0.25 μg or 5 μg intramuscularly twice with 3 weeks apart. This vaccine was recovered from vaccination centers at the Mount Sinai hospital and had undergone one freeze thaw. Quality control tests were performed to guarantee vaccine immunogenicity and effectiveness in mice before the start of this study. Approximately 12 months after the second mRNA vaccine dose, animals were boosted with NDV-based vaccine via the intranasal route.
Viral challenge and passive serum transfer
For SARS-CoV-2 challenge studies, female B6.cg-Tg (K18-ACE2)2Prlmn/J mice (RRID: IMSR_JAX:034860) were intranasally infected with 3 × 104 plaque-forming units (PFU) of SARS-CoV-2 BA.1 virus in 30 μl of total volume. For the passive transfer, serum was pooled and 150 μl was administered per mouse via the intraperitoneal route 2 h before the challenge. Lung and nasal turbinates were harvested five days post challenge and homogenised in 1 mL of sterile PBS. Viral titers in the tissue homogenates were measured by plaque assay on VERO E6 TMPRSS2 T2A ACE2 cells.
Enzyme-linked immunosorbent assay (ELISA)
ELISAs were performed as described previously to measure spike-, receptor binding domain (RBD)- or whole inactivated NDV-specific IgG and IgA in the serum, nasal washes (NW), bronchoalveolar lavage fluid (BALF), vaginal lavages (VL), and intestinal lavages (IL) of mice vaccinated with NDV-HXP-S.30,31 Briefly, 96-well polystyrene microtiter plates (Immulon 4HBX #3855; Thermo Fisher Scientific) were coated with 50 μL/well of recombinant spike or RBD (2 μg/mL) or whole inactivated ND virus preparations (5 μg/mL) in 1x coating buffer diluted from 10 x KPL coating solution (SeraCare Life Sciences Inc., Cat. No.: 5150) overnight at 4 °C. The plates were then washed three times with 220 μL PBS containing 0.1% (v/v) Tween-20 (PBST). The plates were subsequently blocked with 220 μL blocking solution (3% goat serum, 0.5% dry milk, 96.5% PBST) per well for 1 h at RT. After discarding the blocking buffer, mouse sera were threefold serially diluted in blocking solution starting at 1:30 dilution while clarified NW, BALF, VL, and IL were twofold serially diluted starting at 1:2 or undiluted. The plates were subsequently incubated at RT for 2 h. Afterwards, the plates were washed three times with 220 μl PBST and 50 μL of Amersham ECL sheep anti-mouse IgG Horseradish Peroxidase (HRP)-linked whole secondary antibody (Cytiva Cat# NA931, RRID: AB_772210) at 1:3000 or HRP-conjugated goat anti-mouse IgA secondary antibody (Bethyl Cat# A90-103 P, RRID: AB_67140) at 1:2000 was added. The plates were incubated for 1 h at RT and washed 3 times with 220 μL PBST. One hundred μL of o-phenylenediamine dihydrochloride (SigmaFast OPD, Sigma, P9187) substrate was added per well. The plates were developed for 10 min and the reaction was stopped by adding 50 μL of 3 M hydrochloric acid (HCl, Thermo Fisher Scientific, S25856) to each well. The optical density (OD) was measured at 492 nm on a Synergy 4 plate reader (BioTek) or similar instrument. An average of OD values for blank wells plus three standard deviations was used to set a cutoff value for each plate. The endpoint titers were calculated and graphed using GraphPad Prism 9.5.1 (GraphPad Software).
rcVSV-eGFP-CoV2-S neutralization assay
Replication-competent vesicular stomatitis virus carrying an enhanced green fluorescent protein reporter and expressing ancestral, BA.1, BA.5, BQ.1.1, or XBB.1.5 SARS-CoV-2 spike (rcVSV-eGFP-CoV2-S) in place of VSV-G were produced as described previously.42 The virus stocks were titrated on BHK-ACE2 cells. One day prior to use in the neutralization assay, 2 × 104 BHK-ACE2 cells were seeded in flat bottom 96 well plates (Corning Falcon, REF 353072). Pooled serum samples were heat inactivated at 56 °C for 30 min before usage. Virus stocks were pre-mixed with 4-fold serially diluted serum, starting at 1:5, in DMEM-10%FBS-1%P/S and incubated for 15 min at RT. The virus plus serum mix was subsequently transferred onto the BHK-ACE2 target cells. At 12 h post-infection, GFP counts were measured using the Celigo imaging cytometer (Nexcelom Biosciences, version 4.1.3.0). Each assay was done in duplicates. For calculation of the inhibitory dilution at which 50% neutralization is achieved (ID50), GFP counts from “no serum” conditions were set to 100%; GFP counts of each condition (serum treated) were normalised to no serum control wells. Inhibition curves were generated using GraphPad Prism 9.5.1 (GraphPad Software) with “log (inhibitor) vs normalised response-variable slope” settings.
SARS-CoV-2 plaque assay
The plaque assay was performed in the BSL-3 facility of the Icahn School of Medicine at Mount Sinai. The day before the assay, 3 × 105 VERO E6 TMPRSS2 T2A ACE2 cells were seeded in 12-well plates (Thermo Fisher Scientific, REF 130185). Lung and nasal turbinate homogenates were 10-fold serially diluted in infection medium (DMEM + 2% FBS + 1% P/S + 10 mM HEPES). After removing the media from the cells, 200 μL of each tissue homogenate dilution were inoculated onto each well. The dilutions used range from 10−1 to 10−6. The plates were incubated at 37 °C for 1 h with rocking every 15 min. Subsequently, the inoculum was removed and 1 mL of agar overlay consisting of 0.7% agar in 2x MEM +2% FBS was placed onto each well. Once the agar was solidified, the plates were incubated at 37 °C with 5% CO2. Two days later, the plates were fixed with 4% formaldehyde in PBS overnight before being taken out of the BSL-3 facility for subsequent staining in BSL-2 facility. The cells were permeabilised with 0.1% Triton X-100 in PBS for 15 min at room temperature. The plaques were immuno-stained with an anti-SARS-CoV-2 NP primary mouse monoclonal antibody 1C7C7 kindly provided by Dr. James Duty at ISMMS. Amersham ECL sheep anti-mouse IgG Horseradish Peroxidase (HRP)-linked whole secondary antibody (Cytiva Cat# NA931, RRID: AB_772210) was used at 1:2000 and the plaques were visualised using TrueBlue Peroxidase Substrate (SeraCare Life Sciences Inc., REF 5510-0030).
Lung and spleen processing
For IV (intravascular) labelling, mice were anesthetised with a ketamine/xylazine cocktail. Non-responsive, immobile animals were injected with 3 μg of BV421 Rat Anti-mouse CD45 antibody (BD Horizon, Cat: 563890, clone 30-F11, RRID: AB_2651151) in 50 μl sterile PBS into the retro-orbital venous sinus.43 After 5 min of labelling, tissues were harvested and digested immediately, protected from light.
For perfusion, a small incision in the left heart ventricle was made. Using a small syringe, PBS was injected into the right ventricle of the heart and colour change of lungs and heart tissue was observed.
Lungs were dispersed into single-cell suspension by mechanical disruption via a gentleMACS Octo Dissociator with C-tubes (Miltenyi Biotec, Order no.:130096334) as well as enzymatic digestion. The digestion mix consists of liberase (Roche, REF 05401127001) and DNaseI (Roche, REF 10104159001) in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco, REF 11875) and the lungs were incubated for 30 min at 37 °C in the C-tubes. The cell suspension was run through 70 μm cell strainer (Fisher Scientific, Cat. No.: 22-363-548) into cold Hanks’ Balanced salt solution (HBSS, Gibco, REF 14025092) 1% bovine serum albumin (BSA) and 2 mM EDTA. After 5 min of centrifugation at 500×g 4 °C, supernatant was discarded and red blood cells were lysed using ammonium-chloride-potassium (ACK) Lysing Buffer (Gibco, A1049201). Cells were washed with HBSS 1% BSA 2 mM EDTA. Cells were pelleted and re-suspended in FACS (fluorescence-activated cell sorting) buffer (PBS containing 1% BSA and 2 mM EDTA). Cells were counted using the CellDrop FL automated cell counter (DeNovix).
The spleens were dispersed into single-cell suspensions by mechanical disruption using a gentleMACS Octo Dissociator with C-tubes filled with R10 media (RPMI-1640 – 10% FBS – 1 x P/S – 1x GlutaMAX). The homogenised tissues were run through 40 μm cell strainer (Fisher Scientific, Cat. No.: 22-363-547) into cold HBSS 1% BSA 2 mM EDTA. After 5 min of centrifugation at 500×g 4 °C, supernatant was discarded and red blood cells were lysed using ACK Lysing Buffer. Cells were washed with HBSS 1% BSA 2 mM EDTA and cell debris was removed by passing samples through 40 μm cell strainer. Cells were pelleted and re-suspended in FACS buffer. Cells were counted using the CellDrop FL automated cell counter (DeNovix).
Lung and bronchoalveolar lavage fluid (BALF) surface staining
Single cell suspensions from digested, red blood cell (RBC)-lysed lung or BALF were transferred to 96-well V-bottom plates (ThermoFisher) and centrifuged at 400×g for 5 min at room temperature. Supernatants were discarded and cell pellets were resuspended with 50 μL/well of Fc Block (BD, clone 2.4G2, RRID: AB_394656) diluted 1:100 in Staining Buffer (1% BSA with 2 mM EDTA in PBS), then incubated at room temperature for 5 min. After incubation with diluted Fc Block, 50 μL/well of the following surface staining cocktail diluted in Staining Buffer was added: CD3ε FITC (1:100, clone 145-2C11, BD, RRID: AB_394595), IgA PE (1:100, clone mA-6E1, Invitrogen, RRID: AB_465917), CD44 PE-CF594 (1:100, clone IM7, BD, RRID: AB_11153123), CD8α PerCP (1:100, clone 53-6.7, BD, RRID: AB_394573), CD69 PE-Cy7 (1:100, clone H1.2F3, BD, RRID: AB_394508), CD103 APC (1:100, clone 2E7, Invitrogen, RRID: AB_1106992), CD19 Alexa Fluor 700 (1:100, clone 1D3, Invitrogen, RRID: AB_837083), MHC II APC-Cy7 (1:100, clone M5/115.15.2, BD, RRID: AB_1659252), and Fixable Viability Dye eFluor 450 (1:200, Invitrogen). Cells were surface stained for 20 min at room temperature in the dark. After surface staining, 120 μL/well of Staining Buffer was added to all wells as the first wash step, then cells were centrifuged for 400×g for 5 min at room temperature. To wash a second time, supernatants were discarded, cells were resuspended with 200 μL/well of staining buffer, then centrifuged for 400×g for 5 min at room temperature. Supernatants were discarded and washed cell pellets were resuspended in 200 μL/well of staining buffer, then transferred to fluorescence-activated cell sorting (FACS) tubes for analysis. For quantification of absolute numbers, 5 μL of CountBright Absolute Counting Beads (Invitrogen) were added to each sample, barring unstained cell controls. For each single-stained compensation control, 1 μL of antibody was added to 1 drop of UltraComp eBeads Plus (BD).
Lung SARS-CoV-2 S-specific TRM surface staining
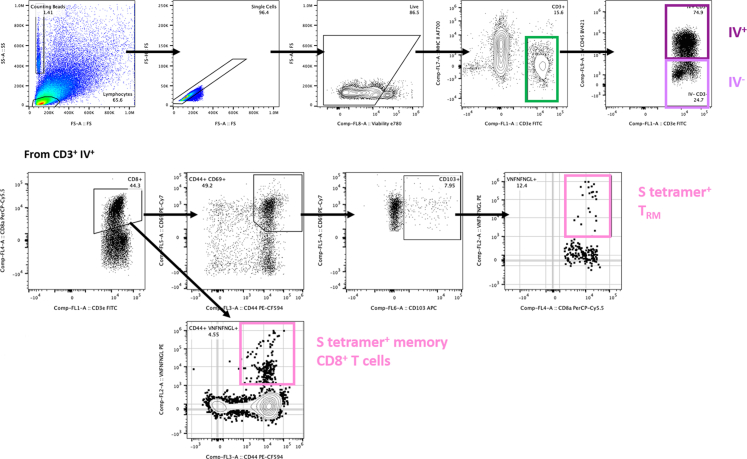
Single-cell suspensions from CD45 IV-labelled, digested and RBC-lysed lungs were transferred to 96-well V-bottom plates (ThermoFisher), centrifuged at 400×g for 5 min at room temperature, supernatants were decanted, then pellets were resuspended with 50 μL/well of diluted Fc Block, as described above. Cells were surface stained at room temperature for 20 min in the dark by adding 50 μL/well of the following surface staining cocktail diluted in staining buffer: CD3ε FITC (1:80, clone 145-2C11, BD, RRID: AB_394595), PE-conjugated H–2K(b) VNFNFNGL Tetramer (1:100, NIH Tetramer Core Facility), CD44 PE-CF594 (1:320, clone IM7, BD, RRID: AB_11153123), CD8α PerCP (1:160, clone 53-6.7, BD, RRID: AB_394573), CD69 PE-Cy7 (1:80, clone H1.2F3, BD, RRID: AB_394508), CD103 APC (1:80, clone 2E7, Invitrogen, RRID: AB_1106992), MHC II Alexa Fluor 700 (1:160, clone M5/115.15.2, Invitrogen, RRID: AB_494009), and Fixable Viability Dye eFluor 780 (1:800, Invitrogen). After surface staining, cells were washed twice with staining buffer, as described above. Following the wash steps, cells were resuspended with 100 μL/well of Cytofix Fixation Buffer (BD) and incubated at room temperature for 5 min in the dark. Cells were washed twice with staining buffer then resuspended with 200 μL/well of staining buffer. Plates were foil sealed and stored at 4 °C overnight in the dark. On the day of acquisition, cells were resuspended then transferred to FACS tubes for analysis. A fluorescence minus one (FMO) control was generated for PE and a strain, sex, and age-matched naïve mouse that did not receive CD45 BV421 IV-labelling was used to generate the unstained lung cell control. Addition of counting beads and preparation of compensation controls are as described above.
Blood SARS-CoV-2 S-specific T cell surface staining
One volume of blood was collected into four volumes of 0.5 M EDTA in centrifuge tubes to prevent coagulation, then pooled by group. Whole blood was stained with the direct addition of 1 μL of PE-conjugated H–2K(b) VNFNFNGL tetramer (1:100, NIH Tetramer Core Facility). Tubes were flicked to mix reagents then incubated for 40 min at room temperature in the dark. After incubation, the following antibodies or fluorescent dyes were added directly to tubes containing tetramer-stained whole blood: 1 μL of Fc Block (BD, clone 2.4G2, RRID: AB_394656), 1 μL of CD8α PerCP (clone 53-6.7, BD, RRID: AB_394573), 1 μL of CD3ε Alexa Fluor 700 (clone 17A2, BioLegend, RRID: AB_493696), 0.5 μL of MHC II eFluor 450 (clone M5/115.15.2, Invitrogen, RRID: AB_1272204), and 0.5 μL of Fixable Viability Dye eFluor 450 (Invitrogen). Tubes were flicked to mix and incubated at room temperature for 20 min in the dark. After incubation, 1 mL of staining buffer was added to each tube to quench staining, then tubes were centrifuged at 400×g for 5 min at 4 °C. Supernatants were removed using a pipette, then to fix and lyse RBCs, cell pellets were resuspended with 200 μL of 1X eBioscience Foxp3/Transcription Factor Fixation/Permeabilization Buffer (Invitrogen), prepared according to manufacturer instructions. Cells were incubated for 10 min at room temperature in the dark. After incubation, cells were washed twice by adding 1 mL of staining buffer to each tube, centrifuging at 400×g for 5 min at 4 °C, then removing supernatants. After the final wash, cell pellets were resuspended with 200 μL of staining buffer. Compensation controls were prepared as described above.
All flow cytometry samples were measured on a Beckman Coulter Gallios flow cytometer equipped with Kaluza data acquisition software. Analysis was performed using FlowJo 10.8.1 (Treestar) and compensated using the built-in AutoSpill algorithm.
Intracellular cytokine staining (ICS)
Splenocytes were resuspended in complete RPMI-1640 media (cRPMI, Gibco, Thermo Fisher Scientific) supplemented with 10% w/v FBS (Gibco), 100 U/mL penicillin, 100 μg/mL streptomycin (Gibco), and 2 mM L-Glutamine (Gibco). Cells were seeded in V-bottom 96-well plates (CELLSTAR, Greiner Bio-One North America Inc.) at an average of 2 × 106 cells/well in cRPMI media containing anti-mouse CD28 (1:500, BD, Cat# 557393, Clone 37.51, RRID: AB_394764), brefeldin A (1:1000, GolgiPlug™, BD, Cat# 555029), and monensin (1:1,000, GolgiStop™, BD, Cat# 554724). Splenocytes were ex vivo stimulated with PepMix™ SARS-CoV-2 or PepMix™ SARS-Cov-2 (Spike B.1.1.529/BA.1/Omicron), a pool of 315 (158 + 157) peptides (15-mer peptides with 11 amino acid overlap) spanning the full length of the ancestral or BA.1 spike glycoprotein (Cat# PM-WCPV-S-2, PM-SARS2-SMUT08-1; JPT Peptides), at a final individual peptide concentration of 5 μg/mL at 37 °C with 5% CO2 for 8–10 h. Negative control cells were stimulated with an equivalent volume of Dimethylsulfoxide (DMSO). Positive control cells were stimulated with a cocktail containing phorbol 12-myristate 13-acetate (0.5 mg/mL, Sigma–Aldrich) and ionomycin (1 mg/mL, Sigma–Aldrich). The unstimulated control cells were only treated with cRPMI media. After stimulation, cells were washed with PBS containing 2% FBS and centrifuged at 350×g for 5 min and then stained with Zombie Red™ diluted in PBS (1:500, BioLegend, Cat# 423109) for 15 min at RT in the dark. Cells were washed in PBS containing 2% FBS (350×g for 5 min) and incubated with surface staining cocktail containing Fc Block CD16/CD32 (1:50, BD, Cat# 553141, RRID: AB_394656) and the anti-mouse antibodies BV 711 CD3 (1:300, clone 17A2, BD, Cat# 100241, RRID: AB_2563945), Pacific Blue CD4 (1:400, clone GK1.5, BioLegend, Cat# 100428, RRID: AB_493647), PerCP/Cy5.5 CD8 (1:200, clone 53-6.7, BioLegend, Cat#100734, RRID: AB_2075238), BUV737 anti-CD44 (1:400, clone IM7, BD, Cat# 612799, RRID: AB_2870126), and anti-CD154 (1:100, clone MR1, Invitrogen, Cat# 12–154, RRID: AB_465887) for 30 min at 4 °C in FACS buffer. Cells were washed in FACS buffer and then incubated in fixation/permeabilization buffer (CytoFix, BD, Cat# 554714) for 5 min at 4 °C. After fixation, cells were washed in 1 × permeabilization buffer (CytoPerm, BD, Cat# 554714), then incubated with the intracellular staining cocktail containing anti-mouse antibodies Alexa Fluor 647 IFN-γ (1:400, clone XMG1.2, BioLegend, Cat# 505814, RRID: AB_493314), Alexa Fluor 488 TNF-α (1:300, clone MP6-XT22, BioLegend, Cat# 506313, RRID: AB_493328), PE/Cy7 IL-2 (1:300, clone JES6-5H4, BioLegend, Cat# 503832, RRID: AB_2561750), APC IL-4 (1:400, clone 11B11, BioLegend, Cat# 504106, RRID: AB_315320), and BV650 IL-17 (1:400, clone TC11-18H10, BD, Cat# 564170, RRID: AB_2738641) in 1 × permeabilization buffer for 1 h at 4 °C. Samples were then washed in 1 × permeabilization buffer and resuspended in PBS buffer for acquisition. Samples were measured on an Aurora spectral cytometer (Cytek) using SpectroFlo® software (Cytek), with the relevant single fluorochrome compensation controls set by the daily acquisition of Cytometer Setup and Tracking beads. Analysis was performed with FCS Express 7 (DeNovo Software) and GraphPad Prism 9.5.1 (GraphPad Software).
B and T cell flow cytometry
To measure SARS-CoV-2 spike antigen-specific B cells, tetrameric B cells probes were generated. Specifically, spike (S) or receptor-binding domain (RBD) proteins of the ancestral or BA.1 strains, biotinylated at the carboxyl-terminal avitag were purchased from R&D Systems (ancestral spike: AVI10549-050; BA.1 spike: AVI11060-050; ancestral RBD: AVI10500-050; BA.1 RBD: AVI11056-050). Streptavidin (SAV) that was fluorescently labelled with R-phycoerythin (PE), allophycocyanin (APC) or brilliant violet 711 (BV711) was purchased from BioLegend (PE Steptavidin: Catalog No. 405204; APC Steptavidin: Catalog No. 405243; BV711 Steptavidin: 405241). Fifty microliters of the biotinylated S was mixed with PE-SAV and APC-SAV at a molar ratio of 6:1, or fifty microliters of the biotinylated RBD was mixed with APC-SAV and BV711-SAV at a molar ratio of 4:1. The mixtures were incubated at 4 °C for 1 h with rocking and the reaction was quenched by an equal volume of 4 mM free biotin with another 30 min rocking at 4 °C. Free biotin was removed with Amicon ultra-0.5 (50Kd molecular weight cut-off) spin column at 14,000×g for 5 min at 4 °C. The samples were washed once with cold PBS and were concentrated via spinning at 14,000×g for 5 min at 4 °C. The volume was brought up with cold sterile PBS to reach a spike protein concentration of 200 ng/mL and a RBD protein concentration of 50 ng/mL. For surface staining of the splenocytes, 1 × 106 live cells were seeded into 96-well V-bottom plate. Cells were spun at 500×g at 4 °C for 5 min and were first stained with 1 μL of each of the two or three probes (PE-SAV-S vs APC-SAV-S; PE-SAV-S vs APC-SAV-S vs BV711-SAV-RBD; APC-SAV-RBD vs BV711-SAV-RBD) in a total of 50 μL of the FACS buffer for 1 h at 4 °C in the dark. One hundred microliters of the FACS buffer were added to each well and cells were spun down and washed once with another 150 μL FACS buffer. One hundred microliters of FACS buffer containing Fc block (anti-CD16/32, Catalog # 14-0161-82 eBioscience) at 0.5 mg/test were added to each well for 10 min at 4 °C before staining the cells with 50 μL of the antibody cocktail for 30 min in the dark at 4 °C. The antibody cocktail contains Zombie Red (1:400, BioLegend, Cat# 423109) as the viability dye, Pacific Blue anti-mouse CD3 (1:200, clone 17A2, BioLegend, Cat# 100214, RRID: AB_493645), Alexa Fluor 700 anti-mouse/human CD45R/B220 (1:200, clone RA3-6B2, BioLegend, Cat# 103232, RRID: AB_493717), Brilliant Violet 785 anti-mouse CD19 (1:100, clone 6D5, BioLegend, Cat# 115543, RRID: AB_11218994), Brilliant Violet 605 anti-mouse IgD (1:200, clone 11-26c.2a, BioLegend, Cat# 405727, RRID: AB_2562887), APC/Cyanie7 anti-mouse IgM (1:400, clone RMM-1, BioLegend, Cat# 406516, RRID: AB_10660305), PerCP/Cyanine 5.5 anti-MU/HU GL7 antigen (1:50, clone GL7, BioLegend, Cat# 144610, RRID: AB_2562979), and PE/Cyanine 7 anti-mouse CD38 (1:400, clone 90, BioLegend, Cat# 102718, RRID: AB_2275531). The antibody cocktail was decanted and cells were washed once with FACS buffer. To fix the cells, 50 μL of the 4% methanol-free paraformaldehyde (PFA) was added and incubated for 30 min in the dark at 4 °C. PFA was then decanted and cells were washed once with FACS buffer. Cells were then resuspended in FACS buffer and kept in the dark at 4 °C before acquisition. Samples were measured on an Aurora spectral cytometer (Cytek, Fremont, CA, USA) using SpectroFlo® software (Cytek), with the relevant single fluorochrome unmixing reference controls set by the daily acquisition of Cytometer Setup and Tracking beads. Analysis was performed with FCS Express 7 (DeNovo Software) and GraphPad Prism 9.5.1 (GraphPad Software). Total antigen-specific memory B cells (MBC) were gated as live CD3-B220+CD19+IgD−GL7−CD38hi double-probe positive population and isotype switched (sw) antigen-specific memory B cells (swMBC) were gated as live CD3-B220+CD19+IgD−IgM-GL7−CD38hi double-probe positive population. The frequency of the antigen-specific MBC or swMBC out of the total MBC or swMBC respectively was calculated and graphed.
Statistical analysis
The details about the type of statistical analysis conducted for each experiment are listed in the respective figure legends. Briefly, the statistical analysis was performed using GraphPad Prism 9.5.1 (GraphPad Software). One-way analysis of variance (ANOVA) or Kruskal–Wallis test was used when only one factor in form of different vaccination groups was analysed as in T and B cell studies or plaque assays. Two-way ANOVA was chosen (sometimes after log transforming the data to normalise it), when two factors in form of different vaccination groups, different time points, different cytokines, different virus variants or different antigens were analysed as in ELISAs or ICS. ANOVAs and Kruskal–Wallis tests were followed by Tukey’s or Dunn’s multiple comparisons test. For Tukey’s multiple comparison test, every mean was compared with every other mean. For Dunn’s multiple comparisons test, the mean of each group was compared with the mean of a control group (like PBS). Significance was considered with p values equal or less than 0.05.
Role of funders
The funding sources for this study had no role in the study design, data collection, analysis, interpretation, or writing of the manuscript.
Results
Design and production of NDV-HXP-S Omicron BA.1 and BA.5 variant vaccines
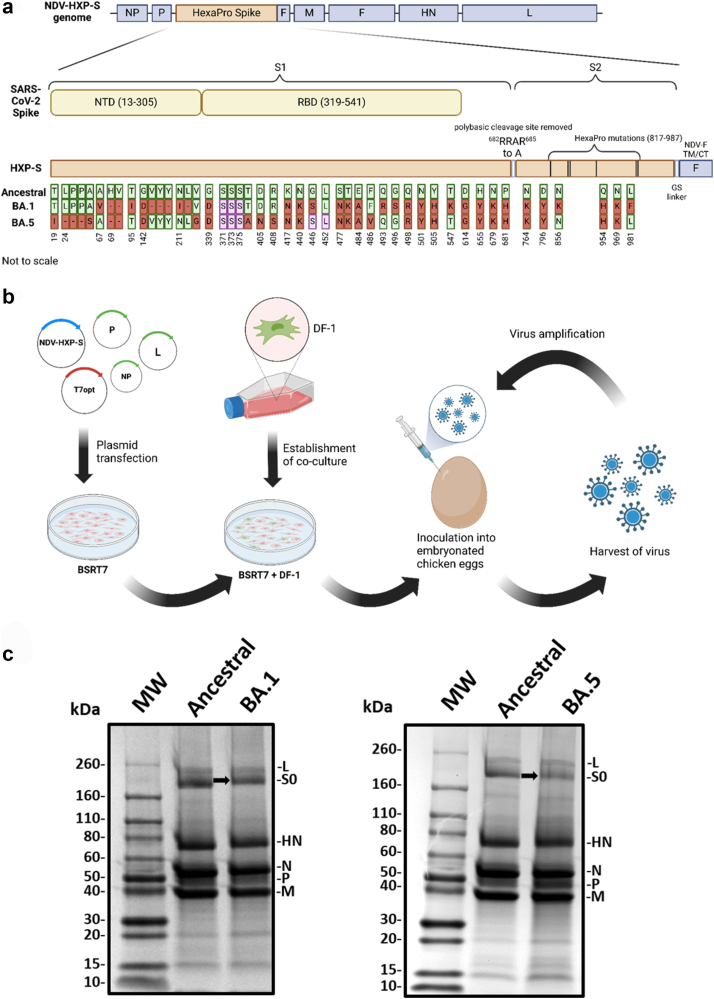
NDV-HXP-S vaccines expressing a modified SARS-CoV-2 spike of BA.1 or BA.5 variants were generated.32 The SARS-CoV-2 spike was stabilised in its prefusion (S0) conformation by introduction of six proline mutations into the spike S2 region (F817P, A892P, A899P, A942P, K986P, and V987P) and elimination of the polybasic cleavage site (682RRAR685 to 682--A-685) (Fig. 1a).40 The HXP-S was anchored in the membrane of the NDV virion by replacing the transmembrane domain (TM) and cytoplasmic tail (CT) of the spike with those from the fusion (F) protein of LaSota NDV. The transgene was inserted between the P and the M genes. The HXP-S nucleotide sequence was codon-optimised for mammalian host expression and 36 mutations specific for VOC BA.1 or 35 mutations specific for VOC BA.5 were introduced compared to the ancestral NDV-HXP-S (Fig. 1a). However, both Omicron spikes proved to be less stable than the ancestral strain when the Omicron specific mutations S371L, S373P, and S375F were included.44 Genetic engineering was performed by keeping those amino acid positions as the ancestral serines to increase spike stability of BA.1. Furthermore, position R452 in the BA.5 was changed back to 452 L of the ancestral strain to further enhance stability.45 Additionally, G446S was included for better T cell recognition in the BA.5 spike (Fig. 1a).46
Fig. 1.
Design and production of NDV-HXP-S Omicron BA.1 and BA.5 variant vaccines. (a) Structure and design of the NDV-HXP-S genome. The SARS-CoV-2 HexaPro spike sequences were inserted between the P and the M genes of the LaSota NDV strain. The ectodomain of the spike was connected to the transmembrane domain and the cytoplasmic tail (TM/CT) of the F protein of NDV (ectodomain of the spike in beige; NDV components in blue). The original polybasic cleavage site was deleted by mutating 682RRAR685 to 682--A-685. The spike-stabilizing HexaPro (F817P, A892P, A899P, A942P, K986P, and V987P) mutations were introduced. The sequence was codon-optimised for mammalian host expression. The ancestral HXP-S sequence aligned with the new Omicron variants BA.1 and BA.5 is depicted. Amino acids of the ancestral original HXP-S sequence are shown in green while changes for BA.1 or BA.5 are depicted in red. Amino acids that have been genetically engineered are shown in purple. (b) Experimental scheme of NDV-HXP-S variant rescue. BSRT7 cells were transfected with pNDV-HXP-S and the helper plasmids pTM1-NP, pTM1-P, pTM1-L, and pCI-T7opt.26 The next day, transfected BSRT7 cells were co-cultured with DF-1 cells. After two or three days of incubation, co-cultures were inoculated into 8- or 9-day old specific pathogen-free (SPF) embryonated chicken eggs. Following three days of incubation at 37 °C, eggs were cooled at 4 °C overnight and subsequently virus containing allantoic fluid was harvested. The recombinant virus was amplified and passaged multiple times to confirm genetic stability. (c) Protein analysis of NDV-HXP-S BA.1 and BA.5. NDV-HXP-S BA.1, BA.5 and ancestral were purified from allantoic fluid via ultra-centrifugation through a sucrose cushion and resuspended in PBS. The purified viruses were resolved on 4–20% SDS page and the viral proteins visualised by Coomassie blue staining (L, S0, HN, N, P, and M). The uncleaved SARS-CoV-2 S0 spike protein is highlighted with an arrow.
The NDV-HXP-S variant vaccines were rescued using reverse genetics in BSRT7-DF-1 co-cell cultures and further amplified in specific pathogen-free (SPF) embryonated chicken eggs as described previously (Fig. 1b).32,33 The vaccine virus was concentrated and cleared of soluble egg protein by subjecting the allantoic fluid to ultra-centrifugation through a 20% sucrose cushion. Subsequently, the virus was analysed by sodium dodecyl-sulfate polyacrylamide gel electrophoresis (SDS-PAGE) with Coomassie blue staining to verify spike expression and stability. NDV-HXP-S BA.1 as well as BA.5 displayed a band below the NDV L protein, between the 260 kDa and 160 kDa weight markers, in accordance with the size of the uncleaved S0 protein (Fig. 1c).32 The BA.1 as well as the BA.5 construct grew to similar titers and presented similar spike expression levels compared to that of the ancestral spike vaccine virus.
Intranasal administration of NDV-HXP-S vaccine induces mucosal humoral and T cell memory
To highlight the potential of live NDV-HXP-S IN vaccination for the induction of mucosal immunity, BALB/c mice were vaccinated once or twice IN with 105 or 106 fifty percent of egg embryo infectious dose (EID50) of NDV-HXP-S expressing ancestral spike four weeks apart (Fig. 2a). A vector-only control group was included using 106 EID50 of WT LaSota NDV, while a negative control group was mock-vaccinated with PBS. Serum and nasal washes were collected four weeks after the prime, eight weeks after the prime and four weeks after the boost. Additionally, bronchoalveolar lavage fluid (BALF) was collected eight weeks after the prime and four weeks after the boost and lungs were collected four weeks after the boost (Fig. 2a).
Fig. 2.
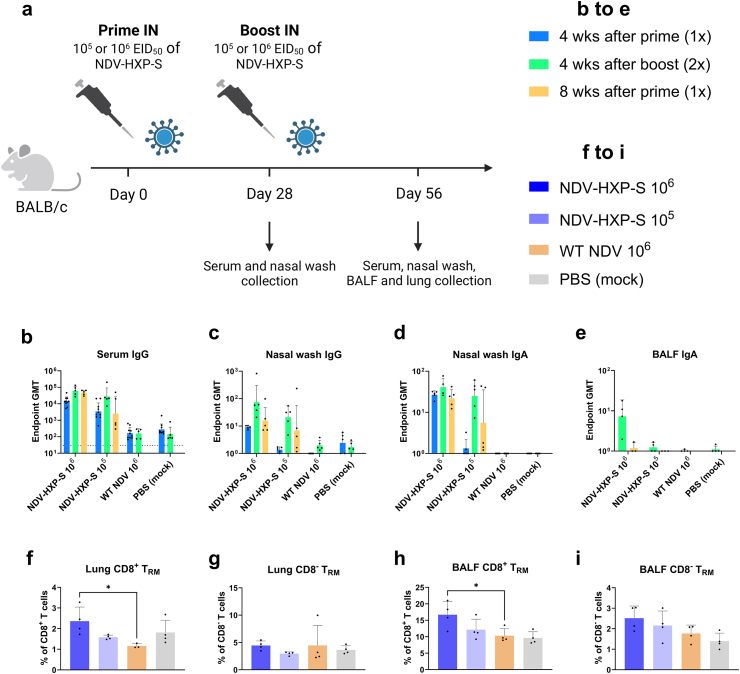
Intranasal administration of NDV-HXP-S vaccine induces mucosal humoral and T cell memory. (a) Design of the study. Six to eight week-old female BALB/c mice were intranasally immunised with 105 or 106 fifty percent of egg embryo infectious dose (EID50) of NDV-HXP-S expressing ancestral spike. Four weeks later, half of the mice were boosted. The empty WT LaSota NDV vector (106 EID50) and PBS were used for vector-only and negative control groups, respectively. Twenty-eight days after prime, sera (n = 10) and nasal washes (n = 5 or 4) were collected and 56 days after prime (28 days after boost), serum (n = 5), nasal washes (n = 5 or 4), and bronchoalveolar lavage fluid (BALF) (n = 4 or 3) were collected to assess binding antibody responses. Lung tissues and BALF were collected for T cell analysis (n = 5 or 4). (b–e) Measurement of ancestral spike-specific (b) serum IgG, (c) nasal wash IgG, (d) nasal wash IgA, and (e) BALF IgA in naïve mice, mice immunised with empty vector, mice immunised with 105 EID50 or mice immunised with 106 EID50 of NDV-HXP-S ancestral; at four weeks after prime, four weeks after boost and eight weeks after prime. The geometric mean titer (GMT) endpoint titers of the ELISA were graphed (an endpoint titer of one was assigned to negative samples). The dashed line indicates the limit of detection (for nasal wash and BALF x-axis equals limit of detection). The error bars represent geometric standard deviation (SD). (f–i) Staining for CD3, CD8, CD44, CD69, and CD103 was used to assess tissue resident memory cells (TRM) (f) in the lungs and positive for CD8, (g) in the lungs and negative for CD8, (h) in the BALF and positive for CD8 and (i) in the BALF and negative for CD8. Mean with SD was graphed. Statistical significance was calculated by means of [(b)–(e)] log transforming the data set to normalise followed by a two-way analysis of variance (ANOVA) or [(f)–(i)] one-way ANOVA followed by Tukey’s multiple comparisons test. The experiment was conducted twice, pooled data are shown ∗p < 0.05.
Even a single IN vaccination with the higher dose of 106 EID50 induced a strong systemic immune response with comparable levels of serum IgG against the ancestral SARS-CoV-2 spike between single and double immunization groups (with low levels of non-specific binding encountered in the negative control groups) (Fig. 2b). However, the lower vaccination dose needed a boost to reach serum IgG titers of similar magnitude. Corroborating results were obtained in nasal wash antibody levels, as only mice receiving two doses developed high levels of anti–SARS-CoV-2 spike IgA and IgG (Fig. 2c and d). In our hands, only low levels of IgA were detected in BALF (Fig. 2e). This might be due to the low vaccination volume of 10 μl used, which typically prevents the mice from deeply inhaling the vaccine and thereby keeping the NDV-HXP-S in the upper airways (not the lungs). In our opinion, keeping the vaccine in the upper respiratory tract of the mice closely imitates intranasal vaccination techniques in humans.
Interestingly, we observed an increase in titers between collection time points four and eight weeks post prime even without an additional immunization. This could indicate prolonged germinal center (GC) reactions.47, 48, 49
Although memory T cells do not provide sterilizing immunity, they are key in overcoming viral infections once the virus has escaped neutralizing humoral immunity. Furthermore, memory T cells have been associated with improved clinical outcome in SARS-CoV-2 infection.50, 51, 52 Therefore, we assessed tissue-resident memory T cells in the lungs of mice vaccinated twice. An overall increase of CD8+ T cells with the canonical markers of T resident memory (TRM) cells, CD69 and CD103, was observed in lung tissues after vaccination with 106 EID50 of NDV-HXP-S (Fig. 2f and g). A similar increase in TRM cells was detected in BALF of animals that received 106 EID50 of NDV-HXP-S (Fig. 2h and i).
Thus, intranasal administration of NDV-HXP-S induces local T cell responses in the lungs as well as high IgA levels in the upper respiratory tract. Overall, vaccinating twice three or four weeks apart proved to be the most effective of the tested regimens, especially for inducing mucosal immunity, and was therefore utilised going forward.
NDV-HXP-S BA.1 and BA.5 variant vaccinations increase strain specific humoral immunity
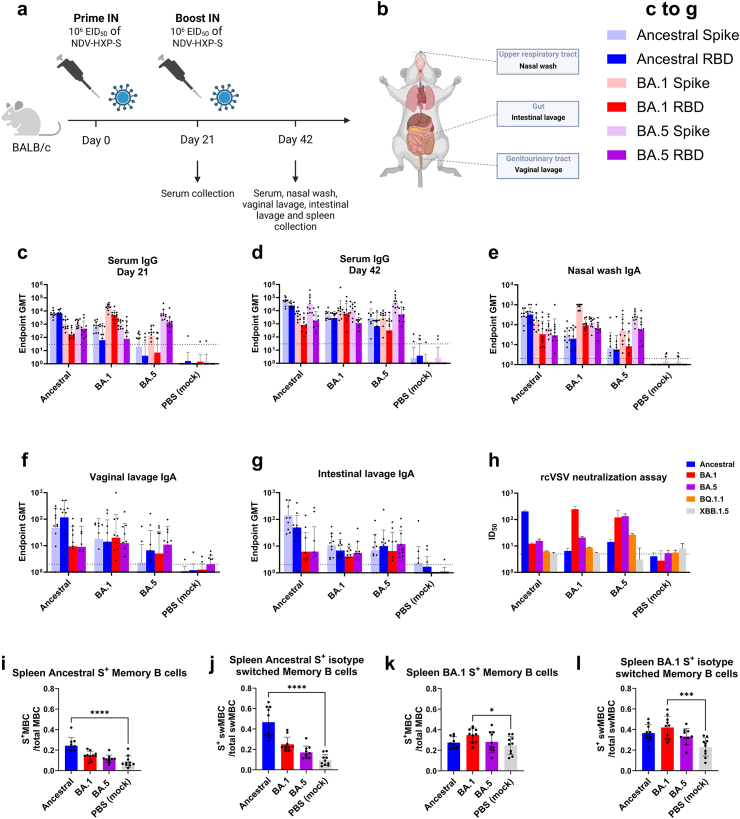
In order to test the immunogenicity of the newly developed NDV-HXP-S BA.1 and BA.5 vaccine candidates, BALB/c mice were vaccinated twice three weeks apart IN with 106 EID50 of NDV-HXP-S expressing ancestral, BA.1 or BA.5 spike (Fig. 3a). Mice mock-vaccinated with PBS were included as negative control group. On day 21 post-prime serum was collected before administering the booster. Three weeks later, serum, nasal washes, vaginal lavages, and intestinal lavages were harvested (Fig. 3b). Subsequently, antibody titers in serum, nasal washes, vaginal lavages, and intestinal lavages were tested against ancestral spike, ancestral RBD, BA.1 spike, BA.1 RBD, BA.5 spike, and BA.5 RBD (Fig. 3c–g). At 21 days after the prime, serum IgG levels of the ancestral spike vaccination group against ancestral spike and RBD were already very high with an endpoint titer of around 104 (Fig. 3c). Similarly, the BA.1 and BA.5 vaccination groups had high IgG titers against homologous spikes and RBDs (the same spike/RBD as the vaccine) (Fig. 3c). However, IgG titers against heterologous spikes and RBDs (spikes/RBDs different from the vaccine) were comparatively low (with for example the BA.1 vaccination group having 30 times higher IgG titers against the BA.1 spike than the ancestral spike). At 21 days after the booster, binding titers against homologous spikes and RBDs were further increased (Fig. 3d). Additionally, serum IgG levels against heterologous spikes and RBDs were greatly improved, with endpoint titers increasing around tenfold after boosting (Fig. 3d). While the broadening of the serum antibody response via boosting improved the titers of the ancestral spike vaccination group against Omicron antigens, the serum antibodies of the Omicron vaccine groups were still superior at binding the corresponding Omicron RBDs (Fig. 3c and d). This is due to a majority of the antibody evading mutations being located in the RBD region. Similarly, BA.1 vaccination induced the highest IgA titers against BA.1 antigens in the tested nasal washes, while BA.5 vaccination elicited the highest IgA titers against BA.5 antigens (Fig. 3e). The same trend was observed in vaginal and intestinal lavages (Fig. 3f and g; due to the limited quantity of vaginal and intestinal lavages, not all ELISAs could be run against all antigens). In summary, the Omicron variant vaccine candidates show an increased ability to induce IgG and IgA against BA.1 and BA.5 RBD. This advantage in binding variant RBDs correlated with an increase in serum neutralization titers against BA.1 and BA.5 pseudotyped virus, respectively (Fig. 3h). The antigenically distant variants BQ.1.1 and XBB.1.5 were only neutralised at the highest concentrations by the mismatched vaccine sera.
Fig. 3.
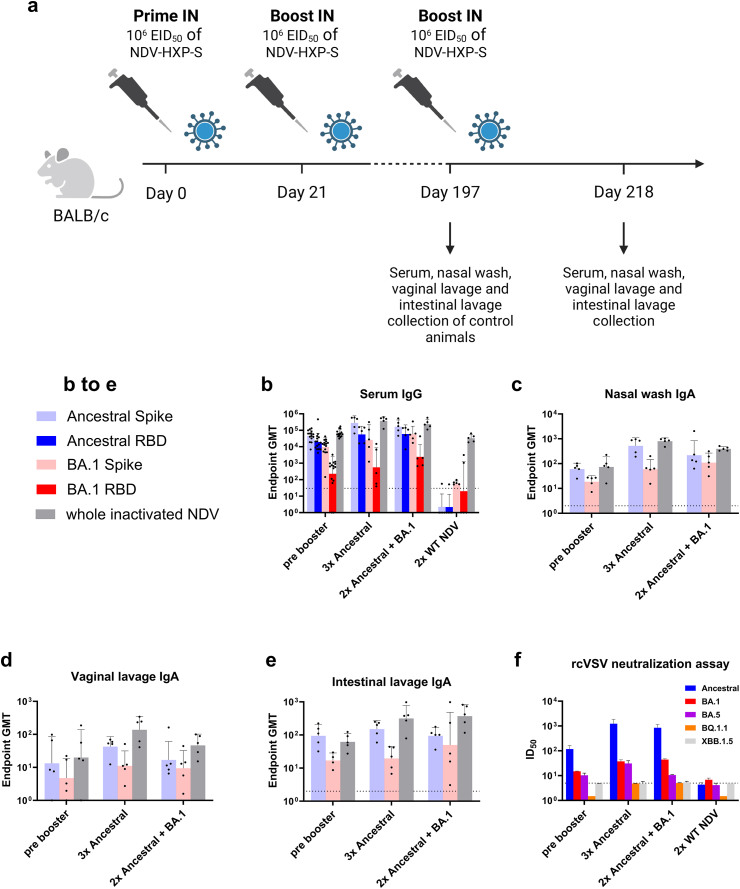
NDV-HXP-S BA.1 and BA.5 variant vaccinations increase strain specific humoral immunity. (a) Design of the study. Six to eight week-old female BALB/c mice were intranasally immunised twice 21 days apart with 106 EID50 of NDV-HXP-S ancestral, BA.1 or BA.5. PBS was used for a negative control group. Twenty-one days after prime, serum was collected and 42 days after prime (21 days after boost), serum, nasal washes, vaginal lavages, and intestinal lavages were collected to assess binding and neutralizing antibody response. Spleens were collected for memory B cell analysis. (n = 10) (b) Anatomical scheme of harvesting nasal wash, intestinal lavage, and vaginal lavage. From each animal, nasal washes from the upper respiratory tract, intestinal lavages from the gut and vaginal lavages from the genitourinary tract were harvested for IgA measurement. (c–e) Measurement of ancestral spike, ancestral RBD, BA.1 spike, BA.1 RBD, BA.5 spike, and BA.5 RBD-specific (c) serum IgG at the 1st time point (21 days post prime), (d) serum IgG at the 2nd time point (42 days post prime, 21 days post booster) and (e) nasal wash IgA in mice (mock) immunised with PBS or 106 EID50 of NDV-HXP-S ancestral, BA.1 or BA.5. (f and g) Measurement of ancestral spike, ancestral RBD, BA.1 RBD, and BA.5 RBD-specific (f) vaginal lavage IgA and (g) intestinal lavage IgA in mice (mock) immunised with PBS or 106 EID50 of NDV-HXP-S ancestral, BA.1 or BA.5. The GMT endpoint titers of the ELISA assay were graphed (an endpoint titer of one was assigned to negative samples). The dashed line indicates the limit of detection. The error bars represent geometric SD. (h) Neutralization titers against SARS-CoV-2 ancestral, BA.1, BA.5, BQ.1.1, or XBB.1.5 spike-pseudotyped vesicular stomatitis virus expressing GFP (rcVSVeGFP-CoV-2-S) were measured in technical duplicates from pooled sera. Geometric mean with geometric SD of the inhibitory dilution at which 50% neutralization is achieved (ID50) was graphed (ID50 of 1.5 was assigned to negative samples). The dashed line indicates the limit of detection. (i–l) Using two separate tetrameric B cell probes for ancestral or BA.1 spike respectively, B cell subsets in the spleens were measured. This included (i) ancestral spike specific memory B cells (CD3- Live+ B220+ CD19+ IgD− GL7- CD38+), (j) isotype switched memory B cells specific for ancestral spike (CD3- Live+ B220+ CD19+ IgM− IgD− GL7- CD38+), (k) BA.1 spike specific memory B cells and (l) isotype switched memory B cells specific for BA.1. Mean with SD was graphed. Statistical significance was calculated by means of [(c)–(g)] log transforming the data set to normalise followed by a two-way ANOVA or [(i)–(l)] one-way ANOVA followed by Tukey’s multiple comparisons test. ∗p < 0.05; ∗∗p < 0.01; ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001.
To further determine the protective capability of serum antibodies induced by IN NDV-HXP-S vaccination, a passive transfer experiment was conducted (Supplementary Fig. S1a). To this end, sera from each vaccination group was pooled and 150 μl was administered intraperitoneally (IP) to each naïve K18-hACE2 mouse. Two hours post transfer, the mice were challenged with 3 × 104 plaque-forming units (PFU) of SARS-CoV-2 BA.1 virus. A healthy control group was kept uninfected. Infectious viral titers in the lungs and nasal turbinates were measured five days after the challenge as readout of protection (Supplementary Fig. S1b and c). Limited by the titer of our BA.1 virus stock, the titers detected in tissues were relatively low. While the poor virus replication diminished the effect of vaccine-specific protection, we did observe a reduction of viral titers in the lung homogenates of BA.1 serum recipient mice compared to the PBS serum recipients (Supplementary Fig. S1b). Additionally, a trend of reduced viral titers in nasal turbinates was observed in mice receiving BA.1 or BA.5 serum (Supplementary Fig. S1c).
In secondary immune responses, memory B lymphocytes provide rapid and efficient immune protection and are therefore of utmost importance to long lasting immunity.53 A big cluster of that population can be found in the spleen.54, 55, 56, 57 To measure antigen-specificity of memory B cells (MBCs) in the spleens, two separate tetrameric B cell probes for ancestral or BA.1 spike, respectively, were used to stain memory B cells (Supplementary Fig. S1). Additionally, we gated for class-switched memory B cells that lack IgM and IgD (therefore expressing IgG, IgA or IgE). It was observed that vaccination with NDV-HXP-S expressing ancestral spike led to significantly increased frequencies of ancestral spike-specific memory B cells as well as isotype switched memory B cells in spleens (Fig. 3i and j). Likewise, administration of NDV-HXP-S BA.1 induced a pool of BA.1 spike-specific memory B cells, many of which were isotype switched (Fig. 3k and l).
Thus, the NDV-HXP-S variant vaccines induce potent mucosal and systemic antibody responses against the respective VOC as well as antigen-specific memory B cells.
Intranasal immunization with NDV-HXP-S variant vaccines protects mice from homologous infection
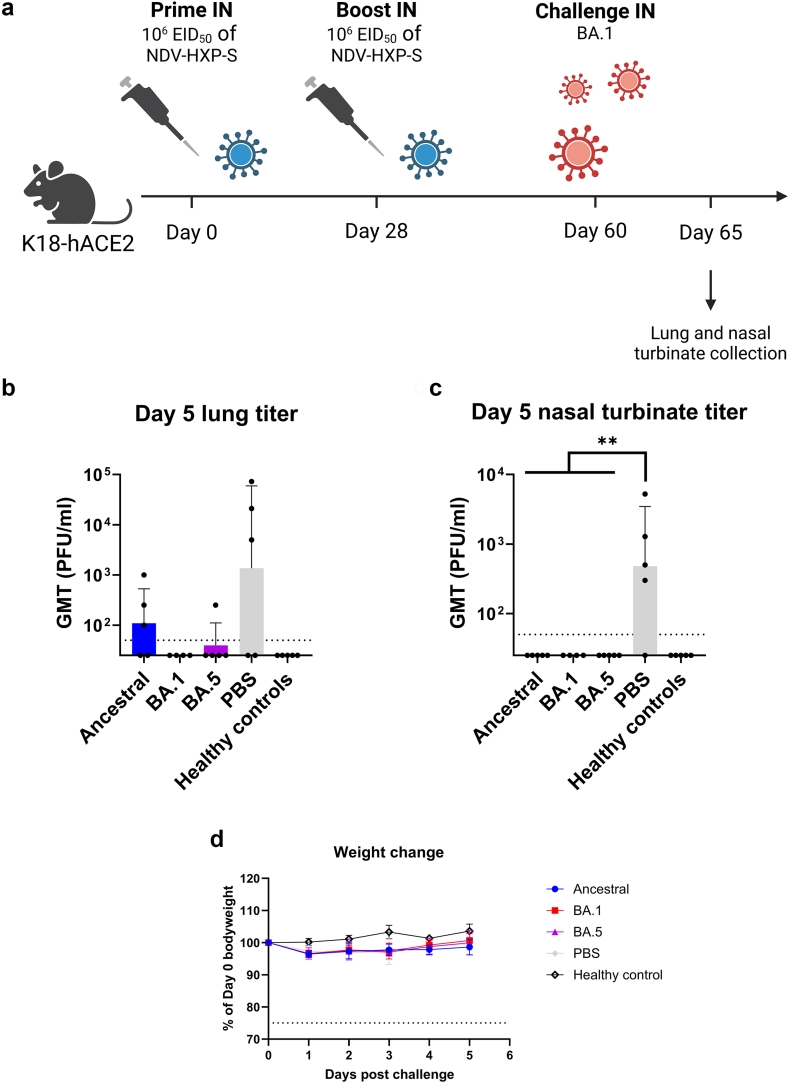
A challenge experiment in K18-hACE2 mice was conducted to test the in vivo protection provided by IN NDV-HXP-S variant vaccination. To that end, mice were vaccinated twice four weeks apart IN with 106 EID50 of NDV-HXP-S expressing ancestral, BA.1 or BA.5 spike (Fig. 4a). In addition, a group mock-vaccinated with PBS was included as negative control. One month after the boost, the mice were challenged with 3 × 104 PFU of SARS-CoV-2 BA.1 virus (Fig. 4a). A healthy control group was kept uninfected. Five days after the challenge, lungs and nasal turbinates were collected and infectious viral titers in the tissues were determined (Fig. 4b and c). No infectious virus was detected in lung homogenates of mice immunised with the NDV-HXP-S BA.1 variant vaccine matching the challenge virus used (Fig. 4b). One animal of the BA.5 vaccination group presented with low infectious titers in the lungs compared to three out of five mice receiving the ancestral vaccine (Fig. 4b). All NDV-HXP-S vaccines prevented virus replication in the nasal turbinates (Fig. 4c). No significant weight loss was detected in any of the animals (Fig. 4d), as has been described by others.58,59 Therefore, mice IN vaccinated with NDV-HXP-S BA.1 were protected from infection with SARS-CoV-2 BA.1 with no virus found in the tissues tested.
Fig. 4.
Intranasal immunization with NDV-HXP-S variant vaccines protects mice from homologous infection. (a) Design of the study. Six to eight week-old female K18-hACE2 mice were intranasally immunised twice 28 days apart with 106 EID50 of NDV-HXP-S ancestral, BA.1 or BA.5. PBS was used for a negative control group. Thirty-two days after the boost, the mice were challenged with 3 × 104 plaque-forming units (PFU) of SARS-CoV-2 BA.1 (n = 5 or 4). One animal of the BA.1 vaccination group died during the challenge process unrelated to the vaccination or virus. A healthy control group was kept uninfected (n = 5). (b and c) Measurement of viral titers in the (b) lungs and (c) nasal turbinates. Tissues of the animals were collected on day 5 post challenge. The whole lungs or nasal turbinates were homogenised in 1 mL of PBS. The viral load was measured by plaque assay on VERO E6 TMPRSS2 T2A ACE2 cells and graphed as GMT of PFU/ml (a titer of 25 PFU/mL was assigned to negative samples). The dashed line indicates the limit of detection. The error bars represent geometric SD. (d) Body weight of mice infected with SARS-CoV-2 BA.1. Data are mean with SD. Statistical significance was calculated by means of [(b) and (c)] Kruskal–Wallis test followed by Dunn’s multiple comparisons test. ∗p < 0.05; ∗∗p < 0.01.
Comparing these results to the passive transfer data (Supplementary Fig. S1), fewer of the IN vaccinated animals had measurable viral titers than the animals receiving sera. This emphasises the importance of the mucosal aspect of immunity induced by IN vaccination.
A third vaccination increases mucosal as well as serum antibody titers with no interference due to vector immunity
An often-discussed topic when using viral vector vaccines is the immunity developing against the vector itself and how it could interfere with subsequent boosting. In order to address these concerns, we vaccinated mice twice IN with 106 EID50 of NDV-HXP-S expressing ancestral spike three weeks apart (Fig. 5a). WT LaSota NDV was used for a vector-only control group. After 197 days, serum, nasal washes, vaginal lavages, and intestinal lavages of the pre booster group were collected. The other groups were vaccinated a third time IN with 106 EID50 of NDV-HXP-S expressing ancestral or BA.1 spike (at the time of this experiment, BA.5 was not yet available). Three weeks later, serum, nasal washes, vaginal lavages, and intestinal lavages of the 3x ancestral and 2x ancestral + BA.1 group were collected.
Fig. 5.
A third vaccination increases mucosal as well as serum antibody titers with no interference due to vector immunity. (a) Design of the study. Six to eight week-old female BALB/c mice were intranasally immunised twice 21 days apart with 106 EID50 of NDV-HXP-S ancestral or WT LaSota NDV vector. After 197 days, serum of the WT NDV control group as well as serum, nasal washes, vaginal lavages, and intestinal lavages of the animals of the control group pre booster (vaccinated twice with NDV-HXP-S expressing ancestral spike) were collected to assess binding and neutralizing antibody response. The other mice were boosted IN with 106 EID50 of NDV-HXP-S ancestral or BA.1. Twenty-one days later, on day 218, serum, nasal washes, vaginal lavages, and intestinal lavages of booster groups 3x ancestral and 2x ancestral + BA.1 were collected to assess binding and neutralizing antibody response. (n = 5 or 15) (b) Measurement of ancestral spike, ancestral RBD, BA.1 spike, BA.1 RBD, and whole inactivated NDV-specific serum IgG. (c–e) Measurement of ancestral spike, BA.1 spike, and whole inactivated NDV-specific (c) nasal wash IgA, (d) vaginal lavage IgA and (e) intestinal lavage IgA in mice immunised twice with 106 EID50 of NDV-HXP-S ancestral, three times with NDV-HXP-S ancestral or immunised twice with NDV-HXP-S ancestral and boosted once with BA.1. The GMT endpoint titers of the ELISA assay were graphed (an endpoint titer of one was assigned to negative samples). The dashed line indicates the limit of detection (for vaginal lavage x-axis equals limit of detection). The error bars represent geometric SD. (f) Neutralization titers against SARS-CoV-2 ancestral, BA.1, BA.5, BQ.1.1, or XBB.1.5 spike-pseudotyped vesicular stomatitis virus expressing GFP (rcVSVeGFP-CoV-2-S) were measured in technical duplicates from pooled sera. Geometric mean with geometric SD of the ID50 was graphed (ID50 of 1.5 was assigned to negative samples). The dashed line indicates the limit of detection. Statistical significance was calculated by means of [(b)–(e)] log transforming the data set to normalise followed by a two-way ANOVA followed by Tukey’s multiple comparisons test.
We only observed a small decline in serum IgG as well as nasal wash IgA titers of mice over 170 days after their last vaccination (compare pre-booster mice Fig. 5b with 2 b and 5c with 2 d), potentially indicating prolonged GC reactions.47, 48, 49 While antibody titers against whole inactivated NDV increased in all tested samples after boosting, IgG and IgA levels against SARS-CoV-2 spikes increased to a similar degree (Fig. 5b–e). A third vaccination with the ancestral spike-specific vaccine further increased serum IgG levels against ancestral spike and RBD (Fig. 5b). The BA.1 booster increased the immunity against ancestral antigens to a lesser extent, but improved anti-BA.1 spike and RBD IgG levels (Fig. 5b). A similar increase was observed for IgA levels in nasal washes, vaginal, and intestinal lavages (Fig. 5c–e). Accordingly, both boosters improved serum neutralization titers against the ancestral pseudotyped virus substantially (Fig. 5f). However, while the BA.1 booster improved ancestral specific neutralization, it only increased BA.1 specific neutralization titers threefold. We suspect that this is due to antigenic imprinting, as similar results have been reported for a variety of SARS-CoV-2 vaccines.48,60, 61, 62 This is comparable to the phenomenon of original antigenic sin observed in anti-influenza virus immune responses.63,64
In summary, titers against ancestral spike can be increased by three separate vaccinations with NDV as a vector (3x ancestral or 2x ancestral + BA.1). These results support multiple vaccinations with the same NDV vector to increase protective antibody titers.
A mucosal boost with NDV-HXP-S in mice after high-dose mRNA vaccinations improves cellular and mucosal immunity
To assess the potential of IN NDV-HXP-S booster after IM vaccinations, mimicking the real-world vaccination scenario, K18-hACE2 mice were vaccinated twice with 5 μg of mRNA-lipid nanoparticles (LNPs, Pfizer/BioNTech BNT162b2) 21 days apart (Fig. 6a). Over a year later, the mice were boosted with 106 EID50 of NDV-HXP-S expressing ancestral or BA.1 spike. An IM mRNA-LNP vaccinated group received PBS IN as a mock booster and was used as no booster control. An additional group received PBS throughout the whole vaccination schedule representing the naïve control group. One week after boosting, spleens were collected for memory B cell analysis. Four weeks post-boost, serum, nasal washes, vaginal lavages, intestinal lavages, spleens, and lungs were collected (Fig. 6a).
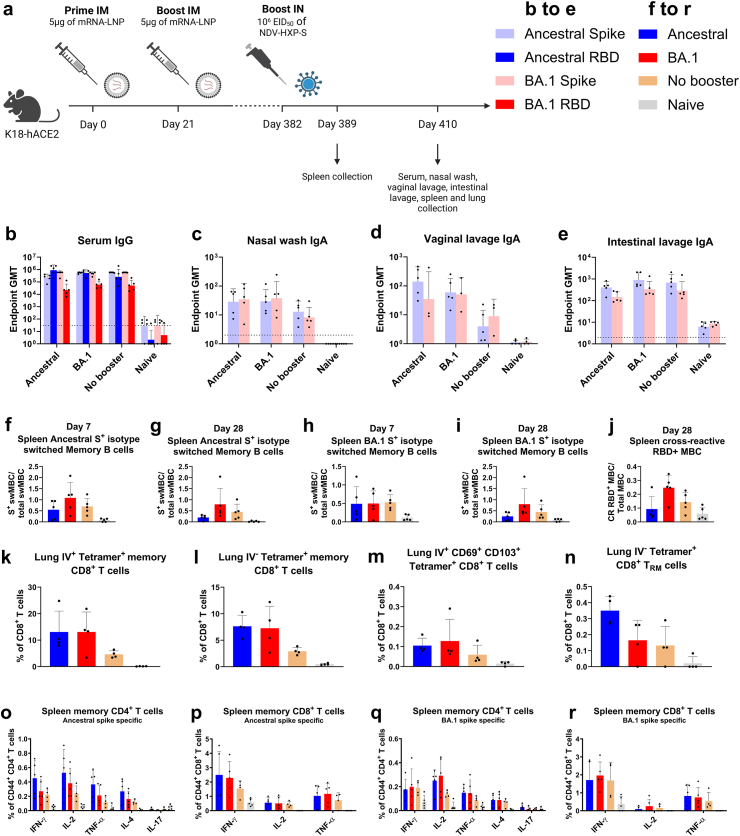
Fig. 6.
A mucosal boosting with NDV-HXP-S in mice with ceiling serum antibodies induced by high-dose mRNA vaccinations improves cellular and mucosal immunity. (a) Design of the study. Six to eight week-old female K18-hACE2 mice were intramuscularly immunised twice 21 days apart with 5 μg of mRNA-lipid nanoparticles (LNPs, Pfizer/BioNTech BNT162b2). After 382 days, the mice were boosted IN with 106 EID50 of NDV-HXP-S ancestral or BA.1. A no booster group received PBS. A naïve control group received PBS throughout the whole vaccination schedule. After 389 days (seven days after boosting), spleens were collected. After 410 days (28 days after boosting), serum, nasal washes, vaginal lavages, and intestinal lavages were collected to assess binding antibody response. Spleens were collected to analyse memory B and T cells. Lungs were collected to analyse CD8+ T cells. (n = 5 or 4) (b) Measurement of ancestral spike, ancestral RBD, BA.1 spike and BA.1 RBD-specific serum IgG harvested 28 days after the booster. (c–e) Measurement of ancestral spike and BA.1 spike-specific (c) nasal wash IgA, (d) vaginal lavage IgA and (e) intestinal lavage IgA harvested 28 days after boosting mice with 106 EID50 of NDV-HXP-S ancestral or BA.1 or PBS (no booster and naïve control groups). The GMT endpoint titers of the ELISA assay were graphed (an endpoint titer of one was assigned to negative samples). The dashed line indicates the limit of detection (for vaginal lavage x-axis equals limit of detection). The error bars represent geometric SD. (f–j) Using two separate tetrameric B cell probes for ancestral or BA.1 spike respectively as well as an ancestral RBD probe, B cell subsets in the spleens were measured. This included (f) isotype switched memory B cells specific for ancestral spike seven days after boosting, (g) isotype switched memory B cells specific for ancestral spike 28 days after boosting, (h) isotype switched memory B cells specific for BA.1 spike seven days after boosting, (i) isotype switched memory B cells specific for BA.1 spike 28 days after boosting (CD3- Live+ B220+ CD19+ IgM− IgD− GL7- CD38+) and (j) cross-reactive memory B cells (gated ancestral RBD vs BA.1 spike, CD3- Live+ B220+ CD19+ IgD− GL7- CD38+). Mean with SD was graphed. (k–n) Using CD45 intravascular (IV) labelling as well as tetramer staining, intravascular (IV+) and extravascular (IV−) CD8+ T cells specific for SARS-CoV-2 spike were measured in the lungs 28 days after boosting. This included (k) IV+ memory CD8+ T cells, (l) IV− memory CD8+ T cells (CD3+ MHC II− CD44+), (m) IV+ CD69+ CD103+ CD8+ T cells (CD3+ MHC II− CD44+ CD69+ CD103+), and (n) IV− CD8+ TRM cells (CD3+ MHC II− CD44+ CD69+ CD103+). Mean with SD was graphed. (o–r) Twenty-eight days after boosting, splenocytes were isolated and restimulated with an ancestral or BA.1 spike specific peptide pool. Intracellular cytokine staining was used to measure antigen-specific production of IFN-γ, TNF-α, IL-2, IL-4, and IL-17 by CD44+ CD4+ or CD44+ CD8+ T cells out of total CD44+ CD4+ or CD44+ CD8+ T cells. Mean with SD was graphed. Statistical significance was calculated by means of [(b)–(e)] log transforming the data set to normalise followed by a two-way ANOVA, or [(f)–(n)] one-way ANOVA, or [(o)–(r)] two-way ANOVA followed by Tukey’s multiple comparisons test.
The strong systemic immunogenicity of mRNA-LNPs and the high dosage used led to high serum IgG titers that were not significantly boosted by IN administration of NDV-HXP-S (Fig. 6b). However, even though serum antibody levels apparently reached a ceiling, IN boosting with NDV-HXP-S increased IgA levels in nasal washes and vaginal lavages, when comparing the endpoint titers of the no booster group to ancestral and BA.1 groups (Fig. 6c and d). Interestingly, there was no significant difference detected between intestinal lavage groups as the animals of the no booster control group still reached high intestinal lavage IgA titers comparable to booster groups (Fig. 6e). These high levels of IgA found in the intestinal lavages could be due to the close proximity of the intestines to the injection sites (hind limb) in combination with a high enough vaccination dosage enabling antigen to circulate to multiple lymph nodes. The high-dose mRNA vaccines may have provoked a modest IgA-secreting B cell responses within gut-associated lymphoid tissues.
Spleens were harvested on day 7 and day 28 after the booster and antigen-specific memory B cells were subsequently analysed using flow cytometry with tetrameric B cell probes (Supplementary Fig. S2). Unexpectedly, the NDV-HXP-S ancestral booster led to fewer isotype switched memory B cells specific for the ancestral spike than no booster (Fig. 6f and g). While the ancestral spike-specific memory B cells seemed to be depleted in the ancestral booster group, the memory B cells were increased in the BA.1 booster group. The frequency of isotype switched memory B cells specific for BA.1 spike was similar across all vaccination groups 7 days after the booster (Fig. 6h). However, three weeks later the amount of memory B cells had increased for the BA.1 booster group, but stayed similar for the ancestral and no booster groups (Fig. 6i). These data suggest that the antibody responses observed after boosting stem from a memory B cell recall response, not naïve B cell development. Recent publications have observed a similar phenomenon in humans after boosting with mRNA Omicron vaccines.62 Furthermore, an increase in cross-reactive RBD binding memory B cells was detected 28 days after boosting with NDV-HXP-S BA.1 (Fig. 6j). These changes of the memory B cell pool induced by the BA.1 booster could lead to better protection against BA.1 upon future exposure.
In order to assess the effect of the IN NDV-HXP-S booster on eliciting spike-specific circulating and extravascular CD8+ T cells, intravascular (IV) labelling with an anti-CD45 antibody was performed.65 To identify antigen-specific CD8+ T cells, the harvested lung cells were labelled with major histocompatibility complex (MHC) class I tetramers loaded with spike epitope VNFNFNGL (Supplementary Fig. S3), which is conserved between the ancestral, the BA.1 and recent strains like XBB.1.5. Four weeks after boosting, the ancestral and the BA.1 booster group displayed an increase in IV+ memory CD8+ T cells compared to the no booster group (Fig. 6k). This population of memory CD8+ T cells recognizing the SARS-CoV-2 spike is in the pulmonary vasculature (IV+). Furthermore, both NDV-HXP-S boosters expanded the pool of spike-specific memory CD8+ T cells in the parenchyma (Fig. 6l), identifiable by their lack of IV CD45 labelling (IV−). This pool consists mostly of peripheral tissue-circulating effector memory T cells (TEM).66,67 Additionally, a slight increase in the population of vasculature-associated or circulating (IV+) CD69+ CD103+ CD8+ T cells was detected (Fig. 6m). These cells are not bona fide CD8+ TRM cells (IV− CD69+ CD103+ CD8+ T) but precursors in vasculature prior to moving to their dedicated niche.66 Finally, we saw a boost in TRM cells 4 weeks after IN administration of NDV-HXP-S expressing ancestral spike (Fig. 6n). It appears that priming with the same antigen (ancestral spike via IM mRNA-LNPs) strengthens the lung TRM recruitment by the IN NDV-HXP-S booster.22,68,69 Overall, as the majority of antigen-specific CD8+ T cells are IV+, it is also in these populations where we observed the biggest improvements after boosting.65,70
To further characterise the systemic memory T cell response, intracellular cytokine staining was performed on harvested splenocytes after overnight ex vivo stimulation with an ancestral or BA.1 spike peptide pool (Supplementary Fig. S4). Based on an increase in the frequency of IFN-γ+, TNF-α+, and IL-2+ T cells, we found that IN administration of NDV-HXP-S led to a higher number of ancestral spike-specific CD4+ T cells phenotypically resembling T helper type 1 cells (TH1) four weeks after boosting (Fig. 6o and q). Only a slight increase in TH2 (IL-4) and no increase in TH17 (IL-17) CD4+ T cells was observed. Furthermore, the NDV-HXP-S booster induced an expansion of cytotoxic T lymphocyte-like CD8+ T cell populations (Fig. 6p and r).
Thus, in an experimental setup where the serum antibodies have already reached a ceiling after primary IM mRNA-LNP vaccination series, boosting IN with NDV-HXP-S can still improve other arms of immune responses. Specifically, mucosal IgA and spike-specific CD8+ memory T cells are induced in the lungs and the systemic T cell response is improved as well. Finally, boosting with NDV-HXP-S BA.1 has the potential to reshape the memory B cell pool for better Omicron coverage.
A mucosal boost with NDV-HXP-S in mice after low-dose mRNA vaccinations improves multiple arms of adaptive immunity
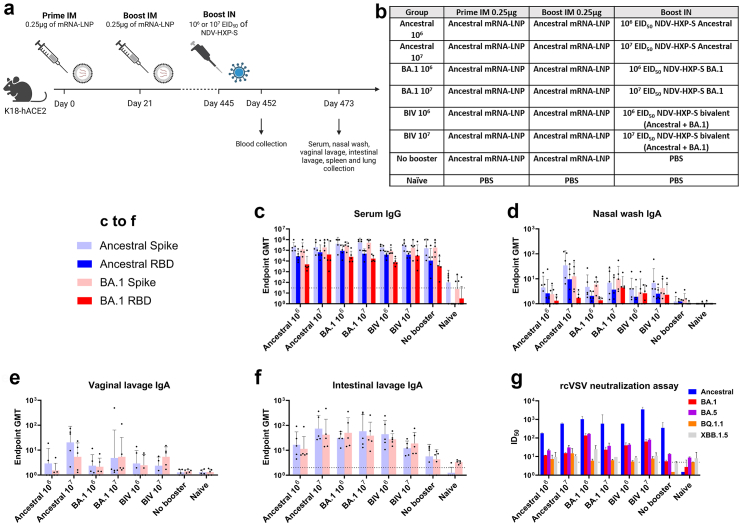
In order to further explore the antibody changes induced by IN NDV-HXP-S booster after IM mRNA vaccinations, we used K18-hACE2 mice that received lower dose (0.25 μg) of mRNA-LNPs (Pfizer/BioNTech BNT162b2) than those in the previous study (Fig. 7a vs Fig. 6a). After over 14 months, mice were boosted IN with 106 or 107 EID50 of NDV-HXP-S. The mice received either an ancestral, BA.1 or bivalent (BIV, equal amounts of ancestral and BA.1 spike expressing NDV-HXP-S to a total of the stated EID50) formulation (Fig. 7b). For animals of the no booster control group, PBS was used instead of NDV-HXP-S. The sex- and age-matched naïve control group received PBS throughout the whole vaccination regimen. One week after IN boosting, the mice were bled via submandibular vein (Fig. 7a). Four weeks after the NDV-HXP-S administration, serum, nasal washes, vaginal lavages, intestinal lavages, spleens, and lungs were collected and analysed.
Fig. 7.
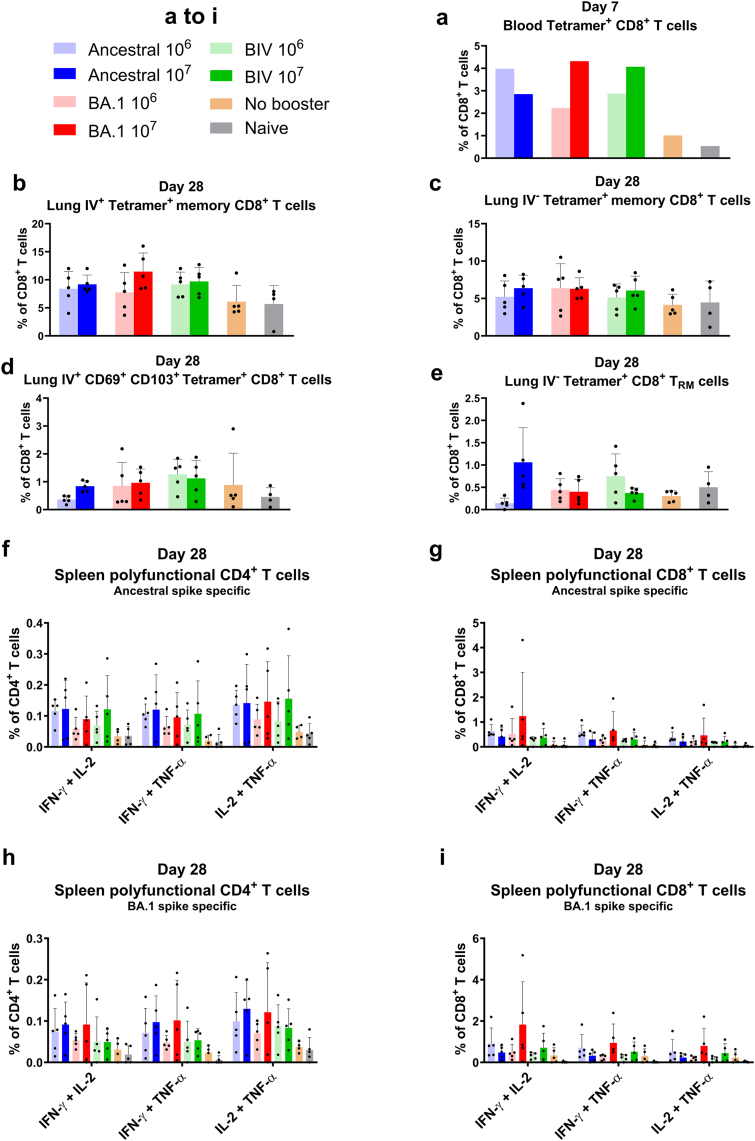
A mucosal boost with NDV-HXP-S in mice after low-dose mRNA vaccinations improves serum antibody binding and neutralization titers and induces mucosal IgA responses. (a and b) Design of the study and groups. Six to eight week-old female K18-hACE2 mice were intramuscularly immunised twice 21 days apart with 0.25 μg of mRNA-LNPs (Pfizer/BioNTech BNT162b2). After 445 days, the mice were boosted IN with 106 or 107 EID50 of NDV-HXP-S ancestral, BA.1 or bivalent (BIV, equal amounts of ancestral and BA.1 to a total of 106 or 107 EID50). A no booster group received PBS. A naïve control group received PBS throughout the whole vaccination schedule. After 452 days (seven days after boosting), serum was collected to assess blood circulating CD8+ T cells. After 473 days (28 days after boosting) serum, nasal washes, vaginal lavages, and intestinal lavages were collected to assess binding and neutralizing antibody response. Spleens were collected to analyse T cells. Lungs were collected to analyse CD8+ T cells. (n = 5) (c and d) Measurement of ancestral spike, ancestral RBD, BA.1 spike, and BA.1 RBD-specific (b) serum IgG and (c) nasal wash IgA. (e and f) Measurement of ancestral spike and BA.1 spike-specific (d) vaginal lavage IgA and (e) intestinal lavage IgA harvested 28 days after boosting mice with 106 or 107 EID50 of NDV-HXP-S ancestral, BA.1 or BIV or PBS (no booster and naïve control groups). The GMT endpoint titers of the ELISA assay were graphed (an endpoint titer of one was assigned to negative samples). The dashed line indicates the limit of detection (for nasal wash and vaginal lavage x-axis equals limit of detection). The error bars represent geometric SD. (g) Neutralization titers against SARS-CoV-2 ancestral, BA.1, BA.5, BQ.1.1, or XBB.1.5 spike-pseudotyped vesicular stomatitis virus expressing GFP (rcVSVeGFP-CoV-2-S) were measured in technical duplicates from pooled sera. Geometric mean with geometric SD of the ID50 was graphed (ID50 of 1.5 was assigned to negative samples). The dashed line indicates the limit of detection. Statistical significance was calculated by means of [(c)–(f)] log transforming the data set to normalise followed by a two-way ANOVA followed by Tukey’s multiple comparisons test.
Four weeks after the booster, the serum IgG levels against ancestral and BA.1 spike were still very high in all vaccination groups with endpoint titers of 105 or higher (Fig. 7c). Therefore, only slight improvements in spike binding titers were observed after high dosage (EID50 of 107) IN boosting. Interestingly, we detected an improvement in ancestral and BA.1 RBD binding titers upon boosting with every NDV-HXP-S formulation (Fig. 7c). It has been reported before that NDV-HXP-S vaccination elicits an RBD-focused antibody response and this seems also to be the case for utilizing NDV-HXP-S as a booster.37 No IgA against spike or RBD was detected in nasal washes of animals not receiving an IN booster as opposed to the booster groups which all had good IgA titers (Fig. 7d). Animals receiving higher dosages of NDV-HXP-S also developed higher nasal wash IgA levels. Similarly, only animals receiving NDV-HXP-S IN produced ancestral and BA.1 spike binding IgA in their genitourinary tract (Fig. 7e). While some spike binding IgA was found in the intestinal lavages of animals only receiving IM mRNA-LNP vaccination, all IN boosters improved intestinal lavage IgA titers (Fig. 7f). Finally, the previously described improvements in serum RBD-binding IgG levels translated into an increase of pseudotyped virus neutralization titers (Fig. 7g). As expected from animals only immunised with the ancestral spike, the no booster group neutralised ancestral pseudotyped virus well, but failed at efficiently neutralizing the Omicron variants (Fig. 7g). In comparison, all the NDV-HXP-S formulations improved Omicron neutralization titers (Fig. 7g). Nonetheless, the boosters containing the NDV-HXP-S BA.1 component had the biggest impact, inducing neutralization titers against BA.1 as well as BA.5 pseudotyped virus. This suggests the induction or boosting of cross-reactive antibodies, some of which are neutralizing. This is consistent with our findings for memory B cells, where we observed that a mucosal monovalent BA.1 booster can increase BA.1-specific and RBD-targeting cross-reactive memory B cells (Fig. 6i and j). However, variants known to be substantially antigenically distant, like BQ.1.1 and XBB.1.5, were not efficiently neutralised. The mechanism of such observations needs further investigation. Overall, the low dosage monovalent NDV-HXP-S BA.1 booster induced the highest level of BA.1 neutralization antibodies, while the high dosage bivalent booster showed a compelling capacity to improve both ancestral as well as BA.1 neutralization titers (Fig. 7g).
Regarding cellular immunity, mice were bled 7 days after the booster and the blood analysed for antigen-specific CD8+ T cells with MHC I tetramer staining (Fig. 8a). All formulations of the NDV-HXP-S IN booster induced an increase in CD8+ T cells circulating in the blood compared to the no booster and naïve controls.
Fig. 8.
A mucosal boost with NDV-HXP-S in mice after low-dose mRNA vaccinations improves T cell responses in blood, lungs and spleens. (a) Using tetramer staining, CD8+ T cells specific for SARS-CoV-2 spike were measured in the blood seven days after boosting (CD3+ MHC II−). Blood of individual animals was pooled for each group. Mean was graphed. (b–e) Using CD45 IV labelling as well as tetramer staining, intravascular (IV+) and extravascular (IV−) CD8+ T cells specific for SARS-CoV-2 spike were measured in the lungs 28 days after boosting (n = 5 or 4). This included (b) IV+ memory CD8+ T cells, (c) IV− memory CD8+ T cells (CD3+ MHC II− CD44+), (d) IV+ CD69+ CD103+ CD8+ T cells (CD3+ MHC II− CD44+ CD69+ CD103+), and (e) IV− CD8+ TRM cells (CD3+ MHC II− CD44+ CD69+ CD103+). Mean with SD was graphed. (f–i) Twenty-eight days after boosting, splenocytes were isolated and restimulated with an ancestral or BA.1 spike specific peptide pool (n = 5 or 4). Intracellular cytokine staining was used to measure antigen-specific production of IFN-γ, TNF-α, and IL-2 by CD4+ or CD8+ T cells out of total CD4+ or CD8+ T cells. Polyfunctional T cell populations positive for two markers are shown. Mean with SD was graphed. Statistical significance was calculated by means of [(b)–(e)] one-way ANOVA or [(f)–(i)] two-way ANOVA followed by Tukey’s multiple comparisons test.
Four weeks after IN administration of NDV-HXP-S, the lungs were examined for antigen-specific CD8+ T cells in lung vasculature and parenchyma (Fig. 8b–e). In line with our previous finding of increased CD8+ T cells in the blood seven days after boosting, we also detected a rise in the number of circulating memory CD8+ T cells in the lungs 28 days after boosting (Fig. 8b). Furthermore, we observed a slight trend of an increase of IV− memory CD8+ T cells binding the antigen tetramer after IN NDV-HXP-S administration (Fig. 8c). However, no improvement in the pool of IV+ CD69+ CD103+ CD8+ T cells was detected (Fig. 8d). Interestingly, only the high dosage ancestral booster was able to increase the frequency of CD8+ TRM cells in the lungs (Fig. 8e). As discussed previously, priming with the same antigen might be needed for efficient TRM induction.22,68,69
Polyfunctional T cells have been shown to be important parameters of immune protection due to their increased cytokine production efficiency.71,72 Therefore, we analysed the harvested splenocytes by intracellular cytokine staining after peptide ex vivo stimulation (Fig. 8f–I; Supplementary Fig. S5). We found that IN boosting with NDV-HXP-S elicited a TH1 like CD4+ polyfunctional T cell response with production of IFN-γ, TNF-α, and IL-2 by spike-specific cells (Fig. 8f and h). A higher dosage of NDV-HXP-S led to more polyfunctional CD4+ T cells compared to the lower dosage of the same formulation (Fig. 8f and h). The bivalent NDV-HXP-S booster induced similar amounts of polyfunctional CD4+ T cells specific for the ancestral spike as the ancestral booster (Fig. 8f). However, the bivalent booster did not have the same impact on BA.1 spike-specific T cells, with the BA.1 as well as the ancestral formulation being more effective at increasing the numbers of BA.1 spike-specific T cells (Fig. 8h). The BA.1 booster led to the biggest surge in polyfunctional T cells specific for the BA.1 spike. None of the boosters induced big increases of CD8+ polyfunctional T cells compared to the no booster group (Fig. 8g and i).
Thus, boosting mice with IN NDV-HXP-S after IM vaccinations improved systemic humoral immunity as displayed by increased serum neutralization titers. Furthermore, mucosal immunity was induced in the upper respiratory tract, the gut, and the genitourinary tract. The booster also increased CD8+ T cell levels in the blood and lungs as well as polyfunctional T cell numbers in the spleens.
Discussion
In this study, we constructed variant vaccines of the Omicron BA.1 and BA.5 strains utilizing the NDV vaccine platform and evaluated intranasal administration in mice. Vaccination with one or two doses of NDV-HXP-S elicited robust systemic as well as mucosal humoral immunity. The variant vaccines induced systemic neutralizing antibody responses against BA.1 and BA.5 and protected against variant matched challenge. Additionally, isotype switched memory B cells specific for the spike antigen were found in the spleens. Furthermore, we observed high IgA titers in nasal washes, vaginal, and intestinal lavages. The neutralizing activity of mucosal antibodies was not measured and is a limitation of this study. While vaccination occurred in the upper respiratory tract, immunity was induced at all mucosal sites tested. This observation suggests that IN NDV-HXP-S vaccination promotes homing of IgA memory B cells to a variety of different mucosal sites.73 This could be leveraged in the future for targeting other pathogens that infect via the mucosal route like human papillomavirus (HPV), human immunodeficiency virus (HIV) or poliovirus.
Recent preclinical studies have shown that IN SARS-CoV-2 vaccines not only prevent disease but also reduce viral shedding and transmission.14, 15, 16, 17, 18, 19, 20, 21, 22 The majority of currently utilised and highly efficacious COVID-19 vaccines is based on mRNA technology, administered IM.74 Therefore, developing an IN booster for administration after IM mRNA vaccinations, providing all the advantages that come with strong mucosal immunity, is key. We show that NDV-HXP-S used as an IN booster has great efficacy at inducing robust mucosal immunity after IM mRNA-LNP vaccination. Even in mice of old age, IgA responses were detected in the upper respiratory tract, the gut, and the genitourinary tract after boosting IN with NDV-HXP-S. This could have major implications in future booster administrations of elderly individuals, who do not generate as robust of an immune response to IM vaccinations or boosters, are at high-risk of severe disease following respiratory viral infections, and could greatly benefit from elicitation of local, protective immunity in the respiratory tract. Furthermore, CD8+ T cell populations increased in the lungs of mice boosted IN with NDV-HXP-S. While the majority of induced CD8+ T cells in the lungs were intravascular, a substantial increase in extravascular antigen-specific TRM cells was achieved upon boosting with NDV-HXP-S expressing ancestral spike. We speculate that circulating CD8+ T cells induced by the IM priming expand during the recall response triggered by the booster. Subsequently, these circulating memory CD8+ T cells are recruited to the lungs. There, they may differentiate into TRM cells. It appears that an antigen similar or identical to the antigen used for priming is necessary in order to guide pre-existing CD8+ T cell populations to the site of boosting and potentially amplify them there. A similar prime-pull effect has been reported in the female genital tract of mice.68 Taken together, this provides a rationale to implement IN boosters into current SARS-CoV-2 vaccination schedules, and potentially for other respiratory pathogens.
Many of the current clinical trials of mucosal vaccines utilise live-attenuated or replication-deficient viral vectors. It has been established that vector immunity can lead to reduced immunogenicity, severely inhibiting the potential of viral vector vaccines.75, 76, 77 We demonstrated, that while immunity against the NDV vector is induced, it does not prevent further boosting of neutralizing anti-spike and RBD antibody titers. NDV-antigen-specific cellular immunity was not explored and represents a limitation of this study that we plan to address in future work.
The WHO Technical Advisory Group on COVID-19 Vaccine Composition (TAG–CO–VAC) has published their recommendations on the antigen composition of future COVID-19 vaccines, suggesting the use of a monovalent XBB.1 descendent lineage as the vaccine antigen.78 This recommendation was supported by The European Medicines Agency (EMA), arguing that a monovalent vaccine against the XBB.1 lineage is reasonably effective at protecting against circulating strains and possibly further variants.79 The FDA’s Vaccines and Related Biological Products Advisory Committee (VRBPAC) argued similarly, voting unanimously for a monovalent XBB-lineage based COVID-19 vaccine, preferably XBB.1.5.80 As a proof of principle, we demonstrated boosting with monovalent IN NDV-HXP-S BA.1 after priming with ancestral antigen using mRNA-LNPs. We measured an increase in neutralizing serum IgG titers against BA.1 and BA.5. Interestingly, when repeatedly vaccinating with the same vaccine via the same route (IN NDV-HXP-S) we saw antigenic imprinting/sin. In contrast, upon heterologous boosting (IM mRNA LNP followed by IN NDV-HXP-S) we observed a prominent Omicron specific neutralizing antibody response. Furthermore, mucosal IgA against BA.1 was detected. In accordance with the antibody data, a de novo B cell response against BA.1 epitopes as well as an increase in cross-reactive memory B cells was induced. Additionally, the NDV-HXP-S BA.1 booster elicited a polyfunctional CD4+ TH1 response against BA.1 spike. These changes of the humoral and cellular immunity towards a BA.1 specific response after IN boosting with NDV-HXP-S BA.1 could lead to better protection against BA.1 and could be translated to other variant antigens. These results support the WHO’s, EMA’s and VRBPAC’s decision to use monovalent boosters moving forward, especially considering the modest benefits reported for the bivalent BA.5 booster.61,62,81,82
Here, we demonstrate the broad versatility of NDV-HXP-S variant vaccines by utilizing them for primary immunizations as well as boosting. The efficient induction of systemic as well as mucosal immunity prevented infection and shows promise to reduce transmission. Future studies will be performed in hamster models to test the ability of IN NDV-HXP-S variant vaccines to interrupt the transmission of the virus as well as its effectiveness against a heterologous challenge. Clinical trials using IN administered NDV-HXP-S will illuminate how these pre-clinical findings compare to human data (United States: phase I NCT05181709; Mexico: phase II, NCT05205746).
Racing against viral evolution has rarely been a winning strategy, so in order to mitigate the impact of future SARS-CoV-2 variants and other respiratory viruses, new approaches will be needed. Vaccines and boosters able to induce mucosal immunity may help to protect individuals against breakthrough infections and reduce transmission of the virus in the population.
Contributors
Conceptualization, S.S., P.P. and W.S.; Data curation, S.S., I.G.D. and W.S.; Formal analysis, S.S., I.G.D., L.A.C., M.N. and W.S.; Funding acquisition, M.S., A.G.S, P.P.; Investigation, S.S., I.G.D., L.A.C., N.L., T.Y.L., J.L.M., V.D., A.A., M.N., W.S.; Methodology, S.S., I.G.D., L.A.C., S.K., W.S.; Resources, S.K., G.S., P.W., G.S., A.G.S, B.L., F.K., M.S., P.P., W.S.; Supervision, M.S., P.P, W.S.; Validation, P.P, W.S.; Visualization, S.S; Writing – original draft, S.S., P.P, W.S.; Writing – review & editing, all authors. All authors read and approved the final version of the manuscript.
Data sharing statement
All data are available in the main text or Supplementary materials. Raw data will be shared and distributed for research purposes upon request to the corresponding author.
Declaration of interests
The Icahn School of Medicine at Mount Sinai has filed patent applications entitled “RECOMBINANT NEWCASTLE DISEASE VIRUS EXPRESSING SARS-COV-2 SPIKE PROTEIN AND USES THEREOF” which names P.P., A.G.S, F.K. and W.S. as inventors. Mount Sinai is seeking to commercialise this vaccine; therefore, the institution and its faculty inventors could benefit financially.
I.G.D. has co-chaired at the ninth ESWI Influenza conference, which has no competing interest with this work.
The M.S. laboratory has received unrelated research funding in sponsored research agreements from 7Hills Pharma, ArgenX N.V., Moderna and Phio Pharmaceuticals, which has no competing interest with this work.
F.K. has consulted for Merck, Seqirus, Curevac and Pfizer, and is currently consulting for Pfizer, Third Rock Ventures, GSK and Avimex. The FK laboratory is also collaborating with Pfizer on animal models of SARS-CoV-2.
The A.G.-S. laboratory has received research support from GSK, Pfizer, Senhwa Biosciences, Kenall Manufacturing, Blade Therapeutics, Avimex, Johnson & Johnson, Dynavax, 7Hills Pharma, Pharmamar, ImmunityBio, Accurius, Nanocomposix, Hexamer, N-fold LLC, Model Medicines, Atea Pharma, Applied Biological Laboratories and Merck. A.G.S. has consulting agreements for the following companies involving cash and/or stock: Castlevax, Amovir, Vivaldi Biosciences, Contrafect, 7Hills Pharma, Avimex, Pagoda, Accurius, Esperovax, Farmak, Applied Biological Laboratories, Pharmamar, CureLab Oncology, CureLab Veterinary, Synairgen, Paratus and Pfizer. A.G.S. has been an invited speaker in meeting events organised by Seqirus, Janssen, Abbott and Astrazeneca. A.G.S. is inventor on patents and patent applications on the use of antivirals and vaccines for the treatment and prevention of virus infections and cancer, owned by the Icahn School of Medicine at Mount Sinai, New York. Specifically, A.G.S., a member of the faculty of the Icahn School of Medicine at Mount Sinai (Mount Sinai) is an inventor of a novel COVID-19 vaccine currently being investigated in clinical trials. Mount Sinai is advancing the development of this vaccine and related technologies for potential commercial use. Mount Sinai has created CastleVax Inc., a Mount Sinai company, and has licensed the applicable IP to it. Mount Sinai will receive financial compensation from CastleVax Inc. pursuant to that license if vaccine development proceeds and as an owner of the company subject to the sale of its ownership interest in the future. Subject to Mount Sinai receiving such financial consideration, A.G.S. will receive a portion of that consideration pursuant to the terms of the Mount Sinai Intellectual Property Policy.
All other authors declare no competing interests.
Acknowledgements
All figures created with BioRender.com and/or GraphPad.
We appreciate the work of Randy Albrecht, Carlos Franco and the CCMS team overseeing the CBSL-3 and ABSL-3 facilities at Mount Sinai. We would also like to thank the expertise and assistance of the Dean's Flow Cytometry CORE at Mount Sinai.
This work was partially funded by the NIAID Centers of Excellence for Influenza Research and Response (CEIRR) contract 75N93021C00014 (to P.P. and A.G.S.) and by the NIAID Collaborative Vaccine Innovation Centers contract 75N93019C00051 (to P.P.), by NIAID grant R01 AI145870 (to P.P.) and by institutional funding from the Icahn School of Medicine at Mount Sinai (to P.P.).
SARS-CoV-2 work in the Schotsaert lab is funded by NIH/NIAID R01AI160706, NIH/NIAID R21AI176069 and NIH/NIDDK R01DK130425 (to M.S.).
The Conventional Biocontainment Facility (CBF) is a NIHBSL3/BSL3 facility that is part of the BSL-3 Biocontainment CoRE. This core is supported by subsidies from the ISMMS Dean’s Office and by investigator support through a cost recovery mechanism. Research reported in this publication was supported by the National Institute of Allergy and Infectious Diseases of the National Institutes of Health under Award Number G20AI174733 (R.A. Albrecht). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Footnotes
Supplementary data related to this article can be found at https://doi.org/10.1016/j.ebiom.2024.105185.
Contributor Information
Peter Palese, Email: peter.palese@mssm.edu.
Weina Sun, Email: weina.sun@mssm.edu.
Appendix ASupplementary data
Supplementary Fig. S1.
Supplementary Figure S1. Serum passive transfer decreases viral load in the lungs. (a) Design of the study and groups. Sera from sequentially intranasally vaccinated BALB/c mice were pooled and administered intraperitoneally (IP) to six to eight week-old female K18-hACE2 mice. After two hours, the mice were challenged with 3 x 104 plaque-forming units (PFU) of SARS-CoV-2 BA.1 (n = 5). A healthy control group was kept uninfected (n = 5). (b and c) Measurement of viral titers in the (b) lungs and (c) nasal turbinates. Tissues of the animals were collected on day 5 post challenge. The whole lungs or nasal turbinates were homogenised in 1 mL of PBS. The viral load was measured by plaque assay on VERO E6 TMPRSS2 T2A ACE2 cells and graphed as GMT of PFU/ml (a titer of 25 PFU/ml was assigned to negative samples). The dashed line indicates the limit of detection. The error bars represent geometric SD. Statistical significance was calculated by means of Kruskal-Wallis test followed by Dunn’s multiple comparisons test.
Supplementary Fig. S2.
Supplementary Figure S2. Memory B cells gating strategy and S protein probe staining. (a) Representative gating strategy for the identification of memory B cells (MBC) and isotype switched memory B cells (swMBC) in the spleen. (b) Representative flow cytometry plots of ancestral S protein probe staining (PE and APC) for MBC and swMBC. (c) Representative flow cytometry plots of BA.1 S protein probe staining (PE and APC) for memory B cells (MBC) and isotype switched memory B cells (swMBC).
Supplementary Fig. S3.
Supplementary Figure S3. Gating strategy for S tetramer specific memory CD8+T cells in the lung. Representative gating strategy for the identification of lung IV+/- tetramer+ memory CD8+ T cells or TRM (CD69+ CD103+) cells. In the specific example shown, the IV+ population is further analysed, but the same gating strategy was applied for IV- populations.
Supplementary Fig. S4.
Supplementary Figure S4. Gating strategy for intracellular cytokine staining of T cells. Representative gating strategy of splenocytes for the identification of CD4+ CD44+ T cells producing IFN-γ, TNF-α, IL-2, IL-4, or IL-17 and CD8+ CD44+ T cells producing IFN-γ, TNF-α or IL-2 upon restimulation with an ancestral or BA.1 spike specific peptide pool. The specific examples shown are positive control samples.
Supplementary Fig. S5.
Supplementary Figure S5. A mucosal boost with NDV-HXP-S in mice after low-dose mRNA vaccinations increases the frequency of IFN-γ+CD8+T cells. (a–d) Twenty-eight days after boosting, splenocytes were isolated and restimulated with an ancestral or BA.1 spike specific peptide pool. Intracellular cytokine staining was used to measure antigen-specific production of IFN-γ, TNF-α, and IL-2 by CD44+ CD4+ or CD44+ CD8+ T cells out of total CD44+ CD4+ or CD44+ CD8+ T cells. Mean with SD was graphed. Statistical significance was calculated by means of two-way ANOVA followed by Tukey’s multiple comparisons test.
References
- 1.Carvalho T., Krammer F., Iwasaki A. The first 12 months of COVID-19: a timeline of immunological insights. Nat Rev Immunol. 2021;21:245–256. doi: 10.1038/s41577-021-00522-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Gupta R.K., Topol E.J. COVID-19 vaccine breakthrough infections. Science. 2021;374:1561–1562. doi: 10.1126/science.abl8487. [DOI] [PubMed] [Google Scholar]
- 3.Lucas C., Vogels C.B.F., Yildirim I., et al. Impact of circulating SARS-CoV-2 variants on mRNA vaccine-induced immunity. Nature. 2021;600:523–529. doi: 10.1038/s41586-021-04085-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tartof S.Y., Slezak J.M., Fischer H., et al. Effectiveness of mRNA BNT162b2 COVID-19 vaccine up to 6 months in a large integrated health system in the USA: a retrospective cohort study. Lancet. 2021;398:1407–1416. doi: 10.1016/S0140-6736(21)02183-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chalkias S., Harper C., Vrbicky K., et al. A bivalent omicron-containing booster vaccine against covid-19. N Engl J Med. 2022;387:1279–1291. doi: 10.1056/NEJMoa2208343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Chalkias S., Eder F., Essink B., et al. Safety, immunogenicity and antibody persistence of a bivalent Beta-containing booster vaccine against COVID-19: a phase 2/3 trial. Nat Med. 2022;28:2388–2397. doi: 10.1038/s41591-022-02031-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lee I.T., Cosgrove C.A., Moore P., et al. Omicron BA.1-containing mRNA-1273 boosters compared with the original COVID-19 vaccine in the UK: a randomised, observer-blind, active-controlled trial. Lancet Infect Dis. 2023;23:1007–1019. doi: 10.1016/S1473-3099(23)00295-5. [DOI] [PubMed] [Google Scholar]
- 8.Laczkó D., Hogan M.J., Toulmin S.A., et al. A single immunization with nucleoside-modified mRNA vaccines elicits strong cellular and humoral immune responses against SARS-CoV-2 in mice. Immunity. 2020;53:724–732.e7. doi: 10.1016/j.immuni.2020.07.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Goel R.R., Painter M.M., Apostolidis S.A., et al. mRNA vaccines induce durable immune memory to SARS-CoV-2 and variants of concern. Science. 2021;374 doi: 10.1126/science.abm0829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Turner J.S., O’Halloran J.A., Kalaidina E., et al. SARS-CoV-2 mRNA vaccines induce persistent human germinal centre responses. Nature. 2021;596:109–113. doi: 10.1038/s41586-021-03738-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sheikh-Mohamed S., Isho B., Chao G.Y.C., et al. Systemic and mucosal IgA responses are variably induced in response to SARS-CoV-2 mRNA vaccination and are associated with protection against subsequent infection. Mucosal Immunol. 2022;15:799–808. doi: 10.1038/s41385-022-00511-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Sano K., Bhavsar D., Singh G., et al. SARS-CoV-2 vaccination induces mucosal antibody responses in previously infected individuals. Nat Commun. 2022;13:5135. doi: 10.1038/s41467-022-32389-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tang J., Zeng C., Cox T.M., et al. Respiratory mucosal immunity against SARS-CoV-2 after mRNA vaccination. Sci Immunol. 2022;7 doi: 10.1126/sciimmunol.add4853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hassan A.O., Kafai N.M., Dmitriev I.P., et al. A single-dose intranasal Chad vaccine protects upper and lower respiratory tracts against SARS-CoV-2. Cell. 2020;183:169–184.e13. doi: 10.1016/j.cell.2020.08.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hassan A.O., Feldmann F., Zhao H., et al. A single intranasal dose of chimpanzee adenovirus-vectored vaccine protects against SARS-CoV-2 infection in rhesus macaques. Cell Reports Med. 2021;2 doi: 10.1016/j.xcrm.2021.100230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bricker T.L., Darling T.L., Hassan A.O., et al. A single intranasal or intramuscular immunization with chimpanzee adenovirus-vectored SARS-CoV-2 vaccine protects against pneumonia in hamsters. Cell Rep. 2021;36 doi: 10.1016/j.celrep.2021.109400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fischer R.J., van Doremalen N., Adney D.R., et al. ChAdOx1 nCoV-19 (AZD1222) protects Syrian hamsters against SARS-CoV-2 B.1.351 and B.1.1.7. Nat Commun. 2021;12:5868. doi: 10.1038/s41467-021-26178-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lu M., Dravid P., Zhang Y., et al. A safe and highly efficacious measles virus-based vaccine expressing SARS-CoV-2 stabilized prefusion spike. Proc Natl Acad Sci USA. 2021;118(12) doi: 10.1073/pnas.2026153118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen J., Wang P., Yuan L., et al. A live attenuated virus-based intranasal COVID-19 vaccine provides rapid, prolonged, and broad protection against SARS-CoV-2. Sci Bull. 2022;67(13):1372–1387. doi: 10.1016/j.scib.2022.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.An D., Li K., Rowe D.K., et al. Protection of K18-hACE2 mice and ferrets against SARS-CoV-2 challenge by a single-dose mucosal immunization with a parainfluenza virus 5-based COVID-19 vaccine. Sci Adv. 2021;7(27) doi: 10.1126/sciadv.abi5246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Warner B.M., Santry L.A., Leacy A., et al. Intranasal vaccination with a Newcastle disease virus-vectored vaccine protects hamsters from SARS-CoV-2 infection and disease. iScience. 2021;24(11) doi: 10.1016/j.isci.2021.103219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Mao T., Israelow B., Peña-Hernández M.A., et al. Unadjuvanted intranasal spike vaccine elicits protective mucosal immunity against sarbecoviruses. Science. 2022;378(6622) doi: 10.1126/science.abo2523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lavelle E.C., Ward R.W. Mucosal vaccines - fortifying the frontiers. Nat Rev Immunol. 2022;22:236–250. doi: 10.1038/s41577-021-00583-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Belshe R.B., Nichol K.L., Black S.B., et al. Safety, efficacy, and effectiveness of live, attenuated, cold-adapted influenza vaccine in an indicated population aged 5-49 years. Clin Infect Dis. 2004;39:920–927. doi: 10.1086/423001. [DOI] [PubMed] [Google Scholar]
- 25.Li J.X., Wu S.P., Guo X.L., et al. Safety and immunogenicity of heterologous boost immunisation with an orally administered aerosolised Ad5-nCoV after two-dose priming with an inactivated SARS-CoV-2 vaccine in Chinese adults: a randomised, open-label, single-centre trial. Lancet Respir Med. 2022;10(8):739–748. doi: 10.1016/S2213-2600(22)00087-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li J.X., Hou L.H., Gou J.B., et al. Safety, immunogenicity and protection of heterologous boost with an aerosolised Ad5-nCoV after two-dose inactivated COVID-19 vaccines in adults: a multicentre, open-label phase 3 trial. Lancet Infect Dis. 2023;23(10):1143–1152. doi: 10.1016/S1473-3099(23)00350-X. [DOI] [PubMed] [Google Scholar]
- 27.Zhu F., Huang S., Liu X., et al. Safety and efficacy of the intranasal spray SARS-CoV-2 vaccine dNS1-RBD: a multicentre, randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Respir Med. 2023;11(12):1075–1088. doi: 10.1016/S2213-2600(23)00349-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sunagar R., Prasad S.D., Ella R., Vadrevu K.M. Preclinical evaluation of safety and immunogenicity of a primary series intranasal COVID-19 vaccine candidate (BBV154) and humoral immunogenicity evaluation of a heterologous prime-boost strategy with COVAXIN (BBV152) Front Immunol. 2022;13 doi: 10.3389/fimmu.2022.1063679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ying B., Darling T.L., Desai P., et al. Mucosal vaccine-induced cross-reactive CD8+ T cells protect against SARS-CoV-2 XBB.1.5 respiratory tract infection. Nat Immunol. 2024;25(3):537–551. doi: 10.1038/s41590-024-01743-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sun W., Leist S.R., McCroskery S., et al. Newcastle disease virus (NDV) expressing the spike protein of SARS-CoV-2 as a live virus vaccine candidate. EBioMedicine. 2020;62 doi: 10.1016/j.ebiom.2020.103132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sun W., McCroskery S., Liu W.-C., et al. A Newcastle disease virus (NDV) expressing a membrane-anchored spike as a cost-effective inactivated SARS-CoV-2 vaccine. Vaccines. 2020;8:771. doi: 10.3390/vaccines8040771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sun W., Liu Y., Amanat F., et al. A Newcastle disease virus expressing a stabilized spike protein of SARS-CoV-2 induces protective immune responses. Nat Commun. 2021;12:6197. doi: 10.1038/s41467-021-26499-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.González-Domínguez I., Martínez J.L., Slamanig S., et al. Trivalent NDV-HXP-S vaccine protects against phylogenetically distant SARS-CoV-2 variants of concern in mice. Microbiol Spectr. 2022;10 doi: 10.1128/spectrum.01538-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pitisuttithum P., Luvira V., Lawpoolsri S., et al. Safety and immunogenicity of an inactivated recombinant Newcastle disease virus vaccine expressing SARS-CoV-2 spike: interim results of a randomised, placebo-controlled, phase 1 trial. EClinicalMedicine. 2022;45 doi: 10.1016/j.eclinm.2022.101323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Duc Dang A., Dinh Vu T., Hai Vu H., et al. Safety and immunogenicity of an egg-based inactivated Newcastle disease virus vaccine expressing SARS-CoV-2 spike: interim results of a randomized, placebo-controlled, phase 1/2 trial in Vietnam. Vaccine. 2022;40:3621–3632. doi: 10.1016/j.vaccine.2022.04.078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ponce-de-León S., Torres M., Soto-Ramírez L.E., et al. Interim safety and immunogenicity results from an NDV-based COVID-19 vaccine phase I trial in Mexico. NPJ Vaccines. 2023;8(1):67. doi: 10.1038/s41541-023-00662-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Carreño J.M., Raskin A., Singh G., et al. An inactivated NDV-HXP-S COVID-19 vaccine elicits a higher proportion of neutralizing antibodies in humans than mRNA vaccination. Sci Transl Med. 2023;15 doi: 10.1126/scitranslmed.abo2847. [DOI] [PubMed] [Google Scholar]
- 38.Beaty S.M., Park A., Won S.T., et al. Efficient and robust paramyxoviridae reverse genetics Systems. mSphere. 2017;2 doi: 10.1128/mSphere.00376-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Buchholz U.J., Finke S., Conzelmann K.K. Generation of bovine respiratory syncytial virus (BRSV) from cDNA: BRSV NS2 is not essential for virus replication in tissue culture, and the human RSV leader region acts as a functional BRSV genome promoter. J Virol. 1999;73:251–259. doi: 10.1128/JVI.73.1.251-259.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hsieh C., Goldsmith J.A., Schaub J.M., et al. Structure-based design of prefusion-stabilized SARS-CoV-2 spikes. Science. 2020;369:1501–1505. doi: 10.1126/science.abd0826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ayllon J., García-Sastre A., Martínez-Sobrido L. Rescue of recombinant Newcastle disease virus from cDNA. J Vis Exp. 2013;(80):50830. doi: 10.3791/50830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Ikegame S., Siddiquey M.N.A., Hung C.-T., et al. Neutralizing activity of Sputnik V vaccine sera against SARS-CoV-2 variants. Nat Commun. 2021;12:4598. doi: 10.1038/s41467-021-24909-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Yardeni T., Eckhaus M., Morris H.D., Huizing M., Hoogstraten-Miller S. Retro-orbital injections in mice. Lab Anim (NY) 2011;40:155–160. doi: 10.1038/laban0511-155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lin S., Chen Z., Zhang X., et al. Characterization of SARS-CoV-2 Omicron spike RBD reveals significantly decreased stability, severe evasion of neutralizing-antibody recognition but unaffected engagement by decoy ACE2 modified for enhanced RBD binding. Signal Transduct Target Ther. 2022;7:56. doi: 10.1038/s41392-022-00914-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Motozono C., Toyoda M., Zahradnik J., et al. SARS-CoV-2 spike L452R variant evades cellular immunity and increases infectivity. Cell Host Microbe. 2021;29:1124–1136.e11. doi: 10.1016/j.chom.2021.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Motozono C., Toyoda M., Tan T.S., et al. The SARS-CoV-2 Omicron BA.1 spike G446S mutation potentiates antiviral T-cell recognition. Nat Commun. 2022;13:5440. doi: 10.1038/s41467-022-33068-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sokal A., Chappert P., Barba-Spaeth G., et al. Maturation and persistence of the anti-SARS-CoV-2 memory B cell response. Cell. 2021;184:1201–1213.e14. doi: 10.1016/j.cell.2021.01.050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Röltgen K., Nielsen S.C.A., Silva O., et al. Immune imprinting, breadth of variant recognition, and germinal center response in human SARS-CoV-2 infection and vaccination. Cell. 2022;185:1025–1040.e14. doi: 10.1016/j.cell.2022.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Wang C., Hart M., Chui C., et al. Germinal center B cell and T follicular helper cell responses to viral vector and protein-in-adjuvant vaccines. J Immunol. 2016;197:1242–1251. doi: 10.4049/jimmunol.1502472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Israelow B., Mao T., Klein J., et al. Adaptive immune determinants of viral clearance and protection in mouse models of SARS-CoV-2. Sci Immunol. 2021;6 doi: 10.1126/sciimmunol.abl4509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Tan A.T., Linster M., Tan C.W., et al. Early induction of functional SARS-CoV-2-specific T cells associates with rapid viral clearance and mild disease in COVID-19 patients. Cell Rep. 2021;34 doi: 10.1016/j.celrep.2021.108728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Moss P. The T cell immune response against SARS-CoV-2. Nat Immunol. 2022;23:186–193. doi: 10.1038/s41590-021-01122-w. [DOI] [PubMed] [Google Scholar]
- 53.Riedel R., Addo R., Ferreira-Gomes M., et al. Discrete populations of isotype-switched memory B lymphocytes are maintained in murine spleen and bone marrow. Nat Commun. 2020;11:2570. doi: 10.1038/s41467-020-16464-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Takemori T., Kaji T., Takahashi Y., Shimoda M., Rajewsky K. Generation of memory B cells inside and outside germinal centers. Eur J Immunol. 2014;44:1258–1264. doi: 10.1002/eji.201343716. [DOI] [PubMed] [Google Scholar]
- 55.Anderson S.M., Tomayko M.M., Ahuja A., Haberman A.M., Shlomchik M.J. New markers for murine memory B cells that define mutated and unmutated subsets. J Exp Med. 2007;204:2103–2114. doi: 10.1084/jem.20062571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Mamani-Matsuda M., Cosma A., Weller S., et al. The human spleen is a major reservoir for long-lived vaccinia virus–specific memory B cells. Blood. 2008;111:4653–4659. doi: 10.1182/blood-2007-11-123844. [DOI] [PubMed] [Google Scholar]
- 57.Tangye S.G., Tarlinton D.M. Memory B cells: effectors of long-lived immune responses. Eur J Immunol. 2009;39:2065–2075. doi: 10.1002/eji.200939531. [DOI] [PubMed] [Google Scholar]
- 58.Halfmann P.J., Iida S., Iwatsuki-Horimoto K., et al. SARS-CoV-2 Omicron virus causes attenuated disease in mice and hamsters. Nature. 2022;603(7902):687–692. doi: 10.1038/s41586-022-04441-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Shuai H., Chan J.F., Hu B., et al. Attenuated replication and pathogenicity of SARS-CoV-2 B.1.1.529 Omicron. Nature. 2022;603(7902):693–699. doi: 10.1038/s41586-022-04442-5. [DOI] [PubMed] [Google Scholar]
- 60.Reincke S.M., Prüss H., Wilson I.A., Kreye J. Antigenic imprinting in SARS-CoV-2. Clin Transl Med. 2022;12 doi: 10.1002/ctm2.923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Carreño J.M., Singh G., Simon V., Krammer F. Bivalent COVID-19 booster vaccines and the absence of BA.5-specific antibodies. Lancet Microbe. 2023;4(8) doi: 10.1016/S2666-5247(23)00118-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Alsoussi W.B., Malladi S.K., Zhou J.Q., et al. SARS-CoV-2 Omicron boosting induces de novo B cell response in humans. Nature. 2023;617:592–598. doi: 10.1038/s41586-023-06025-4. [DOI] [PubMed] [Google Scholar]
- 63.Francis T. On the doctrine of original antigenic sin. Proc Am Philos Soc. 1960;104:572–578. [Google Scholar]
- 64.Zhang A., Stacey H.D., Mullarkey C.E., Miller M.S. Original antigenic sin: how first exposure shapes lifelong anti–influenza virus immune responses. J Immunol. 2019;202:335–340. doi: 10.4049/jimmunol.1801149. [DOI] [PubMed] [Google Scholar]
- 65.Anderson K.G., Mayer-Barber K., Sung H., et al. Intravascular staining for discrimination of vascular and tissue leukocytes. Nat Protoc. 2014;9:209–222. doi: 10.1038/nprot.2014.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Masopust D., Soerens A.G. Tissue-resident T cells and other resident leukocytes. Annu Rev Immunol. 2019;37:521–546. doi: 10.1146/annurev-immunol-042617-053214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Takamura S., Yagi H., Hakata Y., et al. Specific niches for lung-resident memory CD8+ T cells at the site of tissue regeneration enable CD69-independent maintenance. J Exp Med. 2016;213:3057–3073. doi: 10.1084/jem.20160938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Çuburu N., Kim R., Guittard G.C., et al. A prime-pull-amplify vaccination strategy to maximize induction of circulating and genital-resident intraepithelial CD8+ memory T cells. J Immunol. 2019;202:1250–1264. doi: 10.4049/jimmunol.1800219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lapuente D., Fuchs J., Willar J., et al. Protective mucosal immunity against SARS-CoV-2 after heterologous systemic prime-mucosal boost immunization. Nat Commun. 2021;12:6871. doi: 10.1038/s41467-021-27063-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Anderson K.G., Sung H., Skon C.N., et al. Cutting edge: intravascular staining redefines lung CD8 T cell responses. J Immunol. 2012;189:2702–2706. doi: 10.4049/jimmunol.1201682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Boyd A., Almeida J.R., Darrah P.A., et al. Pathogen-specific T cell polyfunctionality is a correlate of T cell efficacy and immune protection. PLoS One. 2015;10 doi: 10.1371/journal.pone.0128714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Darrah P.A., Patel D.T., De Luca P.M., et al. Multifunctional TH1 cells define a correlate of vaccine-mediated protection against Leishmania major. Nat Med. 2007;13:843–850. doi: 10.1038/nm1592. [DOI] [PubMed] [Google Scholar]
- 73.Van Splunter M., Van Hoffen E., Floris-Vollenbroek E.G., et al. Oral cholera vaccination promotes homing of IgA+ memory B cells to the large intestine and the respiratory tract article. Mucosal Immunol. 2018;11:1254–1264. doi: 10.1038/s41385-018-0006-7. [DOI] [PubMed] [Google Scholar]
- 74.https://ourworldindata.org/grapher/covid-vaccine-doses-by-manufacturer n.d.
- 75.Barouch D.H., Pau M.G., Custers J.H.H.V., et al. Immunogenicity of recombinant adenovirus serotype 35 vaccine in the presence of pre-existing anti-ad5 immunity. J Immunol. 2004;172:6290–6297. doi: 10.4049/jimmunol.172.10.6290. [DOI] [PubMed] [Google Scholar]
- 76.Sumida S.M., Truitt D.M., Kishko M.G., et al. Neutralizing antibodies and CD8 + T lymphocytes both contribute to immunity to adenovirus serotype 5 vaccine vectors. J Virol. 2004;78:2666–2673. doi: 10.1128/jvi.78.6.2666-2673.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Sanchez S., Palacio N., Dangi T., Ciucci T., Penaloza-Macmaster P. Fractionating a COVID-19 Ad5-vectored vaccine improves virus-specific immunity. Sci Immunol. 2021;6 doi: 10.1126/sciimmunol.abi8635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.WHO https://www.who.int/news/item/18-05-2023-statement-on-the-antigen-composition-of-covid-19-vaccines
- 79.EMA EMA and ECDC statement on updating COVID-19 vaccines to target new SARS-CoV-2 virus variants n.d. https://www.ema.europa.eu/en/news/ema-ecdc-statement-updating-covid-19-vaccines-target-new-sars-cov-2-virus-variants
- 80.FDA’s Vaccines and Related Biological Products Advisory Committee (VRBPAC) Updated COVID-19 vaccines for use in the United States beginning in fall 2023 - VRBPAC 2023. https://www.fda.gov/vaccines-blood-biologics/updated-covid-19-vaccines-use-united-states-beginning-fall-2023
- 81.Collier A.Y., Miller J., Hachmann N.P., et al. Immunogenicity of BA.5 bivalent mRNA vaccine boosters. N Engl J Med. 2023;388:565–567. doi: 10.1056/nejmc2213948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Wang Q., Bowen A., Valdez R., et al. Antibody response to Omicron BA.4–BA.5 bivalent booster. N Engl J Med. 2023;388:567–569. doi: 10.1056/nejmc2213907. [DOI] [PMC free article] [PubMed] [Google Scholar]