ABSTRACT
The majority of newly discovered archaeal lineages remain without a cultivated representative, but scarce experimental data from the cultivated organisms show that they harbor distinct functional repertoires. To unveil the ecological as well as evolutionary impact of Archaea from metagenomics, new computational methods need to be developed, followed by in-depth analysis. Among them is the genome-wide protein fusion screening performed here. Natural fusions and fissions of genes not only contribute to microbial evolution but also complicate the correct identification and functional annotation of sequences. The products of these processes can be defined as fusion (or composite) proteins, the ones consisting of two or more domains originally encoded by different genes and split proteins, and the ones originating from the separation of a gene in two (fission). Fusion identifications are required for proper phylogenetic reconstructions and metabolic pathway completeness assessments, while mappings between fused and unfused proteins can fill some of the existing gaps in metabolic models. In the archaeal genome-wide screening, more than 1,900 fusion/fission protein clusters were identified, belonging to both newly sequenced and well-studied lineages. These protein families are mainly associated with different types of metabolism, genetic, and cellular processes. Moreover, 162 of the identified fusion/fission protein families are archaeal specific, having no identified fused homolog within the bacterial domain. Our approach was validated by the identification of experimentally characterized fusion/fission cases. However, around 25% of the identified fusion/fission families lack functional annotations for both composite and split states, showing the need for experimental characterization in Archaea.
IMPORTANCE
Genome-wide fusion screening has never been performed in Archaea on a broad taxonomic scale. The overlay of multiple computational techniques allows the detection of a fine-grained set of predicted fusion/fission families, instead of rough estimations based on conserved domain annotations only. The exhaustive mapping of fused proteins to bacterial organisms allows us to capture fusion/fission families that are specific to archaeal biology, as well as to identify links between bacterial and archaeal lineages based on cooccurrence of taxonomically restricted proteins and their sequence features. Furthermore, the identification of poorly characterized lineage-specific fusion proteins opens up possibilities for future experimental and computational investigations. This approach enhances our understanding of Archaea in general and provides potential candidates for in-depth studies in the future.
KEYWORDS: large-scale screening, comparative genomics, archaeal evolution, archaeal biology, bacterial fusions
INTRODUCTION
Recent advancements in sequencing techniques have revolutionized our ability to explore the prokaryotic world, especially Archaea (1–4), by providing vast amounts of metagenomic data at a cost-effective rate. Despite the daily deposition of thousands of metagenome-assembled genomes (MAGs) in public databases (5), the isolation and cultivation of these newly discovered microbial lineages remains challenging and time-consuming (6–10). In the absence of cultivated representatives, the functional repertoire of the discovered lineages remains largely unknown, with metabolic predictions being made based only on their genomic repertoire. Archaea are of particular interest due to their less studied biology as compared to Bacteria (11), with many unknowns to be discovered.
Unlike Bacteria, Archaea were discovered much later, in the 1970s (12), and were recognized as a separate domain of life in the 1990s (13). They thrive in diverse habitats and often dominate in extreme environments (11). Their cultivation, similar to some bacteria, imposes some challenges such as long generation times and metabolite limitations in the absence of syntropic/symbiotic partners outside of their natural habitat (6, 8, 9, 14). As a consequence, the isolation and cultivation of pure cultures and co-cultures for Archaea have been successful only for a few high-rank lineages out of the 24 known phyla (15). Moreover, microbiologists have been working on bacteria longer than with archaea which can also explain why there are more bacterial isolates than archaeal ones. However, existing (meta)genome-based metabolic reconstructions (8, 16–20), growth experiments (21–24), and experimental validation of key proteins (25–28) have allowed us to estimate the notable archaeal impact on carbon, nitrogen, and sulfur cycles, three major biogeochemical cycles on early and modern Earth (11).
Computational approaches are routinely used to decipher microbial physiology and evolution within MAGs (29–34), with comparative genomics methods remaining the state-of-the-art for homology-driven functional assignments. Well-established homology detection methods entail structure/sequence similarity searches (e.g., Dali, BLAST, HMMER) (35–38), context methods (gene synteny, ontology) (39), and reciprocal best BLAST hits with Markov Clustering algorithms (40) to link sequences to known functions. However, homologous families often contain both orthologs and paralogs, whose functions can be quite diverse (41). Furthermore, evolutionary rates vary between and within protein families such that floating thresholds are required to group homologs or separate them into their respective orthologs. Nevertheless, functional predictions can still rely on orthology detection methods.
Genomic reorganizations and fusion of functionally linked genes, in particular, impose yet another challenge on homology detection (42). The Rosetta stone method (43–45) for the identification of gene fusions states that two or more genes (split genes) encoded in one genome can be found together (composite gene) in the same genome or any other. The process of gene merging corresponds to a fusion event, while the opposite process of gene splitting corresponds to a fission event. This can hinder the grouping of proteins into their natural ortholog families since a fused protein can have higher sequence similarity to one of its split counterparts, with the other marked as absent. Hence, fusions although providing undeniable functional clues, also complicate the sorting of proteins into their natural families.
Using small data sets of (mainly) closed bacterial genomes, previous studies have identified fusion/fission events in prokaryotic databases, revealing that fusion events are more common within the same functional category (42), as a result of modular genome organization common for prokaryotes (46, 47) and that the rate of fusion events is at least four times higher than fissions (42). Also, direct correlations between genome size and number of fusion events were proposed (45, 48). These studies were pivotal to establish the baseline criteria for fusion/fission events identification, their evaluation in prokaryotes and moreover, support the hypothesis of complex protein families’ evolution through fusions of smaller ones (49). More importantly, they show that fusion identification can be a tool to aid in phylogenetic and metabolic reconstructions. As genomic data expands, so do the attempts to integrate fusion predictions systematically, for instance, in the case of STRING (50), SEED (51), and the Integrated Microbial Genomes (IMG) databases (52). So far, these predictive annotations relied exclusively on hidden Markov model (HMM) assignments and on genomic colocalization criteria which can lead to false-positive (FP) identifications. To reduce false positives, in more recent studies (45, 53), a system of rigorous model coverage cutoffs, together with attempts to remove large families of promiscuous domains as fusion partners was implemented.
Herein, we integrated local alignment searches and HMM profiling, alongside genomic colocalization features, to analyze an extensive data set of archaeal genomes. Unlike previous studies that focused solely on archaeal model organisms, our screening includes uncultivated lineages to enhance the understanding of archaeal functional diversity. The identified fusion families were then compared with a large bacterial data set to explore potential evolutionary scenarios. In addition, our results were cross-referenced with existing predictions and experimentally characterized fusions to validate our findings. This comprehensive approach provides valuable insights into the evolution and functional diversity of fusion and fission events in archaeal genomes.
RESULTS
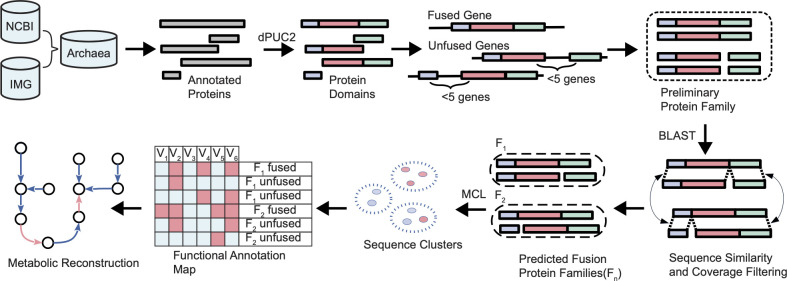
A predictive pipeline to screen for fusion/fission proteins was developed and applied to a data set of 1,678 archaeal genomic assemblies (Table S1). The predictions were based on several criteria (see Materials and Methods). Briefly, the genomic assemblies were run against the PFAM database and the initial candidate protein fusions were defined as the ones having two or more non-overlapping PFAM domains. Based on the Rosetta stone method, the protein fusion candidate (composite protein) would only be retained if, within the genomic data set, the corresponding split protein set (matching each of the single domains of the initial composite candidate) was identified. Given that fusion/fission events tend to occur between functionally related proteins (46, 47), often encoded in proximity to each other, we implemented a genomic colocalization criterion for at least one split protein set per candidate family. Finally, at least 60% of the composite protein had to be covered by the respective split components (Fig. 1).
Fig 1.
Fusion/fission protein screening pipeline.
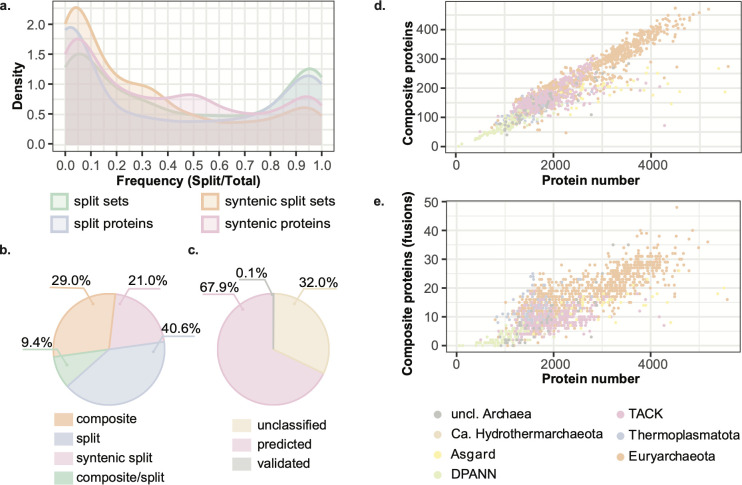
As a result, we obtained 2,183 fusion/fission protein families grouped by domain architecture. The predicted composite proteins were clustered into 1,927 clusters, with five clusters further divided based on the assigned functional annotations (e.g., 14_1). this approach led to a total of 1,932 clusters. As clusters can contain multiple domain architectures, a cluster subindex was assigned for each one of those (e.g., 1.1, Table S2). Of those, 558 only contained a single protein (singletons, Table S2) and were excluded from classification (see below). The frequency of syntenic split protein sets in regard to the total number of composite and syntenic split sets per family shows a left-skewed distribution with a weak sign for bimodality (right orange peak, Fig. 2a). Since in every split set at least two proteins are present (versus only one in the composite side), the frequency distribution considering the total number of syntenic proteins (instead of sets) is shifted to the right by 0.2. (Fig. 2a, in pink). At the same time, the frequency of total split proteins and the corresponding split sets indicates no skew for either of the two modes (Fig. 2a, in blue and green). Despite split proteins accounting for 61.6% of proteins within our data set, the total number of composite proteins almost doubles the number of the exclusively syntenic ones (Fig. 2b). The functional annotation of the proteins revealed that although experimentally characterized proteins comprise only 0.1% of the protein data set, almost 70% of protein families had KEGG orthology (KO) annotations (Fig. 2c; Table S2) enriching metabolic reconstructions. The composite proteins were used to screen the bacterial data set (Table S3) for proteins with identical domain composition and a set of ~12 million composite bacterial proteins was obtained (see Materials and Methods and supplemental table deposit at reference 54.
Fig 2.
Composite (fused) and split (unfused) protein quantification. (a) Frequency distribution (smoothed density) between composite and split proteins, split syntenic/nonsyntenic sets, and composite proteins. (b)) Proportion of composite and split proteins identified. (c) Proportion of validated function (BRENDA or SwissProt), predicted function (significant KO), and unclassified. (d) Correlation between the number of composite proteins and the number of proteins per assembly. (e)) Correlation between the total number of fusion proteins and the number of proteins per assembly.
Caveats
This screening strategy relies on sequence similarity, domain identifications, as well as syntenic analysis. Thus, several artifacts can mask the results and those will be discussed below. It is possible that throughout time, homologous sequences diverged beyond the level of 25% identity, defined as our cut-off. Also, some of the proteins might fail below PFAM model thresholds and thus, have no PFAM assignment. So, some cases might have not been identified by the method. However, our comparison with experimentally validated archaeal fusions and the ones reported in a fusion study in E. coli (45) has shown that in the majority of reported cases (Table S4), the composite and/or split components were identified. Some exceptions are for instance, the case of leader peptidase HopD where the N-terminus is not covered by PFAM models, or in the case of the bifunctional enzyme Fae/Hps (55), where no single Hps domain was found. In addition, the method does not consider homology at the family level and the hits are assigned to the most similar (higher identity) protein. An example would be the case of the fusion of the siroheme domain with the F420 domain as happens in the F420-dependent sulfite reductase, Fsr, found in methanogens (56) which, although homologous to DsrA and DsrB proteins from the Dsr-dependent dissimilatory sulfur metabolism (57), has higher local homology to the AsrC(Dsr-LP group III) and thus was attributed to this family. This closer relationship can also be observed in the phylogenies of the entire family (57). Moreover, we have imposed a synteny filtering, in which, a fusion/fission protein is only kept if, in at least one genome, the split components are within five genes from each other. Although in biology, genomic rearrangements are common, the probability of this having occurred in all of the genomes with the genes of interest is low, considering that genes with functional relationships typically exhibit proximity in a genome, and fusions often happen between functionally associated genes (41). This filter criterium also gives strength to the identified cases, since it reduces false positives in the split component side that could arise from the presence of highly distributed and frequent domains in the genomic data set (e.g., flavin or iron-sulfur cluster domain). As an example, the biotin-dependent pyruvate carboxylase has its subunits fused in some bacteria (PycAB); however, split genes have lost operonic organization in some lineages (58). In archaea, the PycAB fusion is not identified, but instead, in Methanocella, PycA is found fused to biotin ligase (BirA/Bpl), as its gene is often colocalized with pyc genes. Using the approach of adding non-syntenic split sets if the syntenic sets are present, we recovered an additional 296 split proteins, including the Bpl and the PycA of M. jannaschii (59). In addition, Eukaryotes are not included in the analysis. Since the commonly accepted view is that eukaryotes evolved after the diversification of the two prokaryotic domains (60, 61) and, possibly having as host a lineage most closely related with the Asgard group (8, 62, 63), including eukaryotes would allow us to potentially identified events that happened in this domain, and not within Archaea, the focus of our study.
Classification of protein fusion/fission families
The general lack of experimentally validated proteins for composite and split states prevents from using many classification algorithms. Therefore, a simple heuristic approach based on functional and genomic criteria was implemented to classify protein clusters as containing fusions or fissions. The syntenic split set frequency in the family was used for the initial separation between fission and fusion events (less than 0.1 for a fission, more than 0.9 for a fusion) (Fig. 2a). For high confidence fission assignments, besides a split frequency below 0.1, we relied on the presence of the respective composite protein in high-quality archaeal genomes and within bacterial lineages (see methods). KO model coverage (partial versus full) of the split proteins in comparison to the composite one was considered as well. Cases in which the split frequency was above 0.1 but the remaining criteria fulfilled were assigned as “high confidence” fission families. Clusters with a frequency above 0.1, where only some of the criteria were fulfilled, were assigned as probable fission. The presence of complete genomes, with full KO model coverage for the split proteins, and a limited distribution within bacteria were the criteria for a fusion assignment. Based on their presence/absence pattern, in combination with split frequency, the assignments of “high confidence” or probable fusion were given. As in fissions, cases in which, the split frequency was below 0.9 but the remaining criteria fulfilled were assigned as “high confidence” fusion families.
Often, proteins sharing a cluster can be both composite and split at the same time (Fig. 2b). Such overlap can be the result of concurrent fusion and fission processes or a series of fusion events that happened within that homologous family. In this case, if the split frequency is low, and split proteins are in low assembly level genomes (contig or scaffold), the protein cluster would be classified as probable fission. If within a cluster, a composite protein is itself part of a split set for a protein with more complex domain architecture also present in the cluster, then the cluster has the “fusion and fission” assignment. In the second largest “fusion and fission” cluster, besides “canonical” two domain peptide/nickel transport ATP-binding cassette (ABC) proteins and the gene products corresponding to a fusion of two peptide/nickel transport ATP-binding cassette (ABC) subunits from Halobacteria, also split products corresponding to the single ATP-binding subunit are found. Finally, when neither frequency nor any of the other criteria were supported, the cluster would remain unclassified.
As a result, 395 (28,8%) high confidence fissions and 289 (21,0%) fusions were identified. The remaining protein clusters were distributed among probable fission (20,4%) /fusion families (13.8%) or were a combination of both fusion and fission features in the same protein cluster (4.9%) with 11.1% remaining unclassified. The number of fusion/fission composite proteins correlates (r > 0.9) with the number of proteins per assembly, corresponding approximately to 10% of the total number of proteins per assembly. However, the correlation gets weaker in the case of fusions (r ~ 0.8), with fusion-derived composite proteins comprising ~1% of the total number of proteins in a genome (Fig. 2c through e). Regarding the split state, the number of sets highly correlates with the total number of proteins per assembly (Fig. S1). However, the correlation is lost for the high confidence fissions where only 119 assemblies contributed more than five syntenic split protein sets (Fig. S2). This high number of syntenic split proteins could be an indication of assembly fragmentation, particularly in the case of two Thermoproteales genomes. One of them, Vulcanisaeta souniana (IMG_2681813013) (64), has 148 out of 264 protein sets attributed to high confidence fission events, while on average, Thermoproteales assemblies only contain 56 split syntenic sets.
In some cases, the classification was difficult to assess, especially when homologous proteins were present in the same cluster or, when the split and composite state was inconsistent between archaeal and bacterial lineages (see Supplementary Information).
Taxonomic distribution and functional annotation of fusion/fission families
Except DPANN and Ca. Hydrothermoarchaeota lineages, fusions and fission proteins are widely spread across archaeal lineages (Fig. 3; Fig. S2; Table S5). High confidence fission protein clusters contain on average 12 times more proteins than high confidence fusion clusters. This apparent counterintuitive finding can be attributed to the occurrence of fission proteins within families that contain a higher number of composite proteins where only a minority of proteins are split into separate entities. In fact, fissions are more prevalent in protein families associated with housekeeping processes (e.g., informational processes) than with metabolic processes (see below). However, it can be observed that some lineages have a higher prevalence of protein families in which such events (fusion or fission) occurred. This applies to the halobacterial orders, Methanosarcinales and Methanomicrobiales in the case of fusions (Fig. 3) and Methanosarcinales, Thermoproteales, and Thermoplasmatales in the case of fissions (Fig. 3; Table S6). As the number of representatives per lineage is not uniform, we considered the distribution of families in the composite and split state on the genome level (Fig. 3; Fig. S3). For instance, Methanosarcinales and Halobacteria retain elevated numbers of fusion-fission events per genome (~180 families per genome, Fig. S3), but, on average, genomes of Methanophagales and Ca. Poseidoniia are more enriched in high-confidence fusions. The full representation of fusion/fission family diversity per lineage can be achieved with at most 75 assemblies, while for some lineages it takes only ~25 (Fig. S4). Performing an analogy with pangenomic analysis (65), it seems with regard to fusions, a closed “pan-fusion” behavior is observed. Several events of fission and fusion were identified in unclassified Archaea or unclassified Euryarchaeota. However, in MAGs with unclear taxonomic placement, the quality of the assembly can lead to technical artifacts in genome-wide applications.
Fig 3.
Taxonomic distribution of fusions across 1,678 archaeal assemblies. The taxonomic level, represented on the vertical axis, is grouped by order, phyla, or superphyla (indicated in bold). Protein clusters/families are represented on the horizontal axis, with functional category indication underneath. High-confidence fusions are represented on the left side, with probable fusions on the right side. Black indicates the presence of split syntenic sets, green of split sets (where syntenic representatives are absent), red of composite proteins, and white absence of any within the taxonomic rank. The top bar chart shows the percentage of split proteins over the total number of proteins per family. Singletons were excluded from the figure. On the right, the row annotation bar charts show (a) the average number of fusion events per genome (composite protein count); (b) the average number of probable fusion events per genome (composite protein count); and (c) the average number of probable fission events per genome (split protein count).
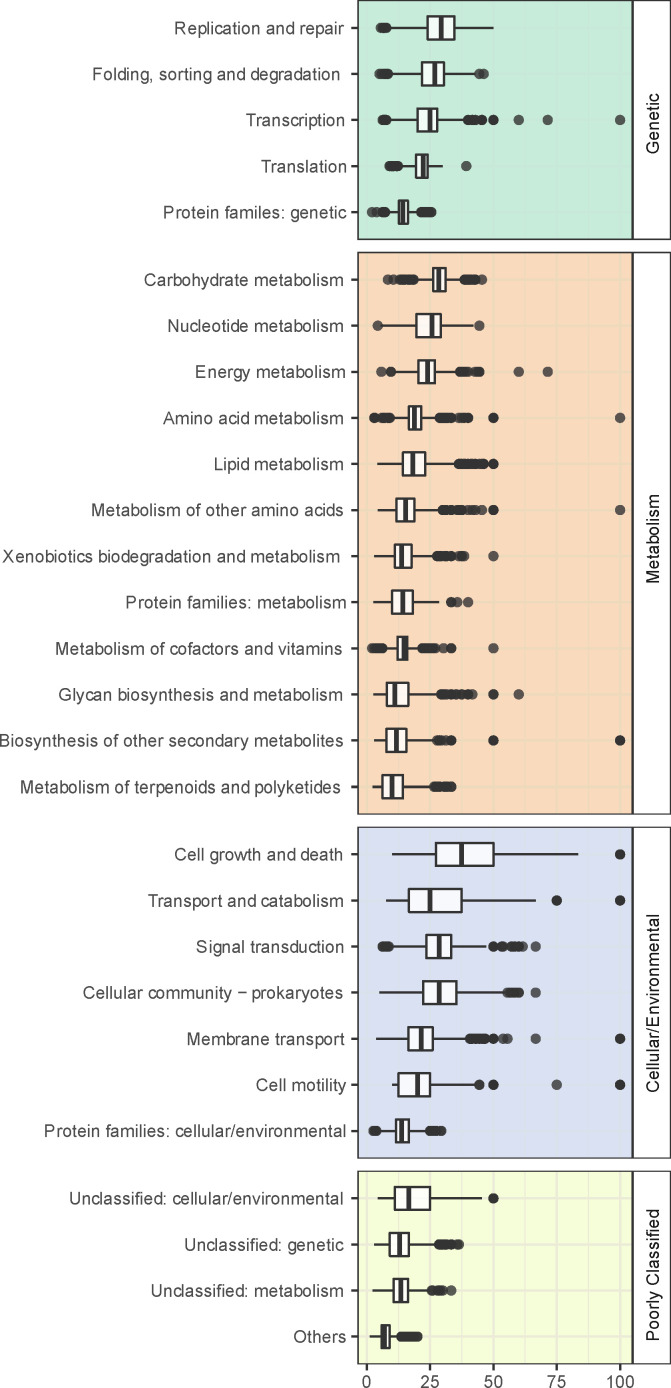
The functional annotations indicated that fissions tend to prevail in proteins related to genomic information processing and carbohydrate metabolism, while fusions are mainly found in quorum sensing, cell motility, chemotaxis, and poorly characterized proteins. (Fig. S2; Fig. 4). When compared to other categories, energy metabolism and amino acid biosynthesis categories are enriched in fission and fusion events (Fig. 4), both represented equally. Notably, a fusion of two or more proteins usually affects only the C- or N-terminus, with possible length reduction, however, does not have an impact on the active or binding sites (Fig. S5).
Fig 4.
Relative abundance of fusion/fission proteins versus the total number of proteins identified per functional category (KEGG categories and KO annotations per assembly were utilized to calculate the ratio).
Functionally, the largest fusion/fission clusters were assigned to ABC transporters, accommodating ~8% of the total number of composite proteins. Among those, 34% are annotated as ABC-2 type, a largely uncharacterized family, universally present in Bacteria. Among the highly populated clusters, three others comprise sequences classified as ABC-transporters, specialized in transporting nickel/peptide, amino acids, and branched sugars, respectively. However, due to their high sequence homology, the functional affiliations between these clusters partially overlap. The lack of experimental validation regarding the specificity of ABC transporters does not allow us to apply the existing ABC-transporter structural classification (66) to the data set.
Apart from transporters, sequences from clusters universally distributed among this data set often belong to the oxidoreductases category. The remaining protein families with large taxonomic representation are affiliated to central carbon metabolism, DNA/RNA processing, or two-component system. Both taxonomical and functional affiliation of the clusters containing mapped split components indicate that these families were mainly retrieved due to scattered cases of fission. A closer inspection of these cases showed common trends regarding the low level of genomic assembly (scaffold or contig) and the lack (or low coverage level) of KO assignments for the unfused components, suggesting their retrieval due to a technical rather than biological fission. This once again indicates that the level of gene fragmentation or frameshifts is not always captured by standard genome quality assessment criteria.
Among archaeal lineages, Methanosarcinales, Methanomicrobiales, Methanobacteriales, and Ca. Poseidoniales contribute to approximately half of the lineage-specific protein clusters (162/347 clusters). Notably, functional categories such as O-Antigen and lipopolysaccharide biosynthesis proteins, glycosyltransferases, and transporters are highly represented in these identified protein families. A considerable fraction (~50% corresponding to 173 clusters) of the clusters consist of poorly characterized enzymes and proteins with unknown function.
When focusing on class-specific fusions, halobacterial proteins stand out, being present in 138 protein clusters (Fig. S6). However, no clear patterns of either fusion or fission or functional module enrichment are observed, apart from poorly classified signaling and cellular processes, to which 16 of these clusters are affiliated. In addition, approximately 50% of the clusters have no defined function according to the KEGG database. Within methanogens, 200 unique protein clusters containing fusion/fission proteins were identified, some of which are associated with energy metabolism, in particular methane metabolism.
The mapping of the identified archaeal composite proteins to bacterial assemblies revealed that although many of the protein families have homologs widely distributed across bacterial groups (Table S7), for 162 cases, no bacterial composite homologs were identified. If we considered a “minimum of two assemblies per lineage” rule, this number would increase to 228 non-singleton clusters (Fig. S7). These clusters correspond to unique archaeal composite proteins. These unique archaeal fusion/fission families belong to the genetic information processing (eight clusters) and metabolism (19 clusters) categories (Table S2). Within the metabolism category, archaeal-specific fusion clusters were found to have occurred among genes from folate, methane, and amino acid metabolism.
Fusion/fissions events in archaeal metabolism
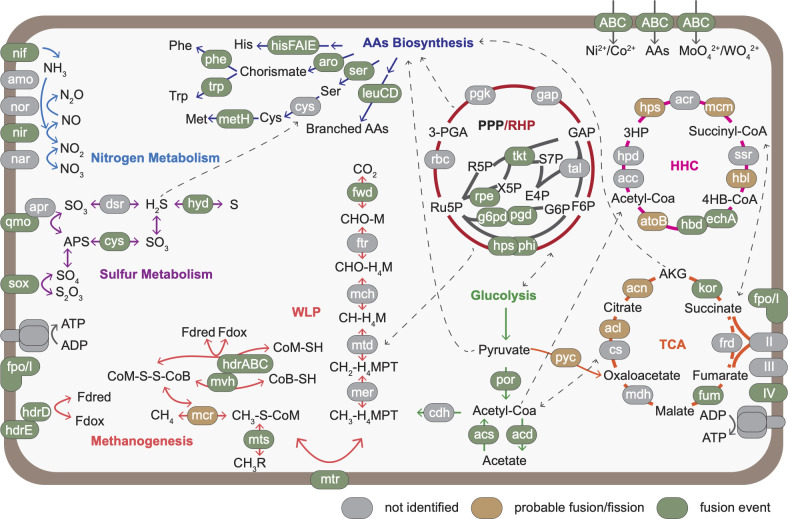
The impact of fusion and fission events in archaeal metabolism is represented in Fig. 5 and will be discussed in detail below.
Fig 5.
Fusion events in archaeal metabolism. The listed abbreviations do not occur in the text. Protein abbreviations. HHC: Acc, acetyl-CoA carboxylase; Hhps, hydroxypropionyl-coenzyme A synthetase; Acr, acryloyl-CoA reductase; Ssr, succinate semialdehyde reductase; Hbl, 4-hydroxybutyrate-CoA ligase; AtoB, acetyl-CoA acetyltransferase; Hpd, hydroxypropionyl-CoA dehydratase. PPP/RHP: Pgk, phosphoglycerate kinase; Gap, glyceraldehyde-3-phosphate dehydrogenase; Tal, transaldolase. TCA: Cs, citrate synthase; Frd, fumarate reductase. Respiratory chain: Nuo, NADH-quinone oxidoreductase; Qcr, quinol-cytochrome c reductase; Cyo, cytochrome-c oxidase. WLP: Fwd: formylmethanofuran dehydrogenase; Ftr, formylmethanofuran–tetrahydromethanopterin N-formyltransferase; mch, methenyltetrahydromethanopterin cyclohydrolase; Mtd, methylenetetrahydromethanopterin dehydrogenase; Mer, 5,10-methylenetetrahydromethanopterin reductase. Methanogenesis: Mcr, methyl-coenzyme M reductase; Mvh, F420-non-reducing hydrogenase; Fpo, F420H2: phenazine/quinone oxidoreductase. Sulfur metabolism: Sox, sulfur-oxidation system; Qmo, quinone oxidoreductase; Apr, adenosine 5′-phosphosulfate reductase; Dsr, dissimilatory sulfite reductase; Cys, sulfate assimilation enzymes. Nitrogen metabolism: Nar, nitrate reductase; Nir, nitrite reductase; Nor, nitric oxide reductase; Amo, ammonia monooxygenase; Nif, nitrogenase. AAs biosynthesis proteins: HisFAIE, histidine; Aro, chorismate; Cys, cysteine; Phe, phenylalanine; Trp, tryptophane; MetH, methyltetrahydrofolate-homocysteine methyltransferase; LeuCD, leucine; Ser, serine. Others: Por, pyruvate-ferredoxin/flavodoxin oxidoreductase; Cdh, carbon-monoxide dehydrogenase; Acs, acetyl-CoA synthetase; Pyc: pyruvate carboxylase; Compound abbreviations. HHC: 3HP, 3-hydroxypropionate; 4HB, 4-hydroxybutyrate. PPP/RHP: Ru5P, ribulose 5-phosphate; 3-PGA, 3-Phospho-D-glycerate; GAP, glyceraldehyde 3-phosphate; F6P, fructose-6-phosphate; R5P, ribose-5-phosphate; X5P, D-xylulose 5-phosphate; S7P, sedoheptulose 7-phosphate; E4P, erythrose 4-phosphate; G6P, glucose-6-phosphate. TCA: AGK, alpha-ketoglutarate; WLP: MF, methanofuran; H4MPT, tetrahydromethanopterin; Fd, ferredoxin; Sulfur metabolism: AAs biosynthesis: His, histidine; Ser, serine; Cys, cysteine; Phe, phenylalanine; Trp, tryptophan; Met, methionine.
Within carbohydrate metabolism, both fission and fusion proteins were identified (14 fusions, 47 fissions, and 18 probable fusions/fissions). Notably, while in glycolysis 9 fissions were identified, carbon fixation and methane metabolism (assigned to Energy metabolism) show a more balanced distribution of both fusion and fission events (23 fusions and 13 fissions, in total, with “fusion and fission” families counting for both states) (Fig. 5). Within the tricarboxylic acid cycle (TCA), several fusion events were identified in both reductive and oxidative versions. For instance, a fusion was observed in the dihydrolipoamide dehydrogenase (LpdA, cluster 14_1), part of the aerobic pyruvate dehydrogenase complex, responsible for converting pyruvate to acetyl-CoA (67). In halophilic archaea, some LpdA proteins are fused to a biotin-dependent domain (cluster 14_1.2). On the other hand, anaerobic archaea use the alternative enzyme pyruvate oxidoreductase (Por) for the conversion of pyruvate to acetyl-CoA (68). A fusion between the alpha (PorA) and beta subunit (PorB) was identified in nine genomes, mainly in Ca. Bathyarchaeota and Ca. Korarchaeota (cluster 45.2). The enzyme aconitase A (AcnA) (69), responsible for converting citrate to isocitrate, can be the result of a fusion between two subunits from a methanogenic homocitrate aconitase (70) or subunits from the widely spread isopropylmalate dehydratase involved in leucine biosynthesis (70) (cluster 139). Two fusion events, involving 2-oxoglutarate oxidoreductase Kor subunits, a complex responsible for converting oxoglutarate into succinyl (68), were identified: one between korA and korC (cluster 75.1) genes, and another between korB and korC genes (cluster 368). Fusions tend to occur between closely related genes, and for this enzyme, there is a high conservation of synteny across the archaeal phyla. Moreover, the split components of this family include the experimentally validated KorB and KorC proteins from M. marburgensis (68), corroborating our prediction. In the case of fumarate hydratase (Fum) class 1, the subunits were identified as fused in most bacteria but in the split state in the majority of Archaea (71). Exceptions include two Desulfurococcales and two ANME assemblies, in which the fused version was identified (cluster 926). The subunits alpha (SucC) and beta (SucD) from succinyl-CoA synthetase, an enzyme that converts succinyl to succinate (72), were found in the split syntenic configuration in four and two assemblies, respectively (cluster 120). The sucD gene is also involved in a probable fusion event with a citrate synthase protein, forming the composite protein ATP citrate (pro-S)-lyase (Acl) (73, 74) which is found in some Methanosarcinales species (cluster 120). This enzyme is a part of the reductive TCA, converting citrate to acetyl-CoA, and was previously described in the split state in Aquificae (74) (cluster 547). Fissions were also identified in the flavoprotein subunit of succinate dehydrogenase (SdhA) (six assemblies, cluster 127_1) and between the two domains of malate dehydrogenase (eight split proteins, four assemblies, cluster 142). While in the first case, our analysis favors these to be the result of assembly artifacts (technical fission), the second represents a candidate for further experimental investigations.
The oxidative phase of the pentose phosphate pathway (PPP) is not widespread among archaea, and within this pathway, a single fusion event occurring between glucose-6-phosphate 1-dehydrogenase (G6pd) and 6-phosphogluconate dehydrogenase (Pgd) enzymes was identified in Diapherotrites (DPANN phylum, cluster 1367, singleton). Interestingly, the fused form is also present in 60 bacterial candidate phyla assemblies. By contrast, within the biosynthesis of ribose from fructose (the dominant route in Archaea), the pipeline identified the experimentally validated bifunctional enzyme (Hps-Phi) (55, 75), which is the result of a fusion between 3-hexulose-6-phosphate synthase (Hps) and 6-phospho-3-hexuloisomerase (Phi) (cluster 220, Fig. S5). While in bacteria, Hps and Phi were identified as two separate proteins, the distribution pattern in archaea suggests that it is likely a result of an archaeal fusion event (76). Fusions were also identified for proteins from the non-oxidative branch of the pentose phosphate pathway (nPPP). Among them was the three-domain bacterial transketolase (Tkt) found as a split protein pair in 441 archaeal assemblies (cluster 341.2). In a few representatives from the DPANN superphylum and Ca. Thorarchaeota, the protein was identified in the fused state. Notably, in Thaumarchaeota (73 out of 98 assemblies), the N-terminus of Tkt is fused to a ribulose-phosphate 3-epimerase (Rpe) enzyme (cluster 341.1). This fusion event suggests the potential presence of a complete functional nPPP in Thaumarchaeota, differentiating it from the typical pathway configuration found in most other archaeal lineages, as previously suggested (77–79). The identification of composed and split proteins suggests that the non-oxidative branch of the pentose phosphate pathway is more widely distributed in archaea (77–79). This indicates a greater prevalence and functional significance of nPPP in various archaeal lineages, contributing to a more comprehensive understanding of carbohydrate metabolism in these organisms.
Regarding the reductive hexulose-phosphate (RHP) pathway, several fissions of multidomain proteins have been observed, including the large chain of ribulose-1,5-bisphosphate carboxylase (RuBisCo/Rbc, cluster 236). However, no fusions were detected in this pathway. On the contrary, both fusion and fission events were observed in the 3-hydroxypropionate/4-hydroxybutyrate (3HP/4HB) carbon fixation cycle, a widely utilized pathway by diverse, mostly non-methanogenic archaeal lineages (80, 81). The pathway involves a series of carboxylation and reduction events and reuses the same domain architectures in different enzymes. Multiple enzymes of the 3HP/4HB cycle were found to be part of the same fusion/fission cluster, which also contain homologs involved in carbohydrate and fatty acid metabolism. In ammonia-oxidizing archaea (AOA), fusion events promoted the divergence and functional adaptation of the homologous enzymes, 3-hydroxypropionyl-CoA synthetase (Hps) and 4-hydroxybutyrate-CoA ligase (Hbl) (81). While the Hps enzyme, besides the shared domain with Hbl, contains an ATP-grasp domain fused to its N-terminus, the Hbl enzyme has the ATP-grasp domain at its C-terminus. Both Hps and Hbl enzymes cluster with acetate-CoA ligase (AcdAB) enzymes, which catalyze a similar CoA-dependent reaction (81, 82) (cluster 20). The corresponding split versions of the three enzymes were found in several extremophilic lineages, including Desulfurococcales (17/30) and Thermoproteales (24/35). In Sulfolobales, this enzyme is absent, since the organisms use alternative enzymes for carbon fixation (83). Phylogenetic reconstructions of this extended family could give insights and aid in the clarification of the precise order of these events. An additional fusion between 3-hydroxybutyryl-CoA dehydrogenase (Hbd) and crotonyl(enoyl)-CoA hydratase (EchA), two enzymes involved in consecutive steps of the pathway (84) was also identified (cluster 65, Fig. S5). The EchA-Hbd fusion is widely distributed in the TACK superphyla (186/425), the majority of halophiles (374/412), and in Archaeoglobales (13). On the other hand, split versions of these enzymes occur in Sulfolobales (~97%), ~50% of Halobacteria, and 85% of the AOA assemblies analyzed here. In the unfused Hbd proteins from AOA, a duplication of the C-terminus domain is conserved in all of the fused components identified (cluster 65). In the halobacterial version of Hbd, however, this duplication is absent (cluster 65.3). In addition, a fusion between acetyl-CoA C-acetyltransferase, an enzyme responsible for acetyl-CoA regeneration (81, 84), and an uncharacterized conserved protein was identified in two Thermoplasmatales genomes (cluster 17.2). The corresponding split pairs tend to be syntenic in 12 phyla, predominantly from TACK and Asgard supergroups.
In the context of methane metabolism, protein families display a balanced rate of fused and unfused components (Fig. 5). Many reactions of the pathway rely on multi-subunit complexes (85), and we observe multiple fusion combinations per protein complex. The initial step of methanogenesis, where CO2 reduction is initiated, is catalyzed by formylmethanofuran dehydrogenase (Fwd/Fmd) (86, 87) which contains between 6 and 8 subunits. In 16 Methanomicrobiales assemblies and 14 TACK assemblies, fusions of subunits B and D were identified (cluster 525). In addition, in 3 assemblies of Methanosaeta, a fusion between Fmd subunit E and a transcriptional regulator is present (cluster 663.2–3). In some Methanosarcinales (4) and Archaeoglobales (3), FmdE is fused with a tRNA methyltransferase domain (cluster 663.1). The identified fusions between the only fmdE gene copy and a tRNA methyltransferases domain might indicate the peripherical role of this subunit within the complex, pointing to a possible regulatory function. In fact, in the recently obtained structure of the complex (87), the FmdE subunit is not observed, which can further corroborate this idea. Other events can be found in Table S2.
Three fusion events were identified between the subunits of tetrahydromethanopterin S-methyltransferase (Mtr), a key enzyme involved in methyl-coenzyme M formation (88). These include a singular fusion of subunit A to B in unclassified Thaumarchaeota (cluster 797.1), between subunit A and H in unclassified Archaea and between subunit A and F in 12 Methanococcales and 2 Methanocellales assemblies (cluster 797.2).
The Methyl-coenzyme M reductase (Mcr) enzyme is responsible for catalyzing the reduction of methyl group to methane through a reaction between coenzyme M and B (89). In the assembly of Methanoculleus marisnigri, its subunit A is split into two proteins (cluster 306.1). This finding is particularly important since the subunit is widely used for phylogenetic reconstructions (90), and proper recognition of fission events is crucial for accurate analyses.
Finally, within the heterodisulfide reductase complexes (HdrABC and the HdrED present in cytochrome containing methanogens) that catalyze the reduction of coenzyme B (CoB) and M (CoM) (91, 92), several fusions between their subunits were identified (Table S2; Fig. S5). These are widely distributed across taxa, can involve more than two subunits, and may include fusion of duplications, as seen in the alpha subunit (HdrA). For instance, Ca. Thorarchaeota (16/18 assemblies) and Ca. Lokiachaeota (9/10 assemblies) have multiple copies of the hdrA gene per assembly (cluster 77). In all of these lineages, at least one of these HdrA proteins is the short version of the enzyme, having on an average of 650 amino acids. In addition, proteins resulting from a fusion of two HdrA domains are further fused with an F420-non-reducing hydrogenase iron-sulfur subunit D (mvhD) that is often attached to its C terminus (17/54 duplicated HdrA fusions) (cluster 77). These longer versions of HdrA proteins were identified to be in up to six copies per assembly. Except Thaumarchaeota where only in two assemblies four HdrA fusions were detected, a high number of fused HdrA domains are found within the TACK phyla. Methanogens, however, have only one or two HdrA copies in the cluster, most of them corresponding to the “canonical” shorter form or instead, fused with the mvhD domain.
Fusions are also observed in the enzymes responsible for the biosynthesis of cofactors related to methanogenesis. For instance, fusions were identified between sulfopyruvate decarboxylase subunits (ComE and ComD), an enzyme involved in the biosynthesis of coenzyme M (93) (cluster 348). This fusion was identified in 175 assemblies from 16 taxa, excluding uncharacterized Archaea and uncharacterized Euryarchaeota. These findings highlight the diversity and evolutionary adaptations in carbon fixation pathways and its associated cofactor biosynthesis enzymes among archaeal lineages, shedding light on the complex metabolic strategies employed by different archaea in response to their environmental niches, in particular in terms of carbon metabolism.
Nitrogen and sulfur metabolism
Among archaeal fusion/fission protein families, there are notable representatives involved in nitrogen metabolism (seven clusters) and sulfur metabolism (11 clusters). However, it is worth mentioning that these fusion events predominantly occur within assimilatory routes, being scarce in dissimilatory ones. Archaeal fusion/fission protein families are spread across diverse aspects of the nitrogen cycle, primarily in assimilatory nitrogen reduction, denitrification, and nitrogen fixation (11). However, it is important to note that the majority of these protein clusters are the result of fission events. One way to assimilate nitrogen is the uptake of nitrate and its conversion to ammonia via nitrite (11). The subsequent nitrite reduction step is catalyzed by the ferredoxin-nitrite reductase (NirA) (94) which is homologous to the sulfite reductase (Sir) (94), involved in sulfur metabolism. NirA is a four-domain protein, arranged in an A-B-A-B manner. NirA is widely present in Halobacteria where 19 out of the 529 identified proteins are fused to a rhodanase domain (cluster 195.2). Rhodaneses are sulfur carriers involved in sulfur metabolism (95), and these halobacterial composite proteins might perform a role in sulfur metabolism. The four-domain NirA itself might be the result of a fusion between the two domain proteins (A-B), identified in Methanosarcinales. Sir and NirA (94) are found in the same cluster. Both could have emerged from the fusion of already duplicated proteins before their functional specialization.
A fusion event involving a NO-forming, copper-containing nitrite reductase nirK (96, 97), and a plastocyanin, is present in four Halobacteria (cluster 505). In the majority of Halobacteria, however, the NirK and plastocyanin proteins are found in the split version, conserving synteny. A similar copper-containing protein architecture with a plastocyanin extension is observed across Thaumarchaeota (18), including AOA, where NirK is hypothesized to be involved in N2O production (98) (cluster 505). In addition, one high-confidence fusion between NifH and NifD/E proteins in Methanomassiliicoccales was identified, while in Methanosarcinales and Methanomicrobiales both composite and split forms are found (cluster 443).
In sulfur metabolism, only a few fusion/fission events were identified, with fission events prevailing, especially in the assimilatory reduction pathway. Among the proteins involved in the assimilatory pathway, three distinct protein clusters were identified. The first corresponds to sulfate adenylyltransferase (Sat), which catalyzes the initial activation step of sulfate to adenosine 5′-phosphosulfate(APS) (99). The Sat cluster was retrieved due to a single fission event in the Vulcanisaeta sounina IMG assembly (cluster 407.1). A single case of Sat protein fusion with histidine phosphatase was also registered in a Sulfolobus genome (cluster 407.2). The second cluster corresponds to phosphoadenosine 5′-phosphosulfate reductase (cysH), which is responsible for the third step in the pathway (100) (cluster 536). This cluster is exclusively composed of Methanomicrobiales proteins. Interestingly, the domain architectures suggest a fusion of CysH with a cysteine desulfurase (SufS), an enzyme involved in amino acid biosynthesis (101, 102) (cluster 536). However, only in four assemblies, the unfused gene sets were observed, hindering a clear fusion/fission classification. In addition, a fusion between cysH and the threonine synthase (thrC) gene is observed in three Thermoplasmata assemblies (cluster 1057).
In contrast to the assimilatory reduction pathway, no fusion or fission events were identified in enzymes responsible for the conversion of sulfite to sulfide in the dissimilatory sulfate/sulfite reductive pathway. Only in the case of V. souniana, the adenylylsulfate reductase(Apr) that catalyzes the reduction of APS to sulfite (103) was observed as a split set. This was previously reported and currently, it is not clear if the Dsr-cascade is operational or not in this organism (23). Of note, a fusion of the siroheme domain also present in DsrAB and AsrC proteins with the F420 domain as seen in the case of the F420-dependent sulfite reductase, Fsr, was observed. This protein is present in methanogens and has a higher identity to the anaerobic sulfide reductase (AsrC) rather than with DsrAB proteins. AsrC, as well as Fsr, has an additional iron-sulfur cluster (57, 104, 105), and it is proposed that Fsr originated from a fusion between a siroheme-domain containing AsrC ancestral domain and the FrhB domain (56).
Lastly, a fusion between the sulfur hydrogenase beta and gamma subunits, which are involved in sulfur/sulfide reduction, was observed. This composite protein has so far only identified in 11 archaeal assemblies, mainly in Thermoplasmatales, Ca. Bathyarchaeota, as well as several bacteria.
Complexes from mitochondrial-like respiratory chains
The mitochondrial electron transport chain is characterized by the presence of four complexes (I to IV) and an ATP synthase. Each one of these complexes has homologues within the prokaryotic domain, namely in Archaea. Complex I is part of group 4 of membrane-bound [NiFe] hydrogenases and within Archaea, due to the inexistence of the electron input module NuoEFG, denominated as Fpo (F420H2:phenazine oxidoreductase) (106), or Fqo, F420H2 quinone oxidoreductase (107). A fusion between the subunits B and C of this complex was identified in 47 assemblies with diverse taxonomic affiliations (cluster 482), with 10 of these cases being further fused to subunit D (cluster 482). Importantly, in the initial group (fqoBC), the protein from Archaeoglobus fulgidus is present, as previously suggested to be a fusion of those subunits (107). Regarding the terminal heme-copper oxygen reductase (Complex IV), the fusion between subunits I and III was identified (cluster 251), in which, the experimentally characterized S. acidocaldarius protein is present. The possibly identified fission event regarding complex II or succinate dehydrogenase was already discussed above (TCA cycle).
Amino acid metabolism
The majority of fusion events related to amino acid metabolism are present in the biosynthesis of histidine and aromatic amino acids pathways. The initial steps of histidine biosynthesis are catalyzed by the bifunctional fusion protein phosphoribosyl-AMP cyclohydrolase/phosphoribosyl-ATP pyrophosphohydrolase (HisIE) (108). While initially thought to be widely distributed only in Thermococcales and Thermoplasmatales (109), this fused form has now been found in some novel methanogenic lineages, including Methanomassilococcales, Methanofastidiosa, and Methanomethyliales (cluster 384). On the other hand, split syntenic pairs of HisIE are present in Thermoproteales, Ca. Bathyarchaeaota, Ca. Hadesarchaea, and Desulforococcales (cluster 384). The emerging distribution of HisIE suggests that the fusion might have originated within the archaeal clade itself, providing a possible alternative to the previously suggested horizontal gene transfer from bacterial partners (109). Another intriguing fusion event involves two subunits of imidazole glycerol phosphate synthase (HisH and HisF), found fused mostly in eukaryotic species (110). The fused HisHF is present in 2 Archaeoglobales species and has approximately 70% identity to HisHF from sulfate-reducing bacteria (cluster 787.3). However, the identified HisF may also function as a phosphoribosylformimino-5-aminoimidazole carboxamide ribotide isomerase (HisA) since they are homologous and have evolved from an ancient duplication (109). Moreover, HisA/F fusions with HisIE (phosphoribosyl-AMP cyclohydrolase/phosphoribosyl-ATP pyrophosphohydrolase) have been identified in four metagenomic assemblies, scarcely distributed across DPANN and unclassified Archaea (cluster 787.1). Interestingly, the bifunctional imidazoleglycerol-phosphate dehydratase/histidinol-phosphatase (HisB) (111) and the fusion of HisIE with histidinol dehydrogenase (HisD) (112), present in Bacteria and Eukarya (109), were not identified in the Archaea domain.
In the biosynthesis of chorismate, a key precursor of aromatic amino acids, several composite proteins have been identified (Fig. 5). A fusion between 3-dehydroquinate dehydratase (aroD) and shikimate dehydrogenase (aroE) genes, first discovered in plants (113), was also identified in some archaea, including 8 Methanofastidiosa and 10 Methanomassiliicoccales assemblies, as well as in two bacterial phyla (198 assemblies of Planctomycetes and 88 assemblies of Acidobacteria) (cluster 359.1). A fusion involving aroE and the shikimate kinase aroK (114) genes has been identified in Methanomicrobiales (42/55), one unclassified Methanomicrobia, and two bacterial assemblies (cluster 319).
In the tryptophan biosynthesis pathway, the composite protein anthranilate synthase/phosphoribosyltransferase (TrpGD) catalyzes the initial steps of the pathways in a range of bacteria (115) (cluster 373). Interestingly, trpGD was found fused to a nitric oxide reductase gene (nor) in one Methanocella genome, indicating a unique evolutionary event in this archaeal lineage or a result from interdomain LGT (cluster 373.2). A fusion between trfD and trfC genes was also identified in some organisms (cluster 940). In addition, a fusion between phosphoribosylanthranilate isomerase (trpF) and tryptophan synthase beta chain (trpB) (116), although rare, has been identified in both bacterial and archaeal domains (cluster 830.1, single fusion protein trpAB in 830.2). Finally, in two assemblies, a fusion between trpF and trpC was identified (cluster 1136).
In the biosynthesis of phenylalanine, a fusion between chorismate mutase and prephenate dehydratase (PheA) (117) is common among bacteria (present in 9,768 bacterial assemblies). However, within archaea, this fusion only occurs in a few representatives of the DPANN and TACK superphyla (cluster 245.2). Moreover, in Archaeoglobus and Ca. Huberarchaea, a second step fusion has been identified, where chorismate mutase/prephenate dehydratase is further fused to a prephenate dehydrogenase (118), an enzyme involved in the first step of tyrosine biosynthesis (cluster 693). This multi-fusion event appears to be unique to these two archaeal lineages.
Cofactor biosynthesis
Cofactor biosynthesis plays a vital role in cellular metabolism, as these essential molecules can be recycled and used multiple times by enzymes with divergent functions and evolutionary histories. Thus, understanding the evolution of cofactor biosynthesis enzymes can provide valuable insights into the early evolution of life. Through comprehensive screening, numerous fusion events have been identified in various biosynthesis pathways, including those for thiamin, biotin, riboflavin, tetrahydrofolate, molybdopterin, and heme biosynthesis. Below, we give the details of the latter two pathways.
Molybdopterin biosynthesis is a crucial pathway involved in the synthesis of a cofactor that binds to a broad range of oxidoreductases (119). In this biosynthetic process, several fusion and fission events have been identified for its enzymes. Notable fusion events involve the cyclic pyranopterin monophosphate synthase MoaC (120), which has been independently fused on multiple occasions with itself (cluster 625.1) as well as to enzymes of two downstream biosynthesis reactions, namely MoaE and MoaB (120) (cluster 625.2 and 765, respectively). These fusions are not limited to specific taxonomic groups and could be a result of frameshifts in the conserved syntenic blocks. Interestingly, no fusion between MoaC and GTP 3′,8-cyclase (MoaA) (121), an enzyme that initializes the pathway, was identified. On the other hand, fusions involving MoaD, a sulfur-carrier protein (120, 121), are lineage specific. For example, the MoaD-MoaE fusion (subunits of molybdopterin synthase) has been identified in (18 out of 35) Thermoproteales and (5 out of 30) Desulforococcales, as well as in a small fraction of bacterial genomes (775/33957) (cluster 419.1). However, the protein was proposed to be non-functional in its fused form, as the C-terminus of MoaD needs to remain open to bind the sulfur group, requiring post-translation cleavage (122). The fusion between MoaD and the molybdopterin-synthase adenylyltransferase (MoeB), the protein responsible for the MoaD adenylation (123), is present in Thaumarchaeota (61/98) and unclassified Crenarchaeota (4/246) and two other assemblies (cluster 356.1). In bacteria, this fusion is limited to Ca. Rokubacteria and Ca. division NC10, both of which are hypothesized to be nitrite reducers (124). Moreover, the moaD fusions with moeBR gene, where moeB is fused with a rhodonase domain (cluster 356.3, 14 assemblies) and moeBR (cluster 356.2, 25 assemblies) were identified mainly in unclassified Euryarchaeota and Ca. Poseidoniales. This suggests the possibility of a gene transfer event from Acidobacteria to Archaea, given the relatively high identity (over 55%) between the proteins from these two groups. The last enzyme in the molybdopterin biosynthesis pathway, MoeA, is responsible for catalyzing the insertion of molybdenum into molybdopterin (125). The protein appears to have fissions(cluster 27.1) as well as periplasmic domain extension (cluster 27.2), classified as fission with both cluster subgroups lacking split components. The taxonomic distribution of the fused and split enzymes is not uniform in Archaea, indicating a complex evolutionary history for molybdopterin biosynthesis across different taxa.
Heme is a crucial iron-containing porphyrin cofactor involved in electron transfer, essential for both aerobic and anaerobic respiration. Prokaryotes utilize three distinct pathways for heme biosynthesis, converging at uroporphyrinogen-3 as their last common intermediate (126). Fission and fusion events are common occurrences in the enzymes catalyzing early reactions (HemABLCD) of the pathways from glutamate to uroporphyrinogen-3. Our analysis revealed the existence of several fusion proteins in this pathway. One such fusion event was found between glutamyl-tRNA reductase (HemA) and porphobilinogen synthase (HemB), in a single Sulfolobus assembly. In addition, two fusion events were identified between hydroxymethylbilane synthase (HemC) and the composite protein uroporphyrinogen III methyltransferase/synthase (CobA-HemD) in Euryarchaeota (cluster 192.3). CobA-HemD (127) is a fusion protein in itself, where CobA is responsible for catalyzing precorrin-2 biosynthesis, representing the first divergent reaction in the alternative heme biosynthesis pathway. This fusion protein was initially characterized in sulfur-dependent bacteria (127) and based on our screening results, it is also present in Actinobacteria (3448/5525), Firmicutes (1974/5564), and around 100 ammonia-oxidizing bacteria (cluster 192). However, although the synteny of the split proteins was universally conserved across other archaeal lineages (cluster 605.1) within Archaea, the fused CobA-HemD proteins were mainly identified in the genus Methanothrix and Ca. Methanofastidiosa.
Fission and fusion events were also identified in the alternative heme biosynthesis pathway, where siroheme serves as an intermediate (126). In 14 lineages of TACK and the majority of Euryarchaeota, siroheme decarboxylase genes ahbA and ahbB were found fused, while they remained split in the majority of Methanosarcinales, as previously reported (128), and also in Methanomicrobiales and Methanonatronarchaeales genomes (cluster 230). Interestingly, a fusion between ahbAB and ahbD (AdoMet-dependent heme synthase) genes was identified in five genomic assemblies affiliated with Methanomicrobia (cluster 36.2). These fusion and fission events in the heme biosynthesis pathway illustrate the dynamic nature of enzyme evolution and how organisms can utilize different strategies to adapt and regulate their cellular processes, particularly in the context of synthesizing essential cofactors like heme.
DISCUSSION
The identification of fusions and fissions in prokaryotic genomes has raised several technical and biological questions. We observed that the number of identified fissions increases with the growing number of incomplete genomes, leading to challenges in elucidating biological meaning from the noise in the data. Thus, we speculate that many of our identified fissions and fusions might be due to the incorrect assembly of public metagenomic records. Here, we chose to include records with heterogenous data quality to explore their impact in this and other analyses.
The distribution of bacterial and archaeal protein families often shows polar patterns, which can be attributed to various factors like the absence of certain proteins, interdomain horizontal gene transfers, poor alignment coverage of split pairs from distant lineages, or misidentification of protein domains. Efforts have been made to address some of these issues through clan mapping, but many protein domain models need to be updated with diversified archaeal sequences. Even the currently available archaeal clusters of orthologue proteins are based on a reduced number of archaeal lineages and sometimes contain multiple paralogs within a group (129).
This screening also revealed a lack of functional characterization in the archaeal domain. In some cases, poorly characterized proteins are fused with experimentally validated ones, providing insights into their potential functions. However, in many instances, both split proteins have unknown functions, making functional deductions challenging. To fill these knowledge gaps, more experimental characterizations, especially in what concerns protein from the archaeal domain need to be performed.
The set of candidate fusion families was constructed by combining protein domain and full sequence similarity between composite and split proteins. Additional classification was applied to distinguish between fusion and fission events, with fissions typically having a wider taxonomic distribution and being involved in genetic information processing, while fusions were more commonly associated with taxonomically restricted metabolic processes. These results highlighted unique archaeal fusions and revealed underestimated the capacity for energy metabolism in certain lineages (carbon fixation, e.g., cluster 65), as well as the importance of fusion processes in halobacterial signaling and cellular processes (two-component system/chemotaxis proteins in multiple clusters; NO signaling e.g., cluster 835; flagellins, e.g., cluster 575, cluster 89).
In conclusion, the identification of fusion and fission events in prokaryotic genomes has shed light on various aspects of microbial physiology and evolution. The study has underlined the need for integrating new steps into genome assembly quality assessment pipelines to improve the accuracy of fusion identification. Overall, the findings contribute to a deeper understanding of the functional diversity and metabolic capabilities of prokaryotic lineages, particularly Archaea, and highlight the importance of experimental validation to fill gaps in our knowledge.
MATERIALS AND METHODS
Data acquisition
A total of 2,693 archaeal assembly records were obtained from NCBI Archaea (Jan. 2020). In addition, 1,000 publicly available genomes of Archaea were retrieved from IMG (January 2020). Genome completeness and redundancy (contamination) levels were estimated on *faa files containing the annotated protein sequences using the set of archaeal marker protein HMMs (130). Both estimates were used as criteria for the data set filtering. The genome redundancy cutoff was set to 10%, while the completeness cutoff was optimized according to the taxonomic affiliation, to account for organisms known to have reduced genomes. In cases where strains were represented by multiple assemblies, the assembly level (complete genome, scaffold, etc.), completeness, and redundancy were used to select a single reference assembly. To reduce the total number of genomes, bacterial genomes were further filtered for cases of overrepresented species. In total, 1,678 archaeal and 33,957 bacterial genomes, downloaded from NCBI in January 2020 and November 2019, respectively, were used in this analysis.
Fusion/fission family identification
Proteins from the archaeal genome data set were searched for protein domains (pfams) against Pfam32.0 (131) using HMMER3 (132). The HMMER3 output was processed with dPUC2 (133) to produce sets of protein domains with the highest pairwise directional probabilities. Next, the archaeal protein domain architectures were matched against each other to construct the preliminary set of fission/fusion protein families. When possible, and to expand the fusion protein search space, pfams were mapped to their respective clans, which contain two or more homologous families (134). For the assignments, the multidomain architecture of a single protein (composite protein) had to match the sum of the domains of the split protein pairs/sets, and, in the case of multidomain split components, also the order of domains had to be retained. Finally, among the genes encoding candidate unfused proteins at least one set should be genomically colocalized, in this case, a maximum of five genes apart (Fig. 1). The last condition has been derived from the operonic organization of prokaryotic genomes making the fusion between colocalized genes more probable. Candidate fusion proteins with identical domain architectures were grouped into preliminary fusion families, together with the corresponding subsets of unfused components.
Similarity searches
The preliminary families were aligned with each other using DIAMOND blastp (38). To remain as the part of fusion family, candidate proteins had to fulfill several criteria. First, pairwise alignment between composite and split protein should have a sequence identity of more than 25% and an E-value less than 1e−9. The overlapping coverage between the candidate fusion protein and the potential split candidate should cover at least 60% of the latter. Neither of the proteins in the set should overlap with each other by more than 100 amino acids in the alignment to the composite subject. Moreover, at least 60% of a candidate protein sequence had to be covered by the split proteins. Finally, a fusion/fission family had to have at least one composite protein and one set of split proteins encoded by syntenic genes passing all listed filtering criteria (Fig. 1). When genes encoding the split proteins are not colocalized but have the same domain combination and cover the same composite protein as the ones in synteny, they were added to the split side of the fusion/fission family.
MCL clustering and functional annotation
The composite proteins from predicted fusion/fission families were subjected to global alignment using the Needleman-Wunsch algorithm and clustered using the Markov algorithm (MCL) (135) with the inflation parameter set to 1.2. Inflation of 1.2 results in lower cluster granularity which can potentially lead to poorly related paralogues sequences being attributed to the same cluster. To split paralogs into orthologous groups, the clusters with more than 10 members were further subclustered at 1.6 inflation. If resulting subclusters demonstrated clear separation by the combination of existing functional annotations or/and sequence length while keeping maximum inter-cluster identities below 40% then they were kept, otherwise, it remained at 1.2. Apart from existing PFAM annotations, the cluster proteins were annotated using KofamScan, Kyoto Encyclopedia of Genes and Genomes (KEGG) (136) based software to retrieve orthology (KO) mappings. For KO models with no threshold to be considered, they had to be the protein’s best hit and with an E-value below 1e−10. The EggNOG database tool, eggNOG-mapper, was used to retrieve Clusters of Orthologous Groups of proteins (COGs), including arCOGs (137, 138). Finally, the transporter classification database (TCDB) (139) was used to refine transporter assignments.
Bacterial mapping
The bacterial proteins were searched for protein domains with the same combination of tools as archaeal proteins. The proteins with the same composite domain architecture were annotated with KofamScan and aligned to the archaeal genomes the fusion/fission proteins came from using DIAMOND blastp. The output was filtered for the best one-directional hit. If the best hit was the predicted archaeal composite protein and both bacterial and archaeal proteins had identical domain architectures, the bacterial protein was mapped to the mcl cluster of the archaeal fusion protein. If the domain architectures differed, then it was counted separately. For each candidate, the bacterial protein with multiple best hits and the hit with the highest score per identical and non-identical domain architecture were picked to represent the final affiliation to a fusion/fission protein family.
Fusion/fission classification
Protein clusters were classified as fusion, fission, or both based on the distribution of composite and syntenic split states, genome quality, taxonomic, and functional affiliation, using a simple heuristic approach. First, the frequency of split protein sets per family was used as an initial criterion for the class separation (less than 0.1 for fission, more than 0.9 for a fusion). The second criterion was the taxonomic distribution of composite proteins in bacteria. A fission assignment would prevail if the composite state was present in more than two lineages (phylum) or at least 1,000 bacterial genomes. On the other hand, the absence of bacterial diversity favored a fusion classification. The presence of composite and split proteins from closed genomes, the frequency in the case of split and composite proteins, was the third criterion to classify a protein cluster. When the majority of complete genomes in a protein family contained composite genes, the classification would favor fissions. On the opposite, the prevalence of split proteins in high-quality genomes would favor fusions. When available, an additional criterion was considering the median KO model coverage of split and composite proteins. In cases where fused proteins had full KO coverage and unfused proteins had it halved, this would indicate a fission. However, as many sequences used to build models are of bacterial origin, other criteria had the advantage. If the majority of criteria agreed either on fusion or on fission, then the assignment would be “high confidence,” otherwise a protein cluster would get the “probable” one. If no features were found to support either of the assignments a protein cluster would remain unclassified. Finally, if there is a high frequency of proteins that are composite and split partners mapped to more complex proteins within the same cluster then the assignment would be “fusion and fission.” Furthermore, all protein families were manually analyzed.
ACKNOWLEDGMENTS
We acknowledge support from the Wiener Wissenschafts, Forschungs- und Technologiefonds (Austria) through the grant VRG15-007.
All authors thank members of the Genome Evolution and Ecology group for fruitful discussions.
F.L.S. designed the analysis; G.N. and A.P. contributed to the fusion/fission screening method development; A.P. and F.L.S. analyzed the data; and A.P. and F.L.S. wrote the initial draft of the paper. All authors have seen and approved the final version submitted.
Contributor Information
Filipa L. Sousa, Email: filipa.sousa@univie.ac.at.
Saheed Imam, LifeMine Therapeutics, Cambridge, Massachusetts, USA.
DATA AVAILABILITY
The mappings are at the basis of Fig. 1 to 5; Fig. S1 to 3 are available in supplemental tables. Bacterial mappings are deposited at Figshare (54) ( https://doi.org/10.6084/m9.figshare.24084189). The code will be made available upon request.
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/msystems.00948-23.
Supplemental discussion, supplemental figures, and captions for supplemental tables.
Archaeal data set: taxonomy, completeness, and contamination.
Archaeal fission/fusion families distribution and functional annotations of the composite and split components, including domain assignments.
Bacterial data set: taxonomy, completeness, and contamination.
E. coli fusions in archaea.
Presence or absence of fusion/fission composite proteins per assembly.
Counts of fusion/fission families per taxonomic lineage.
Presence or absence of archaeal composite proteins and corresponding bacterial mappings.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Castelle CJ, Banfield JF. 2018. Major new microbial groups expand diversity and alter our understanding of the tree of life. Cell 172:1181–1197. doi: 10.1016/j.cell.2018.02.016 [DOI] [PubMed] [Google Scholar]
- 2. Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, Butterfield CN, Hernsdorf AW, Amano Y, Ise K, Suzuki Y, Dudek N, Relman DA, Finstad KM, Amundson R, Thomas BC, Banfield JF. 2016. A new view of the tree of life. Nat Microbiol 1:16048. doi: 10.1038/nmicrobiol.2016.48 [DOI] [PubMed] [Google Scholar]
- 3. Adam PS, Borrel G, Brochier-Armanet C, Gribaldo S. 2017. The growing tree of Archaea: new perspectives on their diversity, evolution and ecology. ISME J 11:2407–2425. doi: 10.1038/ismej.2017.122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Parks DH, Rinke C, Chuvochina M, Chaumeil P-A, Woodcroft BJ, Evans PN, Hugenholtz P, Tyson GW. 2017. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nat Microbiol 2:1533–1542. doi: 10.1038/s41564-017-0012-7 [DOI] [PubMed] [Google Scholar]
- 5. Sayers EW, Cavanaugh M, Clark K, Pruitt KD, Schoch CL, Sherry ST, Karsch-Mizrachi I. 2021. Genbank. Nucleic Acids Research 49:D92–D96. doi: 10.1093/nar/gkaa1023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Huber H, Hohn MJ, Rachel R, Fuchs T, Wimmer VC, Stetter KO. 2002. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417:63–67. doi: 10.1038/417063a [DOI] [PubMed] [Google Scholar]
- 7. Murugkar PP, Collins AJ, Chen T, Dewhirst FE. 2020. Isolation and cultivation of candidate phyla radiation saccharibacteria (TM7) bacteria in coculture with bacterial hosts. J Oral Microbiol 12:1814666. doi: 10.1080/20002297.2020.1814666 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Imachi H, Nobu MK, Nakahara N, Morono Y, Ogawara M, Takaki Y, Takano Y, Uematsu K, Ikuta T, Ito M, Matsui Y, Miyazaki M, Murata K, Saito Y, Sakai S, Song C, Tasumi E, Yamanaka Y, Yamaguchi T, Kamagata Y, Tamaki H, Takai K. 2020. Isolation of an Archaeon at the prokaryote–eukaryote interface. Nature 577:519–525. doi: 10.1038/s41586-019-1916-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Hu H, Natarajan VP, Wang F. 2021. Towards enriching and isolation of uncultivated Archaea from marine sediments using a refined combination of conventional microbial cultivation methods. Mar Life Sci Technol 3:231–242. doi: 10.1007/s42995-021-00092-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Yakimov MM, Merkel AY, Gaisin VA, Pilhofer M, Messina E, Hallsworth JE, Klyukina AA, Tikhonova EN, Gorlenko VM. 2022. Cultivation of a vampire: candidatus absconditicoccus praedator. Environ Microbiol 24:30–49. doi: 10.1111/1462-2920.15823 [DOI] [PubMed] [Google Scholar]
- 11. Offre P, Spang A, Schleper C. 2013. Archaea in biogeochemical cycles. Annu Rev Microbiol 67:437–457. doi: 10.1146/annurev-micro-092412-155614 [DOI] [PubMed] [Google Scholar]
- 12. Balch WE, Magrum LJ, Fox GE, Wolfe RS, Woese CR. 1977. An ancient divergence among the bacteria. J Mol Evol 9:305–311. doi: 10.1007/BF01796092 [DOI] [PubMed] [Google Scholar]
- 13. Springer E, Sachs MS, Woese CR, Boone DR. 1995. Partial gene sequences for the a subunit of methyl-coenzyme M reductase (mcrI) as a phylogenetic tool for the family methanosarcinaceae. Int J Syst Bacteriol 45:554–559. doi: 10.1099/00207713-45-3-554 [DOI] [PubMed] [Google Scholar]
- 14. Sun Y, Liu Y, Pan J, Wang F, Li M. 2020. Perspectives on cultivation strategies of Archaea. Microb Ecol 79:770–784. doi: 10.1007/s00248-019-01422-7 [DOI] [PubMed] [Google Scholar]
- 15. Baker BJ, De Anda V, Seitz KW, Dombrowski N, Santoro AE, Lloyd KG. 2020. Diversity, ecology and evolution of Archaea. Nat Microbiol 5:887–900. doi: 10.1038/s41564-020-0715-z [DOI] [PubMed] [Google Scholar]
- 16. Ulas T, Riemer SA, Zaparty M, Siebers B, Schomburg D. 2012. Genome-scale reconstruction and analysis of the metabolic network in the hyperthermophilic archaeon sulfolobus solfataricus. PLoS One 7:e43401. doi: 10.1371/journal.pone.0043401 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Feist AM, Scholten JCM, Palsson BØ, Brockman FJ, Ideker T. 2006. Modeling methanogenesis with a genome-scale metabolic reconstruction of methanosarcina barkeri. Mol Syst Biol 2:1–14. doi: 10.1038/msb4100046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Elkins JG, Podar M, Graham DE, Makarova KS, Wolf Y, Randau L, Hedlund BP, Brochier-Armanet C, Kunin V, Anderson I, Lapidus A, Goltsman E, Barry K, Koonin EV, Hugenholtz P, Kyrpides N, Wanner G, Richardson P, Keller M, Stetter KO. 2008. A korarchaeal genome reveals insights into the evolution of the Archaea. Proc Natl Acad Sci U S A 105:8102–8107. doi: 10.1073/pnas.0801980105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. McKay LJ, Dlakić M, Fields MW, Delmont TO, Eren AM, Jay ZJ, Klingelsmith KB, Rusch DB, Inskeep WP. 2019. Co-occurring genomic capacity for anaerobic methane and dissimilatory sulfur metabolisms discovered in the korarchaeota. Nat Microbiol 4:614–622. doi: 10.1038/s41564-019-0362-4 [DOI] [PubMed] [Google Scholar]
- 20. Berghuis BA, Yu FB, Schulz F, Blainey PC, Woyke T, Quake SR. 2019. Hydrogenotrophic methanogenesis in archaeal phylum verstraetearchaeota reveals the shared ancestry of all methanogens. Proc Natl Acad Sci U S A 116:5037–5044. doi: 10.1073/pnas.1815631116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA. 2005. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437:543–546. doi: 10.1038/nature03911 [DOI] [PubMed] [Google Scholar]
- 22. Dridi B, Fardeau M-L, Ollivier B, Raoult D, Drancourt M. 2012. Methanomassiliicoccus luminyensis gen. nov., sp. nov., a methanogenic archaeon isolated from human faeces. Int J Syst Evol Microbiol 62:1902–1907. doi: 10.1099/ijs.0.033712-0 [DOI] [PubMed] [Google Scholar]
- 23. Chernyh NA, Neukirchen S, Frolov EN, Sousa FL, Miroshnichenko ML, Merkel AY, Pimenov NV, Sorokin DY, Ciordia S, Mena MC, Ferrer M, Golyshin PN, Lebedinsky AV, Cardoso Pereira IA, Bonch-Osmolovskaya EA. 2020. Dissimilatory sulfate reduction in the archaeon ‘candidatus vulcanisaeta moutnovskia’ sheds light on the evolution of sulfur metabolism. Nat Microbiol 5:1428–1438. doi: 10.1038/s41564-020-0776-z [DOI] [PubMed] [Google Scholar]
- 24. Ettwig KF, Zhu B, Speth D, Keltjens JT, Jetten MSM, Kartal B. 2016. Archaea catalyze iron-dependent anaerobic oxidation of methane. Proc Natl Acad Sci U S A 113:12792–12796. doi: 10.1073/pnas.1609534113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Dahl C, Kredich NM, Deutzmann R, Trüper HG. 1993. Dissimilatory sulphite reductase from Archaeoglobus fulgidus: physico-chemical properties of the enzyme and cloning, sequencing and analysis of the reductase genes. J Gen Microbiol 139:1817–1828. doi: 10.1099/00221287-139-8-1817 [DOI] [PubMed] [Google Scholar]
- 26. Krüger M, Meyerdierks A, Glöckner FO, Amann R, Widdel F, Kube M, Reinhardt R, Kahnt J, Böcher R, Thauer RK, Shima S. 2003. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically. Nature 426:878–881. doi: 10.1038/nature02207 [DOI] [PubMed] [Google Scholar]
- 27. Setzke E, Hedderich R, Heiden S, Thauer RK. 1994. H2: heterodisulfide oxidoreductase complex from methanobacterium thermoautotrophicum. Eur J Biochem 220:139–148. doi: 10.1111/j.1432-1033.1994.tb18608.x [DOI] [PubMed] [Google Scholar]
- 28. Grabarse W, Mahlert F, Shima S, Thauer RK, Ermler U. 2000. Comparison of three methyl-coenzyme M reductases from phylogenetically distant organisms: unusual amino acid modification, conservation and adaptation. J Mol Biol 303:329–344. doi: 10.1006/jmbi.2000.4136 [DOI] [PubMed] [Google Scholar]
- 29. Radivojac P, Clark WT, Oron TR, Schnoes AM, Wittkop T, Sokolov A, Graim K, Funk C, Verspoor K, Ben-Hur A, et al. 2013. A large-scale evaluation of computational protein function prediction. Nat Methods 10:221–227. doi: 10.1038/nmeth.2340 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Goberna M, Verdú M. 2016. Predicting microbial traits with phylogenies. ISME J 10:959–967. doi: 10.1038/ismej.2015.171 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Gligorijević V, Renfrew PD, Kosciolek T, Leman JK, Berenberg D, Vatanen T, Chandler C, Taylor BC, Fisk IM, Vlamakis H, Xavier RJ, Knight R, Cho K, Bonneau R. 2021. Structure-based protein function prediction using graph convolutional networks. Nat Commun 12:3168. doi: 10.1038/s41467-021-23303-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kulmanov M, Hoehndorf R. 2020. DeepGOPLus: improved protein function prediction from sequence. Bioinformatics 36:422–429. doi: 10.1093/bioinformatics/btz595 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Bileschi ML, Belanger D, Bryant DH, Sanderson T, Carter B, Sculley D, Bateman A, DePristo MA, Colwell LJ. 2022. Using deep learning to annotate the protein universe. Nat Biotechnol 40:932–937. doi: 10.1038/s41587-021-01179-w [DOI] [PubMed] [Google Scholar]
- 34. Rives A, Meier J, Sercu T, Goyal S, Lin Z, Liu J, Guo D, Ott M, Zitnick CL, Ma J, Fergus R. 2021. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proc Natl Acad Sci U S A 118:e2016239118. doi: 10.1073/pnas.2016239118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Holm L, Laakso LM. 2016. Dali server update. Nucleic Acids Res 44:W351–W355. doi: 10.1093/nar/gkw357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Eddy SR. 1998. Profile hidden markov models. Bioinformatics 14:755–763. doi: 10.1093/bioinformatics/14.9.755 [DOI] [PubMed] [Google Scholar]
- 37. Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. 1990. Basic local alignment search tool. J Mol Biol 215:403–410. doi: 10.1016/S0022-2836(05)80360-2 [DOI] [PubMed] [Google Scholar]
- 38. Buchfink B, Xie C, Huson DH. 2015. Fast and sensitive protein alignment using DIAMOND. Nat Methods 12:59–60. doi: 10.1038/nmeth.3176 [DOI] [PubMed] [Google Scholar]
- 39. Huynen M, Snel B, Lathe W, Bork P. 2000. Predicting protein function by genomic context: quantitative evaluation and qualitative inferences. Genome Res 10:1204–1210. doi: 10.1101/gr.10.8.1204 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Nelson-Sathi S, Sousa FL, Roettger M, Lozada-Chávez N, Thiergart T, Janssen A, Bryant D, Landan G, Schönheit P, Siebers B, McInerney JO, Martin WF. 2015. Origins of major archaeal clades correspond to gene acquisitions from bacteria. Nature 517:77–80. doi: 10.1038/nature13805 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Gabaldón T, Koonin EV. 2013. Functional and evolutionary implications of gene orthology. Nat Rev Genet 14:360–366. doi: 10.1038/nrg3456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Kummerfeld SK, Teichmann SA. 2005. Relative rates of gene fusion and fission in multi-domain proteins. Trends Genet 21:25–30. doi: 10.1016/j.tig.2004.11.007 [DOI] [PubMed] [Google Scholar]
- 43. Enright AJ, Iliopoulos I, Kyrpides NC, Ouzounis CA. 1999. Protein interaction maps for complete genomes based on gene fusion events. Nature 402:86–90. doi: 10.1038/47056 [DOI] [PubMed] [Google Scholar]
- 44. Date SV. 2008. The rosetta stone method, p 169–180. In Keith JM (ed), Bioinformatics: structure, function and applications [DOI] [PubMed] [Google Scholar]
- 45. Henry CS, Lerma-Ortiz C, Gerdes SY, Mullen JD, Colasanti R, Zhukov A, Frelin O, Thiaville JJ, Zallot R, Niehaus TD, Hasnain G, Conrad N, Hanson AD, de Crécy-Lagard V. 2016. Systematic identification and analysis of frequent gene fusion events in metabolic pathways. BMC Genomics 17:473. doi: 10.1186/s12864-016-2782-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Yanai I, Derti A, DeLisi C. 2001. Genes linked by fusion events are generally of the same functional category: a systematic analysis of 30 microbial genomes. Proc Natl Acad Sci U S A 98:7940–7945. doi: 10.1073/pnas.141236298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Pasek S, Risler JL, Brézellec P. 2006. Gene fusion/fission is a major contributor to evolution of multi-domain bacterial proteins. Bioinformatics 22:1418–1423. doi: 10.1093/bioinformatics/btl135 [DOI] [PubMed] [Google Scholar]
- 48. Snel B, Bork P, Huynen M. 2000. Genome evolutiongene fusion versus gene fission. Trends Genet 16:9–11. doi: 10.1016/s0168-9525(99)01924-1 [DOI] [PubMed] [Google Scholar]
- 49. Buljan M, Bateman AG. 2009. The evolution of protein domain families. Biochem Soc Trans 37:751–755. doi: 10.1042/BST0370751 [DOI] [PubMed] [Google Scholar]
- 50. Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C. 2021. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 49:D605–D612. doi: 10.1093/nar/gkaa1074 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, Edwards RA, Gerdes S, Parrello B, Shukla M, Vonstein V, Wattam AR, Xia F, Stevens R. 2014. The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucl Acids Res 42:D206–D214. doi: 10.1093/nar/gkt1226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Markowitz VM, Chen I-M, Palaniappan K, Chu K, Szeto E, Grechkin Y, Ratner A, Jacob B, Huang J, Williams P, Huntemann M, Anderson I, Mavromatis K, Ivanova NN, Kyrpides NC. 2012. IMG: the integrated microbial genomes database and comparative analysis system. Nucleic Acids Res 40:D115–D122. doi: 10.1093/nar/gkr1044 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Reid AJ, Ranea JAG, Clegg AB, Orengo CA. 2010. CODA: accurate detection of functional associations between proteins in eukaryotic genomes using domain fusion. PLoS ONE 5:e10908. doi: 10.1371/journal.pone.0010908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Padalko A, Nair G, Sousa FL. 2024. Bacterial Mappings. doi: 10.6084/m9.figshare.24084189 [DOI]
- 55. Grochowski LL, Xu H, White RH. 2005. Ribose-5-phosphate biosynthesis in Methanocaldococcus jannaschii occurs in the absence of a pentose-phosphate pathway. J Bacteriol 187:7382–7389. doi: 10.1128/JB.187.21.7382-7389.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Jespersen M, Wagner T. 2023. Assimilatory sulfate reduction in the marine methanogen methanothermococcus thermolithotrophicus. Nat Microbiol 8:1227–1239. doi: 10.1038/s41564-023-01398-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Susanti D, Mukhopadhyay B. 2012. An intertwined evolutionary history of methanogenic Archaea and sulfate reduction. PLoS One 7:e45313. doi: 10.1371/journal.pone.0045313 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Mukhopadhyay B, Patel VJ, Wolfe RS. 2000. A stable archaeal pyruvate carboxylase from the hyperthermophile Methanococcus jannaschii. Arch Microbiol 174:406–414. doi: 10.1007/s002030000225 [DOI] [PubMed] [Google Scholar]
- 59. Mukhopadhyay B, Purwantini E, Kreder CL, Wolfe RS. 2001. Oxaloacetate synthesis in the methanarchaeon methanosarcina Barkeri: pyruvate carboxylase genes and a putative Escherichia coli-type bifunctional biotin protein ligase gene (BPL/birA) exhibit a unique organization. J Bacteriol 183:3804–3810. doi: 10.1128/JB.183.12.3804-3810.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Spang A, Stairs CW, Dombrowski N, Eme L, Lombard J, Caceres EF, Greening C, Baker BJ, Ettema TJG. 2019. Proposal of the reverse flow model for the origin of the eukaryotic cell based on comparative analyses of asgard archaeal metabolism. Nat Microbiol 4:1138–1148. doi: 10.1038/s41564-019-0406-9 [DOI] [PubMed] [Google Scholar]
- 61. López-García P, Moreira D. 2020. The syntrophy hypothesis for the origin of eukaryotes revisited. Nat Microbiol 5:655–667. doi: 10.1038/s41564-020-0710-4 [DOI] [PubMed] [Google Scholar]
- 62. Zaremba-Niedzwiedzka K, Caceres EF, Saw JH, Bäckström D, Juzokaite L, Vancaester E, Seitz KW, Anantharaman K, Starnawski P, Kjeldsen KU, Stott MB, Nunoura T, Banfield JF, Schramm A, Baker BJ, Spang A, Ettema TJG. 2017. Asgard Archaea illuminate the origin of eukaryotic cellular complexity. Nature 541:353–358. doi: 10.1038/nature21031 [DOI] [PubMed] [Google Scholar]
- 63. Liu Y, Makarova KS, Huang W-C, Wolf YI, Nikolskaya AN, Zhang X, Cai M, Zhang C-J, Xu W, Luo Z, Cheng L, Koonin EV, Li M. 2021. Expanded diversity of asgard Archaea and their relationships with eukaryotes. Nature 593:553–557. doi: 10.1038/s41586-021-03494-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Vulcanisaeta Souniana JCM 11219. 2002. Available from: https://img.jgi.doe.gov/cgi-bin/m/main.cgi?section=TaxonDetail&page=taxonDetail&taxon_oid=2681813013
- 65. Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R. 2005. The microbial pan-genome. Curr Opin Genet Dev 15:589–594. doi: 10.1016/j.gde.2005.09.006 [DOI] [PubMed] [Google Scholar]
- 66. Thomas C, Aller SG, Beis K, Carpenter EP, Chang G, Chen L, Dassa E, Dean M, Duong Van Hoa F, Ekiert D, et al. 2020. Structural and functional diversity calls for a new classification of ABC transporters. FEBS Lett 594:3767–3775. doi: 10.1002/1873-3468.13935 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Vettakkorumakankav NN, Stevenson KJ. 1992. Dihydrolipoamide dehydrogenase from haloferax volcanii: gene cloning, complete primary structure, and comparison to other dihydrolipoamide dehydrogenases. Biochem Cell Biol 70:656–663. doi: 10.1139/o92-101 [DOI] [PubMed] [Google Scholar]
- 68. Tersteegen A, Linder D, Thauer RK, Hedderich R. 1997. Structures and functions of four anabolic 2-oxoacid oxidoreductases in methanobacterium thermoautotrophicum. Eur J Biochem 244:862–868. doi: 10.1111/j.1432-1033.1997.00862.x [DOI] [PubMed] [Google Scholar]
- 69. Prodromou C, Artymiuk PJ, Guest JR. 1992. The aconitase of Escherichia coli. nucleotide sequence of the aconitase gene and amino acid sequence similarity with mitochondrial aconitases, the iron-responsive-element-binding protein and isopropylmalate isomerases. Eur J Biochem 204:599–609. doi: 10.1111/j.1432-1033.1992.tb16673.x [DOI] [PubMed] [Google Scholar]
- 70. Drevland RM, Jia Y, Palmer DRJ, Graham DE. 2008. Methanogen homoaconitase catalyzes both hydrolyase reactions in coenzyme B biosynthesis. J Biol Chem 283:28888–28896. doi: 10.1074/jbc.M802159200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Heim S, Künkel A, Thauer RK, Hedderich R. 1998. Thiol:fumarate reductase (Tfr) from methanobacterium thermoautotrophicum. identification of the catalytic sites for fumarate reduction and thiol oxidation. Eur J Biochem 253:292–299. doi: 10.1046/j.1432-1327.1998.2530292.x [DOI] [PubMed] [Google Scholar]
- 72. Buck D, Spencer ME, Guest JR. 1986. Cloning and expression of the succinyl-CoA synthetase genes of Escherichia coli K12. J Gen Microbiol 132:1753–1762. doi: 10.1099/00221287-132-6-1753 [DOI] [PubMed] [Google Scholar]
- 73. Fatland BL, Ke J, Anderson MD, Mentzen WI, Cui LW, Allred CC, Johnston JL, Nikolau BJ, Wurtele ES. 2002. Molecular characterization of a heteromeric ATP-citrate Lyase that generates cytosolic acetyl-coenzyme A in arabidopsis. Plant Physiol 130:740–756. doi: 10.1104/pp.008110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Verschueren KHG, Blanchet C, Felix J, Dansercoer A, De Vos D, Bloch Y, Van Beeumen J, Svergun D, Gutsche I, Savvides SN, Verstraete K. 2019. Structure of ATP citrate Lyase and the origin of citrate synthase in the Krebs cycle. Nature 568:571–575. doi: 10.1038/s41586-019-1095-5 [DOI] [PubMed] [Google Scholar]
- 75. Martinez-Cruz LA, Dreyer MK, Boisvert DC, Yokota H, Martinez-Chantar ML, Kim R, Kim S-H. 2002. Crystal structure of MJ1247 protein from M. ˚ resolution infers a molecular jannaschii at 2. 0 A function of 3-hexulose-6-phosphate isomerase physical BioSciences division of Lawrence. Structure 10:195–204. doi: 10.1016/s0969-2126(02)00701-3 [DOI] [PubMed] [Google Scholar]
- 76. Bräsen C, Esser D, Rauch B, Siebers B. 2014. Carbohydrate metabolism in archaea: current insights into unusual enzymes and pathways and their regulation. JAMA Ophthalmol 132:326–331. doi: 10.1111/j.1462-2920.2012.02893.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Spang A, Poehlein A, Offre P, Zumbrägel S, Haider S, Rychlik N, Nowka B, Schmeisser C, Lebedeva EV, Rattei T, Böhm C, Schmid M, Galushko A, Hatzenpichler R, Weinmaier T, Daniel R, Schleper C, Spieck E, Streit W, Wagner M. 2012. The genome of the ammonia-oxidizing candidatus nitrososphaera gargensis: insights into metabolic versatility and environmental adaptations. Environ Microbiol 14:3122–3145. doi: 10.1111/j.1462-2920.2012.02893.x [DOI] [PubMed] [Google Scholar]
- 78. Reji L, Francis CA. 2020. Metagenome-assembled genomes reveal unique metabolic adaptations of a basal marine thaumarchaeota lineage. ISME J 14:2105–2115. doi: 10.1038/s41396-020-0675-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Zhong H, Lehtovirta-Morley L, Liu J, Zheng Y, Lin H, Song D, Todd JD, Tian J, Zhang X-H. 2020. Novel insights into the thaumarchaeota in the deepest oceans: their metabolism and potential adaptation mechanisms. Microbiome 8:78. doi: 10.1186/s40168-020-00849-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Berg IA, Kockelkorn D, Buckel W, Fuchs G. 2007. A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea. Science 318:1782–1786. doi: 10.1126/science.1149976 [DOI] [PubMed] [Google Scholar]
- 81. Könneke M, Schubert DM, Brown PC, Hügler M, Standfest S, Schwander T, Schada von Borzyskowski L, Erb TJ, Stahl DA, Berg IA. 2014. Ammonia-oxidizing Archaea use the most energy-efficient aerobic pathway for CO2 fixation. Proc Natl Acad Sci U S A 111:8239–8244. doi: 10.1073/pnas.1402028111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Musfeldt M, Selig M, Schönheit P. 1999. Acetyl coenzyme A synthetase (ADP forming) from the hyperthermophilic archaeon pyrococcus furiosus: Identification, cloning, separate expression of the encoding genes, acdAI and acdBI, in Escherichia coli, and in vitro reconstitution of the active heterot. J Bacteriol 181:5885–5888. doi: 10.1128/JB.181.18.5885-5888.1999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Fuchs G. 2011. Alternative pathways of carbon dioxide fixation: insights into the early evolution of life?. Annu Rev Microbiol 65:631–658. doi: 10.1146/annurev-micro-090110-102801 [DOI] [PubMed] [Google Scholar]
- 84. Ramos-Vera WH, Weiss M, Strittmatter E, Kockelkorn D, Fuchs G. 2011. Identification of missing genes and enzymes for autotrophic carbon fixation in crenarchaeota. J Bacteriol 193:1201–1211. doi: 10.1128/JB.01156-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Thauer RK, Kaster AK, Seedorf H, Buckel W, Hedderich R. 2008. Methanogenic Archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol 6:579–591. doi: 10.1038/nrmicro1931 [DOI] [PubMed] [Google Scholar]
- 86. Vorholt JA, Vaupel M, Thauer RK. 1996. A polyferredoxin with eight [4Fe-4S] clusters as a subunit of molybdenum formylmethanofuran dehydrogenase from methanosarcina barkeri. Eur J Biochem 236:309–317. doi: 10.1111/j.1432-1033.1996.t01-1-00309.x [DOI] [PubMed] [Google Scholar]
- 87. Watanabe T, Pfeil-Gardiner O, Kahnt J, Koch J, Shima S, Murphy BJ. 2021. Three-megadalton complex of methanogenic electron-bifurcating and CO2-fixing enzymes. Science 373:1151–1156. doi: 10.1126/science.abg5550 [DOI] [PubMed] [Google Scholar]
- 88. Harms U, Weiss DS, Gärtner P, Linder D, Thauer RK. 1995. The energy conserving N5-methyltetrahydromethanopterin:coenzyme M methyltransferase complex from methanobacterium thermoautotrophicum is composed of eight different subunits. Eur J Biochem 228:640–648. doi: 10.1111/j.1432-1033.1995.0640m.x [DOI] [PubMed] [Google Scholar]
- 89. Rospert S, Linder D, Ellermann J, Thauer RK. 1990. Two genetically distinct methyl-coenzyme M reductases in methanobacterium thermoautotrophicum strain marburg and delta H. Eur J Biochem 194:871–877. doi: 10.1111/j.1432-1033.1990.tb19481.x [DOI] [PubMed] [Google Scholar]
- 90. Evans PN, Boyd JA, Leu AO, Woodcroft BJ, Parks DH, Hugenholtz P, Tyson GW. 2019. An evolving view of methane metabolism in the Archaea. Nat Rev Microbiol 17:219–232. doi: 10.1038/s41579-018-0136-7 [DOI] [PubMed] [Google Scholar]
- 91. Buan NR, Metcalf WW. 2010. Methanogenesis by methanosarcina acetivorans involves two structurally and functionally distinct classes of heterodisulfide reductase. Mol Microbiol 75:843–853. doi: 10.1111/j.1365-2958.2009.06990.x [DOI] [PubMed] [Google Scholar]
- 92. Appel L, Willistein M, Dahl C, Ermler U, Boll M. 2021. Functional diversity of prokaryotic HdrA(BC) modules: role in flavin-based electron bifurcation processes and beyond. Biochim Biophys Acta Bioenerg 1862:148379. doi: 10.1016/j.bbabio.2021.148379 [DOI] [PubMed] [Google Scholar]
- 93. Graupner M, Xu H, White RH. 2000. Identification of the gene encoding sulfopyruvate decarboxylase, an enzyme involved in biosynthesis of coenzyme M. J Bacteriol 182:4862–4867. doi: 10.1128/JB.182.17.4862-4867.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Schnell R, Sandalova T, Hellman U, Lindqvist Y, Schneider G. 2005. Siroheme- and [Fe4-S4]-dependent NirA from Mycobacterium tuberculosis is a sulfite reductase with a covalent Cys-Tyr bond in the active site. J Biol Chem 280:27319–27328. doi: 10.1074/jbc.M502560200 [DOI] [PubMed] [Google Scholar]
- 95. Bordo D, Bork P. 2002. The rhodanese/Cdc25 phosphatase superfamily. sequence-structure-function relations. EMBO Rep 3:741–746. doi: 10.1093/embo-reports/kvf150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Ichiki H, Tanaka Y, Mochizuki K, Yoshimatsu K, Sakurai T, Fujiwara T. 2001. Purification, characterization, and genetic analysis of cu-containing dissimilatory nitrite reductase from a denitrifying halophilic archaeon, haloarcula marismortui . J Bacteriol 183:4149–4156. doi: 10.1128/JB.183.14.4149-4156.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Miralles-Robledillo JM, Bernabeu E, Giani M, Martínez-Serna E, Martínez-Espinosa RM, Pire C. 2021. Distribution of denitrification among haloarchaea: a comprehensive study. Microorganisms 9:1669. doi: 10.3390/microorganisms9081669 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Lund MB, Smith JM, Francis CA. 2012. Diversity, abundance and expression of nitrite reductase (nirK)-Like genes in marine thaumarchaea. ISME J 6:1966–1977. doi: 10.1038/ismej.2012.40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Hipp WM, Pott AS, Thum-Schmitz N, Faath I, Dahl C, Trüper HG. 1997. Towards the phylogeny of APS reductases and sirohaem sulfite reductases in sulfate-reducing and sulfur-oxidizing prokaryotes. Microbiology (Reading) 143 (Pt 9):2891–2902. doi: 10.1099/00221287-143-9-2891 [DOI] [PubMed] [Google Scholar]
- 100. Berendt U, Haverkamp T, Prior A, Schwenn JD. 1995. Reaction mechanism of thioredoxin: 3′-phospho-adenylylsulfate reductase investigated by site-directed mutagenesis. Eur J Biochem 233:347–356. doi: 10.1111/j.1432-1033.1995.347_1.x [DOI] [PubMed] [Google Scholar]
- 101. Mihara H, Kurihara T, Yoshimura T, Soda K, Esaki N. 1997. Cysteine sulfinate desulfinase, a NIFS-like protein of Escherichia coli with selenocysteine lyase and cysteine desulfurase activities. Gene cloning, purification, and characterization of a novel pyridoxal enzyme. J Biol Chem 272:22417–22424. doi: 10.1074/jbc.272.36.22417 [DOI] [PubMed] [Google Scholar]
- 102. Zafrilla B, Martínez-Espinosa RM, Esclapez J, Pérez-Pomares F, Bonete MJ. 2010. Sufs protein from haloferax volcanii involved in Fe-S cluster assembly in haloarchaea. Biochim Biophys Acta 1804:1476–1482. doi: 10.1016/j.bbapap.2010.03.001 [DOI] [PubMed] [Google Scholar]
- 103. Lampreia J, Moura I, Teixeira M, Peck HD, Legall J, Huynh BH, Moura JJG. 1990. The active centers of adenylylsulfate reductase from desulfovibrio gigas. Characterization and spectroscopic studies. Eur J Biochem 188:653–664. doi: 10.1111/j.1432-1033.1990.tb15447.x [DOI] [PubMed] [Google Scholar]
- 104. Huang CJ, Barrett EL. 1991. Sequence analysis and expression of the Salmonella Typhimurium asr operon encoding production of hydrogen sulfide from sulfite. J Bacteriol 173:1544–1553. doi: 10.1128/jb.173.4.1544-1553.1991 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Dhillon A, Goswami S, Riley M, Teske A, Sogin M. 2005. Domain evolution and functional diversification of sulfite reductases. Astrobiology 5:18–29. doi: 10.1089/ast.2005.5.18 [DOI] [PubMed] [Google Scholar]
- 106. Marreiros BC, Batista AP, Duarte AMS, Pereira MM. 2013. A missing link between complex I and group 4 membrane-bound [NiFe] hydrogenases. Biochim Biophys Acta 1827:198–209. doi: 10.1016/j.bbabio.2012.09.012 [DOI] [PubMed] [Google Scholar]
- 107. Brüggemann H, Falinski F, Deppenmeier U. 2000. Structure of the F420H2:guinone oxidoreductase of Archaeoglobus fulgidus identification and overproduction of the F420H2-oxidizing subunit. Eur J Biochem 267:5810–5814. doi: 10.1046/j.1432-1327.2000.01657.x [DOI] [PubMed] [Google Scholar]
- 108. Fujimori K, Ohta D. 1998. Isolation and characterization of a histidine biosynthetic gene in arabidopsis encoding a polypeptide with two separate domains for phosphoribosyl-ATP pyrophosphohydrolase and phosphoribosyl-AMP cyclohydrolase. Plant Physiol 118:275–283. doi: 10.1104/pp.118.1.275 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. Fani R, Brilli M, Fondi M, Lió P. 2007. The role of gene fusions in the evolution of metabolic pathways: the histidine biosynthesis case. BMC Evol Biol 7 Suppl 2:S4. doi: 10.1186/1471-2148-7-S2-S4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110. Kuenzler M, Balmelli T, Egli CM, Paravicini G, Braus GH. 1993. Cloning, primary structure, and regulation of the HIS7 gene encoding a bifunctional glutamine amidotransferase:cyclase from saccharomyces cerevisiae. J Bacteriol 175:5548–5558. doi: 10.1128/jb.175.17.5548-5558.1993 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Rangarajan ES, Proteau A, Wagner J, Hung M-N, Matte A, Cygler M. 2006. Structural snapshots of Escherichia coli histidinol phosphate phosphatase along the reaction pathway. J Biol Chem 281:37930–37941. doi: 10.1074/jbc.M604916200 [DOI] [PubMed] [Google Scholar]
- 112. Carlomagno MS, Chiariotti L, Alifano P, Nappo AG, Bruni CB. 1988. Structure and function of the Salmonella Typhimurium and Escherichia coli K-12 histidine operons. J Mol Biol 203:585–606. doi: 10.1016/0022-2836(88)90194-5 [DOI] [PubMed] [Google Scholar]
- 113. Singh SA, Christendat D. 2006. Structure of arabidopsis dehydroquinate dehydratase-shikimate dehydrogenase and implications for metabolic channeling in the shikimate pathway. Biochemistry 45:7787–7796. doi: 10.1021/bi060366+ [DOI] [PubMed] [Google Scholar]
- 114. Løbner-Olesen A, Marinus MG. 1992. Identification of the gene (aroK) encoding shikimic acid kinase I of Escherichia coli. J Bacteriol 174:525–529. doi: 10.1128/jb.174.2.525-529.1992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Nichols BP, Miozzari GF, van Cleemput M, Bennett GN, Yanofsky C. 1980. Nucleotide sequences of the trpG regions of Escherichia coli, Shigella dysenteriae, Salmonella Typhimurium and serratia marcescens. J Mol Biol 142:503–517. doi: 10.1016/0022-2836(80)90260-0 [DOI] [PubMed] [Google Scholar]
- 116. Tang X, Ezaki S, Fujiwara S, Takagi M, Atomi H, Imanaka T. 1999. The tryptophan biosynthesis gene cluster trpCDEGFBA from pyrococcus kodakaraensis KOD1 is regulated at the transcriptional level and expressed as a single mRNA. Mol Gen Genet 262:815–821. doi: 10.1007/s004380051145 [DOI] [PubMed] [Google Scholar]
- 117. Kleeb AC, Kast P, Hilvert D. 2006. A monofunctional and thermostable prephenate dehydratase from the archaeon Methanocaldococcus jannaschii. Biochemistry 45:14101–14110. doi: 10.1021/bi061274n [DOI] [PubMed] [Google Scholar]
- 118. Xu S, Yang Y, Jin R, Zhang M, Wang H. 2006. Purification and characterization of a functionally active Mycobacterium tuberculosis prephenate dehydrogenase. Protein Expr Purif 49:151–158. doi: 10.1016/j.pep.2006.05.020 [DOI] [PubMed] [Google Scholar]
- 119. Mendel RR. 2013. The molybdenum cofactor. J Biol Chem 288:13165–13172. doi: 10.1074/jbc.R113.455311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120. Schwarz G, Mendel RR, Ribbe MW. 2009. Molybdenum cofactors, enzymes and pathways. Nature 460:839–847. doi: 10.1038/nature08302 [DOI] [PubMed] [Google Scholar]
- 121. Rivers SL, McNairn E, Blasco F, Giordano G, Boxer DH. 1993. Molecular genetic analysis of the moa operon of Escherichia coli K‐12 required for molybdenum cofactor biosynthesis. Mol Microbiol 8:1071–1081. doi: 10.1111/j.1365-2958.1993.tb01652.x [DOI] [PubMed] [Google Scholar]
- 122. Yang Y-M, Won Y-B, Ji C-J, Kim J-H, Ryu S-H, Ok Y-H, Lee J-W. 2018. Cleavage of molybdopterin synthase MoaD-MoaE linear fusion by JAMM/MPN+ domain containing metalloprotease DR0402 from deinococcus radiodurans. Biochem Biophys Res Commun 502:48–54. doi: 10.1016/j.bbrc.2018.05.117 [DOI] [PubMed] [Google Scholar]
- 123. Leimkühler S, Wuebbens MM, Rajagopalan KV. 2001. Characterization of Escherichia coli MoeB and its involvement in the activation of molybdopterin synthase for the biosynthesis of the molybdenum cofactor. J Biol Chem 276:34695–34701. doi: 10.1074/jbc.M102787200 [DOI] [PubMed] [Google Scholar]
- 124. Ivanova AA, Oshkin IY, Danilova OV, Philippov DA, Ravin NV, Dedysh SN. 2022. Rokubacteria in northern peatlands: habitat preferences and diversity patterns. Microorganisms 10:11. doi: 10.3390/microorganisms10010011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125. Hasona A, Ray RM, Shanmugam KT. 1998. Physiological and genetic analyses leading to identification of a biochemical role for the moeA (molybdate metabolism) gene product in Escherichia coli. J Bacteriol 180:1466–1472. doi: 10.1128/JB.180.6.1466-1472.1998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126. Dailey HA, Dailey TA, Gerdes S, Jahn D, Jahn M, O’Brian MR, Warren MJ. 2017. Prokaryotic heme biosynthesis: multiple pathways to a common essential product. Microbiol Mol Biol Rev 81:e00048-16. doi: 10.1128/MMBR.00048-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Lobo SAL, Brindley A, Warren MJ, Saraiva LM. 2009. Functional characterization of the early steps of tetrapyrrole biosynthesis and modification in desulfovibrio vulgaris hildenborough. Biochem J 420:317–325. doi: 10.1042/BJ20090151 [DOI] [PubMed] [Google Scholar]
- 128. Kühner M, Haufschildt K, Neumann A, Storbeck S, Streif J, Layer G. 2014. The alternative route to heme in the methanogenic archaeon methanosarcina barkeri. Archaea 2014:327637. doi: 10.1155/2014/327637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129. Makarova KS, Wolf YI, Koonin EV. 2015. Archaeal clusters of orthologous genes (arCOGs): an update and application for analysis of shared features between thermococcales, methanococcales, and methanobacteriales. Life (Basel) 5:818–840. doi: 10.3390/life5010818 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130. Rinke C, Lee J, Nath N, Goudeau D, Thompson B, Poulton N, Dmitrieff E, Malmstrom R, Stepanauskas R, Woyke T. 2014. Obtaining genomes from uncultivated environmental microorganisms using FACS-based single-cell genomics. Nat Protoc 9:1038–1048. doi: 10.1038/nprot.2014.067 [DOI] [PubMed] [Google Scholar]
- 131. El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer ELL, Hirsh L, Paladin L, Piovesan D, Tosatto SCE, Finn RD. 2019. The Pfam protein families database in 2019. Nucleic Acids Res 47:D427–D432. doi: 10.1093/nar/gky995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132. Eddy SR. 2009. A new generation of homology search tools based on probabilistic inference. Genome Informatics 23:205–211. doi: 10.1142/9781848165632_0019 [DOI] [PubMed] [Google Scholar]
- 133. Ochoa A, Singh M. 2017. Domain prediction with probabilistic directional context. Bioinformatics 33:2471–2478. doi: 10.1093/bioinformatics/btx221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134. Mistry J, Finn RD, Eddy SR, Bateman A, Punta M. 2013. Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res 41:e121. doi: 10.1093/nar/gkt263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135. Dongen S. 2000. Graph clustering by flow simulation, PhD thesis, Cent Math Comput Sci [Google Scholar]
- 136. Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. 2016. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44:D457–D462. doi: 10.1093/nar/gkv1070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137. Huerta-Cepas J, Szklarczyk D, Heller D, Hernández-Plaza A, Forslund SK, Cook H, Mende DR, Letunic I, Rattei T, Jensen LJ, von Mering C, Bork P. 2019. EggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47:D309–D314. doi: 10.1093/nar/gky1085 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138. Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV, Krylov DM, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, Smirnov S, Sverdlov AV, Vasudevan S, Wolf YI, Yin JJ, Natale DA. 2003. The COG database: an updated vesion includes eukaryotes. BMC Bioinformatics 4:1–14. doi: 10.1186/1471-2105-4-41 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139. Saier MH, Reddy VS, Moreno-Hagelsieb G, Hendargo KJ, Zhang Y, Iddamsetty V, Lam KJK, Tian N, Russum S, Wang J, Medrano-Soto A. 2021. The transporter classification database (TCDB): 2021 update. Nucleic Acids Res 49:D461–D467. doi: 10.1093/nar/gkaa1004doi:33170213 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental discussion, supplemental figures, and captions for supplemental tables.
Archaeal data set: taxonomy, completeness, and contamination.
Archaeal fission/fusion families distribution and functional annotations of the composite and split components, including domain assignments.
Bacterial data set: taxonomy, completeness, and contamination.
E. coli fusions in archaea.
Presence or absence of fusion/fission composite proteins per assembly.
Counts of fusion/fission families per taxonomic lineage.
Presence or absence of archaeal composite proteins and corresponding bacterial mappings.
Data Availability Statement
The mappings are at the basis of Fig. 1 to 5; Fig. S1 to 3 are available in supplemental tables. Bacterial mappings are deposited at Figshare (54) ( https://doi.org/10.6084/m9.figshare.24084189). The code will be made available upon request.