Abstract
The phytochrome superfamily comprises three groups of photoreceptors sharing a conserved GAF (cGMP-specific phosphodiesterases, cyanobacterial adenylate cyclases, and formate hydrogen lyase transcription activator FhlA) domain that uses a covalently attached linear tetrapyrrole (bilin) chromophore to sense light. Knotted red/far-red phytochromes are widespread in both bacteria and eukaryotes, but cyanobacteria also contain knotless red/far-red phytochromes and cyanobacteriochromes (CBCRs). Unlike typical phytochromes, CBCRs require only the GAF domain for bilin binding, chromophore ligation, and full, reversible photoconversion. CBCRs can sense a wide range of wavelengths (ca. 330–750 nm) and can regulate phototaxis, second messenger metabolism, and optimization of the cyanobacterial light-harvesting apparatus. However, the origins of CBCRs are not well understood: we do not know when or why CBCRs evolved, or what selective advantages led to retention of early CBCRs in cyanobacterial genomes. In the current work, we use the increasing availability of genomes and metagenome-assembled-genomes from early-branching cyanobacteria to explore the origins of CBCRs. We reaffirm the earliest branches in CBCR evolution. We also show that early-branching cyanobacteria contain late-branching CBCRs, implicating early appearance of CBCRs during cyanobacterial evolution. Moreover, we show that early-branching CBCRs behave as integrators of light and pH, providing a potential unique function for early CBCRs that led to their retention and subsequent diversification. Our results thus provide new insight into the origins of these diverse cyanobacterial photoreceptors.
Keywords: phytochrome, photosynthesis, chromatic acclimation, phycocyanobilin, phototaxis
INTRODUCTION
Phototrophic organisms face the need to carry out photosynthesis under constantly changing light conditions in the presence of competition. These challenges are often met through the use of photoreceptors to evaluate the light environment, generating photochemical signals that are transduced into a range of biological responses. For example, land plants measure the ratio of red light to far-red light using phytochromes [1, 2]. If a plant is shaded by competitors, this ratio will be low due to depletion of photosynthetically active red light and the plant will trigger a shade avoidance response [3]. Phytochromes use a knotted, N-terminal PAS-GAF-PHY photosensory core module containing a covalently linked linear tetrapyrrole (bilin) chromophore [4, 5]. 15,16–photoisomerization of the bilin reversibly photoconverts plant phytochrome between a red-absorbing 15Z dark-adapted state (Pr) and a far-red-absorbing 15E photoproduct (Pfr). This initial photoisomerization triggers a cascade of structural rearrangements starting in the conserved bilin-binding pocket in the GAF domain that can ultimately regulate the expression of thousands of plant genes controlling almost every aspect of plant development [6–10].
Phytochromes are also found in some fungi, eukaryotic algae, and bacteria [11]. Such phytochromes can use a range of different bilins with light sensitivities varying from approximately 600–780 nm [12–14], but all retain the knotted PAS-GAF-PHY photosensor. Phytochromes are common in cyanobacteria, the oldest known oxygenic photosynthetic organisms [15, 16]. Uniquely, the phytochrome superfamily of bilin-binding GAF photoreceptors have diversified in cyanobacteria, giving rise to three distinct families (Fig. S1a). Conventional knotted phytochromes are present, including red/far-red examples such as the model phytochrome Cph1 and unique cyanobacterial variations able to sense blue or even near-UV light [17, 18]. However, cyanobacteria also contain knotless phytochromes lacking the N-terminal PAS domain [19–21]. These GAF-PHY photoreceptors retain photoconversion between Pr and Pfr. However, the more distantly related cyanobacteriochromes (CBCRs) have a minimal photosensor comprising only the bilin-binding GAF domain [22–24].
CBCRs do not exhibit the phytochrome red/far-red photocycle. Instead, they exhibit a diverse range of photocycles between different photostates ranging from 330–750 nm [17, 23, 25, 26]. This photosensory diversity arises from several tuning mechanisms that have evolved during the diversification of CBCRs. For example, red/green CBCRs such as AnPixJg2 and NpR6012g4 typically photoconvert between a red-absorbing 15Z state similar to that of phytochromes and a green-absorbing 15E photoproduct (Fig. S1b) in which the bilin is trapped in a twisted configuration [27–31]. In this case, the 15Z phycocyanobilin (PCB) chromophore adduct is equivalent to that of the knotted phytochrome Cph1 and the knotless phytochrome Cph2, with protons on all four bilin nitrogen atoms and a cationic π system (Fig. S1c; [32, 33]). By contrast, green/red CBCRs such as CcaS and RcaE, familiar regulators of complementary chromatic acclimation (CA; [34–38]), exhibit a reversed protochromic photocycle rather than a trapped-twist mechanism. In such proteins, the 15Z chromophore in green/red CBCRs is deprotonated and absorbs green light [37]. Absorption of a green photon triggers photoisomerization and then proton transfer, resulting in a protonated, red-absorbing 15E photoproduct [37, 39]. These photosensors also belong to distinct CBCR lineages: red/green CBCRs are part of the extended red/green (XRG) clade, which evolved late in the diversification of CBCRs [40, 41], whereas green/red CBCRs belong to the greater green/red (GGR) clade that appeared earlier [40, 42, 43]. However, full-length photoreceptors can contain one or more photosensory domains of different types and lineages. Some cases have tandem arrays of two or more CBCR domains [44–46], but other cases can combine CBCRs with knotless phytochromes [47] or can combine different CBCR domains from distinct lineages with other domains (Fig. S1d).
The evolutionary history of CBCR photoreceptors is thus a complicated interplay between the evolution of cyanobacterial signaling proteins and that of the CBCRs themselves. CBCRs are thought to have arisen from knotless phytochromes via progessive reduction (Fig. S1a). Knotless phytochromes have an intermediate GAF-PHY photosensory module simpler than that of knotted phytochromes but more complex than that of CBCRs, they share a common chromophore precursor with most CBCRs, and they are also exclusively found in cyanobacteria [11]. However, the initial steps and selective pressures underlying this process are not well understood. Put simply, we do not know when or why CBCRs evolved from phytochromes. This transition would have been accompanied by loss of the PHY domain and by changes in a number of conserved residues in close proximity to the chromophore [48, 49]. However, loss of chromophorylation during this process could result in loss of the nascent CBCR rather than its retention in ancient cyanobacterial genomes: although signaling can proceed in phytochromes in the absence of chromophore in other bacteria [50], it is also possible that the loss of photosensory function in such a “proto-CBCR” sequence would result in the loss of any selective advantage.
Studies of CBCR evolution have provided some insight into potential early branches in CBCR evolution. An early phylogenetic analysis identified GGR CBCRs such as RcaE as the earliest branch [40]. More recently, an updated phylogenetic analysis was used in inferring ancestral CBCR sequences, which were shown to exhibit green/red photocycles similar to those of RcaE or CcaS [43]. This work also placed the GGR clade as the second branch in CBCR evolution, with an earlier branch considered a possible artifact due to an anomalously long branch length. Only two sequences belonging to this potential early branch were available at the time of the previous analysis, apparently resulting in incorrect placement of those sequences in the earlier phylogeny. This discrepancy thus highlights a potential pitfall of studying CBCR sequences based on phylogenetic analysis alone, because the placement of such sequences of possible interest might change as new genomic and metagenomic data become available for cyanobacteria. The consensus view that the emergence of DXCF CBCRs [40, 51–53] and subsequent diversification of a range of lineages including the XRG clade [27, 28, 41, 54] followed the appearance of GGR CBCRs is well corroborated by the most current data.
Recent advances in understanding cyanobacterial evolution provide an alternative approach to studying the origins of CBCRs. Until recently, the earliest branch in cyanobacterial evolution was thought to be solely represented by Gloeobacter spp., which lack the thylakoid membranes found in all other oxygenic photosynthetic organisms [55]. Gloeobacter violaceus PCC 7421 has a single CBCR of the DXCF type, which does not seem to be ancestral [40]. All phytochromes and CBCRs are absent in G. kilaueensis [56], so the CBCR found in G. violaceus and related isolates [56–59] may have been acquired by horizontal gene transfer (HGT). This would then indicate that CBCRs arose after the split between Gloeobacterales and all other cyanobacteria. However, it has recently become clear that Gloeobacterales is a more species-rich, diverse lineage than previously thought [60–63]. Studies have identified at least two branches within Gloeobacterales [61, 62]: a branch including several metagenome-assembled genomes (MAGs), with Gloeobacter spp. being derived organisms within that branch, and a second branch including Aurora vandensis, Anthocerotibacter panamensis, and the polar species Sivonenia alaskensis (Fig. S2; [63]). Phylogenetic analysis of 16S rRNA sequences indicates that both branches contain additional diversity [64]. Interestingly, Gloeobacter spp. have apparently lost the GUN4 protein [64], which is present in A. vandensis and A. panamensis and which is thought to have been present in the last common cyanobacterial ancestor (LCCA). The dearth of CBCRs in Gloeobacter spp. could thus reflect a similar loss of ancestral capabilities during the evolution of these organisms. Conserved CBCRs within Gloeobacterales would provide reliable insight into the timing of CBCR evolution by virtue of being from the earliest known branch of cyanobacteria regardless of subsequently appearing genomes. For example, a CBCR lineage shown to have been ancestrally present in both branches of Gloeobacterales would have evolved before those branches split 1.5 billion years ago [62]. Analysis of CBCRs from Gloeobacterales thus would focus on CBCRs from organisms of known interest rather than on CBCR sequences of possible interest.
In the current work, we use the increasing availability of genomic and metagenomic resources for Gloeobacterales and other early-branching cyanobacterial lineages to reexamine the question of CBCR evolution and diversification. Phylogenetic analysis of the CBCR domain confirms the previous identification of the earliest branch in CBCR evolution [43], which we designate as the “earliest extant branch in CBCR evolution as of 2023” (the ee23 CBCRs). The ee23 lineage is followed by the GGR clade. Characterization of heterologously expressed CBCR domains demonstrates that both ee23 and GGR CBCRs can integrate light and pH cues, in contrast to DXCF and XRG CBCRs. Examination of full-length photoreceptor sequences demonstrates that Gloeobacterales contain characteristic lineages of DXCF and XRG CBCRs associated with conserved photoreceptors, including a candidate phototaxis locus. Our results thus provide evidence for early evolution of CBCRs relative to cyanobacterial diversification and for integration of light with pH as a driving force for appearance of early CBCR lineages.
RESULTS
Phylogenetic analysis of the CBCR domain identifies early-branching CBCR lineages.
We began by constructing a sequence alignment which incorporated recently described CBCR lineages [26, 43, 65, 66], CBCR sequences from Gloeobacterales identified by BLAST searches [60–62, 67], and a small number of knotless phytochrome GAF domains to provide an outgroup for phylogenetic analysis. The resulting alignment was used to infer a maximum-likelihood phylogeny (Figs. 1a and S3–S6) as described in the Methods. The CBCR domain is small for this purpose, with only 156 amino acids remaining for phylogenetic inference after removal of gap-enriched columns in our alignment. Nevertheless, the transition from the phytochrome outgroup to early CBCR branches is well resolved, with good support for ee23 CBCRs and GGR CBCRs as the first two branches in CBCR diversification (Figs. 1a & S3). Notably, we identified three early-diverging members of the ee23 CBCR lineage (Figs. 1b & S3), resulting in a much shorter branch length for this CBCR clade than in previous work [43]. Two of these sequences were from Gloeobacterales, with the third (MBD2102707) found in the genome of Leptolyngbya sp. FACHB-261. The latter genome is associated with a recent large-scale study of cyanobacterial evolution and diversification [68], but this strain is not described in this work. We therefore constructed a small catenation of ribosomal proteins and translation elongation factor 4 to provide a tentative placement of this organism with respect to recent advances in cyanobacterial branching order [69, 70]. Phylogenetic analysis using the resulting catenation did not place this organism within Gloeobacterales (Fig. S7). Instead, it was placed as sister to the filamentous genus Pseudanabaena and related organisms, consistent with the definition of Leptolyngbya as a filamentous genus. The ee23 CBCRs were associated with similar domain architectures typically including C-terminal histidine kinase bidomains (Fig. S8), PAS domains, and GAF domains from outside the phytochrome GAF lineage. One protein from Gloeobacterales, MBC8123355, instead combines a histidine kinase with a DXCF CBCR and an ee23 CBCR as a tandem pair. The other exceptional case was MBD2102707, which is predicted to be a much smaller protein lacking a complete histidine kinase. These studies thus provide good support for the ee23 lineage as the earliest known CBCR clade and identify members of this lineage from Gloeobacterales (Figs. 1b & S3).
Figure 1. Phylogenetic analysis of the CBCR domain.

(a) A maximum-likelihood phylogenetic tree is shown as a collapsed view for the CBCR domain, with a small group of GAF domains from knotless phytochromes as an outgroup. Root placement is between the outgroup and all CBCRs. This analysis provides good support for the ee23 clade (“earliest extant as of 2023”) as the earliest known branch in CBCR evolution, followed by the GGR clade. The tree was inferred as described in the Methods, using an alignment of 302 sequences and 156 characters. The complete tree is presented in Figs. S3–S6. Bold italic, lineages from which CBCRs were chosen for characterization in this study. (b) A detailed view is shown for the ee23 lineage. The complete ee23 and GGR clades are presented in context in Fig. S3. Bold, characterized CBCRs. ‡‡, CBCRs characterized in this study.
After the ee23 CBCRs, the next CBCR branch to emerge is the GGR clade (Figs. 1a & S3). GGR CBCRs were not found in Gloeobacterales or in the Thermostichales, the earliest unambiguously placed branch of cyanobacteria known to contain thylakoids [64, 69–72]. This is consistent with the behavior of G. violaceus, which regulates phycoerythrin in response to culture age rather than via CA [73], and with the absence of phycoerythrin biosynthesis genes in Thermostichales. The Aegeococcales are tentatively placed as sister to Thermostichales or as a basal branch within this lineage ([64, 70]; Fig. S2), and the only complete genome for this group, that of Synechococcus sp. PCC 7336, does contain a GGR CBCR (MBD2102481). This protein is not found in an early branch within the GGR CBCRs, but is instead related to RcaE (Fig. S3). This result is thus consistent with a later acquisition of MBD2102481 in Synechococcus sp. PCC 7336 and also with the presence of type III CA in this organism [74].
After GGR CBCRs, several lineages of DXCF CBCRs were recovered as the next branches in CBCR evolution (Figs. 1a & S3–S6). DXCF CBCRs from Gloeobacter spp. (as opposed to Gloeobacterales in general) were recovered in one of these. Consistent with previous work [40], these Gloeobacter representatives are not early-branching members of that early DXCF CBCR lineage (Fig. S3), suggesting their acquisition by more recent HGT. Subsequent DXCF lineages have given rise to other CBCR groups (Figs. S4–S5), such as the DXCIP CBCRs and the NpR3784 group [65, 75]. These observations are consistent with prior studies recovering the DXCF group as a grade that has repeatedly given rise to other lineages [40, 41, 43, 46, 76]. Most of these derived groups were largely absent from Gloeobacterales, excepting a single DXCIP example likely to have been acquired via HGT (Fig. S5). However, one lineage of DXCF CBCRs was almost entirely comprised of sequences from Gloeobacterales (Figs. 1a & S6). The only members of this Gloeo-DXCF CBCR clade not found in Gloeobacterales were found in MAGs from South African stromatolites [77] that have recently been assigned to an early, mesophilic branch of the Thermostichales [64]. Finally, we also observed two small clades of XRG CBCRs that were similarly enriched for Gloeobacterales sequences (Fig. S6). One of these was entirely comprised of red/green CBCRs from Gloeobacterales. The other clade was characterized by a large insertion reminiscent of that seen in insert-Cys (ins-Cys) CBCRs, but having more diversity and only partial conservation of the characteristic Cys residue defining the ins-Cys CBCRs. These “ins-X” sequences were found in the same organisms as Gloeo-DXCF CBCRs: Gloeobacterales and early-branching, mesophilic members of the Thermostichales (Fig. S6). Taken together, our analysis thus identifies multiple clades of CBCRs that are uniquely found in early cyanobacterial branches.
Spectral diversity of early-branching CBCRs.
We next characterized several ee23 CBCRs to learn more about the behavior of this lineage. The only characterized member of this group, Oscil6304_4203 from Oscillatoria acuminata PCC 6304, was previously mis-assigned to a DXCF lineage because only two ee23 CBCR sequences were known at that time [40]. Not an early-diverging member of the ee23 CBCR lineage (Fig. 1b), Oscil6304_4203 was shown to have a green/blue photocycle [40]. We began by examining four of the close relatives of this protein: NIES2119_03185 from Phormidium ambiguum NIES-2119, WP_083263321 from Desertifilum tharense IPPAS B-1220, OSC10802_3032 from Oscillatoria sp. PCC 10802, and MBF2026838 from a hot springs MAG. NIES2119_03185 exhibited a green-absorbing 15Z state with PCB chromophore (Fig. 2a & Table 1); photoconversion yielded a 15E state with blue-absorbing and orange-absorbing populations, similar to the photocycle of Oscil6304_4203 [40]. WP_083263321 exhibited very similar behavior but with lower yield (Fig. 2b & Table 1). By contrast, the other two proteins exhibited red-absorbing 15Z states that gave rise to similar photoproducts (Fig. 2c–d). In these cases, the orange-absorbing species did not contribute significantly to the difference spectrum, resulting in an effective red/blue photocycle. We also examined the two known ee23 CBCRs from Gloeobacterales. MBC8123355g2 is from Gloeobacterales sp. ES-bin-141 (Fig. S2; [61]). This protein exhibited a complex absorption spectrum in the 15Z state, with peaks in the red and yellow regions of the spectrum (Fig. 2e). However, photoconversion of this apparently mixed state gave rise to a simpler photoproduct with only a blue-absorbing peak. By contrast, HGZ86378 from Gloeobacterales sp. SpSt-379 [58, 61] exhibited a red/green photocycle without the blue-absorbing photoproduct population seen in the other examples (Fig. 2f). The three examples with green-absorbing 15Z states thus have very similar difference spectra, differing in the amount of the orange-absorbing 15E population (Fig. 2g), but the three examples with red-absorbing 15Z states have more variation in photoproduct absorption (Fig. 2h). All six ee23 CBCRs examined in this study used PCB adducts as chromophore as judged by acid denaturation (Table 1), consistent with the known properties of Oscil6304_4203 [40]. We conclude that the ee23 CBCRs are a diverse lineage of functional photoreceptors.
Figure 2. Characterization of the early-branching ee23 CBCR lineage.

(a) Absorption and difference spectra are presented for NIES2119_03185. The dashed line indicates zero absorbance. Blue, 15Z state; orange, 15E state; green, (15Z – 15E) photochemical difference spectrum. (b) Absorption and difference spectra are presented for WP_083263321, using the conventions of panel (a). (c) Absorption and difference spectra are presented for OSC10802_3032, using the conventions of panel (a). (d) Absorption and difference spectra are presented for MBF2026838, using the conventions of panel (a). (e) Absorption and difference spectra are presented for MBC8123355g2, using the conventions of panel (a). (f) Absorption and difference spectra are presented for HGZ86378, using the conventions of panel (a). (g) Normalized photochemical difference spectra are shown for NIES2119_03185 (dark blue), WP_083263321 (rose), and Oscil6304_4203 (grey; [40]). (h) Normalized photochemical difference spectra are shown for OSC10802_3032 (forest green), MBF2026838 (bronze), and HGZ86378 (aquamarine). Panels (g) and (h) were normalized by dividing ΔAbsorbance by the maximum observed value. All data taken at pH 7.5.
Table 1:
Spectroscopic parameters for proteins in this study1
| Protein | Variant | SARdenat | 15Z λdenat | 15E λdenat | 15Z λnative | 15E λnative |
|---|---|---|---|---|---|---|
| AnPixJg22 | wild-type | 0.4 | 674 | 580 | 648 | 540 |
| Cri9333_0815 | wild-type | 0.2 | 674 | 580 | 626 | 672 |
| HGZ86378 | wild-type | 0.2 | 672 | 574 | 642 | 522 |
| M595_1144 | wild-type | 0.3 | 674 | 578 | 532 | 668 |
| MBC8123355g2 | wild-type | 0.3 | 676 | 576 | 638, 586 | 450 |
| MBF2026838 | wild-type | 0.6 | 674 | 580 | 652 | 446 |
| Mic7113_1774g1 | wild-type | 0.1 | 674 | 578 | 620 | 678 |
| MP9P1_10_2A | wild-type | 0.6 | 674 | 578 | 424 | 568 |
| MP9P1_10_2B | wild-type | 0.3 | 674 | 578 | 644 | 426 |
| NIES2119_03185 | wild-type | 0.5 | 672 | 578 | 544 | 424, 616 |
| NJL97683g1 | wild-type | 0.4 | 678 | 580 | 440 | 576 |
| NJL97683g2 | wild-type | 0.1 | 674 | 584 | 408 | 564 |
| NpR4776 GAF-PHY3 | wild-type | n/r | n/r | n/r | 644 | 712 |
| NpR4776g1 | wild-type | 0.05 | 678 | 580 | 644 | 694 |
| NpR4776g1-PADCIP | D86C | 0.3 | 676 | 586 | 646 | 534 |
| OSC10802_3032 | wild-type | 0.4 | 674 | 578 | 650 | 442 |
| RfpAg13 | wild-type | n/r | n/r | n/r | 645 | 695 |
| RfpAg1-PAECIP | G79E D80C | 0.6 | 672 | 578 | 646 | 536 |
| WP_006508324 | wild-type | 0.1 | 678 | 582 | 656, 692 | 428, 434 |
| WP_027842869 | wild-type | 0.1 | 674 | 576 | 418 | 568 |
| WP_083263321 | wild-type | 0.4 | 674 | 588 | 554 | 424 |
| WP_218079506g3 | wild-type | 0.7 | 676 | 578 | 428 | 580 |
Reported parameters are SARdenat (specific absorbance ratio under acid denatured conditions), λdenat for both 15Z and 15E photostates (peak wavelengths from the photochemical difference spectrum of the denatured sample), and λnative (peak wavelengths for the native sample at pH 7.5, again from the photochemical difference spectrum). SAR and difference spectra were calculated as described in the Methods. All wavelengths are in nm. n/r not reported.
Preparation reported in [87].
Wavelengths and preparation reported in [21].
Retention of chromophorylation during the initial appearance of CBCRs.
Absorption of blue, violet, or near-UV light by CBCRs proceeds via two-Cys photocycles in which the chromophore is covalently attached to the protein not only by the usual ligation of a conserved ‘canonical’ Cys to the C3 side chain (Fig. S1c) but by another thioether linkage from a second Cys residue to C10 of the chromophore [17, 52, 53, 78–80]. If this linkage is absent in the 15Z state but present in the 15E state, then the green/blue or red/blue photocycles observed with some ee23 CBCRs would result [53, 78]. The sequences of ee23 CBCRs contain both the canonical Cys and a candidate second Cys residue that aligns with an Asp residue that is part of a conserved DIP motif in phytochromes (Fig. 3a; [81, 82]). In ee23 CBCRs, this motif retains the Pro residue, with the Ile matching a conserved hydrophobic residue. The resulting CϕP motif (where ϕ = hydrophobic) is clearly distinct from both the DIP motif and the equivalent EVFP motif of GGR CBCRs (Fig. 3a), and it is largely conserved in the ee23 CBCRs. Notably, the CϕP motif is also conserved in HGZ86378 despite the absence of a blue-absorbing species in this protein (Fig. 2f); however, variant motifs lacking the Cys are present in MBD2102707 and in the late-branching ee23 CBCR MBD2180124 (Figs. 3a & S3).
Figure 3. Characterization of knotless phytochromes mimicking ee23 CBCRs.

(a) The conserved phytochrome DIP motif [81, 82] is compared to the equivalent CϕP (Cys-hydrophobic-Pro) motif of the ee23 CBCRs and the EVFP motif of selected GGR CBCRs. The region around this motif and that around the canonical Cys are shown. A selection of knotless phytochromes are shown, including sequences used in the outgroup (Fig. S3) and characterized examples [19–21]. bold, residues modified by site-directed mutagenesis. Light orange, conserved residues of knotless phytochromes; slate blue, conserved residues of ee23 CBCRs; light green, conserved residues of GGR CBCRs. (b) The domain structure of NpR4776 (Npun_R4776), a complex photoreceptor also containing red/green CBCR domains [28], is shown. Truncations expressing individual photosensory modules are indicated; NpR4776-GAF-PHY has been previously described [21]. (c) Absorption and difference spectra are presented for NpR4776g1, using the conventions of Fig. 2a. (d) Absorption and difference spectra are presented for the D86C variant of NpR4776g1 (NpR4776g1-PADCIP), using the conventions of Fig. 2a. (e) Normalized photochemical difference spectra are shown for wild-type NpR4776g1 (aquamarine), NpR4776g1-PADCIP (brick red), and the NpR4776-GAF-PHY bidomain [21]. (f) Absorption and difference spectra are presented for the isolated GAF domain of knotless phytochrome RfpA, RfpAg1, with G79E D80C substitutions (RfpAg1-PAECIP). Spectra are presented using the conventions of Fig. 2a. (g) Normalized photochemical difference spectra are shown for wild-type RfpAg1 (aquamarine; [21]), and RfpAg1-PAECIP (brick red). (h) Normalized absorption spectra are shown for wild-type OSC10802_3032 (coral) and RfpAg1-PAECIP (brick red) in the light-activated 15E state. Panels (e), (g), and (h) were normalized by dividing Absorbance or ΔAbsorbance by the maximum observed value.
The evolution of the DIP motif into CϕP would thus seem to be one step that occurred during the transition from knotless phytochromes to ee23 CBCRs. However, the Asp residue of the DIP motif is a highly conserved residue in the bilin-binding pockets of both knotted and knotless phytochromes [4, 5, 8, 48], and substitutions for this residue in the model phytochrome Cph1 or for an equivalent Asp in the model red/green CBCR NpR6012g4 can have profound effects on chromophorylation or on the photocycle [83, 84]. We therefore tested the effects of Asp-to-Cys substitutions in GAF-only constructs of knotless phytochromes, mimicking a possible step in early CBCR evolution. We began with NpR4776g1 from Nostoc punctiforme PCC 73102 (Fig. 3b), which is a further truncation relative to the previously characterized GAF-PHY construct exhibiting a normal red/far-red photocycle [21]. Like the GAF1-only construct of Cph2 from Synechocystis sp. PCC 6803 [19], NpR4776g1 exhibited greatly reduced Pfr formation in the absence of the PHY domain (Fig. 3c). Unlike the green-absorbing 15Z state of NIES2119_03185, however, the D86C variant of NpR4776g1 exhibited a red-absorbing 15Z state (Fig. 3d), despite their shared PADCIP motifs (Fig. 3a). Photoconversion of the D86C variant resulted in formation of a bleached 15E photoproduct similar to that of wild-type NpR4776g1, albeit lacking the small amount of 15E far-red absorption present in the GAF-only wild-type (Figs. 3d–e).
We also examined the GAF-only construct of another knotless phytochrome, the FaRLiP regulator RfpA [85]. RfpA has been shown to exhibit efficient Pfr formation even in the absence of the PHY domain [21]. In this case, we constructed a doubly substituted G79E D80C variant protein containing a PAECIP sequence matching that of the red-absorbing OSC10802_3032 (Fig. 3a). This variant protein again exhibited a red-absorbing 15Z state, but Pfr formation was again ablated (Fig. 3f–g & Table 1). Instead, this doubly substituted G79E D80C variant formed an orange-absorbing species similar to one of the two 15E populations seen with OSC10802_3032 (Fig. 3h). We conclude that loss of the conserved Asp of the DIP motif need not result in loss of chromophorylation or of photoconversion in knotless phytochromes, even in the absence of the PHY domain.
Spectral diversity and conserved chromophore configuration in GGR CBCRs.
After ee23 CBCRs, the next branch in our phylogeny is the GGR clade. This group comprises the green/red CBCRs, RcaE and CcaS, as well as far-red-sensing CBCRs like Anacy_2551g3 [26, 35, 37, 42]. Here, we found that M595_1144 from Lyngbya aestuarii exhibits a green/red photocycle (Fig. 4a) despite its placement as an early-branching member of the clade including Anacy_2551g3 and other far-red CBCRs (Fig. S3). This is consistent with an ancestral green/red photocycle within the GGR clade, but we also found previously unappreciated diversity in this clade. Mic7113_1774g1 from Microcoleus sp. PCC 7113 exhibited a red-absorbing 15Z state along with a 15E state similar to those of green/red CBCRs, giving a red/red photocycle (Fig. 4b). Cri933_0815 from Crinalium epipsammum PCC 9333 had multiple peaks in the 15Z state (Fig. 4c), reminiscent of the ee23 CBCR MBC8123355g2. However, photoconversion of this species resulted in a red-absorbing photoproduct and in a photochemical difference spectrum very similar to that of Mic7113_1774g1 (Fig. 4c–d). These results thus underscore similarities between the ee23 and GGR CBCRs, with both lineages having examples with green- or red-absorbing 15Z states using PCB chromophores.
Figure 4. Characterization of additional diversity in the GGR CBCR lineage.

(a) Absorption and difference spectra are presented for M595_1144, using the conventions of Fig. 2a. (b) Absorption and difference spectra are presented for Mic7113_1774g1, using the conventions of Fig. 2a. (c) Absorption and difference spectra are presented for Cri9333_0815, using the conventions of Fig. 2a. (d) Normalized photochemical difference spectra are shown for Mic7113_1774g1 (dark blue) and Cri9333_0815 (rose). Difference spectra were normalized by dividing by the maximum observed value. (e) Circular dichroism (CD) spectra are shown for RfpAg1-PAECIP, using the conventions of Fig. 2a. (f) CD spectra are shown for OSC10802_3032, using the conventions of Fig. 2a. (g) CD spectra are shown for HGZ86378, using the conventions of Fig. 2a. (h) CD spectra are shown for Mic7113_1774g1, using the conventions of Fig. 2a. All data taken at pH 7.5.
We next sought to examine potential conservation of chromophore configurations during the early evolution of CBCRs using CD spectroscopy. Bilin chromophores can be CD active, providing insight into the 3-dimensional disposition of the chromophore about the approximate plane of the B- and C-rings [86–88]. RfpA-PAECIP exhibited negative CD on the red to orange (S0–S1) band in both photostates (Fig. 4e), in contrast to the inversion of CD upon Pfr formation in knotless phytochromes [89] but similar to a range of CBCRs [17, 28, 37, 78]. The same behavior was also seen in the 15Z photostates of five ee23 CBCRs, regardless of whether they exhibited green, red, or mixed peak absorption in the dark-adapted 15Z state (Figs. 4f–g & S9a–c). The orange- or green-absorbing photoproduct populations of these proteins also gave clearly negative CD signals, but signals from the blue-absorbing populations were variable: this population was not resolved in NIES2119_03185 (Fig. S9a), gave weak negative signals in OSC10802_3032 and MBF2026838 (Figs. 4f & S9b), and gave a robust negative signal in MBC8123355g2 (Fig. S9c). The GGR CBCRs Mic7113_1174g1 (Fig 4h), Cri9333_0815 (Fig. S9d), and M595_1144 (Fig. S9e) also exhibited robust negative CD for the long-wavelength bands in both photostates. Taken together, these experiments demonstrate that knotless phytochromes and early CBCR branches exhibit a conserved chromophore configuration in the 15Z state and that the variable spectral tuning mechanisms found in ee23 and GGR CBCRs do not result from gross changes in that configuration.
Integration of light and pH cues by early-branching CBCRs.
The presence of ee23 and GGR CBCRs with either green or red absorption in the 15Z dark-adapted state is reminiscent of the protochromic green/red photocycle of GGR CBCRs such as RcaE, in which changes in protonation state give rise to the changes in peak absorption upon photoconversion [37]. We reasoned that changes in protonation might thus explain the observed variation in dark-adapted states in these CBCR lineages. For example, the green-absorbing ee23 CBCR NIES2119_03185 might have a deprotonated 15Z chromophore like RcaE, whereas the red-absorbing GGR CBCR Mic7113_1774g1 might instead have a protonated 15Z chromophore despite its closer relationship to RcaE (Fig. S3).
We therefore tested this hypothesis for representative ee23 CBCRs. CBCRs dialyzed against standard TKKG buffer at pH 7.5 in the 15Z photostate were diluted into 10 volumes of 400 mM buffers ranging from pH 5 to pH 10 (see Methods). Were the green-absorbing state of NIES2119_03185 to be deprotonated, it would be expected to become protonated at lower pH values and should shift to the red. This was indeed the case, with a red-absorbing species appearing at pH 6 and becoming the majority species at pH 5 (Fig. 5a). Diluting this protein into buffer at higher pH did not change the spectral properties of the green-absorbing state, again consistent with a deprotonated bilin in the green-absorbing state (Fig. 5a). By contrast, the red-absorbing 15Z state of OSC10808_3032 was stable at pH 7; however, this species titrated to a green-absorbing state at higher pH (Fig. 5b). The red-absorbing MBF2026838 behaved similarly, with a stable red-absorbing species still observed at pH 6 (Fig. 5c). Estimated pKa values for these proteins were also consistent with protochromic spectral tuning in these proteins: 15Z NIES2119_03185 had an estimated pKa of 5.7, similar to that of 15Z RcaE [37], whereas OSC10802_3032 and MBF2026838 had higher values of 8.4 and 7.5, respectively (Figs. 5d & S10a–b; Table 2). The two peaks observed in 15Z MBC8123355g2 (Fig. 2d) might thus reflect a mixture of protonated and deprotonated species. Consistent with this interpretation, this protein exhibited increasing red absorption at pH 7 and a simple red-absorbing state at pH 6; however, the same material collapsed to a single green-absorbing state at pH 9 or pH 10 (Fig. 5e). By contrast, HGZ86378 was similar to MBF2026838 (Fig. 5f). MBC8123355g2 exhibited an intermediate pKa of 6.9, whereas HGZ86378 had a higher pKa of 8.1 (Figs. 5g & S10c; Table 2). We favor the simplest interpretation of these results [37]: bilin protonation is an important tuning mechanism for ee23 CBCRs in the dark-adapted state.
Figure 5. Titration of early-branching CBCRs in the 15Z dark-adapted state.

(a) Absorption spectra are shown for NIES2119_03185 after 10-fold dilution into buffers at pH 5 (red), pH 6 (orange), and pH 9 (aquamarine). (b) Absorption spectra are shown for OSC10802_3032 after 10-fold dilution into buffers at pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (c) Absorption spectra are shown for MBF2026838 after 10-fold dilution into buffers at pH 6 (orange), pH 9 (aquamarine), and pH 10 (dark blue). (d) For estimation of pKa values, spectra were normalized on the aromatic amino acid band at ca. 280 nm, and chromophore absorbance was then plotted for different pH values (olive open circles, NIES2119_03185; brick red filled squares, OSC10802_3032). Data were fit under the assumption of a single titratable group using the procedure applied to RcaE [37]. Additional data are in Fig. S10, and pKa values are in Table 2. (e) Absorption spectra are shown for MBC8123355g2 after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 9 (aquamarine), and pH 10 (dark blue). (f) Absorption spectra are shown for HGZ86378 after 10-fold dilution into buffers at pH 6 (orange), pH 9 (aquamarine), and pH 10 (dark blue). (g) Normalized absorption of MBC8123355g2 (mahogany open circles) and HGZ86378 (purple filled squares) was analyzed as in panel (d). (h) Absorption spectra are shown for RfpAg1-PAECIP after 10-fold dilution into buffers at pH 5 (red), pH 6 (orange), pH 9 (aquamarine), and pH 10 (dark blue). (i) Absorption spectra are shown for Mic7113_1774g1 after 10-fold dilution into buffers at pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (j) Absorption spectra are shown for Cri9333_0815 after 10-fold dilution into buffers at pH 7 (khaki) and pH 10 (dark blue), with concentration-corrected data from pH 7.5 shown for comparison (dashed grey). (k) Normalized absorption of Mic7113_1774g1 (mauve filled circles) and Cri9333_0815 (forest green open squares) was analyzed as in panel (d). (l) Absorption spectra are shown for the red/green CBCR AnPixJg2 [27] after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue).
Table 2:
Estimated pKa values for CBCRs
| Protein | photostate | pKa |
|---|---|---|
| NIES2119_03185 | 15Z | 5.7±0.2 |
| OSC10802_3032 | 15Z | 8.4±0.1 |
| MBF2026838 | 15Z | 7.5±0.3 |
| MBC8123355g2 | 15Z | 6.9±0.1 |
| HGZ86378 | 15Z | 8.1±0.2 |
| Mic7113_1774g1 | 15Z | 8.6±0.4 |
| Cri9333_0815 | 15Z | 7.3±0.2 |
| MP9P1_10_2A | 15E | 8.6±0.3 |
| WP_218079506g3 | 15E | 8.4±0.2 |
We next sought to test pH responses in RfpAg1-PAECIP. Unfortunately, this variant knotless phytochrome was unstable outside the range of pH 7–8, as shown by marked scattering in the absorption spectra (Fig. 5h). Scattering continued to increase during illumination, also precluding examination of any pH effects on the 15E photostate. By contrast, the atypical red/red GGR CBCR Mic7113_1774g1 was more stable and exhibited spectral changes at different pH values similar to those of the ee23 CBCR OSC10802_3032 (Fig. 5i), consistent with a protonated 15Z bilin. Like MBC8123355g2, the GGR CBCR Cri9333_0815 behaved as though the 15Z absorption at pH 7.5 arose due to a mix of protonated and deprotonated species (Fig. 5j). However, these two proteins show an important difference in photoconversion: both the protonated and deprotonated states give rise to the 15E blue-absorbing state in MBC8123355g2 (Fig. 2e), whereas the protonated red-absorbing population seems to photoconvert much more efficiently than the deprotonated species in Cri9333_0815 (Fig. 4c–d). The estimated pKa value for Mic7113_1774g1 was similar to that of OSC10802_3032, whereas that for Cri9333_0815 was instead similar to that of MBC8123355g2 (Figs. 5k & S10c; Table 2). It is also important to note that such changes in bilin protonation are not ubiquitous in CBCRs, as shown by the broad stability of the DXCF CBCR Oscil6304_2705 in the 15Z blue-absorbing state ([90]; see Fig. S4 for phylogenetic placement of this protein). We also tested the model red/green CBCR AnPixJg2 [27, 49, 91, 92] in our current procedure (Fig. 5l). Its 15Z red-absorbing state was stable over pH 6–9 and remained the majority species even at pH 10, in marked contrast to the red-absorbing states of ee23 and GGR CBCRs. Hence, ee23 and GGR CBCRs show specific responses to pH change that are not seen in other CBCRs.
We next examined the effects of pH upon photoconversion of ee23 CBCRs. We measured photochemical difference spectra after dilution into different buffers and normalized them using the actual 15Z absorbance observed in the same buffer. This procedure should correct for changes in extinction coefficient that coincide with changes in protonation and should also correct for possible loss of activity. The results are presented in Fig. 6. The green-absorbing ee23 CBCR NIES2119_03185 retained activity even at pH 10, but the blue-absorbing photoproduct did not form at this pH. Instead, higher amounts of the long-wavelength species were formed. By contrast, the blue-absorbing photoproduct population was efficiently formed at pH 5.5 (Fig. 6a). OSC10802_3032 retained a normal red/blue photocycle at pH 8 and showed depletion of both protonated and deprotonated species with red light at pH 9, but photoconversion was lost at pH 10 (Fig. 6b). Photoconversion of MBF2026838 was similar (Fig. 6c), with reduced formation of the blue-absorbing photoproduct at pH 6, formation of a photoactive deprotonated 15Z species at higher pH, and then loss of activity at pH 10. MBC8123355g2 retained a robust red/blue photocycle at pH 6, but photoconversion became less efficient at higher pH as the deprotonated species formed (Fig. 6d). Hence, these three ee23 CBCRs exhibit very similar red-absorbing 15Z states at pH 6 but give rise to different photoproducts (Fig. 6e), whereas there is more variation in the 15Z state at pH 9 but similar formation of a blue-absorbing 15E population (Fig. 6f). Despite these differences, all three of these proteins lost activity at higher pH, with photoconversion essentially lost at pH 10 (Fig. 6b–d). This loss of activity was not observed in the GGR CBCRs Mic7113_1774g1 and Cri9333_0815 (Fig. 6g–h), but in these cases the photoproduct was blue-shifted relative to that observed at pH 7.5 and hence was probably also deprotonated. RcaE exhibited a similar photoproduct blue shift in this experiment (Fig. 6i), as expected [37], and indeed its photochemical difference spectrum at pH 10 was very similar to those of Mic7113_1774g1 and Cri9333_0815 under the same conditions (Fig. 6j). We also examined the ee23 CBCR HGZ86378, which lacks the 15E blue-absorbing species of other members of this lineage. In this protein, the photochemical difference spectrum still exhibited red/green character at pH 9 (Fig. 6k), even though the 15Z chromophore was substantially deprotonated under this condition (Fig. 5f & Table 2). The deprotonated species thus seems less photochemically active in this protein, and indeed photoconversion was reduced at pH 10 (Fig. 6k). Such changes in the photocycle are not ubiquitous in CBCRs, because the photocycle of AnPixJg2 remained remarkably consistent over pH 6–9 and exhibited only a minor loss of activity at pH 10 (Fig. 6l). These results thus establish both ee23 and GGR CBCRs as exhibiting pH-dependent photocycles.
Figure 6. Integration of light and pH sensing in early-branching CBCRs.

(a) Normalized photochemical difference spectra are shown for NIES2119_03185 at pH 5.5 (red), pH 7.5 (dashed grey; see Fig. 2a) and pH 10 (dark blue). (b) Normalized photochemical difference spectra are shown for OSC10802_3032 at pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (c) Normalized photochemical difference spectra are shown for MBF2026838 at pH 6 (orange), pH 7.75 (moss green), pH 8.5 (blue-green), and pH 10 (dark blue). (d) Normalized photochemical difference spectra are shown for MBC8123355g2 at pH 6 (orange), pH 7.75 (moss green), pH 9 (aquamarine), and pH 10 (dark blue). (e) Normalized photochemical difference spectra are shown for MBC8123355g2 (mauve), OSC10802_3032 (forest green), and MBF2026838 (bronze) at pH 6. (f) Normalized photochemical difference spectra are shown for the same proteins at pH 9. (g) Normalized photochemical difference spectra are shown for Mic7113_1774g1 at pH 8 (teal) and pH 10 (dark blue). (h) Normalized photochemical difference spectra are shown for Cri9333_0815 at pH 7 (khaki), pH 8 (teal), and pH 10 (dark blue). (i) Normalized photochemical difference spectra are shown for RcaE at pH 7 (khaki), pH 9 (aquamarine), and pH 10 (dark blue). (j) Normalized photochemical difference spectra are shown for Mic7113_1774g1 (dark blue), Cri9333_0815 (rose), and RcaE (khaki) at pH 10. (k) Normalized photochemical difference spectra are shown for HGZ86378 at pH 6.5 (brown), pH 9 (aquamarine), and pH 10 (dark blue). (l) Normalized photochemical difference spectra are shown for AnPixJg2 at pH 6 (orange), pH 9 (aquamarine), and pH 10 (dark blue). For panels (a)-(d), (g)-(i), and (k)-(l), difference spectra were normalized by dividing the difference spectrum by the peak absorbance measured for the long-wavelength chromophore band at that pH (Fig. 4), yielding the ΔAbsorbance as a fraction of the total to visualize efficiency of photoconversion. For panels (e), (f), and (j), difference spectra were instead divided by the maximum observed value to facilitate comparison of lineshapes.
Late-branching CBCRs from early-branching cyanobacteria.
Integration of light and pH by early CBCRs could provide an advantage relative to conventional red/far-red phytochromes. However, current data for the ee23 and GGR lineages do not provide reliable insight into the timing of CBCR evolution. The ee23 CBCRs currently known from Gloeobacterales (see above) are found in only one of the two branches of these organisms (Figs. 1b & S2–S3), so it is not clear whether these were vertically inherited from the last common ancestor of Gloeobacterales (LCAG) or were instead acquired later in only that branch via HGT. However, the process of constructing a sequence alignment for phylogenetic analysis did make it clear that the paucity of bilin-binding photoreceptors in Gloeobacter spp. is not representative of Gloeobacterales as a whole. As previously reported [25], there is a rough correlation between the number of phytochromes and CBCR open reading frames (ORFs) encoded in a given cyanobacterial genome and the size of that genome. This correlation successfully predicts the absence of such photoreceptors in the reduced genomes of Cyanobium, Prochlorococcus, and related Synechococcus strains (Fig. 7a). However, Gloeobacter spp. are outliers in this analysis: they possess fewer photoreceptors than expected based on their genome size. Plotting the density of phytochrome or CBCR open reading frames per Mb (ORFs/Mb) for different cyanobacterial lineages shows that Gloeobacter spp. are also not representative of Gloeobacterales as a whole, with the latter instead being comparable to other early unicellular branches such as Thermostichales or Gloeomargaritales (Fig. 7b). Given the frequent occurrence of CBCRs in tandem arrays, as in the NpPtxD phototaxis receptor from N. punctiforme ([16, 28, 44]; Fig. S1d), we also calculated a tandem index for these lineages, defined as the average number of predicted bilin-binding GAF domains per open reading frame for a given genome or MAG. This parameter again underscores the difference between Gloeobacter spp. and other members of the Gloeobacterales (Fig. 7c), suggesting that Gloeobacter does not reflect the ancestral CBCR complement of LCAG.
Figure 7. A conserved lineage of DXCF CBCRs in Gloeobacterales.

(a) The number of full-length proteins containing one or more phytochrome and/or CBCR photoreceptor is plotted against assembly size in Mb for diverse cyanobacterial genomes and MAGs (filled blue diamonds) and for those from Prochlorococcaceae ([70]; open red circles) and Gloeobacter spp. (Fig. S7; open violet squares). (b) The photoreceptor density (calculated as phytochrome and/or CBCR open reading frames per Mb, or ORFs/Mb) was calculated for genomes and MAGs from panel (a) and is plotted by lineage. Open blue circles, Gloeobacter spp. (defined as in Fig. S7); filled violet circles, all other Gloeobacterales; open aquamarine triangles, Thermostichales; open green rectangles, Pseudanabaenales; filled dark blue diamonds, Gloeomargaritales; filled brick red squares, more derived cyanobacteria. Prochlorococcaceae lack phytochromes and CBCRs entirely and are omitted. (c) The tandem index (calculated as the average number of phytochrome or CBCR photosensors in a single open reading frame) is plotted as in panel (b). (d) The Gloeo-DXCF CBCR lineage is shown in detail (see Fig. 1a and Fig. S6). Bold, CBCRs characterized in this study. ‡, CBCRs from Thermostichales. (e) Absorption and difference spectra are presented for MP9P1_10_2A, using the conventions of Fig. 2a. (f) Absorption and difference spectra are presented for NJL97683g1, using the conventions of Fig. 2a. (g) Absorption and difference spectra are presented for WP_218079506g3, using the conventions of Fig. 2a.
A CBCR domain vertically inherited from the last common cyanobacterial ancestor (LCCA) or from LCAG should satisfy multiple criteria in a phylogenetic analysis. First, it should be found in a discrete clade. Second, such a domain should be present in both lineages of Gloeobacterales and should mimic the overall pattern of diversification in these organisms (Fig. S2). It is possible for such a sensor to be transferred from Gloeobacterales to other cyanobacteria by HGT, so the presence of non-Gloeobacterales members of such a clade is not disqualifying. Using this reasoning, we identified three CBCR clades that could have been vertically inherited from LCAG in modern Gloeobacterales genomes and MAGs (Figs. 1a & S6). Interestingly, these photosensors do not appear early in CBCR evolution: both the ins-X CBCRs and a small group of red/green CBCRs from Gloeobacterales belong to the late-emerging XRG clade, and the Gloeo-DXCF CBCRs are one of the later DXCF branches to emerge (Fig. S6). Remarkably, these three groups of CBCRs are associated with only two full-length photoreceptors: a histidine kinase with a tandem Gloeo-DXCF:ins-X pair found in both Gloeobacterales and Thermostichales, and a candidate phototaxis receptor specific to Gloeobacterales which had a tandem array of 2–3 Gloeo-DXCF CBCRs followed by 0–2 red/green domains and then by a C-terminal MCP domain (Fig. S11a). Both full-length photoreceptors are present in the two known branches of Gloeobacterales (Fig. S11b), consistent with the presence of both in LCAG. This analysis also identified an additional MCP protein in Gloeobacterales sp. ES-bin-313 that was apparently split across two scaffolds and hence is annotated as two different proteins, despite a region of perfect overlap in the region between the first and second GAF domains (Fig. S11c–d). If one assumes that this overlap reflects a single, authentic photoreceptor, the result is a CBCR-MCP protein matching that observed in the closely related Gloeobacterales sp. SpSt-379 (Fig. S11a; [61]), so we propose that this is a single photoreceptor. Extant Gloeo-DXCF domains seem to have originated as part of the tandem pair and then were transferred to the MCP protein, with subsequent duplications of the domain in the MCP proteins (Fig. 7d). Taken together, these analyses provide evidence for at least two CBCR-based photoreceptors and three types of CBCR domain in LCAG.
We next characterized the properties of Gloeo-DXCF domains. MP9P1_10_2A is the Gloeo-DXCF domain found in a tandem pair with an ins-X CBCR (Fig. S11a) on scaffold 10 of the MAG for A. vandensis MP9P1 [60]. It belongs to the earliest branch of the Gloeo-DXCF proteins (Fig. 7d). This CBCR exhibited a blue-absorbing 15Z state which photoconverted to a yellow-absorbing 15E state, using a PCB adduct (Fig. 7e & Table 1). NJL97683g1 is from a similar photoreceptor but is found in SM2_3_2, a MAG belonging to a mesophilic branch of the Thermostichales [64]. Its properties were very similar to those of MP9P1_10_2A (Fig. 7f & Table 1). Lastly, we examined WP_218079506g3, the third in a tandem array of three Gloeo-DXCF domains in a candidate phototaxis receptor from A. panamensis ([62]; Fig. S11a). This protein again exhibited very similar behavior (Fig. 7g & Table 1). All three proteins also exhibited very similar behavior in CD spectroscopy (Fig. S9f–h). These results thus demonstrate that Gloeo-DXCF domains from different organisms and/or different photoreceptors nevertheless exhibit remarkably similar properties.
In contrast, we found that the photocycles of ins-X CBCRs were more variable. The ins-X group has large insertions roughly corresponding to those seen in ins-Cys CBCRs, some of which have candidate second Cys residues matching those of the ins-Cys group (Fig. 8a). However, the two CBCR lineages are apparently distinct phylogenetically (Fig. S6). We characterized two ins-X CBCRs, both of which are found in tandem pairs with characterized Gloeo-DXCF domains (see above). MP9P1_10_2B from A. vandensis exhibited robust photoconversion between a 15Z red-absorbing dark state and a 15E blue-absorbing photoproduct, using a PCB chromophore (Fig. 8b & Table 1). NJL97683g2 from SM2_3_2, known as a MAG from a South African stromatolite [77], instead exhibited a violet-absorbing 15Z state that gave rise to a broad photoproduct peaking in the green, again with a PCB chromophore (Fig. 8c & Table 1). This variation contrasts not only with the conserved Gloeo-DXCF photocycles from the same full-length photoreceptors (Fig. 7e–f) but also with characterized ins-Cys CBCRs, which can have variable tuning in the 15E state but which exhibit dark-adapted 15Z states ranging from the near-UV to the blue region of the spectrum [17, 25, 76]. However, we observed similar variation in two hybrid CBCRs that were recovered as sister to each other in our phylogenetic analysis (Fig. S6). Like ins-X and ins-Cys CBCRs, hybrid CBCRs belong to the XRG clade [41, 54]. However, they lack the large insertion of the other two groups, instead having reacquired the DXCF Cys residue (Fig. 8a). Hybrid CBCRs have evolved repeatedly within the XRG clade [41]. We found that one such hybrid CBCR domain, WP_027842869 from the euendolithic (rock-boring) cyanobacterium Mastigocoleus testarum BC008, exhibited a photocycle very similar to that of NJL97683g2 (Fig. 8d & Table 1). However, its sister CBCR, WP_06508324 from Xenococcus sp. PCC 7305 (Fig. S6), did not have a similar photocycle. WP_006508324 instead exhibited a red/blue photocycle (Fig. 8e & Table 1), in this case with a small far-red absorbing population consistent with binding a small amount of a verdin chromophore [76, 92, 93]. The ins-X CBCRs thus seem to combine the photosensory variation of hybrid CBCRs with structural adaptations similar to those of the ins-Cys CBCRs. We do not currently understand the reasons for such variable photocycles in relatively closely related proteins, but it is interesting to note that NJL97683g2 and WP_027842869 have similar violet-absorbing dark-adapted states and are from organisms growing in rocky or mineralized locations, whereas MP9P1_10_2B and WP_006508324 do not and were not found in such environments.
Figure 8. Conserved XRG CBCRs in Gloeobacterales.

(a) A subset of the sequence alignment used to infer the CBCR phylogeny is shown. Second Cys residues of DXCF [25, 51, 53, 94, 136], hybrid [41], ins-Cys [17], and ins-X CBCRs are shown, as is the canonical Cys typically used for chromophore attachment [16, 22]. Sequences are shown for selected red/green, hybrid, ins-Cys, and ins-X CBCRs (see Fig. S6 for phylogenetic placement). (b) Absorption and difference spectra are presented for MP9P1_10_2B, using the conventions of Fig. 2a. (c) Absorption and difference spectra are presented for NJL97683g2, using the conventions of Fig. 2a. (d) Absorption and difference spectra are presented for WP_027842869, using the conventions of Fig. 2a. (e) Absorption and difference spectra are presented for WP_06508324, using the conventions of Fig. 2a.
The identification of these stable, well-chromophorylated, late-evolving CBCRs from Gloeobacterales also allowed us to test their ability to integrate light and pH. The 15Z blue-absorbing state of MP9P1_10_2A from A. vandensis was quite robust, exhibiting no significant changes over pH 6–10 (Fig. 9a). Photoconversion revealed changes in the difference spectrum at pH 9–10 (Fig. 9b) which could reflect either loss of activity or a change in extinction coefficient. Absorption spectra taken after illumination with blue light exhibited robust depletion of the 15Z photostate (Fig. 9c), indicating a change in 15E extinction coefficient at high pH. Plotting absorbance at 568 nm versus pH for this photostate (Fig. 9d) suggested that this change arose via a simple titration with a single pKa value of approximately 8.5, which is notably higher than that observed for a 15E PCB adduct under denaturing conditions [37] and is thus likely to reflect a titrating amino acid moiety such as a Cys side-chain. Similar changes were observed in WP_218079506g3 from A. panamensis (Fig. 9e–f, Fig. S10d, & Table 2). We also examined pH responses in the red/blue ins-X CBCR MP9P1_10_2B from A. vandensis. In this case, the red-absorbing 15Z state behaved very similarly to that of AnPixJg2, with nearly identical spectra from pH 6–9 and retention of red absorbance even at pH 10 (Figs. 5l & 9g). Formation of the blue-absorbing 15E photoproduct of MP9P1_10_2B proceeded readily at pH 6–9 but was less efficient at pH 10 (Fig. 9h), again resembling the behavior of the XRG CBCR AnPixJg2 (Fig. 6l). Hence, these late-branching DXCF and XRG CBCRs from Gloeobacterales showed reduced sensitivity at high pH, but they did not change their spectral tuning or lose photoconversion. This behavior thus contrasts with that seen in the early-branching ee23 and GGR CBCRs.
Figure 9. Characterization of pH responses in CBCR lineages from Gloeobacterales.

(a) Absorption spectra are shown for MP9P1_10_2A in the 15Z state after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (b) Normalized photochemical difference spectra are shown for MP9P1_10_2A after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (c) Absorption spectra are shown for MP9P1_10_2A in the 15E state after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (d) Absorbance at 568 nm for MP9P1_10_2A in the 15E state (panel (c)) was normalized by peak absorption on the UV band and plotted as a function of pH. Values were fit to a model describing a single titrating group [37] as described in the Methods, yielding an estimated pKa of 8.5 ± 0.2 (Table 2). (e) Absorption spectra are shown for WP_218079506g3 in the 15Z state after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (f) Normalized photochemical difference spectra are shown for MP9P1_10_2A after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (g) Absorption spectra are shown for MP9P1_10_2B in the 15Z state after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). (h) Normalized photochemical difference spectra are shown for MP9P1_10_2A after 10-fold dilution into buffers at pH 6 (orange), pH 7 (khaki), pH 8 (teal), pH 9 (aquamarine), and pH 10 (dark blue). All difference spectra were normalized as in Fig. 5.
Evolution of CBCR signaling systems in Gloeobacterales.
These studies have demonstrated the presence of functional CBCR domains in Gloeobacterales, with at least two photoreceptors apparently vertically inherited from LCAG (see above). The signaling “output” domains of these two photoreceptors are a histidine kinase bidomain and an MCP domain (Fig. S11a), which are also common C-terminal effector domains for a broad range of other CBCRs [22, 25, 28, 53, 94]. We did not find evidence for CBCR-GGDEF, CBCR-EAL, or CBCR-adenylate cyclase proteins in Gloeobacterales, in contrast to crown cyanobacteria [46, 95–97]. We used phylogenetic analysis to test the interplay between the evolution of these signaling domains and that of the Gloeobacterales CBCR domains. Phylogenetic analysis of histidine kinase bidomains (Figs. 10a & S12) demonstrated that the Gloeo-DXCF:ins-X tandem pairs found in Gloeobacterales were associated with a single clade of His kinases. However, the equivalent tandem pairs from Thermostichales (e.g., NJL97683g1:NJL97683g2) were associated with a distinct His kinase. Similarly, ee23 CBCRs from Gloeobacterales and those from more derived organisms were again associated with distinct His kinase domains (Figs. 10a & S12), indicating that domain shuffling events have played a significant role in the evolution of these photoreceptors.
Figure 10. Phylogenetic analysis of signaling domains from Gloeobacterales CBCRs.
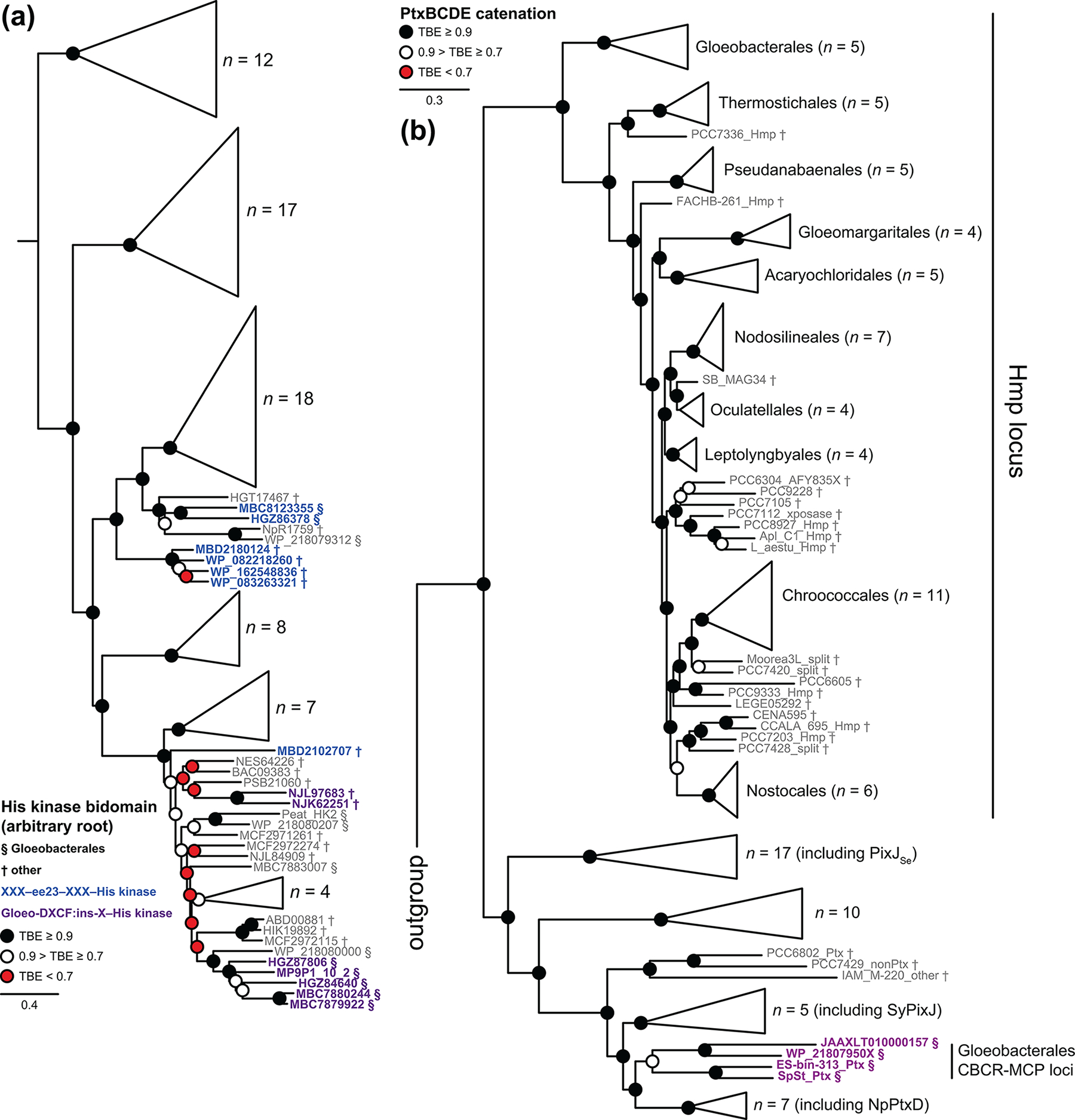
(a) A maximum-likelihood phylogenetic tree is shown as a collapsed view for selected His kinase bidomains. Root placement is arbitrary, and His kinases associated with ee23 CBCRs (blue) or with Gloeo-DXCF:ins-X tandem CBCR pairs (purple) are shown in detail. (b) A maximum-likelihood phylogenetic tree is shown as a collapsed view for a catenation of candidate HmpBCDE/PtxBCDE sequences, using only the MCP region of HmpD/PtxD proteins. PtxBCDE candidates associated with Gloeo-DXCF CBCRs are highlighted in purple, and the clade of candidate Hmp loci is indicated. Sequences from Gloeobacterales (§) and all other cyanobacteria (†) are indicated.
Functional CBCR-based phototaxis receptors typically combine one or more CBCR domains with a C-terminal MCP domain [44, 45, 98], as is the case for those found in Gloeobacterales (Fig. S11a). However, such proteins are also typically found in the context of larger, stereotyped loci that also include several other proteins similar to those found in other bacterial taxis loci [44, 98, 99]. Functional analysis of the Pix locus in Synechocystis sp. 6803, the Ptx locus in N. punctiforme PCC 73102, and the PixJSe photoreceptor in Synechococcus elongatus UTEX 3055 has demonstrated that the SyPixJ, NpPtxD/NpF2164, and PixJSe proteins are functional photoreceptors in vivo [44, 45, 98, 100], and biochemical characterization has established their spectroscopic properties [17, 28, 45, 101, 102]. However, a second taxis locus is required for general motility in Synechocystis and N. punctiforme [44, 99, 100]. This Hmp locus again has an MCP protein and homologs for PatA, CheY, CheW, and CheA proteins, but the MCP protein does not have CBCR domains. Motility in cyanobacteria requires type IV pili, so a large “parts list” of proteins for biogenesis of these pili is required for motility in addition to the taxis loci [99, 103, 104]. Several of these proteins have recently been shown to be vertically inherited from LCCA [64]. BLAST searches of Gloeobacterales genomes and MAGs detected apparent orthologs for each of these proteins, the proteins of the Hmp locus, and the proteins of the Pix/Ptx locus in at least three organisms spanning both branches of Gloeobacterales (Figs. S2 & S13). The absence of some required proteins in MAGs is not conclusive, but complete genomes of Gloeobacter spp. are missing both the Hmp locus and the phototaxis locus (Fig. S13). The absence of these loci is consistent with the absence of motility in G. violaceus [55] and raises the possibility that motility may have been present in LCAG but lost in Gloeobacter spp. at a later date.
For phylogenetic analysis of the MCP proteins, we used a catenation of PtxB/HmpB/CheY, PtxC/HmpC/CheW, the MCP domain, and PtxE/HmpE/CheA proteins. Unlike other components of the Hmp locus, the hmpA gene (PatA homolog) is not required for motility in Synechocystis and N. punctiforme [99, 103, 104], so this protein and its paralogs from other loci were omitted. Phylogenetic analysis of the resulting catenation (Figs. 10b & S14) clearly resolved a Hmp clade whose branching pattern closely matches that seen for catenated cyanobacterial translation proteins (Fig. S7), supporting vertical inheritance of the Hmp locus from LCCA. Other taxis loci were recovered in several clades. The candidate phototaxis loci from Gloeobacterales were recovered as part of a derived clade also including the known phototaxis loci from Synechocystis and N. punctiforme, with that from S. elongatus being only distantly related (Fig. 10b). Taken together, this analysis supports motility as a vertically inherited, ancestral characteristic of cyanobacteria but favors a more complex scenario for evolution of phototaxis.
DISCUSSION
In the current work, we have leveraged the increasing availability of genomic and metagenomic resources for early-branching cyanobacteria to examine the evolution of CBCRs. We confirm the identity of the earliest CBCR branches and identify conserved CBCR photoreceptors in Gloeobacterales. We also characterize both early-branching CBCRs and CBCRs from early-branching cyanobacteria. Our results provide new insight into the timing of CBCR evolution and into potential factors underlying the successful diversification of CBCRs.
CBCRs evolved early in cyanobacterial evolution.
We identified three clades of CBCRs that were apparently present in the last common ancestor of Gloeobacterales (LCAG). These CBCR clades are not early-branching photoreceptors. Indeed, two of them arose within the XRG CBCRs, which are well-established as a derived branch that arose late in CBCR evolution [17, 40, 41, 94]. However, these CBCR groups were present in Gloeobacterales before the divergence of the two branches within this cyanobacterial clade (Fig. S2), which has been estimated to have occurred 1.5 billion years ago [62]. Therefore, the XRG CBCRs must have arisen and diversified into different clades before this time, requiring the initial emergence of CBCRs to be earlier still. The presence of Gloeo-DXCF:ins-X tandem pairs in both Gloeobacterales and Thermostichales can be explained multiple ways based on the current data. This photosensory pair could have been acquired by Thermostichales via HGT from Gloeobacterales, or it could have been present in the last common ancestor of these two taxa. However, Gloeobacterales are the earliest branch in cyanobacterial evolution, so the last common ancestor of Gloeobacterales and Thermostichales is also LCCA. Hence, it is possible that Gloeo-DXCF:ins-X tandem pairs were present in LCCA, which would imply that the initial evolution of CBCRs and much of their subsequent diversification took place very early in cyanobacterial evolution, at least 2 billion years ago [62].
Our examination of taxis loci is also consistent with this conjecture. The Hmp locus is recovered as a single, vertically inherited clade (Figs. 10B & S14), implying motility in LCCA. However, functional phototaxis loci with CBCR-MCP proteins do not form a single clade, much less a vertically inherited one. The observed pattern is instead consistent with repeated duplications of taxis loci and with the appearance of CBCR-MCP proteins in those loci multiple times via some type of domain shuffling. The Gloeo-DXCF domain provides an example of this process: it first appeared as a tandem pair with ins-X CBCRs and C-terminal His kinases, and then was transferred to CBCR-MCP proteins (Figs. 7d & S11a–b). It then duplicated in those MCP proteins to form tandem arrays, as is also the case for PixJSe [45]. Were the taxis loci to have undergone repeated duplications and shuffling prior to the split between Gloeobacterales and other cyanobacteria, then such loci could have been present in LCCA and the current situation would represent billions of years of duplication and deletion as well as vertical inheritance. Such a scenario would also explain the absence of GGR CBCRs and other lineages in Gloeobacterales: they have simply been lost. Taken together, our work provides new information on the timing of CBCR evolution but not complete understanding of this process, and we expect new insights to emerge as additional genomes and MAGs become available.
Spectral tuning and the transition from phytochromes to early CBCRs.
Our work also provides new insight into the behavior of the earliest known CBCR branches, the ee23 and GGR clades. Both clades exhibit considerable diversity in spectral tuning, so we cannot identify an ancestral CBCR photocycle with confidence based on our own work. The green/red photocycles observed in inferred ancestral sequences [43] are not found in ee23 CBCRs characterized to date, but we cannot rule out that such CBCRs will be found in the future. The hypothetical ancestral sequences inferred in such analysis could also be changed by inclusion of the ee23 CBCRs from Gloeobacterales. However, it is clear that chromophore protonation plays a key role in the photocycles of both the ee23 and GGR CBCRs. Deprotonated bilins give rise to green-absorbing 15Z photostates in both lineages, as typified by NIES2119_03185 (Fig. 2a) and RcaE [37]. However, we have now shown that both lineages also contain examples of red-absorbing 15Z states with protonated bilins, such as Mic7113_1774g1 and OSC10802_3032 (Figs. 2b & 4b), along with the mixed cases MBC8123355g2 and Cri9333_0815 (Figs. 2e & 4c).
The photoproducts of ee23 CBCRs also show considerable variation at a “standard” pH of 7.5. Several such proteins give rise to a mix of blue- and orange-absorbing species, but this is not universal (Fig. 2). MBC8123355g2 from Gloeobacterales sp. ES-bin-141 forms the blue-absorbing species much more efficiently, whereas HGZ86378 from Gloeobacterales sp. SpSt-379 does not form this species at all and instead exhibits a red/green photocycle. Interestingly, formation of the blue-absorbing photoproduct is sensitive to pH in ee23 CBCRs, but not in a conserved pattern (Fig. 6): NIES2119_03185 fails to form this species at high pH, whereas OSC10802_3032 and MBF2026838 instead fail to form this species at low pH. In phytochromes and CBCRs, such blue-absorbing species have been shown to arise from nucleophilic addition of a second Cys residue to C10 of the bilin chromophore [17, 52, 66, 78–80, 105], and the CϕP motif of ee23 CBCRs contains a candidate Cys residue (Fig. 3a). Variation in the photoproduct tuning of ee23 CBCRs could thus be interpreted in the context of such a reaction. The loss of the blue-absorbing species at low pH in OSC10802_3032 and MBF2026838 would arise due to protonation of the Cys residue, making it less nucleophilic. The loss of this species at high pH in NIES2119_03185 would instead arise to deprotonation of the 15E bilin, making it less electrophilic. The absence of this species in HGZ86378 could arise because its green-absorbing photoproduct is trapped in a conformation that is not well positioned relative to the nucleophile, resulting in the observed red/green photocycle. Future studies are thus likely to elucidate the protein-chromophore interactions determining this variation and permit its engineering, as has proven the case for different properties other CBCR lineages [17, 37, 41, 51–53, 65, 84, 87, 91, 92, 105–110].
Our work also demonstrates that the initial transition from knotless phytochromes to CBCRs may not have involved a loss of photosensory function. Knotless phytochromes are able to bind chromophore and carry out photoisomerization as isolated GAF domains, like CBCRs (Fig. 3 & Table 1). In the current work, we show that one of the key amino acid substitutions required to evolve from a knotless phytochrome into an ee23 CBCR does not prevent either process but does ablate Pfr formation. Moreover, such variant phytochromes exhibit CD signatures similar to those of CBCRs rather than those of the parent phytochromes able to make Pfr (Fig. 4e–h & Fig. S9a–e). Thus, it is possible that ancestrally red-absorbing knotless phytochromes gave rise to ancestral CBCRs retaining red absorbance in the 15Z state, as observed in ee23 proteins from Gloeobacterales and in the GGR CBCR Mic7113_1774g1. Retention of photosensory function could have provided a selective advantage during this process and hence could have provided a selective pressure to retain such transitional photoreceptors in ancient cyanobacterial genomes.
What advantages did early CBCRs offer to cyanobacteria?
CBCRs from the ee23 and GGR lineages show major changes in spectral tuning and/or photoconversion at high pH (Figs. 5–6). Other CBCRs from Gloeobacterales show much more minor effects (Fig. 8), as do the model red/green CBCR AnPixJg2 (Figs. 5–6), the DXCF CBCR Oscil6304_2705 [90], and the model cyanobacterial phytochrome Cph1 [111]. Therefore, the pH responses of ee23 and GGR CBCRs do not arise from some fundamental limitation of bilin-based photoreceptors. We propose that this behavior instead reflects an ancient function as integrative sensory proteins that detect both the pH and the light environment.
We note that there are subtle differences between the red-absorbing states observed in some ee23 and GGR CBCRs (Figs. 2 & 4; Table 1). We speculate that those differences may be related to the spectral properties of cyanobacterial light-harvesting pigments, allowing optimization of light-harvesting strategies in the presence of competing anoxygenic photosynthetic organisms. Cyanobacterial cells exhibit absorption peaks at ca. 680 and 440 nm due to chlorophyll a (Chl a) and ca. 620 nm due to phycocyanin (PC), as shown in Fig. 11a for Synechocystis cells [112]. Both phycobiliproteins and PCB biosynthesis were present in LCCA [88], so its spectral features are likely to have been broadly similar. Cyanobacteria also contain a smaller peak at ca. 650–660 nm due to allophycocyanin (APC; Fig. 11b). Some cyanobacteria also harvest green light via phycoerythrin (PE) or phycoerythrocyanin (PEC; [113]), resulting in a peak at 550–570 nm (Fig. 11a; [36]). Before oxygenation of the atmosphere, cyanobacteria would have competed for light energy with anoxygenic photosynthetic bacteria and with each other. Anoxygenic photosynthesis relies on bacteriochlorophyll (Bchl). The simplest competing case could perhaps be viewed as the ancient homodimeric reaction center of heliobacteria such as Heliobacterium modesticaldum, which contains Bchl g and 81-OH-Chl aF [114] and which is not associated with a peripheral light-harvesting complex [115]. The more complex green and purple photosynthetic bacteria employ distinct light-harvesting strategies relying on Bchl a or Bchl c, respectively [116], resulting in different spectral properties for such cells [117, 118]. Comparison of these competitor spectra with that of the modern cyanobacteria Synechocystis sp. PCC 6803 and N. punctiforme (Fig. 11a; [112]) shows that there are overlaps, such as the peaks at 550–600 nm in H. modesticaldum and R palustris that overlap with PE and the minor peak in H. modesticaldum at 670 nm that overlaps with Chl a. However, there is also a window from 600–650 nm (red and orange light) in which cyanobacteria enjoy little competition from such competitors. CBCR-based optimization of light harvesting pigments via CA is well established for PE and PEC in modern cyanobacteria [34, 38], allowing efficient usage of green light that is not utilized by modern competitors such as land plants. It thus does not seem implausible that early cyanobacteria could have used similar processes to optimize usage of red and orange light that is not utilized by anoxygenic photosynthetic bacteria.
Figure 11. Early-branching CBCR lineages and cyanobacterial light-harvesting pigments.

(a) Approximate absorption spectra are shown for cells of Synechocystis sp. PCC 6803 ([112]; solid cyan), Nostoc punctiforme grown under green light ([36]; solid brown), and Chlorobium tepidum ([118]; dashed dark green), for intracytoplasmic membranes of Rhodopseudomonas palustris ([117]; dashed purple), and for the anoxygenic, homodimeric reaction center of Heliobacterium modesticaldum ([114]; dashed grey), with wavelength ranges and perceived colors indicated underneath. (b) Approximate peak wavelengths are shown for major cyanobacterial light-harvesting pigments [112, 113]: chlorophyll a (Chl a; 440 and 680 nm), allophycocyanin (APC; 650 nm), phycocyanin (PC; 620 nm); phycoerythrin (PE; 564 nm), and phycoerythrocyanin (PEC; 568 nm). (c) Cartoon red/far-red photocycles are shown for representative cyanobacterial phytochromes using PCB (Fig. S1c) or biliverdin IXα (BV). (d) Cartoon photocycles are shown for selected CBCRs from the ee23 and GGR lineages.
In contrast, cyanobacterial phytochromes are not well suited for the regulation of light harvesting in under-utilized regions of the EM spectrum prior to the genesis of oxygenic photosynthesis. Cyanobacterial phytochromes can use either biliverdin IXα (BV) or PCB as the physiological chromophore precursor [119, 120]. Those using BV are quite similar to bacteriophytochromes from anoxygenic photosynthetic organisms [119–121] and overlap with some Bchl absorption bands, whereas those using PCB have Pr states approximately matching APC and Pfr states that also overlap Bchl bands (Fig. 11a & c). Such phytochromes can provide a means for modern cyanobacteria to detect far-red light for use in the FaRLiP response [21], but they could also provide a means to detect anoxygenic competitors that would specifically deplete far-red light. However, they are not well suited to measure the red and orange region matching PC.
By contrast, the photocycles of some ee23 and GGR CBCRs are very well suited to this task (Table 1). The red/blue difference spectra of OSC10802_3032 and MBF2026838 are excellent matches for APC and for the high-energy Soret peak of Chl a (Figs. 2b–c, 11a–b, & 11d). Blue light can be an important energy source in diverse light environments [122, 123], but cyanobacteria are intrinsically inefficient at using blue light for photosynthesis [112, 124] and would face competition in this spectral window (Fig. 11a). The ability to detect the presence of an alternative energy source could thus have been advantageous for ancient cyanobacteria. Similarly, the red/red difference spectra of Mic7113_1774g1 and Cri9333_0815 (Fig. 3d) are excellent matches for PC and for the low-energy Chl a peak (Fig. 11b & d). The photocycles of these early-branching CBCRs thus provide a means for optimizing the ratios of these three pigments at pH 7.5 to pH 8, matching the normal physiological range for cyanobacterial cytosol during active photosynthesis [125–127]. This is also consistent with the known benefits of and photobiological responses to orange light in modern cyanobacteria [124, 128, 129].
However, at higher pH these photoreceptors all shift to the green region of the spectrum, with some of them also losing photoconversion (Figs. 5–6). This behavior could provide a self-regulating mechanism for cells that have abnormally high intracellular pH, which might have been particularly advantageous for ancient cyanobacteria having more rudimentary mechanisms for pH homeostasis. Indeed, transient changes in pH accompanying changes in light fluence are thought to be an important physiological cue for regulation of photosynthesis in the modern chloroplast [130]. In this regard, it is interesting to note that an ee23 CBCR from Gloeobacterales also exhibits a red/blue photocycle at lower pH (Figs. 2d & 5d), which may reflect the reduced capacity of such organisms to regulate intracellular pH in the absence of thylakoids [125]. Although we cannot hope to observe the function of ancestral CBCRs in ancient cyanobacteria, it seems possible that such photoreceptors could have provided such organisms with a competitive advantage relative to anoxygenic photosynthetic organisms and to other cyanobacteria lacking such mechanisms. This selective advantage could have led to the retention of CBCRs in ancient genomes, permitting their subsequent diversification into the sweeping palette of photosensors seen today.
MATERIALS AND METHODS
Bioinformatics.
Multiple sequence alignments were constructed in MAFFT v7.450 [131] using the command-line settings --genafpair --maxiterate 16 --clustalout –reorder. For maximum-likelihood phylogenetic analysis, the resulting alignment was processed with an in-house script to remove positions having ≥5% gaps. Phylogenies were inferred with PhyML-3.1 [132] with 100 bootstraps, using the command-line settings -m WAG -d aa -s SPR -a e -c 4 -v e -o tlr -b 100. Statistical robustness was assessed using the transfer bootstrap expectation (TBE) as implemented in booster [133]. Completeness of sequences after gap removal was assessed using the —length option of alnfilter, a utility available as part of the homolmapper distribution [134]. The final sequence alignment used for phylogenetic analysis of CBCRs had 302 sequences and 156 characters, with root placement between the knotless phytochrome outgroup and all CBCRs. The final alignment used for histidine kinase bidomains had 96 sequences and 216 characters, with arbitrary root placement. In this case, two sequences were less than 90% complete by length in the final tree input files and are indicated as such in Fig. S12. The catenation for translation proteins (elongation factor 4 plus ribosomal proteins) had 96 sequences and 1282 characters; that for taxis proteins (HmpBCDE/PtxBCDE) had 125 sequences and 1346 characters. Root placement for both was between the outgroup of non-cyanobacterial sequences and all cyanobacterial sequences. The presence or absence of Pix/Ptx and Hmp loci was assessed using the protein sequences encoded by the characterized loci from N. punctiforme [44, 99] as queries in BLAST searches. The presence or absence of PilM, PilN, and the two PilT isoforms PilT1 and PilT2 in Gloeobacterales and Thermostichales has recently been evaluated [64].
Protein expression and characterization.
Expression and purification of AnPixJg2, RcaE, and RfpAg1 followed published procedures for these constructs [21, 37, 87]. Expression and purification of NpR4776g1 followed procedures previously published for the GAF-PHY construct [21]. Other CBCRs were obtained via commercial gene synthesis as His-tagged expression clones in pET28a (Genscript; Piscataway, New Jersey). Expression and purification of these CBCRs followed our recently published procedure [87] and used pKT271 [135] to supply PCB biosynthesis enzymes, with the variation that final dialysis was into TKKG buffer (25 mM TES-KOH, 100 mM KCl, 10% (v/v) glycerol) at pH 7.5 rather than pH 7.8. This buffer was also used to dilute samples for spectroscopic characterization as needed. For pH responses, 0.1 ml of CBCR solution in TKKG buffer was added to 1 ml of 0.4 M buffer (pH 5, sodium acetate; pH 6, MES; pH 6.5, PIPES; pH 7–8, TES; pH 8.5, Tricine; pH 9–10, CHES). Other pH values were generated by combining 0.5 ml each of two of these buffers (pH 5.5, pH 5 + pH 6; pH 7.75, pH 7 + pH 8.5). For denaturation analysis, 0.1 ml of CBCR solution in TKKG buffer was added to 1 ml of 6M guanidinium chloride/1% HCl (v/v). Denaturation analysis was used to assign 15Z and 15E configurations [26, 52], and all difference spectra are reported as (15Z – 15E). Both experiments included buffer blanks at each condition. All absorption spectra were acquired using Cary50 or Cary60 spectrophotometers at 25°C (250–900 nm, sampled every 2 nm for 0.125 s). CD spectra were acquired using an Applied Photophysics Chirascan at 25°C (250–850 nm).
Analysis of spectroscopic data.
Estimation of pKa values used a modification of the fitting procedure previously used for RcaE [37] under the assumption that the observed changes arose due to a single titrating group. Spectra at each pH value (including the buffer blank) were rezeroed using the mean observed absorbance at 850–900 nm, and the buffer was then subtracted. Each spectrum was then normalized using the peak absorbance on the aromatic amino acid band (250–300 nm) to correct for possible differences in dilution factor, and normalized absorbance values were used for nonlinear regression analysis to estimate pKa values. Photochemical difference spectra at different pH conditions were divided by the peak bilin absorbance on the long-wavelength (S0–S1) band at that pH (ΔAbs/Abs). This normalization condition will detect changes in extinction coefficient or loss of activity. Other difference spectra were divided by the maximum value of the difference spectrum itself to facilitate comparison of lineshapes.
Supplementary Material
HIGHLIGHTS.
Cyanobacteriochromes (CBCRs) are photoreceptors related to phytochromes
CBCRs from early-branching cyanobacteria identify early-branching CBCR lineages
Early-branching CBCRs integrate light and pH sensing
Early-branching cyanobacteria harbor late-evolving CBCRs
CBCR signaling systems appeared early in cyanobacterial evolution
ACKNOWLEDGEMENTS
We thank Ms. Shelley S. Martin for assistance with expression and purification of CBCR proteins. This work was supported by a grant from the Chemical Sciences, Geosciences, and Biosciences Division, Office of Basic Energy Sciences, Office of Science, U.S. Department of Energy (DOE DE-FG02–09ER16117 to J.C.L.). Characterization of CBCRs from Gloeobacterales, pH studies and phylogenetic analysis were supported by NIH grant 5R35GM139598–02 to J.C.L. The contents of this work are solely the responsibility of the authors and do not necessarily represent the official views of the NIGMS, NIH or DOE.
ABBREVIATIONS
- ΔAbs
change in absorbance
- APC
allophycocyanin
- Bchl
bacteriochlorophyll
- BV
biliverdin IXα
- CBCR
cyanobacteriochrome N-cyclohexyl-2-aminoethanesulfonic acid
- Chl a
chlorophyll a
- DXCF
Asp-Xaa-Cys-Phe motif defining a CBCR lineage
- DXCI
Asp-Xaa-Cys-Ile motif defining other CBCR lineages
- ee23
earliest extant as of 2023 (defining a CBCR clade)
- DXCIP
Asp-Xaa-Cys-Ile-Pro motif defining a CBCR lineage
- EAL
cyclic-di-GMP phosphodiesterase named after a sequence motif
- GAF
domain acronym derived from vertebrate cGMP-specific phosphodiesterases, cyanobacterial adenylate cyclases, and formate hydrogen lyase transcription activator FhlA
- GGDEF
cyclic-di-GMP synthase domain named after a sequence motif
- GGR
greater green/red (designating a CBCR clade)
- Gloeo-DXCF
a lineage of DXCF CBCRs almost exclusively found in Gloeobacterales
- HGT
horizontal gene transfer
- ins-Cys
insert-Cys (defining a CBCR lineage)
- ins-X
a distinct CBCR lineage with a similar insertion
- LCAG
last common ancestor of Gloeobacterales
- LCCA
last common cyanobacterial ancestor
- MAG
metagenome-assembled genome
- ORF
open reading frame
- PC
phycocyanin
- PE
phycoerythrin
- PEC
phycoerythrocyanin
- Pr
red-absorbing state of phytochrome
- Pfr
far-red-absorbing state of phytochrome
- MCP
methyl-accepting chemotaxis protein
- MES
2-(N-morpholino)-ethanesulfonic acid
- PAS
domain acronym derived from Per, ARNT, Sim
- PHY
domain specific to red/far-red phytochromes, C-terminal to the GAF domain
- PCB
phycocyanobilin
- SAR
specific absorbance ratio (peak absorption of the long-wavelength chromophore band divided by that of the aromatic amino acid at ~ 280 nm)
- TES
N-[tris(hydroxymethyl)methyl]-2-aminoethanesulfonic acid
- TKKG buffer
25 mM TES-KOH pH 7.8, 100 mM KCl, 10% (v/v) glycerol
- XRG
extended red/green (designating a CBCR clade)
Footnotes
ACCESSION NUMBERS
Genbank protein accessions OKH39970 (NIES2119_03185), WP_083263321, MBF2026838, WP_082218260 (OSC10802_3032), MBC8123355 (MBC8123355g2), HGZ86378, ERT08869 (M595_1144), AFZ17632 (Mic7113_1774g1), AFZ11732 (Cri9333_0815), WP_027842869, NJL97683 (NJL97683g1 and NJL97683g2); WP_006508324, WP_218079506 (WP_218079506g3), BAB73026 (All1069, AnPixJg2); MBC7882157, MBC7881526, ACC83125 (Npun_R4776 or NpR4776); DOE-IMG accession number CYJSC1_DRAFT_64120 (RfpAg1); Genbank scaffold accessions JACMLN010000107 (encoding MBC7882157), JACMLN010000089 (encoding MBC7881526), JAAXLU010000010 (encoding MP9P1_10_2A and MP9P1_10_2B); Genbank genome accession GCA_014696065.1 (cf. Leptolyngbya sp. FACHB-261, associated with BioPoject accession PRJNA598298).
Data availability.
Raw and normalized absorption spectra, absorption values used in pKa estimation, multiple sequence alignments before and after gap removal, the resulting tree files, and analysis of phytochrome superfamily content in cyanobacterial lineages are available via DataDryad (https://doi.org/doi:10.25338/B8JM11).
REFERENCES
- [1].Rockwell NC, Su YS, Lagarias JC. Phytochrome structure and signaling mechanisms. Annu Rev Plant Biol. 2006;57:837–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Franklin KA, Quail PH. Phytochrome functions in Arabidopsis development. J Exp Bot. 2010;61:11–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Casal JJ. Photoreceptor signaling networks in plant responses to shade. Annu Rev Plant Biol. 2013;64:403–27. [DOI] [PubMed] [Google Scholar]
- [4].Wagner JR, Brunzelle JS, Forest KT, Vierstra RD. A light-sensing knot revealed by the structure of the chromophore binding domain of phytochrome. Nature. 2005;438:325–31. [DOI] [PubMed] [Google Scholar]
- [5].Essen LO, Mailliet J, Hughes J. The structure of a complete phytochrome sensory module in the Pr ground state. Proc Natl Acad Sci USA. 2008;105:14709–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Yang X, Kuk J, Moffat K. Crystal structure of Pseudomonas aeruginosa bacteriophytochrome: photoconversion and signal transduction. Proc Natl Acad Sci USA. 2008;105:14715–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [7].Hu W, Su YS, Lagarias JC. A light-independent allele of phytochrome B faithfully recapitulates photomorphogenic transcriptional networks. Mol Plant. 2009;2:166–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [8].Yang X, Ren Z, Kuk J, Moffat K. Temperature-scan cryocrystallography reveals reaction intermediates in bacteriophytochrome. Nature. 2011;479:428–32. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Takala H, Bjorling A, Berntsson O, Lehtivuori H, Niebling S, Hoernke M, et al. Signal amplification and transduction in phytochrome photosensors. Nature. 2014;509:245–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Kurttila M, Rumfeldt J, Takala H, Ihalainen JA. The interconnecting hairpin extension “arm”: An essential allosteric element of phytochrome activity. Structure. 2023. [DOI] [PubMed] [Google Scholar]
- [11].Rockwell NC, Lagarias JC. Phytochrome evolution in 3D: deletion, duplication, and diversification. New Phytol. 2020;225:2283–300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Brandt S, von Stetten D, Gunther M, Hildebrandt P, Frankenberg-Dinkel N. The fungal phytochrome FphA from Aspergillus nidulans. J Biol Chem. 2008;283:34605–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Rockwell NC, Duanmu D, Martin SS, Bachy C, Price DC, Bhattacharya D, et al. Eukaryotic algal phytochromes span the visible spectrum. Proc Natl Acad Sci USA. 2014;111:3871–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [14].Fortunato AE, Jaubert M, Enomoto G, Bouly JP, Raniello R, Thaler M, et al. Diatom Phytochromes Reveal the Existence of Far-Red-Light-Based Sensing in the Ocean. Plant Cell. 2016;28:616–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [15].Yeh K-C, Wu S-H, Murphy JT, Lagarias JC. A cyanobacterial phytochrome two-component light sensory system. Science. 1997;277:1505–8. [DOI] [PubMed] [Google Scholar]
- [16].Rockwell NC, Lagarias JC. A brief history of phytochromes. ChemPhysChem. 2010;11:1172–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [17].Rockwell NC, Martin SS, Feoktistova K, Lagarias JC. Diverse two-cysteine photocycles in phytochromes and cyanobacteriochromes. Proc Natl Acad Sci USA. 2011;108:11854–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [18].Song JY, Lee HY, Yang HW, Song JJ, Lagarias JC, Park YI. Spectral and photochemical diversity of tandem cysteine cyanobacterial phytochromes. J Biol Chem. 2020;295:6754–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [19].Wu SH, Lagarias JC. Defining the bilin lyase domain: lessons from the extended phytochrome superfamily. Biochemistry. 2000;39:13487–95. [DOI] [PubMed] [Google Scholar]
- [20].Ulijasz AT, Cornilescu G, von Stetten D, Kaminski S, Mroginski MA, Zhang J, et al. Characterization of two thermostable cyanobacterial phytochromes reveals global movements in the chromophore-binding domain during photoconversion. J Biol Chem. 2008;283:21251–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Gan F, Zhang S, Rockwell NC, Martin SS, Lagarias JC, Bryant DA. Extensive remodeling of a cyanobacterial photosynthetic apparatus in far-red light. Science. 2014;345:1312–7. [DOI] [PubMed] [Google Scholar]
- [22].Ikeuchi M, Ishizuka T. Cyanobacteriochromes: a new superfamily of tetrapyrrole-binding photoreceptors in cyanobacteria. Photochem Photobiol Sci. 2008;7:1159–67. [DOI] [PubMed] [Google Scholar]
- [23].Fushimi K, Narikawa R. Cyanobacteriochromes: photoreceptors covering the entire UV-to-visible spectrum. Curr Opin Struct Biol. 2019;57:39–46. [DOI] [PubMed] [Google Scholar]
- [24].Fushimi K, Narikawa R. Phytochromes and cyanobacteriochromes: photoreceptor molecules incorporating a linear tetrapyrrole chromophore. Adv Exp Med Biol. 2021;1293:167–87. [DOI] [PubMed] [Google Scholar]
- [25].Cho SM, Jeoung SC, Song JY, Kupriyanova EV, Pronina NA, Lee BW, et al. Genomic survey and biochemical analysis of recombinant candidate cyanobacteriochromes reveals enrichment for near UV/violet sensors in the halotolerant and alkaliphilic cyanobacterium Microcoleus IPPAS B353. J Biol Chem. 2015;290:28502–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Rockwell NC, Martin SS, Lagarias JC. Identification of cyanobacteriochromes detecting far-red light. Biochemistry. 2016;55:3907–19. [DOI] [PubMed] [Google Scholar]
- [27].Narikawa R, Fukushima Y, Ishizuka T, Itoh S, Ikeuchi M. A novel photoactive GAF domain of cyanobacteriochrome AnPixJ that shows reversible green/red photoconversion. J Mol Biol. 2008;380:844–55. [DOI] [PubMed] [Google Scholar]
- [28].Rockwell NC, Martin SS, Lagarias JC. Red/green cyanobacteriochromes: sensors of color and power. Biochemistry. 2012;51:9667–77. [DOI] [PubMed] [Google Scholar]
- [29].Rockwell NC, Martin SS, Lim S, Lagarias JC, Ames JB. Characterization of red/green cyanobacteriochrome NpR6012g4 by solution nuclear magnetic resonance spectroscopy: a hydrophobic pocket for the C15-E,anti chromophore in the photoproduct. Biochemistry. 2015;54:3772–83. [DOI] [PubMed] [Google Scholar]
- [30].Lim S, Yu Q, Gottlieb SM, Chang C-W, Rockwell NC, Martin SS, et al. Correlating structural and photochemical heterogeneity in cyanobacteriochrome NpR6012g4. Proc Natl Acad Sci USA. 2018;115:4387–92. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Xu X, Port A, Wiebeler C, Zhao KH, Schapiro I, Gärtner W. Structural elements regulating the photochromicity in a cyanobacteriochrome. Proc Natl Acad Sci USA. 2020;117:2432–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].Song C, Psakis G, Lang C, Mailliet J, Gärtner W, Hughes J, et al. Two ground state isoforms and a chromophore D-ring photoflip triggering extensive intramolecular changes in a canonical phytochrome. Proc Natl Acad Sci USA. 2011;108:3842–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Rockwell NC, Martin SS, Lim S, Lagarias JC, Ames JB. Characterization of red/green cyanobacteriochrome NpR6012g4 by solution nuclear magnetic resonance spectroscopy: a protonated bilin ring system in both photostates. Biochemistry. 2015;54:2581–600. [DOI] [PubMed] [Google Scholar]
- [34].Kehoe DM, Gutu A. Responding to color: the regulation of complementary chromatic adaptation. Annu Rev Plant Biol. 2006;57:127–50. [DOI] [PubMed] [Google Scholar]
- [35].Hirose Y, Shimada T, Narikawa R, Katayama M, Ikeuchi M. Cyanobacteriochrome CcaS is the green light receptor that induces the expression of phycobilisome linker protein. Proc Natl Acad Sci USA. 2008;105:9528–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Hirose Y, Narikawa R, Katayama M, Ikeuchi M. Cyanobacteriochrome CcaS regulates phycoerythrin accumulation in Nostoc punctiforme, a group II chromatic adapter. Proc Natl Acad Sci USA. 2010;107:8854–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Hirose Y, Rockwell NC, Nishiyama K, Narikawa R, Ukaji Y, Inomata K, et al. Green/red cyanobacteriochromes regulate complementary chromatic acclimation via a protochromic photocycle. Proc Natl Acad Sci USA. 2013;110:4974–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Hirose Y, Chihong S, Watanabe M, Yonekawa C, Murata K, Ikeuchi M, et al. Diverse chromatic acclimation processes regulating phycoerythrocyanin and rod-shaped phycobilisome in cyanobacteria. Mol Plant. 2019;12:715–25. [DOI] [PubMed] [Google Scholar]
- [39].Gottlieb SM, Kim PW, Rockwell NC, Hirose Y, Ikeuchi M, Lagarias JC, et al. Primary photodynamics of the green/red-absorbing photoswitching regulator of the chromatic adaptation E domain from Fremyella diplosiphon. Biochemistry. 2013;52:8198–208. [DOI] [PubMed] [Google Scholar]
- [40].Rockwell NC, Martin SS, Lagarias JC. Identification of DXCF cyanobacteriochrome lineages with predictable photocycles. Photochem Photobiol Sci. 2015;14:929–41. [DOI] [PubMed] [Google Scholar]
- [41].Rockwell NC, Martin SS, Lagarias JC. There and back again: loss and reacquisition of two-Cys photocycles in cyanobacteriochromes. Photochem Photobiol. 2017;93:741–54. [DOI] [PubMed] [Google Scholar]
- [42].Bandara S, Rockwell NC, Zeng X, Ren Z, Wang C, Shin H, et al. Crystal structure of a far-red-sensing cyanobacteriochrome reveals an atypical bilin conformation and spectral tuning mechanism. Proc Natl Acad Sci USA. 2021;118:e2025094118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Priyadarshini N, Steube N, Wiens D, Narikawa R, Wilde A, Hochberg GKA, et al. Evidence for an early green/red photocycle that precedes the diversification of GAF domain photoreceptor cyanobacteriochromes. Photochem Photobiol Sci. 2023;22:1415–27. [DOI] [PubMed] [Google Scholar]
- [44].Campbell EL, Hagen KD, Chen R, Risser DD, Ferreira DP, Meeks JC. Genetic analysis reveals the identity of the photoreceptor for phototaxis in hormogonium filaments of Nostoc punctiforme. J Bacteriol. 2015;197:782–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [45].Yang Y, Lam V, Adomako M, Simkovsky R, Jakob A, Rockwell NC, et al. Phototaxis in a wild isolate of the cyanobacterium Synechococcus elongatus. Proc Natl Acad Sci USA. 2018;115:E12378–E87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [46].Blain-Hartung M, Rockwell NC, Lagarias JC. Natural diversity provides a broad spectrum of cyanobacteriochrome-based diguanylate cyclases. Plant Physiol. 2021;187:632–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [47].Savakis P, De Causmaecker S, Angerer V, Ruppert U, Anders K, Essen LO, et al. Light-induced alteration of c-di-GMP level controls motility of Synechocystis sp. PCC 6803. Mol Microbiol. 2012;85:239–51. [DOI] [PubMed] [Google Scholar]
- [48].Anders K, Daminelli-Widany G, Mroginski MA, von Stetten D, Essen LO. Structure of the cyanobacterial phytochrome 2 photosensor implies a tryptophan switch for phytochrome signaling. J Biol Chem. 2013;288:35714–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [49].Narikawa R, Ishizuka T, Muraki N, Shiba T, Kurisu G, Ikeuchi M. Structures of cyanobacteriochromes from phototaxis regulators AnPixJ and TePixJ reveal general and specific photoconversion mechanism. Proc Natl Acad Sci USA. 2013;110:918–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Fixen KR, Baker AW, Stojkovic EA, Beatty JT, Harwood CS. Apo-bacteriophytochromes modulate bacterial photosynthesis in response to low light. Proceedings of the National Academy of Sciences of the United States of America. 2014;111:E237–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [51].Rockwell NC, Njuguna SL, Roberts L, Castillo E, Parson VL, Dwojak S, et al. A second conserved GAF domain cysteine is required for the blue/green photoreversibility of cyanobacteriochrome Tlr0924 from Thermosynechococcus elongatus. Biochemistry. 2008;47:7304–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [52].Ishizuka T, Kamiya A, Suzuki H, Narikawa R, Noguchi T, Kohchi T, et al. The cyanobacteriochrome, TePixJ, isomerizes its own chromophore by converting phycocyanobilin to phycoviolobilin. Biochemistry. 2011;50:953–61. [DOI] [PubMed] [Google Scholar]
- [53].Rockwell NC, Martin SS, Gulevich AG, Lagarias JC. Phycoviolobilin formation and spectral tuning in the DXCF cyanobacteriochrome subfamily. Biochemistry. 2012;51:1449–63. [DOI] [PubMed] [Google Scholar]
- [54].Fushimi K, Ikeuchi M, Narikawa R. The expanded red/green cyanobacteriochrome lineage: an evolutionary hot spot. Photochem Photobiol. 2017;93:903–6. [DOI] [PubMed] [Google Scholar]
- [55].Rippka R, Waterbury J, Cohen-Bazire G. A cyanobacterium which lacks thylakoids. Arch Microbiol. 1974;100:419–36. [Google Scholar]
- [56].Saw JH, Schatz M, Brown MV, Kunkel DD, Foster JS, Shick H, et al. Cultivation and complete genome sequencing of Gloeobacter kilaueensis sp. nov., from a lava cave in Kilauea Caldera, Hawai’i. PLoS One. 2013;8:e76376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Nakamura Y, Kaneko T, Sato S, Mimuro M, Miyashita H, Tsuchiya T, et al. Complete genome structure of Gloeobacter violaceus PCC 7421, a cyanobacterium that lacks thylakoids. DNA Res. 2003;10:137–45. [DOI] [PubMed] [Google Scholar]
- [58].Saw JH, Cardona T, Montejano G. Complete genome sequencing of a novel Gloeobacter species from a waterfall cave in Mexico. Genome Biol Evol. 2021;13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Ward RD, Stajich JE, Johansen JR, Huntemann M, Clum A, Foster B, et al. Metagenome sequencing to explore phylogenomics of terrestrial cyanobacteria. Microbiol Resour Announc. 2021;10:e0025821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [60].Grettenberger CL, Sumner DY, Wall K, Brown CT, Eisen JA, Mackey TJ, et al. A phylogenetically novel cyanobacterium most closely related to Gloeobacter. ISME J. 2020;14:2142–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [61].Grettenberger CL. Novel Gloeobacterales spp. from diverse environments across the globe. mSphere. 2021;6:e0006121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [62].Rahmatpour N, Hauser DA, Nelson JM, Chen PY, Villarreal AJ, Ho MY, et al. A novel thylakoid-less isolate fills a billion-year gap in the evolution of cyanobacteria. Curr Biol. 2021;31:2857–67 e4. [DOI] [PubMed] [Google Scholar]
- [63].Pessi IS, Popin RV, Durieu B, Lara Y, Tytgat B, Savaglia V, et al. Novel diversity of polar cyanobacteria revealed by genome-resolved metagenomics. Microb Genom. 2023;9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [64].Rockwell NC, Lagarias JC. GUN4 appeared early in cyanobacterial evolution. PNAS Nexus. 2023;2:pgad131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [65].Fushimi K, Rockwell NC, Enomoto G, Ni Ni W, Martin SS, Gan F, et al. Cyanobacteriochrome Photoreceptors Lacking the Canonical Cys Residue. Biochemistry. 2016;55:6981–95. [DOI] [PubMed] [Google Scholar]
- [66].Hoshino H, Narikawa R. Novel cyanobacteriochrome photoreceptor with the second Cys residue showing atypical orange/blue reversible photoconversion. Photochem Photobiol Sci. 2023;22:251–61. [DOI] [PubMed] [Google Scholar]
- [67].Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [68].Chen MY, Teng WK, Zhao L, Hu CX, Zhou YK, Han BP, et al. Comparative genomics reveals insights into cyanobacterial evolution and habitat adaptation. ISME J. 2021;15:211–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [69].Komárek J, Johansen JR, Šmarda J, Strunecký O. Phylogeny and taxonomy of Synechococcus–like cyanobacteria. Fottea (Praha). 2020;20:171–91. [Google Scholar]
- [70].Strunecký O, Ivanova AP, Mareš J. An updated classification of cyanobacterial orders and families based on phylogenomic and polyphasic analysis. J Phycol. 2023;59:12–51. [DOI] [PubMed] [Google Scholar]
- [71].Sorokovnikova EG, Tikhonova IV, Belykh OI, Klimenkov IV, Likhoshvai EV. Identification of two cyanobacterial strains isolated from the Kotel’nikovskii hot spring of the Baikal rift. Mikrobiologiia. 2008;77:413–20. [PubMed] [Google Scholar]
- [72].Mareš J, Strunecký O, Bučinská L, Wiedermannová J. Evolutionary patterns of thylakoid architecture in cyanobacteria. Front Microbiol. 2019;10:277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [73].Mareš J, Hrouzek P, Kaňa R, Ventura S, Strunecký O, Komárek J. The primitive thylakoid-less cyanobacterium Gloeobacter is a common rock-dwelling organism. PLoS One. 2013;8:e66323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [74].Bryant DA. The photoregulated expression of multiple phycocyanin species. A general mechanism for the control of phycocyanin synthesis in chromatically adapting cyanobacteria. Eur J Biochem. 1981;119:425–9. [DOI] [PubMed] [Google Scholar]
- [75].Rockwell NC, Martin SS, Gan F, Bryant DA, Lagarias JC. NpR3784 is the prototype for a distinctive group of red/green cyanobacteriochromes using alternative Phe residues for photoproduct tuning. Photochem Photobiol Sci. 2015;14:258–69. [DOI] [PubMed] [Google Scholar]
- [76].Moreno MV, Rockwell NC, Mora M, Fisher AJ, Lagarias JC. A far-red cyanobacteriochrome lineage specific for verdins. Proc Natl Acad Sci USA. 2020;117:27962–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [77].Waterworth SC, Isemonger EW, Rees ER, Dorrington RA, Kwan JC. Conserved bacterial genomes from two geographically isolated peritidal stromatolite formations shed light on potential functional guilds. Environ Microbiol Rep. 2021;13:126–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [78].Rockwell NC, Martin SS, Lagarias JC. Mechanistic insight into the photosensory versatility of DXCF cyanobacteriochromes. Biochemistry. 2012;51:3576–85. [DOI] [PubMed] [Google Scholar]
- [79].Burgie ES, Walker JM, Phillips GN Jr., Vierstra RD. A photo-labile thioether linkage to phycoviolobilin provides the foundation for the blue/green photocycles in DXCF-cyanobacteriochromes. Structure. 2013;21:88–97. [DOI] [PubMed] [Google Scholar]
- [80].Lim S, Rockwell NC, Martin SS, Dallas JL, Lagarias JC, Ames JB. Photoconversion changes bilin chromophore conjugation and protein secondary structure in the violet/orange cyanobacteriochrome NpF2163g3. Photochem Photobiol Sci. 2014;13:951–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [81].Burgie ES, Vierstra RD. Phytochromes: an atomic perspective on photoactivation and signaling. Plant Cell. 2014;26:4568–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [82].Takala H, Lehtivuori HK, Berntsson O, Hughes A, Nanekar R, Niebling S, et al. On the (un)coupling of the chromophore, tongue interactions, and overall conformation in a bacterial phytochrome. J Biol Chem. 2018;293:8161–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [83].Hahn J, Strauss HM, Landgraf FT, Gimenez HF, Lochnit G, Schmieder P, et al. Probing protein-chromophore interactions in Cph1 phytochrome by mutagenesis. FEBS J. 2006;273:1415–29. [DOI] [PubMed] [Google Scholar]
- [84].Rockwell NC, Martin SS, Gulevich AG, Lagarias JC. Conserved phenylalanine residues are required for blue-shifting of cyanobacteriochrome photoproducts. Biochemistry. 2014;53:3118–30. [DOI] [PubMed] [Google Scholar]
- [85].Zhao C, Gan F, Shen G, Bryant DA. RfpA, RfpB, and RfpC are the Master Control Elements of Far-Red Light Photoacclimation (FaRLiP). Front Microbiol. 2015;6:1303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [86].Rockwell NC, Shang L, Martin SS, Lagarias JC. Distinct classes of red/far-red photochemistry within the phytochrome superfamily. Proc Natl Acad Sci USA. 2009;106:6123–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [87].Rockwell NC, Moreno MV, Martin SS, Lagarias JC. Protein-chromophore interactions controlling photoisomerization in red/green cyanobacteriochromes. Photochem Photobiol Sci. 2022;21:471–91. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [88].Rockwell NC, Martin SS, Lagarias JC. Elucidating the origins of phycocyanobilin biosynthesis and phycobiliproteins. Proc Natl Acad Sci USA. 2023;120:e2300770120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [89].Anders K, von Stetten D, Mailliet J, Kiontke S, Sineshchekov VA, Hildebrandt P, et al. Spectroscopic and photochemical characterization of the red-light sensitive photosensory module of Cph2 from Synechocystis PCC 6803. Photochem Photobiol. 2011;87:160–73. [DOI] [PubMed] [Google Scholar]
- [90].Sato T, Kikukawa T, Miyoshi R, Kajimoto K, Yonekawa C, Fujisawa T, et al. Protochromic absorption changes in the two-cysteine photocycle of a blue/orange cyanobacteriochrome. J Biol Chem. 2019;294:18909–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [91].Fushimi K, Enomoto G, Ikeuchi M, Narikawa R. Distinctive properties of dark reversion kinetics between two red/green-type cyanobacteriochromes and their application in the photoregulation of cAMP synthesis. Photochem Photobiol. 2017;93:681–91. [DOI] [PubMed] [Google Scholar]
- [92].Fushimi K, Miyazaki T, Kuwasaki Y, Nakajima T, Yamamoto T, Suzuki K, et al. Rational conversion of chromophore selectivity of cyanobacteriochromes to accept mammalian intrinsic biliverdin. Proc Natl Acad Sci USA. 2019;116:8301–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [93].Narikawa R, Nakajima T, Aono Y, Fushimi K, Enomoto G, Ni Ni W, et al. A biliverdin-binding cyanobacteriochrome from the chlorophyll d-bearing cyanobacterium Acaryochloris marina. Sci Rep. 2015;5:7950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [94].Hasegawa M, Fushimi K, Miyake K, Nakajima T, Oikawa Y, Enomoto G, et al. Molecular characterization of DXCF cyanobacteriochromes from the cyanobacterium Acaryochloris marina identifies a blue-light power sensor. J Biol Chem. 2018;293:1713–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [95].Enomoto G, Nomura R, Shimada T, Ni Ni W, Narikawa R, Ikeuchi M. Cyanobacteriochrome SesA is a diguanylate cyclase that induces cell aggregation in Thermosynechococcus. J Biol Chem. 2014;289:24801–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [96].Enomoto G, Ni Ni W, Narikawa R, Ikeuchi M. Three cyanobacteriochromes work together to form a light color-sensitive input system for c-di-GMP signaling of cell aggregation. Proc Natl Acad Sci USA. 2015;112:8082–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [97].Blain-Hartung M, Rockwell NC, Moreno MV, Martin SS, Gan F, Bryant DA, et al. Cyanobacteriochrome-based photoswitchable adenylyl cyclases (cPACs) for broad spectrum light regulation of cAMP levels in cells. J Biol Chem. 2018;293:8473–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [98].Yoshihara S, Suzuki F, Fujita H, Geng XX, Ikeuchi M. Novel putative photoreceptor and regulatory genes required for the positive phototactic movement of the unicellular motile cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 2000;41:1299–304. [DOI] [PubMed] [Google Scholar]
- [99].Risser DD. Hormogonium development and motility in filamentous cyanobacteria. Appl Environ Microbiol. 2023;89:e0039223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [100].Yoshihara S, Geng X, Ikeuchi M. pilG Gene cluster and split pilL genes involved in pilus biogenesis, motility and genetic transformation in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 2002;43:513–21. [DOI] [PubMed] [Google Scholar]
- [101].Yoshihara S, Katayama M, Geng X, Ikeuchi M. Cyanobacterial phytochrome-like PixJ1 holoprotein shows novel reversible photoconversion between blue- and green-absorbing forms. Plant Cell Physiol. 2004;45:1729–37. [DOI] [PubMed] [Google Scholar]
- [102].Yoshihara S, Shimada T, Matsuoka D, Zikihara K, Kohchi T, Tokutomi S. Reconstitution of blue-green reversible photoconversion of a cyanobacterial photoreceptor, PixJ1, in phycocyanobilin-producing Escherichia coli. Biochemistry. 2006;45:3775–84. [DOI] [PubMed] [Google Scholar]
- [103].Khayatan B, Meeks JC, Risser DD. Evidence that a modified type IV pilus-like system powers gliding motility and polysaccharide secretion in filamentous cyanobacteria. Mol Microbiol. 2015;98:1021–36. [DOI] [PubMed] [Google Scholar]
- [104].Harwood TV, Zuniga EG, Kweon H, Risser DD. The cyanobacterial taxis protein HmpF regulates type IV pilus activity in response to light. Proc Natl Acad Sci USA. 2021;118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [105].Narikawa R, Enomoto G, Ni Ni W, Fushimi K, Ikeuchi M. A New Type of Dual-Cys Cyanobacteriochrome GAF Domain Found in Cyanobacterium Acaryochloris marina, Which Has an Unusual Red/Blue Reversible Photoconversion Cycle. Biochemistry. 2014;53:5051–9. [DOI] [PubMed] [Google Scholar]
- [106].Kuwasaki Y, Miyake K, Fushimi K, Takeda Y, Ueda Y, Nakajima T, et al. Protein Engineering of Dual-Cys Cyanobacteriochrome AM1_1186g2 for Biliverdin Incorporation and Far-Red/Blue Reversible Photoconversion. Int J Mol Sci. 2019;20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [107].Fushimi K, Hasegawa M, Ito T, Rockwell NC, Enomoto G, Win NN, et al. Evolution-inspired design of multicolored photoswitches from a single cyanobacteriochrome scaffold. Proc Natl Acad Sci U S A. 2020;117:15573–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [108].Fushimi K, Hoshino H, Shinozaki-Narikawa N, Kuwasaki Y, Miyake K, Nakajima T, et al. The Cruciality of Single Amino Acid Replacement for the Spectral Tuning of Biliverdin-Binding Cyanobacteriochromes. Int J Mol Sci. 2020;21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [109].Fushimi K, Narikawa R. Unusual ring D fixation by three crucial residues promotes phycoviolobilin formation in the DXCF-type cyanobacteriochrome without the second Cys. Biochem J. 2021;478:1043–59. [DOI] [PubMed] [Google Scholar]
- [110].Suzuki T, Yoshimura M, Hoshino H, Fushimi K, Arai M, Narikawa R. Introduction of reversible cysteine ligation ability to the biliverdin-binding cyanobacteriochrome photoreceptor. FEBS J. 2023. [DOI] [PubMed] [Google Scholar]
- [111].Kirpich JS, Mix LT, Martin SS, Rockwell NC, Lagarias JC, Larsen DS. Protonation heterogeneity modulates the ultrafast photocycle initiation dynamics of phytochrome Cph1. J Phys Chem Lett. 2018;9:3454–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [112].Luimstra VM, Schuurmans JM, Verschoor AM, Hellingwerf KJ, Huisman J, Matthijs HCP. Blue light reduces photosynthetic efficiency of cyanobacteria through an imbalance between photosystems I and II. Photosynth Res. 2018;138:177–89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [113].Bryant DA, Glazer AN, Eiserling FA. Characterization and structural properties of the major biliproteins of Anabaena sp. Arch Microbiol. 1976;110:61–75. [DOI] [PubMed] [Google Scholar]
- [114].Chauvet A, Sarrou J, Lin S, Romberger SP, Golbeck JH, Savikhin S, et al. Temporal and spectral characterization of the photosynthetic reaction center from Heliobacterium modesticaldum. Photosynth Res. 2013;116:1–9. [DOI] [PubMed] [Google Scholar]
- [115].Tsukatani Y, Yamamoto H, Mizoguchi T, Fujita Y, Tamiaki H. Completion of biosynthetic pathways for bacteriochlorophyll g in Heliobacterium modesticaldum: The C8-ethylidene group formation. Biochim Biophys Acta. 2013;1827:1200–4. [DOI] [PubMed] [Google Scholar]
- [116].Bryant DA, Canniffe DP. How nature designs light-harvesting antenna systems: design principles and functional realization in chlorophototrophic prokaryotes. J Phys B: At Mol Opt Phys. 2018;51:033001. [Google Scholar]
- [117].Hayashi H, Miyao M, Morita S. Absorption and fluorescence spectra of light-harvesting bacteriochlorophyll-protein complexes from Rhodopseudomonas palustris in the near-infrared region. J Biochem. 1982;91:1017–27. [DOI] [PubMed] [Google Scholar]
- [118].Hohmann-Marriott MF, Blankenship RE. Variable fluorescence in green sulfur bacteria. Biochim Biophys Acta. 2007;1767:106–13. [DOI] [PubMed] [Google Scholar]
- [119].Quest B, Hubschmann T, Sharda S, Tandeau de Marsac N, Gartner W. Homologous expression of a bacterial phytochrome. The cyanobacterium Fremyella diplosiphon incorporates biliverdin as a genuine, functional chromophore. FEBS J. 2007;274:2088–98. [DOI] [PubMed] [Google Scholar]
- [120].Sharda S, Shah R, Gartner W. Domain interaction in cyanobacterial phytochromes as a prerequisite for spectral integrity. Eur Biophys J. 2007;36:815–21. [DOI] [PubMed] [Google Scholar]
- [121].Giraud E, Fardoux J, Fourier N, Hannibal L, Genty B, Bouyer P, et al. Bacteriophytochrome controls photosystem synthesis in anoxygenic bacteria. Nature. 2002;417:202–5. [DOI] [PubMed] [Google Scholar]
- [122].Grébert T, Doré H, Partensky F, Farrant GK, Boss ES, Picheral M, et al. Light color acclimation is a key process in the global ocean distribution of Synechococcus cyanobacteria. Proc Natl Acad Sci USA. 2018;115:E2010–E9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [123].Enomoto G, Ikeuchi M. Blue-/Green-Light-Responsive Cyanobacteriochromes Are Cell Shade Sensors in Red-Light Replete Niches. iScience. 2020;23:100936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [124].Kumar S, Ali Kubar A, Zhu F, Shao C, Cui Y, Hu X, et al. Sunlight filtered via translucent-colored polyvinyl chloride sheets enhanced the light absorption capacity and growth of Arthrospira platensis cultivated in a pilot-scale raceway pond. Bioresour Technol. 2023;386:129501. [DOI] [PubMed] [Google Scholar]
- [125].Belkin S, Mehlhorn RJ, Packer L. Proton gradients in intact cyanobacteria. Plant Physiol. 1987;84:25–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [126].Giraldez-Ruiz N, Mateo P, Bonilla I, Fernandez-Piñas F. The relationship between intracellular pH, growth characteristics and calcium in the cyanobacterium Anahaena sp. strain PCC7120 exposed to low pH. New Phytol. 1997;137:599–605. [DOI] [PubMed] [Google Scholar]
- [127].Nakamura S, Fu N, Kondo K, Wakabayashi KI, Hisabori T, Sugiura K. A luminescent Nanoluc-GFP fusion protein enables readout of cellular pH in photosynthetic organisms. J Biol Chem. 2021;296:100134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [128].Kashimoto T, Miyake K, Sato M, Maeda K, Matsumoto C, Ikeuchi M, et al. Acclimation process of the chlorophyll d-bearing cyanobacterium Acaryochloris marina to an orange light environment revealed by transcriptomic analysis and electron microscopic observation. J Gen Appl Microbiol. 2020;66:106–15. [DOI] [PubMed] [Google Scholar]
- [129].Kumar S, Cheng J, Ali Kubar A, Guo W, Song Y, Liu S, et al. Orange light spectra filtered through transparent colored polyvinyl chloride sheet enhanced pigment content and growth of Arthrospira cells. Bioresour Technol. 2021;319:124179. [DOI] [PubMed] [Google Scholar]
- [130].Trinh MDL, Masuda S. Chloroplast pH Homeostasis for the Regulation of Photosynthesis. Front Plant Sci. 2022;13:919896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [131].Katoh K, Standley DM. MAFFT: iterative refinement and additional methods. Methods Mol Biol. 2014;1079:131–46. [DOI] [PubMed] [Google Scholar]
- [132].Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21. [DOI] [PubMed] [Google Scholar]
- [133].Lemoine F, Domelevo Entfellner JB, Wilkinson E, Correia D, Davila Felipe M, De Oliveira T, et al. Renewing Felsenstein’s phylogenetic bootstrap in the era of big data. Nature. 2018;556:452–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [134].Rockwell NC, Lagarias JC. Flexible mapping of homology onto structure with homolmapper. BMC Bioinf. 2007;8:123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [135].Mukougawa K, Kanamoto H, Kobayashi T, Yokota A, Kohchi T. Metabolic engineering to produce phytochromes with phytochromobilin, phycocyanobilin, or phycoerythrobilin chromophore in Escherichia coli. FEBS Lett. 2006;580:1333–8. [DOI] [PubMed] [Google Scholar]
- [136].Ma Q, Hua HH, Chen Y, Liu BB, Kramer AL, Scheer H, et al. A rising tide of blue-absorbing biliprotein photoreceptors: characterization of seven such bilin-binding GAF domains in Nostoc sp. PCC7120. FEBS J. 2012;279:4095–108. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Raw and normalized absorption spectra, absorption values used in pKa estimation, multiple sequence alignments before and after gap removal, the resulting tree files, and analysis of phytochrome superfamily content in cyanobacterial lineages are available via DataDryad (https://doi.org/doi:10.25338/B8JM11).


