ABSTRACT
Parascedosporium putredinis NO1 is a plant biomass-degrading ascomycete with a propensity to target the most recalcitrant components of lignocellulose. Here we applied proteomics and activity-based protein profiling (ABPP) to investigate the ability of P. putredinis NO1 to tailor its secretome for growth on different lignocellulosic substrates. Proteomic analysis of soluble and insoluble culture fractions following the growth of P. putredinis NO1 on six lignocellulosic substrates highlights the adaptability of the response of the P. putredinis NO1 secretome to different substrates. Differences in protein abundance profiles were maintained and observed across substrates after bioinformatic filtering of the data to remove intracellular protein contamination to identify the components of the secretome more accurately. These differences across substrates extended to carbohydrate-active enzymes (CAZymes) at both class and family levels. Investigation of abundant activities in the secretomes for each substrate revealed similar variation but also a high abundance of “unknown” proteins in all conditions investigated. Fluorescence-based and chemical proteomic ABPP of secreted cellulases, xylanases, and β-glucosidases applied to secretomes from multiple growth substrates for the first time confirmed highly adaptive time- and substrate-dependent glycoside hydrolase production by this fungus. P. putredinis NO1 is a promising new candidate for the identification of enzymes suited to the degradation of recalcitrant lignocellulosic feedstocks. The investigation of proteomes from the biomass bound and culture supernatant fractions provides a more complete picture of a fungal lignocellulose-degrading response. An in-depth understanding of this varied response will enhance efforts toward the development of tailored enzyme systems for use in biorefining.
IMPORTANCE
The ability of the lignocellulose-degrading fungus Parascedosporium putredinis NO1 to tailor its secreted enzymes to different sources of plant biomass was revealed here. Through a combination of proteomic, bioinformatic, and fluorescent labeling techniques, remarkable variation was demonstrated in the secreted enzyme response for this ascomycete when grown on multiple lignocellulosic substrates. The maintenance of this variation over time when exploring hydrolytic polysaccharide-active enzymes through fluorescent labeling, suggests that this variation results from an actively tailored secretome response based on substrate. Understanding the tailored secretomes of wood-degrading fungi, especially from underexplored and poorly represented families, will be important for the development of effective substrate-tailored treatments for the conversion and valorization of lignocellulose.
KEYWORDS: Parascedosporium putredinis NO1, lignocellulose, proteomics, CAZymes, activity-based protein profiling
INTRODUCTION
Lignocellulosic residues from the waste components of established food crops (1–3) and from dedicated biomass crops (4) are a large and underutilized source of biomass for biorefinery-based production of chemicals and biofuels. The major constituents of lignocellulose are cellulose, hemicelluloses, lignin, and minor amounts of pectins and nitrogen compounds (5). This intricate and insoluble network of polysaccharides and aromatic polymers makes lignocellulose a difficult substrate to degrade; however, a full understanding of the deconstruction of this complex substrate can offer a valuable source of sustainable fuels, chemicals, and materials for a wide range of applications (6–9).
Biological pre-treatments in the form of enzyme cocktails can deconstruct plant biomass as an environmentally friendly alternative to traditional physicochemical methods. Deployment of these enzymatic treatments has been limited by the hydrophobic and recalcitrant nature of lignocellulose. The heterogeneity of the substrate demands a diverse array of enzymes and treatment can be limited by the release of inhibitor compounds during biomass breakdown (10). This process is further complicated by the differing structures and compositions between lignocellulose sources, each ideally requiring a tailored enzyme cocktail, something that current commercial cocktails do not address (11).
Much research has focused on natural lignocellulose degrading microorganisms for enzymatic solutions to these issues (12, 13). Of particular significance, fungi are widely recognized as efficient degraders of plant biomass and major contributors to the carbon cycle of forest ecosystems. Across the fungal kingdom, a broad range of strategies are used in the deconstruction of lignocellulose. This includes the secretion of arrays of oxidative and hydrolytic enzymes to attack the polysaccharide components of plant cell walls, the production of high levels of lignin-degrading peroxidase and laccase enzymes to attack the lignin directly, and the generation of strong oxidants, via non-enzymatic radical based reactions to degrade lignocellulose indirectly (14, 15). Ascomycete soft-rot fungi are interesting sources of lignocellulose degrading enzymes because they often rely on a strategy of penetrating plant secondary cell walls where they can secrete large amounts of enzymes directly at the site of attack (16).
Parascedosporium putredinis NO1 is a soft-rot ascomycete identified from a mixed microbial community grown on wheat straw (17). P. putredinis NO1 dominated the fungal population in the later stages of the community culture, suggesting an ability to metabolize the more recalcitrant carbon sources in the substrate. This makes P. putredinis NO1 an interesting candidate for the identification of new lignocellulose-degrading enzymes. Indeed, a new oxidase enzyme was recently identified in the P. putredinis NO1 secretome which cleaves the major β-ether units in lignin to enhance the digestibility of wheat straw releasing tricin, coumaric acid, and vanillic acid in the process (17). Understanding the secretome response of a fungus that produces new lignocellulose-degrading enzymes, and which may target the recalcitrant components of lignocellulose, will be important for the development of effective methods to valorize lignocellulose.
An annotated reference genome for this fungus now provides a resource for a more effective investigation of the P. putredinis NO1 secretome (18). Proteomic analysis revealed an extensive and diverse array of carbohydrate-active enzymes (CAZymes) produced by NO1 during growth. The ability of P. putredinis NO1 to grow on various industrially relevant lignocellulosic substrates or kraft lignin has revealed that it is a versatile lignocellulose, particularly lignin, degrader.
To understand what genes are expressed during complex biomass degradation and to what degree these vary depending on the lignocellulosic substrate, the secretome during the growth of P. putredinis NO1 on six lignocellulosic substrates was investigated using both proteomic and activity-based protein profiling (ABPP) techniques. Proteins were harvested from both the supernatant and substrate-bound fractions of liquid cultures to give a more complete picture of the lignocellulose degrading proteome. The variability of this lignocellulose-degrading protein complement across substrates evidences a highly dynamic and tailored response to substrates by P. putredinis NO1. The enzymes identified here represent the vanguard of biomass-degrading capacity produced by a fungus that is highly effective at degrading lignin-enriched biomass. Understanding this complex enzyme system will aid the design of feedstock-matched enzyme cocktails to enhance biorefining efficiency.
RESULTS AND DISCUSSION
Analysis of the P. putredinis NO1 secretome on different lignocellulosic substrates
To investigate whether the secretome of P. putredinis NO1 varies with lignocellulosic substrates, the fungus was grown on six substrates: oil palm empty fruit bunch, kraft lignin, rice straw, sugar cane bagasse, wheat bran, and wheat straw, and the resulting proteomes were sampled and analyzed. Proteins were harvested from culture supernatants and via a surface protein labeling approach with biotin that harvests proteins bound to substrates (19). It should be noted that kraft lignin is more soluble than the other substrates and therefore a larger portion of the kraft lignin bound fraction likely reflects proteins associated with the fungal cell wall. After harvesting and identifying proteins through label-free liquid chromatography–mass spectrometry (LC–MS), molar percentage values were generated for each protein. Quality control was performed by investigating protein count, performing principal component analysis, and clustering for all replicates. Through this analysis, a bound fraction sugar cane bagasse replicate, a supernatant fraction sugar cane bagasse replicate, a supernatant kraft lignin replicate, and a supernatant fraction wheat bran replicate were identified as outliers and removed from further analysis.
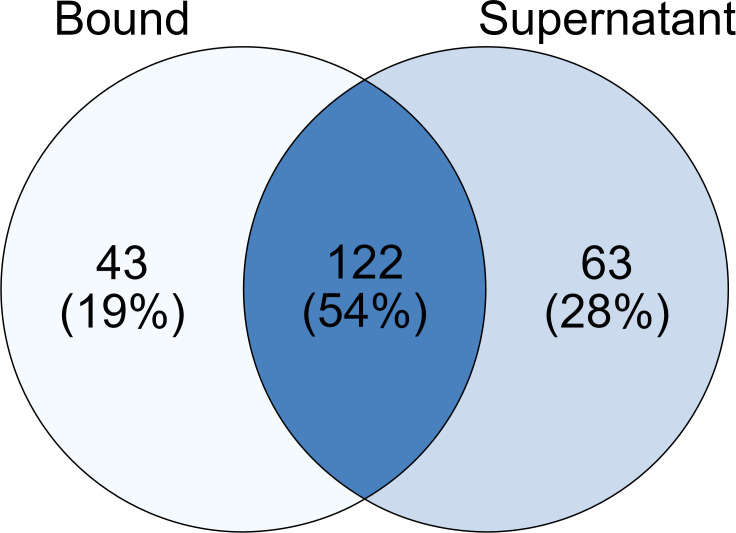
Across all samples 2,014 proteins were identified in at least one replicate, 1,890 proteins were identified among the substrate-bound fractions, and 973 were identified among the supernatant fractions, with 849 proteins shared across both fractions (Fig. S1). Many intracellular proteins, which were likely the results of cell death and lysis, were present in the proteomic data set despite efforts to target the extracellular fractions (20). The proteomic data were therefore filtered to remove the contaminating intracellular proteins according to the secretome isolation bioinformatics workflow presented previously (21). Secretome proteins were predicted to be extracellular using both BUSCA and DeepLoc localization prediction tools (20, 22), or were predicted to encode a secretion signal by SignalP, TargetP, and SecretomeP tools (23–25). The workflow utilizes both localization prediction and secretion signal prediction to capture both conventionally and unconventionally secreted fungal proteins (26). The vesicle-mediated release of lignocellulose degrading enzymes has been reported previously for Trichoderma reesei, another ascomycete degrader of plant biomass, highlighting the importance of capturing proteins secreted through unconventional pathways (27). Proteins that contained more than one predicted transmembrane domain by TMHMM or contained a single predicted transmembrane helix with more than 10 amino acids of this helix occurring in the first 60 amino acids of the protein sequence were then removed (28). The final secretome contained 228 proteins, a significant reduction from the total proteome data set. In the substrate-bound fraction of the secretome 165 proteins were identified in at least a single replicate, in the culture supernatant fraction 185 proteins were identified, and 122 proteins were shared between both fractions (Fig. 1). The now much smaller number of proteins identified exclusively in the bound and supernatant fractions were 43 and 63 proteins, respectively. Considering each substrate individually; for empty fruit bunch 107 proteins were identified in the bound fraction, 109 exclusively in the supernatant fraction, and 58 shared between both; for Kraft lignin, 85 proteins were identified in the bound fraction, 20 in the supernatant fraction, and 14 shared between both; for rice straw 130 proteins were identified in the bound fraction, 100 in the supernatant fraction, and 60 shared between both; for sugar cane bagasse 105 proteins were identified in the bound fraction, 104 in the supernatant fraction, and 66 shared between both; for wheat bran 72 proteins were identified in the bound fraction, 150 in the supernatant fraction, and 47 shared between both; wheat straw 101 proteins were identified in the bound fraction, 144 in the supernatant fraction, and 77 shared between both. The difference between the substrate-bound and culture supernatant fractions highlights the importance of capturing both fractions, in order to fully understand the secretome response. The limitations of this workflow have been addressed previously (21), and it is accepted that some intracellular proteins may remain in the predicted secretome. However, the increased proportion of proteins annotated with extracellular functions achieved through this workflow demonstrates the removal of large numbers of intracellular proteins and therefore improves any interpretation of the P. putredinis NO1 secretome.
Fig 1.
Investigating the distribution of proteins across bound and supernatant fractions in the P. putredinis NO1 secretome. The number of proteins identified in at least one replicate across all substrates for the bound fraction compared to the supernatant fraction within the predicted P. putredinis NO1 secretome from growth on six lignocellulosic substrates for 4 days.
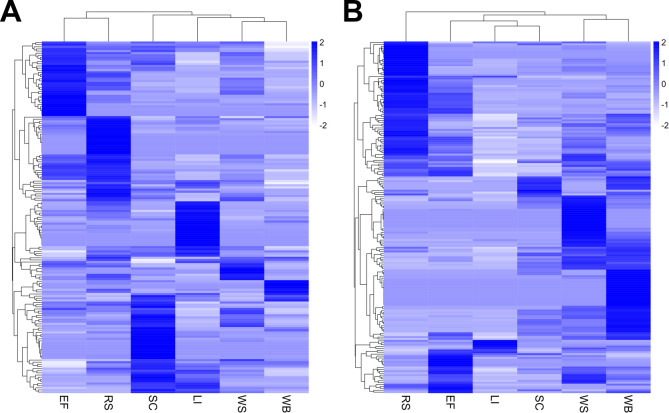
The overall protein abundance profiles of the P. putredinis NO1 secretome after growth on each substrate were visualized by scaling molar percentage abundance for each protein across substrates and clustering (Fig. 2). The substrate-dependent variation in the P. putredinis NO1 secretome was visible, with all substrates displaying substrate exclusive protein profiles to varying extents in both fractions. The differences between substrates were also investigated through principal component analysis, and clustering (Fig. S2 and S3). In the substrate-bound fraction, replicates grouped very well within substrates in principal component analysis (PCA) plots and clustered well within substrates in dendrograms, while demonstrating good separation between substrates in PCA plots and dendrograms also. In the supernatant fraction, the differences between substrates were less clear, with single replicates from some substrates grouping away from other replicates of the same substrate; however, this is likely due to the lower identified protein count for these replicates in the supernatant fraction. The lower number of replicates for the supernatant fractions of some substrates, and the lower protein counts in the supernatant fractions of some replicates reflect the difficulty in applying a single method of proteome harvest to cultures containing different substrates and therefore different conditions. This could also limit the ability to perform statistical comparisons between substrates. However, for this study, these replicates were still suitable for the visualization and comparison of the overall profiles of the P. putredinis NO1 secretome across substrates. The heatmaps in Fig. 2 still demonstrate that clear differences are visible across substrates in both fractions. This is especially interesting considering the somewhat similar proportions of cellulose, hemicellulose, and lignin for the empty fruit bunch from oil palm (29), rice straw (30), sugar cane bagasse (31), and wheat straw (32). Differences in fungal secretomes have been demonstrated previously when grown on compositionally distinct carbon sources (33–35). However, the variation on broadly similar lignocellulosic substrates demonstrated here is important as it may suggest a tailoring of the lignocellulose-degrading enzymes produced by P. putredinis NO1 which could be utilized for substrate-specific cocktail development. When T. reesei was grown on compositionally similar cellulose substrates it was suggested that secretome variation may be influenced by physical and structural differences (33), and the differences between the substrates investigated here may influence the variability of the P. putredinis NO1 secretome. Further study of these more subtle structural, compositional, and physical differences could improve the understanding of how fungal secretomes are influenced and tailored accordingly. Wheat bran has a different overall composition than the other lignocellulosic substrates with an increased starch content, correspondingly decreased proportions of cellulose and hemicellulose, and a reduced lignin content (36). The kraft lignin substrate is expected to contain minimal polysaccharides and instead is composed predominantly of branched polymeric lignin containing native lignin bonding patterns and new structures created during the kraft pulping process (37). The difference in the secretome abundance profiles after the growth of P. putredinis NO1 on wheat bran and kraft lignin was therefore expected.
Fig 2.
Differences in the P. putredinis NO1 secretome across lignocellulosic substrates. Molar percentage values for proteins identified on at least one substrate in the bound (A) and supernatant (B) fractions of the P. putredinis NO1 secretome scaled to Z-scores across substrates. EF, empty fruit bunch; LI, kraft lignin; RS, rice straw; SC, sugar cane bagasse; WB, wheat bran; WS, wheat straw.
An investigation of the functional profile of the P. putredinis NO1 secretome on different lignocellulosic substrates
To investigate the functional profiles of the P. putredinis NO1 secretome across lignocellulosic substrates, proteins were annotated with CAZyme domains and GO categories and terms. The overall functional profiles were investigated by comparing the proportional abundance of proteins annotated with CAZyme domains, if not annotated as CAZymes then assigned to GO categories, or not annotated with either (unknown) (Fig. 3). Clear variation in the proportions of functional category assignments were observed across substrates in both bound and supernatant fractions. The differences between bound and supernatant fractions for individual substrates validate further the importance of capturing both fractions when exploring a secretome in this context, as each fraction contains distinct profiles of enzyme activities. In the bound fraction (Fig. 3A), CAZymes contribute the most to total abundance for all substrates. This contribution is slightly reduced for the kraft lignin substrate, which is to be expected due to the low abundance of polysaccharides on which many CAZymes act (37). In the supernatant fraction (Fig. 3B), CAZymes again contribute the most to total abundance except for rice straw. Rice straw has a lower lignin content than some of the other substrates at 12% which may suggest increased accessibility to polysaccharide components and a reduced requirement for the production of oxidative lignin degraders (30). Structural aspects of lignocellulose such as cellulose structure and accessibility, pore size and distribution, and the extent and nature of lignin-carbohydrate complexes have all been reported to influence the efficiency of enzymatic hydrolysis (38). However, the relationship between such physical characteristics and microbial enzyme production is poorly understood. Surprisingly, CAZymes contributed the most to total abundance in the supernatant fraction for kraft lignin which, although poorly soluble in water, lacks the polysaccharide chains targeted by binding modules (33, 39). Despite bioinformatic filtering of proteomic data to produce the P. putredinis NO1 secretome, some intracellular and membrane proteins may have persisted, as evidenced by the contribution of “cellular component” GO category proteins to abundance for all substrates in both bound and supernatant fractions. The persistence of proteins with predicted intracellular functions was reported previously in the development of the secretome isolation workflow applied here (21). The interference of these potentially intracellular proteins could have been reduced through stricter filtering when obtaining the in silico secretome; however, this could risk removing unknown proteins that are important for lignocellulose degradation. In both the bound and supernatant fractions, unknown proteins contribute similar proportions of abundance as the functional GO categories and again the proportion of these unknown proteins varies across substrates in both fractions investigated.
Fig 3.
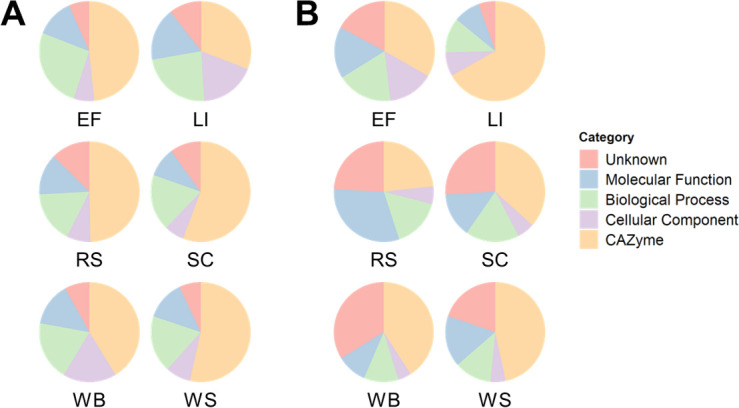
Differences in functional categories of the P. putredinis NO1 secretome across lignocellulosic substrates. The molar percentage abundance of proteins identified as CAZymes, that were not identified as CAZymes but assigned to GO categories, or that were not annotated as CAZymes or with GO categories (unknown) were calculated proportionally for each substrate of the bound (A) and supernatant fractions (B) of the P. putredinis NO1 secretome.
To understand the functional profiles of the P. putredinis NO1 secretome in more detail, the proportional abundance of the ten most abundant GO terms was plotted for each substrate for the bound and supernatant fractions (Fig. 4 and 5). For all substrates in both the bound (Fig. 4) and supernatant (Fig. 5) fractions, the top GO term annotations contributing to total abundance were predominantly extracellular activities associated with the hydrolytic breakdown of lignocellulose, agreeing with the polysaccharide abundance in many of the substrates (29–32, 36). In both fractions, many substrates included membrane annotations in the top ten most abundant GO terms. This may result from the identification of proteins in the workflow which may be targeted to the secretory pathway, but that is not necessarily secreted and instead fulfill roles in membranes (40). Another possibility is the binding and extraction of extracellular cell surface proteins by the biotin labeling technique, as biotinylation is a common technique for surface proteomic studies (41). In the bound fraction, membrane GO annotations contributed the highest abundance of any GO term for kraft lignin (Fig. 4B) and wheat bran (Fig. 4E), which may reflect the reduced CAZyme proportions seen previously for these substrates in this fraction. Once again, kraft lignin and wheat bran, the two most compositionally distinct substrates explored here demonstrate the most distinct secretome activity profiles. Aside from cell surface proteins, abundant intracellular annotations were largely absent from most of the secretomes except for the kraft lignin substrate. Unknown proteins, i.e., with no GO annotations, were also present in the top ten most abundant annotations for all substrates in both fractions except for the supernatant fraction of kraft lignin. In the supernatant fraction, unknown proteins made the largest contribution of any annotation for empty fruit bunch (Fig. 5A), rice straw (Fig. 5C), sugar cane bagasse (Fig. 5D), wheat bran (Fig. 5E), and wheat straw (Fig. 5F). The most abundant annotations for most substrates in both fractions appear to be associated with the breakdown of cellulose and xylan, two abundant polysaccharides in these lignocellulosic substrates (29–32, 36). However, there is a clear difference in profile for the kraft lignin substrate in both the bound (Fig. 4B) and supernatant (Fig. 5B) fractions. In the bound fraction, there appear to be more membrane-targeted GO annotations in the ten most abundant annotations. Finally, the GO annotation profile of the bound fraction for wheat bran (Fig. 4E) distinctly contained chitin binding and catabolizing annotations. Wheat bran has been investigated for its use as a substrate for the induction of chitinase production in fungal species previously and perhaps a similar induction is occurring for P. putredinis NO1 (42).
Fig 4.
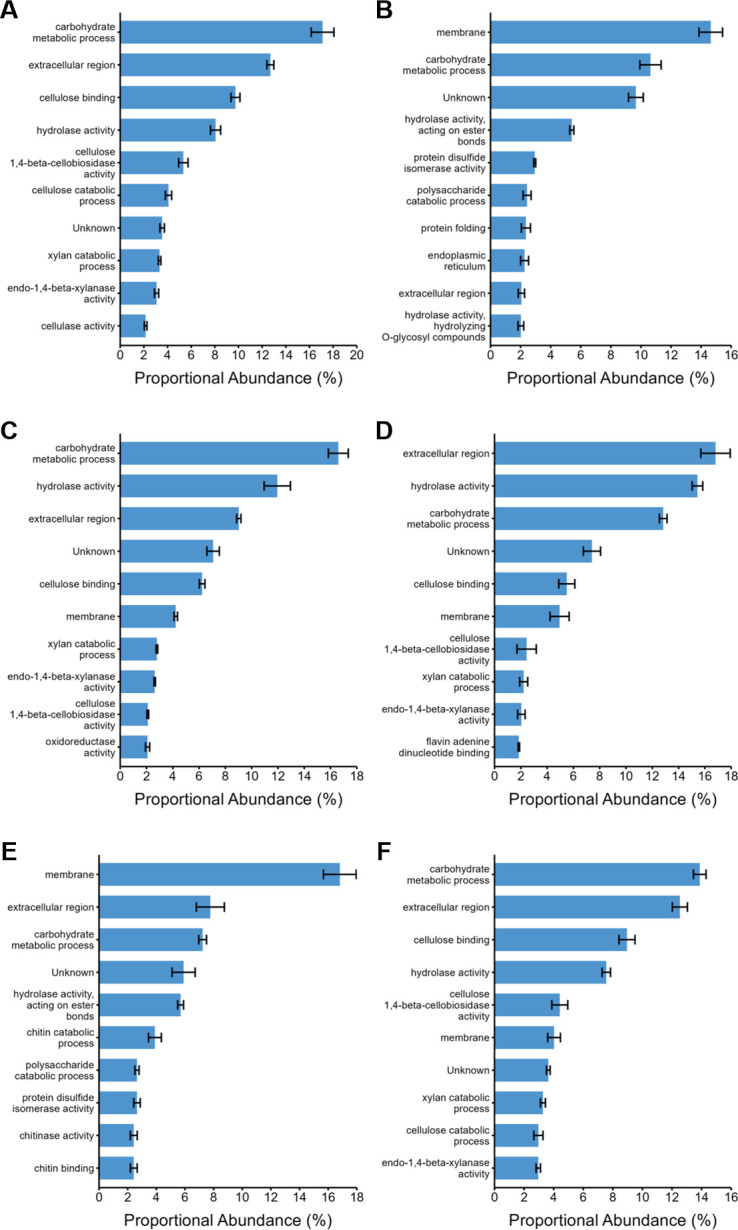
Proportional abundances of enzyme activities in the bound fraction of the P. putredinis NO1 secretome. The molar percentage abundance of proteins assigned to GO terms was calculated proportionally for the bound fraction of the P. putredinis NO1 secretome after 4 days of growth on six lignocellulosic substrates. EF, empty fruit bunch (A); LI, kraft lignin (B); RS, rice straw (C); SC, sugar cane bagasse (D); WB, wheat bran (E); WS, wheat straw (F). Average molar percentage ± standard error (n = 3, n = 2 for SC substrate).
Fig 5.
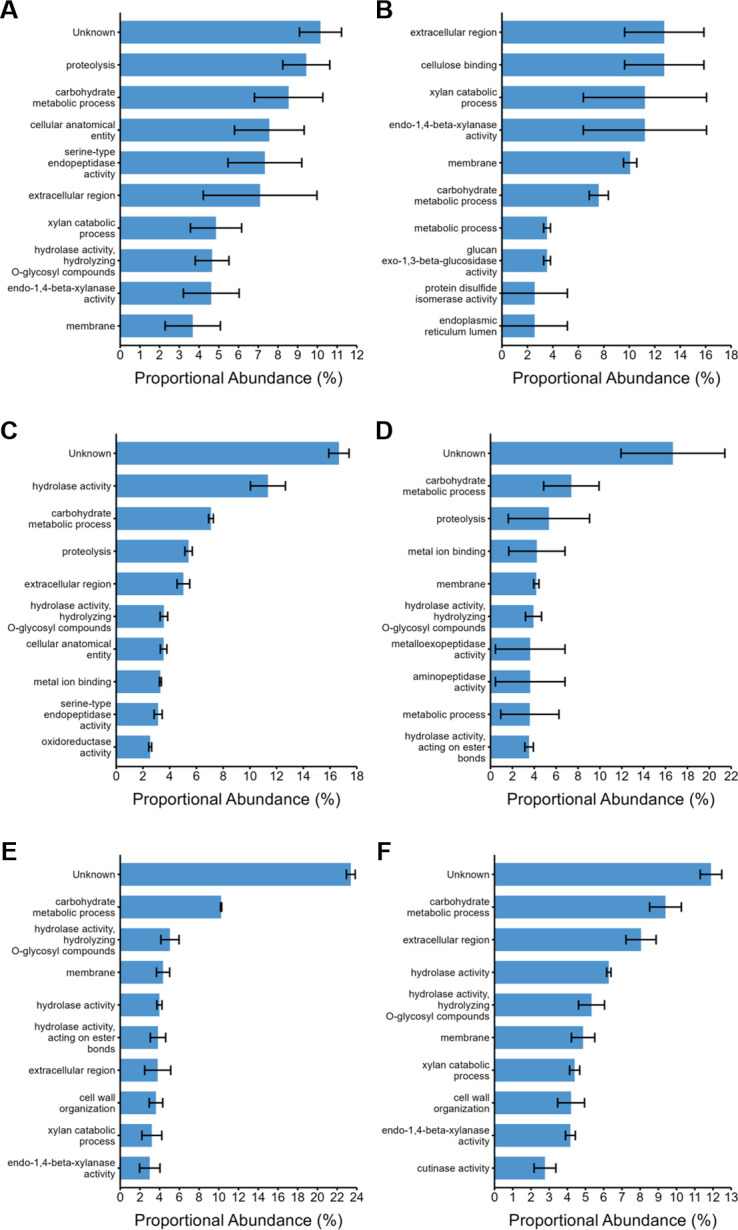
Proportional abundances of enzyme activities in the supernatant fraction of the P. putredinis NO1 secretome. The molar percentage abundance of proteins assigned to GO terms was calculated proportionally for the supernatant fraction of the P. putredinis NO1 secretome after 4 days of growth on six lignocellulosic substrates. EF, empty fruit bunch (A); LI, kraft lignin (B); RS, rice straw (C); SC, sugar cane bagasse (D); WB, wheat bran (E); WS, wheat straw (F). Average molar percentage ± standard error (n = 3, n = 2 for SC, LI, and WB substrates).
The lignocellulose degrading secretome of P. putredinis NO1 varies depending on the growth substrate
To investigate the lignocellulose-degrading enzyme repertoire of the P. putredinis NO1 secretome, protein sequences were annotated for CAZyme domains using the dbCAN server (43). CAZyme proteins are assigned to the following catalytic classes: auxiliary activity (AA), carbohydrate esterase (CE), glycoside hydrolase (GH), glycosyl transferase (GT), and polysaccharide lyase (PL). Additionally, many CAZymes have carbohydrate-binding modules (CBMs) which are also assigned.
Comparing the contribution of proteins assigned to these catalytic classes to total CAZyme abundance across substrates demonstrates similar variation seen for the total secretome and for functional categories in both bound and supernatant fractions (Fig. 6). In the bound fraction AA class CAZymes contribute a high proportion of total CAZyme abundance for the empty fruit bunch, rice straw, sugar cane bagasse, and wheat straw substrate but are low in abundance for wheat bran and kraft lignin substrates (Fig. 6A). Crystalline cellulose is present in lower levels or is absent from wheat bran and kraft lignin substrates, and this may reflect a reduced production of AA class CAZymes involved in lytic polysaccharide monooxygenase (LPMO) systems which degrade crystalline cellulose oxidatively (36, 37, 44, 45). Potentially, the oxidative enzymes responsible for lignin breakdown are still present but in low abundance. The reductions in AA abundances for wheat bran and kraft lignin substrates correspond with increased proportions of all other CAZyme classes in the bound fraction, including GT class CAZymes (46). Some GT enzymes are extracellular and bound to the fungal cell surface which is likely why these proteins have been isolated in the P. putredinis NO1 secretome and they are more likely to play a role in fungal growth, cell wall remodeling, or potentially associate the fungus with the growth substrate (47, 48). PL class enzymes were absent from both fractions of the P. putredinis NO1 secretome during growth on kraft lignin. They were either absent or present in low abundances for all substrates in the bound fraction but were present with varying contributions to total CAZyme abundance in the supernatant fraction. This could be explained by the soluble pectin substrates on which PL enzymes act which are more likely to be present in the culture supernatant (49). In the supernatant fraction, GH class CAZymes dominated the CAZyme abundance profiles for all substrates (Fig. 6B). It can be hypothesized that soluble components of the substrates are present initially and are also released into the supernatant fraction during degradation. These soluble components would then be accessible to the wide array of hydrolytic GH class CAZyme activities which act on diverse substrates, and this potentially explains the abundance of these enzymes in this fraction (50). The similarity in the proportion of GH class CAZymes in the supernatant fraction even for compositionally distinct substrates such as wheat bran and kraft lignin could be the result of the transcriptional activation of GH class CAZymes like cellulases and xylanases. Mono- and disaccharides, which could be present in all substrates investigated here, have been demonstrated to act as inducers of the expression of large numbers of hydrolytic CAZymes in other ascomycete fungi previously (35, 51). This may be occurring for all substrates despite the target polysaccharides of induced enzymes not necessarily being present in similar amounts across all substrates investigated.
Fig 6.
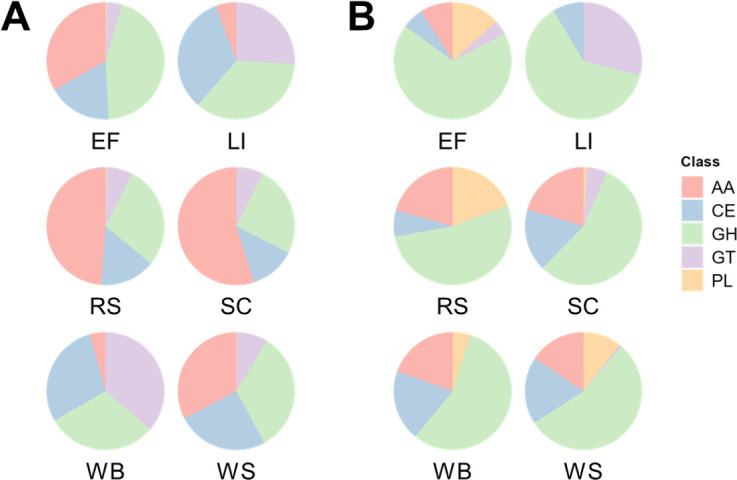
Differences in proportional catalytic CAZyme class abundance of the P. putredinis NO1 secretome across lignocellulosic substrates. The molar percentage abundance of proteins belonging to each catalytic class of CAZyme was calculated proportionally to the total abundance of CAZymes for each substrate for the bound (A) and supernatant fractions (B) of the P. putredinis NO1 secretome.
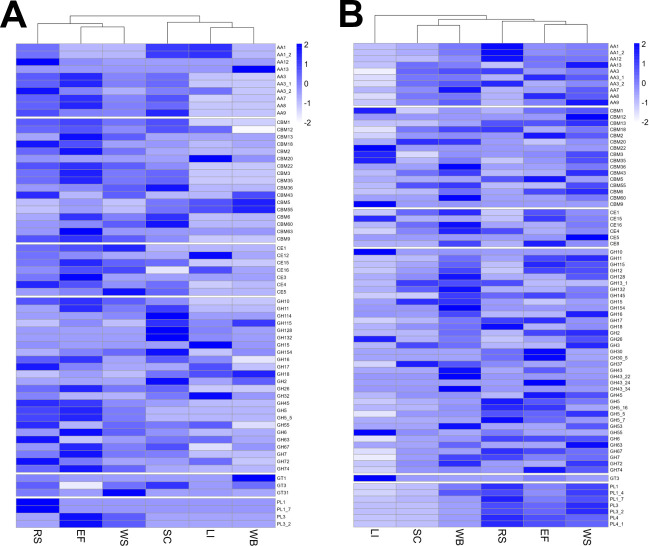
To investigate the CAZyme classes in more detail and to address some of the questions raised by comparing class profiles, the CAZyme family abundances were compared across substrates for the bound and supernatant fraction separately (Fig. 7). Variation across substrates was again clear at the CAZyme family level in both fractions even for substrates which looked similar at the class level. This highlights how similarities at the class level masked these underlying differences at the level of the CAZyme family, which defines an enzyme’s activity more specifically. As expected, the relative abundances of AA9 LPMOs and other AA families, for example, AA3, AA7, and AA12 which act in LPMO systems are reduced for kraft lignin and wheat bran substrates in the bound fraction (52–54). However, the AA1 laccase family enzymes responsible for the oxidative cleavage of lignin structures are present in increased abundance in the bound fraction for the kraft lignin substrate (53). All AA families were muted in abundance in the bound fraction for wheat bran except for the AA13 family of starch-degrading LPMOs, likely resulting from the increased starch content of the substrate (36, 55). Variation in the abundance profiles of proteins with CBMs belonging to different families can also be seen in (Fig. 7). CBMs are non-catalytic domains that predominantly bind to plant cell wall polymers to prolong contact of catalytic CAZyme domains with their substrates to enhance efficiency (56). Harvesting the bound fraction of lignocellulose-degrading secretomes was performed in an attempt to capture proteins with such binding capacity, which may have been missed by harvesting proteins from the soluble supernatant fraction only. A wide range of CBM families provide binding capabilities for different enzymes to a range of cell wall components, and the variation in CBM containing protein abundance observed here may reflect the varying compositions and structures of the substrates (33). Indeed some expected patterns can be seen in the bound fraction, such as the relatively lower abundance of CBM1 proteins, involved in crystalline cellulose binding, to kraft lignin and wheat bran substrates which have reduced proportions of cellulose in comparison to the other substrates investigated (36, 37, 57). For wheat bran, in the bound fraction there is a relatively high abundance of CBM43, CBM5, and CBM56-containing proteins. CBM5 proteins are involved in binding chitin and are likely to be involved in cell-wall remodeling here (58). CBM43 and CBM56 are involved in binding β-1,3-glucans, which are present in fungal cell walls, but also present in the wheat bran substrate and are potentially being targeted for breakdown here (36, 59, 60). The hydrolytic families of CAZymes belonging to GH, CE, and PL classes show variation in abundance profiles across all substrates in both fractions and likely reflect a tailored hydrolytic response to the different polysaccharide compositions of each substrate. Although the cellobiose and xylose-mediated induction of hydrolytic CAZymes is documented for more established ascomycete fungi (35, 51, 61), whether these mechanisms exist in P. putredinis NO1 is not yet known. The variation in other classes of CAZyme in ascomycetes, where regulation has not been well investigated in the context of lignocellulose breakdown, also suggests additional mechanisms of activation of gene expression.
Fig 7.
Differences in CAZyme family abundance of the P. putredinis NO1 secretome across lignocellulosic substrates. Molar percentage values for proteins annotated as CAZymes and identified on at least one substrate of the P. putredinis NO1 secretome scaled to Z-scores across substrates for the bound (A) and supernatant (B) fractions separately.
Generally, the abundances of the hydrolytic CAZymes are lower for the kraft lignin substrate in both fractions, agreeing with the reduced levels of polysaccharides. In the supernatant fraction, the relatively high abundance of some polysaccharide hydrolyzing or binding CAZyme family domains during growth on kraft lignin may reflect enzymes produced irrespective of the lignocellulosic growth substrate. Indeed, it has been observed in T. reesei that spore germination alone can lead to substantial upregulation of polysaccharide-degrading CAZymes to prepare the fungus for a habitat containing plant biomass (62). The GH10, GH26, GH55, and CE15 families produced in relatively high abundance for kraft lignin possibly represent a similar response. It is possible that the action of these enzymes, which may be produced irrespective of the substrate, releases oligo- or mono-saccharides that could lead to induction and tailoring of the secretome (35, 51, 61). Investigating CAZyme family profiles in the supernatant fraction also demonstrates reduced relative abundances for all CAZyme domain-containing proteins for the sugar cane bagasse substrate, and variation across wheat straw, rice straw, and empty fruit bunches from oil palm, despite all these substrates having similar overall biomass compositions (63, 64). This variation demonstrates the complexity of the fungal plant biomass-degrading response to different substrates which is still poorly understood.
The differences in CAZyme family abundance all suggest a potential ability of P. putredinis NO1 to tailor its secretome dependent on its lignocellulosic growth substrate. Empty fruit bunches from oil palm, rice straw, sugar cane bagasse, and wheat straw all show clear differences between the substrates as well, although the overall compositions are more similar (29–32). One limitation of the current study was that it was possible for the observed differences in secretome profiles to be a product of different fungal growth stages on each substrate, as all data were harvested at a single time point. An alternative approach was therefore taken to confirm that differences in the P. putredinis NO1 secretome are maintained over time.
Activity-based protein profiling shows how lignocellulose-degrading enzyme profiles vary over time
Fluorescence-based ABPP with cyclophellitol-derived activity-based probes (ABPs) for retaining β-glucosidases (65), cellulases (66), and xylanases (67) have been used to screen fungal secretomes previously (67, 68). Cellulases, xylanases, and β-glucosidases are broadly distributed and highly expressed in plant-biomass degrading fungi, making them good candidates for ABPP techniques and for comparison across lignocellulolytic systems (68). These probes were also used to display the induction of an array of GH enzymes in basidiomycete secretome when grown on lignocellulosic compared to simple substrates like maltose (68). Therefore, this is an appropriate and effective technique to explore the potentially tailored secretome of the ascomycete P. putredinis NO1 during growth on lignocellulosic substrates.
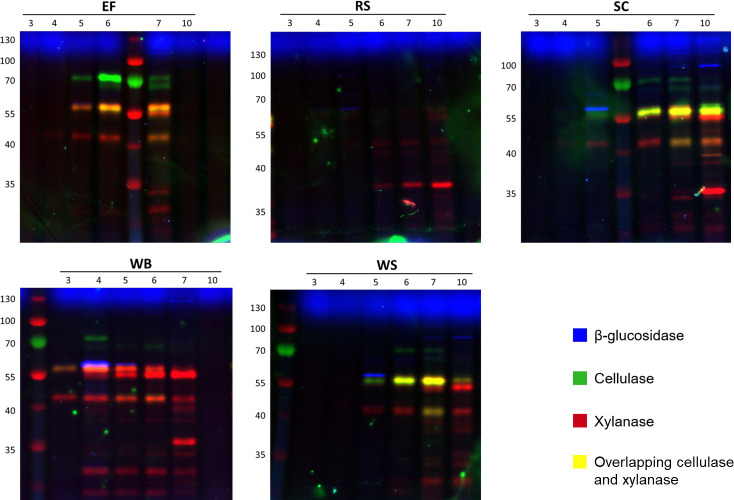
Activity-based probes for GHs were employed to screen samples of supernatants harvested from cultures of P. putredinis NO1 grown on the same substrates as the proteomics experiment at days 3, 4, 5, 6, 7, and 10 (69). The biotin-labeling approach used in the proteomic experiments to sample bound fractions involves protein denaturation and is therefore incompatible with this ABPP technique which requires active site binding. The supernatants were used to screen for and to determine the relative levels of lignocellulose-degrading glycoside hydrolases. The activities were present in the empty fruit bunch from oil palm, rice straw, sugar cane bagasse, wheat bran, and wheat straw samples (Fig. 8). Unfortunately, the high aromatic content of the kraft lignin interfered with staining and meant that this substrate was incompatible with this technique (Fig. S4). Fluorescent gel visualization was performed for triplicate samples from all substrates and good agreement in hydrolase abundance pattern over time was observed within substrates (Fig. S4). For simplicity, single replicate time course fluorescence gels are presented in Fig. 8, and integrated band intensity values are resolved in Fig. 9.
Fig 8.
Differences in P. putredinis NO1 glycoside hydrolase production over time visualized with activity-based probes. Fluorescence imaging following SDS-PAGE is shown for single replicates of samples of culture supernatants taken at days 3, 4, 5, 6, 7, and 10 of growth. They were treated with a triplex probe mixture targeting cellulases, xylanases, and retaining β-glucosidases. EF, RS, SC, WB, WS, alongside PageRuler (Thermo) prestained protein ladder. Secretomes were stained in triplicate, a single replicate is shown here.
Fig 9.
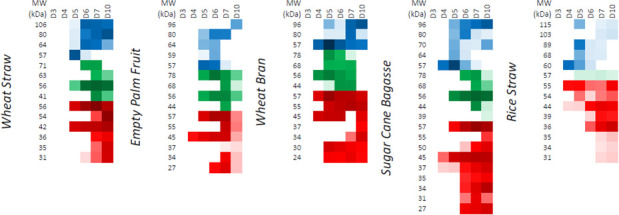
ABPP-determined variation in relative active enzyme levels over time during P. putredinis NO1 growth on various substrates. Resolved bands running at different apparent MW values (left column in each block) were integrated into the Cy2 (β-glucosidase probe), Cy3 (cellulase probe), or Cy5 (xylanase probe) channels. Average band integration values (n = 3) are shown as color intensity varying from white (not detected) to full color (~1,000,000 counts) to black (saturation) on a logarithmic scale. Secretomes were prepared and stained in biological triplicate following different culture times (labels above columns).
As seen previously for screening of basidiomycete secretomes, the P. putredinis NO1 secretome shows differential GH production over time dependent on growth substrate (Fig. 8 and 9) (68). P. putredinis NO1 demonstrated muted production of GHs at day 3 for all substrates. For rice straw, this production remained low for the entire time course with low levels of xylanase and β-glucosidase and only a single cellulase detected at low abundances. Although the production profiles of these GHs were demonstrated to vary by species, it is worth noting that transcriptomic analysis of another ascomycete, Thielavia terrestris, only found a single cellulase gene to be in the most highly expressed genes when grown on rice straw compared to eleven cellulases for growth on Avicel (68, 70). It was also observed that compared to growth on glucose, predominantly oxidative AA LPMO family CAZymes and hemicellulose-active enzymes were up-regulated during T. terrestris growth on rice straw (70). The reason for the lack of cellulose targeting GHs also observed here is unclear, especially considering the reported induction of cellulases from both cellobiose and xylose in other ascomycetes (35, 51, 61), but may suggest a predominantly oxidative approach to rice straw deconstruction instead. Perhaps the distinct fluorescent profile observed here for rice straw may result from the unique structural properties of rice straw, such as its high silica content which may prevent access to polysaccharides (71). Relatively high abundances of AA3 and AA12 family CAZymes were observed in the rice straw secretome supernatant fraction, where the probes are deployed (Fig. 7). These oxidative enzymes act within LPMO systems; however, relatively low abundances of the AA9 LPMOs were also observed. Further investigation specifically into growth on rice straw would be required to fully understand the strategy of lignocellulose breakdown adopted here by P. putredinis NO1 but is warranted considering the abundance and environmental issues posed by the vast amounts of rice straw generated annually and burnt on the field (72, 73).
Substrate-specific xylanase production patterns have been observed previously using the same fluorescent probe to investigate the growth of Aspergillus niger on beechwood xylan (67). The xylanase probe detected similar variation across substrates here, most noticeably for wheat bran where xylanase production was the most dominant of the three types of GH screened (Fig. 8). Cellulase production was detected despite being obscured on the gel images by the high xylanase signals (Fig. 9). However, cellulase production was not sustained until day 10 as it was for wheat straw, sugar cane bagasse, and empty fruit bunch. This reflects the pattern of reduced cellulose degrading CAZyme family abundance observed for wheat bran in the proteomic investigation, a substrate with reduced cellulose content (36). Although wheat bran has been explored as a potential substrate for cellulase production from the ascomycete T. reesei, where the nitrogen-rich substrate induces cellulase production (74, 75), this study demonstrates how varied fungal lignocellulose-degrading enzyme responses can be for the same substrate and how characterization of new lignocellulose-degrading systems are important for understanding different approaches which may adopted industrially for the conversion of biomass.
GH CAZyme production patterns looked similar for wheat straw and sugar cane bagasse substrates aside from some additional low molecular weight xylanases detected for the latter. All substrates except rice straw showed the presence of a 57 kDa glucosidase at high levels on day 5 which then became undetectable by day 7 for wheat straw, empty fruit bunch, and sugar cane bagasse, often giving way to a series of higher MW glucosidases. This two-stage pattern of glucosidase production may represent distinct responses to water-soluble glucosides vs polysaccharide-derived glucosides. It has also been observed previously in microbial communities that β-glucosidase patterns can shift depending on other available carbon sources (76). Perhaps, the two-stage pattern of β-glucosidase production observed here also reflects a change in available carbon after day 5 of growth on these substrates.
Time-dependent analysis of the oil palm empty fruit bunch secretomes showed that induction was slow, with no enzymes detected at day 4. Day 5 showed a strong induction across all samples of the production of cellulases, glucosidases, and xylanases. Especially obvious was the sustained high production of a higher molecular weight cellulase detected across all substrates investigated (Fig. 8 and 9). The production of hemicellulose-degrading enzymes by A. niger on this substrate has been explored previously; however, exploration of cellulase production has not been performed for fungal isolates (77). The growth of the bacterium Paenibacillus macerans on empty fruit bunches from oil palm has been explored where degradation of the substrate was demonstrated (78). However, cellulase activity from this microorganism was not observed. Overall, this analysis demonstrates the time-dependent adaptability of the P. putredinis NO1 secretome to different lignocellulosic substrates and illuminates the different CAZyme specificities deployed by this fungus. Considering the potential lignin-degrading lifestyle of P. putredinis NO1, from which a new lignin-degrading oxidase has been identified previously (17), the investigation of oxidative enzymes in the secretome and their variation dependent on substrate and over time would be of great interest. However, at present, the use of ABPP techniques to probe fungal secretomes as was performed here is limited to hydrolytic enzymes.
Conclusions
Proteomic analysis of the secretome of P. putredinis NO1 grown on multiple lignocellulosic substrates showed a substrate-dependent variation of the CAZyme complement both bound to the biomass and in the culture supernatant. This variation was maintained when comparing the abundances of CAZymes at both the class and family levels. Patterns of abundance of important lignocellulose degrading enzymes were observed, for example, the lack of crystalline cellulose targeting enzymes during growth on substrates with reduced cellulose contents. When investigating CAZymes at the family level, which more specifically defines enzyme activity, the abundance profiles were found to vary greatly across substrates in both fractions of the lignocellulose degrading secretomes. This likely reflects the varying structures and compositions of the lignocellulosic substrates and suggests a potential ability for P. putredinis NO1 to actively tailor its enzymatic response.
It could be argued that differences in proteomic data across substrates reflected different stages of fungal growth on the different substrates. However, by utilizing an ABPP-based approach to visualize secretomes it was demonstrated that the diversity and titers of active cellulases, xylanases, and β-glucosidases in the secretomes varied both with substrate and over time. Therefore, the fungal growth stage is not solely responsible for differences observed between the secretomes. Patterns of production of these GH class CAZymes were also found which both agreed and contradicted previous reports on fungal GH expression on lignocellulosic substrates (35, 51, 61). This highlights how varied and complex lignocellulose degrading systems can be and demonstrates the value of exploring new systems like P. putredinis NO1 to better understand lignocellulose breakdown and to identify novel enzymes.
The complexity of fungal lignocellulose-degrading responses revealed here highlights the shortcoming of generic commercial cocktails for application to different substrates, but also the difficulty in disentangling enzyme effects and designing optimally tailored cocktails. Understanding the adaptability of fungal secretomes will allow increased efficiency in the depolymerization and biorefining of lignocellulosic substrates. Furthermore, the significant number of unknown proteins identified in the secretome after filtering through the bioinformatics pipeline suggests that there are still new lignocellulosic activities to be discovered.
MATERIALS AND METHODS
Strain isolation
P. putredinis NO1 was isolated from a wheat straw enrichment culture and maintained as reported previously (17). A mixed microbial community grown on wheat straw was characterized by 16S ribosomal and ITS region amplicon sequencing over 8 weeks of growth. ITS reads assigned to the genus Graphium dominated the eukaryotic community after 4 weeks of incubation. P. putredinis NO1, a synamorphy of Graphium, was readily cultivated from this community on potato dextrose agar plates.
P. putredinis NO1 cultures for proteomics
Triplicate 500 mL solutions of media containing 1.5% (wt/vol) rice straw, wheat straw, sugarcane bagasse, wheat bran, empty fruit bunches from oil palm or containing 5% (wt/vol) kraft lignin were inoculated to a final concentration of 105 spores/mL of P. putredinis NO1. Cultures were incubated at 30°C at 150 rpm for 4 days before harvesting for proteomic investigation. The optimized media contained KCl 0.52 g/L, KH2PO4 0.815 g/L, K2HPO4 1.045 g/L, MgSO4 1.35 g/L, NaNO3 1.75 g/L, yeast extract 8.85 g/L, and Hutner’s trace elements and was based on A. niger media (79).
Harvesting the P. putredinis NO1 proteomes across substrates
To harvest supernatant proteins, culture supernatants were centrifuged at 12,000 × g for 20 min at 4°C and were then sterilized through 0.22 µm PES filter units. Triplicate 5 mL technical replicates for each culture were combined with 5 volumes of ice-cold 100% acetone and mixed by inverting before incubation overnight at −20°C. Samples were then centrifuged at 4,500 rpm for 20 min at 4°C and the acetone supernatant discarded. The pellets were then washed twice by the addition of ice-cold 80% acetone, vortexing, and centrifuging at 4,500 rpm for 10 min at 4°C. Pellets were air dried and resuspended in 1 mL of 0.5× PBS buffer before transferring to Eppendorfs and snap freezing in liquid nitrogen and storing at −80°C.
To extract proteins bound to substrates, triplicate samples of 2 g of biomass from each culture were washed twice through the addition of 25 mL ice-cold 0.5× PBS and centrifugation at 4,500 rpm for 5 min at 4°C. The supernatant was discarded, and pellets were resuspended in 19 mL of 0.5× PBS with 1 mL of biotin (EZ-link-Sulfo-NHS-SS-biotin, Thermo Scientific) with rotation at 4°C for 1 h. Samples were then centrifuged at 4,500 rpm for 10 min at 4°C and the supernatant discarded. The reaction was quenched by the addition of 25 mL of 50 mM Tris-HCl pH 8.0 with rotation at 4°C for 30 min. The biomass was pelleted as previously, and the supernatants were discarded. Biomass pellets were washed twice with 20 mL 0.5× PBS and were resuspended in 10 mL of 2% SDS pre-heated to 60°C with rotation at 20°C for 1 h. Samples were centrifuged at 4,500 rpm for 10 min and the supernatant was collected. Five volumes of ice-cold 100% acetone were added, and samples were incubated at −20°C overnight. Samples were pelleted and washed in the same way as before for the supernatant fraction but were resuspended after air drying in 1 mL of 0.1% SDS solution before filtering through 0.22 µm PES filters.
Each replicate was loaded onto its own individual HiTrap Streptavidin HP 1 mL column (GE Healthcare) at a flow rate of 0.5 mL/min. Columns were left to incubate for 1 h at 4°C and were then washed with 10 mL of 0.1% SDS in PBS solution at a flow rate of 1 mL/min. Proteins were eluted by loading 1 mL of 50 mM DTT in PBS solution and incubating columns overnight at 4°C. Another 1 mL of of 50 mM DTT/PBS was added, and 1 mL of protein was eluted, the column was incubated for 1 h at 4°C, and then another 1 mL of 50 mM DTT/PBS was added and another 1 mL of protein collected.
Both supernatant and bound fraction proteins were then desalted by spinning the samples through 5 mL Zeba Spin Columns 7 k MWCO. Desalted samples were then lyophilized overnight and resuspended in NuPAGE loading buffer before being loaded onto NuPAGE gels (Invitrogen). Gels were run for 6 min at 180 V.
Peptide identification by LC–MS/MS
Protein samples in gels were then prepared for and subjected to Label-free LC–MS by the Metabolomics and Proteomics Department in the Technology Facility at the University of York.
In-gel tryptic digestion was performed post-reduction with DTE and S-carbamidomethylation with iodoacetamide. Extracted peptides were loaded onto an mClass nanoflow UPLC system (Waters) equipped with a nanoEaze M/Z Symmetry 100 Å, C 18, 5 µm trap column (180 µm × 20 mm, Waters) and a PepMap, 2 µm, 100 Å, C 18, EasyNano nanocapillary column (75 mm × 500 mm, Thermo). The trap wash solvent was aqueous 0.05% (vol:vol) trifluoroacetic acid and the trapping flow rate was 15 µL/min. The trap was washed for 5 min before switching flow to the capillary column. Separation used gradient elution of two solvents: solvent A, aqueous 0.1% (vol:vol) formic acid; solvent B, acetonitrile containing 0.1% (vol:vol) formic acid. The flow rate for the capillary column was 300 nL/min and the column temperature was 40°C. The linear multi-step gradient profile was: 3%–10% B over 8 min, 10%–35% B over 115 min, 35%–99% B over 30 min, and then proceeded to wash with 99% solvent B for 4 min. The column was returned to initial conditions and re-equilibrated for 15 min before subsequent injections.
The nanoLC system was interfaced with an Orbitrap Fusion Tribrid mass spectrometer (Thermo) with an EasyNano ionization source (Thermo). Positive electrospray ionization mass spectrometry (ESI-MS) and MS 2 spectra were acquired using Xcalibur software (version 4.0, Thermo). Instrument source settings were: ion spray voltage, 1,900–2,100 V; sweep gas, 0 Arb; ion transfer tube temperature, 275°C. MS 1 spectra were acquired in the Orbitrap with 120,000 resolution, scan range: m/z 375–1,500; AGC target, 4e5; max fill time, 100 ms. The data-dependent acquisition was performed in top speed mode using a 1 s cycle, selecting the most intense precursors with charge states >1. Easy-IC was used for internal calibration. Dynamic exclusion was performed for 50 s post precursor selection and a minimum threshold for fragmentation was set at 5e3. MS 2 spectra were acquired in the linear ion trap with scan rate, turbo; quadrupole isolation, 1.6 m/z; activation type, HCD; activation energy: 32%; AGC target, 5e3; first mass, 110 m/z; max fill time, 100 ms. Acquisitions were arranged by Xcalibur to inject ions for all available parallelizable time.
Peak picking, database searching, and quantification of proteomic data from Thermo .raw files were performed using FragPipe (v19.1). Data were searched against a custom database of all coding regions of the P. putredinis NO1 genome appended with common contaminants and reversed sequences. The default LFQ-MBR workflow was used with the following modifications: precursor mass tolerance = ±3 ppm; fragment mass tolerance = 0.5 Da; IonQuant, feature detection m/z tolerance = 3 ppm; MBR RT tolerance = 7.5 min; add MaxLFQ, MBR FDR = 0.01, MBR min ions = 2. Final protein-level data were filtered to 1% FDR, a minimum protein probability of 0.99, and a minimum of two peptides. Molar percentage values were calculated for each protein in each sample as a percentage of the sum of MaxLFQ values for each sample.
Quality control of proteomic data
Identified protein count, principal component analysis, and hierarchical clustering with a Canberra distance matrix and ward.D2 clustering was used to investigate proteomic data for all biological replicates for all substrates investigated. This analysis was performed in R studio 4.2.3 using the “ggplot2,” “FactoMineR,” “factoextra,” “ggdendro,” and “dendextend” packages (80–85). From this analysis, outliers were removed to provide a final data set for comparative analysis.
Isolating the P. putredinis NO1 secretome
Proteins were filtered to produce the P. putredinis NO1 secretome using the “strict” filtering workflow developed previously (21). Secretome proteins were predicted to be extracellular by both BUSCA and DeepLoc localization prediction tools (20, 22), or were predicted to encode a secretion signal by SignalP, TargetP, and SecretomeP tools (23–25), and lacked more than one predicted transmembrane domain by TMHMM or contained a single predicted transmembrane helix with more than 10 amino acids of this helix occurring in the first 60 amino acids of the protein sequence (28).
Comparing the P. putredinis NO1 secretome across substrates
All comparative analysis was performed in R studio v 4.2.3 (80), and analysis was repeated for the total proteome and the filtered secretome.
The number of proteins identified in at least one replicate across all substrates was compared between the bound and supernatant fraction using the “ggVennDiagram” package (86). To visualize differences across substrates, heatmaps were created to compare the average molar percentage of proteins across substrates using the “pheatmap” package (87). Principal component analysis was carried out to investigate replicate grouping within and between substrates using the “FactoMineR” and “factoextra” packages (82, 83). Canberra distance matrix calculation with the ward.D2 clustering was used to plot dendrograms to investigate replicate clustering within and between substrates using the “ggdendro”, and “dendextend” packages (84, 85). GO category and GO term annotation was performed using Blast2GO in the OmicsBox package (88, 89) and abundances were visualized and compared using the “ggplot2” package (81). CAZyme domain annotation was performed using the dbCAN server (43), and abundances were visualized and compared using the “ggplot2” package (81).
Fluorescence-based activity-based protein profiling
Two microliters of triplex probe mixture (60 µM JJB376, CB644, SYF230) were added to 18 µL of secretome without added buffer (measured pH ~7.5). The reactions were incubated for 1 h at 30°C then quenched by the addition of 8 µL of 4× Laemmli buffer (Bio-Rad) and heating to 95°C for 2 min. Ten microliters of the resulting solution were separated over either a 10% SDS-PAGE gel or a 4%–20% gradient gel (Bio-Rad) at 200 V alongside PageRuler prestained protein ladder (Thermo Scientific). The resulting gels were transferred to a Typhoon 5 scanner without fixation and imaged using the Cy2, Cy3, and Cy5 laser/filter sets sequentially. The resulting images were integrated using ImageQuant (GE Healthcare).
Supplementary Material
ACKNOWLEDGMENTS
The York Centre of Excellence in Mass Spectrometry was created thanks to a major capital investment through Science City York, supported by Yorkshire Forward with funds from the Northern Way Initiative, and subsequent support from ESPRC (EP/K039660/1; EP/M028127/1).
This work was funded by the Biotechnology and Biological Sciences Research Council (BBSRC), UK (BB/P027717/1, BB/W000695/1, BB/W003309/1, BB/S01196X/1). C.J.R.S. was supported by a CASE studentship from the BBSRC Doctoral Training Programme (BB/M011151/1) with Prozomix Ltd. G.J.D. is funded by the Royal Society Ken Murray Research Professor and G.J.D./H.S.O. though ERC-2020-SyG-951231 “Carbocentre.”
C.J.R.S. conceptualized the investigation, analyzed the proteomic data, harvested culture supernatants for ABPP, and was a major contributor in writing the manuscript. N.G.S.M. performed the ABPP and imaged the gels, analyzed the ABPP results, and was a major contributor in writing the manuscript. D.R.L. aided the analysis and investigation and was a major contributor in writing the manuscript. N.C.O. helped to conceptualize the investigation and performed the proteomic experiment. A.A. aided in performing the proteomic experiment. A.S. aided in performing the proteomic experiment. A.D. performed the proteomic identification and helped in the analysis of the proteomic data. H.S.O. was a major contributor to the development of the ABPP methodology and contributed to writing. G.J.D. was a contributor to supervision and helped in writing the manuscript. N.C.B. was a major contributor to the conceptualization and supervision of the study in addition to making a major contribution to the writing of the manuscript.
Contributor Information
Conor J. R. Scott, Email: cs1535@york.ac.uk.
Neil C. Bruce, Email: neil.bruce@york.ac.uk.
Sudhir P. Singh, Center of Innovative and Applied Bioprocessing, Mohali, Punjab, India
Vinod Kumar, CSIR-Indian Institute of Integrative Medicine, Jammu, India.
Anmoldeep Randhawa, International Centre for Genetic Engineering and Biotechnology, New Delhi, India.
DATA AVAILABILITY
All proteomic data generated during this research are available at MassIVE MSV000092129.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/spectrum.03943-23.
Investigating the distribution of protein.
Variation in the P. putredinis NO1 secretome.
Clustering of replicates of the P. putredinis NO1 secretome.
Differences in P. putredinis NO1 glycoside hydrolase production.
An accounting of the reviewer comments and feedback.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Saha BC, Qureshi N, Kennedy GJ, Cotta MA. 2016. Biological pretreatment of corn stover with white-rot fungus for improved enzymatic hydrolysis. Int Biodeter Biodegr 109:29–35. doi: 10.1016/j.ibiod.2015.12.020 [DOI] [Google Scholar]
- 2. Salvachúa D, Martínez AT, Tien M, López-Lucendo MF, García F, de Los Ríos V, Martínez MJ, Prieto A. 2013. Differential proteomic analysis of the secretome of Irpex lacteus and other white-rot fungi during wheat straw pretreatment. Biotechnol Biofuels 6:115. doi: 10.1186/1754-6834-6-115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Manavalan T, Manavalan A, Thangavelu KP, Heese K. 2012. Secretome analysis of Ganoderma lucidum cultivated in sugarcane bagasse. J Proteomics 77:298–309. doi: 10.1016/j.jprot.2012.09.004 [DOI] [PubMed] [Google Scholar]
- 4. Cianchetta S, Bregoli L, Galletti S. 2017. Microplate-based evaluation of the sugar yield from giant reed, giant miscanthus and switchgrass after mild chemical pre-treatments and hydrolysis with tailored Trichoderma enzymatic blends. Appl Biochem Biotechnol 183:876–892. doi: 10.1007/s12010-017-2470-z [DOI] [PubMed] [Google Scholar]
- 5. Andlar M, Rezić T, Marđetko N, Kracher D, Ludwig R, Šantek B. 2018. Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation. Eng Life Sci 18:768–778. doi: 10.1002/elsc.201800039 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Ou MS, Ingram LO, Shanmugam KT. 2011. L(+)-lactic acid production from non-food carbohydrates by thermotolerant Bacillus coagulans. J Ind Microbiol Biotechnol 38:599–605. doi: 10.1007/s10295-010-0796-4 [DOI] [PubMed] [Google Scholar]
- 7. Demeke MM, Dietz H, Li Y, Foulquié-Moreno MR, Mutturi S, Deprez S, Den Abt T, Bonini BM, Liden G, Dumortier F, Verplaetse A, Boles E, Thevelein JM. 2013. Development of a D-xylose fermenting and inhibitor tolerant industrial Saccharomyces cerevisiae strain with high performance in lignocellulose hydrolysates using metabolic and evolutionary engineering. Biotechnol Biofuels 6. doi: 10.1186/1754-6834-6-89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Schutyser W, Renders T, Van den Bosch S, Koelewijn S-F, Beckham GT, Sels BF. 2018. Chemicals from lignin: an interplay of lignocellulose fractionation, depolymerisation, and upgrading. Chem Soc Rev 47:852–908. doi: 10.1039/c7cs00566k [DOI] [PubMed] [Google Scholar]
- 9. Ragauskas AJ, Beckham GT, Biddy MJ, Chandra R, Chen F, Davis MF, Davison BH, Dixon RA, Gilna P, Keller M, Langan P, Naskar AK, Saddler JN, Tschaplinski TJ, Tuskan GA, Wyman CE. 2014. Lignin valorization: improving lignin processing in the biorefinery. Science 344:1246843. doi: 10.1126/science.1246843 [DOI] [PubMed] [Google Scholar]
- 10. Salvachúa D, Prieto A, López-Abelairas M, Lu-Chau T, Martínez AT, Martínez MJ. 2011. Fungal pretreatment: an alternative in second-generation ethanol from wheat straw. Bioresour Technol 102:7500–7506. doi: 10.1016/j.biortech.2011.05.027 [DOI] [PubMed] [Google Scholar]
- 11. Karnaouri A, Matsakas L, Topakas E, Rova U, Christakopoulos P. 2016. Development of thermophilic tailor-made enzyme mixtures for the bioconversion of agricultural and forest residues. Front Microbiol 7:177. doi: 10.3389/fmicb.2016.00177 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Pellegrin C, Morin E, Martin FM, Veneault-Fourrey C. 2015. Comparative analysis of secretomes from ectomycorrhizal fungi with an emphasis on small-secreted proteins. Front Microbiol 6:1278. doi: 10.3389/fmicb.2015.01278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Miao Y, Chen X, Li T, Zhu H, Tang S, Liu D, Shen Q. 2020. Proteomic analysis reflects an environmental alkalinization-coupled pH-dependent mechanism of regulating lignocellulases in Trichoderma guizhouense NJAU4742. Biotechnol Biofuels 13:6. doi: 10.1186/s13068-020-1651-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Presley GN, Zhang JW, Schilling JS. 2018. A genomics-informed study of oxalate and cellulase regulation by brown rot wood-degrading fungi. Fungal Genet Biol 112:64–70. doi: 10.1016/j.fgb.2016.08.004 [DOI] [PubMed] [Google Scholar]
- 15. Li F, Ma F, Zhao H, Zhang S, Wang L, Zhang X, Yu H. 2019. A lytic polysaccharide monooxygenase from a white-rot fungus drives the degradation of lignin by a versatile peroxidase. Appl Environ Microbiol 85:e02803-18. doi: 10.1128/AEM.02803-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Kameshwar AKS, Qin W. 2018. Molecular networks of Postia placenta involved in degradation of lignocellulosic biomass revealed from metadata analysis of open access gene expression data. Int J Biol Sci 14:237–252. doi: 10.7150/ijbs.22868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Oates NC, Abood A, Schirmacher AM, Alessi AM, Bird SM, Bennett JP, Leadbeater DR, Li Y, Dowle AA, Liu S, Tymokhin VI, Ralph J, McQueen-Mason SJ, Bruce NC. 2021. A multi-omics approach to lignocellulolytic enzyme discovery reveals a new ligninase activity from Parascedosporium putredinis NO1. Proc Natl Acad Sci U S A 118:e2008888118. doi: 10.1073/pnas.2008888118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Scott CJR, Leadbeater DR, Oates NC, James SR, Newling K, Li Y, McGregor NGS, Bird S, Bruce NC. 2023. Whole genome structural predictions reveal hidden diversity in putative oxidative enzymes of the lignocellulose-degrading ascomycete Parascedosporium putredinis NO1. Microbiol spectr 11:e0103523. doi: 10.1128/spectrum.01035-23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Alessi AM, Bird SM, Oates NC, Li Y, Dowle AA, Novotny EH, deAzevedo ER, Bennett JP, Polikarpov I, Young JPW, McQueen-Mason SJ, Bruce NC. 2018. Defining functional diversity for lignocellulose degradation in a microbial community using multi-omics studies. Biotechnol Biofuels 11:166. doi: 10.1186/s13068-018-1164-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Savojardo C, Martelli PL, Fariselli P, Profiti G, Casadio R. 2018. BUSCA: an integrative web server to predict subcellular localization of proteins. Nucleic Acids Res 46:W459–W466. doi: 10.1093/nar/gky320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Scott CJR, Leadbeater DR, Bruce NC. 2023. A bioinformatic workflow for in silico secretome prediction with the lignocellulose degrading ascomycete fungus Parascedosporium putredinis NO1. Mol Microbiol 120:754–762. doi: 10.1111/mmi.15144 [DOI] [PubMed] [Google Scholar]
- 22. Almagro Armenteros JJ, Sønderby CK, Sønderby SK, Nielsen H, Winther O. 2017. DeepLoc: prediction of protein subcellular localization using deep learning. Bioinformatics 33:3387–3395. doi: 10.1093/bioinformatics/btx431 [DOI] [PubMed] [Google Scholar]
- 23. Teufel F, Almagro Armenteros JJ, Johansen AR, Gíslason MH, Pihl SI, Tsirigos KD, Winther O, Brunak S, von Heijne G, Nielsen H. 2022. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol 40:1023–1025. doi: 10.1038/s41587-021-01156-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Almagro Armenteros JJ, Salvatore M, Emanuelsson O, Winther O, von Heijne G, Elofsson A, Nielsen H. 2019. Detecting sequence signals in targeting peptides using deep learning. Life Sci Alliance 2:e201900429. doi: 10.26508/lsa.201900429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Bendtsen JD, Jensen LJ, Blom N, Von Heijne G, Brunak S. 2004. Feature-based prediction of non-classical and leaderless protein secretion. Protein Eng Des Sel 17:349–356. doi: 10.1093/protein/gzh037 [DOI] [PubMed] [Google Scholar]
- 26. Alfaro M, Castanera R, Lavín JL, Grigoriev IV, Oguiza JA, Ramírez L, Pisabarro AG. 2016. Comparative and transcriptional analysis of the predicted secretome in the lignocellulose-degrading basidiomycete fungus Pleurotus ostreatus. Environ Microbiol 18:4710–4726. doi: 10.1111/1462-2920.13360 [DOI] [PubMed] [Google Scholar]
- 27. de Paula RG, Antoniêto ACC, Nogueira KMV, Ribeiro LFC, Rocha MC, Malavazi I, Almeida F, Silva RN. 2019. Extracellular vesicles carry cellulases in the industrial fungus Trichoderma reesei. Biotechnol Biofuels 12:146. doi: 10.1186/s13068-019-1487-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Krogh A, Larsson B, von Heijne G, Sonnhammer EL. 2001. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580. doi: 10.1006/jmbi.2000.4315 [DOI] [PubMed] [Google Scholar]
- 29. Sudiyani Y, Styarini D, Triwahyuni E, Sembiring KC, Aristiawan Y, Abimanyu H, Han MH. 2013. Utilization of biomass waste empty fruit bunch fiber of palm oil for bioethanol production using pilot–scale unit. Energy Procedia 32:31–38. doi: 10.1016/j.egypro.2013.05.005 [DOI] [Google Scholar]
- 30. Van Hung N, Maguyon-Detras MC, Migo MV, Quilloy R, Balingbing C, Chivenge P. 2020. Rice straw overview: availability, properties, and management practices, p 1–13. In Gummert M, Van Hung N, Chivenge P, Douthwaite B (ed), Sustainable rice straw management. Springer International Publishing, Cham. [Google Scholar]
- 31. Mahmud MA, Anannya FR. 2021. Sugarcane bagasse - A source of cellulosic fiber for diverse applications. Heliyon 7:e07771. doi: 10.1016/j.heliyon.2021.e07771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kapoor M, Panwar D, Kaira GS. 2016. Chapter 3 - Bioprocesses for enzyme production using agro-industrial wastes: technical challenges and commercialization potential, p 61–93. In Dhillon GS, Kaur S (ed), Agro-industrial wastes as feedstock for enzyme production. Academic Press, San Diego. [Google Scholar]
- 33. Peciulyte A, Anasontzis GE, Karlström K, Larsson PT, Olsson L. 2014. Morphology and enzyme production of Trichoderma reesei Rut C-30 are affected by the physical and structural characteristics of cellulosic substrates. Fungal Genet Biol 72:64–72. doi: 10.1016/j.fgb.2014.07.011 [DOI] [PubMed] [Google Scholar]
- 34. Juhász T, Szengyel Z, Réczey K, Siika-Aho M, Viikari L. 2005. Characterization of cellulases and hemicellulases produced by Trichoderma reesei on various carbon sources. Process Biochem 40:3519–3525. doi: 10.1016/j.procbio.2005.03.057 [DOI] [Google Scholar]
- 35. Novy V, Nielsen F, Cullen D, Sabat G, Houtman CJ, Hunt CG. 2021. The characteristics of insoluble softwood substrates affect fungal morphology, secretome composition, and hydrolytic efficiency of enzymes produced by Trichoderma reesei. Biotechnol Biofuels 14:105. doi: 10.1186/s13068-021-01955-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Merali Z, Collins SRA, Elliston A, Wilson DR, Käsper A, Waldron KW. 2015. Characterization of cell wall components of wheat bran following hydrothermal pretreatment and fractionation. Biotechnol Biofuels 8:23. doi: 10.1186/s13068-015-0207-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Crestini C, Lange H, Sette M, Argyropoulos DS. 2017. On the structure of softwood kraft lignin. Green Chem 19:4104–4121. doi: 10.1039/C7GC01812F [DOI] [Google Scholar]
- 38. Novy V, Nielsen F, Seiboth B, Nidetzky B. 2019. The influence of feedstock characteristics on enzyme production in Trichoderma reesei: a review on productivity, gene regulation and secretion profiles. Biotechnol Biofuels 12:238. doi: 10.1186/s13068-019-1571-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Maitz S, Schlemmer W, Hobisch MA, Hobisch J, Kienberger M. 2020. Preparation and characterization of a water‐soluble kraft lignin. Adv Sustain Syst 4. doi: 10.1002/adsu.202000052 [DOI] [Google Scholar]
- 40. Sperschneider J, Williams AH, Hane JK, Singh KB, Taylor JM. 2015. Evaluation of secretion prediction highlights differing approaches needed for oomycete and fungal effectors. Front Plant Sci 6:1168. doi: 10.3389/fpls.2015.01168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Jia L-J, Krüger T, Blango MG, von Eggeling F, Kniemeyer O, Brakhage AA. 2020. Biotinylated surfome profiling identifies potential biomarkers for diagnosis and therapy of Aspergillus fumigatus infection. mSphere 5:e00535-20. doi: 10.1128/mSphere.00535-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. López-García CL, Guerra-Sánchez G, Santoyo-Tepole F, Olicón-Hernández DR. 2024. Chitinase induction in Trichoderma harzianum: a solid-state fermentation approach using shrimp waste and wheat bran/commercial chitin for chitooligosaccharides synthesis. Prep Biochem Biotechnol:1–11. doi: 10.1080/10826068.2024.2313631 [DOI] [PubMed] [Google Scholar]
- 43. Zhang H, Yohe T, Huang L, Entwistle S, Wu P, Yang Z, Busk PK, Xu Y, Yin Y. 2018. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 46:W95–W101. doi: 10.1093/nar/gky418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Kracher D, Scheiblbrandner S, Felice AKG, Breslmayr E, Preims M, Ludwicka K, Haltrich D, Eijsink VGH, Ludwig R. 2016. Extracellular electron transfer systems fuel cellulose oxidative degradation. Science 352:1098–1101. doi: 10.1126/science.aaf3165 [DOI] [PubMed] [Google Scholar]
- 45. Levasseur A, Drula E, Lombard V, Coutinho PM, Henrissat B. 2013. Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol Biofuels 6:41. doi: 10.1186/1754-6834-6-41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Lairson LL, Henrissat B, Davies GJ, Withers SG. 2008. Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem 77:521–555. doi: 10.1146/annurev.biochem.76.061005.092322 [DOI] [PubMed] [Google Scholar]
- 47. Bandi CK, Agrawal A, Chundawat SPS. 2020. Carbohydrate-active enZyme (CAZyme) enabled glycoengineering for a sweeter future. Curr Opin Biotechnol 66:283–291. doi: 10.1016/j.copbio.2020.09.006 [DOI] [PubMed] [Google Scholar]
- 48. King R, Urban M, Lauder RP, Hawkins N, Evans M, Plummer A, Halsey K, Lovegrove A, Hammond-Kosack K, Rudd JJ. 2017. A conserved fungal glycosyltransferase facilitates pathogenesis of plants by enabling hyphal growth on solid surfaces. PLoS Pathog 13:e1006672. doi: 10.1371/journal.ppat.1006672 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Mayans O, Scott M, Connerton I, Gravesen T, Benen J, Visser J, Pickersgill R, Jenkins J. 1997. Two crystal structures of pectin lyase A from Aspergillus reveal a pH driven conformational change and striking divergence in the substrate-binding clefts of pectin and pectate lyases. Structure 5:677–689. doi: 10.1016/s0969-2126(97)00222-0 [DOI] [PubMed] [Google Scholar]
- 50. Amin K, Tranchimand S, Benvegnu T, Abdel-Razzak Z, Chamieh H. 2021. Glycoside hydrolases and glycosyltransferases from hyperthermophilic archaea: insights on their characteristics and applications in biotechnology. Biomolecules 11:1557. doi: 10.3390/biom11111557 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Stricker AR, Mach RL, de Graaff LH. 2008. Regulation of transcription of cellulases- and hemicellulases-encoding genes in Aspergillus niger and Hypocrea jecorina (Trichoderma reesei). Appl Microbiol Biotechnol 78:211–220. doi: 10.1007/s00253-007-1322-0 [DOI] [PubMed] [Google Scholar]
- 52. Sützl L, Laurent C, Abrera AT, Schütz G, Ludwig R, Haltrich D. 2018. Multiplicity of enzymatic functions in the CAZy AA3 family. Appl Microbiol Biotechnol 102:2477–2492. doi: 10.1007/s00253-018-8784-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Haddad Momeni M, Fredslund F, Bissaro B, Raji O, Vuong TV, Meier S, Nielsen TS, Lombard V, Guigliarelli B, Biaso F, Haon M, Grisel S, Henrissat B, Welner DH, Master ER, Berrin J-G, Abou Hachem M. 2021. Discovery of fungal oligosaccharide-oxidising flavo-enzymes with previously unknown substrates, redox-activity profiles and interplay with LPMOs. Nat Commun 12:2132. doi: 10.1038/s41467-021-22372-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Várnai A, Umezawa K, Yoshida M, Eijsink VGH. 2018. The pyrroloquinoline-quinone-dependent pyranose dehydrogenase from Coprinopsis cinerea drives lytic polysaccharide monooxygenase action. Appl Environ Microbiol 84:e00156-18. doi: 10.1128/AEM.00156-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Nekiunaite L, Arntzen MØ, Svensson B, Vaaje-Kolstad G, Abou Hachem M. 2016. Lytic polysaccharide monooxygenases and other oxidative enzymes are abundantly secreted by Aspergillus nidulans grown on different starches. Biotechnol Biofuels 9:187. doi: 10.1186/s13068-016-0604-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Hervé C, Rogowski A, Blake AW, Marcus SE, Gilbert HJ, Knox JP. 2010. Carbohydrate-binding modules promote the enzymatic deconstruction of intact plant cell walls by targeting and proximity effects. Proc Natl Acad Sci U S A 107:15293–15298. doi: 10.1073/pnas.1005732107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Miyauchi S, Navarro D, Grisel S, Chevret D, Berrin JG, Rosso MN. 2017. The integrative omics of white-rot fungus Pycnoporus coccineus reveals co-regulated CAZymes for orchestrated lignocellulose breakdown. PLoS One 12:e0175528. doi: 10.1371/journal.pone.0175528 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Manjeet K, Madhuprakash J, Mormann M, Moerschbacher BM, Podile AR. 2019. A carbohydrate binding module-5 is essential for oxidative cleavage of chitin by a multi-modular lytic polysaccharide monooxygenase from Bacillus thuringiensis serovar kurstaki. Int J Biol Macromol 127:649–656. doi: 10.1016/j.ijbiomac.2019.01.183 [DOI] [PubMed] [Google Scholar]
- 59. Hurtado-Guerrero R, Schüttelkopf AW, Mouyna I, Ibrahim AFM, Shepherd S, Fontaine T, Latgé J-P, van Aalten DMF. 2009. Molecular mechanisms of yeast cell wall glucan remodeling. J Biol Chem 284:8461–8469. doi: 10.1074/jbc.M807990200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Krishnan B, Srivastava SS, Sankeshi V, Garg R, Srivastava S, Sankaranarayanan R, Sharma Y. 2019. βγ-crystallination endows a novel bacterial glycoside hydrolase 64 with Ca2+-dependent activity modulation. J Bacteriol 201:e00392-19. doi: 10.1128/JB.00392-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Znameroski EA, Coradetti ST, Roche CM, Tsai JC, Iavarone AT, Cate JHD, Glass NL. 2012. Induction of lignocellulose-degrading enzymes in Neurospora crassa by cellodextrins. Proc Natl Acad Sci U S A 109:6012–6017. doi: 10.1073/pnas.1118440109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Metz B, Seidl-Seiboth V, Haarmann T, Kopchinskiy A, Lorenz P, Seiboth B, Kubicek CP. 2011. Expression of biomass-degrading enzymes is a major event during conidium development in Trichoderma reesei. Eukaryot Cell 10:1527–1535. doi: 10.1128/EC.05014-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Ishola MM, Taherzadeh MJ. 2014. Effect of fungal and phosphoric acid pretreatment on ethanol production from oil palm empty fruit bunches (OPEFB). Bioresour Technol 165:9–12. doi: 10.1016/j.biortech.2014.02.053 [DOI] [PubMed] [Google Scholar]
- 64. Borrero-López AM, Valencia C, Franco JM. 2022. Lignocellulosic materials for the production of biofuels, biochemicals and biomaterials and applications of lignocellulose-based polyurethanes: a review. Polymers (Basel) 14:881. doi: 10.3390/polym14050881 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Jiang JB, Beenakker TJM, Kallemeijn WW, Marel GA, Elst H, Codee JDC, et al. 2015. Comparing cyclophellitol N-alkyl and N-acyl cyclophellitol aziridines as activity-based glycosidase probes. Chemistry 21:10861–10869. doi: 10.1002/chem.201501313 [DOI] [PubMed] [Google Scholar]
- 66. de Boer C, McGregor NGS, Peterse E, Schröder SP, Florea BI, Jiang J, Reijngoud J, Ram AFJ, van Wezel GP, van der Marel GA, Codée JDC, Overkleeft HS, Davies GJ. 2020. Glycosylated cyclophellitol-derived activity-based probes and inhibitors for cellulases. RSC Chem Biol 1:148–155. doi: 10.1039/d0cb00045k [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Schröder SP, de Boer C, McGregor NGS, Rowland RJ, Moroz O, Blagova E, Reijngoud J, Arentshorst M, Osborn D, Morant MD, Abbate E, Stringer MA, Krogh K, Raich L, Rovira C, Berrin J-G, van Wezel GP, Ram AFJ, Florea BI, van der Marel GA, Codée JDC, Wilson KS, Wu L, Davies GJ, Overkleeft HS. 2019. Dynamic and functional profiling of xylan-degrading enzymes in Aspergillus secretomes using activity-based probes. ACS Cent Sci 5:1067–1078. doi: 10.1021/acscentsci.9b00221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. McGregor NGS, de Boer C, Santos M, Haon M, Navarro D, Schroder S, Berrin J-G, Overkleeft HS, Davies GJ. 2022. Activity-based protein profiling reveals dynamic substrate-specific cellulase secretion by saprotrophic basidiomycetes. Biotechnol Biofuels Bioprod 15:6. doi: 10.1186/s13068-022-02107-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Wu L, Armstrong Z, Schröder SP, de Boer C, Artola M, Aerts JM, Overkleeft HS, Davies GJ. 2019. An overview of activity-based probes for glycosidases. Curr Opin Chem Biol 53:25–36. doi: 10.1016/j.cbpa.2019.05.030 [DOI] [PubMed] [Google Scholar]
- 70. Tõlgo M, Hüttner S, Rugbjerg P, Thuy NT, Thanh VN, Larsbrink J, Olsson L. 2021. Genomic and transcriptomic analysis of the thermophilic lignocellulose-degrading fungus Thielavia terrestris LPH172. Biotechnol Biofuels 14:131. doi: 10.1186/s13068-021-01975-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Oladosu Y, Rafii MY, Abdullah N, Magaji U, Hussin G, Ramli A, Miah G. 2016. Fermentation quality and additives: a case of rice straw silage. Biomed Res Int 2016:7985167. doi: 10.1155/2016/7985167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Saini JK, Saini R, Tewari L. 2015. Lignocellulosic agriculture wastes as biomass feedstocks for second-generation bioethanol production: concepts and recent developments. 3 Biotech 5:337–353. doi: 10.1007/s13205-014-0246-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Toan N-S, Hanh DH, Dong Phuong NT, Thuy PT, Dong PD, Gia NT, Tam LD, Thu TTN, Thanh DTV, Khoo KS, Show PL. 2022. Effects of burning rice straw residue on-field on soil organic carbon pools: environment-friendly approach from a conventional rice paddy in central Viet Nam. Chemosphere 294:133596. doi: 10.1016/j.chemosphere.2022.133596 [DOI] [PubMed] [Google Scholar]
- 74. Singhania RR, Sukumaran RK, Pandey A. 2007. Improved cellulase production by Trichoderma reesei RUT C30 under SSF through process optimization. Appl Biochem Biotechnol 142:60–70. doi: 10.1007/s12010-007-0019-2 [DOI] [PubMed] [Google Scholar]
- 75. Verma N, Kumar V. 2020. Impact of process parameters and plant polysaccharide hydrolysates in cellulase production by Trichoderma reesei and Neurospora crassa under wheat bran based solid state fermentation. Biotechnol Rep (Amst) 25:e00416. doi: 10.1016/j.btre.2019.e00416 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Zhang X, Chen X, Li S, Bello A, Liu J, Gao L, Fan Z, Wang S, Liu L, Ma B, Li H. 2022. Mechanism of differential expression of β-glucosidase genes in functional microbial communities in response to carbon catabolite repression. Biotechnol Biofuels Bioprod 15:3. doi: 10.1186/s13068-021-02101-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Ottenheim C, Verdejo C, Zimmermann W, Wu JC. 2014. Hemicellulase production by Aspergillus niger DSM 26641 in hydrothermal palm oil empty fruit bunch hydrolysate and transcriptome analysis. J Biosci Bioeng 118:696–701. doi: 10.1016/j.jbiosc.2014.05.014 [DOI] [PubMed] [Google Scholar]
- 78. Uke A, Sornyotha S, Baramee S, Tachaapaikoon C, Pason P, Waeonukul R, Ratanakhanokchai K, Kosugi A. 2023. Genomic analysis of Paenibacillus macerans strain I6, which can effectively saccharify oil palm empty fruit bunches under nutrient-free conditions. J Biosci Bioeng 136:1–6. doi: 10.1016/j.jbiosc.2023.03.016 [DOI] [PubMed] [Google Scholar]
- 79. Hutner S, Provosoli L, Schatz A, Haskins C. 1950. Some approaches to the study of the role of metals in the metabolism of microorganisms. Proc Am Philos Soc 94:152–170. doi: 10.2307/3143215 [DOI] [Google Scholar]
- 80. R Development Core Team . 2022. R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. [Google Scholar]
- 81. Villanueva RAM, Chen ZJ. 2019. ggplot2: elegant graphics for data analysis (2nd ed.). Meas Interdiscip Res Perspect 17:160–167. doi: 10.1080/15366367.2019.1565254 [DOI] [Google Scholar]
- 82. Le S, Josse J, Husson F. 2008. FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18. doi: 10.18637/jss.v025.i01 [DOI] [Google Scholar]
- 83. Kassambra A, Mundt F. Factoextra: extract and visualize the results of multivariate data analyses. R package version 1.0.7 ed2020
- 84. de Vries A, Ripley BD. ggdendro: create dendrograms and tree diagrams using 'ggplot2'. R package version 0.1.23 Ed2022
- 85. Galili T. 2015. dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31:3718–3720. doi: 10.1093/bioinformatics/btv428 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Gao C-H, Yu G, Cai P. 2021. ggVennDiagram: an intuitive, easy-to-use, and highly customizable R package to generate Venn Diagram. Front Genet 12:706907. doi: 10.3389/fgene.2021.706907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Kolde R. 2012. Pheatmap: pretty heatmaps. R package version
- 88. Götz S, García-Gómez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talón M, Dopazo J, Conesa A. 2008. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435. doi: 10.1093/nar/gkn176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Bioinformatics B . 2019. OmicsBox - bioinformatics made easy
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Investigating the distribution of protein.
Variation in the P. putredinis NO1 secretome.
Clustering of replicates of the P. putredinis NO1 secretome.
Differences in P. putredinis NO1 glycoside hydrolase production.
An accounting of the reviewer comments and feedback.
Data Availability Statement
All proteomic data generated during this research are available at MassIVE MSV000092129.