Abstract
The placenta plays a crucial role in pregnancy success. ΔNp63α (p63), a transcription factor from the TP53 family, is highly expressed in villous cytotrophoblasts (CTBs), the epithelial stem cells of the human placenta, and is involved in CTB maintenance and differentiation. We examined the mechanisms of action of p63 by identifying its downstream targets. Gene expression changes were evaluated following overexpression and knockdown of p63 in the JEG3 choriocarcinoma cell line, using microarray-based RNA profiling. High-temperature requirement A4 (HTRA4), a placenta-specific serine protease involved in trophoblast differentiation and altered in preeclampsia, was identified as a gene reciprocally regulated by p63, and its expression was characterized in primary human placental tissues by RNA-sequencing and in situ hybridization. Potential p63 DNA-binding motifs were identified in the HTRA4 promoter, and p63 occupancy at some of these sites was confirmed using chromatin immunoprecipitation, followed by quantitative PCR in both JEG3 and trophoblast stem cells. These data begin to identify members of the transcriptional network downstream of p63, thus laying the groundwork for probing mechanisms by which this important transcription factor regulates trophoblast stemness and differentiation.
The placenta is a transient organ critical for proper fetal growth and development.1 The epithelial compartment of the placenta originates from the trophectoderm, the outer layer of the blastocyst-stage embryo, which expands following implantation to form the cytotrophoblastic shell.2 This shell is composed of stem cells, called cytotrophoblasts (CTB), which differentiate into either multinucleated syncytiotrophoblast (STB) within floating chorionic villi to form the gas/nutrient exchange interface of the placenta or extravillous trophoblast (EVT) within anchoring villi to anchor the placenta to the uterine wall.3,4 CTBs are characterized by several surface markers, including epidermal growth factor receptor (EGFR), which is lost as the cells undergo differentiation into EVT within the cell columns of anchoring villi, acquiring instead the non-classic major histocompatibility complex class I molecule, human leukocyte antigen G (HLAG).5 Although many studies have focused on pathways driving EVT differentiation,6, 7, 8, 9, 10 less is known about the transcriptional program(s) involved in maintenance of CTB stemness.11,12
ΔNp63α, an isoform of TP63, a transcriptional regulator, and member of the TP53 family, is a specific marker of CTB within the human placenta.13 ΔNp63α, heretofore referred to simply as p63, is a known oncogene, produced by differential promoter use and alternative splicing.14 p63 is highly expressed in the proliferative CTB compartment, diminishing on differentiation into either STB or EVT.13,15 Forced overexpression of p63 in primary first-trimester and term CTB maintains cyclin B1 expression and inhibited differentiation.15,16 In JEG3 human choriocarcinoma cells, p63 overexpression enhances cell proliferation and reduces EVT-associated marker expression and function; conversely, p63 knockdown reduces cell proliferation and enhances EVT-associated features of these cells.15 Finally, p63 knockdown in human pluripotent stem cells interferes with bone morphogenetic protein 4 (BMP4)–induced initiation of CTB differentiation.16 Taken together, these results suggest a master regulatory role for p63 in both initiation and maintenance of CTB stem cell fate.
We investigated the molecular machinery downstream of p63. Differential gene expression changes were examined in JEG3 cells following p63 overexpression and knockdown, using microarray-based RNA profiling. The findings demonstrated that ΔNp63 is epistatic to numerous genes, and confirmed one such downstream target, high-temperature requirement protein A (HTRA4), by chromatin immunoprecipitation (ChIP). These results begin to define the regulatory mechanisms by which p63 functions in the development and function of the human placenta.
Materials and Methods
Human Placental Samples
Human placental tissues were collected under a protocol approved by the Human Research Protections Program Committee Institutional Review Board of the University of California. All patients provided informed consent for the collection and use of these tissues.
Cell Lines and Culture Conditions
The human choriocarcinoma cell line JEG3 was obtained from ATCC (Manassas, VA) and cultured in Dulbecco's modified Eagle's medium (Thermo Fisher Scientific, Waltham, MA) supplemented with 10% fetal bovine serum (Sigma, St. Louis, MO). The full-length ΔNp63α pCMV lentivirally enforced overexpression and control (pCMV-lentivirus without any gene insert) vectors and the pLKO.1-based nontargeting scramble and p63-specific shRNA lentiviral constructs were previously described.15 A male human trophoblast stem cell (TSC) line (1000P), derived from 6-week gestational age placenta in July 2018, was cultured, as previously described,8 and used for ChIP experiments.
Microarray-Based Gene Expression Profiling and Analysis
For gene expression microarray, total RNA was purified using the mirVana RNA Isolation Kit (Ambion, Carlsbad, CA), quantified using the Ribogreen reagent (Lifetech, Inc., Carlsbad, CA) and a Qubit fluorometer (Thermo Fisher Scientific), and quality controlled on an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA). Only those RNA samples with an RNA integrity number >8.0 were subjected to further gene expression analysis. Total input RNA (200 ng) was amplified and labeled using the TotalPrep kit (Ambion). The labeled product was then hybridized to Illumina HT12 arrays and scanned on a BeadArray Reader (Illumina, Inc., San Diego, CA), according to the manufacturer's instructions. This microarray assay profiled 48,000 predefined probes through hybridization using optimized and standardized protocols. Three independent pCMV-ΔNp63 overexpression and pCMV-vector alone cell lines were subjected to microarray hybridization following confirmation of p63 enrichment.
With a detection P value cutoff of 0.01, probes were filtered, and normalized by using the LUMI package in R version 3.0.0 (https://www.r-project.org) with the robust spline normalization method. Qlucore Omics Explorer software version 3.1 (Qlucore AB, Lund, Sweden) was used to explore and analyze microarray gene expression data sets and to generate heat maps. Gene expression profiles of JEG3 p63 overexpression and knockdown cells were compared with cells transduced with the empty pCMV lentivirus or scramble shRNA lentivirus, respectively. Specific gene signatures were defined with a variance <0.01, two-group comparison (equivalent to paired t-test) q < 0.05, and fold change ≥1.5. The microarray data in this publication have been deposited in the National Center for Biotechnology Information's Gene Expression Omnibus and are accessible (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE251742; accession number GSE251742).
RNA-Sequencing Data Analysis
RNA-sequencing data used in this study were described in detail by Morey et al.8 Sequencing data from first-trimester CTB (n = 10) and EVT (n = 10) from this data set were re-analyzed for expression of p63, EGFR, ITGA6, HTRA4, HLAG, and MMP9. Similarly, RNA-sequencing data were obtained from Okae et al,17 for expression analysis of p63 and HTRA4 from first-trimester CTB, STB, and EVT. Unpaired t-test and one-way analysis of variance for multiple comparison tests were performed using GraphPad Prism version 10.1.0 (GraphPad Software, Boston, MA), with statistical significance set at 0.05.
In Situ Hybridization of Cells and Tissues
To examine HTRA4 expression in placenta, early gestation and term placental tissues (one each from gestational ages of 5, 7, 8, and 20 weeks, and term) were fixed in neutral-buffered formalin, dehydrated, embedded in paraffin, and divided into sections (5 μm thick). Sections were deparaffinized, rehydrated, and subjected to antigen retrieval and protease treatment as described by the manufacturer (ACD-Bio, Hayward, CA). In situ hybridization was performed using the RNAscope method (ACD-Bio) with mRNA probes specific to human HTRA4. After amplification steps, the probes were visualized with 3,3′-diaminobenzidine, and the slides were counterstained with hematoxylin. Slides were evaluated by conventional light microscopy on an Olympus BX43 microscope (Olympus, Waltham, MA).
In Silico Analysis and DNA Motif Comparisons
The p63 transcription factor DNA-binding sites were identified using the TFBIND tool software version 1.18 The DNA motif comparison analysis was performed for each promoter p63 DNA-binding site using Tomtom Motif Comparison algorithm version 5.5.3.19 The TFBIND software uses a cutoff value of 0.70, the same cutoff score as TRANSFAC R.3.4 (https://tfbind.hgc.jp/cutoff_table.html).
Chromatin Immunoprecipitation
ChIP was performed on JEG3 cells and TSC, as previously described.20 Briefly, approximately 2 × 106 to 4 × 106 cells (JEG3 or TSC) for each immunoprecipitation were cross-linked with formaldehyde, followed by micrococcal nuclease digestion and sonication to 100 to 500 bp. Before ChIP, chromatin was precleared with a mixture of protein A (Sigma). Selective immunoprecipitation was performed using 6 μg of anti-p63 [D2K8X (XP); Cell Signaling, Danvers, MA] or anti-IgG (ab133469; Abcam, Waltham, MA) antibody control. DNA recovered from 1% of precleared sheared chromatin was used as an input control. Quantitative real-time ChIP-PCR analysis was performed using the ChIP primers listed in Table 1 and SYBR Green dye (Applied Biosystems/Thermo Fisher, Waltham, MA) on the Applied Biosystems StepOnePlus Real-Time PCR System. Each PCR was performed in triplicate. The fold enrichment was calculated relative to the IgG antibody control using the ΔΔCT method. Data are expressed as the means ± SEM, and differences between groups were determined using one-sided t-tests with paired variance using Excel for Mac version 16.84 (Microsoft, Redmond, WA), with statistical significance set at 0.05.
Table 1.
List of Primers Used for Quantitative ChIP
| Primer name | Primer sequence |
|---|---|
| ΔNp63 A forward | 5′-CCTTCTGTAAATCGTGGTGGTC-3′ |
| ΔNp63 A reverse | 5′-CCCAGGAGACAGACAGGTAAA-3′ |
| ΔNp63 B forward | 5′-GAAACCACTGGCATTTCTCTG-3′ |
| ΔNp63 B reverse | 5′-TTAGGAGAGAGTACGCAAATCC-3′ |
| ΔNp63 C forward | 5′-GACCGGTGGTTTAACTTGTTATTG -3′ |
| ΔNp63 C reverse | 5′-CCACTGCCTGATAGCTAAAGAG-3′ |
| ΔNp63 D forward | 5′-CAGATTCTACATGAATGTTGGTACG-3′ |
| ΔNp63 D reverse | 5′-GCACGTGATGCATCTATGTAAA-3′ |
| HTRA4 A forward | 5′- GGGGATGTACGGGTGTGTAT-3′ |
| HTRA4 A reverse | 5′-TCCTCGCTTCCACTCTTCAG-3′ |
| HTRA4 B forward | 5′-TGGAAACTGTTACGCTTCTCA-3′ |
| HTRA4 B reverse | 5′-GTCTCTAGCCCTACCAG-3′ |
| HTRA4 C forward | 5′-TGCATGGAAACTGTTACGCT-3′ |
| HTRA4 C reverse | 5′-CCCATGGTCAGAAGGTCAGT-3′ |
| HTRA4 D forward | 5′-GGGATGTACGGGTGTGTATATG-3′ |
| HTRA4 D reverse | 5′-GACCTGCAAACCCTCCTAAA-3′ |
| HTRA4 E forward | 5′-AAATCTTGTCTGCCAGCACC-3′ |
| HTRA4 E reverse | 5′-AGCGTAACAGTTTCCATGCA-3′ |
| HTRA4 F forward | 5′-AAATGTCGGAAGTGGACAGG-3′ |
| HTRA4 F reverse | 5′-CAGGCAATAACACAGAGAGGAA-3′ |
| HTRA4 G forward | 5′-CAGTAGTTGAGGAGTGGAGTTTAG-3′ |
| HTRA4 G reverse | 5′-CCTGTCCACTTCCGACATTT-3′ |
ChIP, chromatin immunoprecipitation; HTRA4, high-temperature requirement A4.
Results
p63 Governs a Cytotrophoblast Gene Signature
To gain insight into the downstream targets of ΔNp63, a microarray-based platform was used to identify differentially expressed genes, following overexpression or knockdown of p63 in JEG3 cells. Overexpression of p63 significantly altered gene expression (Figure 1A and Supplemental Table S1), including up-regulation of 30 genes and down-regulation of 34 genes. Genes whose expression increased with p63 overexpression included CXXC5 (a zinc finger protein involved in growth of breast cancer cells)21; SLC22A11 (a solute carrier, involved in transporting harmful organic anions away from the fetal circulation)22; SERPINF1 (member of the serpin family of serine protease inhibitors enriched in glioma stem cells)23; and MAFB (a basic leucine zipper transcription factor associated with growth of esophageal squamous cell carcinoma).24 Conversely, genes down-regulated following p63 overexpression consisted of several STB-associated genes, including five chorionic gonadotropin β (CGB) subunit genes and HOPX (a homeodomain-containing transcription factor implicated in STB differentiation)25; as well as several EVT-associated genes, including FSTL326 and CDH5.27
Figure 1.
Identification of genes downstream of ΔNp63α (p63). A: Heat map of gene expression changes following ΔNp63α overexpression in JEG3 cells. B: Heat map of gene expression changes following p63 knockdown in JEG3 cells. The color scale represents the measured gene expression intensity, with red representing high and green representing low gene expression. Selected genes are highlighted to the right of each heat map; HTRA4 (boldfaced), a gene reciprocally regulated by p63 overexpression and downregulation, is further evaluated in this study. CMV, cytomegaloviral promoter; sh-p63, p63-specific short hairpin RNA; sh-Scramble, nonspecific short hairpin RNA.
Genes with altered expression following p63 knockdown in JEG3 cells were evaluated next. The p63 knockdown led to down-regulation of only 6 genes, but caused up-regulation of 19 genes, among which were MSLN (mesothelin; a glycophosphotidylinositol-anchored cell adhesion molecule), MYC (a proto-oncogene involved in cell cycle progression and apoptosis), as well as GAGE4 (a member of the family of cancer-testis antigens with a possible role in protection against stress-induced cell death)28 (Figure 1B and Supplemental Table S1).
Interestingly, HTRA4 was reciprocally regulated downstream of p63, with approximately 1.8-fold down-regulation in p63-overexpressing cells and approximately 1.8-fold up-regulation following p63 knockdown (Supplemental Table S1). HTRA4 is a serine protease that is highly expressed in the human placenta, is implicated in trophoblast differentiation and function,29, 30, 31 and is altered in the setting of preeclampsia.32,33 Therefore, this study further focused on the HTRA4 gene as a possible direct target of p63.
HTRA4 Is Expressed in Differentiated Trophoblast in Human Placental Tissues
To further evaluate the reciprocal expression of p63 and HTRA4 in placental tissues, the study first queried recently published RNA-sequencing data from CTB and EVT, isolated from first-trimester placental tissues, and purified based on surface expression of EGFR and HLAG, respectively.8 As shown in Figure 2A, TP63, EGFR, and ITGA6 were highly enriched in CTB, whereas HLAG and MMP9 were enriched in EVT. Similar to the latter group, HTRA4 showed the lowest expression in CTB, with 43-fold enrichment in EVT. In addition, published RNA-sequencing data from Okae et al17 were re-analyzed to query the levels of TP63 and HTRA4 in STB as well as CTB and EVT. As shown in Figure 2B, TP63 levels were highest in CTB and significantly diminished 25-fold in EVT and 80-fold in STB. In contrast, compared with CTB, HTRA4 was enriched 25-fold in EVT and 6-fold in STB (Figure 2B).
Figure 2.
High-temperature requirement A4 (HTRA4) expression in primary human placental cells and tissues. A: Tumor protein p63 (TP63) and HTRA4 show an inverse gene expression pattern by RNA-sequencing (RNA-seq) analysis in primary cytotrophoblasts [CTBs; identified by high epidermal growth factor receptor (EGFR) and integrin alpha-6 (ITGA6) expression] compared with extravillous trophoblasts [EVTs; identified by high human leukocyte antigen G (HLAG) and matrix metalloproteinase 9 (MMP9) expression], isolated from first-trimester placenta. B: Similarly, re-analysis of RNA-seq data from Okae et al17 shows TP63 to be highly expressed in CTB, whereas HTRA4 is more highly expressed in differentiated trophoblast [EVT and syncytiotrophoblast (STB)] of the first-trimester placenta. Unpaired t-test and one-way analysis of variance was applied for cell expression comparison tests, and P values are shown. C:In situ hybridization of primary human placental tissues across gestation at indicated weeks (w) of gestation, using HTRA4-specific probes. HTRA4 expression is enriched in both STB of floating villi (V) and cell column EVT (CC) in first trimester, but remained only in EVT (basal plate) at later gestational ages (20 weeks and beyond). All panels show tissue at a total magnification of ×100, except for the top right panel, which is a magnified version of the red boxed area in the top left panel (magnified ×400); within the top right panel, arrows show HTRA4 expression in villous STB and arrowheads highlight absence of HTRA4 in villous CTB. ∗P < 0.05, ∗∗∗P < 0.001, and ∗∗∗∗P < 0.0001.
Next, in situ hybridization was performed with HTRA4-specific probes, on human placental tissue sections across gestation. HTRA4 RNA was enriched in both STB and EVT early in gestation but remained enriched only in EVT as gestation progressed (Figure 2C). Collectively, these data confirm that HTRA4 and p63 have an inverse expression pattern, with HTRA4 being confined to differentiated trophoblast, and particularly enriched in EVT.
p63 Transcription Factor Targets Both the ΔNp63α and HTRA4 Promoters
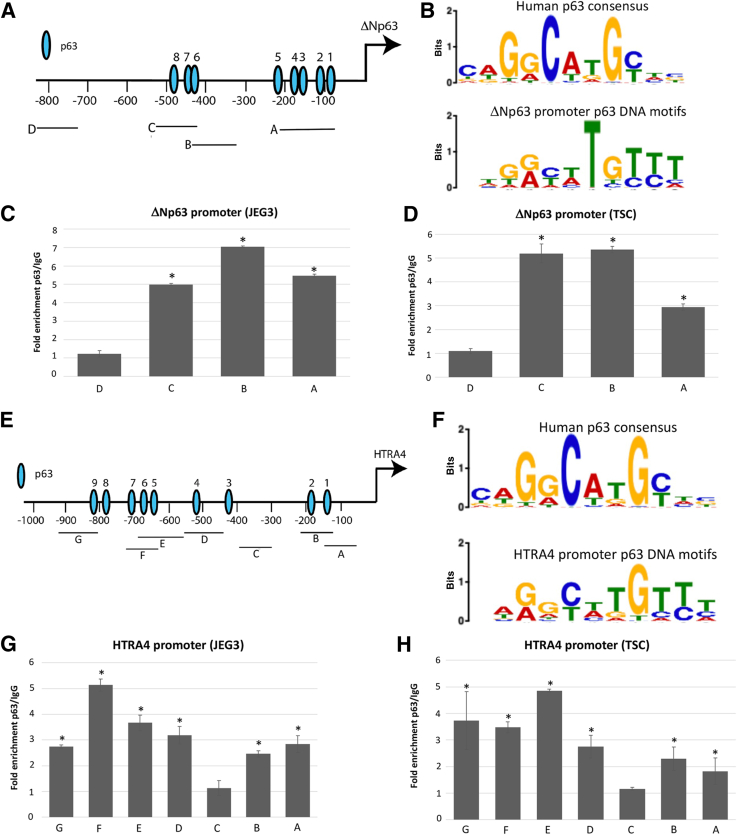
Whether p63 bound directly to the p63 and HTRA4 promoters was investigated next. First, p63 itself was evaluated, as previous studies in keratinocytes34 have suggested autoregulation. An in silico analysis was used to survey the ΔNp63 promoter for p63 transcription factor binding sites (Figure 3A). Eight consensus p63 DNA motifs were identified within the approximately 1.0-kb region upstream of the transcriptional start site (Figure 3B). Whether p63 occupied the ΔNp63 promoter by performing ChIP in JEG3 and trophoblast stem cells was addressed next. Real-time PCR using four primers sets (A through D), three of which represented the eight potential p63 binding sites, predicted by the in silico analysis, flanking the p63 promoter, was performed to quantitate the amount of ΔNp63 occupancy. The p63 bound the ΔNp63 promoter in three of the three regions (represented by primer sets A to C) with signal above background (IgG control), and not in the negative control region (represented by primer set D), in both JEG3 and TSC lines (Figure 3, C and D).
Figure 3.
The ΔNp63α (p63) transcription factor occupies the ΔNp63α and high-temperature requirement A4 (HTRA4) promoters. A: Map of p63 transcription factor binding sites (1 to 8) within the ΔNp63 promoter; primer sets spanning these sites are shown under the map, labeled A through D (D was used as a negative control). B: p63 Binding motif consensus within ΔNp63α promoter. Top panel: Human p63 binding motif consensus. Bottom panel:ΔNp63α promoter p63 consensus. P value for the p63 DNA motifs at ΔNp63 promoter is 1.48 × 10–3. C and D: Chromatin immunoprecipitation (ChIP) analysis showing fold enrichment of p63 binding (over IgG control) at each region in the TP63 promoter in JEG3 (C) and trophoblast stem cells (TSCs; D). E: Map of p63 transcription factor binding sites (1 to 9) within the HTRA4 promoter. Primer sets spanning these sites are shown under the map, labeled (A to G (C was used as a negative control). F: The p63 binding motif consensus within the HTRA4 promoter. Top panel: Human p63 binding motif consensus. Bottom panel: HTRA4 promoter p63 binding motif consensus. P value for p63 DNA motifs at HTRA4 promoter is 1.48 × 10–4. G and H: ChIP analysis showing fold enrichment of p63 binding (over IgG control) at each region in the HTRA4 promoter in JEG3 (G) and TSC (H). ∗P < 0.05.
HTRA4 was evaluated as a differentially expressed gene downstream of p63. An in silico analysis was used to evaluate the approximately 1.0-kb region upstream of the HTRA4 transcriptional start site for p63 DNA-binding sites. The query yielded nine possible p63 binding sites (Figure 3E) and identified a p63 consensus binding motif for the HTRA4 promoter (Figure 3F). Following ChIP for p63 in JEG3 and TSC, real-time PCR was performed with seven primer sets (A through G), six of which represented the nine potential p63 binding sites, predicted by the in silico analysis, flanking the HTRA4 promoter. p63 bound all regions with predicted p63 binding sites (represented by primer sets A, B, and D through G) with signal above background (IgG control) (Figure 3, G and H), and not the negative control region (represented by primer set C), in both JEG3 and TSC lines. These results show that p63 occupies the HTRA4 promoter region, suggesting that HTRA4 is a direct target of p63 in both JEG3 and TSC.
Discussion
p63 (specifically the ΔNp63α isoform) is a specific marker of proliferative CTB in the human placenta, with a role in both induction and maintenance of the CTB stem cell fate.13,15,16 Much is known about how p63 functions in other tissues, particularly those composed of stratified epithelia, including skin, cervix, and esophagus14,35,36; however, virtually nothing is known about how this protein performs its various functions in trophoblasts, the epithelial cells of the placenta. This study used microarray-based RNA profiling to evaluate genes up-regulated or down-regulated downstream of p63. JEG3 cells were used, as previously established stable lines that either overexpressed ΔNp63α or knocked it down using lentiviral shRNA were available.15 One gene, HTRA4, whose expression was reciprocally altered in both the overexpressing and knockdown cells and is known to play a role in trophoblast differentiation and function as a direct target of p63, was confirmed using quantitative real-time ChIP-PCR, in both JEG3 and TSC lines.
Similar to proliferating CTB in the placenta, ΔNp63α is the most abundant isoform in stem cells of stratified epithelia, such as skin, and has been termed the master regulator of epithelial stem cell identity.37 The villous trophoblast layer of the human placenta is akin to a stratified epithelium in the first trimester, with CTB as the basal cells, adjacent to the villous stromal core, either fusing to form STB in floating villi or undergoing epithelial-mesenchymal transition through the cell column in anchoring villi to form EVT.13 p63 exerts a positive regulatory effect/autoregulation on its own promoter.35 Here, p63 occupation was identified at multiple sites within its own proximal promoter in both JEG3 and primary trophoblast stem cells. These results suggest that p63 may function as a pioneer transcription factor in trophoblast, able to target its own promoter and remodel inaccessible chromatin as described for human epidermal development.38 Further studies are needed to address this particular role for ΔNp63α in the placenta.
The microarray data in this study show several genes up-regulated or down-regulated following p63 overexpression and knockdown. This suggests that p63 may function in the trophoblast as both an activator and a repressor, depending on its transcriptional context. Interestingly, in the overexpression experiment, the numbers of genes up-regulated and down-regulated were similar (30 versus 34). However, in the knockdown experiment, the number of genes derepressed was approximately three times the number of repressed genes (19 versus 6). This may indicate a dominant repressive role for p63 in trophoblast differentiation. This role as a transcriptional repressor, including at the HTRA4 promoter, can occur through several mechanisms. One such mechanism would be for p63 to recruit a transcriptional repressor or repressor complex to the HTRA4 promoter. In fact, p63 associates with histone deacetylases, HDAC1 and HDAC2, to promote cellular survival in squamous cell carcinoma.39 p63 might also recruit the polycomb group repressor complexes (PRC1 or PRC2) to the HTRA4 promoter. The PRC1 subunit, chromobox homolog 4, is a transcriptional repressor that acts downstream of p63 to maintain epithelial stem cell identity and proliferation in human40 and mouse epidermis41 and mouse thymus.42 Alternatively, p63 may inhibit binding of a transcriptional activator to the HTRA4 promoter. This may be direct or indirect repression through another transcription factor, such as glial cells missing-1, a master transcriptional regulator of trophoblast differentiation,43 which is directly upstream of HTRA4 in trophoblast.29 Recently, Wang et al44 described a functional antagonism between p63 and glial cells missing-1 in term CTB, showing that TSC culture conditions stabilized p63, which, in turn, reduced glial cells missing-1 transcriptional activity. Further studies are needed to probe binding partners of p63 in CTB/TSC.
Although significantly more work remains to be done to identify gene networks downstream of p63 in the placenta, these data begin to elucidate the mechanisms by which p63 exerts its effects on trophoblast differentiation and function. A major limitation of this study is the use of JEG3 cells, which are not fully representative of CTB/TSC in vivo.45 Studies of p63 targets in other tissues have identified integrins (specifically ITGA6 and ITGB4) as direct transcriptional targets of p6335; of note, villous CTBs are characterized by expression of α6β4 integrin,46 which is an essential component of hemidesmosomes, required for the stable adhesion of basal epithelial cells to the underlying basement membrane. These integrins were not identified as direct targets of p63 in JEG3 cells. While this may be a distinction between trophoblast and other epithelia, it is also possible that, as choriocarcinoma cells, JEG3 cells are suboptimal for these studies. Therefore, future work should focus on identifying both p63 DNA-binding sites and p63 binding partners in primary CTB and TSC.
Disclosure Statement
None declared.
Footnotes
Supported by NIH grants R01-HD104805 (M.M.P.) and R01-HD096260 (F.S.). Data management, storage, and analysis were performed using the Extreme Science and Engineering Discovery Environment Expanse at the San Diego Supercomputing Center through allocation MCB140074.
Supplemental material for this article can be found at http://doi.org/10.1016/j.ajpath.2024.03.011.
Supplemental Data
References
- 1.Knöfler M., Haider S., Saleh L., Pollheimer J., Gamage T.K.J.B., James J. Human placenta and trophoblast development: key molecular mechanisms and model systems. Cell Mol Life Sci. 2019;76:3479–3496. doi: 10.1007/s00018-019-03104-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.James J.L., Carter A.M., Chamley L.W. Human placentation from nidation to 5 weeks of gestation: part I: what do we know about formative placental development following implantation? Placenta. 2012;33:327–334. doi: 10.1016/j.placenta.2012.01.020. [DOI] [PubMed] [Google Scholar]
- 3.Turco M.Y., Moffett A. Development of the human placenta. Development. 2019;146 doi: 10.1242/dev.163428. [DOI] [PubMed] [Google Scholar]
- 4.Cartwright J.E., Fraser R., Leslie K., Wallace A.E., James J.L. Remodelling at the maternal-fetal interface: relevance to human pregnancy disorders. Reproduction. 2010;140:803–813. doi: 10.1530/REP-10-0294. [DOI] [PubMed] [Google Scholar]
- 5.Farah O., Nguyen C., Tekkatte C., Parast M.M. Trophoblast lineage-specific differentiation and associated alterations in preeclampsia and fetal growth restriction. Placenta. 2020;102:4–9. doi: 10.1016/j.placenta.2020.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.DaSilva-Arnold S., James J.L., Al-Khan A., Zamudio S., Illsley N.P. Differentiation of first trimester cytotrophoblast to extravillous trophoblast involves an epithelial-mesenchymal transition. Placenta. 2015;36:1412–1418. doi: 10.1016/j.placenta.2015.10.013. [DOI] [PubMed] [Google Scholar]
- 7.Wakeland A.K., Soncin F., Moretto-Zita M., Chang C.-W., Horii M., Pizzo D., Nelson K.K., Laurent L.C., Parast M.M. Hypoxia directs human extravillous trophoblast differentiation in a hypoxia-inducible factor-dependent manner. Am J Pathol. 2017;187:767–780. doi: 10.1016/j.ajpath.2016.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Morey R., Farah O., Kallol S., Requena D.F., Meads M., Moretto-Zita M., Soncin F., Laurent L.C., Parast M.M. Transcriptomic drivers of differentiation, maturation, and polyploidy in human extravillous trophoblast. Front Cell Dev Biol. 2021;9 doi: 10.3389/fcell.2021.702046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Vu H.T.H., Scott R.L., Iqbal K., Soares M.J., Tuteja G. Core conserved transcriptional regulatory networks define the invasive trophoblast cell lineage. Development. 2023;150 doi: 10.1242/dev.201826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Haider S., Lackner A.I., Dietrich B., Kunihs V., Haslinger P., Meinhardt G., Maxian T., Saleh L., Fiala C., Pollheimer J., Latos P.A., Knöfler M. Transforming growth factor-[beta] signaling governs the differentiation program of extravillous trophoblasts in the developing human placenta. Proc Natl Acad Sci U S A. 2022;119 doi: 10.1073/pnas.2120667119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Saha B., Ganguly A., Home P., Bhattacharya B., Ray S., Ghosh A., Rumi M.A.K., Marsh C., French V.A., Gunewardena S., Paul S. TEAD4 ensures postimplantation development by promoting trophoblast self-renewal: an implication in early human pregnancy loss. Proc Natl Acad Sci U S A. 2020;117:17864–17875. doi: 10.1073/pnas.2002449117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Meinhardt G., Haider S., Kunihs V., Saleh L., Pollheimer J., Fiala C., Hetey S., Feher Z., Szilagyi A., Than N.G., Knöfler M. Pivotal role of the transcriptional co-activator YAP in trophoblast stemness of the developing human placenta. Proc Natl Acad Sci U S A. 2020;117:13562–13570. doi: 10.1073/pnas.2002630117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lee Y., Kim K.-R., McKeon F., Yang A., Boyd T.K., Crum C.P., Parast M.M. A unifying concept of trophoblastic differentiation and malignancy defined by biomarker expression. Hum Pathol. 2007;38:1003–1013. doi: 10.1016/j.humpath.2006.12.012. [DOI] [PubMed] [Google Scholar]
- 14.Pecorari R., Bernassola F., Melino G., Candi E. Distinct interactors define the p63 transcriptional signature in epithelial development or cancer. Biochem J. 2022;479:1375–1392. doi: 10.1042/BCJ20210737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Li Y., Moretto-Zita M., Leon-Garcia S., Parast M.M. p63 Inhibits extravillous trophoblast migration and maintains cells in a cytotrophoblast stem cell-like state. Am J Pathol. 2014;184:3332–3343. doi: 10.1016/j.ajpath.2014.08.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li Y., Moretto-Zita M., Soncin F., Wakeland A., Wolfe L., Leon-Garcia S., Pandian R., Pizzo D., Cui L., Nazor K., Loring J.F., Crum C.P., Laurent L.C., Parast M.M. BMP4-directed trophoblast differentiation of human embryonic stem cells is mediated through a [Delta]Np63+ cytotrophoblast stem cell state. Development. 2013;140:3965–3976. doi: 10.1242/dev.092155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Okae H., Toh H., Sato T., Hiura H., Takahashi S., Shirane K., Kabayama Y., Suyama M., Sasaki H., Arima T. Derivation of human trophoblast stem cells. Cell Stem Cell. 2018;22:50–63.e6. doi: 10.1016/j.stem.2017.11.004. [DOI] [PubMed] [Google Scholar]
- 18.Tsunoda T., Takagi T. Estimating transcription factor bindability on DNA. Bioinformatics. 1999;15:622–630. doi: 10.1093/bioinformatics/15.7.622. [DOI] [PubMed] [Google Scholar]
- 19.Gupta S., Stamatoyannopoulos J.A., Bailey T.L., Noble W.S. Quantifying similarity between motifs. Genome Biol. 2007;8:R24. doi: 10.1186/gb-2007-8-2-r24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Donohoe M.E., Zhang L.-F., Xu N., Shi Y., Lee J.T. Identification of a Ctcf cofactor, Yy1, for the X chromosome binary switch. Mol Cell. 2007;25:43–56. doi: 10.1016/j.molcel.2006.11.017. [DOI] [PubMed] [Google Scholar]
- 21.Wang W., Zhang Z., Zhao M., Wang Y., Ge Y., Shan L. Zinc-finger protein CXXC5 promotes breast carcinogenesis by regulating the TSC1/mTOR signaling pathway. J Biol Chem. 2023;299 doi: 10.1016/j.jbc.2022.102812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lofthouse E.M., Brooks S., Cleal J.K., Hanson M.A., Poore K.R., O'Kelly I.M., Lewis R.M. Glutamate cycling may drive organic anion transport on the basal membrane of human placental syncytiotrophoblast. J Physiol. 2015;593:4549–4559. doi: 10.1113/JP270743. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Song L., Huo X., Li X., Xu X., Zheng Y., Li D., Zhang J., Wang K., Wang L., Wu Z. SERPINF1 mediates tumor progression and stemness in glioma. Genes. 2023;14:580. doi: 10.3390/genes14030580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Chen B., Feng M., Yao Z., Zhang Z., Zhang K., Zhou L. Hypoxia promotes thyroid cancer progression through HIF1[alpha]/FGF11 feedback loop. Exp Cell Res. 2022;416 doi: 10.1016/j.yexcr.2022.113159. [DOI] [PubMed] [Google Scholar]
- 25.Asanoma K., Matsuda T., Kondo H., Kato K., Kishino T., Niikawa N., Wake N., Kato H. NECC1, a candidate choriocarcinoma suppressor gene that encodes a homeodomain consensus motif. Genomics. 2003;81:15–25. doi: 10.1016/s0888-7543(02)00011-3. [DOI] [PubMed] [Google Scholar]
- 26.Xie J., Zhu H., Chang H.-M., Klausen C., Dong M., Leung P.C.K. GDF8 promotes the cell invasiveness in human trophoblasts by upregulating the expression of follistatin-like 3 through the ALK5-SMAD2/3 signaling pathway. Front Cell Dev Biol. 2020;8 doi: 10.3389/fcell.2020.573781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Haider S., Meinhardt G., Saleh L., Fiala C., Pollheimer J., Knöfler M. Notch1 controls development of the extravillous trophoblast lineage in the human placenta. Proc Natl Acad Sci U S A. 2016;113:E7710–E7719. doi: 10.1073/pnas.1612335113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zeng Y., He Y., Yang F., Mooney S.M., Getzenberg R.H., Orban J., Kulkarni P. The cancer/testis antigen prostate-associated gene 4 (PAGE4) is a highly intrinsically disordered protein. J Biol Chem. 2011;286:13985–13994. doi: 10.1074/jbc.M110.210765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wang L.-J., Cheong M.-L., Lee Y.-S., Lee M.-T., Chen H. High-temperature requirement protein A4 (HtrA4) suppresses the fusogenic activity of syncytin-1 and promotes trophoblast invasion. Mol Cell Biol. 2012;32:3707–3717. doi: 10.1128/MCB.00223-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Mansilla M., Wang Y., Lim R., Palmer K., Nie G. HtrA4 is up-regulated during trophoblast syncytialization and BeWo cells fail to syncytialize without HtrA4. Sci Rep. 2021;11 doi: 10.1038/s41598-021-93520-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pei C.-Z., Choi B.-C., Park J.-H., Park H.Y., Paek J., Lee K.-J., Yun B.-S., Kim Y.J., Baek K.-H. Cellular functions of high-temperature requirement factor A4 in placenta. Cells. 2023;12:1459. doi: 10.3390/cells12111459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Singh H., Zhao M., Chen Q., Wang Y., Li Y., Kaitu’u-Lino T.J., Tong S., Nie G. Human HtrA4 expression is restricted to the placenta, is significantly up-regulated in early-onset preeclampsia, and high levels of HtrA4 cause endothelial dysfunction. J Clin Endocrinol Metab. 2015;100:E936–E945. doi: 10.1210/jc.2014-3969. [DOI] [PubMed] [Google Scholar]
- 33.Liu C., Xing F., He Y., Zong S., Luo C., Li C., Duan T., Wang K., Zhou Q. Elevated HTRA1 and HTRA4 in severe preeclampsia and their roles in trophoblast functions. Mol Med Rep. 2018;18:2937–2944. doi: 10.3892/mmr.2018.9289. [DOI] [PubMed] [Google Scholar]
- 34.Antonini D., Rossi B., Han R., Minichiello A., Di Palma T., Corrado M., Banfi S., Zannini M., Brissette J.L., Missero C. An autoregulatory loop directs the tissue-specific expression of p63 through a long-range evolutionarily conserved enhancer. Mol Cell Biol. 2006;26:3308–3318. doi: 10.1128/MCB.26.8.3308-3318.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Viganò M.A., Mantovani R. Hitting the numbers: the emerging network of p63 targets. Cell Cycle. 2007;6:233–239. doi: 10.4161/cc.6.3.3802. [DOI] [PubMed] [Google Scholar]
- 36.Trink B., Osada M., Ratovitski E., Sidransky D. p63 Transcriptional regulation of epithelial integrity and cancer. Cell Cycle. 2007;6:240–245. doi: 10.4161/cc.6.3.3803. [DOI] [PubMed] [Google Scholar]
- 37.Melino G., Memmi E.M., Pelicci P.G., Bernassola F. Maintaining epithelial stemness with p63. Sci Signal. 2015;8 doi: 10.1126/scisignal.aaa1033. [DOI] [PubMed] [Google Scholar]
- 38.Yu X., Singh P.K., Tabrejee S., Sinha S., Buck M.J. [Delta]Np63 is a pioneer factor that binds inaccessible chromatin and elicits chromatin remodeling. Epigenetics Chromatin. 2021;14:20. doi: 10.1186/s13072-021-00394-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ramsey M.R., He L., Forster N., Ory B., Ellisen L.W. Physical association of HDAC1 and HDAC2 with p63 mediates transcriptional repression and tumor maintenance in squamous cell carcinoma. Cancer Res. 2011;71:4373–4379. doi: 10.1158/0008-5472.CAN-11-0046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Luis N.M., Morey L., Mejetta S., Pascual G., Janich P., Kuebler B., Cozutto L., Roma G., Nascimento E., Frye M., Di Croce L., Benitah S.A. Regulation of human epidermal stem cell proliferation and senescence requires polycomb-dependent and -independent functions of Cbx4. Cell Stem Cell. 2011;9:233–246. doi: 10.1016/j.stem.2011.07.013. [DOI] [PubMed] [Google Scholar]
- 41.Mardaryev A.N., Liu B., Rapisarda V., Poterlowicz K., Malashchuk I., Rudolf J., Sharov A.A., Jahoda C.A., Fessing M.Y., Benitah S.A., Xu G.-L., Botchkarev V.A. Cbx4 maintains the epithelial lineage identity and cell proliferation in the developing stratified epithelium. J Cell Biol. 2016;212:77–89. doi: 10.1083/jcb.201506065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Liu B., Liu Y.-F., Du Y.-R., Mardaryev A.N., Yang W., Chen H., Xu Z.-M., Xu C.-Q., Zhang X.-R., Botchkarev V.A., Zhang Y., Xu G.-L. Cbx4 regulates the proliferation of thymic epithelial cells and thymus function. Development. 2013;140:780–788. doi: 10.1242/dev.085035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jeyarajah M.J., Jaju Bhattad G., Kelly R.D., Baines K.J., Jaremek A., Yang F.-H.P., Okae H., Arima T., Dumeaux V., Renaud S.J. The multifaceted role of GCM1 during trophoblast differentiation in the human placenta. Proc Natl Acad Sci U S A. 2022;119 doi: 10.1073/pnas.2203071119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wang L.-J., Chen C.-P., Lee Y.-S., Ng P.-S., Chang G.-D., Pao Y.-H., Lo H.-F., Peng C.-H., Cheong M.-L., Chen H. Functional antagonism between ΔNp63[alpha] and GCM1 regulates human trophoblast stemness and differentiation. Nat Commun. 2022;13:1626. doi: 10.1038/s41467-022-29312-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Apps R., Murphy S.P., Fernando R., Gardner L., Ahad T., Moffett A. Human leucocyte antigen (HLA) expression of primary trophoblast cells and placental cell lines, determined using single antigen beads to characterize allotype specificities of anti-HLA antibodies. Immunology. 2009;127:26–39. doi: 10.1111/j.1365-2567.2008.03019.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Aplin J.D. Expression of integrin alpha 6 beta 4 in human trophoblast and its loss from extravillous cells. Placenta. 1993;14:203–215. doi: 10.1016/s0143-4004(05)80261-9. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.