Abstract
Treponematoses encompass a group of chronic and debilitating bacterial diseases transmitted sexually or by direct contact and attributed to Treponema pallidum. Despite being documented since as far back as 1963, the epidemiology of treponematoses in wild primates has remained an uninvestigated territory due to the inherent challenges associated with conducting examinations and obtaining invasive biological samples from wild animals. The primary aim of this study was to investigate the presence of treponemal infections in the critically endangered Western chimpanzees in Senegal, utilizing an innovative non-invasive stool serology method. We provide compelling evidence of the existence of anti-Treponema-specific antibodies in 13 out of 29 individual chimpanzees. Our study also underscores the significant potential of stool serology as a valuable non-invasive tool for monitoring and surveilling crucial emerging diseases in wild animals. We recognize two major implications: (1) the imperative need to assess the risks of treponematosis in Western chimpanzee populations and (2) the necessity to monitor and manage this disease following a holistic One Health approach.
Keywords: Treponema pallidum, Chimpanzees, Stool, Immunoglobulins, Serology
1. Introduction
The proportion of human infections of zoonotic origin is thought to be about 60%, and about 72% of new human pathogens detected in the last decades have originated in animals [1]. This phenomenon is particularly pronounced when the original host of the pathogen is a non-human primate (NHP).
Wild NHPs were detected to be infected by T. pallidum in Africa since 1963 [2]. Up to date, T. pallidum infections have been reported in NHPs in sub-Saharan Africa, specifically in Guinea, Senegal, Gambia, and Tanzania [[2], [3], [4], [5]] (Fig. 1). The first evidence of a non-human reservoir of T. pallidum was reported from Guinea baboons, Papio papio, without genital lesions [6]. A specific strain (Fribourg-Blanc) has been isolated from popliteal lymph nodes of Guinea baboons; it was found to be very close to T. pallidum pertenue [7], and pathogenic for hamsters and humans [8]. Since this discovery, baboons (P. papio, P. cynocephalus, P. anubis), mangabeys (Cercocebus sp.), green monkeys (Chlorocebus sabaeus), western red colobus monkeys (Piliocolobus badius) and patas monkeys (Erythrocebus patas) across Africa were reported to be positive for T. pallidum infection by serology, immunohistochemistry and molecular biology tests [9]. In 1969 Fribourg-Blanc and Mollaret also reported [6] serological evidences of treponematosis in wild chimpanzees from Central Africa (Pan troglodytes troglodytes) and observational studies from gorillas (Gorilla gorilla) suggested that they may be also affected by yaws-like illness [9]. Interestingly, among 2272 primate sera from Africa and Asia tested in 1969 [10] only West African monkeys and several Central African chimpanzees were positive when tested for syphilis serology. In addition, numerous suspicions of treponemal infection based on physical features (genital and orofacial lesions) have been reported for NHPs in the African continent [4].
Fig. 1.
Geographical map illustrating reported detections of T. pallidum in non-human primates in West Africa from 1963 to the present.
Recently, the opportunity to perform a post-mortem autopsy of a female Western chimpanzee (P. t. verus) in the Republic of Guinea by Mubemba et al., permitted to reveal a severe infection with T. pallidum pertenue [5]. The chimpanzee was euthanized, as her lesions were so severe that she would likely have died from the infection. Although this finding provided evidence of a severe treponemal infection in a critically endangered chimpanzee subspecies, the real impact of treponematosis in chimpanzees is still unclear mostly due to the difficulties of screening for infectious diseases in wild populations.
Epidemiological studies of epidemic diseases in great apes pose unique challenges. It is ethically and logistically impossible to systematically collect blood samples from wild ape populations. Instead, we employ diagnostic techniques that utilize fecal samples [11]. Initially, we relied on PCR sequences, but we have since advanced to stool serology techniques, applicable to humans, monkeys, and apes, for diagnosing infections or tracing their origins [11]. This approach has been instrumental in understanding diseases like HIV, malaria, and their transmission dynamics.
A prominent advantage of this method lies in its non-invasiveness and its absolute harmlessness to the animal subjects [11]. Furthermore, it equips us with the capability to genetically pinpoint the origin of the fecal material, thereby obviating the necessity for redundant sample collection and repetitive testing. In our research endeavors, we harnessed this technique to diagnose treponematosis. Initially, we honed our methodology using human feces as a proof-of-concept before employing it on a repository of chimpanzee feces amidst a well-documented epidemic outbreak in West Africa.
2. Materials and methods
2.1. Ethical considerations
Permits to collect wild chimpanzee fecal samples were granted by the Ministry of Environment and Sustainable Development of Senegal (permit 001914/DEF/DGF from the Direction des Eaux, Forêts, Chasses et de la Conservation des Sols, June, 5, 2016). The authorization to access and use genetic resources according to the Nagoya Protocol on Access to Genetic Resources (№001042, September 2021) was also granted by the Ministry of Environment and Sustainable Development of Senegal. Chimpanzee fecal samples were collected from the substrate in a non-invasive manner, when the apes were not present, in accordance with the animal welfare guidelines of the Jane Goodall Institute Spain and Senegal hygiene and safety protocol, ZooParc de Beauval (letter ZPB/CVCR/2020–87, October 22, 2020) and La Vallée des Singes animal park (letter VDS/Eth/EchFec/2020_10, October 24, 2020).
Two human stools from syphilis-positive patients collected during patient care and sent for diagnostic purposes to the clinical microbiology laboratory of our Public University Hospitals (AP-HM, located at the IHU Méditerranée Infection, Marseille, France) were also retrospectively analyzed as allowed by French law (Article L.1211–2 of the CSP). Patients had been informed of the possible reuse of their samples and their personal data collected during care for research purpose. They had the possibility to oppose this by reporting it to the DPO of the AP-HM. Neither patient expressed any objection.
2.2. Observations and stool sample collection
The Jane Goodall Institute Spain and Senegal conducts a long-term identification of individual chimpanzees from camera trap videos and photographs taken during opportunistic direct observations at the Dindefelo Community Nature Reserve (hereafter Dindefelo), in southeastern Senegal. We analyzed images obtained from July 2013 to December 2016 to identify individuals with orofacial depigmentation. Chimpanzee fecal samples were collected from the substrate in a non-invasive manner, when the apes were not present, in accordance with the animal welfare guidelines of the Jane Goodall Institute Spain and Senegal's hygiene and safety protocol. During this period, camera traps were deployed at 32 locations within and around Dindefelo in places frequented by the chimpanzees (e.g. tool use sites such as termite mounds, and paths).
In August 2016, we collected 48 chimpanzee fecal samples in Dindefelo. During sample collection, we observed some individuals with marked depigmentation spots on the face, strikingly similar to treponematosis-associated lesions (Fig. 2). We also obtained 30 chimpanzee stool samples from the ZooParc de Beauval (Saint-Aignan, France), four chimpanzee stool samples from La Vallée des Singes animal park (Romagne, France), and two human stools from syphilis-positive patients. These samples served as negative and positive controls respectively in this study. All stool samples were collected in individual jars (Labelians, Nemours, France) and 50 mL Falcon tubes (Dutscher, Brumath, France) with an extraction buffer consisting of a phosphate-buffered saline solution supplemented with 4-(2-aminoethyl)-benzenesulphonyl fluoride 4 mM, bestatin 0.26 mM, E-64 28 μM, leupeptin 2 μM and aprotinin 0.6 μM, pH 7.4 and SIGMAFAST™ protease inhibitor tablets (Sigma-Aldrich, St Louis, MO, USA) based on volume/weight (V/W), stored in dry ice and transported to our laboratory where they were maintained at −80 °C prior to analysis.
Fig. 2.
Geographical map of the wild chimpanzee sample collection site (red star) and chimpanzee with depigmentation and orofacial lesions at the Dindefelo Community Nature Reserve (Senegal). (For interpretation of the references to color in this figure legend, the reader is referred to the web version of this article.)
2.3. DNA extraction and qPCR assay
For DNA extraction, 40 mg of each stool sample were mixed in 1.5 mL tubes with 360 μL of G2 lysis buffer from EZ1®DNA Tissue Kit (Qiagen, Courtabeuf, France). After a mechanic lysis with tungsten beads (Qiagen) using FastPrep-24TM 5G Grinder for 40 s, the mixtures were incubated 10 min at 100 °C. The tubes were centrifuged at 10,000 ×g for 1 min and 200 μL of the supernatant was enzymatically digested using 20 μL of proteinase K (20 mg/mL, Qiagen) and incubated overnight at 56 °C. DNA extraction was performed using 200 μL with the EZ1®DNA Tissue Kit on BIOROBOT EZ1 (Qiagen), according to the manufacturer's instructions. Elution was performed in 200 μL volume, dilutions to 1:10 and to 1:100 were performed.
Pure and diluted DNA extracts were screened for the presence of T. pallidum using two qPCR assays. Assays were performed in 20 μL final volume containing 10 μL of Master Mix Roche (Eurogentec, Seraing, Belgium), 0.5 μL each primer per reaction at the concentration of 20 μM, 0.5 μL UDG, 0.5 μL of each probe at the concentration of 5 μM and 5 μL of the DNA template. The amplifications were performed in a CFX96 Real-Time system (Biorad Laboratories, Foster City, USA) using the following program: 2 hold steps at 50 °C for 2 min, followed by 95 °C for 15 min and 40 cycles of 2 steps each (95 °C for 30 s and 60 °C for 30 s). DNA of T. pallidum strain DAL-1 was used as positive control and master mixture as a negative control for each reaction.
2.4. Microsatellite typing of chimpanzees
In order to identify the true number of chimpanzee individuals studied and correspondence of each sample to an individual we performed the microsatellite typing. DNA extracts were amplified by PCR targeting the Patr-B gene (~400-bp) following a standard protocol. Fragments of this size were successfully amplified in the microsatellite typing system previously applied on chimpanzee fecal DNA. Then, PCR amplification and sequencing targeting exons 2 and 3 that encode the most polymorphic and functionally relevant parts of the Patr-B gene were performed as reported previously. Exons 2 and 3 sequences were aligned using the CLUSTALW software. Phylogenetic inferences were performed with the Maximum Likelihood and Neighbor-Joining methods under the Kimura 2-parameter models and complete deletion with the MEGA 6.0 software [12]. The list of primers used is shown in Table 3.
Table 3.
qPCR systems used for T. pallidum screening according to sequences previously described.
| Gene | Name | Sequences |
|---|---|---|
| PolA | SyphT_F1 | GTCGAGACTGAAAAGGAGTGCA |
| SyphT_R1 | GTGAGCGTCTCATCATTCCAAAG | |
| SyphT_P1 | FAM-TGCTGTGCAGGATCCGGCATATGTCC-TAMRA | |
| FlaA | Tpal_flaA_F | GCGGTTGCACAGTGGGAG |
2.5. Protein extraction from stool samples
Stool protein extracts were obtained using a new protocol developed by our team [11]. This protocol combines (1) a protein extraction step with an extraction buffer consisting of a phosphate-buffered saline solution supplemented with 4-(2-aminoethyl)-benzenesulphonyl fluoride 4 mM, bestatin 0.26 mM, E-64 28 μM, leupeptin 2 μM and aprotinin 0.6 μM, pH 7.4 and SIGMAFAST™ protease inhibitor tablets (Sigma-Aldrich, St Louis, MO, USA) and (2) a concentration step of the extracted proteins. Stool protein extraction was performed using 2 g of stool or 2 mL of the stool mixture and extraction buffer. The stool-buffer mixture was incubated for 15 min at room temperature, vortexed for 1 min, then centrifuged at 1500 rpm for 15 min at 4 °C. The supernatant was lyophilised for 24 h and then resolubilised in 1 mL distilled water. Next, 500 μL of resolubilised stool was concentrated using Amicon ® Ultra-15 3 k centrifugal filters (Merck, Saint-Romain, France) for further studies.
2.6. T. pallidum antigens preparation
T. pallidum strain DAL-1 was extracted from minced rabbit testicular tissue, which was infected in the Department of Biology, Faculty of Medicine at Masaryk University (Brno, Czech Republic). We carried out the Treponema purification, from the suspension tissue following the protocol developed by Hanff et al. A Percoll density gradient centrifugation was performed to separate the Treponema cell from tissues debris, and multiple washes were done to eliminate the Percoll from the bacteria. After bacteria purification, the Treponema antigens were extracted. The Treponema suspension was centrifuged at 10,000 rpm and the pellet was suspended in TS buffer vortexed and sonicated three times (keeping it on ice between each sonication). The concentrated antigen protein was measured by Bradford protein assay and an SDS-PAGE was performed with 6 μg of the purified antigen and 12% of polyacrylamide.
The commercial recombinant proteins of Treponema, p15, p17, p47 (ABIN934736) were obtained from the manufacturer (Antibodies-Online GmbH, Aachen, Germany). This recombinant protein is obtained from the fusion of three major T. pallidum proteins (p15, p17, p47) 6,49. This is a 77 kDa molecular weight protein used for syphilis diagnostic. The SDS-PAGE was performed with a concentration of 10 μg of recombinant protein, using 12% polyacrylamide for confirmation.
2.7. Total IgA and T. pallidum specific IgA, IgG, IgM quantification
Total IgA was measured by ELISA using immunoassay kits (R&D systems and Abcam manufacturers, Cambridge, UK) according to the manufacturer's protocols. The sensitivity of the assays was 1.0 pg/mL. The quantification of T. pallidum specific IgG, IgM, and IgA was performed by ELISA according to the manufacturer's instructions. We used the IgG ELISA (ABIN2683268) and IgM ELISA (ABIN2683269) kits (Antibodies-Online, Aachen, Germany). For the detection of specific IgA, we used a T. pallidum IgG kit by replacing the secondary antibody with anti-IgA coupled to HRP, diluted 1:1000.
2.8. Western blot assays
To evaluate anti-Treponema IgA, IgM, and IgG, we performed a western blot using the Jess™ Simple Western System (ProteinSimple, San Jose, USA) as previously described [13]. We followed the manufacturer's standard method for the 12–230 kDa Jess™ separation module (SM-W004). The western blot was performed with the Treponema strain proteins (0.4 μg/μL), the recombinant protein (0.8 μg/μL), and the rabbit serum (0.4 μg/μL) (to evaluate probable non-specific detection of antibodies attachment to leftover rabbit proteins in T. pallidum cell antigens) as antigens in separate assays under the same conditions. Chimpanzee and human stool extracts were incubated for 60 min, and strips were revealed with anti-human IgG, IgA and IgM antibodies conjugated to goat HRP (Immunology Jackson ImmunoResearch, Ely, UK) diluted at 1:500 for 30 min. Finally, chemiluminescent revelations of the scale and samples were established with peroxide/luminol-S (ProteinSimple). The digital image was analyzed with Compass for SW software (v4.1.0, ProteinSimple) and quantified data of the detected proteins were reported as molecular weight, chemiluminescence intensity, and signal-to-noise ratio. We performed different performances for each antigen using separate assays.
2.9. Statistical analysis
Statistical analysis was performed using R statistical software (http://www.R-project.org). Results were expressed as median and 25–75 percentiles. The median responses for each parameter were compared by the Student or Kruskal-Wallis test, a two-tailed p-value <0.05 was calculated for each parameter.
3. Results
3.1. Field observations and identification of individual chimpanzees from stool samples
Following field observations, we could identify a total of 17 individual chimpanzees (11 adult males and 6 adults females) with orofacial depigmentation marks at Dindefelo from camera trap data and photographs obtained during direct observations from July 2013 to December 2016.
All 48 collected chimpanzee stool samples we subjected to the identification of individuals by sequencing the MHC-B partial gene. Results revealed that in some cases up to four stool samples belonged to the same individual (Fig. 3). Thus, the 48 stool samples collected in Senegal belong in fact to 29 chimpanzee individuals (Fig. 3).
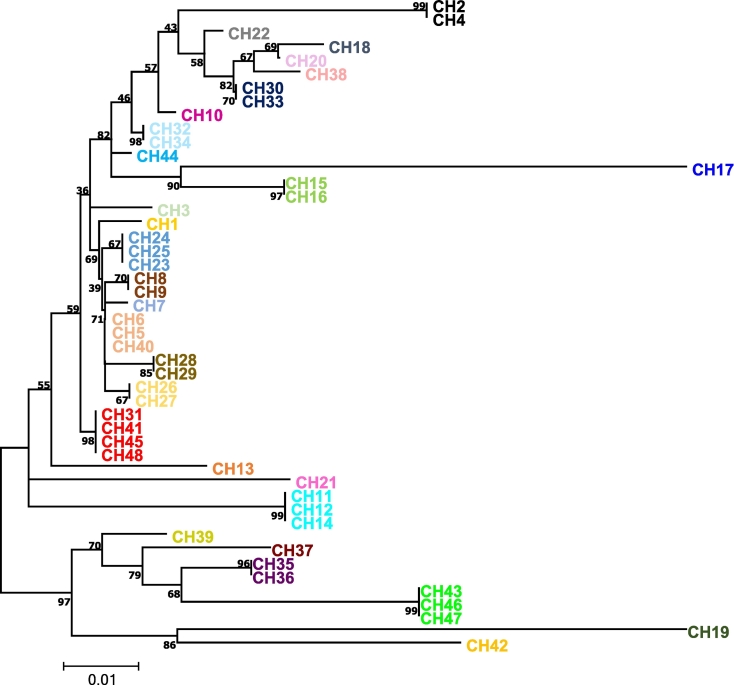
Fig. 3.
Phylogenetic tree based on partial MHC-B gene's sequencing from 48 feces of wild chimpanzees (CH from 1 to 48) allowing the identification of 29 individuals. Each color indicates an individual. The evolutionary distances were computed using the Tamura-Nei method and are in the units of the number of base substitutions per site. The differences in the composition bias among sequences were considered in evolutionary comparisons. The analysis involved 48 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. All positions containing gaps and missing data were eliminated. There was a total of 328 positions in the final dataset. Analysis were performed used MEGA 6.0..
The individuals from the zoo and the animal park are identified because the stool samples were collected during surveillance of the animals in these institutions. The 30 stool samples from the ZooParc de Beauval corresponded to 15 chimpanzee individuals. The four stool samples from La Vallée des Singes animal park corresponded to four chimpanzee individuals.
3.2. Detection of T. pallidum in chimpanzee stool samples
All stool samples were negative for T. pallidum using specific real-time qPCRs targeting two different genes in the samples.
3.3. Total and specific IgA for T. pallidum in stool samples
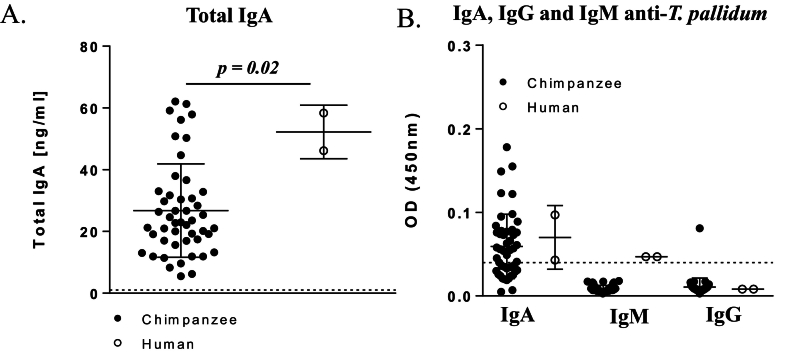
As depicted in Fig. 4A, IgA levels were detectable in all samples tested with median and IQRs of 22.88 ng/mL (17–32 ng/mL) for wild chimpanzee stools, 23.07 ng/mL (23.072–23.32 ng/ml) for captive chimpanzees, and 52.26 ng/mL (42.2–55.33 ng/mL) for human stools (p = 0.02) (Table 1).
Fig. 4.
Immunoglobulin concentrations.
(A) Evaluation of total IgA concentrations in stools of 48 chimpanzees and 2 humans by ELISA. (B) Evaluation of IgG, IgA and IgM specific for T. pallidum by ELISA. Graph representing the specific IgG, IgA and IgM (OD) level of all samples measured at a wavelength of 450 nm. A sample was considered positive for IgG, IgA and IgM when the OD value was >0.042.
Table 1.
Total IgA concentrations from chimpanzee and human stools.
| IgA (ng/ml) | n (%) | Median IQR |
|---|---|---|
| Chimpanzee | 48 (100) | 22.88 (17–32) |
| Human | 2 (100) | 52.26 (42.2–55.33) |
| Captive chimpanzee | 34 (100) | 23.07 (23.07–23.32 |
IgA concentrations were analyzed as median and interquartile ranges (IQR). n (%): number of samples in which the parameter was detected (relative frequency of detection). The stool concentration factor, median and IQR were not included in our calculations, and the median and IQR were calculated by restricting the results above the lower limit of quantitation for each analysis.
We investigated IgG, IgA, and IgM specific for T. pallidum in wild and captive chimpanzees and two human stool samples from syphilis-positive patients. The presence of all three investigated immunoglobulin classes was confirmed (Fig. 4B). Statistical analysis revealed an OD value >5 times the negative control value for this kit (positive OD > 0.042). The value we obtained also corresponds to the OD measurement of the control sample for the positivity threshold value.
Overall, 20 out of 48 (41.67%) wild chimpanzee stool samples and all human stool control samples were detected positive for specific IgA. In accordance with the identification of individual chimpanzees from the stool samples, this corresponds to 13 out of 29 individuals (44.8%). IgM were present in only human samples and were not detected in chimpanzees. In contrast, IgG were not found in human samples but only in one chimpanzee sample. All samples from the captive chimpanzee group were negative all three IgA, IgG and IgM (Fig. 4B).
Overall, these results show that almost half of the wild chimpanzees studied had immune response for T. pallidum.
3.4. Western blot identification of specific antibodies in stool samples
In order to evaluate the presence of specific antibodies against T. pallidum in chimpanzee stool samples, a Jess™ western blotting was performed using two different antigens (recombinant and extracted from T. pallidum strain). The selected antigens were first evaluated by silver stain SDS-PAGE. A single band was observed for the recombinant protein indicating the presence of a recombinant protein originating from fusion of the three treponemal proteins (p15, p17, p47), as indicated by the manufacturer (Fig. 5A). In contrast, several bands ranging from 26 to 250 kDa were observed that corresponded to the proteins from the T. pallidum strain DAL-1 (Fig. 5B), as previously reported [14,15].
Fig. 5.
Silver stain SDS-PAGE analyses for (A) recombinant and (B) T. pallidum strain DAL-1 antigens. In both profiles, the left lane corresponds to the molecular weight standard and the right lane to the antigen; 5 μg of recombinant protein and 6 μg of T. pallidum strain DAL-1 were running in a 12% polyacrylamide gel.
The presence of antibodies against T. pallidum was then evaluated in chimpanzee stool samples using the two different antigens. Among the 48 stool samples from wild chimpanzees, 20 were found positive for both antigens (Table 2). This corresponded to 13 of the 29 (44.8%) wild individuals in our cohort. On the other hand, of the two samples from the same individual (individual Z), one was positive and the other dubious, demonstrating that the degree of positivity may rely on the sample's quality (Table 2).
Table 2.
Table illustrating Ig anti T. pallidum detection by Western Blot using two different antigen sources. Each colored cell represents a stool sample that tested positive for an antigen, with a blue gradient indicating (1) low, (2) medium and (3) high intensity band. Samples were determined as positive according to the recognition for both antigens, recombinant protein and T. pallidum strain DAL-1.
Different protein profiles for the two antigen sources were identified by SDS-PAGE. The bands detected with the recombinant antigen corresponded to those described by the manufacturer (77 kDa) (Fig. 6A). The main band detected by Western Blot with the T. pallidum strain DAL-1 antigen was evaluated to attach the protein of approximately 67 kDa molecular weight (Fig. 6B). It is important to note that all samples showing bands with both antigens by western Blot also showed specific IgA levels by ELISA assay.
Fig. 6.
Immunoglobulins (IgG, IgM, and IgA) detection by western blot for 48 stool samples from 29 chimpanzee individuals living at the Dindefelo Community Nature Reserve in Senegal. (A) Recombinant antigen; (B) T. pallidum strain DAL-1 antigen.
Among the 34 chimpanzee stool samples from the ZooParc de Beauval (30 samples) and La Vallée des Singes animal park (4 samples) included as a healthy group control, no antibodies against T. pallidum were detected (Supplementary Fig. 1). All chimpanzees were born in captivity and are regularly checked by veterinary specialists. Their medical history did not report any clinically evident treponemal disease.
4. Discussion
This study underscores the application of stool serology in screening and diagnosing emerging infectious diseases in wild animals. Based on our prior research [11], we have elucidated the methodology for extracting and purifying Ig from fecal samples obtained from wild animals, accompanied by the visualization of these antibodies through automated western blotting with diverse antigens. Our approach, characterized by the use of minimal fecal quantities per sample and employing non-invasive collection techniques, offers several distinct advantages. It facilitates the unequivocal detection of specific antibodies targeting various proteins of the targeted microorganism, permits comparison of their affinities with different antigens, and enables quantification of the immune response. Although previous studies have reported the presence of specific antibodies in animal stool samples across different species, including great apes [[16], [17], [18], [19]], many of these investigations relied on non-concentrated and/or non-purified fecal samples in immunoassay or immunofluorescence-based assays, which may carry inherent limitations and the potential for generating false-positive or false-negative results.
Our research conclusively demonstrates that the proposed approach is a straightforward and robust method for definitively detecting specific antibodies in fecal samples. Even when analyzing distinct stool samples from the same individual collected in the wild (as exemplified by individuals K, O, and S in Fig. 6), we consistently observed highly similar western blot patterns, thus attesting to the method's remarkable reproducibility and reliability.
Current syphilis diagnosis in humans predominantly relies on serological methods [20,21] due to their high specificity. Alternative approaches such as bacteriological and molecular methods have limited utility, primarily because of the challenges associated with sampling bacterium-containing specimens, such as chancre biopsy. Consequently, the presence of specific anti-T. pallidum antibodies in Western chimpanzees, as demonstrated in this study, unequivocally signifies a treponemal infection that exhibits a persistent, chronic nature, akin to human treponematoses.
Remarkably, we observed elevated concentrations of total IgA in the analyzed stool samples, including those obtained from captive chimpanzees. It is well-established that IgA plays a pivotal role in maintaining immune homeostasis and fostering reciprocal interactions between gut B cells and intestinal bacteria [14,15]. Notably, we successfully detected T. pallidum-specific IgA through ELISA in 41.67% of the wild chimpanzee stool samples of 13/29 individuals, as well as in the two stool samples from known human syphilis-positive cases. These findings were further validated through western blot analysis, where we identified specific bands using both the natural antigen from the T. pallidum strain and a recombinant antigen comprising three major antigenic proteins (p15, p17, and p47) [14,15].
Although facial depigmentation in adult individuals can occur naturally in the Western chimpanzee subspecies [22] and be even the marks of leprosy naturally occurring in chimpanzee in the same region [23], we cannot exclude that this could be, at least in part, the consequence of the recovered orofacial lesions of treponematosis.
The identification of T. pallidum-specific antibodies in wild Western chimpanzees from Dindefelo serves as compelling evidence of the ongoing circulation of T. pallidum within this reserve. Our previous research had already documented treponema circulation in clinically ill monkeys within the nearby Niokolo-Koba National Park in Senegal [2], situated just north of where our current study was conducted. Drawing on indirect evidence from these prior investigations, we hypothesize that the treponemal subspecies afflicting Western chimpanzees is most likely T. pallidum ssp. pertenue, in all likelihood identical to the Fribourg-Blanc strain.
This hypothesis gains further weight from a recent report detailing a female chimpanzee in Guinea afflicted with severe leave syphilis-like lesions [5], which, in conjunction with our findings, underscores the emerging health crisis among wild Western chimpanzees. This critical issue could significantly impact the survival of this already critically endangered subspecies, especially considering the severe progression of treponemal infections and their adverse effects on fetal development [24].
With only approximately 53,000 chimpanzees remaining in West Africa [25], and this subspecies classified as critically endangered on the IUCN Red List [26], the potential spread of the documented treponemal infection across multiple populations of this subspecies could yield devastating consequences. Consequently, there is an urgent need to conduct a comprehensive screening of wild chimpanzee populations across West Africa to assess the risk of treponemal disease.
If veterinary intervention is deemed necessary, it is imperative to evaluate safe and effective measures. Intravenous benzathine penicillin is the preferred treatment for humans at all stages of treponematoses. Assessing the feasibility of safely treating unhabituated chimpanzees is crucial.
Another crucial aspect to consider is the evaluation of wild non-human primates (NHPs) as potential reservoirs for human yaws following a holistic One Health approch. The T. pallidum variant found in African NHPs closely resembles the human yaws agent, raising questions about whether NHPs represent a non-human reservoir for yaws, potentially complicating efforts at eradication. To date, there is no documented evidence of animal-to-human or human-to-animal transmission of T. pallidum in the literature.
This work is original and represents a significant contribution to the understanding of treponematoses in wild Western chimpanzees from Senegal. However, we prefer to mention some study limitations related to field work challenges especially on wild animals: (i) It would be interesting to study additional West African chimpanzees from African zoos. Information from captive populations residing in proximity to human communities raises concerns about the zoonotic potential of the identified pathogens. (ii) We do not possess any information on the age and sex of the wild chimpanzees we studied. This information could also be important. Despite its limitations, our study is to our knowledge the first to report a prevalent treponemal infection in wild apes using a new non-invasive stool serology method.
5. Conclusion
It seems clear that treponematosis in NHP, at least in West Africa, is caused by the Fribourg-Blanc strain of T. pallidum. Almost identical to multiple human yaws agents, this bacterium is also pathogenic for humans and thus its permanent circulation in NHPs may present a non-human reservoir of yaws, making its eradication questionable. Our study stands as the first to document a prevalent treponemal infection in wild apes through the visualization of T. pallidum-specific antibodies in stool samples. We strongly urge conservation organizations and researchers dedicated to safeguarding critically endangered chimpanzees in West Africa to take note of this perilous infection, enabling them to screen populations and assess the risk of the disease using a collaborative One Health approach. Furthermore, the innovative technique we have developed for detecting treponema infections in chimpanzees can be extended to other primates and wildlife for the surveillance of various infectious agents, broadening its utility in wildlife conservation efforts.
Funding
This work was supported by the French Government under the “Investissements d'avenir” (Investments for the future) program managed by the “Agence Nationale de la Recherche” (reference: 10-IAHU-03). YS, SZM, HM are supported by “Fondation Méditerranée Infection” doctoral positions. SM was supported by a post-doctoral “Fondation Méditerranée Infection” fellowship. This work was partially funded by the National Institute of Virology and Bacteriology (Programme EXCELES, ID Project No. LX22NPO5103, Funded by the European Union - Next Generation EU).
Ethical statement
The ethics committee at Masaryk University in Czech Republic approved rabbit maintenance and tissue extraction procedures carried out at the Department of Biology, Faculty of Medicine.
Author contributions
YS, SMZ and HM conducted the experiments, analyzed the data and wrote the paper. SM, JV and NO helped to conduct the experiments and participated in the writing of the paper. CS, BD, GD, DD, AB, AL, and BM contributed to the collection of samples. RAHA, LD and AB reviewed and edited the paper. MN and DŠ provided the strain. OM, DR and FF designed the project, participated in the writing of the paper and provided great support carrying out the experiments.
Declaration of competing interest
All authors declare that they have no conflict of interest with this work.
Acknowledgements
We thank La Vallée des Singes animal park personnel for help and advice. We are grateful to the Ministry of Environment and Sustainable Development of Senegal as well as to the Direction des Eaux et Forêts, Chasses et de la Conservation des Sols for permission to collect samples. From the Institute Jane Goodall Spain and Senegal, we thank Paula Alvarez Varona, Daouda Diallo and Mamadou F. Diallo for help with field work in the Dindefelo Community Nature Reserve (Senegal) and Nadia Mirghani for contributing to identification of wild chimpanzee individuals from camera trap footage.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.onehlt.2024.100694.
Appendix A. Supplementary data
: Supplementary Fig. 1. Immunoglobulins (IgG, IgM, and IgA) detection by western blot for 30 feces of chimpanzees living at the ZooParc de Beauval (1−30) and 4 at La Vallée des Singes animal park (31–34). (A) Recombinant antigen; (B) T. pallidum strain DAL-1 antigen.
Data availability
Data will be made available on request.
References
- 1.Jones K.E., et al. Global trends in emerging infectious diseases. Nature. 2008;451(7181):990–993. doi: 10.1038/nature06536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Mediannikov O., et al. Epidemic of venereal treponematosis in wild monkeys: a paradigm for syphilis origin. New Microbes New Infect. 2020;35 doi: 10.1016/j.nmni.2020.100670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Devaux C.A., Mediannikov O., Medkour H., Raoult D. Infectious disease risk across the growing human-non human primate interface: a review of the evidence. Front. Public Health. 2019;7:305. doi: 10.3389/fpubh.2019.00305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Knauf S., Liu H., Harper K.N. Treponemal infection in nonhuman primates as possible reservoir for human yaws. Emerg. Infect. Dis. 2013;19(12):2058–2060. doi: 10.3201/eid1912.130863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Mubemba B., et al. Yaws disease caused by Treponema pallidum subspecies pertenue in wild chimpanzee, Guinea, 2019. Emerg. Infect. Dis. 2020;26(6):1283–1286. doi: 10.3201/eid2606.191713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Fribourg-Blanc A., Mollaret H.H. Natural treponematosis of the African primate. Primates Med. 1969;3:113–121. [PubMed] [Google Scholar]
- 7.Sepetjian M., Guerraz F.T., Salussola D., Thivolet J., Monier J.C. Contribution to the study of the treponeme isolated from monkeys by a. Fribourg-Blanc. Bull. World Health Organ. 1969;40(1):141–151. [PMC free article] [PubMed] [Google Scholar]
- 8.Smith J.L., et al. Neuro-ophthalmological study of late yaws and pinta. II. The Caracas project. Br. J. Vener. Dis. 1971;47(4):226–251. doi: 10.1136/sti.47.4.226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Harper K.N., Knauf S. In: Primates, Pathogens, and Evolution. Brinkworth J.F., Pechenkina K., editors. Springer New York; New York, NY: 2013. Treponema pallidum infection in Primates: Clinical manifestations, epidemiology, and evolution of a stealthy pathogen; pp. 189–219. [DOI] [Google Scholar]
- 10.Baylet R., Thivolet J., Sepetjian M., Bert J. Seroepidemiological studies on primate treponematosis in Senegal. Bull. Soc. Pathol. Exot. Filiales. 1971;64(6):836–841. [PubMed] [Google Scholar]
- 11.Sereme Y., et al. Stool serology: development of a non-invasive immunological method for the detection of enterovirus-specific antibodies in Congo gorilla faeces. Microbiology. 2020 doi: 10.1101/2020.11.28.402230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Tamura K., Stecher G., Peterson D., Filipski A., Kumar S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013;30(12):2725–2729. doi: 10.1093/molbev/mst197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wu J., Haitjema C.H., Heger C.D., Boge A. A proof-of-concept analysis of carbohydrate-deficient transferrin by imaged capillary isoelectric focusing and in-capillary immunodetection. BioTechniques. 2020;68(2):85–90. doi: 10.2144/btn-2019-0058. [DOI] [PubMed] [Google Scholar]
- 14.Suzuki K., Ha S., Tsuji M., Fagarasan S. Intestinal IgA synthesis: a primitive form of adaptive immunity that regulates microbial communities in the gut. Semin. Immunol. 2007;19(2):127–135. doi: 10.1016/j.smim.2006.10.001. [DOI] [PubMed] [Google Scholar]
- 15.Woof J.M., Kerr M.A. The function of immunoglobulin a in immunity. J. Pathol. 2006;208(2):270–282. doi: 10.1002/path.1877. [DOI] [PubMed] [Google Scholar]
- 16.Seo S. Won, Sunwoo S. Young, Hyun B. Hoon, Lyoo Y.S. Detection of antibodies against classical swine fever virus in fecal samples from wild boar. Vet. Microbiol. 2012;161(1–2):218–221. doi: 10.1016/j.vetmic.2012.07.010. [DOI] [PubMed] [Google Scholar]
- 17.Nieto-Pelegrín E., Rivera-Arroyo B., Sánchez-Vizcaíno J.M. First detection of antibodies against African swine fever virus in Faeces samples. Transbound. Emerg. Dis. 2015;62(6):594–602. doi: 10.1111/tbed.12429. [DOI] [PubMed] [Google Scholar]
- 18.Pearson M., et al. Mucosal immune response to feline enteric coronavirus infection. Viruses. 2019;11(10) doi: 10.3390/v11100906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mombo I.M., et al. Detection of Ebola virus antibodies in fecal samples of great apes in Gabon. Viruses. 2020;12(12):1347. doi: 10.3390/v12121347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Morshed M.G., Singh A.E. Recent trends in the serologic diagnosis of syphilis. Clin. Vaccine Immunol. CVI. 2015;22(2):137–147. doi: 10.1128/CVI.00681-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Luo Y., Xie Y., Xiao Y. Laboratory diagnostic tools for syphilis: current status and future prospects. Front. Cell. Infect. Microbiol. 2021;10 doi: 10.3389/fcimb.2020.574806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.J. F. Oates, Primates of West Africa: a field guide and natural history, Conservation International, Arlington, 2011.
- 23.Hockings K.J., et al. Leprosy in wild chimpanzees. Nature. 2021;598(7882):652–656. doi: 10.1038/s41586-021-03968-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rac M.W.F., Revell P.A., Eppes C.S. Syphilis during pregnancy: a preventable threat to maternal-fetal health. Am. J. Obstet. Gynecol. 2017;216(4):352–363. doi: 10.1016/j.ajog.2016.11.1052. [DOI] [PubMed] [Google Scholar]
- 25.Heinicke S., et al. Advancing conservation planning for Western chimpanzees using IUCN SSC A.P.E.S.- the case of a taxon-specific database. Environ. Res. Lett. May 2019;14(6):064001. doi: 10.1088/1748-9326/ab1379. [DOI] [Google Scholar]
- 26.IUCN . In: The IUCN Red List of Threatened Species 2016: e.T15935A102327574’. International Union for Conservation of Nature. Humle T., Boesch C., Campbell G., Junker J., Koops K., Kuehl H., Sop T., editors. 2016. Pan troglodytes ssp. verus. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
: Supplementary Fig. 1. Immunoglobulins (IgG, IgM, and IgA) detection by western blot for 30 feces of chimpanzees living at the ZooParc de Beauval (1−30) and 4 at La Vallée des Singes animal park (31–34). (A) Recombinant antigen; (B) T. pallidum strain DAL-1 antigen.
Data Availability Statement
Data will be made available on request.