Abstract
Plant defense responses to the soil-borne fungus Verticillium longisporum causing stem stripe disease on oilseed rape (Brassica napus) are poorly understood. In this study, a population of recombinant inbred lines (RILs) using the Arabidopsis accessions Sei-0 and Can-0 was established. Composite interval mapping, transcriptome data, and T-DNA mutant screening identified the NITRATE/PEPTIDE TRANSPORTER FAMILY 5.12 (AtNPF5.12) gene as being associated with disease susceptibility in Can-0. Co-immunoprecipitation revealed interaction between AtNPF5.12 and the MAJOR LATEX PROTEIN family member AtMLP6, and fluorescence microscopy confirmed this interaction in the plasma membrane and endoplasmic reticulum. CRISPR/Cas9 technology was applied to mutate the NPF5.12 and MLP6 genes in B. napus. Elevated fungal growth in the npf5.12 mlp6 double mutant of both oilseed rape and Arabidopsis demonstrated the importance of these genes in defense against V. longisporum. Colonization of this fungus depends also on available nitrates in the host root. Accordingly, the negative effect of nitrate depletion on fungal growth was less pronounced in Atnpf5.12 plants with impaired nitrate transport. In addition, suberin staining revealed involvement of the NPF5.12 and MLP6 genes in suberin barrier formation. Together, these results demonstrate a dependency on multiple plant factors that leads to successful V. longisporum root infection.
Keywords: Arabidopsis, Brassica napus, major latex protein, MLP6, nitrate, NPF5.12, Verticillium longisporum
Recognition of the fungal pathogen Verticillium longisporum triggers nitrate starvation and MLP6-mediated defenses in Arabidopsis, together reducing the suberin barrier.
Introduction
Plant roots grow in a complex soil matrix while competing for space, water, and nutrients with a plethora of organisms. Microbial activities are most intensive in the zone surrounding the roots, known as the rhizosphere, where numerous processes occur that are important for the availability and distribution of nutrients and their subsequent uptake into the plant root. While many components of the rhizosphere microbiome are beneficial to plant growth, several plant pathogenic microorganisms are attracted to this nutrient-rich environment. Soil-borne plant pathogens comprise vast organism groups and species of bacteria, fungi, oomycetes, plasmodiophorids, insects, and nematodes, all of which use survival in the bulk soil as a part of their disease cycle. During their biologically active phase, these soil organisms frequently feed on or infect plant roots to start new rounds of multiplication. These general characteristics apply to members of the fungal genus Verticillium. Verticillium dahliae and V. albo-atrum are common fungi with a broad host range. Together, these two fungi induce disease in more than 200 dicotyledonous plant species (Pegg and Brady, 2002). In contrast, V. longisporum primarily infects plants of the family Brassicaceae, including Arabidopsis (Tjamos et al., 2005; Johansson et al., 2006b). Verticillium longisporum produces hardy melanized microsclerotia. They are released into the soil from infected plant residues at the end of the disease cycle, where they remain dormant for many years until suitable germination conditions occur. Based on the current understanding of plant infection processes, hyphae from germinating microsclerotia colonize root tissues, followed by penetration of root epidermal cells and entry into xylem elements (Zhou et al., 2006; Eynck et al., 2007). The xylem is nutrient poor, and to adapt to such an environment, V. longisporum may acquire necessary nutrients via digestion of host cell walls and induction of ion leakage from neighboring cells (Singh et al., 2010; Klosterman et al., 2011; Yadeta and Thomma, 2013). In the xylem, fungal metabolites and possibly occlusion of xylem tissues by the fungus may lead to premature senescence (Zhou et al., 2006; Eynck et al., 2007). Fungal infection and progression in Brassica species are processes known to be slow and to advance without obvious external symptoms (Depotter et al., 2016). Occasionally, one-sided chlorosis of the leaves is observed before abscission. When the flowering stage is initiated in the host plant, the fungus starts to produce microsclerotia that protrude into the plant tissue and become visible as black spots, particularly on stems or stubble after harvest (Heale and Karapapa, 1999; Johansson et al., 2006a). This long period of latent or invisible infection and the clear disease symptoms at late growth stages contribute to the underestimation of the disease incidence of this pathogen. Data reported thus far on losses in oilseed rape range between negligible and 50% (Dunker et al., 2008; Depotter et al., 2019). In geographic regions where blackleg (Leptoshpaeria maculans) is also a problem, additive losses of oilseed rape are observed (Wang et al., 2023).
Transport of nitrate, the main nitrogen source for plants, from the soil into root cells involves the activity of several specific membrane transporters (Miller et al., 2007). Nitrate is stored in the vacuole or further processed into ammonium and amino acids followed by translocation via xylem and phloem tissues to aboveground sink organs (Islam et al., 2022). Transmembrane nitrate and peptide transporters in plants are divided into three families: the ATP-BINDING CASSETTE (ABC) superfamily (Kang et al., 2011), the NITRATE TRANSPORTER/PEPTIDE TRANSPORTER (NTR/PTR) family (Rentsch et al., 2007), and the OLIGOPEPTIDE TRANSPORTER (OPT) family (Lubkowitz, 2011). Members of the ABC transporter family, which in Arabidopsis consists of more than 120 proteins, hydrolyse ATP to drive transport of substrates ranging from small ions to large macromolecules predominantly out of the cytoplasm. In contrast, NTRs/PTRs and OPTs are proton-coupled symporters that transport substrates in the opposite direction (Schaaf et al., 2004; Osawa et al., 2006; Kurt and Filiz, 2022). The OPT family comprises 17 members in Arabidopsis, transporting tetra- and pentapeptides as well as glutathione (Lubkowitz, 2011). The NTR/PTR family is more complex and has over time been named PROTON-COUPLED OLIGOPEPTIDE TRANSPORTER (POT), PEPTIDE TRANSPORTER (PepT/PTR), or SOLUTE CARRIER 15 (SLC15). The revised nomenclature is now based on the phylogenetic relationships of NRT1/PTR family members in 31 sequenced plant genomes (Léran et al., 2014): NPF (NRT1/PTR FAMILY), with individual members identified by numbers based on their position among the eight identified subfamilies. Arabidopsis NPFs are phylogenetically divided into four clades (Nour-Eldin et al., 2012; Léran et al., 2014). At least 20 NPFs are characterized as low-affinity nitrate transporters, including NPF5.12 (He et al., 2017; Kanstrup and Nour-Eldin, 2022). All NPF proteins contain 12 transmembrane domains connected by short protein loops, as revealed by 3D crystal structures of prokaryotic homologs (Solcan et al., 2012; Guettou et al., 2013) and the structure of Arabidopsis NRT 1.1 (Parker and Newstead, 2014; Sun et al., 2014).
In this study, we performed crossings between two contrasting Arabidopsis accessions in combination with single nucleotide polymorphism (SNP) genotyping, transcriptome data, and T-DNA insertion mutant screen to identify the nitrate/peptide transporter gene AtNPF5.12 as participating in defense responses to V. longisporum. Co-immunoprecipitation experiments and bimolecular fluorescence complementation (BiFC) microscopy revealed interaction between AtNPF5.12 and a MAJOR LATEX PROTEIN (AtMLP6) in the plasma membrane and the endoplasmic reticulum. Arabidopsis and B. napus plants with NPF5.12 and MLP6 mutations showed enhanced susceptibility to V. longisporum. The two genes were also found to participate in processes that strengthen the endodermal suberin barrier, suggesting multiple MLP6 functions.
Materials and methods
Arabidopsis materials and growth conditions
Arabidopsis accessions Col-0 (N1092), Can-0 (N1064), Sei-0 (N1504), and homozygous T-DNA insertion mutants (Alonso et al., 2003; Supplementary Table S1) were grown hydroponically (Fradin et al., 2011) or in soil (Bohman et al., 2004) at a light intensity of 100 µmol m−2 s−1 and a temperature of 21 °C (light) and 16 °C (dark).
Fungal isolates, Arabidopsis inoculation, and quantification
Verticillium longisporum isolate VL1 (Fogelqvist et al., 2018) was used for all inoculations, except for monitoring of fungal colonization, for which a green fluorescent protein (GFP)-tagged isolate of V. longisporum Vl43 (Eynck et al., 2007) was used. Soil-based inoculation was performed as previously described (Roos et al., 2014). Disease symptoms were monitored for up to 4 weeks. In gene expression experiments, roots of 2-week-old plants grown in vitro were dipped in a 104 conidium ml−1 suspension and transferred to fresh Murashige and Skoog (MS) plates without sucrose. Root materials were collected at 2 days post-inoculation (dpi) in biological replicates with ≥20 plants in each sample. For fungal quantification, 2-week-old hydroponically grown Arabidopsis plants were dipped in 104 conidium ml−1 suspension for 30 min in a separate container, and transferred back to the hydroponic culture, which was continued. At 14 dpi, the roots were rinsed in water and 70% ethanol to remove potential external fungal growth, snap frozen in liquid nitrogen, and stored at −70 °C. Total DNA was extracted from the root materials using GeneJET Plant Genomic DNA Purification Kit (Thermo Fisher Scientific, Waltham, MA, USA). qPCR primer efficiency and specificity were assessed according to Schmittgen and Livak (2008). The sequences of the primers used are listed in Supplementary Table S2.
Recombinant inbred lines and plant phenotyping
The parental lines Can-0 (susceptible) and Sei-0 (resistant) were crossed, and individual F2 offspring were self-fertilized until the F8 generation. Recombinant inbred line (RIL) plants (F8) growing in soil were infected with V. longisporum as described above, and disease symptoms were scored as follows: 0, no symptoms, control; 1, discoloration of leaf vascular tissues starting to appear; 2, plants reduced in size and chlorosis starting to develop compared with control; 3, plants chlorotic and significantly stunted compared with control (Supplementary Fig. S1). Each RIL was infected at least three times, and the disease phenotype was determined by using a minimum of 20 inoculated plants per replicate.
SNP genotyping, QTL, and array analyses
DNA from each RIL were extracted using the cetyltrimethylammonium bromide extraction method (Doyle and Doyle, 1987). SNP markers capable of distinguishing between Can-0 and Sei-0 were previously published (Kover et al., 2009) or designed from 1001 Genomes 250k SNP data (Horton et al., 2012) and used in a GoldenGate Genotyping Assay (Illumina, San Diego, CA, USA) with SNP Technology Platform at Uppsala University Hospital (Uppsala, Sweden). The resulting SNP genotypes were combined into a genetic map using MAPMAKER/EXP software (Lander et al., 1987). Quantitative trait locus (QTL) analysis was performed with composite interval mapping implemented in QGene software (Joehanes and Nelson, 2008). A permutation test of 1000 replicates was run to determine the significance of the QTLs identified. Col-0, Can-0, and Sei-0 genome sequences were compared using Genome Express Browser 3.0 (http://signal.salk.edu/atg1001/3.0/gebrowser.php). Genes present in the QTL region were compared with genes differentially expressed in an Affymetrix ATH1 genome array based on mock (water) and V. longisporum-inoculated Col-0 and ndr1-1 mutant plants at 2 dpi (GEO accession GSE62537) (Roos et al., 2015). Microarray analysis was performed in R (R Core Team, 2016) and normalized and background corrected using the robust multiarray analysis method (Irizarry et al., 2003), as implemented in the package affy v1.50.0 (Gautier et al., 2004). Low-signal and low-variance probe sets were filtered. In the final step, the probe sets were required to have at least one sample with a normalized intensity above log2(100) and an interquartile range of log2 intensities of at least 0.2. Differentially expressed genes were determined using Student’s t-test with empirical Bayes moderation of standard errors, as implemented in the R package limma v3.28.21 (Ritchie et al., 2015). The significance threshold was set at P<0.05 (P-values adjusted for multiple testing using the correction of Benjamini and Hochberg, 1995). A heatmap was produced using the ggplot2 v2.1.0 package (Wickham, 2009).
RNA isolation and quantitative real-time PCR
Total RNA was isolated from Arabidopsis plants using Qiagen RNeasy Plant Mini Kit (Qiagen Sciences Inc., Germantown, MD, USA). cDNA was synthesized with a qScript cDNA Synthesis kit (Quanta Biosciences, Gaithersburg, MD, USA). Quantitative real-time PCR was performed with Fermentas Maxima SYBR Green/Fluorescein qPCR Master Mix (Thermo Fisher Scientific). Gene-specific primers were designed using Primer3 (Rozen and Skaletsky, 2000), and expression was normalized to the ACTIN2 gene in Arabidopsis (Supplementary Fig. S2). All primers used are listed in Supplementary Table S2. Transcript data were analysed with the comparative CT method (Livak and Schmittgen, 2001), and quantitative reverse transcription PCR efficiency correction was determined from the slope of standard curves.
Plasmid construction and Arabidopsis transformation
Arabidopsis cDNA or gDNA target sequences were PCR amplified with Phusion DNA polymerase (Thermo Fisher Scientific) and cloned into the pCR8/GW/TOPO cloning vector (Thermo Fisher Scientific) followed by sequencing. Confirmed inserts were introduced into suitable destination vectors using the Gateway system (Thermo Fisher Scientific). The primers and vectors used are provided in Supplementary Table S3. Final p35S:AtNPF5.12, p35S:AtNPF5.12-His, p35S:AtNPF5.12-GFP, pAtNPF5.12:AtNPF5.12-His, pAtNPF5.12:AtNPF5.12-GFP, and p35S:AtMLP6 constructs were transformed into Agrobacterium tumefaciens strain C58, followed by transformation of Col-0, Atnpf5.12 or Atmlp6 plants using the floral-dip method (Davis et al., 2009). Confirmed T2 lines were used for GFP analyses; T3 homozygous complementation lines were used for inoculation assays. At least three independent transgenic lines per construct were used for the analyses. F2 plants were selected on MS medium with 50 μg ml−1 kanamycin. Data on independent transgenic line 2 and 3 for each construct are compiled in Supplementary Table S4.
Protein extraction, immunoprecipitation, and mass spectrometry
Total proteins were extracted from leaves of p35S:NPF5.12-His transgenic Arabidopsis plants using extraction buffer [50 mM Tris–HCl pH 7.4, 10 mM EDTA, 0.1% Triton X-100, and 1 μl ProteoBlock protease inhibitor cocktail (Thermo Fisher Scientific)]. For western blotting, 10 μg of total protein crude extract was separated by 12% SDS-PAGE, followed by electrotransfer to a polyvinylidene difluoride membrane. A primary anti-His antibody (Thermo Fisher Scientific) and peroxidase-conjugated secondary antibodies (goat anti-rabbit IgG, rabbit anti-mouse IgG, Dako, Glostrup, Denmark) were used, followed by chemiluminescence detection (GE Healthcare, Pittsburgh, PA, USA). For immunoprecipitation, crude extracts were incubated with the anti-His antibody overnight at 4 °C, followed by a 90 min incubation with Protein A Sepharose 4 Fast flow beads (GE Healthcare) and subsequent washes with immunoprecipitation washing buffers A (50 mM Tris pH 8.0, 150 mM NaCl, 0.1% Triton X-100) and B (50 mM Tris pH 8.0, 0.1% Triton X-100). The washed beads were separated by 12% SDS-PAGE followed by colloidal Coomassie staining. Candidate protein bands were excised and analysed by matrix-assisted laser desorption/ionization tandem mass spectrometry (MALDI-MS/MS) using an Ultraflex III TOF/TOF (Bruker Daltonics, Coventry, UK). The Mascot program (http://www.matrixscience.com/) was used to compare the resulting MS/MS spectra against an Arabidopsis subset in the NCBI database for identification.
In silico analyses and protein modeling
The Arabidopsis amino acid sequence of AtNPF5.12 was analysed using TMHMM v.2 software (Krogh et al., 2001). The NetSurfP (Petersen et al., 2009) and Phyre2 (Kelley and Sternberg, 2009) servers were used to build three-dimensional structure predictions of AtNPF5.12 and AtMLP6 using default settings.
Cellular localization and bimolecular fluorescence complementation
Cellular localization of AtNPF5.12 was monitored using fluorescence microscopy of transgenic p35S:AtNPF5.12-GFP Arabidopsis plant roots and 2-week-old Nicotiana benthamiana leaves infiltrated with Agrobacterium harboring the pAtNPF5.12:AtNPF5.12-GFP construct. Co-infiltration with mCherry-tagged markers (Takara) for the plasma membrane and endoplasmic reticulum (Nelson et al., 2007) was used to support data on subcellular localization. For BiFC analysis (Li et al., 2010), AtNPF5.12 and AtMLP6 cDNAs were PCR amplified with the primers listed in Supplementary Table S3. Fragments were ligated into the pCR8/GW/TOPO entry vector and sequenced, followed by transformation into Gateway-compatible BiFC vectors (pSITE:nEYFP-C1 and pSITE:cEYFP-C1). The final pSITE:cEYFP-AtNPF5.12 and pSITE:nEYFP-AtMLP6 plasmids were transformed into Agrobacterium strain GV3101. Agrobacteria harboring the two different plasmids were co-infiltrated into N. benthamiana leaves at a 1:1 (v/v) ratio and imaged at 4 d post-infiltration (Schütze et al., 2009).
Brassica napus genome editing
Orthologous genes in B. napus were located using NCBI BLASTP (https://blast.ncbi.nlm.nih.gov/Blast.cgi) with AtNPF5.12 (NP_177359.1) and AtMLP6 (NP_194098.1) as queries. Conserved regions in exon 1 for BnMLP6 loci and exons 3 and 4 for BnNPF5.12 loci were targeted by single-guide RNAs (sgRNAs) (Supplementary Table S5). CRISPys (Hyams et al., 2018) and CRISPR MultiTargeter (Prykhozhij et al., 2015) were used to locate suitable sgRNAs, and off-targets were predicted by Cas-OFFinder (Bae et al., 2014). Four mutant combinations were created: two independent lines of quadruple BnNPF5.12, two independent lines of 14-fold BnMLP6 and one quadruple BnNPF5.12/octuple BnMLP6 line and a quadruple BnNPF5.12/14-fold BnMLP6 line. Bnnpf5.12-1, Bnnpf5.12-2, Bnmlp6-1, Bnmlp6-2, Bnnpf5.12/Bnmlp6-1, and Bnnpf5.12/Bnmlp6-2 were used (Supplementary Table S6). Dual sgRNA cassettes were cloned into the destination vector pHSE401 (Xing et al., 2014). Phusion High-Fidelity PCR Master Mix with HF Buffer (Thermo Fisher Scientific) was used to amplify sgRNA and plasmid adapters. The final plasmids were transformed into Agrobacterium Gv3101 carrying the disarmed Ti plasmid pMP90 with the freeze‒thaw method (Weigel and Glazebrook, 2006).
Brassica napus transformation
Transformation of B. napus hypocotyls (cv. Kumily) and plant regeneration were based on Schröder et al. (1994), with the modifications listed below. Information about different media is provided in Supplementary Table S7. Etiolated hypocotyl segments were dissected in M1 medium, rapidly transferred to M1 medium containing 4 × 108Agrobacterium cells ml−1 and incubated for 30 min at room temperature. The hypocotyls were left overnight at 28 °C followed by 2 d in a 24 °C darkroom on callus-inducing medium (M2). Antibiotics were then added for 3 weeks to allow for selection (M3). Calli were maintained on shoot regeneration medium (M4) with antibiotics 4–8 weeks and selection-free M4 medium for an additional 8 weeks. Shoots were transferred to glass jars with rooting medium (M5) for 10 d prior to soil transfer. Plants of generation T0 were screened for the presence of the Hyg transgene by PCR, and T1 plants were sequenced at targeted loci (Supplementary Table S8) and used for fungal inoculation. Gene-specific sequencing primers were produced with NCBI Primer-BLAST applied to B. napus genome assembly AST_PRJEB5043_v1. The primers used for cloning, PCR and sequencing are listed in Supplementary Table S9.
Fungal inoculation of Brassica napus
Brassica napus plants were grown hydroponically (Jambagi and Dixelius, 2023) at a constant temperature of 22 °C with 120 µmol m−2 s−1 light intensity. The optimal V. longisporum inoculum for hydroponic culture of B. napus was determined to be 4 × 105 ml−1 (Supplementary Fig. S3). Two-week-old plants were dipped in the fungal spore suspension for 30 min and transferred to the hydroponic culture. Root materials were collected after 7 d, washed in 70% ethanol, and frozen in liquid nitrogen. At least four biological replicates, each with 10 plants, were prepared. The DNA extraction, qPCR settings, and calculations were performed as previously described (Martin et al., 2011). All primers used are listed in Supplementary Table S2.
Nitrogen depletion
Arabidopsis plants were grown in full-strength hydroponic medium containing nitrogen or in nitrogen free-medium in which KNO3 was replaced with 1.75 mmol l−1 K2SO4. Ammonium nitrate (NH4NO3) was removed.
Suberin staining and quantification
Histochemical staining of suberin was performed on 21-day-old Arabidopsis and 14-day-old B. napus roots at 7 dpi or with water (mock) treatment. The staining was performed according to Barberon et al. (2016), except that a lower concentration (0.02% w/v) of Fluorol Yellow 088 (Santa Cruz Biotechnology, Dallas, TX, USA) was used. Fluorescence intensity data were collected from confocal laser scanning microscope images using ImageJ v1.53t. Corrected total cell fluorescence (CTCF) was calculated as , where is the integrated density of fluorescence intensity, is the area measured, and is the mean background fluorescence intensity of the selected area (McCloy et al., 2014).
Salicylic acid measurements
Salicylic acid (SA) was quantified using [13C1]SA as an internal standard following the protocol by Ratzinger et al. (2009). Free SA content in hydroponically grown Col-0, Atnpf5.12, and Atmlp6 plants inoculated with VL1 or mock-treated (water) were analysed at 2 dpi.
Confocal microscopy
Fluorescence microscopy images were captured with a Zeiss 780 confocal scanning microscope using Zen2011 software. The excitations/emissions for each channel were as follows: GFP (488/493–530 nm), Fluorol Yellow 088 (488/410–534 nm), yellow fluorescent protein (YFP) (514/518–560 nm), chlorophyll (633/647–721 nm), and mCherry (561/600–650 nm). The red color of mCherry was replaced with magenta in all images.
Statistical analyses
Statistical analyses for transcript accumulation, fungal DNA content and Fluorol Yellow 088 signal intensity were performed in R (https://www.r-project.org/). Statistical differences were determined using Student’s t-test, except that one-way ANOVA and Tukey’s multiple comparisons of means were applied for multiple group comparisons. The significance level was set at 95%.
Results
A QTL on Arabidopsis chromosome 1 is associated with V. longisporum disease resistance
In an effort to identify defense genes against V. longisporum, genetic variation between two Arabidopsis accessions was utilized in a gene mapping approach. In a previous screen with V. longisporum (Johansson et al., 2006b), the accessions Can-0 from the Canary Islands (Spain) and Sei-0 from Seis am Schlern (Italy) were identified as being highly susceptible and most resistant, respectively (Fig. 1A, B). Can-0 and Sei-0 were therefore used as parental genotypes to produce RILs. This approach resulted in a set of 119 RIL individuals screened for their response to V. longisporum. To identify genomic regions linked to disease symptom differences, the RILs and the parental accessions were genotyped with a set of 143 SNP markers (Supplementary Dataset S1) using an Illumina Golden Gate assay. The genetic mapping and QTL analyses revealed a single QTL (logarithm of the odds=11) located on chromosome 1 to be associated with disease resistance (Supplementary Fig. S4A, B).
Fig. 1.
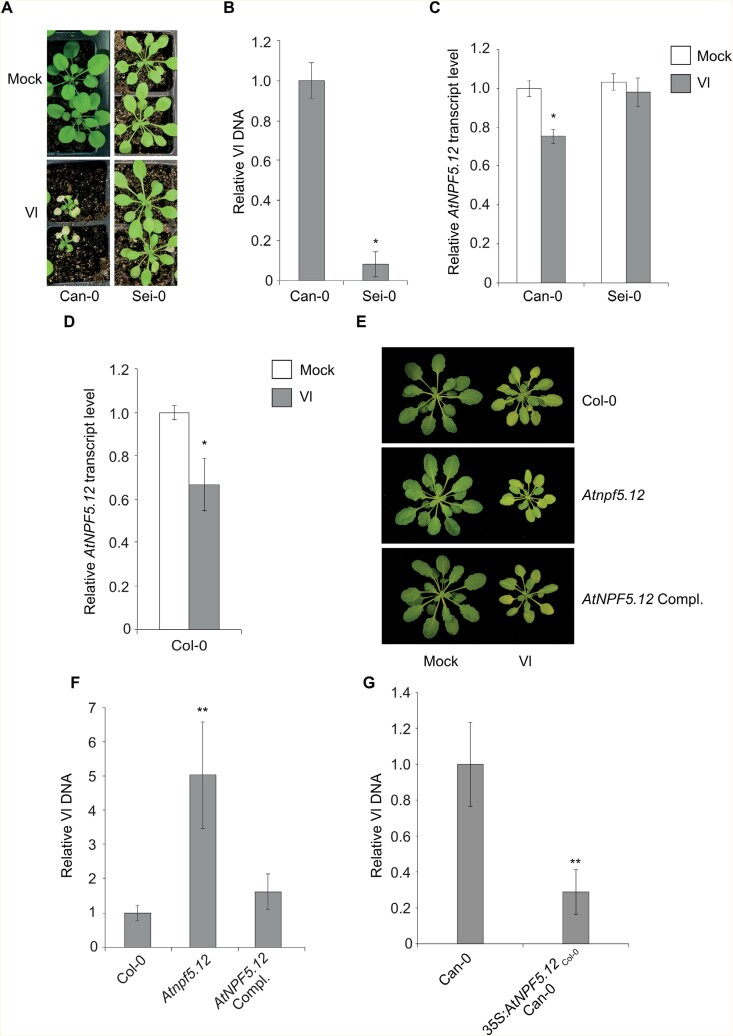
Phenotype and fungal DNA quantification in different Arabidopsis genotypes. (A) Disease symptoms of soil-grown Arabidopsis Can-0 and Sei-0 plants. Photos taken 21 days post-inoculation (dpi) with V. longisporum VL1. (B) Relative fungal DNA content in Sei-0 (0.1-fold) plant roots compared with Can-0 at 14 dpi. Bar chart represents means ±SE (n=6 biological replicates, 20 plants for each plant line and replicate). (C) Relative transcript levels of AtNPF5.12 in roots of Can-0 (0.8-fold) and Sei-0 plants. Data from V. longisporum (Vl) inoculated plants are relative to data from mock-treated plants at 2 dpi. Bar chart represents means ±SE (n=6 biological replicates of >25 plants for each plant line and treatment, repeated twice). (D) Relative transcript levels of V. longisporum (Vl) inoculated AtNPF5.12 roots (0.7-fold) compared with mock-treated plants at 2 dpi. Bar chart represents means ±SE (n=6 replicates of >20 plants for each treatment, repeated twice). (E) Disease symptoms of soil-grown Arabidopsis Col-0, Atnpf5.12, and Atnpf5.12 Compl. plants at 21 dpi. Atnpf5.12 Compl. plants are complemented with the native gene and promoter (pAtNPF5.12Col-0:AtNPF5.12Col-0). (F) Relative fungal DNA content in in vitro grown roots of V. longisporum inoculated Atnpf5.12 (5-fold) and Atnpf5.12 complemented lines (Compl. plants). The data are relative to Col-0 at 14 dpi. Bar chart represents means ±SE (n=6 biological replicates of 20 plants for each plant line). (G) Relative V. longisporum DNA content in roots of inoculated p35S:AtNPF5.12Col-0 complemented plants (0.3-fold). The data are relative to Can-0 at 14 dpi. Bar chart represents means ±SE (n=6 biological replicates of 20 plants for each plant line). Plants in (B–G) were grown in a hydroponic system. All transcription data were normalized to AtACTIN2. Asterisks represent significant difference by Student’s t-test: *P≤0.05; **P≤0.01.
Chromosomal comparison between Can-0 and Col-0
We compared the genome sequence of Can-0 (Gan et al., 2011) with Col-0 in the mapped region between At1g71697 and At1g80640 to find divergent sequences. A total of 918 genes are present in this region in Col-0. BLAST searches against the Can-0 genome with these genes as query sequences revealed strong homology between the two genomes; the differences comprise 78 genes absent in the Can-0 genome compared with Col-0. Extended BLAST searches and comparison with RNA sequence data from Can-0 (Gan et al., 2011) revealed that 75 of the 78 genes are present in other locations outside the mapped region in the Can-0 genome. Two adjacent genes coding for typical resistance proteins of the Toll/interleukin-1 receptor–nucleotide binding site–leucine-rich repeat category (At1g72840 and At1g72850) are absent in the genomes of Can-0 and Sei-0 (https://1001genomes.org). A third gene (At1g76960) with unknown function but with a WRKY40 binding motif harbors indels in Can-0 and Sei-0 compared with Col-0. No differential expression of the three genes was detected in Col-0 samples, with transcripts being entirely absent in Can-0 and Sei-0 plant materials.
AtNPF5.12 contributes to V. longisporum defense
To identify important gene candidates, we continued the analysis by exploiting an array dataset (GEO accession GSE62537) between mock- and fungus-inoculated Col-0 and the NON-RACE SPECIFIC DISEASE RESISTANCE 1 (Atndr1) mutant (Roos et al., 2015). Twelve differentially expressed genes were identified in the QTL region on chromosome 1 (Supplementary Fig. S5). When analysing the transcript levels of these potential candidates, the nitrate peptide transporter AtNPF5.12 (At1g72140) was non-responsive in Sei-0 at 2 dpi and down-regulated in Can-0 (Fig. 1C), indicating that this gene might contribute to the differential V. longisporum response observed in the two RIL parents. Similarly, AtNPF5.12 transcription was down-regulated in Col-0 at 2 dpi (Fig. 1D). Transcript accumulation of the closely related genes AtNPF5.11, AtNPF5.13, AtNPF5.14, and AtNPF5.16 was not altered at 2 dpi compared with mock-treated plants (Supplementary Fig. S6). Three independent T-DNA insertion lines were examined (Supplementary Table S1), and SAIL_168_G10, hereafter denoted Atnpf5.12, was selected for further analysis. When the Atnpf5.12 mutant was inoculated with V. longisporum, only slightly enhanced disease symptoms compared with Col-0 were observed (Fig. 1E). However, a 5-fold elevated level of fungal DNA was detected at 14 dpi. This molecular phenotype reverted to wild-type levels in a genetic complementation line harboring pAtNPF5.12Col-0:AtNPF5.12Col-0, which suggests that a recognition factor is impaired in the mutant (Fig. 1F). To demonstrate whether the low transcription level of AtNPF5.12 in the Can-0 accession is responsible for the susceptible phenotype, we developed transgenic Can-0 plants expressing p35S:AtNPF5.12Col-0. Quantification of V. longisporum DNA revealed reduced fungal growth in the overexpression line compared with the wild-type Can-0 (Fig. 1G). Together, these results show that attenuated NPF5.12 expression in Can-0 background may play a role in promoting colonization and growth of V. longisporum in Arabidopsis. Next, sequence comparisons of AtNPF5.12 between the three Arabidopsis accessions based on information in the 1001 Genomes database revealed a short indel (9 nt) in the promoter sequence of At1g72140 in Sei-0 at position Chr1:27141089. We believe this is the main reason why Sei-0 exhibits a resistant phenotype compared with Col-0 and Can-0. Furthermore, Col-0 harbors an SNP in exon 4 of At1g72140 compared with Can-0 and Sei-0. This SNP causes a leucine to valine transversion at position Chr1:27144326. Because Col-0 was used as a template for the Can-0 complementation experiments, this amino acid substitution may be responsible for the intermediate response of Col-0 compared with Can-0 and Sei-0.
NPF5.12 localizes to the plasma membrane
Several characterized NPF proteins localize to the plasma membrane (Kanno et al., 2012; Nour-Eldin et al., 2012) and the vacuole tonoplast (Weichert et al., 2011; He et al., 2017). In silico analysis with TMHMM software predicted 12 transmembrane regions in the AtNPF5.12 protein, and the Phyre2 server suggested NRT1.1/NPF6.3 as the closest structural homolog (Supplementary Fig. S7A). To support the prediction and to clarify its subcellular localization, p35S:NPF5.12-GFP transgenic Arabidopsis plants were produced. When examined under a confocal fluorescence microscope, these plants showed a strong GFP signal in the plasma membrane in both roots and leaves (Fig. 2A). Further evidence of plasma membrane and tonoplast localization was indicated by transient expression of an NPF5.12:NPF5.12-GFP construct together with an mCherry-tagged plasma membrane marker in Nicotiana benthamiana (Fig. 2B).
Fig. 2.
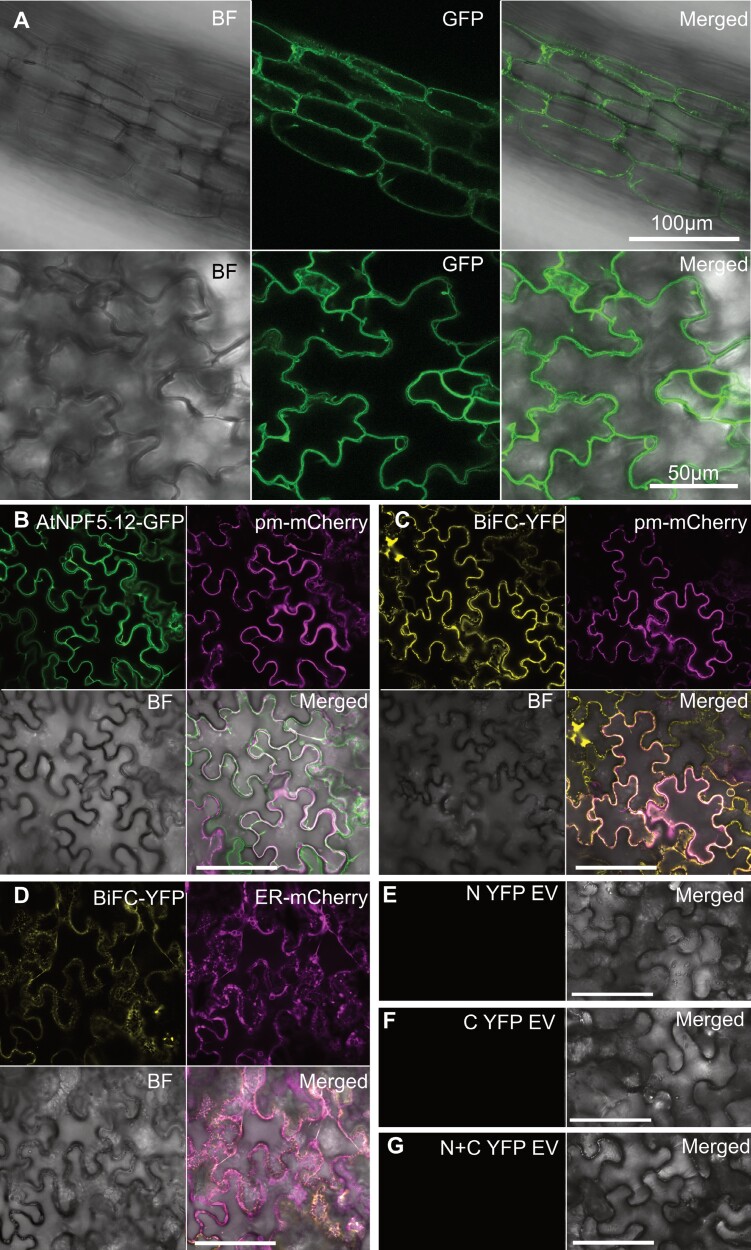
Cellular localization and interaction between NPN5.12 and MLP6 proteins. (A) Fluorescence microscopy images of roots (upper panel) and leaves (lower panel) from p35S:AtNPF5.12-GFP transgenic Arabidopsis plants. The AtNPF5.12–GFP protein fusion localizes in the plasma membrane. (B) Confocal microscope image of leaves from N. benthamiana plants, 4 d post-co-infiltration with a pAtNPF5.12:AtNPF5.12-GFP construct and a plasma membrane-localized mCherry marker. (C, D) Reconstituted yellow fluorescent protein (YFP) signal in N. benthamiana plants, 4 d post-infiltration with pSITE-cEYFP-AtNPF5.12 and pSITE-nEYFP-AtMLP6 BiFC constructs, together with a plasma membrane-localized (C) or endoplasmic reticulum-localized (D) mCherry marker. (E–G) empty vector pSITE-nEYFP, pSITE-cEYFP and pSITE-nEYFP coinfiltrated with pSITE-cEYFP as negative controls. BF, bright field; Merged, composite image; GFP, green fluorescence; YFP, yellow fluorescence; pm-mCherry, plasma membrane mCherry marker; ER-Cherry, endoplasmic reticulum mCherry marker. Scale bar (B–G) = 100 µm.
AtNPF5.12 interacts with the major latex protein AtMLP6
To identify plant proteins interacting with AtNPF5.12, transgenic p35S:AtNPF5.12-His Arabidopsis plants were generated and used for co-immunoprecipitation. Pull-down experiments with anti-His antibodies revealed a smaller protein of ~14 kDa in addition to the ~60 kDa protein corresponding to AtNPF5.12-His (Supplementary Figs S7B, S8A). MALDI-MS/MS analyses identified this protein as At4g23670 (Supplementary Fig. S8B) referring to previously known MAJOR LATEX PROTEIN 6 or AtMLP6 (Guo et al., 2011).
BiFC experiments were performed to visualize interaction between AtNPF5.12 and AtMLP6 in planta. Fluorescence microscopy showed a reconstituted signal in the plasma membrane and in the endoplasmic reticulum, supporting interaction between AtNPF5.12 and AtMLP6 in the two compartments. These intracellular localization observations were validated by co-infiltration with endoplasmic reticulum- and plasma membrane-specific mCherry-tagged markers (Fig. 2C, D). No signal was detected in empty vector negative controls (Fig. 2E–G).
NPF5.12 and MLP6 jointly contribute to reinforced defense against V. longisporum
To investigate whether the AtNPF5.12-interacting protein AtMLP6 is an important player in defense against V. longisporum, three independent T-DNA insertion mutants were evaluated for responses to fungal inoculation (Supplementary Table S1). The promoter mutant SALK_088249 (Atmlp6) was chosen for further studies due to its clearer responses. The fungal colonization level of the Atmlp6 mutant was similar to the responses in Atnpf5.12 plants at 14 dpi compared with Col-0. Furthermore, a double mutant between Atmlp6 and Atnpf5.12 was generated, and the amount of fungal DNA in these plants was clearly higher than that in the Atnpf5.12 and Atmlp6 single-mutant plants (Fig. 3A). A p35S:AtMLP6 complementation line in the Atmlp6 background displayed reversion of the susceptible phenotype observed in the Atmlp6 mutant (Supplementary Fig. S9). Transcript levels of AtMLP6 also decreased in both Col-0 and in the Atnpf5.12 mutant in response to fungal challenge (Fig. 3B), with only a marginal increase in disease phenotype (Fig. 3C), suggesting a possible link between the AtNPF5.12 and AtMLP6 genes in this specific defense response. Transcript accumulation for the highly sequence similar AtMLP6-related gene, At4g23680, was not altered by fungal inoculation (Supplementary Fig. S10).
Fig. 3.
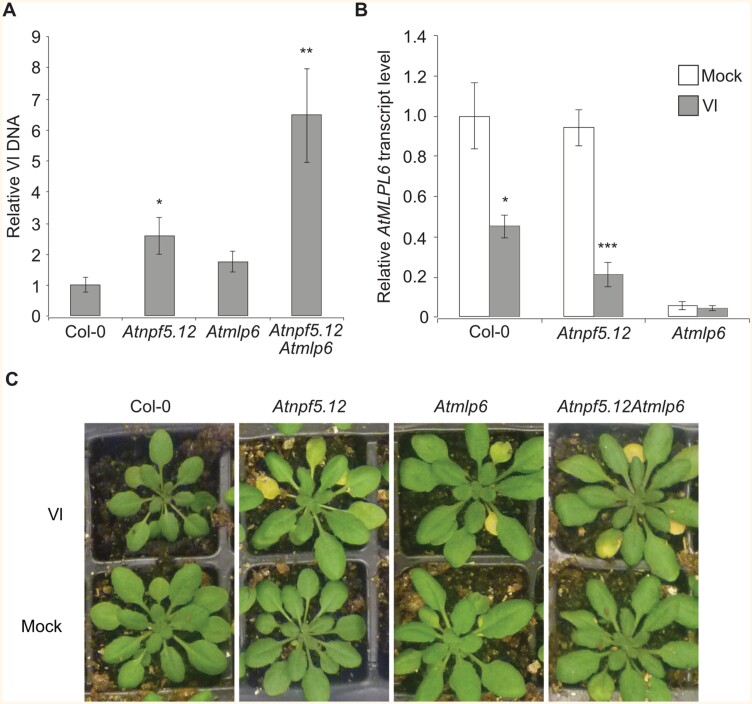
Phenotypes and fungal quantification in Arabidopsis mutants. (A) Relative fungal DNA content in roots of in vitro grown V. longisporum inoculated Atnpf5.12 (2.5-fold), Atmlp6 (1.8-fold), and Atnpf5.12 x Atmlp6 (6.5-fold) plants at 14 dpi. The data are relative to inoculated Col-0. Bar chart represents means ±SE (n=6 biological replicates of 20 plants for each plant line). (B) Relative AtMLP6 transcript levels in V. longisporum (Vl) inoculated roots of Col-0 (0.45-fold), Atnpf5.12 (0.2-fold), and Atmlp6 plants. The data are relative to mock-treated control plants at 2 dpi. Bar chart represents means ±SE (n=5 biological replicates of >20 plants for each plant line and treatment). Asterisks represent significant difference by Student’s t-test: *P≤0.05; **P≤0.01; ***P≤0.001. (C) Disease symptoms of soil-grown Col-0, Atnpf5.12, Atmlp6, Atnpf5.12 × Atmlp6 plants, 18 dpi with V. longisporum.
Salicylic acid-dependent responses are associated with AtNPF5.12 and AtMLP6
Previous reports indicate an increase in SA accumulation in B. napus 7 d after inoculation with V. longisporum (Ratzinger et al., 2009). To generate more information on the molecular functions of NPF5.12 and MLP6, we quantified SA levels in Col-0, Atnpf5.12, and Atmlp6 plants. Increased levels of SA were found in Col-0 and Atmlp6 2 d after inoculation compared with mock-treated Col-0, while SA levels in Atnpf5.12 were unaffected (Supplementary Fig. S11A). Transcription of WRKY70 and PATHOGENESIS-RELATED 1 (PR1) genes was negatively regulated in inoculated Atnpf5.12 and Atmlp6 plants compared with Col-0 (Supplementary Fig. S11B, C). Elevated levels of the jasmonic acid responsive gene PLANT DEFENSIN 1.2 (PDF1.2) were found as a response to fungal inoculation in both mutants (Supplementary Fig. S11D). These results suggest that NPF5.12 is an important activator of SA biosynthesis. NPF5.12 and MLP6 most likely excite the WRKY70 transcription factor impacting PR1 and PDF1.2 transcription. WRKY70-mediated activation of PR1 and repression of PDF1.2 is in agreement with previous reports (Li et al., 2004; Shim et al., 2013).
Defense responses of orthologous NPF5.12 and MLP6 genes in oilseed rape
Oilseed rape (Brassica napus) is the main crop suffering from stem stripe disease caused by V. longisporum (Depotter et al., 2016). Therefore, we were interested in clarifying any impact on the defense response by mutating the BnNPF5.12 and BnMLP6 orthologous genes through CRISPR/Cas9 technology. Four loci for BnNPF5.12 and 14 loci for BnMLP6 were identified with at least 98% and 87% amino acid sequence similarity, respectively (Supplementary Table S5). All identified loci were targeted with single-guide RNA in different combinations to generate the four multiple mutant types. Fifty-seven seed-setting transgenic lines were confirmed by PCR analyses. Two individual lines for each mutant type with a wild-type phenotype were chosen for further experiments. Target loci were sequenced to confirm mutations (Supplementary Table S7) and then used in disease screening. No visible disease phenotype was observed at 7 dpi, but mycelia had started to colonize roots (Fig. 4A, B). The multiple B. napus mutants Bnnpf5.12-1, Bnnpf5.12-2, Bnmlp6-1, Bnmlp6-2 and Bnnpf5.12/Bnmlp6-1 and Bnnpf5.12/Bnmlp6-2 all had significantly more fungal DNA relative to wild-type (Fig. 4C). Together with the data from Arabidopsis, this information demonstrates that functional NPF5.12 and MLP6 genes are required for defense against V. longisporum in both species.
Fig. 4.
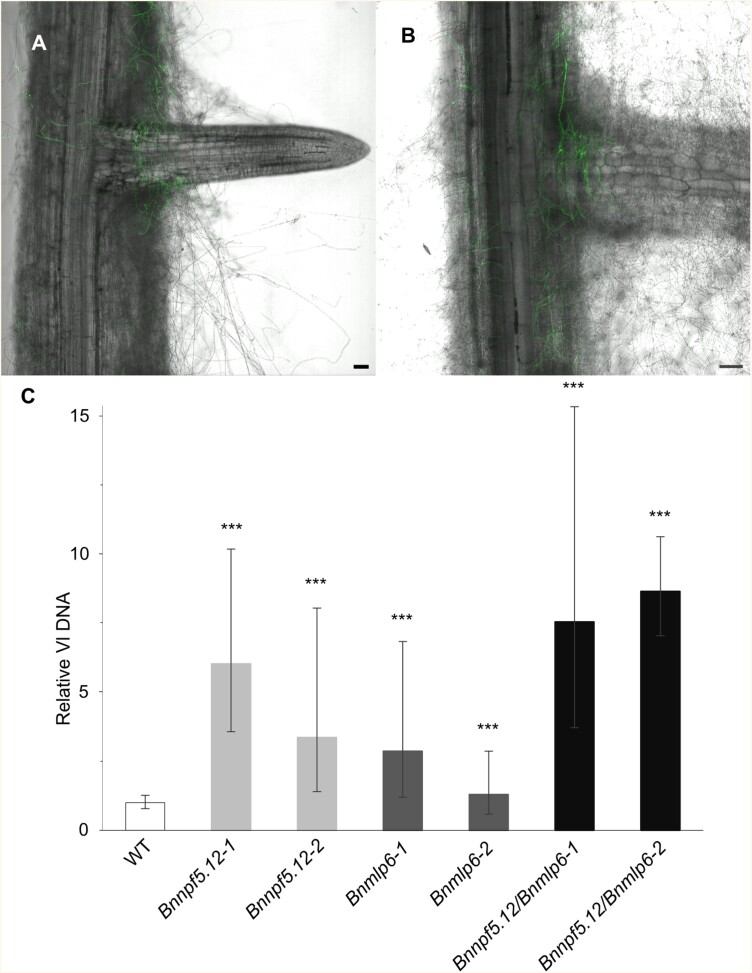
Responses to V. longisporum inoculation of Brassica napus and genome edited BnNPF5.12 and BnMLP lines. (A, B) Confocal microscopy image of 14-day-old B. napus cv. Kumily (WT) (A), and Bnnpf5.12-1 (B) inoculated by V. longisporum (Vl43:GFP). Photos taken 7 d post-infection. Scale bars: 50 µm. (C) Quantification of V. longisporum (Vl) DNA in Bnnpf5.12-1 (6-fold), Bnnpf5.12-2 (3.3-fold), Bnmlp6-1 (2.8-fold), Bnmlp6-2 (1.3-fold), Bnnpf5.12/Bnmlp6-1 (7.6-fold), and Bnnpf5.12/Bnmlp6-2 (8.7-fold) relative to WT. Bar chart represents mean fold change ±SD Vl DNA (n≥4 biological replicates of 10 roots for each plant line). The experiment was repeated twice. Asterisks represent statistical significance compared with WT (Student’s t-test: *P≤0.05; **P≤0.01; ***P≤0.001).
Colonization and growth of V. longisporum is affected by nitrogen availability
To clarify the importance of available nitrate for the infection process, Col-0, Atnpf5.12 and Atmlp6 plants growing in nitrogen-depleted and nitrogen-rich media were inoculated. The plants were sampled and quantified for responses to V. longisporum at two time points. Although colonization was severely limited due to a lack of nitrogen in all plant lines, both Atnpf5.12 and Atmlp6 showed a significant decrease in V. longisporum DNA at 14 dpi. The negative effect of nitrate depletion on fungal colonization was greatest in Col-0 (Supplementary Fig. S12). These results suggest that nitrate availability is an important factor for V. longisporum root colonization and that NPF5.12 has a substantial impact on the amount of nitrate accessible to the fungus.
Mutations in NPF5.12 and MLP6 reduce suberin deposition in the vascular system
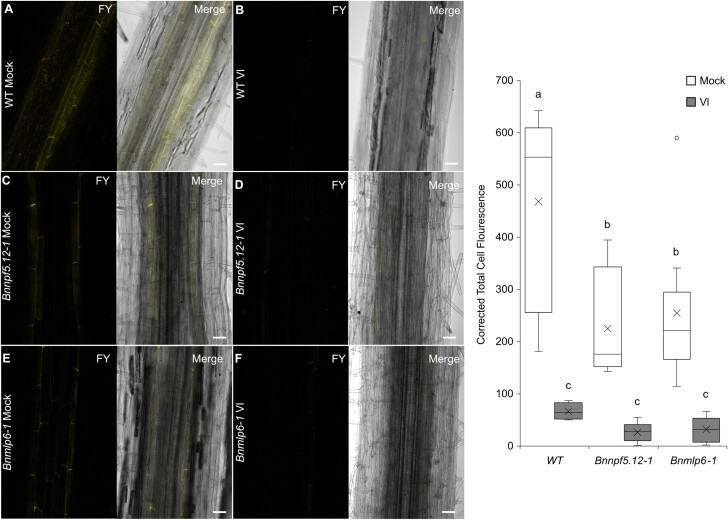
The root cellular architecture plays an important role in hindering pathogens from invading and colonizing intracellular structures (Kawa and Brady, 2022). The Casparian strip surrounds the vascular stele, and suberin lamellae are deposited around endodermal cells (Andersen et al., 2018), together controlling water and nutrient transport in a plant. Down-regulation of genes involved in Casparian strip formation and suberin deposition has been observed at 2 dpi of V. longisporum in Arabidopsis (Fröschel et al., 2021). To enhance our understanding of the importance of the vascular tissue during fungal colonization, we closely monitored the early infection phase in the roots of oilseed rape. Invasive hyphae swiftly penetrated the lateral cell layers of the primary root followed by growth in the cortex layer in a shoot-ward direction (Fig. 4A, B). At 7 dpi, there was a clear reduction in suberin in all inoculated materials compared with mock-treated roots (P≤0.001) (Fig. 5A–G). Bnnpf5.12-1 and Bnmlp6-1 plants had less suberin deposition than wild-type plants, suggesting a role for NPF5.12 and MLP6 in the polymerization processes of suberin. Similar attenuation of suberin was found in our Arabidopsis material (Supplementary Fig. S13A–G), which could explain the increased susceptibility of the npf5.12 and mlp6 mutant lines.
Fig. 5.
Suberin staining and quantification in genome edited Bnnpf5.12-1 and Bnmlp6-1 B. napus roots. (A, B) Fluorol Yellow 088 stained 14-day old roots of mock-treated and (A) V. longisporum inoculated (B) B. napus cv. Kumily (WT). (C, D) Mock-treated (C) and fungal inoculated (D) Bnnpf5.12-1 mutant. (E, F) Mock-treated (E) and fungal inoculated (F) Bnmlp6-1 mutant. (G) Quantification of Fluorol Yellow 088 fluorescence intensity in fungal infected (Vl) and mock-treated B. napus cv. Kumily (WT) and Bnnpf5.12-1 and Bnmlp6-1 mutant roots. Statistical significances are based on ANOVA/Tukey’s multiple comparisons of means (n≥4 roots per line and treatment, P≤0.001). Photographs were taken and quantification was performed 7 d post-treatment. Scale bars: 50 µm. FY, Fluorol Yellow 088; Merge, brightfield and FY composite image. The experiment was performed twice.
Discussion
Access to nitrate determines the degree of V. longisporoum infection
Plants use numerous importers and exporters to facilitate proper exchange of diverse metabolites and ions, either out of or into roots, events that are crucial for cellular functions and proper growth. Nitrate, the main source of nitrogen for plants, is involved in numerous processes (Fredes et al., 2019). Upon pathogen root colonization and external or internal growth, plant cells counteract nutrient loss by restricting nutrient transfer via reprogramming of nutrient metabolism and transport (Tünnermann et al., 2022). In this study, we identified the nitrate transporter NPF5.12 as important in the defense against V. longisporum. Nitrate is taken up by roots and translocated to vacuoles for use in metabolism (Martinoia et al., 2007). NPF5.12 was previously evaluated with regard to uptake of glucosinolates, but no such function was detected (Nour-Eldin et al., 2012). More recently, it was found that NPF5.12 is a tonoplast transporter and functions in allocation of nitrate from vacuoles to the cytosol (He et al., 2017). Overexpression of NPF5.12 results in reduced nitrate content in roots due to reallocation to shoot organs. Our data show that NPF5.12 localizes to the plasma membrane and that it may transport nitrate to the extracellular space. This result would explain the previously reported long-distance transport of nitrate in the NPF5.12 overexpression line (He et al., 2017). Accordingly, the p35S:NPF5.12Col-0 construct reduced fungal growth in the Can-0 genomic background (Fig. 1G). When we compared responses in nitrate dependency experiments, the relative difference in fungal growth was greater in Col-0 than in the npf5.12 mutant, which suggests the involvement of additional factors. Down-regulation of NPF5.12 after infection may be a starvation tactic employed by the plant. Similar responses have been found in other pathosystems, but the details are unclear (Sun et al., 2020). Functional redundancy among nitrate transporters has been assessed (Lu et al., 2022). Unexpectedly, nitrate uptake was higher in a sextuple mutant than in wild-type; additional transporters, such as chloride channels, were examined, and AtCLCa was found to be a major nitrate transporter candidate. We presume that the vast numbers of nitrogen and amino acid transporters are highly redundant to support the essential metabolic role of nitrogen in almost all developmental, growth, and metabolic plant processes. We believe that V. longisporum to some extent hijacks parts of the host’s nitrogen metabolism for colonization and growth.
Do major latex-like proteins function as intracellular shuttles activating defense responses?
Major latex-like proteins (MLPs) are plant specific and were first identified in the latex of opium poppy (Papaver somniferium), in which they are located in specific secretory cells: laticifers (Nessler et al., 1985). Laticifers are a large and heterogeneous group of cells that contain a wide range of compounds and secondary metabolites (Ramos et al., 2019; Ozber et al., 2022). Few plant species produce latex (Gracz-Bernaciak et al., 2021). In contrast, MLP genes are common in most plant genomes (25 in Arabidopsis), and it is thought that they exert divergent biological roles. Several MLP genes are implicated in defense against soil-borne pathogens in addition to activation of other stress agents and associations with hormone responses (Fujita and Inui, 2021). The data generated thus far are not conclusive, and gaps in biosynthesis and biological functions remain to be clarified.
A common feature of the 3D structure of MLP proteins is the hydrophobic cavity acting as a potential binding site for various ligand molecules (Li et al., 2023). AtMLP6 was previously detected in the phloem sap of Arabidopsis and thought to bind a hydrophobic systemic acquired resistance (SAR) signaling molecule (Carella et al., 2016). The role of SA, a main player in local and SAR responses in other plant defense systems, has not been clarified in regard to V. longisporum. Unlike many other pathosystems, there is a positive correlation between SA in the shoot extracts of inoculated B. napus and the biomass of V. longisporum (Ratzinger et al., 2009). In a follow-up study, B. napus infection stages were divided into two phases: early, with no symptoms, and late, when disease symptoms started to be visible (Zheng et al., 2019). The SA content declined after an initial infection stage, and the phenylpropanoid pathway in B. napus was activated in the resistant genotypes from 7 dpi and later, leading to a range of enhanced activation of enzymes, including those involved in lignin synthesis and ferulic acid and peroxidase. The different physiology between B. napus and Arabidopsis may explain divergent reported results. Differences in the spatio-temporal control of reproductive plant architecture illustrate the delicate balance of resources and related signals in the two species (Walker et al., 2021).
Early infection events and defense responses to V. longisporum
The major birch pollen allergen Bet v I is the prototypical member of the Bet v I superfamily to which MLP6 belongs (Morris et al., 2021). Its natural ligand was identified as quercetin-3-O-sophoroside (Q3SO), a glycosylated flavonoid that serves as an important signal for pollen germination (von Loetzen et al., 2014). Several MLPs closely related to MLP6, including the ZUSAMMEN CIS-CINNAMIC ACID ENHANCED genes ZCE1 and ZCE2, were previously implicated in flavonoid responses and plant development (Guo et al., 2011). Flavonoids might thus constitute an important part of the signaling response downstream of NPF5.12 and MLP6. Lipid-binding domains are also present in MLP6 and Bet v I, and the latter binds reversibly to the cytosolic side of the plasma membrane (Mogensen et al., 2007). Whether Bet v I is present as a monomer or dimer or whether ligand binding facilitates membrane translocation is currently unknown.
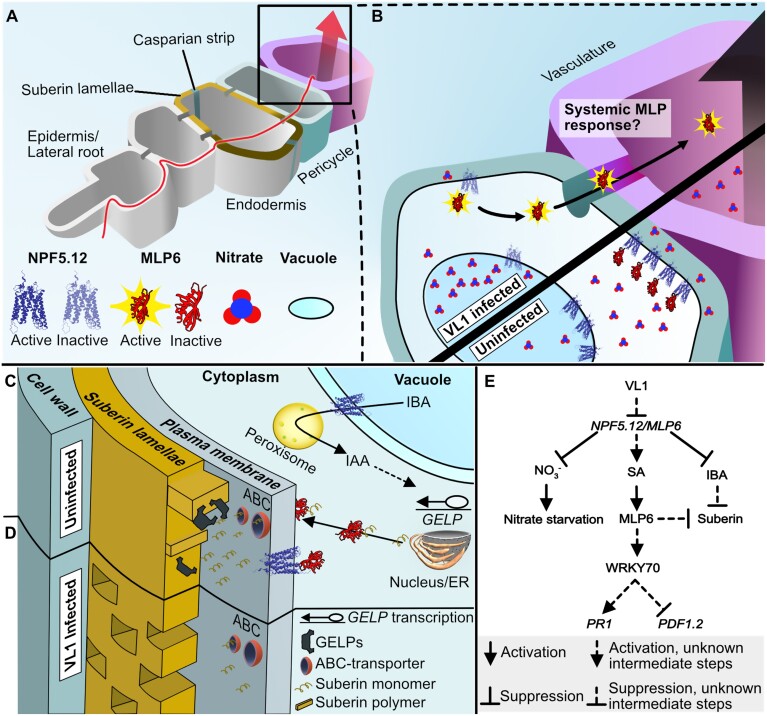
Many diverse factors have been implicated in defense against V. longisporum, though a clear picture that links them all together has not yet been presented. Here we summarize results from this investigation and earlier studies on the V. longisporum–plant interaction (Fig. 6). The root infection is triggered by secretion of volatile monoterpenes by the host (Roos et al., 2015), followed by hyphal entry at lateral root emergence sites (Fig. 6A). Disruption of the endodermal suberin lamellae enables hyphal entry into the vasculature. NPF5.12 proteins transport nitrates from the vacuole to the apoplastic space. NPF5.12 and MLP6 transcriptionally reprogram within 2 d post-infection in susceptible plants. This change limits available nitrates to the pathogen and restricts fungal growth (Fig. 6B). NPF5.12 and MLP6 contribute to suberin deposition in the endodermis (Fig. 6C, D). Gibberellin (GA), abscisic acid (ABA) and the auxin indole acetic acid (IAA) all regulate the suberization process (Woolfson et al., 2022). The AtNPF2.12 and AtNPF2.13 transporters have been implicated in GA and ABA accumulation and translocation from the shoot to the root (Binenbaum et al., 2023). A third GA and ABA transporter, AtNPF2.14, regulates the formation of suberin in the root.
Fig. 6.
A proposed working model for the plant–V. longisporum interaction. (A) Verticillium longisporum (VL1, red line) hyphae enter the root at lateral root emergence sites. Disruption of the endodermal suberin lamellae enables hyphal entry into the vasculature. (B) NPF5.12 proteins transport nitrates from the vacuole to the apoplastic space. Two days post-infection, NPF5.12 and MLP6 transcriptionally reprogram. This change limits available nitrates to the pathogen and restricts fungal growth in susceptible plants. MLP6 migrate systemically though the vasculature. (C) NPF5.12 and MLP6 contribute to suberin deposition in the endodermis. The MLP6 carrier protein transports suberin monomers from the endoplasmic reticulum (ER) to the cell periphery. ABC-proteins transport suberin monomers to the apoplast. NPF5.12 contributes by transporting indole butyric acid (IBA), which is converted to indole acetic acid (IAA) in peroxisomes. Accumulation of endodermal IAA activates several Gly–Asp–Ser–Leu (GDSL)-type esterase/lipase proteins (GELPs), which polymerizes the suberin. (D) This process is interrupted by VL1 infection, causing loss of suberin polymerization. (E) VL1 infection and NPF5.12/MLP6 reprogramming trigger salicylic acid (SA) accumulation at two dpi. MLP6 mRNA and proteins migrate through plasmodesmata and phloem to increase WKRY70 transcription. WRKY70 contributes to transcriptional PR1 gene activation, and at the same time PDF1.2 repression.
It has been suggested that carrier proteins transport suberin monomers from the endoplasmic reticulum toward the plasma membrane (Serra and Geldner, 2022). ABC-type transporters then shuttle the monomers to the apoplast. MLP6 may contribute to suberin deposition by delivering the hydrophobic suberin monomers to the cell periphery (Fig. 6C). NPF5.12 transports indole butyric acid (IBA) to the cytoplasm (Michniewicz et al., 2019), and IBA is β-oxidized to IAA in the peroxisomes (Strader et al., 2010). This information links NPF5.12 to the regulation of suberin polymerization. Accumulation of IAA in the endodermis activates transcription of several key suberin esterase/lipase enzymes responsible for polymerization of the suberin monomers and degradation of polymers (Ursache et al., 2021). The latter process is interrupted by V. longisporum infection, causing loss of suberin polymerization (Fig. 6D). We assume that MLP6 migrates in the phloem sap and influences transcription factors, genes, and/or metabolites such as peroxide and flavonoids that affect suberin (Floerl et al., 2012; Woolfson et al., 2022). Activation of peroxisomal processes in the plant host could generate peroxide and jasmonic acid (Roos et al., 2014). The latter is negatively regulated by the Rab GTPase-activating protein RabGAP22. In addition, to the components discussed above, phytohormones are induced (Fig. 6E). NPF5.12 and MLP6 are involved in induction of SA-dependent defense responses during early stages of infection. Colonization of this fungus also depends on available nitrates in the host root. Accordingly, the negative effect of nitrate depletion on fungal growth was less pronounced in Atnpf5.12 plants with impaired nitrate transport. MLP6 triggers WRKY70, which regulates PR1 and PDF1.2 in opposite directions. PR1 is promoted whereas PDF1.2 is repressed. Much remains to be elucidated among these factors and their implications on defense against V. longisporum.
Supplementary data
The following supplementary data are available at JXB online.
Fig. S1. Disease phenotype scoring scale.
Fig. S2. Transcription level of ACTIN2 in all experimental conditions.
Fig. S3. Fungal DNA accumulation in Brassica napus.
Fig. S4. Linkage map and QTL analysis of Can-0 and Sei-0 accessions.
Fig. S5. Differentially expressed genes in the mapped region of chromosome 1.
Fig. S6. Transcription profile of genes with high sequence similarity to AtNPF5.12.
Fig. S7. Predicted protein structure of AtNPF5.12 and AtMLP6.
Fig. S8. Immunoprecipitation of AtNPF5.12 and MALDI-MS/MS analysis.
Fig. S9. Relative fungal DNA content in Atmlp6.
Fig. S10. Transcription profile of At4g23680.
Fig. S11. Salicylic acid-dependent responses associated with NPF5.12 and MLP6.
Fig. S12. Fungal DNA content in nitrogen-depleted conditions.
Fig. S13. Suberin staining in Arabidopsis.
Table S1. T-DNA insertion mutants.
Table S2. qPCR and qRT-PCR primer sequences.
Table S3. Arabidopsis construct information.
Table S4. Fungal DNA content in additional transgenic Arabidopsis lines.
Table S5. Single-guide RNA (sgRNA) information.
Table S6. Genes mutated in B. napus.
Table S7. Brassica napus tissue culture media.
Table S8. Genotype of CRISPR/Cas9-edited B. napus.
Table S9. Primers used in cloning and sequencing B. napus.
Dataset S1. Can-0 and Sei-0 SNP markers.
Acknowledgements
We thank Dr Tina Olsson for help with the plant material. Lantmännen Lantbruk, Svalöv provided the B. napus seeds. We thank Dr Åke Engström for the MALDI-MS/MS analyses and the Science for Life Laboratory (SciLife, Uppsala) for support with the SNP genotyping.
Glossary
Abbreviations
- BiFC
bimolecular fluorescence complementation
- dpi
days post-inoculation
- MLP
major latex protein
- NPF
nitrate/peptide transporter
- RIL
recombinant inbred line
- SA
salicylic acid
Contributor Information
Fredrik Dölfors, Swedish University of Agricultural Sciences, Department of Plant Biology, Uppsala BioCenter, Linnean Center for Plant Biology, P.O. Box 7080, S-75007 Uppsala, Sweden.
Jonas Ilbäck, Swedish University of Agricultural Sciences, Department of Plant Biology, Uppsala BioCenter, Linnean Center for Plant Biology, P.O. Box 7080, S-75007 Uppsala, Sweden.
Sarosh Bejai, Swedish University of Agricultural Sciences, Department of Plant Biology, Uppsala BioCenter, Linnean Center for Plant Biology, P.O. Box 7080, S-75007 Uppsala, Sweden.
Johan Fogelqvist, Swedish University of Agricultural Sciences, Department of Plant Biology, Uppsala BioCenter, Linnean Center for Plant Biology, P.O. Box 7080, S-75007 Uppsala, Sweden.
Christina Dixelius, Swedish University of Agricultural Sciences, Department of Plant Biology, Uppsala BioCenter, Linnean Center for Plant Biology, P.O. Box 7080, S-75007 Uppsala, Sweden.
Monica Höfte, University of Ghent, Belgium.
Author contributions
CD, FD, and JI conceived the research. FD, JI, and SB performed the experiments, FD, JI, and JF performed the data analysis. FD, JI, and CD wrote the manuscript.
Conflict of interest
The authors declare that there are no conflicts of interest.
Funding
This work was supported by research grants from the following foundations: Nilsson-Ehle, Helge Ax:son Johnson (F20-0361), Memory of Oscar and Lili Lamm, the Swedish Oilseed Rape Foundation, the Research Council Formas (2017-00827), the Swedish University of Agricultural Sciences, and support with SNP genotyping at the Science for Life Laboratory (SciLife, Uppsala) from VR and Wallenberg Foundation.
Data availability
All data supporting the findings of this study, including supplementary materials, are available from the corresponding author upon request.
References
- Alonso JM, Stepanova AN, Leisse TJ, et al. 2003. Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301, 653–657. [DOI] [PubMed] [Google Scholar]
- Andersen TG, Naseer S, Ursache R, Wybouw B, Smet W, De Rybel B, Vermeer JEM, Geldner N.. 2018. Diffusible repression of cytokinin signalling produces endodermal symmetry and passage cells. Nature 555, 529–533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bae S, Park J, Kim JS.. 2014. Cas-OFFinder: a fast and versatile algorithm that searches for potential off-target sites of Cas9 RNA-guided endonucleases. Bioinformatics 30, 1473–1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barberon M, Vermeer J, De Bellis D, et al. 2016. Adaptation of root function by nutrient-induced plasticity of endodermal differentiation. Cell 164, 447–459. [DOI] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y.. 1995. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B 57, 289–300. [Google Scholar]
- Binenbaum J, Wulff N, Camut L, et al. 2023. Gibberellin and abscisic acid transporters facilitate endodermal suberin formation in Arabidopsis. Nature Plants 9, 785–802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bohman S, Staal J, Thomma BPHJ, Wang M, Dixelius C.. 2004. Characterisation of an Arabidopsis-Leptosphaeria maculans pathosystem: resistance partially requires camalexin biosynthesis and is independent of salicylic acid, ethylene and jasmonic acid signalling. The Plant Journal 37, 9–20. [DOI] [PubMed] [Google Scholar]
- Carella P, Merl-Pham J, Wilson DC, Dey S, Hauck SM, Vlot AC, Cameron RK.. 2016. Comparative proteomics analysis of phloem exudates collected during the induction of systemic acquired resistance. Plant Physiology 171, 1495–1510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis AM, Hall A, Millar AJ, Darrah C, Davis SJ.. 2009. Protocol: streamlined sub-protocols for floral-dip transformation and selection of transformants in Arabidopsis thaliana. Plant Methods 5, 3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Depotter JRL, Deketelaere S, Inderbitzin P, Tiedemann AV, Höfte M, Subbarao KV, Wood TA, Thomma BPHJ.. 2016. Verticillium longisporum, the invisible threat to oilseed rape and other brassicaceous plant hosts. Molecular Plant Pathology 17, 1004–1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Depotter JRL, Thomma BPHJ, Wood TA.. 2019. Measuring the impact of Verticillium longisporum on oilseed rape (Brassica napus) yield in field trials in the United Kingdom. European Journal of Plant Pathology 153, 321–326. [Google Scholar]
- Doyle JJ, Doyle JL.. 1987. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin 19, 11–15. [Google Scholar]
- Dunker S, Keunecke H, Steinbach P, von Tiedemann A.. 2008. Impact of Verticillium longisporum on yield and morphology of winter oilseed rape (Brassica napus) in relation to systemic spread in the plant. Journal of Phytopathology 156, 698–707. [Google Scholar]
- Eynck C, Koopmann B, Grunewaldt-Stoecker G, Karlovsky P, Tiedemann A.. 2007. Differential interactions of Verticillium longisporum and V. dahliae with Brassica napus detected with molecular and histological techniques. European Journal of Plant Pathology 118, 259–274. [Google Scholar]
- Floerl S, Majcherczyk A, Possienke M, Feussner K, Tappe H, Gatz C, Feussner I, Kües U, Polle A.. 2012. Verticillium longisporum infection affects the leaf apoplastic proteome, metabolome, and cell wall properties in Arabidopsis thaliana. PLoS ONE, 7, e31435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fogelqvist J, Tzelepis G, Bejai S, Ilbäck J, Schwelm A, Dixelius C.. 2018. Analysis of the hybrid genomes of two field isolates of the soil-borne fungal species Verticillium longisporum. BMC Genomics 19, 14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fradin EF, Abd-El-Haliem A, Masini L, van den Berg GCM, Joosten MHAJ, Thomma BPHJ.. 2011. Interfamily transfer of tomato Ve1 mediates Verticillium resistance in Arabidopsis. Plant Physiology 156, 2255–2265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fredes I, Moreno S, Díaz FP, Gutiérrez RA.. 2019. Nitrate signaling and the control of Arabidopsis growth and development. Current Opinion in Plant Biology 47, 112–118. [DOI] [PubMed] [Google Scholar]
- Fröschel C, Komorek J, Attard A, et al. 2021. Plant roots employ cell-layer specific programs to respond to pathogenic and beneficial microbes. Cell Host & Microbe 29, 299–310.e7. [DOI] [PubMed] [Google Scholar]
- Fujita K, Inui H.. 2021. Review: Biological functions of major latex-like proteins in plants. Plant Science 306, 110856. [DOI] [PubMed] [Google Scholar]
- Gan X, Stegle O, Behr J, et al. 2011. Multiple reference genomes and transcriptomes for Arabidopsis thaliana. Nature 477, 419–423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautier L, Cope L, Bolstad BM, Irizarry RA.. 2004. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315. [DOI] [PubMed] [Google Scholar]
- Gracz-Bernaciak J, Mazur O, Nawrot R.. 2021. Functional studies of plant latex as a rich source of bioactive compounds: focus on proteins and alkaloids. International Journal of Molecular Sciences 22, 12427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guettou F, Quistgaard EM, Trésaugues L, Moberg P, Jegerschöld C, Zhu L, Jong AJO, Nordlund P, Löw C.. 2013. Structural insights into substrate recognition in proton-dependent oligopeptide transporters. EMBO Reports 14, 804–810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo D, Wong WS, Xu WZ, Sun FF, Qing DJ, Li N.. 2011. Cis-cinnamic acid-enhanced 1 gene plays a role in regulation of Arabidopsis bolting. Plant Molecular Biology 75, 481–495. [DOI] [PubMed] [Google Scholar]
- He YN, Peng J-S, Cai Y, Liu DF, Guan Y, Yi HY, Gong JM.. 2017. Tonoplast-localized nitrate uptake transporters involved in vacuolar nitrate efflux and reallocation in Arabidopsis. Scientific Reports 7, 6417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heale JB, Karapapa VK.. 1999. The Verticillium threat to Canada’s major oilseed crop: Canola. Canadian Journal of Plant Pathology 21, 1–7. [Google Scholar]
- Horton M, Hancock A, Huang Y, et al. 2012. Genome-wide patterns of genetic variation in worldwide Arabidopsis thaliana accessions from the RegMap panel. Nature Genetics 44, 212–216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hyams G, Abadi S, Lahav S, Avni A, Halperin E, Shani E, Mayrose I.. 2018. CRISPys: optimal sgRNA design for editing multiple members of a gene family using the CRISPR system. Journal of Molecular Biology 430, 2184–2195. [DOI] [PubMed] [Google Scholar]
- Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP.. 2003. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264. [DOI] [PubMed] [Google Scholar]
- Islam S, Islam R, Kandwal P, Khanam S, Proshad R, Kormoker T, Tusher TR.. 2022. Nitrate transport and assimilation in plants; a potential review. Archives of Agronomy and Soil Science 68, 133–150. [Google Scholar]
- Jambagi S, Dixelius C.. 2023. A robust hydroponic-based system for screening red clover (Trifolium pratense) for Fusarium avenaceum. Legume Science 5, e209. [Google Scholar]
- Joehanes R, Nelson JC.. 2008. QGene 4.0, an extensible Java QTL-analysis platform. Bioinformatics 24, 2788–2789. [DOI] [PubMed] [Google Scholar]
- Johansson A, Goud JKC, Dixelius C.. 2006a. Plant host range of Verticillium longisporum and microsclerotia density in Swedish soils. European Journal of Plant Pathology 114, 139–149. [Google Scholar]
- Johansson A, Staal J, Dixelius C.. 2006b. Early responses in the Arabidopsis-Verticillium longisporum pathosystem are dependent on NDR1, JA- and ET-associated signals via cytosolic NPR1 and RFO1. Molecular Plant-Microbe Interactions 19, 958–969. [DOI] [PubMed] [Google Scholar]
- Kang J, Park J, Choi H, Burla B, Kretzschmar T, Lee Y, Martinoia E.. 2011. Plant ABC transporters. The Arabidopsis Book 9, e0153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanno Y, Hanada A, Chiba Y, Ichikawa T, Nakazawa M, Matsui M, Koshiba T, Kamiya Y, Seo M.. 2012. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proceedings of the National Academy of Sciences, USA 109, 9653–9658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kanstrup C, Nour-Eldin HH.. 2022. The emerging role of the nitrate and peptide transporter family: NPF in plant specialized metabolism. Current Opinion in Plant Biology 68, 102243. [DOI] [PubMed] [Google Scholar]
- Kawa D, Brady SM.. 2022. Root cell types as an interface for biotic interactions. Trends in Plant Science 27, 1173–1186. [DOI] [PubMed] [Google Scholar]
- Kelley LA, Sternberg MJE.. 2009. Protein structure prediction on the web: a case study using the Phyre server. Nature Protocols 4, 363–371. [DOI] [PubMed] [Google Scholar]
- Klosterman SJ, Subbarao KV, Kang S, et al. 2011. Comparative genomics yields insights into niche adaptation of plant vascular wilt pathogens. PLoS Pathogens 7, e1002137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kover PX, Valdar W, Trakalo J, Scarcelli N, Ehrenreich IM, Purugganan MD, Durrant C, Mott R.. 2009. A multiparent advanced generation inter-cross to fine-map quantitative traits in Arabidopsis thaliana. PLoS Genetics 5, e1000551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krogh A, Larsson B, von Heijne G, Sonnhammer EL.. 2001. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. Journal of Molecular Biology 305, 567–580. [DOI] [PubMed] [Google Scholar]
- Kurt F, Filiz E.. 2022. Functional divergence in oligopeptide transporters in plants. In: Upadhyay SK, ed. Cation transporters in plants. Academic Press, 167–183. [Google Scholar]
- Lander E, Green P, Abrahamson J, Barlow A, Daly M, Lincoln S, Newburg L.. 1987. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181. [DOI] [PubMed] [Google Scholar]
- Léran S, Varala K, Boyer J-C, et al. 2014. A unified nomenclature of NITRATE TRANSPORTER 1/PEPTIDE TRANSPORTER family members in plants. Trends in Plant Science 19, 5–9. [DOI] [PubMed] [Google Scholar]
- Li J, Brader G, Palva ET.. 2004. The WRKY70 transcription factor: a node of convergence for jasmonate-mediated and salicylate-mediated signals in plant defense. The Plant Cell 16, 319–331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J, Zeng R, Huang Z, et al. 2023. Genome-wide characterization of major latex protein gene family in peanut and expression analyses under drought and waterlogging stress. Frontiers in Plant Science 14, 1152824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li M, Doll J, Weckermann K, Oecking C, Berendzen KW, Schöffl F.. 2010. Detection of in vivo interactions between Arabidopsis class A-HSFs, using a novel BiFC fragment, and identification of novel class B-HSF interacting proteins. European Journal of Cell Biology 89, 126–132. [DOI] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD.. 2001. Analysis of relative gene expression data using real-time quantitative PCR and the method. Methods 25, 402–408. [DOI] [PubMed] [Google Scholar]
- Lu Y-T, Liu D-F, Wen T-T, Fang Z-J, Chen S-Y, Li H, Gong J-M.. 2022. Vacuolar nitrate efflux requires multiple functional redundant nitrate transporter in Arabidopsis thaliana. Frontiers in Plant Science 13, 926809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lubkowitz M. 2011. The oligopeptide transporters: a small gene family with a diverse group of substrates and functions? Molecular Plant 4, 407–415. [DOI] [PubMed] [Google Scholar]
- Martin T, Biruma M, Fridborg I, Okori P, Dixelius C.. 2011. A highly conserved NB-LRR encoding gene cluster effective against Setosphaeria turcica in sorghum. BMC Plant Biology 11, 151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martinoia E, Maeshima M, Neuhaus HE.. 2007. Vacuolar transporters and their essential role in plant metabolism. Journal of Experimental Botany 58, 83–102. [DOI] [PubMed] [Google Scholar]
- McCloy RA, Rogers S, Caldon CE, Lorca T, Castro A, Burgess A.. 2014. Partial inhibition of Cdk1 in G2 phase overrides the SAC and decouples mitotic events. Cell Cycle 13, 1400–1412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michniewicz M, Ho CH, Enders TA, et al. 2019. TRANSPORTER OF IBA1 links auxin and cytokinin to influence root architecture. Developmental Cell 50, 599–609.e4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller AJ, Fan X, Orsel M, Smith SJ, Wells DM.. 2007. Nitrate transport and signalling. Journal of Experimental Botany 58, 2297–2306. [DOI] [PubMed] [Google Scholar]
- Mogensen JE, Ferreras M, Wimmer R, Petersen SV, Enghild JJ, Otzen DE.. 2007. The major allergen from birch tree pollen, Bet v 1, binds and permeabilizes membranes. Biochemistry 46, 3356–3365. [DOI] [PubMed] [Google Scholar]
- Morris JS, Caldo KMP, Liang S, Facchini PJ.. 2021. PR10/Bet v1-like proteins as novel contributors to plant biochemical diversity. ChemBioChem 22, 264–287. [DOI] [PubMed] [Google Scholar]
- Nelson BK, Cai X, Nebenführ A.. 2007. A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. The Plant Journal 51, 1126–1136. [DOI] [PubMed] [Google Scholar]
- Nessler CL, Allen RD, Galewsky S.. 1985. Identification and characterization of latex-specific proteins in opium poppy. Plant Physiology 79, 499–504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nour-Eldin HH, Andersen TG, Burow M, Madsen SR, Jørgensen ME, Olsen CE, Dreyer I, Hedrich R, Geiger D, Halkier BA.. 2012. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 488, 531–534. [DOI] [PubMed] [Google Scholar]
- Osawa H, Stacey G, Gassmann W.. 2006. ScOPT1 and AtOPT4 function as proton-coupled oligopeptide transporters with broad but distinct substrate specificities. The Biochemical Journal 393, 267–275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ozber N, Carr SC, Morris JS, Liang S, Watkins JL, Caldo KM, Hagel JM, Ng KKS, Faccini PJ.. 2022. Alkaloid binding to opium poppy major latex proteins triggers structural modification and functional aggregation. Nature Communication 13, 6768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker JL, Newstead S.. 2014. Molecular basis of nitrate uptake by the plant nitrate transporter NRT1.1. Nature 507, 68–72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pegg GF, Brady BL.. 2002. Verticillium wilts. Wallingford: CABI Publishing. [Google Scholar]
- Petersen B, Nordahl Petersen T, Andersen P, Nielsen M, Lundegaard C.. 2009. A generic method for assignment of reliability scores applied to solvent accessibility predictions. BMC Structural Biology 9, 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prykhozhij SV, Rajan V, Gaston D, Berman JN.. 2015. CRISPR MultiTargeter: a web tool to find common and unique CRISPR single guide targets in a set of similar sequences. PLoS One 10, e0138634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- R Core Team. 2016. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing. [Google Scholar]
- Ramos VR, Demarco D, da Costa Souza IC, de Freitas CDT.. 2019. Laticifers, latex, and their role in plant defense. Trends in Plant Science 24, 553–567. [DOI] [PubMed] [Google Scholar]
- Ratzinger A, Riediger N, von Tiedemann A, Karlovsky P.. 2009. Salicylic acid and salicylic acid glucoside in xylem sap of Brassica napus infected with Verticillium longisporum. Journal of Plant Research 122, 571–579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rentsch D, Schmidt S, Tegeder M.. 2007. Transporters for uptake and allocation of organic nitrogen compounds in plants. FEBS Letters 581, 2281–2289. [DOI] [PubMed] [Google Scholar]
- Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK.. 2015. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Research 43, e47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roos J, Bejai S, Oide S, Dixelius C.. 2014. RabGAP22 is required for defense to the vascular pathogen Verticillium longisporum and contributes to stomata immunity. PLoS One 9, e88187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roos J, Bejai S, Mozūraitis R, Dixelius C.. 2015. Susceptibility to Verticillium longisporum is linked to mono-terpene production by TPS23/27 in Arabidopsis. The Plant Journal 81, 572–585. [DOI] [PubMed] [Google Scholar]
- Rozen S, Skaletsky HJ.. 2000. Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S, eds. Bioinformatics methods and protocols: methods in molecular biology. Totowa, NJ: Humana Press Inc., 365–386. [DOI] [PubMed] [Google Scholar]
- Schaaf G, Ludewig U, Erenoglu BE, Mori S, Kitahara T, von Wirén N.. 2004. ZmYS1 functions as a proton-coupled symporter for phytosiderophore- and nicotianamine-chelated metals. The Journal of Biological Chemistry 279, 9091–9096. [DOI] [PubMed] [Google Scholar]
- Schmittgen TD, Livak KJ.. 2008. Analyzing real-time PCR data by the comparative CT method. Nature Protocol 3, 1101–1108. [DOI] [PubMed] [Google Scholar]
- Schröder M, Dixelius C, Råhlén R, Glimelius K.. 1994. Transformation of Brassica napus by using the aadA gene as selectable marker and inheritance studies of the marker genes. Physiologia Plantarum 92, 37–46. [Google Scholar]
- Schütze K, Harter K, Chaban C.. 2009. Bimolecular fluorescence complementation (BiFC) to study protein-protein interactions in living plant cells. Methods in Molecular Biology 479, 189–202. [DOI] [PubMed] [Google Scholar]
- Serra O, Geldner N.. 2022. The making of suberin. New Phytologist 235, 848–866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shim JS, Jung C, Lee S, Min K, Lee Y-W, Choi Y, Lee JS, Song JT, Kim J-K, Choi YD.. 2013. AtMYB44 regulates WRKY70 expression and modulates antagonistic interaction between salicylic acid and jasmonic acid signaling. The Plant Journal 73, 483–495. [DOI] [PubMed] [Google Scholar]
- Singh S, Braus-Stromeyer SA, Timpner C,. et al. 2010. Silencing of Vlaro2 for chorismate synthase revealed that the phytopathogen Verticillium longisporum induces the cross-pathway control in the xylem. Applied Microbiology and Biotechnology 85, 1961–1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solcan N, Kwok J, Fowler PW, Cameron AD, Drew D, Iwata S, Newstead S.. 2012. Alternating access mechanism in the POT family of oligopeptide transporters. The EMBO Journal 31, 3411–3421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strader LC, Culler AH, Cohen JD, Bartel B.. 2010. Conversion of endogenous indole-3-butyric acid to indole-3-acetic acid drives cell expansion in Arabidopsis seedlings. Plant Physiology 153, 1577–1586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J, Bankston JR, Payandeh J, Hinds TR, Zagotta WN, Zheng N.. 2014. Crystal structure of the plant dual-affinity nitrate transporter NRT1.1. Nature 507, 73–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Y, Wang M, Mur LAJ, Shen Q, Guo S.. 2020. Unravelling the roles of nitrogen nutrition in plant disease defenses. International Journal of Molecular Sciences 21, 572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tjamos SE, Flemetakis E, Paplomatas EJ, Katinkas P.. 2005. Induction of resistance to Verticillium dahliae in Arabidopsis thaliana by the biocontrol agent K-165 and pathogenesis-related proteins gene expression. Molecular Plant-Microbe Interactions 6, 555–561. [DOI] [PubMed] [Google Scholar]
- Tünnermann L, Colou J, Näsholm T, Gratz R.. 2022. To have or not to have: expression of amino acid transporters during pathogen infection. Plant Molecular Biology 109, 413–425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ursache R, De Jesus Vieira Teixeira C, Dénervaud Tendon V, et al. 2021. GDSL-domain proteins have key roles in suberin polymerization and degradation. Nature Plants 7, 353–364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Loetzen CS, Hoffmann T, Hartl MJ, Schweimer K, Schwab W, Rösch P, Hartl-Spiegelhauer O.. 2014. Secret of the major birch pollen allergen Bet v 1: identification of the physiological ligand. Biochemical Journal 457, 379–390. [DOI] [PubMed] [Google Scholar]
- Walker CH, Wheeldon CD, Bennett T.. 2021. Integrated dominance mechanisms regulate reproductive architecture in Arabidopsis thaliana and Brassica napus. Plant Physiology 186, 1985–2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Strelkov SE, Hwang S-F.. 2023. Blackleg yield losses and interactions with Verticillium stripe in canola (Brassica napus) in Canada. Plants 12, 434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weichert A, Brinkmann C, Komarova NY, Dietrich D, Thor K, Meier S, Suter Grotemeyer M, Remtsch D.. 2011. AtPTR4 and AtPTR6 are differentially expressed, tonoplast-localized members of the peptide transporter/nitrate transporter 1 (PTR/NRT) family. Planta 235, 311–323. [DOI] [PubMed] [Google Scholar]
- Weigel D, Glazebrook J.. 2006. Transformation of Agrobacterium using the freeze-thaw method. Cold Spring Harbor Protocols 2006, pdb.prot4666. [DOI] [PubMed] [Google Scholar]
- Wickham, H. 2009. ggplot2: elegant graphics for data analysis. New York: Springer-Verlag. [Google Scholar]
- Woolfson KN, Esfandiari M, Bernards MA.. 2022. Suberin biosynthesis, assembly, and regulation. Plants 11, 555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xing HL, Dong L, Wang ZP, Zhang HY, Han CY, Liu B, Wang X-C, Chen J.. 2014. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biology 14, 327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yadeta KA, Thomma BP.. 2013. The xylem as battleground for plant hosts and vascular wilt pathogens. Frontiers in Plant Science 4, 97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng X, Koopmann B, von Tiedemann A.. 2019. Role of salicylic acid and components of the phenylpropanoid pathway in basal and cultivar-related resistance of oilseed rape (Brassica napus) to Verticillium longisporum. Plants 8, 491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou L, Hu Q, Johansson A, Dixelius C.. 2006. Verticillium longisporum and V. dahliae: infection and disease in Brassica napus. Plant Pathology 55, 137–144. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data supporting the findings of this study, including supplementary materials, are available from the corresponding author upon request.