Abstract
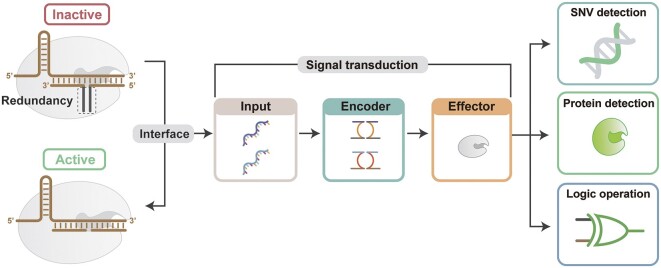
The revolutionary technology of CRISPR/Cas has reshaped the landscape of molecular biology and molecular engineering. This tool is of interest to researchers in multiple fields, including molecular diagnostics, molecular biochemistry circuits, and information storage. As CRISPR/Cas spreads to more niche areas, new application scenarios and requirements emerge. Developing programmability and compatibility of CRISPR/Cas becomes a critical issue in the new phase. Here, we report a redundancy-based modular CRISPR/Cas12a synergistic activation platform (MCSAP). The position, length, and concentration of the redundancy in the split DNA activators can finely regulate the activity of Cas12a. With the redundant structure as an interface, MCSAP serves as a modular plug-in to seamlessly integrate with the upstream molecular network. MCSAP successfully performs three different tasks: nucleic acid detection, enzyme detection, and logic operation. MCSAP can work as an effector for different molecular networks because of its compatibility and programmability. Our platform provides powerful yet easy-to-use tools and strategies for the fields of DNA nanotechnology, molecular engineering, and molecular biology.
Graphical Abstract
Graphical Abstract.
Introduction
Clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated (Cas) proteins come from the acquired immune system of prokaryotes that resist invasion by exogenous genetic elements in phages or plasmids. The CRISPR system can engage the guide RNA (gRNA) such as CRISPR RNA (crRNA) or single-guide RNA (sgRNA) to direct Cas proteins to target and process exogenous nucleic acids (1). Based on the powerful functionality and biological properties of CRISPR/Cas, this revolutionary technology has changed the paradigm of molecular biology, including gene editing (2) and gene therapy (3). In addition, based on the nuclease activity of CRISPR/Cas and the programmability brought about by Watson-crick base-pairing of gRNA, this tool has attracted the attention of researchers in more fields, especially in DNA nanotechnology (4).
Based on high programmability, DNA nanotechnology has already shown promising prospects in many areas, such as molecular detection (5,6), cell imaging (7), logic operations (8), biochemical circuits (9) and information storage (10). Due to their excellent biological compatibility and common applications, the CRISPR/Cas system and DNA nanotechnology are often combined upstream and downstream, demonstrating tremendous potential for joint applications in these fields (4,11–13). However, the inherent performance of CRISPR/Cas is no longer sufficient for the increasing number of niche applications nor the association with other molecular tools. Therefore, a central direction that researchers are currently working on is improving the programmability and controllability of the CRISPR/Cas.
CrRNA is responsible for the recognition function and is the core component of the CRISPR/Cas. A great deal of work has centered around the modification of crRNA. The design of circular crRNAs (14), light-controlled crRNA activation (15,16), G-quadruplex-guided RNAs (17,18), strand displacement gRNAs (19), methylation of gRNA (20) and orthogonal enzyme-inducible crRNAs (21) work to achieve signaling switches and activity-regulated control of the CRISPR/Cas. Although these methods provide new control modes, the scalability of the structures under specific design is often limited and does not easily integrate with upstream molecular networks. On the other hand, most of these control modes are still coarse OFF/ON modes that do not allow for fine activity regulation. The optimal crRNA in the face of different scenarios often requires extensive experimental optimization.
Outside of crRNAs, programmable modification of activators has great scope for development and has recently attracted the attention of researchers. Researchers used a block strand of activators to realize non-nucleic acid target detection (22,23). Toehold-mediated strand displacement (TMSD) was used for crRNA activation, demonstrating the fine regulation ability of CRISPR/Cas (24,25). These works have broadened the scope of applications of CRISPR/Cas, but some limitations and shortcomings have also been revealed. Most of the work relies on complete activators, which limits the design of molecular reaction networks upstream of CRISPR/Cas. Using DNA strands for closure and opening is a common strategy, but most studies realized the regulation with the aptamer (11,26), which is fixed in structure and sequence, reducing programmability.
Two recent works in split activators demonstrate the possibility of non-complete activators, which use the split activator to achieve RNA detection (27) and synergistic activation (28). However, they both use two split activators for direct activation, so the current designs do not enable functionality beyond direct detection of nucleic acid targets, nor are they easily compatible with upstream molecular networks. It is still unknown how programmability can be controlled using split activators. Therefore, developing a programmable regulated synergistic activation strategy can better meet the needs for CRISPR/Cas application in different fields and collaboration with different molecular tools.
In this work, we found that the redundant structure of the two split activators significantly affects CRISPR/Cas activity. Fine control of CRISPR/Cas12a activity can be realized through the position, length, and concentration of the redundancy in the split activators. The activity of CRISPR/Cas12a could be blocked through a specific redundancy structure or in the absence of one split activator. Based on these findings, we reported a redundancy-based modular CRISPR/Cas12a synergistic activation platform (MCSAP). By removing the redundancy structure, we realized single nucleotide variants (SNV) and APE 1 detection using MCSAP. Furthermore, we employed MCSAP as a downstream effector and demonstrated its capability to perform logical operations on arbitrary nucleic acid inputs via the upstream split DNAzyme acting as a converter. The versatility of MCSAP allows for its integration with various molecular tools, enabling a seamless interface with intricate and comprehensive molecular reaction networks, thereby enhancing its accessibility for researchers in the field.
Materials and methods
Materials and apparatus
All oligonucleotides used in this study (sequences in Supplementary Table S1) were synthesized and purified by Sangon Biotech Co., Ltd (Shanghai, China). EnGen Lba Cas12a (Cpf1, LbCas12a) and 10 × NEBuffer 4 were brought from New England Biolabs Inc (Beijing, China). Deionized water (DNase/RNase free) was obtained from Tiangen Biotech (Beijing, China) and used in all experiments. Endo IV was purchased from Thermo Fisher Scientific. APE1, T7 Exonuclease (T7 Exo), Exonuclease I (Exo I), Deoxyribonuclease I (DNase I), and Uracil glycosylase (UDG) were all obtained from New England Biolabs (NEB, MA, USA). Rotor-Gene Q2 plex HRM Instrument (QIAGEN, Hilden, Germany) was used to measure the fluorescence intensity of the samples.
Exploration and construction of synergistically activated control system
To a 200 μl PCR tube, 2 μl of Cas12a (100 nM), 2 μl of crRNA (200 nM), 2 μl of NEBuffer 4 (10×) and 2 μl of ssDNA reporter (1 μM) were added. Then 2 μl of each splitting activator (200 nM) was added and the total volume was brought up to 20 μl by deionized water. The fluorescence intensity was immediately recorded once a cycle (15 s per cycle) at 37°C by Rotor-Gene Q 2plex HRM Instrument.
Single nucleotide variants (SNV) detection
To a 200 μl PCR tube, 2 μl of AP site-Probe a1-i/a2-i (200 nM), 3 μl of the target strands (MT or WT, 200 nM), 1.5 μl of (NH4)2SO4 (400 mM) and 2 μl of Endo IV (0.02 U/μl) were mixed. After 15 min incubation at 37°C, the above solution was heated at 80°C for 20 min to inactive Endo IV. Then, 2 μl of Cas12a (100 nM), 2 μl of crRNA (200 nM), 2.5 μl of NEBuffer 4 (10×), 2 μl of ssDNA reporter(1 μM), and 2 μl of Mn2+ (25 mM) were added. The total volume was brought up to 25 μl by deionized water.
Decoding of drug metabolism gene profiles
To a 200 μl PCR tube, 1 μl of AP site-Probe a1-i or a2-i for per target gene (400 nM), 1 μl of per target strand (target gene 600 nM), 1.5 μl of (NH4)2SO4 (400 mM) and 2 μl of Endo IV (0.02 U/μl) were mixed. After 15 min incubation at 37°C, the above solution was heated at 80°C for 20 min to inactive Endo IV. Then, 2 μl of Cas12a (100 nM), 2 μl of crRNA (200 nM), 2.5 μl of NEBuffer 4 (10×), 2 μl of ssDNA reporter (1 μM), and 2 μl of Mn2+ (25 mM) were added. The total volume was brought up to 25 μl by deionized water. Each tube contains probes corresponding to different genes, and when different target genes are added, the probe corresponding to each target gene produces a signal value (weight coefficient). The weight sum for each channel is obtained by summing the weight. Channels m and n are separated into different tubes for detection.
Apurinic/apyrimidinic endonuclease 1 (APE 1) detection
To a 200 μl PCR tube, 2 μl of AP site-Probe (200 nM), 3 μl of the complementary strands (200 nM), 1.5 μl of NEBuffer 4 (10×), and 2 μl of APE1 (at various concentrations) were mixed. After 30 min incubation at 37°C, the above solution was heated at 80°C for 20 min to inactive APE1. Then, 2 μl of Cas12a (100 nM), 2 μl of crRNA (200 nM), 1 μl of NEBuffer 4 (10×), 2 μl of ssDNA reporter (1 μM), and 2 μl of another split activator strand (200 nM) were added and the total volume was brought up to 25 μl by deionized water.
To study the selectivity of the method, DNase I, UDG, T7 exo, and Exo I were selected as the potential interfering enzymes to compare with APE1. APE1 and other enzymes' activity was detected using the same procedure above. The concentration of all enzymes was 2 U/ml in the solution.
For APE1 detection in biological samples, the method is the same as the previous report. The cell lysate (containing about 40 000 cells/μl) was diluted to 10 cells/μl and 1 cell/μl. 2.5 μl of the diluted cell lysate was used to replace APE1 in the above APE1 detection system. The cell lines were cultured in a humidified incubator at 37°C containing 5% CO2.
Logic operations and DNA circuit construction
Implement the ‘AND’ gate with the full DNAzyme as input: To a 200 μl PCR tube, 2 μl of 200 nM Substrate-1 (10–23 or 8–17), Substrate-2 (10–23 or 8–17), DNAzyme-1 (10–23 or 8–17), DNAzyme-2 (10–23 or 8–17) and 2 μl of Mg2+ (400 mM) were added. Then add the mixture of 2 μl of Cas12a (100 nM), 2 μl of crRNA (200 nM), 2 μl of NEBuffer 4 (10×) and 2 μl of ssDNA reporter (1 μM). The total volume was brought up to 20 μl by deionized water. Other logic gates are constructed under the same conditions as ‘AND’ gates, which are detailed in the supplementary material
Results and discussion
Exploration and construction of synergistically activated control system
CRISPR/Cas12a (Cpf1) belongs to a representative member of the class 2 V-A type CRISPR/Cas family and has a bilobed structure comprising nuclease and recognition domains (29). CRISPR/Cas12a can be activated by single-stranded DNA (ssDNA) without protospacer adjacent motif (PAM) sequence. Upon binding ssDNA activator by crRNA, CRISPR/Cas12a generates non-specific ssDNA cleavage activity known as trans-cleavage activity. The crRNA consisted of a hairpin domain bound with Cas12a protein and a spacer domain with around 20 nucleotides (nt).
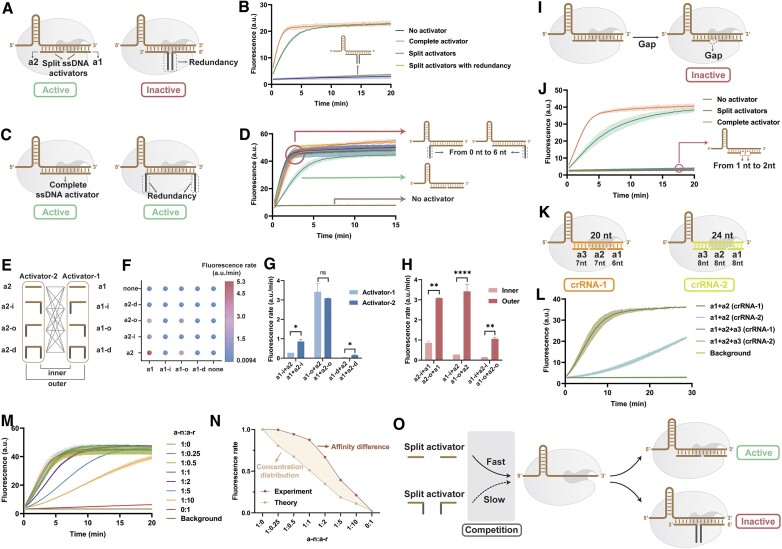
We first explored the effect of the redundant structure in the activator on Cas12a activity. We divided the complete ssDNA activator into two parts with the same 10-nt length. Activator-1 (a1) is at the 5′-end of the spacer domain, and activator-2 (a2) is at the 3′-end of the spacer domain (left of Figure 1A). For comparison, we added redundancy inside the activator, two random sequences of 6 nt, located at the 3′-end of a1 and the 5′-end of a2 (grey region in right of Figure 1A). The 15-nt ssDNA labeled with a fluorophore (FAM) and a quencher (BHQ) was used as the reporter to characterize the trans-cleavage activity of Cas12a (30). The real-time fluorescence plot of Figure 1B demonstrates that both the complete activator (orange in Figure 1B) and split activators (green in Figure 1B) activated Cas12a. The activation efficiency of the split activators is slightly lower than that of the complete activator, which may be due to the split activators’ lower affinity and structural stability. However, activators with redundancy could not activate cas12a (grey in Figure 1B). We verified the effect of the redundant structure on the complete activator (Figure 1C). We set random sequences of different lengths from 0 nt to 6 nt at both ends of the complete activator. As shown in Figure 1D, the redundancy has almost no effect on the complete activator. This suggests that the introduction of redundancy only affects split activators.
Figure 1.
(A) Scheme and (B) fluorescence plot of the synergistic activation pattern of CRISPR/Cas12a based on redundant structures. (C) Scheme and (D) fluorescence plot of the effect of redundancy on complete activators. (E) Scheme and (F) bubble plot of the positional combination pattern of split activators. The fluorescence rate was calculated based on the slope of the fluorescence curves. (G) Comparison of a-1 and a-2 with different redundancy positions. (H) Comparison of inner and outer redundancy positions with different activators. (I) Scheme and (J) fluorescence plot of the effect of crRNA and activator complementary lengths. (K) Scheme and (L) fluorescence plot of the trisection split activators. crRNA-1 and crRNA-2 have different sequences in the spacer domain. (M) Relative rate of fluorescence increase of the competition between activators without (a–n) and with redundancy (a–r). (N) Non-concentration-dominated rate distribution due to affinity differences. (O) scheme of the competition between activators without (a–n) and with redundancy (a–r).
Next, we explored the effect of redundancy's position on split activators. We categorized activators according to the redundancy's position relative to the crRNA spacer region into a–i (inner), a–o (outer) and a–d (double) (Figure 1E). Inner and outer refer to the positions away from and close to the split site of the activator, respectively. By combining four redundancy positions on a1 and a2, we systematically explored the activation degree of 16 combinations (Figure 1E, Supplementary Figures S1, S2). A random sequence of 6 nt was used for all redundancy. As demonstrated in the bubble plot of Figure 1F, the redundancy at each position significantly affects the fluorescence rate compared to a1 and a2. By comparing the data in the same row or column, a–d generally has a lower signal than a–i or a–o (Figure 1F), suggesting that redundancy's inhibition effect on both sides can be superimposed. We next explored the differences in the effects of redundancy on a1 and a2. When the redundancy is inner or double, the redundancy will have more significant inhibition at a1 than at a2 (Figure 1G). However, the difference is insignificant when the redundancy is on the outer side. This may be because the inhibitory effect of the outer redundancy is minor, and this difference could not be highlighted. We compared the difference between redundancy on the inner and outer sides. The inhibitory effect of inner redundancy is more pronounced on a1 or a2 (Figure 1H). This is consistent with the tolerance of CRISPR/Cas12a to bubble structures in double strands, with the middle of the spacer domain having a more significant inhibitory effect (31).
We next explored the effect of the activator and crRNA complementary length (Figure 1I). The crRNA-DNA complex is intolerant of medial base deletions, and a 1 nt gap can completely inhibit Cas12a activation (Figure 1J). We explored two different structures to explore the effect of the activator on CRISPR/Cas, insertion, and mismatch, respectively (Supplementary Figure S3). To demonstrate the generalizability of the results, we selected two crRNAs with spacer regions of different lengths and sequences (Supplementary Table S1). crRNA-1 is the same as the crRNAs used in the other experiments in Figure 1 and subsequent experiments. We also tested to split the activator into three parts (Figure 1K). Figure 1L shows that the activators divided equally into three parts cannot activate Cas12a. The activators split into two halves could also activate Cas12a with crRNA-2 despite some rate loss (Supplementary Figure S4).
The above results show that split activators with redundancy (a-r) can affect CRISPR/Cas12a activity. This could be attributed to two potential factors. First, this inhibition could stem from a–r's inability to bind with the crRNA effectively. Alternatively, the complex formed by a–r and crRNA may be unable to activate Cas12a fully. To investigate this, we combined the activator without redundancy (a–n) and a–r in various n:r ratios. The activity of Cas12a remained unchanged within the ratio range from 1:0 to 1:1 (the standard concentration for ‘1’ is 20 nM). When the ratio increased to 1:2, we observed a decrease in Cas12a activity (Figure 1M). However, the relative fluorescence increase rate changes are smaller than the concentration-dominated competition in theory (Figure 1N). These findings suggest that a-r can also bind to crRNA, though a–r has less affinity for CRISPR/Cas12a than a–n (Figure 1O).
Exploration of the length and location of redundancy
The complementary length of the complete activator and crRNA also affects Cas12a activity. However, the regulable range of lengths is very limited. On the other hand, a sudden drop in activity is caused by single-base length changes, and it is unlikely that ssDNAs below 15 nt can activate Cas12a (32). Thus, the complete activator lacks a method to achieve fine regulation of Cas12a activity. However, MCSAP could achieve fine regulation of Cas12a activity by controlling redundancy length and location.
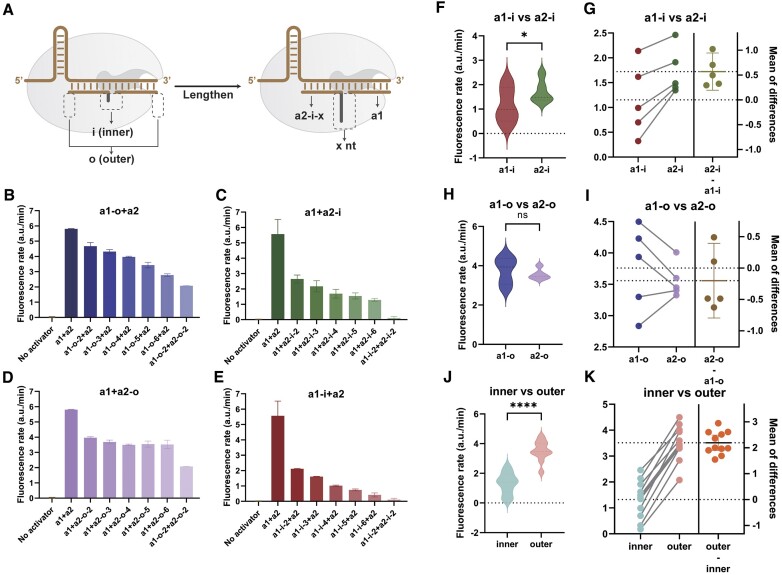
The length of the redundancy was fixed at 6 nt in experiments in Figure 1. In Figure 2A, a1-i-x denotes activator-1 with x nt of inner redundancy. We used an a1 or a2 with no redundancy as a pair and another activator with a constantly extending inner or outer redundancy. As the length of redundancy increases, the fluorescence rate decreases (Figure 2B, C, E, Supplementary Figure S5), except for outer redundancy on a2 (Figure 2D). When a1 and a2 both have redundancy, the activity of Cas12a decreases significantly, even though both ends of the redundancy are relatively short (2 nt, Figure 2B–E). The above results demonstrate the ability to finely regulate Cas12a activity by the position and length of the redundancy.
Figure 2.
(A) Schematic diagram of redundancy at different lengths and positions. a2-i-x denotes the inner side of activator-2 with a x nt of random sequences. (B–E) Fluorescence rates of redundancy of different lengths at four different positions. (F–G) Pairwise comparison of the inner side redundancy of activator-1 and activator-2. (H, I) Pairwise comparison of the outer side redundancy of activator-1 and activator-2. (J, K) Pairwise comparisons of inner and outer redundancy. Two-tailed paired t-tests were used for statistical difference analysis. *P value ≤ 0.05; **P value ≤ 0.01; ***P value ≤ 0.001; ****P value ≤ 0.0001.
Differences between a1 and a2 or inner and outer were systematically investigated by pairings of the same redundancy's length. With inner redundancy, a1 is significantly more inhibited than a2 (Figure 2F). This inhibitory effect was more evident at 4–6 nt, as single base changes in this interval were more pronounced for a1, while a2 was better tolerated (Figure 2G). There is no statistically significant difference in the overall fluorescence signal rate between the a1 and a2 groups with outer redundancy, while the different redundancy lengths on a1 result in the broader rate interval (Figure 2H). Pairwise comparisons by Figure 2I revealed that a1 was more activated than a2 within 2–4 nt redundancy's lengths, and the opposite result was observed when redundancy's length was greater than 4 nt. a1 was more responsive to single-base variations in the redundancy's length, which is consistent with the property of the inner redundancy above. We verified the difference between the inner and outer redundancy similarly. For the same redundancy's length, the redundancy located on the medial side will be more obviously inhibited than the lateral side (Figure 2J, K).
Synergistic activation mode for differential concentration
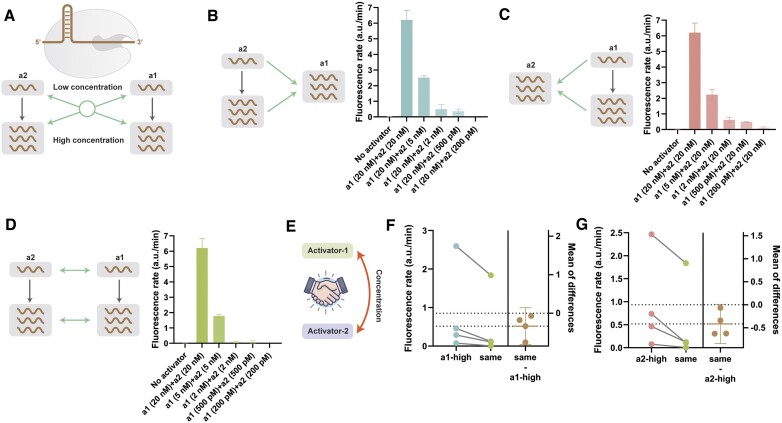
The split activators in MCSAP also allow us to regulate the activator concentrations to control the activity of Cas12a. We set up a series of concentration gradients of a1 and a2 to explore the synergistic effects (Figure 3A). We immobilized 20 nM of a1 and paired it with 20 nM, 5 nM, 2 nM, 500 pM and 200 pM of a2. As the a2 concentration decreases, the fluorescence rate decreases, and the rate is neglectable with 200 pM of a2 (Figure 3B). The same phenomenon was observed for the opposite pairing, where the fluorescence rate decreases with decreasing a1 concentration at a constant a2 concentration (Figure 3C). The same concentration of a1 and a2 from 200 nM to 200 pM was used as a control, and 2 nM of a1 and a2 could hardly activate Cas12a (Figure 3D). These results indicate that in the mode of synergistic activation, the activity of Cas12a is not determined by a particular lowest concentration of activator alone but rather by the concentration of the two activators (Figure 3E). We compared the differences between regimens with the same concentration of both activators and regimens that fixed a high activator concentration (Figure 3F, G). Regardless of whether a high concentration of a1 or a2 was used, there was a higher fluorescence rate than the regimen with the same concentration of activators (Figure 3F, G).
Figure 3.
(A) Synergistic activation mode for differential concentration. (B) A fixed a1 concentration with different a2 concentrations. (C) A fixed a1 concentration with different a2 concentrations. (D) The same concentration for a1 and a2. (E–G) Comparison of fluorescent rate for different concetration activation modes.
Split activators have a synergistic effect on concentration. This mode can be applied to a diverse range of scenarios. For example, when detecting a single target, one of the activators can be converted into an interface for target detection, using another activator as an amplifier to extend the recognition ability and sensitivity of CRISPR/Cas12a. The amplifier can compensate for the rate loss of the split activator in terms of the ability to detect targets at low concentrations, similar to the complete activator (Supplementary Figure S6). In addition, concentration-based differences can be used as a rate regulation scheme to convert upstream molecular information from different concentrations into Cas12a cleavage, resulting in a balanced rate signal. We explored the effect of a higher concentration of the accelerator strand on detection performance enhancement. An excess of accelerator strand does not serve to significantly increase the sensitivity of the detection (Supplementary Figure S7). This upper limit may be restricted by the concentration of crRNA since excess split strands cannot bind to crRNA.
The above studies demonstrate the ability of redundancy to modulate Cas12a activity. We used the redundant structure as an interface between the upstream molecular network and the CRISPR/Cas12a, taking the CRISPR/Cas12a system as a module for downstream reporting. In this way, different tasks can be performed through the upstream molecular network, and the CRISPR/Cas12a acts as a plug-in for the output of results. We next select three tasks, SNV detection, APE 1 detection, and logic circuits, as examples to demonstrate the programmability and modular application of MCSAP.
Single nucleotide variants (SNV) detection
SNV is an important biomarker for the diagnosis of tumor diseases and the selection of therapeutic options (33). In the clinical setting, amplification refractory mutation system (ARMS) PCR has been used to identify SNV (33). However, due to the complexity, high cost, and instrument limitations, researchers have devoted themselves to developing fast, simple, and cost-effective molecular diagnostic tools (34–37). CRISPR/Cas has shown great potential in molecular diagnostics, and many detection systems have achieved better sensitivity and functionality by combining the CRISPR/Cas system with DNA circuits (4,11). Based on the programmability of redundant structure, we explored the potential of the MCSAP for SNV recognition and compatibility with other methods.
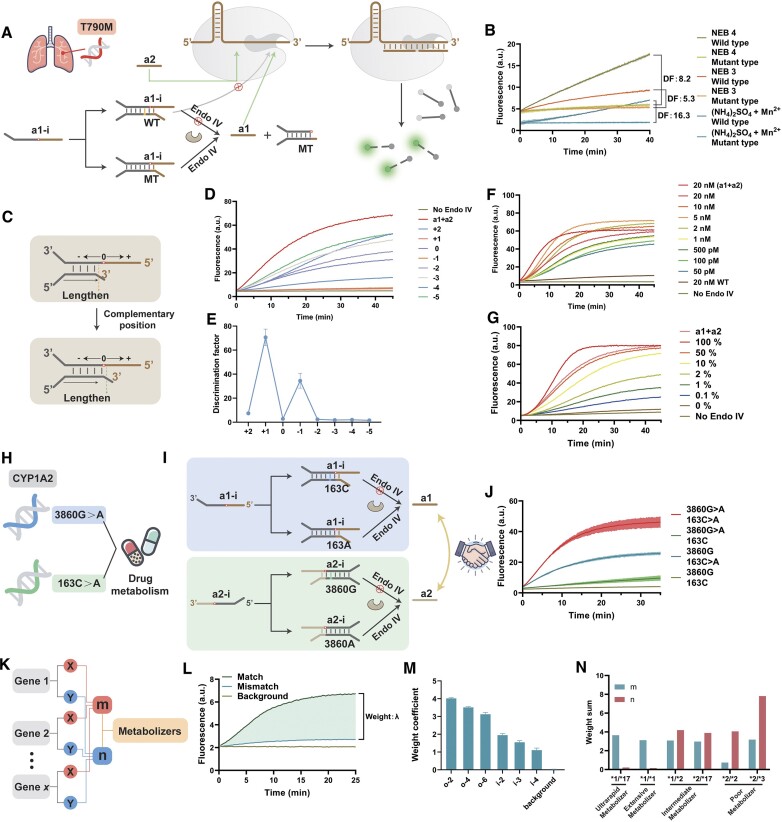
We integrated the Endo IV substrate recognition properties previously discovered by our team with the CRISPR/Cas-based reporter system, selecting the T790M mutation in lung cancer as an example. Endo IV is an apurinic/apyrimidinic (AP) endonuclease (38). It recognizes AP sites in DNA and cleaves the phosphodiester bond at the 5′-end of the damaged site, producing a hydroxyl group at the 3′-end. As shown in Figure 4A, based on the substrate recognition properties of Endo IV, we designed a probe based on the structure of a1-i. The target strand and a1-i form a double-stranded DNA (dsDNA). The wild-type (WT) target strand has a one-base mismatch with probe a1-i, thus forming an unstable structure near the cleavage site that Endo IV is difficult to recognize and cleave. The mutant-type (MT) target strand can be entirely complementary to probe a1-i and can be recognized and cleaved by Endo IV. Endo IV cleavage of a1-i produces a-1 with no redundancy, and a-2 is pre-existing in the system. Combined with a1 and a2, Cas12a exerts trans-cleavage activity and produces a fluorescence signal. Without the target strand or in combination with WT, a1-i produces a potent inhibition because the recognition region at the 3′-end of the probe (gray portion of Figure 4A) has a sufficiently long domain. Cas12a is only slightly activated.
Figure 4.
(A) Schematic diagram of single nucleotide variant detection. (B) Fluorescence plot of Endo IV operating in different buffers. DF (Discrimination factor) was defined as the ratio of fluorescence rate between a1-i/MT and a1-i/WT. (C–E) Optimization of substrate structure of Endo IV. ±x indicates that the target strand is complementary to the probe to the xth base of the 3′-end or 5′-end of the AP site. (F) Fluorescence plot of MCSAP in response to different concentrations of MT. (G) Fluorescence plot of MCSAP in response to different abundances of MT. (H) Diagram and (I) molecular process for the detection of two targets of the CYP1A2 gene. (J) Fluorescence plot of different SNP combinations. (K) Gene polymorphism decoder. (L) Fluorescence plot of weighting coefficients based on redundancy. The weight sum indicates the signal difference between the mismatch and match groups at 25 minutes after the start of the reaction. (M) Weighting coefficients for different redundancy positions and lengths. (N) Decoding of CYP2C19 gene and drug metabolism.
We first tested the working environment of Endo IV and selected (NH4)2SO4+ Mn2+ buffer because of the better discrimination factor (DF) between signal and background (Figure 4B). DF was defined as the ratio of fluorescence increase rate between a1-i/MT and a1-i/WT. Since our group had previously optimized Endo IV in other similar systems (30), we performed experiments with small gradients this time and chose 0.02 U/ul for subsequent experiments (Supplementary Figure S8). Since the selectivity of Endo IV for single base differences in the substrate arises from the structure around the cleavage site, we tested a series of dsDNA structures at different complementary positions to determine the optimal design. We defined the position of the AP site on the probe as 0, with the 5′-end of the AP site in the + direction and the 3′-end in the - direction (Figure 4C). ±x indicates that the target strand is complementary to the probe to the xth base of the 3′ end or 5′ end of the AP site, and 0 indicates that the target strand is aligned to the AP site of the probe. We found that the structures of the +1 and –1 positions were better inhibited (Figure 4D). Correspondingly, we calculated the DF of the different positions. The structure of the +1 position had the best DF to distinguish between MT and WT, which reached 70.5 (Figure 4E). We next evaluated the detection performance of MCSAP for different concentrations of MT. 20 nM a2 is pre-added in the system as an amplifier. MCSAP demonstrated excellent activity and discriminatory ability in this mode. 50 pM MT could still significantly activate Cas12a, whereas 20 nM WT had limited activation ability (Figure 4F). To assess the selectivity of MCSAP, we mixed WT and MT in different ratios to test the fluorescence response. The total concentration of WT and MT was fixed at 20 nM. Figure 4G shows the response of MCSAP to different ratios of MT. MCSAP discriminates well for mutation abundance of 0.1% (Figure 4G). Mutation abundance = cMT/(cMT + cWT) × 100%. To interrogate the performance of MCSAP in a more clinically relevant environment, we tested MCSAP in serum (Supplementary Figure S9). Fetal bovine serum (FBS, Sangon Biotech Co., Ltd) was used. The FBS was stored at –20°C. The active FBS was freshly thawed before use and diluted in TE buffer to prepare 10% FBS. Experiments were performed with 10% FBS to replace TE buffer using the same protocols using TE buffer. We found that, although serum contains many biomolecules, the detection ability of MCSAP has not been significantly affected. Next, we replaced the split activator in our detection flow with a complete activator as in the conventional CRISPR/Cas12a system (Supplementary Figure S10). The inability to exert the ability to detect SNVs in a complete activator system shows that the CRISPR/Cas12a system needs to utilize the redundancy to function as a discrimination tool.
These results demonstrate the detection capability and compatibility of MCSAP. We have realized a modular construction that can decompose tasks and assign appropriate tasks to different molecular tools and circuits. The task of SNV differentiation does not have to be done traditionally through mismatches between crRNAs and activators but can be accomplished through upstream DNA circuits.
Decoding of drug metabolism gene profiles
Over the past decade, with the widespread popularity of human genomic association studies (GWAS), many specific genetic variants have been identified in association with disease diagnoses, therapeutic responses, and physiological functions (39). For example, differences in blood exposure to drugs are an essential cause of oscillations in efficacy, with individual differences in metabolic enzyme activity being the key molecular mechanism. Cytochrome P450 (CYP450) is a superfamily of enzymes that catalyze the oxidation of exogenous substances in the body and is involved in the metabolism of approximately 90% of commonly used clinical drugs. The genes encoding the CYP enzymes are characterized by single nucleotide polymorphisms (SNPs) in the population (40). The realization of gene-guided individualized dosing is of great significance to ensure clinical drug safety (41).
We built a programmable DNA decoding circuit based on MCSAP to convert different genetic polymorphism information into rate-graded signals. MCSAP assigns different weights to corresponding genes, matching the activation capacity. Notably, due to the modular design, sequence recognition tasks are assigned to upstream molecular circuits. Thus, MCSAP avoids sequence dependency and eliminates the need for multiple crRNA designs.
We used two targets of CYP1A2 associated with drug metabolism as examples (Figure 4H). The probe recognizes SNP and is converted to activator redundancy by Endo IV (Figure 4I). Four allele combinations can be distinguished into four fluorescence rates (Figure 4J). With the tunability of redundancy, we designed weighted decoders that perform operations against multiple SNPs (Figure 4K). SNPs of different genes can be recognized and assigned corresponding weight coefficients according to the fluorescence increments (Figure 4K, L, Supplementary Figure S11). We measured the weight coefficients of different redundancies using a2 as a basis (Figure 4M, Supplementary Figure S12). We decoded the drug response of CYP2C19 concerning the coding scheme of Liu et al. (Supplementary Table S2) (41). Different metabolic types can be well distinguished (Figure 4N).
Apurinic/apyrimidinic endonuclease 1 (APE 1) detection
APE1 is a multifunctional protein that plays a vital role in the base-excision repair (BER) pathway (42). APE1 is capable of recognizing and cleaving AP sites in dsDNA. APE1 is abnormally expressed at elevated levels in various cancers, which may promote tumor development by enhancing DNA repair (5).
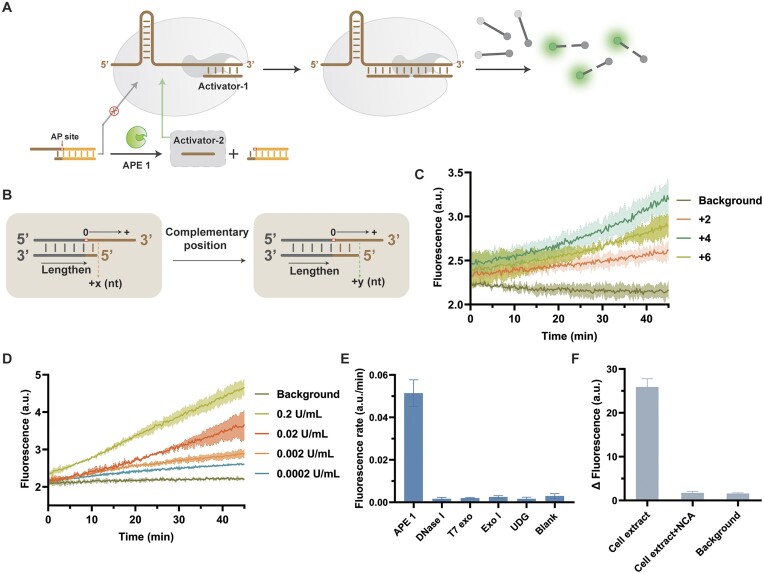
Based on AP site-mediated interfaces, MCSAP has the potential to enable the detection of APE 1. APE 1 can cleave AP sites on probes with redundancy, generating non-redundant a2, which synergistically activates CRISPR/Cas12a with pre-existing a1 (Figure 5A). We first explored the substrate preference of APE 1. We extended the complementary positions of the probes to optimize the best APE 1 substrate (Figure 5B). As shown in Figure 5C, the structure of the +4 position is more favorable for APE 1 cleavage. Based on the optimized structure, we tested the ability of MCSAP to detect different concentrations of APE 1. When the APE 1 concentration was as low as 0.0002 U/ml, MCSAP was still significantly active, creating a clear distinction from the background signal (Figure 5D). The detection ability of generating a2 alone is better compared to generating both a1 and a2 (Figure 5D, Supplementary Figure S13). Additionally, MCSAP has good selectivity for APE 1 compared to other nucleic acid enzymes that may coexist (Figure 5E).
Figure 5.
(A) Schematic diagram of APE 1 detection. (B) Schematic diagram and (C) fluorescence plot of APE 1 preferred substrate structure optimization. (D) Fluorescence plot of MCSAP for detection of different concentrations of APE 1. (E) Selectivity of MCSAP for APE 1 compared with other nucleases. (F) The ability of MCSAP to detect APE 1 in actual biological samples. Extracts of MDA-MB-231 cells that diluted to about one cell was tested. NCA is an APE1-specific inhibitor.
To validate the detection ability of the biosensor in actual biological samples, we used MCSAP to detect APE 1 in extracts of MDA-MB-231 cells (human breast cancer cells). When we diluted the cells to 10 and 1, there was still a clear detection signal in contrast to the background (Figure 5F, Supplementary Figure S14).
Logic operations and DNA circuit construction
Based on high programmability conferred by the Watson-crick base-pairing of nucleic acid materials, DNA nanotechnology has demonstrated outstanding potential for applications in fields such as information processing (43), logic operations (9), and dynamic network construction (44). DNA logic programming based on Boolean operations has been widely used in scenarios such as molecular detection (7), cancer therapy (9), and chemical network construction (44). We try to connect MCSAP with the upstream molecular network to realize the compatibility of the logic operation function (Figure 6A).
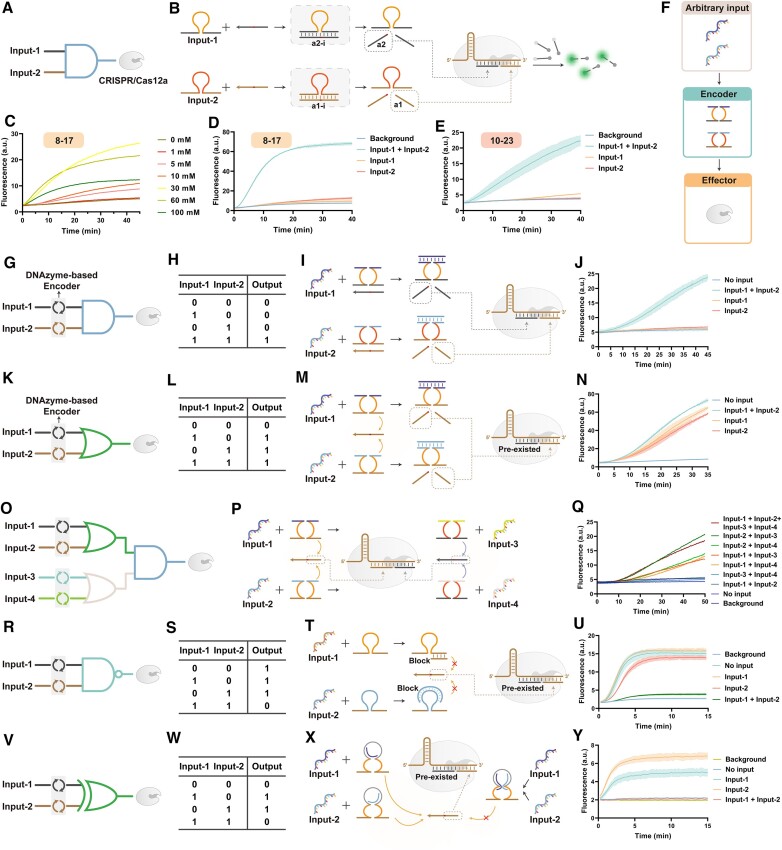
Figure 6.
(A, B) Schematic and molecular implementation of the ‘AND’ gate operation strategy of MCSAP. (C) Optimization of cofactor concentration for 8–17 DNAzyme. (D) 8–17 and (E) 10–23 performing ‘AND’ gate operations. (F) Three-layer structure: Input-Encoder-Effector for processing arbitrary nucleic acid inputs. (G) Schematic diagram, (H) truth table, (I) molecular implementation, and (J) fluorescence plot of ‘AND’ gate. (K) Schematic diagram, (L) truth table, (M) molecular realization, and (N) fluorescence plot of the ‘OR’ gate. (O) Schematic diagram, (P) molecular realization, and (Q) fluorescence plot of a two-level cascade logic gate. The two ‘OR’ gates are the first level, and the ‘AND’ gate is the second level. (R) Schematic diagram, (S) truth table, (T) molecular realization, and (U) fluorescence plot of the ‘NAND’ gate. (V) Schematic diagram, (W) truth table, (X) molecular realization and (Y) fluorescence plot of the ‘XOR’ gate.
We utilized DNAzyme-based encoders (43) as converters between MCSAP and the upstream molecular network. Dnazyme is widely used in molecular detection (11,12), molecular machines (8), and DNA circuits (45,46), owing to its dual nature of enzyme and DNA. Dnazyme can recognize and cleave substrates by base-pairing between the recognition arm and the substrate with cofactors. We first designed two DNAzyme, input-1 and input-2, based on two redundant activators, a1-i and a2-i. Their recognition arms are complementary to a1-i and a2-i, respectively. Only when both DNAzyme are present can a1 and a2 be utterly free from redundancy and have the ability to activate CRISPR/Cas12a (Figure 6B). To demonstrate the versatility of the design of the DNAzyme-based encoder, we used two DNAzyme for the demonstration, 8–17E and 10–23. We used magnesium ions (Mg2+) as cofactors for 8–17E and 10–23. The optimal Mg2+ concentration for the 8–17E and 10–23 was near 30 mM (Figure 6C, Supplementary Figure S15). Through the optimization of smaller concentration gradients, we finally chose 40 mM Mg2+ as the concentration for subsequent experiments (Supplementary Figure S16). The ‘AND’ gate worked well when using 8–17E (Figure 6D, S17) or 10–23 as inputs (Figure 6E, Supplementary Figure S18). Only when both DNAzyme inputs are present is CRISPR/Cas12a activated.
To realize arbitrary nucleic acid information input, we used a split DNAzyme-based encoder as the converter of MCSAP. We constructed MCSAP with a three-layer structure of input–encoder–effector. The encoder can process arbitrary inputs to output signals that crRNA can recognize and activate the effector-Cas12a (Figure 6F). We first constructed the ‘AND’ gate (Figure 6G). Figure 6H shows the truth table of the ‘AND’ gate. In the absence of input, the catalytic cores of the split DNAzyme are far away from each other and cannot cleave the substrate. Input can pair with the binding arms of the split DNAzyme. Then the DNAzyme recognizes and cleaves the substrate, forming the activator without redundancy (Figure 6I). Cas12a can only be activated if inputs 1 and 2 are present at the same time (Figure 6J). Next, the ‘OR’ gate was constructed (Figure 6K). Figure 6L shows the truth table of the ‘OR’ gate. Unlike the ‘AND’ gate, the ‘OR’ gate has a pre-existing a2, and either input can activate the split DNAzyme to produce a1 (Figure 6M). Cas12a can be activated by one or two inputs (Figure 6N). We also tried to generate a2 with DNAzyme and activate Cas12a with pre-existing a1, and the results are similar (Supplementary Figure S19).
On this basis, we performed a cascade of logic gates. We used a two-level cascade of OR-AND as an example for four arbitrary inputs (Figure 6O). The truth table is shown in Supplementary Figure S15. Each split activator constitutes an ‘OR’ gate by split DNAzyme, and two split activators constitute the ‘AND’ relationship (Figure 6P). We tested the cross-combination of different inputs, and the activation of Cas12a matched the truth table (Figure 6Q, Supplementary Figures S20, S21).
We next constructed the ‘NOT’ gate (Supplementary Figure S22). We used three different forms to construct the NOT gate. They are along the same idea. When the input is 0, DNAzyme can work normally and activate the downstream with the output as 1. We designed the input as a block strand. When the input is 1, DNAzyme activity is inhibited and cannot activate the downstream with the output as 0. The input strand in three modes are bound to the two recognition arms and the activity core, respectively. It is experimentally verified that the output signal is suppressed when the input is present (Supplementary Figure S17). ‘NAND’ gates need to be built based on the ‘NOT’ gate (Figure 6R). Figure 6S shows the truth table of the ‘NAND’ gate. The ‘NAND’ gate is a combination of two ‘NOT’ gates and an ‘AND’ gate (Figure 6T). Only when both inputs are present, are both DNAzymes inhibited simultaneously, resulting in a low signal (Figure 6U).
The design of the ‘XOR’ gate requires a process of input annihilation (Figure 6V and W). The molecular structure draws on our previous work (47). Both input strands can cluster allosteric regions to activate the DNAzyme. But when they are present at the same time, each input strand can only bind to a partial region of the DNAzyme due to competition with each other (Figure 6X). A fluorescent signal is generated only when the two inputs are present individually (Figure 6Y).
Through the above experiments, we demonstrated the ability of the MCSAP to perform logic operations and good compatibility with molecular networks, broadening the application scenarios and potential of MCSAP.
Conclusion
In this work, we developed a redundancy-based CRISPR/Cas12a synergistic activation platform with modular access to different upstream molecular networks to perform multiple tasks. We found that redundant structures in split activators can affect Cas12a activity and enable fine-tuning over a large linear range through flexible combinations of length, position, and concentration. We adapted the redundancy as interfaces to bridge with upstream molecular networks and controlled CRISPR/Cas activation. MCSAP became modular reporter units with good compatibility. We performed three tasks: SNV detection, APE 1 detection, and logic operations. Tasks such as recognizing single nucleotide variants and logical operations were handed over to the upstream molecular network for execution without relying on sequences or structures specific to the CRISPR/Cas. Notably, in constructing the DNA circuit, we used split DNAzyme as an encoder for the conversion. Many DNAzyme-based nucleic acids or non-nucleic acids target transformations, and logic algorithms have been developed. These can potentially be seamlessly integrated into MCSAP.
In molecular nanoengineering, programmability is the soul of powerful functionality and ubiquity. Synergistic modulation is the central topic of programmability because it changes the single reaction path. Each realization of synergistic modulation of the basic tools is a significant advancement in the field. Just as the proposal of Associative Toehold Activation changed the traditional toehold-mediated strand displacement (48,49), expanding the basic rule set of DNA circuitry. The proposal of split DNAzyme, based on complete DNAzyme (50), changed the molecular detection field's paradigm. For the first time, we report a split activators-based synergistic control strategy with good programmability and scalability. We believe that MCSAP provides a powerful tool for a wide range of applications of CRISPR/Cas in different fields, such as molecular biology and nanoengineering.
Supplementary Material
Contributor Information
Minghao Hu, School of Pharmacy, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, China; Department of Oncology, Tongji Hospital of Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, China.
Xianzhi Cheng, School of Pharmacy, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, China.
Tongbo Wu, School of Pharmacy, Tongji Medical College, Huazhong University of Science and Technology, Wuhan 430030, China.
Data availability
The data underlying this article are available in the article and in its online supplementary material.
Supplementary data
Supplementary Data are available at NAR Online.
Funding
National Natural Science Foundation of China [82172372]; State Key Laboratory of Digital Medical Engineering [2023-M04]. Funding for open access charge: National Natural Science Foundation of China [82172372].
Conflict of interest statement. None declared.
References
- 1. Brouns S.J.J., Jore M.M., Lundgren M., Westra E.R., Slijkhuis R.J.H., Snijders A.P.L., Dickman M.J., Makarova K.S., Koonin E.V., van der Oost J.. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science. 2008; 321:960–964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Liu G.W., Lin Q.P., Jin S., Gao C.X.. The CRISPR-Cas toolbox and gene editing technologies. Mol. Cell. 2022; 82:333–347. [DOI] [PubMed] [Google Scholar]
- 3. Song X.R., Liu C., Wang N., Huang H., He S.Y., Gong C.Y., Wei Y.Q.. Delivery of CRISPR/Cas systems for cancer gene therapy and immunotherapy. Adv. Drug Deliv/ Rev. 2021; 168:158–180. [DOI] [PubMed] [Google Scholar]
- 4. Li H.M., Xie Y., Chen F.M., Bai H.W., Xiu L.S., Zhou X.N., Guo X.K., Hu Q.Q., Yin K.. Amplification-free CRISPR/Cas detection technology: challenges, strategies, and perspectives. Chem. Soc. Rev. 2023; 52:361–382. [DOI] [PubMed] [Google Scholar]
- 5. Hu Y.Q., Zhang Z., Ye W.C., Zhang W., Hu M.H., Yuan W.Q., Wang H.B., Xiao X.J., Wu T.B.. A DNA structure-mediated fluorescent biosensor for apurinic/apyrimidinic endonuclease 1 activity detection with ultra-high sensitivity and selectivity. Sens. Actuators B. 2021; 330:129332. [Google Scholar]
- 6. Liu J.J., Liu Y., Zhang L.H., Fu S.N., Su X.. Ultra-specific fluorescence detection of DNA modifying enzymes by dissipation system. Biosens. Bioelectron. 2022; 215:114561. [DOI] [PubMed] [Google Scholar]
- 7. Kou Q.N., Wang L., Zhang L.H., Ma L., Fu S.N., Su X.. Simulation-assisted localized DNA logical circuits for cancer biomarkers detection and imaging. Small. 2022; 18:2205191. [DOI] [PubMed] [Google Scholar]
- 8. Hu M.H., Hu Y.Q., Wu T.B.. A multifunctional monolithic interfacial sensor based on gold nanoparticle. Talanta. 2023; 259:124546. [DOI] [PubMed] [Google Scholar]
- 9. Chen J., Fu S.N., Zhang C.Y., Liu H.Y., Su X.. DNA logic circuits for Cancer Theranostics. Small. 2022; 18:2108008. [DOI] [PubMed] [Google Scholar]
- 10. Sadremomtaz A., Glass R.F., Guerrero J.E., LaJeunesse D.R., Josephs E.A., Zadegan R.. Digital data storage on DNA tape using CRISPR base editors. Nat. Commun. 2023; 14:6472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Zhang Y.B., Chen Y., Zhang Q.L., Liu Y.Z., Zhang X.J.. An aM-level sensitive cascade CRISPR-dx system (ASCas) for rapid detection of RNA without pre-amplification. Biosens. Bioelectron. 2023; 230:115248. [DOI] [PubMed] [Google Scholar]
- 12. Lu Y.H., Li F., Bai J.R., Ledesma-Amaro R., Liu D.Y., He Q., Deng R.J.. Rapid screening of antimicrobial probiotics using CRISPR cascade. Biosens. Bioelectron. 2022; 216:114673. [DOI] [PubMed] [Google Scholar]
- 13. Yan H., Wen Y.J., Tian Z.M., Hart N., Han S., Hughes S.J., Zeng Y.. A one-pot isothermal Cas12-based assay for the sensitive detection of microRNAs. Nat. Biomed Eng. 2023; 7:1583–1601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Wu Y.P., Chang D.R., Chang Y.Y., Zhang Q., Liu Y., Brennan J.D., Li Y.F., Liu M.. Nucleic acid enzyme-activated CRISPR-Cas12a with circular CRISPR RNA for biosensing. Small. 2023; 19:2303007. [DOI] [PubMed] [Google Scholar]
- 15. Hu M.L., Liu R.H., Qiu Z.Q., Cao F., Tian T., Lu Y.X., Jiang Y.Z., Zhou X.M.. Light-start CRISPR-Cas12a reaction with caged crRNA enables rapid and sensitive nucleic acid detection. Angew. Chem. Int. Ed. 2023; 62:e202300663. [DOI] [PubMed] [Google Scholar]
- 16. Hu M.L., Qiu Z.Q., Bi Z.R., Tian T., Jiang Y.Z., Zhou X.M.. Photocontrolled crRNA activation enables robust CRISPR-Cas12a diagnostics. Proc. Natl. Acad. Sci. U.S.A. 2022; 119:e2202034119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Deng H.P., Xu H., Wang Y.R., Jia R.Z., Ma X.Q., Feng Y.S., Chen H.M.. G-quadruplex-based CRISPR photoswitch for spatiotemporal control of genomic modulation. Nucleic Acids Res. 2023; 51:4064–4077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Liu X.Y., Cui S.Y., Qi Q.Q., Lei H.J., Zhang Y.T., Shen W., Fu F., Tian T., Zhou X.. G-quadruplex-guided RNA engineering to modulate CRISPR-based genomic regulation. Nucleic Acids Res. 2022; 50:11387–11400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Oesinghaus L., Simme F.C.. Switching the activity of Cas12a using guide RNA strand displacement circuits. Nat. Commun. 2019; 10:2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Hu Z., Sun A., Yang J.L., Naz G., Sun G.W., Li Z.P., Liu J.J.G., Zhang S.C., Zhang X.R.. Regulation of the CRISPR-Cas12a system by methylation and demethylation of guide RNA. Chem. Sci. 2023; 14:5945–5955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Cai W.Q., Liu J., Chen X.H., Mao L.Q., Wang M.. Orthogonal chemical activation of enzyme-inducible CRISPR/Cas9 for cell-selective genome editing. J. Am. Chem. Soc. 2022; 144:22272–22280. [DOI] [PubMed] [Google Scholar]
- 22. Fei X., Lei C., Ren W., Liu X., Liu C.. Regulating the trans-cleavage activity of CRISPR/Cas12a by using an elongation-caged single-stranded DNA activator and the biosensing applications. Anal. Chem. 2023; 95:12169–12176. [DOI] [PubMed] [Google Scholar]
- 23. Liu X., Kang X., Lei C., Ren W., Liu C.. Programming the trans-cleavage activity of CRISPR-Cas13a by single-strand DNA blocker and its biosensing application. Anal. Chem. 2022; 94:3987–3996. [DOI] [PubMed] [Google Scholar]
- 24. Zhao R., Luo W., Wu Y., Zhang L., Liu X., Li J.J., Yang Y.J., Wang L., Wang L.J., Han X.L.et al.. Unmodificated stepless regulation of CRISPR/Cas12a multi-performance. Nucleic Acids Res. 2023; 51:10795–10807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Wu Y., Luo W., Weng Z., Guo Y., Yu H., Zhao R., Zhang L., Zhao J., Bai D., Zhou X.et al.. A PAM-free CRISPR/Cas12a ultra-specific activation mode based on toehold-mediated strand displacement and branch migration. Nucleic Acids Res. 2022; 50:11727–11737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Hu J.Y., Zhou J., Liu R., Lv Y.. Element probe based CRISPR/Cas14 bioassay for non-nucleic-acid targets. Chem. Commun. 2021; 57:10423–10426. [DOI] [PubMed] [Google Scholar]
- 27. Rananaware S.R., Vesco E.K., Shoemaker G.M., Anekar S.S., Sandoval L.S.W., Meister K.S., Macaluso N.C., Nguyen L.T., Jain P.K.. Programmable RNA detection with CRISPR-Cas12a. Nat. Commun. 2023; 14:5409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Li Q., Song Z.L., Zhang Y.X., Zhu L.A., Yang Q., Liu X.F., Sun X.F., Chen X.X., Kong R.M., Fan G.C.et al.. Synergistic incorporation of two ssDNA activators enhances the trans-cleavage of CRISPR/Cas12a. Anal. Chem. 2023; 95:8879–8888. [DOI] [PubMed] [Google Scholar]
- 29. Zetsche B., Gootenberg J.S., Abudayyeh O.O., Slaymaker I.M., Makarova K.S., Essletzbichler P., Volz S.E., Joung J., van der Oost J., Regev A.et al.. Cpf1 Is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015; 163:759–771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Xu J., Liu Z., Zhang Z., Wu T.. Unlocking the full potential of Cas12a: exploring the effects of substrate and reaction conditions on trans-cleavage activity. Anal. Chem. 2023; 95:10664–10669. [DOI] [PubMed] [Google Scholar]
- 31. Zhang W., Shi R., Dong K.J., Hu H., Shu W., Mu Y.Q., Yan B., Li L.J., Xiao X.J., Wang H.B.. The off-target effect of CRISPR-Cas12a system toward insertions and deletions between target DNA and crRNA sequences. Anal. Chem. 2022; 94:8596–8604. [DOI] [PubMed] [Google Scholar]
- 32. Liu Z., Xu J., Huang S., Dai W., Zhang W., Li L., Xiao X., Wu T.. Gene point mutation information translation and detection: leveraging single base extension and CRISPR/Cas12a. Biosens. Bioelectron. 2024; 247:115936. [DOI] [PubMed] [Google Scholar]
- 33. Zhu Y.Z., Guo Z.W., Liu Y., Zheng X.Y., Yang G.H., Zheng G.J.. A novel ARMS-based assay for the quantification of mutations in patients with lung adenocarcinoma. Oncol. Lett. 2018; 15:2905–2912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Chen W., Xu H.Q., Dai S.B., Wang J.Y., Yang Z.Y., Jin Y.W., Zou M.B., Xiao X.J., Wu T.B., Yan W.et al.. Detection of low-frequency mutations in clinical samples by increasing mutation abundance via the excision of wild-type sequences. Nat. Biomed Eng. 2023; 7:1602–1613. [DOI] [PubMed] [Google Scholar]
- 35. Guo C., Deng H., Yang Q.F., Huang D., Shen C.L., Wang G.A., Li F.. Coding intrinsic disorder into DNA hybridization probes enables discrimination of single nucleotide variants over wide and tunable temperature ranges. Angew Chem. Int. Edit. 2023; 62:e202314386. [DOI] [PubMed] [Google Scholar]
- 36. Wang G.A., Xie X.Y., Mansour H., Chen F.F., Matamoros G., Sanchez A.L., Fan C.H., Li F.. Expanding detection windows for discriminating single nucleotide variants using rationally designed DNA equalizer probes. Nat. Commun. 2020; 11:5473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Huang D., Deng H., Zhou J., Wang G.A., Lei Q., Guo C., Peng W., Liang P., Shen C., Ying B.et al.. Mismatch-guided deoxyribonucleic acid assembly enables ultrasensitive and multiplex detection of low-allele-fraction variants in clinical samples. J. Am. Chem. Soc. 2023; 145:20412–20421. [DOI] [PubMed] [Google Scholar]
- 38. Zhang Z., Hu Y.Q., Yuan W.Q., Hu M.H., Deng Y.H., Xiao X.J., Wu T.B.. Endonuclease IV-regulated DNAzyme motor for universal single-nucleotide variation discrimination. Anal. Chem. 2021; 93:9939–9948. [DOI] [PubMed] [Google Scholar]
- 39. Tsimberidou A.M., Kahle M., Vo H.H., Baysal M.A., Johnson A., Meric-Bernstam F.. Molecular tumour boards - current and future considerations for precision oncology. Nat. Rev. Clin. Oncol. 2023; 20:843–863. [DOI] [PubMed] [Google Scholar]
- 40. Samer C.F., Lorenzini K.I., Rollason V., Daali Y., Desmeules J.A.. Applications of CYP450 testing in the clinical setting. Mol. Diagn Ther. 2013; 17:165–184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Liu J.L., Zhang C., Song J.X., Zhang Q., Zhang R.J., Zhang M.Z., Han D., Tan W.H.. Unlocking genetic profiles with a programmable DNA-powered decoding circuit. Adv. Sci. 2023; 10:2206343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Tosolini D., Antoniali G., Dalla E., Tell G.. Role of phase partitioning in coordinating DNA damage response: focus on the Apurinic Apyrimidinic endonuclease 1 interactome. Biomol. Concepts. 2020; 11:209–220. [DOI] [PubMed] [Google Scholar]
- 43. Hu M., Xie T., Hu Y., Li L., Wang T., Wu T.. Allosteric DNAzyme-based encoder for molecular information transfer. Chin. Chem. Lett. 2023; 35:109232. [Google Scholar]
- 44. Liao Y.W., Hu H., Tang X.F., Qin Y., Zhang W., Dong K.J., Yan B., Mu Y.Q., Li L.J., Ming Z.H.et al.. A versatile and convenient tool for regulation of DNA strand displacement and post-modification on pre-fabricated DNA nanodevices. Nucleic Acids Res. 2023; 51:29–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Hu M., Li X., Wu J.-n., Yang M., Wu T.. DNAzyme-based dissipative DNA strand displacement for constructing temporal logic gates. Acs Nano. 2024; 18:2184–2194. [DOI] [PubMed] [Google Scholar]
- 46. Hu M., Yang M., Cheng X., Wu T.. Time-controlled authentication strategies for molecular information transfer. Small. 2024; 20:2400261. [DOI] [PubMed] [Google Scholar]
- 47. Hu Y., Li C., Hu M., Zhang Z., Fu R., Tang X., Wu T.. Allosteric nucleic acid enzyme: a versatile stimuli-responsive tool for molecular computing and biosensing nanodevices. Small. 2023; 19:2300207. [DOI] [PubMed] [Google Scholar]
- 48. Chen X. Expanding the rule set of DNA circuitry with associative toehold activation. J. Am. Chem. Soc. 2012; 134:263–271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Yang X.L., Tang Y.N., Traynor S.M., Li F.. Regulation of DNA strand displacement using an allosteric DNA toehold. J. Am. Chem. Soc. 2016; 138:14076–14082. [DOI] [PubMed] [Google Scholar]
- 50. Mokany E., Bone S.M., Young P.E., Doan T.B., Todd A.V.. MNAzymes, a versatile new class of nucleic acid enzymes that can function as biosensors and molecular switches. J. Am. Chem. Soc. 2010; 132:1051–1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data underlying this article are available in the article and in its online supplementary material.