Abstract
Huntington’s disease (HD) is an autosomal dominant disease caused by the expansion of cytosine-adenine-guanine (CAG) repeats in one copy of the HTT gene (mutant HTT, mHTT). The unaffected HTT gene encodes wild-type HTT (wtHTT) protein, which supports processes important for the health and function of the central nervous system. Selective lowering of mHTT for the treatment of HD may provide a benefit over nonselective HTT-lowering approaches, as it aims to preserve the beneficial activities of wtHTT. Targeting a heterozygous single-nucleotide polymorphism (SNP) where the targeted variant is on the mHTT gene is one strategy for achieving allele-selective activity. Herein, we investigated whether stereopure phosphorothioate (PS)- and phosphoryl guanidine (PN)-containing oligonucleotides can direct allele-selective mHTT lowering by targeting rs362273 (SNP3). We demonstrate that our SNP3-targeting molecules are potent, durable, and selective for mHTT in vitro and in vivo in mouse models. Through comparisons with a surrogate for the nonselective investigational compound tominersen, we also demonstrate that allele-selective molecules display equivalent potency toward mHTT with improved durability while sparing wtHTT. Our preclinical findings support the advancement of WVE-003, an investigational allele-selective compound currently in clinical testing (NCT05032196) for the treatment of patients with HD.
Keywords: MT: Oligonucleotides: Therapies and Applications, Huntington’s disease, stereopure antisense oligonucleotide, allele-selective silencing, HTT, SNP3, mutant huntingtin, wild-type huntingtin, phosphoryl guanidine, SELECT-HD, WVE-003
Graphical abstract

Iwamoto and colleagues develop chemically modified oligonucleotides that selectively lower mutant huntingtin (mHTT) expression by targeting a heterozygous SNP in the transcript. In preclinical models, the molecules are potent, durable, and selective for mHTT. These findings support the advancement of WVE-003, an investigational oligonucleotide for the treatment of Huntington’s disease.
Introduction
Huntington’s disease (HD) is a progressive neurodegenerative disease characterized by motor, cognitive, and psychiatric disability.1 The onset of HD typically occurs in mid-adulthood, and individuals live an average of 15–20 years after the onset of symptoms before succumbing to this invariably fatal disease.1,2 Most therapeutic approaches for HD have focused on the alleviation of symptoms, and there are currently no approved disease-modifying treatments. HD is caused by an autosomal dominant expansion of a cytosine-adenine-guanine (CAG) triplet repeat in exon 1 of the huntingtin (HTT) gene.3 Wild-type HTT protein (wtHTT) is essential for neural development and contributes to several critical cellular functions, including transcription, apoptosis, synaptic activity for long-term potentiation (the molecular correlate for learning and memory), vesicle transport, axonal regeneration, and the formation and function of cilia involved in the clearance of brain catabolites and resorption of cerebrospinal fluid (CSF).2,4,5,6,7,8,9,10,11 Expansion of the CAG repeats (≥36 repeats) in HTT results in the production of a mutated protein (mHTT) that accumulates in the brain.2 The pathology of HD has been attributed to toxic gain-of-function activities of mHTT, which include protein aggregation, transcriptional dysregulation, and dominant negative interference with wtHTT function, as well as the loss of beneficial functions of wtHTT.12 Although HTT is ubiquitously expressed, the most acute neurodegeneration occurs in the striatum and frontal cortex.13,14
Therapeutic strategies to decrease the production of mHTT protein are currently in clinical development. These efforts are supported by preclinical data indicating the reduction of mHTT protein is associated with decreased HD pathology.15,16,17 However, most of these approaches lower both wtHTT and mHTT proteins, and there may be risk in lowering wtHTT.12,18,19,20,21 While some preclinical studies suggest a decrease of up to ∼45% wtHTT protein can be relatively well tolerated in healthy animals (in the absence of mHTT and HD), the safety of long-term suppression of wtHTT in individuals with HD remains unknown.16,22
wtHTT is both important for normal neuronal function in the adult CNS10,23,24,25 and protective against HD.20,26,27 It can protect against stress-induced neurodegeneration in multiple model systems: in cultured neurons, wtHTT is protective against stress-induced apoptosis24,28,29,30; in mice, postnatal deletion of wtHTT leads to progressive neurological phenotypes, neurodegeneration, and premature death,10 whereas overexpression of wtHTT conveys neural protection during stress, including ischemia and other types of CNS injury,21 as well as NMDA-induced excitotoxicity.23 In the YAC128 mouse model of HD, overexpression of wtHTT ameliorates striatal neuropathology,27 whereas loss of the wild-type mouse Htt worsens motor performance, survival, and striatal neuronal size.20 In patients with HD, the A variant of a non-coding single-nucleotide polymorphism (SNP) rs13102260 disrupts a binding site for the transcription factor NF-κB and decreases expression of the associated HTT gene: when the A variant associates with mHTT, disease onset is late (on average, 10 years later than when the G variant associates with mHTT); when the A variant associates with wtHTT, disease onset is earlier (on average, 4 years earlier than when the G variant associates with wtHTT), indicating that increased expression of wtHTT can be protective against HD in patients.26 Together, these studies provide strong evidence that wtHTT is both neural protective during stress and is specifically protective against HD; thus, we believe an allele-selective therapeutic, one that can diminish the production of mHTT while sparing wtHTT, may be ideal.
Multiple approaches to achieve allele-selective silencing of mHTT have been reported, including targeting the CAG-triplet repeat itself and targeting SNPs that are associated with mHTT.31 We have pursued an SNP-targeting approach using antisense oligonucleotides. We and others have reported robust allele-selective suppression of mHTT in model systems using this strategy.32,33,34,35,36 Antisense oligonucleotides are a promising treatment option in part because they can target pre-mRNA (increasing the number of targetable SNP sites), distribute throughout the brain, including the deep regions impacted by HD, and can be taken up by neuronal cell types without the assistance of a delivery vehicle.16,37 In addition, incorporation of phosphoryl guanidine-containing (PN) backbone modifications have been shown to improve the potency, durability, and distribution of stereopure oligonucleotides in the CNS.38
Herein we present the development of allele-selective stereopure oligonucleotides that precisely target SNP variants on the mHTT allele. We evaluate these oligonucleotides’ persistence in tissue, potency and duration of silencing action, and distribution to disease-relevant brain regions, including the striatum. First, we demonstrate that control over oligonucleotide backbone stereochemistry can direct the RNase H enzyme to elicit position-specific cleavage in a target RNA, which when directed at an SNP yields selective targeting of mHTT over wtHTT. We provide evidence that our design strategy may enable the expansion of disease-relevant SNPs amenable to this approach. We then demonstrate that rationally designed SNP-targeting oligonucleotides can elicit potent, durable, and selective reduction of mHTT in patient-derived neurons and two disease-relevant mouse models (BACHD and Hu97/18).39,40 Through direct comparisons with a surrogate for the nonselective HTT-lowering oligonucleotide, tominersen (herein referred to as a pan-silencer), we also demonstrate that the allele-selective molecules match the potency and exceed the durability of this pan-silencing molecule.
Results
Stereochemistry impacts allele selectivity for HTT SNPs
Antisense oligonucleotides elicit their gene silencing activity through the endogenous RNase H1 enzyme (hereafter denoted RNase H). Oligonucleotides typically incorporate chiral phosphorothioate (PS) substitutions in the phosphodiester (PO) backbone to improve their metabolic stability and cellular uptake.41 Traditional synthetic methods for producing PS-modified oligonucleotides generate stereoisomeric mixtures composed of hundreds of thousands of molecules. Because each PS linkage contains a chiral center with the potential to yield one of two configurations, denoted as Rp and Sp, the molecules within a mixture will have distinct stereochemistry, and consequently, distinct pharmacologic properties.41 Chemical modifications that alter the interface between RNase H and the heteroduplex, such as modifications to the sugar at the 2′ position in the DNA, can impact the location and efficiency of cleavage by RNase H.38,42 Recent developments have shown that the RNase H enzyme displays sequence preferences.43 We have demonstrated that RNase H activity is also impacted by stereochemistry of the oligonucleotide backbone.44
The catalytic domain of RNase H (RNase HC) contains a phosphate-binding pocket that recognizes a short stretch of the DNA backbone in a DNA-RNA heteroduplex substrate and a metal-coordinating catalytic tetrad that cleaves the RNA.45 Based on the X-ray crystal structure,46 we proposed that this preference might be driven by stereo-sensitive contacts between a DNA-binding pocket in RNase H and the PS backbone of the DNA strand of the heteroduplex; indeed, we found that a 3′-SSR-5′ stereochemical code in the antisense oligonucleotide gap (denoted SSR motif) increased RNase H activity compared with other stereopure and stereorandom configurations.44
We aimed to test whether control over PS stereochemistry could also be harnessed to enable allele-selective targeting of a heterozygous SNP. We reasoned that by positioning a single Rp PS linkage to direct cleavage to the position of an SNP, we could direct the enzyme’s catalytic domain to sample the perfectly complementary base pair (between the oligonucleotide and the target RNA, mHTT) rather than a mismatched base pair (between the oligonucleotide and the off-target RNA, wtHTT), thereby increasing the mismatch-discriminating power of RNase H47 to be focused specifically on the site of allelic variance on the target RNA. Toward this goal, we performed RNase H biochemical assays with a panel of antisense oligonucleotides with different Rp locations.
To investigate whether the positioning of a single SSR motif (i.e., a single Rp PS linkage within an otherwise all-Sp oligonucleotide) impacts the location of RNase H cleavage, we studied RNase H activity and RNA cleavage position for a stereorandom oligonucleotide and a series of stereopure oligonucleotides hybridized to a complementary surrogate RNA. These molecules all have the same sequence. One stereopure oligonucleotide contains three iterations of the SSR motif in the gap (APO-1920), and one contains an all-Sp backbone (APO-1921). Three additional stereopure oligonucleotides contain a single SSR motif, but the sole Rp linkage is placed in a different position within the gap (APO-543, APO-544, APO-545; Table S1). To assess the impact of these changes, we first hybridized the molecules to a short surrogate RNA (RNA-1, Table S2) and analyzed the rate of RNase H-mediated cleavage. All the stereopure oligonucleotides containing an SSR motif showed increased RNase H cleavage activity compared with a stereorandom control oligonucleotide, APO-424, and the all-Sp stereopure oligonucleotide, APO-1921 (Figure 1A). The extent of RNA cleavage observed with the SSR-containing stereopure oligonucleotides differed, indicating that it varied with the number and/or position of Rp.
Figure 1.
Rp linkages direct the rate, position, and selectivity of RNase H-mediated cleavage
(A) Rates of RNase H-catalyzed RNA cleavage for oligonucleotide-RNA duplexes containing stereorandom or stereopure molecules. The mean percentage of full-length RNA remaining is plotted with respect to time (minutes) for the indicated molecule (n = 3). Data are shown as mean ± SD. Data were fit to nonlinear, one-phase decay using GraphPad Prism software. (B) RNase H cleavage maps in RNA surrogate (RNA-1) when duplexed with the indicated oligonucleotide. Arrows indicate the sites of cleavage. The length of the arrows represents the percentage of product derived from cleavage at that site. The 10-min time point was used to generate cleavage maps. Legend for oligonucleotide cartoons is shown to the right. (C–E) RNase H cleavage activities are shown for the stereorandom molecule HTT-1 (C), stereopure molecule HTT-2 (D), and stereopure molecule HTT-3 (E). The mean percentage of full-length mHTT (RNA-A) and wtHTT (RNA-G) surrogate RNA remaining is plotted with respect to time (minutes) for the indicated oligonucleotide-RNA duplex. The SNP is yellow. Data are presented as mean ± SE, n = 3. PS stereochemistry is indicated by ˆ symbols (Rp = red; Sp = blue; stereorandom = black).
To determine whether the position of RNase H cleavage was affected by the position of the Rp, we characterized and measured the cleavage products from these reactions. The stereorandom oligonucleotide (APO-424) and the all-Sp oligonucleotide (APO-1921) led to the cleavage of the RNA at three adjacent backbone positions at relatively equal rates (5.0%, 4.1%, and 5.7%; and 3.1%, 2.0%, and 3.7, respectively). The stereopure SSR-containing oligonucleotides led to the preferential cleavage of the RNA at a single position. APO-1920, with three SSR motifs, predominately promoted preferential cleavage of the RNA at one position (24.1%, compared with 1.4% and 1.4% at the other positions). The oligonucleotides with a single SSR motif also led to the preferential cleavage of the RNA at a single position (Figure 1B). For APO-543 and APO-545, the most active of these, we only detected products from cleavage at one position (24.3% and 25.5%). For APO-544, we detected two minor products derived from cleavage in the two backbone positions abutting the major cleavage site (1.2%, 16.9%, and 1.4%).
These experiments demonstrate that the placement of a single Rp PS linkage embedded within an otherwise all-Sp PS gap of an oligonucleotide can impact the position of RNase H-mediated cleavage of a target RNA. Although the stereopure oligonucleotide containing three iterations of the SSR motif also yielded one preferred cleavage site, the former design limits our ability to control the preferred position of cleavage. Because the distance between a single Rp and the preferred position of cleavage can be determined, we can deploy a single Rp to define the preferred position of cleavage in an RNA backbone.
Stereochemistry promotes allele selectivity for mHTT SNP rs7685686 (A/G)
To determine whether we can use the positioning of an Rp PS linkage to control cleavage and thereby achieve selective action at an SNP, we performed a parallel set of experiments on an HTT RNA using an exemplary SNP rs7685686 (A/G). The A variant of SNP rs7685686 is associated with the expanded CAG-disease allele, whereas the wild-type nucleotide is G. We generated a series of 20mer antisense oligonucleotides: HTT-1 is a stereorandom molecule (Figure 1C); HTT-2 is a stereopure molecule with a single Rp PS linkage (Figure 1D); HTT-3 is a stereopure molecule with a single Rp PS linkage in a different position from HTT-2 (Figure 1E). To assess allele selectivity, we hybridized the oligonucleotides to RNA-A (20mer synthetic RNA corresponding to the A variant at SNP rs7685686, mHTT) or RNA-G (a 20mer RNA surrogate for the G variant at SNP rs7685686, wtHTT) and analyzed the rate of RNase H cleavage. The stereorandom molecule, HTT-1, was active against both RNA surrogates, with 28% of the mHTT substrate remaining at 60 min and 49% of the wtHTT substrate remaining at the same time (Figure 1C). HTT-1 favored the mHTT substrate ∼1.4-fold over wtHTT. HTT-2, one of the stereopure molecules, was also active against both RNA surrogates, but showed preferential activity against the mHTT substrate. Twenty-five percent of the mHTT substrate remained at 60 min, whereas 90% of wtHTT substrate remained at the same time (Figure 1D). HTT-2 favored the mHTT substrate 7.5-fold over wtHTT, a 5-fold improvement over HTT-1. HTT-3 was only weakly active against the RNA surrogates, with >80% of both substrates remaining after 60 min, indicating that it was not very active and nonselective (Figure 1E). Thus, HTT-2 had the Rp PS linkage positioned to direct the most selective RNase H activity.
Application to SNP3
We applied these findings to SNP3 (SNP rs362273 (A/G)) where the target variant (A) is in linkage disequilibrium with the expanded CAG tract in HTT. SNP3 has historically been difficult to target in an allele-selective manner.33,48,49 We designed a series of oligonucleotides, which all had the same sequence, using PS and PO backbone modifications that differed only in the placement of the Rp PS linkage38,42,44,50 with respect to the SNP. These oligonucleotides were designed to determine the optimal relationship between the Rp PS linkage and the mismatch in SNP3 to achieve allele-selective RNase H activity (Figure 2A). For HTT-956, HTT-957, and HTT-958, the Rp linkage location in the backbone varies from position 10 (P10) through position 12 (P12) when read from the 5′-end of the oligonucleotide. HTT-1233 was included as a stereorandom control and has the same sequence as the other oligonucleotides targeting SNP3. All molecules tested had asymmetric 2′-ribose modifications that were previously demonstrated to enhance the potency of antisense oligonucleotides.42
Figure 2.
PS stereochemistry can promote allele-selective activity at HTT SNP3
Schematic depicting allele-selectivity strategy. The SNP is depicted in both images. In the mHTT complex (yellow SNP), the oligonucleotide is complementary to the RNA. In the wtHTT complex (orange SNP), there is mismatch between that oligonucleotide and the RNA. (A) RNase H cleavage activity (top panels) is shown for stereorandom molecule HTT-1233 (B) and stereopure molecules HTT-956 (C), HTT-957 (D), and HTT-958 (E) where the SNP position varies between position 10 (P10) and position 12 (P12) when read from the 5′-end of the oligonucleotide. For all top panels, the mean percentage of full-length mHTT (RNA-A) and wtHTT (RNA-G) surrogate RNA remaining is plotted with respect to time (minutes) for the indicated antisense oligonucleotide-RNA duplex. Data are presented as mean ± SD. In cases where error bars are not shown, the error was smaller than the symbol. For all bottom panels, the RNA surrogates (depicted in green, 5′-3′) with the SNP shown in yellow are illustrated with respect to each oligonucleotide (shown 3′-5′).
To assess RNase H selectivity and the impact of varying the position of the Rp linkage with respect to the target SNP, we performed cell-free RNase H activity assays and determined the relative cleavage activity as well as the location of cleavage sites in the RNA. We hybridized each oligonucleotide to RNA-A SNP3 (20mer synthetic HTT RNA corresponding to the A variant, mHTT) or RNA-G SNP3 (a 20mer HTT RNA surrogate for the G variant, wtHTT) and analyzed the rate and positions of RNase H cleavage.
The stereorandom molecule, HTT-1233, was active against both RNAs, but it was not selective, with <50% of both the mHTT and wtHTT substrates remaining after 60 min (Figure 2B). HTT-956 and HTT-957, two of the stereopure molecules, were preferentially active against mHTT, with <30% of the mHTT RNA and >70% of the wtHTT RNA remaining after 60 min (Figures 2C and 2D). HTT-958, the other stereopure molecule, was active against both RNAs but was nonselective, with <15% of both substrates remaining after 60 min (Figure 2E). Ultimately, HTT-956 yielded the most selective RNase H activity. To investigate whether cleavage-site preference corresponded with the relative rates of cleavage, we evaluated the products of the RNase H cleavage reactions by mass spectrometry. HTT-1233 (the stereorandom molecule) promoted RNase H cleavage at six positions in the mHTT RNA and four positions in the wtHTT RNA with no position favored compared with the others (Figure S1). The stereopure molecules, HTT-956 and HTT-957, showed preferential cleavage within the mHTT RNA, and these preferred cleavage positions have the same relative position in the RNA with respect to the SSR motif (Figure S1). By contrast, the final stereopure molecule, HTT-958, promoted cleavage of both mHTT and wtHTT RNAs (Figure S1). These data indicate that the location of the SSR motif in HTT-956 and HTT-957 leads to more discrimination at SNP3 than the placement in HTT-958.
PN modification enhances potency and preserves selectivity of SNP3-targeting oligonucleotides in vitro
To further improve the activity of HTT-956, we introduced PN backbone linkages to the asymmetric wings.51,52,53,54,55,56 In our prior work, we evaluated the impact of stereorandom PN, Rp PN, and Sp PN linkages in the wings of antisense oligonucleotides targeting Malat1 and C9orf72 transcripts and noted the following observations: (1) PN linkages do not substantially alter thermal stability; (2) they have no discernable impact on the RNase H-induced cleavage pattern; and (3) they improve in vitro silencing activity compared with oligonucleotides that lack them, and this effect is not substantially impacted by the chirality or pattern of PN linkages.38,57 In mice, PN backbone modifications also improved the potency, durability, and distribution of silencing throughout the CNS.38 Thus, we based the designs of the SNP3-targeting oligonucleotides on these previously published molecules.
We first assessed whether the introduction of PN linkages would impact SNP-variant differentiation using our RNase H activity assay. As in prior figures, these experiments were performed using RNA surrogates representing mHTT (RNA-A SNP3) and wtHTT (RNA-G SNP3). We evaluated the rate of RNase H activity in the presence of stereopure oligonucleotides containing either three (HTT-1533) or four (HTT-1526) PN backbone linkages (Figure 3A). The position and chirality of the PN linkages were based on our prior work.38 These oligonucleotides are identical in sequence and were designed based on HTT-956. For both HTT-1526 and HTT-1533, we observed an enhanced rate of cleavage for mHTT RNA compared with that for wtHTT RNA (Figures 3B and 3C). After 60 min, <10% of mHTT substrate and >70% of the wtHTT substrate remained, suggesting that the inclusion of PN backbone linkages did not affect allele selectivity.
Figure 3.
SNP3-targeting oligonucleotides selectively and dose-dependently decreases mHTT in vitro
Schematic representation of oligonucleotides is shown in (A). RNase H experiments performed with synthetic RNA substrates corresponding to mHTT and wtHTT transcripts. Data are presented as mean ± SEM, n = 3. Mean percentages of the indicated full-length RNA substrate remaining over time are plotted for HTT-1526 (B) and HTT-1533 (C). Patient-derived motor neurons homozygous for SNP3 were treated with HTT-1526, HTT-1533, or HTT-2723 at varying levels up to 60-μM concentration for 7 days. Dose-response curves for HTT normalized to TUBB RNA measured using the bDNA assay are shown in (D). Data shown as mean ± SD, n = 4. (E) Patient-derived motor neurons heterozygous for SNP3 were treated with NTC, HTT-2723, HTT-1526, or HTT-1533 (1.6 μM, 8 μM, or 40 μM). Data depict HTT RNA remaining for wtHTT allele (blue bar) and mHTT allele (red bar). Next-generation sequencing of PCR amplicons covering the SNP3 region was performed to quantify the amount of each allele remaining after treatment. Total HTT lowering was quantified by qPCR and normalized to HPRT1. All treatment conditions were then normalized to mock treatment. Data shown as mean ± SD (mock, n = 30; HTT-2723, n = 2; HTT-1526, n = 2, HTT-1533, n = 2). ∗∗∗p ≤ 0.001, ∗∗∗∗p ≤ 0.0001. p values were calculated via White-adjusted three-way ANOVA followed by two-tailed post hoc tests comparing mutant and wild-type expression per molecule/dose allowing unequal variance.
To assess potency, we tested the activity of HTT-1526 and HTT-1533 in motor neurons derived from induced pluripotent stem cells (iPSCs) from a patient with HD who was homozygous for SNP3 (A/A). We compared the activity of HTT-1526 and HTT-1533 with a pan-silencing comparator oligonucleotide HTT-2723, which was designed based on tominersen,58 in the patient-derived neurons under gymnotic (or free-uptake) conditions. HTT-1526, HTT-1533, and HTT-2723 depleted HTT in a dose-dependent manner, yielding half-maximal inhibitory concentrations (IC50) of 3.14 μM, 1.40 μM, or 4.26 μM, respectively (Figure 3D; Table S3).
To assess selectivity, we tested the activity of HTT-1526 and HTT-1533 in iPSC-derived neurons from a patient heterozygous for SNP3 (A/G) with the A variant on the mHTT allele. We treated these heterozygous neurons with either a non-targeting control (NTC), the pan-silencer HTT-2723, or allele-selective stereopure oligonucleotides HTT-1526 and HTT-1533 under gymnotic conditions (Figure 3E). Compared with NTC, HTT-1526 and HTT-1533 potently and selectively decreased mHTT RNA. After administration of 40 μM of either molecule, <10% of mHTT transcripts remained while >70% of wtHTT transcripts remained. To evaluate allele discrimination, we ran a three-way ANOVA to test the significance of interactions among treatment, concentration, and allele. Post-hoc comparisons between mHTT and wtHTT demonstrated consistently significantly lower mHTT expression than wtHTT expression for HTT-1526 and HTT-1533 at all concentrations except HTT-1526 at 8 μM (p < 0.001). In contrast, there were no significant differences between mHTT and wtHTT expression for HTT-2723 at any concentration. At 40 μM, HTT-2723 decreased mHTT transcripts to 22% and wtHTT transcripts to 18% compared with mock. Taken together, these data demonstrate that both SNP3-targeting, stereopure oligonucleotides lead to a dose-dependent, potent, and selective decrease of mHTT RNA in vitro in patient-derived neurons under free-uptake conditions. The potency of these allele-selective oligonucleotides was roughly equivalent to the potency of the nonselective, stereorandom comparator.
We next assessed the impact of PN chemistry on lipophilicity, a property that can impact the pharmacological properties of oligonucleotide therapeutics, including their activity in the CNS.59,60 For systemically administered oligonucleotides, increasing lipophilicity can improve tissue penetration and in vivo half-life indirectly by promoting binding of serum proteins.61 Although serum-protein binding is not expected to impact the activity of CNS administered oligonucleotides, increasing lipophilicity has also been shown to improve activity in the CNS.59,60 Accordingly, we evaluated the lipophilicity of a small panel of SNP3-targeting oligonucleotides by assessing retention times on a C18 reversed-phase high-performance liquid chromatography column (HPLC). Increased retention times for the SNP3-targeting, PN-containing oligonucleotides HTT-1526 and HTT-1533 compared with the retention time for a comparable oligonucleotide HTT-956, which lacks PN chemistry, shows that the introduction of the PN linkages increases lipophilicity (Table S4), which is consistent with previously reported observations for this modification.53 The increased lipophilicity of PN-containing oligonucleotides may help to explain why this modification improved activity in the CNS.38
SNP3 targeting oligonucleotides outperform first-generation SNP-targeting oligonucleotides
We previously evaluated allele-selective oligonucleotides targeting SNP1 (rs362307) or SNP2 (rs362331) in clinical trials (NCT03225833, NCT03225846). These molecules were discontinued in 2021 because they did not consistently lower CSF mHTT levels in patients with HD (https://ir.wavelifesciences.com/news-releases/news-release-details/wave-life-sciences-provides-update-phase-1b2a-precision-hd). We also wanted to assess the potency and selectivity of SNP3-targeting oligonucleotides compared with these first-generation molecules (Figure 4A).
Figure 4.
SNP3-targeting HTT-1533 performs better in patient iPSC-derived neurons than first-generation oligonucleotides
Schematic representation of oligonucleotides is shown in (A). HTT-164 targets the U variant of SNP1 (rs362307). HTT-273 targets the U variant of SNP2 (rs362331). HTT-1533 targets the A variant of SNP3 (rs362273). Results of genotyping and phasing for iPS-109Q cells are shown in (B). The HTT gene contains both the CAG repeat sequence (yellow) and the SNP (blue box), which need to be in-phase for allele-selective silencing to work. The SNP is heterozygous (blue and green boxes), with the targeted mutant variant (blue) on same copy as the expanded CAG repeat (long yellow box). The wtHTT allele contains a non-expanded CAG tract (short yellow box) and the non-targeted variant at the SNP (green). Genotyping and phasing results for iPS-109Q cells are summarized. (C) Patient iPSC-derived neurons heterozygous for SNP1, SNP2, and SNP3 were treated with HTT-164, HTT-273, HTT-1533 or the corresponding NTC at increasing concentrations (1.6, 8.0, or 40 μM) for 7 days. Percentage of mHTT (red) and wtHTT (blue) transcript expression relative to mock-treated control cells is shown. Data shown as mean ± SEM, n = 3. Stats: three-way ANOVA with two-tailed post hoc comparisons of mHTT to wtHTT per molecule per dose assuming equal variance ∗∗∗p < 0.001; ∗∗∗∗p < 0.0001.
To perform these experiments, we needed to first identify a suitable cellular model for testing selectivity at all three SNPs. We identified iPS-109Q cells (Figure 4B) that are heterozygous for SNP1, SNP2, and SNP3 and have the desired phasing between the SNP variants and the CAG-repeat region. We evaluated the activity of HTT-164 (targets SNP1), HTT-273 (targets SNP2), and HTT-1533 (targets SNP3) in the patient-derived neurons under gymnotic (or free-uptake) conditions. At a mid-range concentration (8 μM), HTT-164 preserved wtHTT expression levels at >90% of mock-treated and led to limited lowering of mHTT expression (mean ∼15%) (Figure 4C). HTT-273 preferentially lowered mHTT levels (>25% mean lowering at 8 μM), while sparing wtHTT (∼100% mean expression at 8 μM). At the same concentration, HTT-1533 exhibited substantially greater lowering of mHTT (mean ∼90%) while preserving expression of wtHTT (>75% mean expression). Indeed, the amount of mHTT lowering observed with HTT-1533 at the lowest concentration (1.6 μM, mean 70% lowering) exceeded mHTT lowering obtained with HTT-164 or HTT-273 at the highest concentration tested (40 μM), which further supports the benefit of the asymmetric wing chemistry and the inclusion of PN chemistry, two advances in oligonucleotide design that were incorporated only in the SNP3-targeting molecules. We also evaluated the data from HTT-1533 to confirm lowering of mHTT and preservation of wtHTT compared with the NTC. At all concentrations, mHTT levels after treatment with HTT-1533 were significantly lower than in sample treated with NTC, whereas wtHTT levels were similar between the treatments (Table S5). This experiment was performed with two different probes, and there is excellent agreement between the datasets (Figures 4C and S2). Taken together, these data demonstrate the SNP3-targeting molecule more potently lowers mHTT expression than either of the SNP1- or SNP2-targeting molecules while preserving the expression of wtHTT.
SNP3-targeting oligonucleotides dose-dependently decrease mHTT in cortex and striatum of BACHD mice
To assess the activity of HTT-1526 and HTT-1533 in vivo, we used the BACHD transgenic model (Figure S3A).39 This bacterial artificial chromosome (BAC)-mediated transgenic mouse model expressing full-length human mHTT (BACHD mouse) contains 97 glutamine (mixed CAA-CAG) encoding repeats under control of the endogenous regulatory machinery.39 This human mHTT transgene also contains the targeted variant of SNP3.39 Quantitative PCR analyses demonstrated that the original BACHD mouse line had approximately five copies of the transgene.39 Because the copy number, integrity and expression of BAC transgenes can vary, with some BAC inserts comprising partial transgenes and with the potential for transgene copy number to change through generations,62,63,64 we evaluated the copy number in our BACHD colony (Figure S3B) using droplet digital PCR (ddPCR).65 The copy number varied across the transgene, with the 5′ region containing the glutamine-encoding repeat and the pan-targeting sequence at an estimated 13 copies, whereas the SNP3-containing region of the transgene was present at an estimated nine copies, indicating that while all copies of the transgene contain the pan-targeting sequences, not all copies of the transgene contain SNP3 (Figure S3B). Although the mouse Htt gene is present in this model, mouse Htt does not contain SNP3. Thus, the SNP3-targeting oligonucleotides should not bind to mouse Htt. The PCR probe set used in these experiments is human-specific, so it will only assess engagement of oligonucleotide with human mHTT.
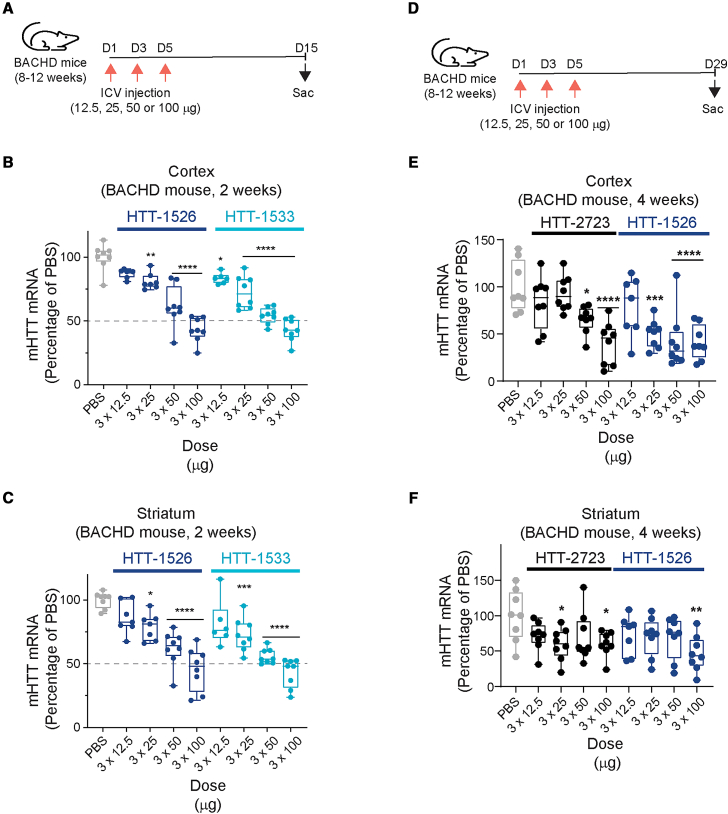
We administered phosphate-buffered saline (PBS) or oligonucleotide via three intracerebroventricular (ICV) injections on days 1, 3, and 5 at varying doses up to 100 μg in 8- to 12-week-old BACHD mice (Figure 5A). In the cortex, HTT-1526 and HTT-1533 significantly decreased the levels of human mHTT RNA in a dose-dependent manner, with 3 × 25 μg, 3 × 50 μg, and 3 × 100 μg dose levels decreasing mHTT levels to a mean of 72%–81% (p ≤ 0.01), 54%–62% (p ≤ 0.0001), and 43% (p ≤ 0.0001) compared with PBS, respectively (Figure 5B). In the same experiments, we also observed significant dose-dependent mHTT RNA decreases for both oligonucleotides in the striatum, with decreases comparable to those observed in cortical tissues. Both molecules lowered mHTT transcripts to a mean of 73%–79% (p ≤ 0.05), 56%–62% (p ≤ 0.0001), and 42%–45% (p ≤ 0.0001) compared with PBS at the 3 × 25 μg, 3 × 50 μg, and 3 × 100 μg dose levels, respectively (Figure 5C).
Figure 5.
SNP3-targeting oligonucleotides dose-dependently decrease mHTT in cortex and striatum of BACHD mice
Dosing regimen for administration of PBS or oligonucleotide (12.5 μg, 25 μg, 50 μg, or 100 μg) to BACHD mice and sample collection 15 days after first dose is shown (A). The relative fold change of human mHTT to mouse Tubb3 as a percentage of PBS in the cortex (B) and striatum (C) is shown at 2 weeks after the first administration (n = 7–8 per treatment). Data shown as mean ± SD (∗p ≤ 0.05, ∗∗p ≤ 0.01, ∗∗∗p ≤ 0.001, ∗∗∗∗p ≤ 0.0001 vs. PBS). p values were calculated via one-way ANOVA followed by two-tailed post-hoc comparisons to PBS assuming equal variance. Dosing regimen for administration and sample collection 29 days after first dose is shown in (D). The relative fold change of human mHTT to mouse Tubb3 as a percentage of PBS in the cortex (E) and striatum (F) is shown at 4 weeks after the first administration (n = 7–8 per treatment). Data shown as mean ± SD (∗p ≤ 0.05, ∗∗p ≤ 0.01,∗∗∗p ≤ 0.001, ∗∗∗∗p ≤ 0.0001 vs. PBS). p values were calculated via one-way ANOVA followed by two-tailed post hoc comparisons to PBS assuming equal variance.
We next compared the potency of an allele-selective molecule, HTT-1526, with the pan-silencer HTT-2723 (Figure 5D). Both oligonucleotides decreased mHTT RNA in a dose-dependent manner in the cortex and striatum (Figures 5E and 5F). Decreases in mHTT transcript levels for HTT-1526 and HTT-2723 in cortex and striatum were roughly comparable at the same dose (Cortex: HTT-1526 to 42% of PBS-treated mice [p ≤ 0.0001], HTT-2723 to 40% PBS [p ≤ 0.0001], Striatum: HTT-1526 to 47% PBS [p ≤ 0.001], HTT-2723 to 60% PBS [p ≤ 0.005]) (Figures 5E and 5F). We performed a similar 4-week experiment with a chemically matched NTC (Table S1) to confirm that mHTT lowering resulted from the specific interaction between the SNP3-targeting oligonucleotide and the mHTT RNA (Figure S3C). These data demonstrate that the potencies of the SNP3-targeting, stereopure molecules result from specific antisense silencing and are comparable to that of the pan-silencing, stereorandom HTT-2723 molecule in the cortex and striatum of BACHD mice.
SNP3-targeting oligonucleotides durably reduce mHTT in the cortex and striatum of BACHD mice
To determine the duration of mHTT RNA suppression by HTT-1526 and HTT-1533, we quantified mHTT transcripts remaining 2, 4, 8, and 12 weeks after the first ICV injection of either PBS or oligonucleotide to BACHD mice (Figure 6A, Study 1). We observed a significant decrease in mHTT transcripts at the earliest time point evaluated (2 weeks), and this decrease persisted for 12 weeks in both the cortex and striatum. HTT-1526 and HTT-1533 decreased mHTT transcript levels to 34%–55% (p ≤ 0.0001) and 34%–63% (p ≤ 0.0001) of PBS-treated mice in cortex and striatum, respectively, 12 weeks after dosing (Figures 6B and 6C). These data indicate that treatment with the stereopure oligonucleotides results in durable lowering of mHTT RNA in the brains of BACHD mice.
Figure 6.
SNP3-targeting oligonucleotides durably reduce mHTT in the cortex and striatum of BACHD mice
Dosing regimens and time of sample collection post-first dose for studies 1 and 2 are shown (A). Oligonucleotides or PBS were administered by ICV injection (3 × 100 μg). For study 1, the relative fold change of human mHTT to mouse Tubb3 as a percentage of PBS in the cortex (B) and striatum (C) is shown at 2, 4, 8, and 12 weeks after the first administration (n = 7–8 per treatment). For study 2, the relative fold change of human mHTT to mouse Tubb3 as a percentage of PBS in the cortex (D) and striatum (E) is shown at 2, 4, and 12 weeks after the first administration (n = 8 per treatment). For (B)–(E), data are shown as mean ± SD (∗p ≤ 0.05; ∗∗p ≤ 0.01; ∗∗∗p ≤ 0.001; ∗∗∗∗p ≤ 0.0001 compared with PBS). p values were calculated via two-way ANOVA (B–D), or White-adjusted two-way ANOVA (E) followed by two-tailed post-hoc comparisons to PBS per week with equal (B–D) or unequal (E) variance.
We also compared the duration of response for HTT-1526 to the pan-silencing comparator HTT-2723. For this experiment, we measured mHTT transcripts remaining at 2, 4, and 12 weeks after the first ICV injection of either PBS or oligonucleotide (Figure 6A, Study 2). HTT-2723 decreased HTT transcript levels to ∼62% (p ≤ 0.05) and to 43%–92% (p ≤ 0.001) in the cortex through 2 weeks and striatum through 4 weeks, respectively, compared with PBS-treated mice (Figures 6D and 6E). In the cortex, HTT-1526 decreased mHTT RNA to levels that remained significantly lower than in PBS-treated animals at all time points including 4 and 12 weeks when HTT-2723 was not significantly different from PBS at these time points. In the striatum at 12 weeks, we also observed a significant difference in activity between HTT-1526 and PBS but not HTT-2723 and PBS (HTT-1526: 51% of mHTT remaining, p ≤ 0.001; HTT-2723: 93% of mHTT remaining, p = 1.0) (Figure 6E). These data demonstrate that the mHTT lowering observed with the allele-selective, stereopure HTT-1526 molecule is more durable than that of the pan-silencer HTT-2723 in BACHD mice.
HTT-1526 has a long half-life in the cortex and striatum of BACHD mice
To better understand the kinetics, we quantified levels of HTT-1526 and HTT-2728 in tissues from experiments shown in Figure 6. The concentration of both oligonucleotides declined following first-order kinetics (Figure S4), providing half-life (t1/2) estimates in cortex (t1/2: HTT-1526 = 25.5 days, HTT-2723 = 36.3 days), and striatum (t1/2:HTT-1526 = 29.1 days, HTT-2723 = 41.4 days) (Table 1). Although the t1/2 was slightly longer for HTT-2723, the concentrations were greater. To understand pharmacodynamics (PD), PD parameters were estimated using data from experiments shown in Figures 5, 6, and S4. In both cortex and striatum, the IC50 concentration was ∼4-fold lower for HTT-1526 than HTT-2723 (IC50 cortex: HTT-1526 0.90 μg/g, HTT-2723 = 3.63 μg/g; IC50 striatum: HTT-1526 = 1.4 μg/g, HTT-2723 = 4.90 μg/g Table 1), indicating that the activity profile for HTT-1526 is driven by better potency and greater concentration compared with HTT-2723. Overall, these pharmacologic parameters are consistent with the observed potency and durability of activity for HTT-1526.
Table 1.
SNP3-targeting oligonucleotide PK/PD parameters based on cortical and striatal tissue of BACHD mice
| Parameters | HTT-1526 |
HTT-2723 |
||
|---|---|---|---|---|
| Cortex | Striatum | Cortex | Striatum | |
| t1/2 (days) | 25.5 | 29.1 | 36.3 | 41.4 |
| IC50 (μg/g) | 0.90 | 1.4 | 3.63 | 4.90 |
| Kout (1/day) | 0.36 | 0.63 | 0.53 | 0.16 |
| Imax | 0.63 | 0.80 | 0.68 | 1.00 |
t1/2: half-life; IC50: Half-maximal inhibitory concentration observed in BACHD mice; Kout: first-order rate constant for loss of response; Imax: Maximum inhibition.
SNP3-targeting oligonucleotide selectively decreases mHTT in the cortex, striatum, and hippocampus of Hu97/18 mice
To further characterize the activity of SNP3-targeting oligonucleotides in vivo, we evaluated activity in the Hu97/18 mouse model for HD, which is a fully humanized transgenic mouse model.40 It was generated by crossing BACHD mice39 with YAC18 mice66 mice on the Htt−/− background.40 The YAC transgene in the YAC18 mice contains 18 glutamine-encoding repeats and lacks the targeted variant of SNP3 (Figure 3A).40 Thus, when co-expressed, the BACDH and YAC18 transgenes are heterozygous for the expanded CAG repeat and SNP3, with appropriate phasing between these elements to evaluate allele-selective mHTT silencing.
To evaluate allele-selective activity in vivo, we quantified HTT RNA remaining at 4, 8, and 12 weeks after the first ICV injection of either PBS or oligonucleotide administered to Hu97/18 mice at 8–12 weeks of age (Figures 7A and S4D–S4F). In all three evaluated tissues, we observed a statistically significant difference between mHTT and wtHTT expression after HTT-1533 administration at every time point evaluated (4 weeks: mHTT decreased to 21%–37% of the level of PBS, wtHTT 75%–81% [p ≤ 0.001]; 8 weeks: mHTT 38%–69%, wtHTT 96%–108% [p ≤ 0.01]; 12 weeks: mHTT 51%–78%, wtHTT 87%–103% [p ≤ 0.0001, except in striatum where p = 0.404) (Figures 7B–7D; Table S6). These data showed a significant interaction between treatment and allele type in cortex, striatum, and hippocampus (p < 0.001), indicating that HTT-1533 but not HTT-2723 differentiated between wtHTT and mHTT in all tissues evaluated. These results demonstrate that treatment with the SNP3-targeting stereopure molecule leads to a selective decrease of mHTT RNA across tissue types that is durable, with significant allele discrimination maintained through 12 weeks.
Figure 7.
SNP3-targeting oligonucleotides selectively decrease mHTT RNA and protein in the CNS of Hu97/18 mice
Dosing regimen and timing of sample collection post-first dose for administration of PBS or oligonucleotide (3 × 100 μg) to Hu97/18 mice is shown in (A). The relative fold change of human mHTT (red) or wtHTT (blue) transcript to mouse Tubb3 as a percentage of PBS in the cortex (B), striatum (C), and hippocampus (D) are shown for tissue collected 4, 8, and 12 weeks after the first administration (n = 6–8 per treatment). Data are shown as mean ± SD. ∗∗p ≤ 0.01, ∗∗∗p ≤ 0.001, ∗∗∗∗p ≤ 0.0001. p values were calculated via three-way ANOVA (B and C) or White-adjusted three-way ANOVA (D) followed by two-tailed post-hoc comparisons of wtHTT to mHTT at each time point with equal (B and C) or unequal (D) variance. Representative western blots from cortex collected 4, 8, and 12 weeks after the first administration are shown to the left (E). The fold change of human mHTT protein (yellow) or wtHTT protein (dark blue) with respect to mouse vinculin protein in the cortex is shown (n = 7–8) to the right (E). Data are shown as mean ± SD. p values were calculated via three-way ANOVA followed by two-tailed post-hoc comparisons of wtHTT and mHTT at each time point with equal variance, ∗p ≤ 0.05, ∗∗∗p ≤ 0.001. Uncropped western blots are shown in Figure S5.
By contrast, we observed no statistically significant effect on allele for HTT-2723 in the cortex, striatum, or hippocampus at any time point evaluated (4 weeks: mHTT decreased to 41%–59% compared with PBS, wtHTT 42%–62% p = 1.0; 8 weeks: mHTT 65%–95%, wtHTT 66%–94% p = 1.0; 12 weeks: mHTT 88%–99%, wtHTT 86%–93% p = 1.0) (Figures 7B–7D). This result is consistent with the pan-silencing mechanism of HTT-2723, which involves targeting a region of the HTT RNA that is shared between the wtHTT and mHTT alleles.
In an alternative analysis, we evaluated the changes in expression compared with PBS control. For HTT-2723, significant lowering of mHTT compared with PBS was always accompanied by significant lowering of wtHTT in all three evaluated tissues, with lowering of wtHTT and mHTT enduring below control levels to 8 weeks but not 12 weeks in cortex and hippocampus and to 4 weeks in striatum (Table S6). By contrast, HTT-1533 significantly lowered wtHTT compared with PBS only in cortex and hippocampus at 4 weeks, despite significant lowering of mHTT through at least 12 weeks in these tissues and through at least 8 weeks in striatum (Table S6). This analysis demonstrates that HTT-1533 but not HTT-2723 is selective for mHTT, and HTT-1533 has a more durable effect on silencing.
SNP3-targeting oligonucleotides suppress mHTT protein in cortex of Hu97/18 mice
To investigate mHTT lowering at the protein level, we quantified soluble, full-length human wtHTT and mHTT proteins by western blot from cortical tissues collected during the experiment shown in Figure 7 (Figure S5). For HTT-1533, mHTT protein was decreased to 18% (p < 0.0001) at 4 weeks and remained below 43% through 12 weeks compared with PBS (p < 0.05). In contrast, wtHTT protein levels remained >59% at 4 weeks (p = 0.12) and >74% by 12 weeks compared with PBS (p = 1) (Figure 7E). These observations are consistent with the significant difference between mHTT and wtHTT RNA levels seen at weeks 4 and 8 in HTT-1533-treated animals. Following HTT-2723 treatment, both mHTT and wtHTT proteins were lowered to 17%–34% of PBS at 4 weeks (p < 0.001), and by 8 weeks, there was no longer a significant decrease of either protein compared with PBS (Figure 7E, wtHTT: p = 0.586, mHTT: p = 0.210). As expected based on these observations, there was no significant difference between mHTT and wtHTT protein levels in HTT-2723-treated animals (Figure 7E, p = 1.0). These results are consistent with the known pan-silencing mechanism for HTT-2723 and the allele-selective silencing mechanism for HTT-1533, and they confirm that selective silencing observed at the RNA level is recapitulated at the protein level in mice.
Discussion
Herein, we illustrate how control over chirality of the PS backbone can be applied to elicit allele-selective silencing by directing the activity of RNase H to the mismatched base pair in a duplex formed between an oligonucleotide and transcript with a 1-base pair mismatch (in this case wtHTT). Although others have reported allele-selective antisense oligonucleotides that promote the selective lowering of mHTT by SNP targeting,33,67 these approaches were suitable for SNPs appearing in permissible sequences but not more challenging sequences, including the SNP described herein.33,48 While chemical modification to oligonucleotides, such as the introduction of constrained ethyl (cEt) or 2′-O-methoxyethyl (MOE) modifications, reportedly impacted the activity and tolerability of molecules,33 no governing principles have been defined to aid the design of antisense oligonucleotides for an allele-selective approach. With the ability to control the chemical and stereochemical features of an oligonucleotide, we have defined design principles that allowed us to develop potent and allele-selective molecules targeting multiple sequences, including a reportedly difficult SNP sequence.33,48
We leveraged our ability to synthesize stereopure PS and PN oligonucleotide backbone linkages to enable allele-selective mHTT lowering by targeting a SNP. By optimally placing the Rp PS linkage, we can achieve selectivity for a target RNA (which is fully complementary to the oligonucleotide), while largely sparing the off-target RNA (which is mismatched to the oligonucleotide at the SNP). We applied this strategy, in combination with our PN chemistry,38 to design the stereopure, antisense oligonucleotides HTT-1526 and HTT-1533 with allele-selective activity at HTT SNP3. These PN-containing, allele-selective oligonucleotides potently decreased the expression of mHTT RNA-containing SNP3, while largely sparing wtHTT in vitro, in both biochemical and cellular assays, and in vivo. Importantly, the activity benefits of PN chemistry are evident in comparison with prior generation allele-selective molecules HTT-164 (WVE-120101) and HTT-273 (WVE-120102) that were discontinued because they did not consistently lower CSF mHTT levels in patients with HD in clinical trials (https://ir.wavelifesciences.com/news-releases/news-release-details/wave-life-sciences-provides-update-phase-1b2a-precision-hd), which further supports the activity benefits conveyed by the inclusion of PN chemistry in the SNP3-targeting molecules.
HTT-1526 and HTT-1533 also led to dose-dependent lowering of human mHTT transcripts in BACHD mice that was comparable in potency and duration to the tominersen surrogate. Unlike the surrogate for tominersen, a stereopure SNP3-targeting molecule selectively lowered mHTT in Hu97/18 mice at the RNA and protein level, while leaving wtHTT protein and RNA expression relatively intact. Importantly, this gain in selectivity does not come with a loss of potency or durability, as the decrease in expression of mHTT RNA and protein expression in response to the allele-selective molecules matched or exceeded the tominersen surrogate in all experiments. Accordingly, we have advanced HTT-1526 (WVE-003) into clinical development to evaluate its potential as a disease-modifying therapy for the treatment of individuals with HD. WVE-003 will enable us to test the hypothesis that the selective lowering of mHTT and relative sparing of wtHTT will provide a safe and efficacious therapeutic option for HD.
Others have provided evidence to support the general hypothesis that lowering of mHTT RNA and protein can impact the progression of HD in preclinical models.16,17,68,69,70,71 In a conditional mouse model for HD in which expression of the disease-causing transgene can be turned off with small-molecule analogs of tetracycline, administration of a “gene-off” small molecule after the onset of disease-like symptoms arrested neuropathology and reversed motor dysfunction (i.e., progressive clasping of the limbs).68 In multiple transgenic mouse models for HD, partial lowering of striatal or striatal and cortical mHTT expression via RNA interference (RNAi) improved neuropathology, as well as motor function in behavioral assays.68,69,70 Similarly, depletion of mHTT expression by infusion of an antisense oligonucleotide to the CNS improved neuropathology and motor function in symptomatic BACHD and YAC128 mouse models for HD.16 In addition, AAV-delivered zinc finger protein transcription factors targeting expanded CAG repeats selectively repressed mHTT RNA expression, and this ameliorated disease phenotypes in three animal models for HD.71 Hence, multiple, independent experiments support the hypothesis that decreasing the expression of mHTT after the onset of symptoms can slow or stop the progression of HD in preclinical studies.
Some insight into the impact of nonselective lowering of HTT in patients with HD was provided through the recent clinical data readouts for tominersen, a pan-silencing antisense oligonucleotide. While no major safety concerns were reported during the phase 1/2 testing period,58 in the subsequent larger phase 3 study an unfavorable benefit-risk profile emerged, which led to premature discontinuation of dosing and study termination.72 While the exact cause of these outcomes cannot be disentangled, the sponsor has acknowledged that activity related to loss of wtHTT may have been a factor.72,73
In 2021, we initiated clinical development of investigational WVE-003, which is currently being tested in the SELECT-HD phase 1b/2a clinical trial (NCT05032196). Our precision approach may provide a personalized treatment option for patients with HD and SNP3. To support patient enrollment, we assess each patient’s genotype using a method that links the SNP3 variant to the expanded CAG allele to determine their eligibility for SELECT-HD.74 Based on literature reports, we anticipate up to 40% of the HD population will be eligible for SELECT-HD.33,49,75 Early data from the ongoing SELECT-HD study (https://ir.wavelifesciences.com/news-releases/news-release-details/wave-life-sciences-announces-positive-update-phase-1b2a-select) provide preliminary evidence that WVE-003 may be lowering mHTT in patients while preserving wtHTT expression. An allele-selective strategy that spares wtHTT, preserving its neuroprotective activity, may represent the best option for delivering a disease-modifying therapy.
Materials and methods
Oligonucleotides
We obtained PO-based and stereorandom PS-modified oligonucleotides from Integrated DNA Technologies or by using a standard solid-phase oligonucleotide synthesis protocol. We synthesized and purified chemically modified, stereopure oligonucleotides as described with minor modifications.38,44 We characterized stereopure oligonucleotides by LC-HRMS and HPLC. The sequences of all oligonucleotides used in this study are shown in Table S1.
RNase H activity assays and cleavage-site analysis
These experiments were performed based on previously reported methods.38,44 In a 96-well plate, we combined 25 μL DNA/RNA duplex (20 μM), 10 μL of 10X RNase H buffer (500 mM Tris-HCl, 750 mM KCl, 30 mM MgCl2, 100 mM DTT), and 55 μL water and incubated the mixture at 37°C for 10 min. Then, we added 10 μL of RNase HC solution for a final substrate:enzyme concentration of 5 μM:0.01 μM (500:1) for RNase H assays with oligonucleotides targeting surrogate RNA and 5 μM: 0.005 μM (1,000:1) for oligonucleotides targeting SNP rs7685686. For oligonucleotides targeting SNP rs362273, a final substrate: enzyme ratio of 100:1 was used. We incubated this reaction mixture at 37°C and quenched it at different time points using 8 μL of 500 mM ethylenediaminetetraacetic acid disodium (Na2EDTA) solution in water. Na2EDTA solution was added to the reaction mixture before the addition of enzyme for the 0-min time point. Ten microliters of each reaction mixture was injected onto an Agilent ESI-TOF mass spectrometer coupled to an Agilent 1290 HPLC system with a UV detector. UV absorbance was recorded at 254 nm and 280 nm. Peak areas from the 254-nm chromatograms, corresponding to full-length RNA oligomer, were integrated and normalized against the ASO. The amount of RNA at 0 min time point was considered as 100%, and the remaining RNA at all other time points was plotted against time to show relative rates of RNA cleavage (n = 3). Error bars in graphs indicate standard deviation.
We characterized the products of the RNase H assay by LC-MS to determine the cleavage sites. The extent of cleavage at each site was determined by relative ratio of the UV peak area of a fragment to the extinction coefficient (http://www.scripps.edu/california/research/dna-protein-Research/forms/biopolymercalc2.html) of that fragment normalized to that of ASO (ratio of UV peak area of ASO to extinction coefficient of the ASO). Normalization to control oligonucleotides was performed to account for variation in the amount of duplex used across reactions. The normalized ratio of each RNA fragment was then expressed as a percentage of the total normalized RNA (RNA fragments and uncleaved RNA) present in the reaction. The percentage amount of fragment (preferably 3′-OH RNA cleavage product or 5′-phosphorylated RNA cleavage product if former was not detected) was used to determine extent of cleavage at a particular site.
Thermal denaturation
Equimolar amounts of oligonucleotide and RNA were combined and annealed in 1× PBS (pH 7.2) to obtain a final concentration of 1 μM of each strand (3 mL). UV absorbance at 254 nm was recorded at intervals of 30 s as the temperature was raised from 15°C to 95°C at a rate of +0.5°C per min, using a Cary Series UV-Vis spectrophotometer (Agilent Technologies). Absorbance was plotted against the temperature, thermal denaturation (Tm) values were calculated by taking the first derivative of each curve.
Lipophilicity
Method 1
HPLC analysis was performed using ACQUITY Premier Oligonucleotide C18 Column, 130 Å (1.7 μm, 2.1 × 100 mm) (Part number 186009485) at 55°C using buffer A (100 mM HFIP, 10 mM TEA) and mobile phase B (50% water, 50% acetonitrile) as eluents using the gradient in Table S7 with UV absorbance at 260 nm.
Method 2
HPLC analysis was performed using ACQUITY UPLC Oligonucleotide BEH C18 Column, 130Å (1.7 μm, 2.1 mm × 500 mm) (Part number 186003949) at 55°C using buffer A (100 mM HFIP, 10 mM TEA) and mobile phase B (100% acetonitrile) as eluents using the gradient in Table S8 with UV absorbance at 260 nm.
iPSC-derived neurons
iPSC-derived neurons from patients with HD (homozygous and heterozygous for SNP3) ND40536-1 iPSC-derived motor neurons homozygous for SNP3 and ND50036 iPSC-derived medium spiny neurons heterozygous for SNP3 in phase with the mHTT allele were obtained from BrainXell. iPSC-derived medium spiny neurons were seeded in 1:1 DMEM/F12 (Life Technologies) + Neurobasal medium (Life Technologies) supplemented with 1x B27 (Life Technologies), 1x N2 (Life Technologies), 0.5 mM GlutaMAX (Life Technologies), 1x seeding supplement (BrainXell), 10 ng/mL BDNF (Peprotech), 10 ng/mL GDNF (Peprotech), 1 ng/mL TGF-B1 (Peprotech), and 15 μg/mL Geltrex (Life Technologies). On day 4 post seeding, media was changed to include day 4 supplement (BrainXell) without the addition of seeding supplement and Geltrex. iPSC-derived motor neurons were cultured in media as above with the addition of Supplement K (BrainXell). On day 4 post seeding, oligonucleotides were gymnotically delivered for 7 days.
Quantification of RNA
bDNA assay
To determine in vitro IC50, ND40536-1 iPSC-derived motor neurons were plated on 384-well PDL-coated plates (Corning) and treated with oligonucleotides under gymnotic conditions. On day 7 after treatment, cells were lysed, and mRNA was quantified using a QuantiGene Singleplex branched DNA (bDNA) assay (Thermo Fisher Scientific) following the manufacturer’s instructions. HTT mRNA was quantified using bDNA probes set HTT (SA-50339) and normalized to TUBB (SA-25525).
Allele selectivity
To measure in vitro allele selectivity, ND50036 iPSC-derived medium spiny neurons were plated on PDL-coated 96-well plates (Corning) at 30,000 cells/well and treated with oligonucleotides for 7 days. Next-generation sequencing (see below) of amplicons generated by SNP3-specific PCR was performed to quantify the amount of each allele remaining after treatment.
The treated cells were harvested, and total RNA was extracted and purified followed by the cDNA production using High-Capacity cDNA Reverse Transcription kit (Thermo Fisher Scientific). The amplicons are then generated by PCR using multiplexed primers. The SNP3 or SNP2 primers with linker sequence to barcode kits (Table S9) were pre-mixed to make 10x working stock at 5 μM of each primer. The PCR reaction mixture was generated as follows: cDNA – 2 μL, Distilled water – 8.8 μL, HF Buffer – 4 μL, 10 mM dNTP – 0.4 μL, Forward primer – 2 μL, Reverse primer – 2 μL, DMSO – 0.6 μL, Phusion Polymerase – 0.2 μL. Following Touchdown PCR protocol was then used: 98 C × 30 s, (98 C × 10 s, 74 C × 30 s) × 5, (98 C × 10 s, 72 C × 30 s) × 5, (98 C × 10 s, 70 C × 30 s) × 5, (98 C × 10 s, 60 C × 15 s, 72 C × 30 s) × 25 cycles, 72 C × 5 min, 4 C forever.
Next-generation sequencing
After PCR, 2 μL of PCR samples were assessed by Fragment Analysis (AATI, DNF-900, 35-500bp) TapeStation (Agilent) for quality control. Bar coding for next-generation sequencing using Invitrogen Platinum II Taq Hot-Start DNA Polymerase and Nextera XT Index Kit v2 Set A-D (FC-131-2001 to FC-131-2004, Illumina) was performed with the following PCR protocol: Primary PCR template – 2 μL, N7xx Index – 2 μL, N5xx Index – 2 μL, 10 μM dNTP – 0.4 μL, PCR Buffer – 4 μL, Taq Polymerase – 0.16 μL, GC Enhancer – 4 μL, Distilled water – 5.44 μL. The following PCR protocol was then used for barcode attachment: 94 C × 2 min (94 C × 30 s, 55 C × 30 s, 72 C × 30 s) × 25 cycles, 72 C × 5 min, 4 C forever. After bar-coding PCR, amplicons were purified by Agencourt AMPure XP beads according to manufacturer’s protocol. Two microliters of final pure amplicon samples were quality-controlled and quantified by Fragment Analysis (AATI, DNF-473, 1-6,000 bp) TapeStation (Agilent). Equal molar of amplicons was mixed and submitted to MiSeq pair-ended 150 bp read with PhiX spike-in (10%). Knockdown of total HTT was measured by qPCR analysis. The allelic ratio of mHTT to wtHTT was normalized to percent total HTT remaining quantified by qPCR after treatment to determine allele-selective knockdown.
qPCR
Total RNA was Trizol extracted followed by purification using the RNeasy 96 Kit (Qiagen). cDNA was produced using the High-Capacity cDNA Reverse Transcription kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. qPCR was performed in the CFX System using iQ Multiplex Powermix (Bio-Rad Laboratories) and the following probes HTT: Hs00918174_m1 (Thermo Fisher Scientific) and HPRT1: Hs02800695_m1 (Thermo Fisher Scientific).
IC50 calculations in vitro
IC50 values were determined by four-parameter curve fitting of oligonucleotide concentration vs. percentage of mRNA remaining.
Animals and ICV injections
Unless otherwise noted, animal experiments were performed at Biomedical Research Models, Inc. dba Biomere (Worcester, MA) in compliance with Biomere’s Institutional Animal Care and Use Committee guidelines for the care and use of animals. Mice were on a 12-h light-dark cycle. Food (lab diet 5001) and water were available ad libitum. Housing rooms were maintained at 20°C–26°C and relative humidity was 30%–70%. The transgenic lines BACHD (bacterial artificial chromosome-mediated transgenic Huntington’s disease, Jackson Laboratory Cat. No. 008197) and Hu97/18 used for in vivo studies have been described previously.39,40
Male and female mice were 8–12 weeks of age at the time of dosing. Mice were balanced across treatment groups by gender and age. For ICV cannulation, mice were anesthetized using avertin, were placed on a rodent stereotaxic apparatus, and were implanted with a stainless-steel guide cannula in the right lateral ventricle (coordinates: −0.3 mm posterior, +1.0 mm lateral, and −2.2 mm vertically from bregma), which was secured in place using dental cement. Mice were allowed a 1-week recovery period prior to dosing.
ddPCR
Genomic DNA was extracted from two BACHD mouse tails (BACHD-1 and BACHD-2) and HEK cells using PureLink Genomic DNA mini kit (Thermo Fisher Scientific) according to the manufacturer’s protocol. The purified genomic DNA was then treated with HindIII-HF (New England BioLabs) at 37o C for 30 min according to the manufacture’s protocol. The ddPCR was performed by MOgene (St. Louis, MO) using a QX200 AutoDG Droplet Digital PCR System (Bio-Rad Laboratories), with ddPCR Supermixes for Probes (no dUTP) (Bio-Rad Laboratories) and the probe sets listed in Table S10 ddPCR was performed using eight HTT probe sets and two human-specific probes (hMalat1-FAM and hRPP30-HEX) on genomic DNA from HEK cells as a control, and eight HTT probe sets and two mouse-specific probes (mRPP-HEX and mTRFC-Hex) on genomic DNA from BACHD mice. The copy number for each probe set, corresponding to various locations along the HTT BAC clone, were normalized to the average counts of mRPP and mTRFC from the samples.
Quantification of oligonucleotides by hybridization ELISA
Cortex, striatum, and hippocampal tissues were processed and quantified as previously described.38 Twenty-five to 45 μL of tissue lysate was saved at −80°C in 96-well plates for pharmacokinetic (PK) measurement. The remaining Trizol lysates were used for RNA extraction. The same Taqman probes used for ALS motor neurons were used for qPCR. The following probes were designed to selectively quantify the oligonucleotide used in this study by hybridization ELISA: capture probe: “WV-21522-CAP”/5AmMC12/GCGG+TGGCG+A; detection probe: “WV-21522-Det”: G+TGGG+TG+AGT/3BioTEG/. Samples were read on the Molecular Device, M5 fluorescence channel plate reader: Ex435 nm, Em555 nm. The oligonucleotide in samples were calculated according to standard curve by 4-parameter regression.
In vivo IC50 and half-life calculations
The in vivo IC50 values and half-life (t1/2) were estimated from PK and PD data from experiments performed in BACHD mice (dose-response and duration studies results were pooled). A one compartmental PK model with first-order absorption and elimination was used to describe the HTT-1526 concentration in both cortex and striatum for the mouse BACHD model. The PK model was fitted to the individual HTT-1526 concentration data for cortex and striatum and the half-life (t1/2) was estimated using a naive pooled approach in Phoenix NLME 8.3 (Certara USA, Inc., Princeton, NJ, USA). An indirect response PK/PD model characterizing the relationship between HTT-1526 concentration and the remaining mHTT mRNA relative to PBS control group for estimation of IC50 were developed for cortex and striatum from dose-response and duration studies using a naive pooled approach in Phoenix NLME 8.3 (Certara USA, Inc., Princeton, NJ, USA).
HTT protein lysate preparation and western blot
HTT protein quantitation was performed using traditional western blot methods. Briefly, dissected cortical tissue from each animal was pulverized and lysed in RIPA buffer with proteinase inhibitor (Roche). The amount of protein was quantified using a BCA assay (Pierce/Thermo Fisher Scientific) according to the manufacturer’s instructions, and an equivalent amount of protein for each sample was added to NuPAGE LDS sample buffer containing reducing agent (Thermo Fisher Scientific). All protein samples were resolved using 3%–8% gradient Tris-Acetate 1.5 mm gels and cassette system (Thermo Fisher Scientific) at a constant 120V for 3 h while on ice, and the polyacrylamide gel was transferred to nitrocellulose membrane using the iBlot 2 gel transfer device (Thermo Fisher Scientific) at 25 V for 10 min according to the manufacturer’s manuals, prior to incubation with TBS blocking solution (Licor) for 1 h. The following primary and secondary antibodies were used: rabbit anti-HTT EPR5526 (1:1,000, Abcam), mouse anti-vinculin MA5-11690 (1:5,000, Thermo Fisher Scientific), IRDye 800CW donkey anti-rabbit IgG (1:10,000, Licor), and IRDye 680LT goat anti-mouse IgG (1:10,000, Licor). Primary antibody incubation was done at 4o C overnight, followed by three washes of TBS+0.05% Tween 20. This was followed by secondary antibody incubation for 1 h at room temperature. After three washes of TBS, the blot was then imaged by the ODYSSEY CLx Imaging System, and the fluorescent intensity of protein bands was analyzed with Image Studio Lite version 5.2 software. The normalized protein expression was analyzed by dividing band intensities of each HTT band (wtHTT and mHTT) by the vinculin band intensity. The amount of wtHTT and mHTT from the PBS-treated group of animals were averaged and used to generate fold change of wtHTT and mHTT amount in each treatment group at various time points.
Statistical analyses
Statistical analyses were run via the R computing environment (v 3.6.0)76 and the KNIME (Konstanz Information Miner) platform (v 4.5.0).77 For all analyses, assumptions of equal variance and normality were tested using Levene’s tests across all experimental factors and Shapiro-Wilk tests of model residuals, respectively (R package rstatix, v 0.7.0.999).78 For multi-factor experiments, type III two- and three-way ANOVAs were run to investigate all possible interactions between factors (R package car, v 3.0–7). In the event of unequal variance (Levene’s test p < 0.05), Welch’s one-way ANOVAs were run in place of typical one-way ANOVAs, while two- and three-way ANOVAs were White-adjusted to allow for unequal variance. To compare groups in each experiment, two-tailed post hoc tests were extracted from the relevant regression model via the multcomp R package (v 1.4–13) for pairwise comparisons or comparisons to a control group. Post hoc p values were Bonferroni-corrected for multiple hypotheses. For experiments featuring unequal variance, robust HC3 covariance estimation was implemented in post hoc comparisons to allow heteroscedasticity (R package sandwich, v 2.5-1).79,80 For experiments featuring mutant and wild-type allele expression, two analyses were run: one comparing each allele’s expression to its designated control (NTC/PBS) per factor level and one directly comparing mutant and wild-type expression per treatment category for relevant compounds.
Data and code availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplemental information.
Acknowledgments
The authors are grateful to UCLA for the BACHD line developed by Xiangdong W. Yang; Dr. Amber Southwell for the Hu97/18 mouse model; Susovan Mohapatra (Wave Life Sciences) for early work on allele-selective targeting; Jeff Brown and Shaunna Berkovitch (Wave Life Sciences) for early work on SNP3 molecules; Tomomi Kawamoto and Megan Cannon (Wave Life Sciences) for support with mouse experiments and tissue processing; Alexandra Walen, Allison Molski, Brooke Koshel and Frank Favaloro (Wave Life Sciences) for HPLC analyses to assess lipophilicity; and Brittany Mayweather Amy Donner (Wave Life Sciences) and Eric Smith for writing and figure generation. This work was funded by Wave Life Sciences.
Author contributions
N.I. and C.V. designed chemistry and stereochemistry of the oligonucleotides. N.I. and J.D.S. designed and performed the in vitro biochemical RNase H experiments. Y.L., A.M., A.A., and A.Z. designed and performed the in vitro cellular experiments. Y.L., M.F.-K., A.M., W.C.T., K.T., J.M., and E.P.-E. designed and performed the in vivo experiments. N.K., W.C.T., and H.U. performed the western blots. F.L., Q.P., H.Y., and Y.Y. quantified oligonucleotides in mouse tissues and performed next-generation sequencing. P.K., K.B., R.L., M.S., and S.S. led synthetic chemistry and process development efforts enabling synthesis of PN-containing stereopure oligonucleotides and formulation of oligonucleotides for application in this work. H.Y., M.B., K.L., E.D., and C.V. coordinated and supervised the project. N.I., Y.L., A.M., W.C.T., H.Y., H.G.J., M.B., E.D., and C.V. interpreted the data. A.L., K.L., and P.S.P. performed statistical analysis of the data. H.G.J. and X.S.H. performed modeling to calculate in vivo PK parameters for WVE-003. N.I., M., C.F., G.L.V., and C.V. conceived the strategy for allele-selective targeting of SNPs. All authors reviewed, provided feedback on, and approved the manuscript.
Declaration of interests
N.I., Y.L., M.F.-K., A.M., W.C.T., K.T., N.K., A.A., K.B., J.D.S., A.L., X.S.H., H.G.J., P.K., F.L., K.L., R.L., M., J.M., Q.P., E.P.-E., M.S., P.S.P., S.S., H.Y., A.Z., C.F., M.B., E.D., and C.V. were employees of Wave Life Sciences during the completion of this work. M. is currently an employee of Stoke Therapeutics. G.L.V. is on the board of directors and is a consultant and shareholder for Wave Life Sciences.
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.omtn.2024.102246.
Supplemental information
References
- 1.Bates G., Dorsey R., Gusella J.F., Hayden M.R., Kay C., Leavitt B.R., Nance M., Ross C.A., Scahill R.I., Wetzel R., Wild E.J., et al. Huntington’s disease. Nat. Rev. Dis. Primers. 2015;1:15005. doi: 10.1038/nrdp.2015.5. [DOI] [PubMed] [Google Scholar]
- 2.Sturrock A., Leavitt B.R. The clinical and genetic features of Huntington disease. J. Geriatr. Psychiatry Neurol. 2010;23:243–259. doi: 10.1177/0891988710383573. [DOI] [PubMed] [Google Scholar]
- 3.Macdonald M. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell. 1993;72:971–983. doi: 10.1016/0092-8674(93)90585-e. [DOI] [PubMed] [Google Scholar]
- 4.Altar C.A., Cai N., Bliven T., Juhasz M., Conner J.M., Acheson A.L., Lindsay R.M., Wiegand S.J. Anterograde transport of brain-derived neurotrophic factor and its role in the brain. Nature. 1997;389:856–860. doi: 10.1038/39885. [DOI] [PubMed] [Google Scholar]
- 5.Cattaneo E., Zuccato C., Tartari M. Normal huntingtin function: an alternative approach to Huntington's disease. Nat. Rev. Neurosci. 2005;6:919–930. doi: 10.1038/nrn1806. [DOI] [PubMed] [Google Scholar]
- 6.Caviston J.P., Ross J.L., Antony S.M., Tokito M., Holzbaur E.L.F. Huntingtin facilitates dynein/dynactin-mediated vesicle transport. Proc. Natl. Acad. Sci. USA. 2007;104:10045–10050. doi: 10.1073/pnas.0610628104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Wong Y.C., Holzbaur E.L.F. The regulation of autophagosome dynamics by huntingtin and HAP1 is disrupted by expression of mutant huntingtin, leading to defective cargo degradation. J. Neurosci. 2014;34:1293–1305. doi: 10.1523/JNEUROSCI.1870-13.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu J.P., Zeitlin S.O. The long and the short of aberrant ciliogenesis in Huntington disease. J. Clin. Invest. 2011;121:4237–4241. doi: 10.1172/JCI60243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Karam A., Tebbe L., Weber C., Messaddeq N., Morlé L., Kessler P., Wolfrum U., Trottier Y. A novel function of Huntingtin in the cilium and retinal ciliopathy in Huntington's disease mice. Neurobiol. Dis. 2015;80:15–28. doi: 10.1016/j.nbd.2015.05.008. [DOI] [PubMed] [Google Scholar]
- 10.Dragatsis I., Levine M.S., Zeitlin S. Inactivation of Hdh in the brain and testis results in progressive neurodegeneration and sterility in mice. Nat. Genet. 2000;26:300–306. doi: 10.1038/81593. [DOI] [PubMed] [Google Scholar]
- 11.Schaeffer J., Vilallongue N., Decourt C., Blot B., El Bakdouri N., Plissonnier E., Excoffier B., Paccard A., Diaz J.J., Humbert S., et al. Customization of the translational complex regulates mRNA-specific translation to control CNS regeneration. Neuron. 2023;111:2881–2898.e12. doi: 10.1016/j.neuron.2023.06.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Saudou F., Humbert S. The Biology of Huntingtin. Neuron. 2016;89:910–926. doi: 10.1016/j.neuron.2016.02.003. [DOI] [PubMed] [Google Scholar]
- 13.Hedreen J.C., Peyser C.E., Folstein S.E., Ross C.A. Neuronal loss in layers V and VI of cerebral cortex in Huntington's disease. Neurosci. Lett. 1991;133:257–261. doi: 10.1016/0304-3940(91)90583-f. [DOI] [PubMed] [Google Scholar]
- 14.Vonsattel J.P., Myers R.H., Stevens T.J., Ferrante R.J., Bird E.D., Richardson E.P., Jr. Neuropathological classification of Huntington's disease. J. Neuropathol. Exp. Neurol. 1985;44:559–577. doi: 10.1097/00005072-198511000-00003. [DOI] [PubMed] [Google Scholar]
- 15.Southwell A.L., Kordasiewicz H.B., Langbehn D., Skotte N.H., Parsons M.P., Villanueva E.B., Caron N.S., Østergaard M.E., Anderson L.M., Xie Y., et al. Huntingtin suppression restores cognitive function in a mouse model of Huntington's disease. Sci. Transl. Med. 2018;10 doi: 10.1126/scitranslmed.aar3959. [DOI] [PubMed] [Google Scholar]
- 16.Kordasiewicz H.B., Stanek L.M., Wancewicz E.V., Mazur C., McAlonis M.M., Pytel K.A., Artates J.W., Weiss A., Cheng S.H., Shihabuddin L.S., et al. Sustained therapeutic reversal of Huntington's disease by transient repression of huntingtin synthesis. Neuron. 2012;74:1031–1044. doi: 10.1016/j.neuron.2012.05.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yamamoto A., Lucas J.J., Hen R. Reversal of Neuropathology and Motor Dysfunction in a Conditional Model of Huntington's Disease. Cell. 2000;101:57–66. doi: 10.1016/s0092-8674(00)80623-6. [DOI] [PubMed] [Google Scholar]
- 18.Rook M.E., Southwell A.L. Antisense Oligonucleotide Therapy: From Design to the Huntington Disease Clinic. BioDrugs. 2022;36:105–119. doi: 10.1007/s40259-022-00519-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kaemmerer W.F., Grondin R.C. The effects of huntingtin-lowering: what do we know so far? Degener. Neurol. Neuromuscul. Dis. 2019;9:3–17. doi: 10.2147/DNND.S163808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Van Raamsdonk J.M., Pearson J., Rogers D.A., Bissada N., Vogl A.W., Hayden M.R., Leavitt B.R. Loss of wild-type huntingtin influences motor dysfunction and survival in the YAC128 mouse model of Huntington disease. Hum. Mol. Genet. 2005;14:1379–1392. doi: 10.1093/hmg/ddi147. [DOI] [PubMed] [Google Scholar]
- 21.Zhang Y., Li M., Drozda M., Chen M., Ren S., Mejia Sanchez R.O., Leavitt B.R., Cattaneo E., Ferrante R.J., Hayden M.R., Friedlander R.M. Depletion of wild-type huntingtin in mouse models of neurologic diseases. J. Neurochem. 2003;87:101–106. doi: 10.1046/j.1471-4159.2003.01980.x. [DOI] [PubMed] [Google Scholar]
- 22.McBride J.L., Pitzer M.R., Boudreau R.L., Dufour B., Hobbs T., Ojeda S.R., Davidson B.L. Preclinical safety of RNAi-mediated HTT suppression in the rhesus macaque as a potential therapy for Huntington's disease. Mol. Ther. 2011;19:2152–2162. doi: 10.1038/mt.2011.219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Leavitt B.R., van Raamsdonk J.M., Shehadeh J., Fernandes H., Murphy Z., Graham R.K., Wellington C.L., Raymond L.A., Hayden M.R. Wild-type huntingtin protects neurons from excitotoxicity. J. Neurochem. 2006;96:1121–1129. doi: 10.1111/j.1471-4159.2005.03605.x. [DOI] [PubMed] [Google Scholar]
- 24.Rigamonti D., Sipione S., Goffredo D., Zuccato C., Fossale E., Cattaneo E. Huntingtin's neuroprotective activity occurs via inhibition of procaspase-9 processing. J. Biol. Chem. 2001;276:14545–14548. doi: 10.1074/jbc.C100044200. [DOI] [PubMed] [Google Scholar]
- 25.Zhang Y., Leavitt B.R., van Raamsdonk J.M., Dragatsis I., Goldowitz D., MacDonald M.E., Hayden M.R., Friedlander R.M. Huntingtin inhibits caspase-3 activation. EMBO J. 2006;25:5896–5906. doi: 10.1038/sj.emboj.7601445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Becanovic K., Norremolle A., Neal S.J., Kay C., Collins J.A., Arenillas D., Lilja T., Gaudenzi G., Manoharan S., Doty C.N., et al. A SNP in the HTT promoter alters NF-kappaB binding and is a bidirectional genetic modifier of Huntington disease. Nat. Neurosci. 2015;18:807–816. doi: 10.1038/nn.4014. [DOI] [PubMed] [Google Scholar]
- 27.Van Raamsdonk J.M., Pearson J., Murphy Z., Hayden M.R., Leavitt B.R. Wild-type huntingtin ameliorates striatal neuronal atrophy but does not prevent other abnormalities in the YAC128 mouse model of Huntington disease. BMC Neurosci. 2006;7:80. doi: 10.1186/1471-2202-7-80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Rigamonti D., Bauer J.H., De-Fraja C., Conti L., Sipione S., Sciorati C., Clementi E., Hackam A., Hayden M.R., Li Y., et al. Wild-Type Huntingtin Protects from Apoptosis Upstream of Caspase-3. J. Neurosci. 2000;20:3705–3713. doi: 10.1523/jneurosci.20-10-03705.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ho L.W., Brown R., Maxwell M., Wyttenbach A., Rubinsztein D.C. Wild type Huntingtin reduces the cellular toxicity of mutant Huntingtin in mammalian cell models of Huntington's disease. J. Med. Genet. 2001;38:450–452. doi: 10.1136/jmg.38.7.450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gauthier L.R., Charrin B.C., Borrell-Pagès M., Dompierre J.P., Rangone H., Cordelières F.P., De Mey J., MacDonald M.E., Lessmann V., Humbert S., Saudou F. Huntingtin controls neurotrophic support and survival of neurons by enhancing BDNF vesicular transport along microtubules. Cell. 2004;118:127–138. doi: 10.1016/j.cell.2004.06.018. [DOI] [PubMed] [Google Scholar]
- 31.Kay C., Skotte N.H., Southwell A.L., Hayden M.R. Personalized gene silencing therapeutics for Huntington disease. Clin. Genet. 2014;86:29–36. doi: 10.1111/cge.12385. [DOI] [PubMed] [Google Scholar]
- 32.Monteys A.M., Wilson M.J., Boudreau R.L., Spengler R.M., Davidson B.L. Artificial miRNAs Targeting Mutant Huntingtin Show Preferential Silencing In Vitro and In Vivo. Mol. Ther. Nucleic Acids. 2015;4 doi: 10.1038/mtna.2015.7. [DOI] [PubMed] [Google Scholar]
- 33.Carroll J.B., Warby S.C., Southwell A.L., Doty C.N., Greenlee S., Skotte N., Hung G., Bennett C.F., Freier S.M., Hayden M.R. Potent and selective antisense oligonucleotides targeting single-nucleotide polymorphisms in the Huntington disease gene/allele-specific silencing of mutant huntingtin. Mol. Ther. 2011;19:2178–2185. doi: 10.1038/mt.2011.201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ostergaard M.E., Gerland B., Escudier J.M., Swayze E.E., Seth P.P. Differential effects on allele selective silencing of mutant huntingtin by two stereoisomers of alpha,beta-constrained nucleic acid. ACS Chem. Biol. 2014;9:1975–1979. doi: 10.1021/cb5003027. [DOI] [PubMed] [Google Scholar]
- 35.van Bilsen P.H.J., Jaspers L., Lombardi M.S., Odekerken J.C.E., Burright E.N., Kaemmerer W.F. Identification and allele-specific silencing of the mutant huntingtin allele in Huntington's disease patient-derived fibroblasts. Hum. Gene Ther. 2008;19:710–719. doi: 10.1089/hum.2007.116. [DOI] [PubMed] [Google Scholar]
- 36.Xu D., Ejebe K., Viglietta V., Dale E., Hu X.S., Boyanapalli R., Lake S.L., Panzara M. F39 Design of an adaptive randomized controlled phase 1B/2A trial of WVE-003 in participants with huntington’s disease. J. Neurol. Neurosurg. Psychiatr. 2021;92:A35. [Google Scholar]
- 37.Jafar-Nejad P., Powers B., Soriano A., Zhao H., Norris D.A., Matson J., DeBrosse-Serra B., Watson J., Narayanan P., Chun S.J., et al. The atlas of RNase H antisense oligonucleotide distribution and activity in the CNS of rodents and non-human primates following central administration. Nucleic Acids Res. 2021;49:657–673. doi: 10.1093/nar/gkaa1235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kandasamy P., Liu Y., Aduda V., Akare S., Alam R., Andreucci A., Boulay D., Bowman K., Byrne M., Cannon M., et al. Impact of guanidine-containing backbone linkages on stereopure antisense oligonucleotides in the CNS. Nucleic Acids Res. 2022;50:5401–5423. doi: 10.1093/nar/gkac037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gray M., Shirasaki D.I., Cepeda C., André V.M., Wilburn B., Lu X.H., Tao J., Yamazaki I., Li S.H., Sun Y.E., et al. Full-length human mutant huntingtin with a stable polyglutamine repeat can elicit progressive and selective neuropathogenesis in BACHD mice. J. Neurosci. 2008;28:6182–6195. doi: 10.1523/JNEUROSCI.0857-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Southwell A.L., Warby S.C., Carroll J.B., Doty C.N., Skotte N.H., Zhang W., Villanueva E.B., Kovalik V., Xie Y., Pouladi M.A., et al. A fully humanized transgenic mouse model of Huntington disease. Hum. Mol. Genet. 2013;22:18–34. doi: 10.1093/hmg/dds397. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Eckstein F. Phosphorothioates, essential components of therapeutic oligonucleotides. Nucleic Acid Ther. 2014;24:374–387. doi: 10.1089/nat.2014.0506. [DOI] [PubMed] [Google Scholar]
- 42.Liu Y., Dodart J.C., Tran H., Berkovitch S., Braun M., Byrne M., Durbin A.F., Hu X.S., Iwamoto N., Jang H.G., et al. Variant-selective stereopure oligonucleotides protect against pathologies associated with C9orf72-repeat expansion in preclinical models. Nat. Commun. 2021;12:847. doi: 10.1038/s41467-021-21112-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kielpinski L.J., Hagedorn P.H., Lindow M., Vinther J. RNase H sequence preferences influence antisense oligonucleotide efficiency. Nucleic Acids Res. 2017;45:12932–12944. doi: 10.1093/nar/gkx1073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Iwamoto N., Butler D.C.D., Svrzikapa N., Mohapatra S., Zlatev I., Sah D.W.Y., Standley S.M., Standley S.M., Lu G., Apponi L.H., et al. Control of phosphorothioate stereochemistry substantially increases the efficacy of antisense oligonucleotides. Nat. Biotechnol. 2017;35:845–851. doi: 10.1038/nbt.3948. [DOI] [PubMed] [Google Scholar]
- 45.Wu H., Lima W.F., Crooke S.T. Investigating the structure of human RNase H1 by site-directed mutagenesis. J. Biol. Chem. 2001;276:23547–23553. doi: 10.1074/jbc.M009676200. [DOI] [PubMed] [Google Scholar]
- 46.Nowotny M., Gaidamakov S.A., Ghirlando R., Cerritelli S.M., Crouch R.J., Yang W. Structure of human RNase H1 complexed with an RNA/DNA hybrid: insight into HIV reverse transcription. Mol. Cell. 2007;28:264–276. doi: 10.1016/j.molcel.2007.08.015. [DOI] [PubMed] [Google Scholar]
- 47.Lima W.F., Rose J.B., Nichols J.G., Wu H., Migawa M.T., Wyrzykiewicz T.K., Siwkowski A.M., Crooke S.T. Human RNase H1 discriminates between subtle variations in the structure of the heteroduplex substrate. Mol. Pharmacol. 2007;71:83–91. doi: 10.1124/mol.106.025015. [DOI] [PubMed] [Google Scholar]
- 48.Skotte N.H., Southwell A.L., Østergaard M.E., Carroll J.B., Warby S.C., Doty C.N., Petoukhov E., Vaid K., Kordasiewicz H., Watt A.T., et al. Allele-specific suppression of mutant huntingtin using antisense oligonucleotides: providing a therapeutic option for all Huntington disease patients. PLoS One. 2014;9 doi: 10.1371/journal.pone.0107434. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Miller J.R.C., Pfister E.L., Liu W., Andre R., Träger U., Kennington L.A., Lo K., Dijkstra S., Macdonald D., Ostroff G., et al. Allele-Selective Suppression of Mutant Huntingtin in Primary Human Blood Cells. Sci. Rep. 2017;7 doi: 10.1038/srep46740. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Byrne M., Vathipadiekal V., Apponi L., Iwamoto N., Kandasamy P., Longo K., Liu F., Looby R., Norwood L., Shah A., et al. Stereochemistry Enhances Potency, Efficacy, and Durability of Malat1 Antisense Oligonucleotides In Vitro and In Vivo in Multiple Species. Transl. Vis. Sci. Technol. 2021;10:23. doi: 10.1167/tvst.10.1.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Golyshev V.M., Pyshnyi D.V., Lomzov A.A. Effects of Phosphoryl Guanidine Modification of Phosphate Residues on the Structure and Hybridization of Oligodeoxyribonucleotides. J. Phys. Chem. B. 2021;125:2841–2855. doi: 10.1021/acs.jpcb.0c10214. [DOI] [PubMed] [Google Scholar]
- 52.Lomzov A.A., Kupryushkin M.S., Shernyukov A.V., Nekrasov M.D., Dovydenko I.S., Stetsenko D.A., Pyshnyi D.V. Data for isolation and properties analysis of diastereomers of a mono-substituted phosphoryl guanidine trideoxyribonucleotide. Data Brief. 2019;25 doi: 10.1016/j.dib.2019.104148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Dyudeeva E.S., Kupryushkin M.S., Lomzov A.A., Pyshnaya I.A., Pyshnyi D.V. Physicochemical Properties of the Phosphoryl Guanidine Oligodeoxyribonucleotide Analogs. Russ. J. Bioorg. Chem. 2019;45:709–718. doi: 10.1134/S1068162019060153. [DOI] [Google Scholar]
- 54.Zhukov S.A., Pyshnyi D.V., Kupryushkin M.S. Synthesis of Novel Representatives of Phosphoryl Guanidine Oligonucleotides. Russ. J. Bioorg. Chem. 2021;47:380–389. doi: 10.1134/S1068162021020291. [DOI] [Google Scholar]
- 55.Bazhenov M.A., Shernyukov A.V., Kupryushkin M.S., Pyshnyi D.V. Study of the Staudinger Reaction and Reveal of Key Factors Affecting the Efficacy of Automatic Synthesis of Phosphoryl Guanidinic Oligonucleotide Analogs. Russ. J. Bioorg. Chem. 2019;45:699–708. doi: 10.1134/S1068162019060074. [DOI] [Google Scholar]
- 56.Kupryushkin M.S., Pyshnyi D.V., Stetsenko D.A. Phosphoryl guanidines: a new type of nucleic Acid analogues. Acta Naturae. 2014;6:116–118. [PMC free article] [PubMed] [Google Scholar]
- 57.Kandasamy P., McClorey G., Shimizu M., Kothari N., Alam R., Iwamoto N., Kumarasamy J., Bommineni G.R., Bezigian A., Chivatakarn O., et al. Control of backbone chemistry and chirality boost oligonucleotide splice switching activity. Nucleic Acids Res. 2022;50:5443–5466. doi: 10.1093/nar/gkac018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Tabrizi S.J., Leavitt B.R., Landwehrmeyer G.B., Wild E.J., Saft C., Barker R.A., Blair N.F., Craufurd D., Priller J., Rickards H., et al. Targeting Huntingtin Expression in Patients with Huntington's Disease. N. Engl. J. Med. 2019;380:2307–2316. doi: 10.1056/NEJMoa1900907. [DOI] [PubMed] [Google Scholar]
- 59.Nagata T., Dwyer C.A., Yoshida-Tanaka K., Ihara K., Ohyagi M., Kaburagi H., Miyata H., Ebihara S., Yoshioka K., Ishii T., et al. Cholesterol-functionalized DNA/RNA heteroduplexes cross the blood-brain barrier and knock down genes in the rodent CNS. Nat. Biotechnol. 2021;39:1529–1536. doi: 10.1038/s41587-021-00972-x. [DOI] [PubMed] [Google Scholar]
- 60.Brown K.M., Nair J.K., Janas M.M., Anglero-Rodriguez Y.I., Dang L.T.H., Peng H., Theile C.S., Castellanos-Rizaldos E., Brown C., Foster D., et al. Expanding RNAi therapeutics to extrahepatic tissues with lipophilic conjugates. Nat. Biotechnol. 2022;40:1500–1508. doi: 10.1038/s41587-022-01334-x. [DOI] [PubMed] [Google Scholar]
- 61.Prakash T.P., Mullick A.E., Lee R.G., Yu J., Yeh S.T., Low A., Chappell A.E., Østergaard M.E., Murray S., Gaus H.J., et al. Fatty acid conjugation enhances potency of antisense oligonucleotides in muscle. Nucleic Acids Res. 2019;47:6029–6044. doi: 10.1093/nar/gkz354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Chandler K.J., Chandler R.L., Broeckelmann E.M., Hou Y., Southard-Smith E.M., Mortlock D.P. Relevance of BAC transgene copy number in mice: transgene copy number variation across multiple transgenic lines and correlations with transgene integrity and expression. Mamm. Genome. 2007;18:693–708. doi: 10.1007/s00335-007-9056-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Van Keuren M.L., Gavrilina G.B., Filipiak W.E., Zeidler M.G., Saunders T.L. Generating transgenic mice from bacterial artificial chromosomes: transgenesis efficiency, integration and expression outcomes. Transgenic Res. 2009;18:769–785. doi: 10.1007/s11248-009-9271-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Goodwin L.O., Splinter E., Davis T.L., Urban R., He H., Braun R.E., Chesler E.J., Kumar V., van Min M., Ndukum J., et al. Large-scale discovery of mouse transgenic integration sites reveals frequent structural variation and insertional mutagenesis. Genome Res. 2019;29:494–505. doi: 10.1101/gr.233866.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Nakagaki A., Urakawa A., Hirano S., Anami T., Kishino T. Application of droplet digital PCR in the analysis of genome integration and organization of the transgene in BAC transgenic mice. Sci. Rep. 2018;8:6638. doi: 10.1038/s41598-018-25001-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Hodgson J.G., Agopyan N., Gutekunst C.A., Leavitt B.R., LePiane F., Singaraja R., Smith D.J., Bissada N., McCutcheon K., Nasir J., et al. A YAC mouse model for Huntington's disease with full-length mutant huntingtin, cytoplasmic toxicity, and selective striatal neurodegeneration. Neuron. 1999;23:181–192. doi: 10.1016/s0896-6273(00)80764-3. [DOI] [PubMed] [Google Scholar]
- 67.Southwell A.L., Skotte N.H., Kordasiewicz H.B., Østergaard M.E., Watt A.T., Carroll J.B., Doty C.N., Villanueva E.B., Petoukhov E., Vaid K., et al. In vivo evaluation of candidate allele-specific mutant huntingtin gene silencing antisense oligonucleotides. Mol. Ther. 2014;22:2093–2106. doi: 10.1038/mt.2014.153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Harper S.Q., Staber P.D., He X., Eliason S.L., Martins I.H., Mao Q., Yang L., Kotin R.M., Paulson H.L., Davidson B.L. RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model. Proc. Natl. Acad. Sci. USA. 2005;102:5820–5825. doi: 10.1073/pnas.0501507102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.DiFiglia M., Sena-Esteves M., Chase K., Sapp E., Pfister E., Sass M., Yoder J., Reeves P., Pandey R.K., Rajeev K.G., et al. Therapeutic silencing of mutant huntingtin with siRNA attenuates striatal and cortical neuropathology and behavioral deficits. Proc. Natl. Acad. Sci. USA. 2007;104:17204–17209. doi: 10.1073/pnas.0708285104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Boudreau R.L., McBride J.L., Martins I., Shen S., Xing Y., Carter B.J., Davidson B.L. Nonallele-specific silencing of mutant and wild-type huntingtin demonstrates therapeutic efficacy in Huntington's disease mice. Mol. Ther. 2009;17:1053–1063. doi: 10.1038/mt.2009.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Zeitler B., Froelich S., Marlen K., Shivak D.A., Yu Q., Li D., Pearl J.R., Miller J.C., Zhang L., Paschon D.E., et al. Allele-selective transcriptional repression of mutant HTT for the treatment of Huntington's disease. Nat. Med. 2019;25:1131–1142. doi: 10.1038/s41591-019-0478-3. [DOI] [PubMed] [Google Scholar]
- 72.Schobel S. CHDI 16th annual HD therapuetics conference. 2021. Preliminary results from GENERATION HD1, a phase III trial of tominersen in individuals with manifest HD. [Google Scholar]
- 73.McColgan P. HD2022: Milton Wexler Biennial Symposium. 2022. Towards an understanding of the treatment and mechanistic effects of tominersen. [Google Scholar]
- 74.Sharma A., Laosinchai-Wolf W., Mitchen E., Eveleigh D., Wilkinson-Busha K., Turner J., Church M., Lara A., Gentry C., Goyal J., et al. eP242: Analytical validation of a PCR/CE assay that phases SNPs with CAG-expanded alleles for selecting Huntington disease patients for allele-selective treatments. Genet. Med. 2022;24:S154. doi: 10.1016/j.gim.2022.01.277. [DOI] [Google Scholar]
- 75.Pfister E.L., Kennington L., Straubhaar J., Wagh S., Liu W., DiFiglia M., Landwehrmeyer B., Vonsattel J.P., Zamore P.D., Aronin N. Five siRNAs targeting three SNPs may provide therapy for three-quarters of Huntington's disease patients. Curr. Biol. 2009;19:774–778. doi: 10.1016/j.cub.2009.03.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.R Development C.T. R Foundation for Statistical Computing; 2019. R: A Language and Environment for Statistical Computing.https://www.R-project.org/ [Google Scholar]
- 77.Berthold M.R., Cebron N., Dill F., Gabriel T.R., Kรถtter T., Meinl T., Ohl P., Sieb C., Thiel K., Wiswedel B. In: Data Analysis, Machine Learning and Applications. Preisach C., Burkhardt H., Schmidt-Thieme L., Decker R., editors. Springer; 2008. KNIME: The Konstanz Information Miner. [Google Scholar]
- 78.Kassambara A. 2023. rstatix: Pipe-Friendly Framework for Basic Statistical Tests.https://rpkgs.datanovia.com/rstatix/ R package version 0.7.2. [Google Scholar]
- 79.Zeileis A. Econometric Computing with HC and HAC Covariance Matrix Estimators. J. Stat. Softw. 2004;11:1–17. doi: 10.18637/jss.v011.i10. [DOI] [Google Scholar]
- 80.Zeileis A. Object-oriented Computation of Sandwich Estimators. J. Stat. Softw. 2006;16:1–16. doi: 10.18637/jss.v016.i09. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The authors confirm that the data supporting the findings of this study are available within the article and its supplemental information.