Abstract
Background
Network pharmacology has emerged as a powerful tool to understand the therapeutic effects and mechanisms of natural products. However, there is a lack of comprehensive evaluations of network-based approaches for natural products on identifying therapeutic effects and key mechanisms.
Purpose
We systematically explore the capabilities of network-based approaches on natural products, using Panax ginseng as a case study. P. ginseng is a widely used herb with a variety of therapeutic benefits, but its active ingredients and mechanisms of action on chronic diseases are not yet fully understood.
Methods
Our study compiled and constructed a network focusing on P. ginseng by collecting and integrating data on ingredients, protein targets, and known indications. We then evaluated the performance of different network-based methods for summarizing known and unknown disease associations. The predicted results were validated in the hepatic stellate cell model.
Results
We find that our multiscale interaction-based approach achieved an AUROC of 0.697 and an AUPR of 0.026, which outperforms other network-based approaches. As a case study, we further tested the ability of multiscale interactome-based approaches to identify active ingredients and their plausible mechanisms for breast cancer and liver cirrhosis. We also validated the beneficial effects of unreported and top-predicted ingredients, in cases of liver cirrhosis and gastrointestinal neoplasms.
Conclusion
our study provides a promising framework to systematically explore the therapeutic effects and key mechanisms of natural products, and highlights the potential of network-based approaches in natural product research.
Keywords: Network-based approach, Therapeutic effects, Panax ginseng
Graphical abstract
Highlights
-
•
We utilized network-based approach to systematically investigate the therapeutic effects and mechanisms of Panax ginsengs.
-
•
We found that a multiscale interaction-based approach outperformed other methods in identifying therapeutic effects of Panax ginseng's ingredients.
-
•
We found that the key mechanisms of ginseng's effects on breast cancer and liver cirrhosis are mediated by cell proliferation and apoptosis.
-
•
We found that Ginsenoside-Rg3 and (20R)-Ginsenoside Rh1 have beneficial effects on liver cirrhosis.
1. Introduction
Network pharmacology, utilizing methods from systems biology, has emerged as a useful tool to understand the therapeutic effect of drugs in interconnected biological systems [1]. In particular, it has been widely employed to analyze and elucidate the therapeutic effects and mechanisms of herbal medicines, and has been actively applied in various studies, such as identifying active compounds and predicting potential targets [2,3]. Our previous study also highlighted the methodological trends in traditional herbal medicine network pharmacology research, revealing a dramatic increase in studies and the utilization of databases/tools [4]. One of the underlying reasons for this is that network pharmacology approaches is well-suited to herbal medicines, which are composed of multiple ingredients that act on multiple targets [5]. For example, one study utilized a network-based approach to identify the therapeutic effects of a polyphenol based on the proximity between the drug target and a disease-related protein [6]. Another researcher used network pharmacological analysis to prioritize and validate flavonoids for nonalcoholic fatty liver disease, or to identify systems-level mechanisms for the antioxidant properties of Bupleuri Radix and its active compound [7,8]. As such, network pharmacology is expected to be useful in discovering the therapeutic effect of herbal medicine and presenting testable hypotheses.
Researchers have increasingly relied on network-based algorithms to identify the therapeutic effects of natural products [9,10]. Unlike other computational approaches such as machine learning, network-based algorithms can suggest reliable, verifiable hypotheses based on associations between drug targets and disease proteins in complex biological networks [11]. The most widely utilized approach is to consider the overlap between natural products and disease-related proteins [[12], [13], [14]]. However, this approach totally neglects protein-protein interactions and cannot account for the propagation of the effects of a drug on other proteins. Recently developed network-based methods have reported that considering the network between various biological systems will provide more accurate results in predicting the therapeutic effect of drugs. Specifically, Güney et al. developed network proximity and applied it to discover new therapeutic effects of drugs [15]. Network proximity is a method that measures the proximity between drug targets and disease-related proteins in a protein-protein interaction network, and is used to predict new therapeutic effects by evaluating the connectivity between existing drug targets and proteins in related diseases. Meanwhile, Ruiz et al. applied propagation effects on a multiscale interactome composed of proteins and biological functions to identify the therapeutic effect of a drug [16]. The study found that simulating the propagation of a compound or disease on a multiscale interactome demonstrated state-of-the-art performance in predicting therapeutic effects and uncovered its key mechanisms. Despite these possibilities, there has not yet been a systematic attempt to compare and evaluate network methods for natural products.
In this study, we aim to systematically explore the efficacy of network-based approaches for natural products, using Panax ginseng C. A. Meyer (P. Ginseng) as a case study. P. Ginseng, a plant widely used for thousands of years in East Asia, has been recognized for its unique therapeutic value worldwide and is currently one of the most consumed medicinal herbs globally. According to Donguibogam, a traditional Korean medicine book registered on the UNESCO World Heritage list, P. ginseng is known to have tonifying effects on internal energy, stabilizing effects on the mind, and replenishing effects on deficiencies. It can be applied to various ailments such as fatigue, fright palpitations, and forgetfulness. Western countries have also drawn attention to the potential therapeutic effects of ginseng, and the European Committee for Medicinal Plants states that it is effective for symptoms associated with tiredness and weakness [17]. Furthermore, current clinical studies have indicated that ginseng could potentially benefit patients with Alzheimer's disease, chronic heart failure, hypertension, and glucose metabolism [[18], [19], [20], [21]]. Despite this interest in ginseng's efficacy, most research on ginseng is limited to reinterpreting prior knowledge. Given the vast number of possible indications and mechanisms, relying solely on experimental screens is both costly and time-consuming. Additionally, ginseng's various effects are exerted by multiple ingredients, making it challenging to identify active compounds and their key mechanisms. Therefore, there is an urgent need for a novel approach to systematically investigate the therapeutic effects and key mechanisms of P. ginseng.
In this work, we employ a network-based approach to investigate the therapeutic effects of P. ginseng and dissect its potential mechanisms (Fig. 1). We comprehensively compiled ingredients of P. ginseng, its protein targets, and known indications. We constructed a compound-target network and tested whether the constructed network could reflect the known mechanisms of P. ginseng. The systematic analysis was performed to evaluate which network-based method can best discriminate between known and unknown disease associations for P. ginseng. As a case study, we further tested whether a network-based approach can identify active ingredients and their plausible mechanisms for breast cancer and liver cirrhosis. We experimentally validated several candidate ginseng ingredients for which there was no prior evidence for liver cirrhosis. Taken together, we believe that this study presents a systematic strategy to dissect the disease treatment mechanism of P. ginseng by elucidating new indications for P. ginseng and its core mechanisms. It also provides a promising framework for the systematic investigation of therapeutic effects and key mechanisms of natural products.
Fig. 1.
Integrated workflow for investigating novel indication of P. ginseng and their potential mechanisms. The workflow includes three main steps: A. compiling P. ginseng ingredients, their protein targets, and known indications; B. evaluating the performance of network-based methods to on recapulating therapeutic effects of P. ginseng; and C. performing case studies to validate the predicted indications and mechanisms of P. ginseng.
2. Materials and methods
2.1. Ingredients and protein targets of panax ginseng
The ingredients of P. ginseng were retrieved from a literature and SymMap (https://www.symmap.org) database [22]. SymMap integrated ingredient information of herbal medicine from the TCMID (version 2015), TCMSP (version 2.3) and TCM-ID (version 1.0) databases according to common identifiers of the compounds. The active ingredients of P. ginseng mentioned in the literature were additionally included [23]. For the analysis, we only considered compounds that 1) could be mapped in PubChem IDs, and 2) had protein target information with experimental evidence. The compound id and its representative name were mapped and curated using pubchem.
The experimentally validated targets of ginseng ingredients were obtained from the data compiled by Huang et al. STITCH, therapeutic target database (TTD), and HIT 2.0 [[24], [25], [26], [27]]. Huang et al. assembled direct and indirect compound-protein interactions of natural products were from several databases. STITCH integrated target information for 430,000 chemicals from disparate data sources. TTD provides comprehensive information about the known and explored targets, the targeted disease, pathway information and the corresponding drugs directed at each of these targets. HIT2.0 is a comprehensive platform for herbal ingredients and their target information based on literature evidences. Target identifiers such as gene symbol or UniProt id were mapped with Entrez gene id using SYNGO [28].
2.2. Compound-target network construction
A compound–target network is a bipartite network, in which nodes in the network are defined as compounds and targets and the edges between compounds and targets are defined as compound-target interactions (Yes or No). The network was constructed and visualized based on information about the compounds, targets, and their interactions using Cytoscape (version 3.8.2) [29]. The biological processes or signaling pathways related to the target proteins were identified by gene set enrichment analysis (GSEA) using Enrichr (http://amp.pharm.mssm.edu/Enrichr/) [30]. Enrichr computes enrichment by the assessment of multiple gene-set libraries (e.g., gene ontology and Kyoto Encyclopedia of Genes and Genomes (KEGG)) and calculates adjusted p-values, z-scores, and combined scores for the gene lists of interest (target genes). The combined score is calculated by the logarithm of the multiplication of the p-value and z-score. For analyzes utilizing the KEGG pathway, we considered pathways related to the organismal system and excluded pathways related to disease (i.e., Hepatitis B).
2.3. Multiscale interactome
The multiscale interactome refers to integrating both physical interactions between proteins and a multiscale hierarchy of biological functions. The multiscale interactome used in this study was obtained from the data built by Ruiz et al. [16]. They built the network by integrating the three types of associations: protein-protein interactions, protein–biological function interactions, and biological function–biological function interactions. For protein-protein interactions, they assembled 387,626 physical interactions between 17,660 proteins from seven major databases such as the Biological General Repository for Interaction Datasets, the Database of Interacting Proteins, and the Human Reference Protein Interactome Mapping Project. For protein-biological function interactions, they compiled 34,777 associations between 7993 proteins and 6387 biological functions from human version of the Gene Ontology. For biological-biological function interactions, they constructed a hierarchy of biological functions consisting of 22,545 associations and 9798 biological functions.
2.4. Network-based methods for predicting disease association
To predict which ginseng ingredient will treat a given disease, we considered three metrics: protein overlap, network proximity, and multiscale interactome (Supplementary Fig. 1). Protein overlap is based on the tendency that ingredients with overlapping targets with disease-related proteins have therapeutic effects. It can be calculated by the Jaccard Simility between the set of drug targets T and the set of disease proteins S:
Network proximity relied on the findings that the closer the targets of a compound are to disease proteins on a human protein-protein interaction network, the more likely that the compound will affect the disease phenotype [15]. It is obtained by calculating a relative score of the shortest path length between drug and disease based on the reference distribution. Let T and S be the set of drug targets and disease proteins, and be the shortest path length between nodes s and t. Network proximity first computes the average closest distance between disease-associated proteins and ingredient targets as follows:
Next, a reference distance distribution is constructed consisting of values of when S and T are randomly permuted to 1000 sets of proteins that match the size and degrees of the original disease proteins and targets in the network. Finally, relative score is computed by taking a z-score of with respect to the reference distribution:
where and denotes the mean and standard deviation of the reference distribution, respectively.
Multiscale interactome models and compares the impact of drug treatment and disease perturbation on the network that integrate interactions between proteins and biological functions. It predict disease association of drugs and natural products by calculating the diffusion profile , and then measures the correlation distance between them. A diffusion profile is computed through a matrix formulation with power iteration as follows:
, where denotes diffusion profiles at k-th state, denotes the probability of the walker continuing its walk at a given step rather than restarting, denotes a restart vector which sets the probability the walker will jump to each node after a restart, and denotes a biased transtion matrix derived from a directed multiscale interactome and set of scalar weights which encode the relative likelihood.
Those procedures are repeated until the convergence of the power iteration computation is as follows:
where denotes the tolerance parameter and was set to , the same value of the previous study.
The correlation of drug and disease diffusion profile is calculated as follows:
where r(c) and r(d) denote the diffusion profiles of the drug and disease, respectively
2.5. Known disease association of ginseng ingredients
Known disease associations of ginseng ingredients were obtained from the Comparative Toxicogenomic Database (CTD) [31]. CTD is a publicly available database that aims to advance understanding of the effects of environmental exposure on human health, providing manually curated information such as chemical–disease associations. Among the chemical-disease associations, we selected associations labeled as “therapeutic”, meaning that the chemical has a known or potential therapeutic role in the condition. The explicit associations between ginseng ingredients-disease were expanded to implicit associations by considering the hierarchical structure of diseases along the Mesh tree. This procedure was performed by propagating associations in the lower branches of the Mesh tree to consider diseases in the higher levels of the same tree branch. Finally, we identified 192 associations between 14 ginseng ingredients and 78 diseases that could be used for ground truth for known disease associations for ginseng ingredients.
2.6. Materials
Anti-p-Smad 2, anti-p-Smad 3 and anti-β-actin antibody were supplied from Cell Signaling Technologies (Danvers, MA, USA). Anti-α-SMA, 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-tetrazolium bromide (MTT) and other reagents were purchased from Sigma–Aldrich (St. Louis, MO, USA).
2.7. Cell culture, MTT assay and western blot analysis
LX-2 cells, an immortalized human HSCs line, were kindly provided by Dr. SL Friedman (Mount Sinai School of Medicine, NY, USA) [32]. AGS cells were purchased from.
American Type Culture Collection (ATCC). LX-2 cells were treated with drugs for the Western blot. Preparation of protein lysates from cell line was performed as previously described protocol [32]. In case of MTT assay, AGS cells were stimulated with compounds for the indicated time [32].
2.8. Data analysis
A Multiple comparison tests for different dose groups were conducted as previously described [32]. Variance homogeneity was determined by using the Levene test [33] The statistical significance of the differences among the treatment groups was verified by one-way analysis of variance (ANOVA). An unpaired t-test was also used to analyze the differences, and a non-parametric test was used (Mann–Whitney U test) if the distribution of data was abnormal.
3. Results
3.1. Construction and analysis of compound-target network
Network construction and analysis was conducted to reveal chemical diversity and experimentally validated protein targets of P. ginseng. We identified 42 ingredients of P. ginseng with experimentally validated protein targets (Fig. 2A). To explain the chemical diversity of ginseng, we identified superclasses and classes of ginseng ingredients using Classyfire [34]. The result showed that ginseng ingredients are distributed across 8 superclasses and 12 classes. Among superclasses, lipids and lipid-like molecules, phenylpropanoids and polyketides, benzenoids are the top three superclasses with 22, 7, and 4 compounds, respectively. Among 12 classes, prenol lipids, fatty Acyls, and flavonoids are the top three classes with 13, 8, and 5 compounds, respectively (Fig. 2B).
Fig. 2.
Selection process for flavonoids evaluated in this study and their chemical distribution. A. The flowchart of selecting the compounds of Panax ginseng. B. Distribution of compounds in Panax ginseng across different chemical superclasses and classes obtained from ClassyFire. The inner circle and outer circle represent the proportions of superclass and classes, respectively.
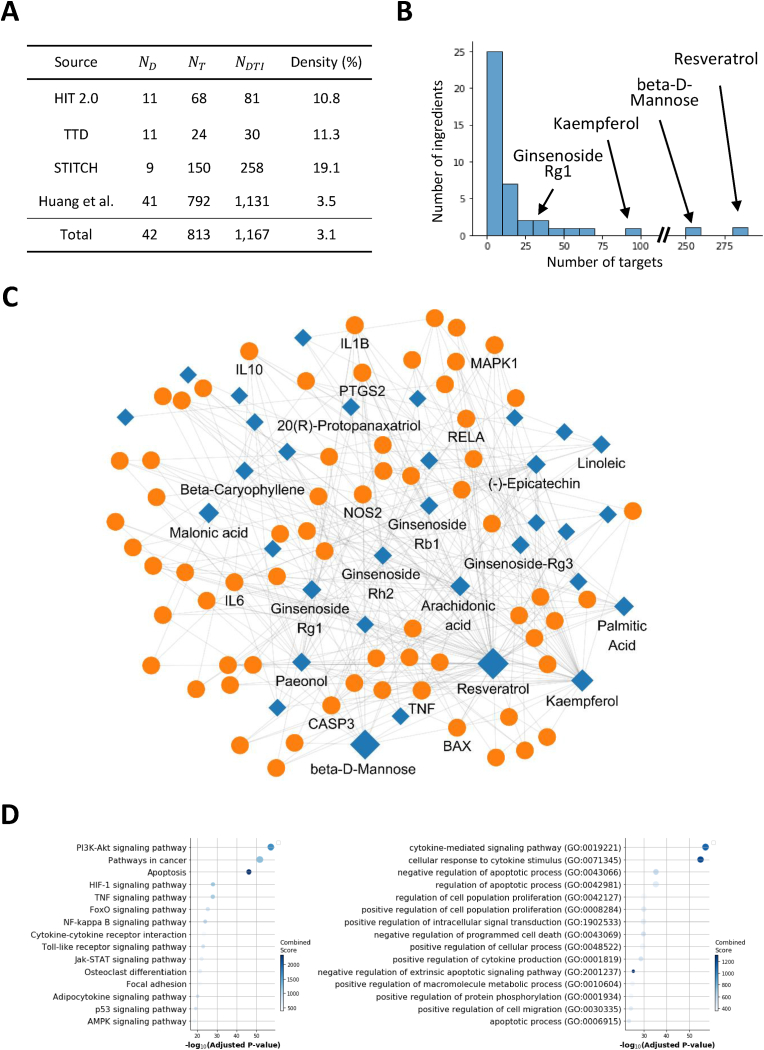
We compiled experimentally validated target information for ginseng ingredients from various databases (Fig. 3A). We find that the majority of ginseng ingredients (25/42) have less than ten protein targets, while a few compounds have an exceptionally large number of targets (Fig. 3B). A compound-target network was constructed and visualized between ginseng ingredients and their representative protein targets (Fig. 3C). To investigate related biological processes and pathways, we conducted the functional enrichment analysis for these targets based on the Gene ontology and KEGG pathway, respectively. Through enrichment analysis on KEGG, we observed that ginseng targets are significantly implicated in various signaling pathways, including the PI3K-Akt pathway and apoptotic processes, as well as pathways involved in cancer. Similar enrichment analysis using Gene Ontology (GO) indicated significant associations with biological functions such as cytokine-mediated signaling and cellular responses to cytokine stimulus. These combined findings suggest that the diverse biological activities of ginseng may be mediated by its interaction with multiple signaling pathways and biological functions. These findings suggest that the therapeutic effects of ginseng could be mediated through its targets' involvement in a diverse range of signaling pathways and biological functions.
Fig. 3.
A compound-target network for Panax ginseng and its property. A. Statistics of experimentally target information for Panax ginseng. , , and represents the number of drugs, targets, and drug-target interactions, respectively. B. Distribution of the number of protein targets of ginseng ingredients. C. A representative compound-target network for Panax ginseng. For efficient visualization, a subnetwork between 66 targets interacting with 3 or more of the protein targets and their ginseng ingredients was visualized. Circles and diamonds denote protein targets and compounds, respectively. Edges denote experimentally validated interactions between them. D. Top (n = 15) enriched pathways (left) and gene ontology terms (Biological Process, right) among all protein targets of Panax ginseng. The x-axis represents the statistical significance associated with each term.
3.2. Recapitulating known ingredient-disease associations for ginseng
The various therapeutic effects of ginseng in treating diverse diseases stem from the different ingredients it contains. Thus, the test to recapitulate known and unknown associations between ginseng ingredients and disease would be an efficient way to evaluate the capacity to identify the therapeutic effects of ginseng. To select the most suitable approaches, we conducted a comprehensive comparison using the conventional method or state-of-the-art network-based methods: protein overlap, network proximity and multi-scale interactome (See methods). For this analysis, we grouped all 12,588 ginseng ingredient-disease associations among 42 ginseng ingredients and 299 diseases into known (192) and unknown (12,396) associations. We assessed how well network-based metrics differentiate between the known and unknown sets by evaluating the area under the curve of the receiver operating characteristic curve (AUROC) and the precision-recall curve (AUPR). It is noteworthy that the performance shown by network-based approaches seem lower than the typical performance shown by other machine learning tasks. This may be due to the inherent difficulty of the problem, stemming from the low proportion of currently known ingredient-disease pairs, and the fact that the unknown ingredient-disease pairs include pairs that have not yet been discovered. Therefore, this task still be efficient in investigating which network-based approach can efficiently identify the therapeutic effect of ginseng.
We observed that all three network-based methods demonstrated performance above the chance level in terms of AUROC and AUPR (Fig. 4A). This supports the notion that considering the association between protein targets and disease-associated proteins could be useful for identifying disease associations in ginseng ingredients. The protein overlap method achieved satisfactory performance in terms of AUROC and AUPR (0.626 and 0.021, respectively) but was only able to priorities for 20 % of the ingredient-disease pairs with overlapping proteins. On the other hand, network proximity and multi-scale interactome (MSI) are not constrained by the limitation of requiring overlapping proteins. In particular, the MSI approach outperformed the other methods, achieving an AUROC of 0.697 and an AUPR of 0.026, demonstrating the highest performance among network-based methods. Taken together, our results suggest that considering the multiscale-level impact on proteins and biological functions between ingredients and diseases is a more effective approach for recapitulating the therapeutic effects of ginseng.
Fig. 4.
Performance Comparison of Network-Based Prediction Methods for Recapitulating Ginseng Component-Disease Associations. A. Performance curve between all ginseng ingredients-disease pairs. B. Performance distribution of predicting the therapeutic effects of ginseng ingredients, evaluated based on both disease levels (n = 78, left penal) and ingredient levels (n = 14, right penal). The figure presents a comparison of the performance of three network-based prediction methods—protein overlap, network proximity, and multiscale interactome—in recapitulating known associations and unknown associations between ginseng ingredients and diseases. Each data point is color-coded based on the method used, the dashed line indicates the chance level performance. MSI: Multiscale interactome; AUC: Area under the curve.
We also investigated how accurately each method predicts which diseases a particular ginseng ingredient may have a therapeutic effect on, and vice versa (Fig. 4B). For convenience, we'll refer to these tasks as ingredient-level tasks and disease-level tasks, respectively. This task allowed us to explore how well each prediction method identifies diseases for a given ginseng ingredient and whether it can pinpoint a ginseng ingredient that can be utilized for a specific disease, respectively. Again, the MSI approach outperformed the other prediction methods in both the disease-level task and the ingredient-level task. We also found that the MSI approach exceeded the chance level for 80 % of both tasks, while the other methods performed below the chance level for almost half of the diseases and ingredients in both tasks. These results suggest that the MSI is consistently superior to other methods for the identification of active ingredients in ginseng for individual diseases, as well as for the identification of therapeutic effects of ingredients on diseases. The protein overlap, network proximity, and multi-scale interactome results for all disease and ginseng ingredient pairs are summarized in Supplementary Table 1.
3.3. Identifying active ingredients and potential mechanisms in various diseases
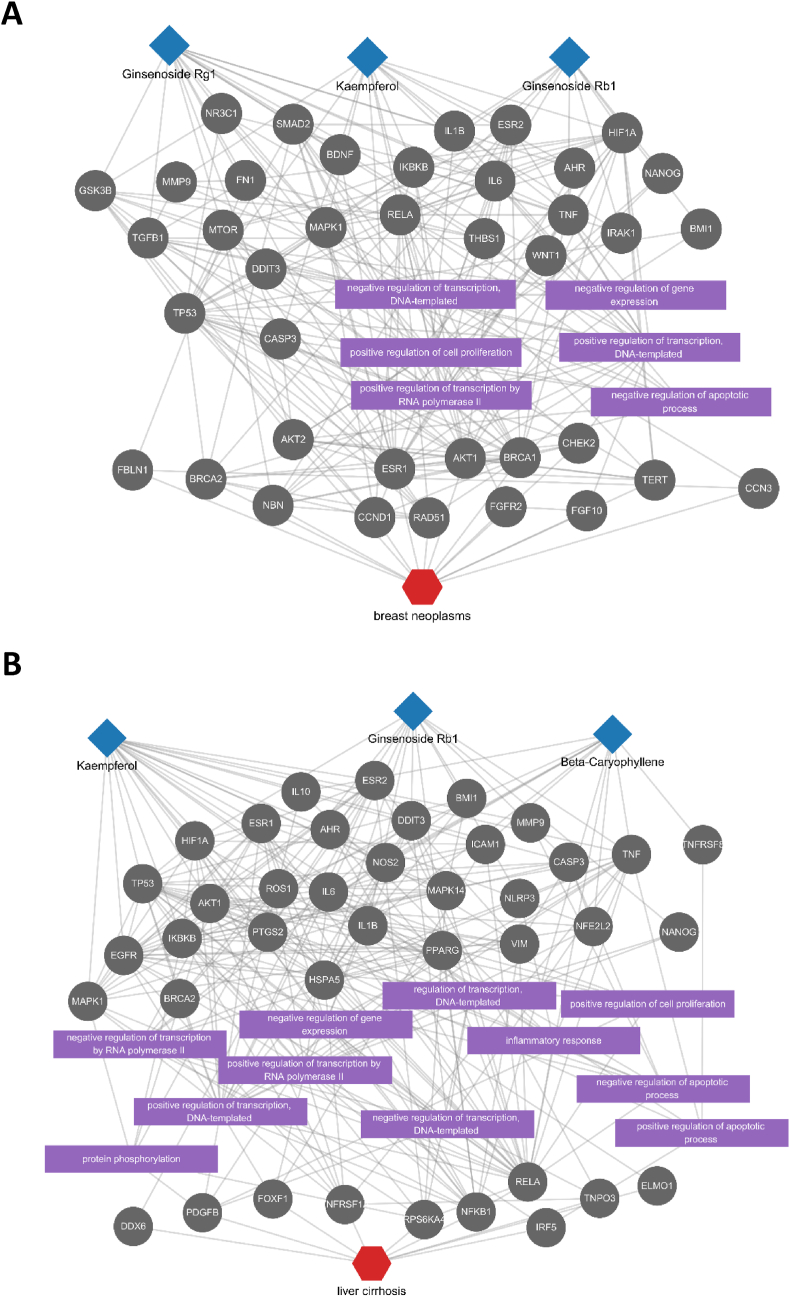
We then investigated whether MSI, which demonstrated the highest performance in previous experiments, could elucidate the active compounds and key mechanisms of P. ginseng for known indications. This was done by prioritizing ingredients for specific diseases and comparing the ingredients' potential mechanisms for the diseases (See Methods). We focused on the detectable compounds in P. ginseng to facilitate the experimental validation of our findings. Our first case study focused on breast cancer, which is one of the most common cancers with a high prevalence rate. A previous cohort study suggested that administration of ginseng increased the survival rate and quality of life in breast cancer patients [35]. Therefore, identifying the key active ingredients and mechanisms of ginseng can facilitate drug discovery for breast cancer. We prioritized ingredients for breast cancer with the MSI approach and found that kaempferol, ginsenoside Rb1, and ginsenoside Rg1 had the highest correlation with the effects of breast cancer (Fig. 5A). Previous studies have reported that these ingredients may indeed be utilized in breast cancer [[36], [37], [38]], and this consistent result indicates that our predictions successfully recapitulate existing studies. To further investigate whether MSI approach can identify the key mechanisms, we constructed a subnetwork consisting of proteins and biological functions that were significantly affected by breast cancer and ginseng ingredients. In the constructed network, there is no direct path of proteins between ginseng ingredients and breast cancer. Instead, the targets of these ingredients directly interact with proteins such as ESR1, BRCA1, and BRCA2, and biological process related to cell proliferation, and apoptotic process.
Fig. 5.
Potential mechanisms of ginseng ingredients for breast neoplasms (A) and liver cirrhosis (B). Each network was constructed by inducing a subgraph consisting of the top 20 nodes in the diffusion profiles of the selected ginseng ingredients and disease. These nodes correspond to the proteins and biological functions most affected by ginseng ingredients or disease.
Following our investigation into breast cancer, we further investigated the effects of ginseng on liver cirrhosis as another case study, given its global health impact and potential benefits for liver-related diseases. A clinical study suggested anti-inflammatory and antifatigue effects of P. ginseng in patients with nonalcoholic fatty liver disease, and in vivo studies also investigated the protective effects against liver fibrosis [39,40]. We found that the diffusion profiles of kaempferol, ginsenoside Rb1, and beta-caryophyllene had the highest correlation scores with that of liver cirrhosis. Again, all of these have reported therapeutic or potentially protective effects in cirrhosis. The results of the subnetwork analysis revealed that their mechanisms of action were suggested to involve the regulation of modulating protein functions related to liver cirrhosis, either directly or through molecular interactions, or by regulating biological functions such as apoptosis, cell proliferation, and inflammation (Fig. 5B). These suggested mechanisms are known to play a crucial role in the onset and progression of liver cirrhosis [41]. Taken together, the findings demonstrated the effectiveness of the network-based approach in successfully recapitulating active ingredients and their key mechanisms against breast cancer and liver cirrhosis.
3.4. Prediction and validation of novel active ingredients
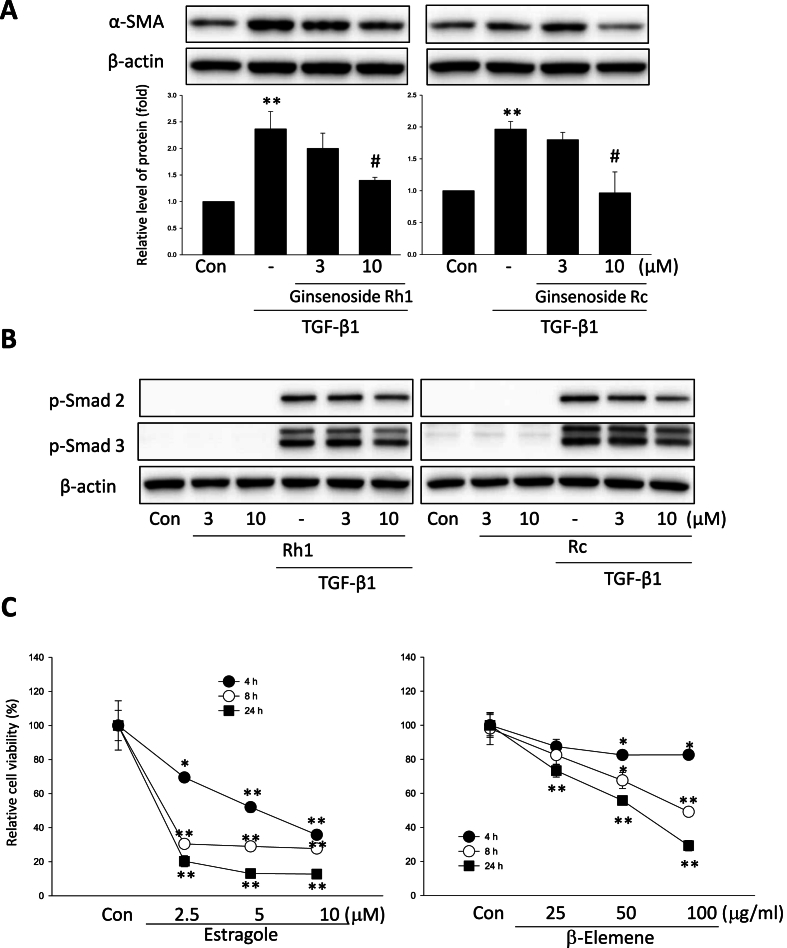
The successful recapitulation of the active ingredients and key mechanisms suggests that the network-based approach has the potential to discover new ginseng ingredients that can be utilized for various diseases. In refining our focus on liver cirrhosis, we catalogued ginseng ingredients known to be detectable through chromatography and classified as endogenous metabolites. We found that ginsenoside-Rc and (20R)-ginsenoside Rh1 had the highest correlation scores for liver cirrhosis among ingredients with no previous evidence (Supplementary Table 2). These findings indicate that these ingredients could be considered novel candidates for exerting beneficial effects on liver cirrhosis. To test these findings, we examined the effects of these compounds on a cell model of liver cirrhosis using hepatic stellate cells (HSCs) stimulated with TGF-β1. Before the investigations, we tested the cytotoxicity of the compounds in LX-2 cells, a human HSC cell line. In an MTT assay, the two compounds showed no significant differences in cell viability up to 10 μM (data not shown). Activation of HSCs is the most important cell mechanism to induce liver cirrhosis, and α-SMA is one of the representative markers of HSC activation. As expected, TGF-β1 significantly induced the level of α-SMA in LX-2 cells (Fig. 6A). Interestingly, both Rc and Rh1 markedly inhibited the induction of α-SMA, which indicates that Rc and Rh1 have the possibility of inhibiting liver cirrhosis. The phosphorylation of Smad 2/3 is most important in the TGF-β1-related signaling to induce α-SMA [42]. Next, we confirmed the effects of Rc and Rh1 on the phosphorylation of Smad 2/3 induced by TGF-β1. Rc and Rh1 inhibited the ability of TGF-β1 to induce the phosphorylation of Smad 2/3 (Fig. 6B). Next, we also shifted our focus to another exemplary disease, gastrointestinal neoplasms. In a similar context to the liver cirrhosis experiments, we discovered that Estragole and β-elemene, though previously without evidence, showed the highest correlation scores (Supplementary Table 3). We determined the effects of the predicted compounds such as estragole and β-Elemene on cell viability of AGS gastric cancer cells (Fig. 6C). As predicted, both estragole and β-Elemene significantly decreased the cell viability of the AGS cells. These findings demonstrate that the network-based approach is not only effective in discovering new therapeutic potentials of ginseng ingredients, but also helpful in validating the mechanisms of action of known ingredients.
Fig. 6.
Prediction and validation of a novel ginseng ingredient candidate for liver cirrhosis. A. Effects of ginsenoside Rc and Rh1 on TGF-β1-induced HSCs activation. Rc and Rh1 suppressed TGF-β1-induced expression of the fibrotic marker protein α-SMA. B. Effects of Rc and Rh1 on TGF-β1/Smad pathway in HSCs. Phosphorylation levels of Smad 2 and 3 were analyzed by immunoblotting with specific antibodies. Western blot performed on the lysates of serum starved LX-2 cells that were treated with Rc and Rh1 in the presence or absence of TGF-β1 (5 ng/ml) for an additional 24 h. C. MTT assay for cancidate ginseng ingredients. AGS cells were treated with β-Elemene (25, 50 and 100 μg/ml) and Estragole (2.5, 5 and 10 μM) for 4, 8 and 24 h. Data represents the mean ± S.E.M. from three separate experiments. *P < 0.05, **P < 0.01 vs vehicle-treated control;#P < 0.05 vs TGF-β1 treated group.
4. Discussion
P. Ginseng is one of the most famous natural products in the world and has been widely prescribed to treat diseases and promote health. In this study, we systematically evaluated and validated the therapeutic effect of ginseng and its active ingredients by utilizing various network-based approaches. We constructed a network for ginseng utilizing experimentally validated information, which well reflected the previously known effects of ginseng. By comparing various network-based methods, we found that MSI provided the most accurate predictions for recapitulating known ingredient-disease associations in P. ginseng. As a case study, we tested whether the active ingredients and key mechanisms of ginseng's therapeutic effects on breast cancer and liver cirrhosis. Finally, we predicted and validated a novel ginseng candidate that can be utilized for the treatment of liver cirrhosis.
By integrating data from multiple databases, we have constructed a network that illuminates the intricate web of signaling pathways and biological functions associated with ginseng's ingredients and targets. Our functional enrichment analysis corroborates ginseng's association with pivotal pathways like PI3K-Akt, HIF-1, and Jak-STAT, and biological processes such as cytokine signaling and apoptosis regulation (Fig. 3D), echoing well-established mechanisms in existing ginseng research [43,44]. Furthermore, the mechanisms we've delineated not only echo known signaling pathways but also emphasize the corroborated therapeutic potential. Notably, P. ginseng is recognized for enhancing Type I collagen synthesis via Smad signaling pathway activation, reflecting its contribution to tissue repair and anti-aging [45]. Similarly, the ginsenoside Rh1 has been observed to enhance the anti-inflammatory effects of dexamethasone, likely through the modulation of the NF-κB signaling pathway [46]. This convergence with known mechanisms substantiates the reliability of our network-based analysis, affirming its value as a robust tool for understanding the therapeutic implications of ginseng.
To the best of our knowledge, our study is the first attempt to systematically comparison to identify active ingredients and new indications for disease treatment of natural products. Current approaches to identifying disease treatments of natural products using network pharmacology mainly relied on protein overlap and network proximity measures. our results show that protein overlap can only be used to compare the priorities of a limited range of component-disease pairs. While network proximity can consider the relationships between proteins on the network, its performance showed results close to random choice. This indicates that simply considering interactions between proteins for natural products can lead to false-positive results. On the other hand, we found that considering the impact on a multiscale interactome between ginseng components and disease proteins most accurately recapitulates the component-disease relationship in ginseng.
The significance of our study lies in its systematic approach to understanding the therapeutic effects of P. ginseng. By leveraging network-based methods, we were able to gain insights into the active ingredients and their underlying mechanisms of action for specific diseases, which could potentially lead to the development of more targeted and effective therapies. Intriguingly, our findings regarding kaempferol, ginsenoside Rb1, and ginsenoside Rg1 align with known mechanisms, suggesting these components modulate key targets such as ESR1, BRCA1, and AKT2 and regulate biological functions associated with apoptosis and cell proliferation in breast cancer. These observations are consistent with reported effects in the literature [38], [47], [48], thereby supporting the therapeutic relevance of these compounds and indicating the predictive capability of our MSI approach. Moreover, our findings can serve as a foundation for future research on natural products and their therapeutic applications, contributing to ongoing efforts to harness the power of traditional medicine for modern healthcare solutions. Our approach could also be extended to explore the synergistic effects or interactions between multiple compounds in natural products, providing valuable insights for the development of combination therapies or understanding the multifaceted nature of herbal medicines.
In this study, there are several potential limitations that warrant further consideration and improvement. First, the data used for constructing the network, including protein targets and their interactions, may not be complete or fully accurate. In the future, incorporating more comprehensive and up-to-date data sources could help enhance the reliability and coverage of our network. Second, the methods used in this study, while promising, might still benefit from further refinement and validation. For instance, alternative network-based approaches or machine learning techniques could be explored to improve the prediction accuracy for active ingredients and their therapeutic effects. Despite these potential limitations, our constructed network effectively reflected the previously known therapeutic effects of ginseng, demonstrating the novelty and robustness of our approach. Furthermore, we experimentally validated the prediction results, providing additional confidence in our findings. Overall, this study lays the groundwork for future research aimed at better understanding the therapeutic potential of natural products and advancing the field of natural product-based drug discovery.
Funding information
This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number HF20C0212), the National Research Foundation of Korea (NRF), funded by the Korean government (MSIT) (grant number 2022R1I1A1A01057053, RS-2023-00243363, RS-2023-00218419), and the Ministry of Food and Drug Safety in 2023 (grant number 21173MFDS561).
Data availability
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding authors.
Ethical statement
Not applicable.
Declaration of competing interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Acknowledgements
Kim YW would like to thank to the Ph.D.‘s program of Kyungpook National University for completing the thesis through this work.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jgr.2024.01.005.
Appendix A. Supplementary data
The following are the Supplementary data to this article.
References
- 1.Hopkins A.L. Network pharmacology: the next paradigm in drug discovery. Nat Chem Biol. 2008;4(11):682–690. doi: 10.1038/nchembio.118. [DOI] [PubMed] [Google Scholar]
- 2.Li S., Zhang B. Traditional Chinese medicine network pharmacology: theory, methodology and application. Chin J Nat Med. 2013;11(2):110–120. doi: 10.1016/S1875-5364(13)60037-0. [DOI] [PubMed] [Google Scholar]
- 3.Zhang R., Zhu X., Bai H., Ning K. Network pharmacology databases for traditional Chinese medicine: Review and assessment. Front Pharmacol. 2019;10:123. doi: 10.3389/fphar.2019.00123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lee W.Y., Lee C.Y., Kim Y.S., Kim C.E. The methodological trends of traditional herbal medicine employing network pharmacology. Biomolecules. 2019;9(8) doi: 10.3390/biom9080362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Li S., Zhang Z.Q., Wu L.J., Zhang X.G., Li Y.D., Wang Y.Y. Understanding ZHENG in traditional Chinese medicine in the context of neuro-endocrine-immune network. IET Syst Biol. 2007;1(1):51–60. doi: 10.1049/iet-syb:20060032. [DOI] [PubMed] [Google Scholar]
- 6.do Valle I.F., Roweth H.G., Malloy M.W., Moco S., Barron D., Battinelli E., Loscalzo J., Barabasi A.L. Network medicine framework shows that proximity of polyphenol targets and disease proteins predicts therapeutic effects of polyphenols. Nat Food. 2021;2(3):143–155. doi: 10.1038/s43016-021-00243-7. [DOI] [PubMed] [Google Scholar]
- 7.Lee W.Y., Lee C.Y., Lee J.S., Kim C.E. Identifying candidate flavonoids for non-Alcoholic fatty liver disease by network-based strategy. Front Pharmacol. 2022;13 doi: 10.3389/fphar.2022.892559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bak S.B., Song Y.R., Bae S.J., Lee W.Y., Kim Y.W. Integrative approach to uncover antioxidant properties of Bupleuri Radix and its active compounds: multiscale interactome-level analysis with experimental validation. Free Radic Biol Med. 2023;199:141–153. doi: 10.1016/j.freeradbiomed.2023.02.016. [DOI] [PubMed] [Google Scholar]
- 9.Wu W., Zhang Z., Li F., Deng Y., Lei M., Long H., Hou J., Wu W. A network-based approach to explore the mechanisms of Uncaria Alkaloids in treating hypertension and alleviating Alzheimer's disease. Int J Mol Sci. 2020;21(5) doi: 10.3390/ijms21051766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Zhang Y.-q., Mao X., Guo Q.-y., Lin N., Li S. Network pharmacology-based approaches capture essence of Chinese herbal medicines. Chinese Herbal Medicines. 2016;8(2):107–116. [Google Scholar]
- 11.Ay M., Goh K.-I., Cusick M.E., Barabasi A.-L., Vidal M. Drug--target network. Nat Biotechnol. 2007;25(10):1119–1127. doi: 10.1038/nbt1338. [DOI] [PubMed] [Google Scholar]
- 12.Hao D.C., Xiao P.G. Network pharmacology: a Rosetta stone for traditional C hinese medicine. Drug Dev Res. 2014;75(5):299–312. doi: 10.1002/ddr.21214. [DOI] [PubMed] [Google Scholar]
- 13.Bu D., Su Z., Zou J., Meng M., Wang C. Study of the mechanism underlying therapeutic effect of Compound Longmaining on myocardial infarction using a network pharmacology-based approach. Biomed Pharmacother. 2019;118 doi: 10.1016/j.biopha.2019.109234. [DOI] [PubMed] [Google Scholar]
- 14.Zhang G.-b., Li Q.-y., Chen Q.-l., Su S.-b. 2013. Network pharmacology: a new approach for Chinese herbal medicine research, Evidence-based complementary and alternative medicine 2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Guney E., Menche J., Vidal M., Barábasi A.-L. Network-based in silico drug efficacy screening. Nat Commun. 2016;7(1) doi: 10.1038/ncomms10331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Ruiz C., Zitnik M., Leskovec J. Identification of disease treatment mechanisms through the multiscale interactome. Nat Commun. 2021;12(1):1796. doi: 10.1038/s41467-021-21770-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Agency E.M. Panax ginseng C.A. Meyer, radix); 2014. Herbal medicine: summary for the public - ginseng root. [Google Scholar]
- 18.Yakoot M., Salem A., Helmy S. Effect of Memo®, a natural formula combination, on Mini-Mental State Examination scores in patients with mild cognitive impairment. Clin Interv Aging. 2013:975–981. doi: 10.2147/CIA.S44777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen W., Yao P., Vong C.T., Li X., Chen Z., Xiao J., Wang S., Wang Y. Ginseng: a bibliometric analysis of 40-year journey of global clinical trials. J Adv Res. 2021;34:187–197. doi: 10.1016/j.jare.2020.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wei H., Wu H., Yu W., Yan X., Zhang X. Shenfu decoction as adjuvant therapy for improving quality of life and hepatic dysfunction in patients with symptomatic chronic heart failure. J Ethnopharmacol. 2015;169:347–355. doi: 10.1016/j.jep.2015.04.016. [DOI] [PubMed] [Google Scholar]
- 21.Vuksan V., Sievenpiper J.L. Herbal remedies in the management of diabetes: lessons learned from the study of ginseng. Nutr Metabol Cardiovasc Dis. 2005;15(3):149–160. doi: 10.1016/j.numecd.2005.05.001. [DOI] [PubMed] [Google Scholar]
- 22.Wu Y., Zhang F., Yang K., Fang S., Bu D., Li H., Sun L., Hu H., Gao K., Wang W. SymMap: an integrative database of traditional Chinese medicine enhanced by symptom mapping. Nucleic Acids Res. 2019;47(D1):D1110–D1117. doi: 10.1093/nar/gky1021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Park S.-Y., Park J.-H., Kim H.-S., Lee C.-Y., Lee H.-J., Kang K.S., Kim C.-E. Systems-level mechanisms of action of Panax ginseng: a network pharmacological approach. Journal of ginseng research. 2018;42(1):98–106. doi: 10.1016/j.jgr.2017.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Huang Y., Fang J., Lu W., Wang Z., Wang Q., Hou Y., Jiang X., Reizes O., Lathia J., Nussinov R. A systems pharmacology approach uncovers wogonoside as an angiogenesis inhibitor of triple-negative breast cancer by targeting hedgehog signaling. Cell Chem Biol. 2019;26(8):1143–1158. doi: 10.1016/j.chembiol.2019.05.004. e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Szklarczyk D., Santos A., Von Mering C., Jensen L.J., Bork P., Kuhn M. Stitch 5: augmenting protein–chemical interaction networks with tissue and affinity data. Nucleic Acids Res. 2016;44(D1):D380–D384. doi: 10.1093/nar/gkv1277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Chen X., Ji Z.L., Chen Y.Z. TTD: therapeutic target database. Nucleic Acids Res. 2002;30(1):412–415. doi: 10.1093/nar/30.1.412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Yan D., Zheng G., Wang C., Chen Z., Mao T., Gao J., Yan Y., Chen X., Ji X., Yu J. Hit 2.0: an enhanced platform for herbal ingredients' targets. Nucleic Acids Res. 2022;50(D1):D1238–D1243. doi: 10.1093/nar/gkab1011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Koopmans F., van Nierop P., Andres-Alonso M., Byrnes A., Cijsouw T., Coba M.P., Cornelisse L.N., Farrell R.J., Goldschmidt H.L., Howrigan D.P. SynGO: an evidence-based, expert-curated knowledge base for the synapse. Neuron. 2019;103(2):217–234. e4. doi: 10.1016/j.neuron.2019.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shannon P., Markiel A., Ozier O., Baliga N.S., Wang J.T., Ramage D., Amin N., Schwikowski B., Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–2504. doi: 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kuleshov M.V., Jones M.R., Rouillard A.D., Fernandez N.F., Duan Q., Wang Z., Koplev S., Jenkins S.L., Jagodnik K.M., Lachmann A. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44(W1):W90–W97. doi: 10.1093/nar/gkw377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Davis A.P., Grondin C.J., Johnson R.J., Sciaky D., McMorran R., Wiegers J., Wiegers T.C., Mattingly C.J. The comparative toxicogenomics database: update 2019. Nucleic Acids Res. 2019;47(D1):D948–D954. doi: 10.1093/nar/gky868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lee J.-H., Jang E.J., Seo H.L., Ku S.K., Lee J.R., Shin S.S., Park S.-D., Kim S.C., Kim Y.W. Sauchinone attenuates liver fibrosis and hepatic stellate cell activation through TGF-β/Smad signaling pathway. Chem Biol Interact. 2014;224:58–67. doi: 10.1016/j.cbi.2014.10.005. [DOI] [PubMed] [Google Scholar]
- 33.Levene A. Pathological factors influencing excision of tumours in the head and neck. Part I. Clin Otolaryngol Allied Sci. 1981;6(2):145–151. doi: 10.1111/j.1365-2273.1981.tb01800.x. [DOI] [PubMed] [Google Scholar]
- 34.Djoumbou Feunang Y., Eisner R., Knox C., Chepelev L., Hastings J., Owen G., Fahy E., Steinbeck C., Subramanian S., Bolton E. ClassyFire: automated chemical classification with a comprehensive, computable taxonomy. J Cheminf. 2016;8:1–20. doi: 10.1186/s13321-016-0174-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cui Y., Shu X.-O., Gao Y.-T., Cai H., Tao M.-H., Zheng W. Association of ginseng use with survival and quality of life among breast cancer patients. Am J Epidemiol. 2006;163(7):645–653. doi: 10.1093/aje/kwj087. [DOI] [PubMed] [Google Scholar]
- 36.Wang X., Yang Y., An Y., Fang G. The mechanism of anticancer action and potential clinical use of kaempferol in the treatment of breast cancer. Biomed Pharmacother. 2019;117 doi: 10.1016/j.biopha.2019.109086. [DOI] [PubMed] [Google Scholar]
- 37.Lau W.S., Chen W.F., Chan R.Y.K., Guo D.A., Wong M.S. Mitogen‐activated protein kinase (MAPK) pathway mediates the oestrogen‐like activities of ginsenoside Rg1 in human breast cancer (MCF‐7) cells. Br J Pharmacol. 2009;156(7):1136–1146. doi: 10.1111/j.1476-5381.2009.00123.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Lee Y.J., Jin Y.R., Lim W.C., Park W.K., Cho J.Y., Jang S., Lee S.K. Ginsenoside-R b1 acts as a weak phytoestrogen in MCF-7 human breast cancer cells. Arch Pharm Res (Seoul) 2003;26:58–63. doi: 10.1007/BF03179933. [DOI] [PubMed] [Google Scholar]
- 39.Hong M., Lee Y.H., Kim S., Suk K.T., Bang C.S., Yoon J.H., Baik G.H., Kim D.J., Kim M.J. Anti-inflammatory and antifatigue effect of Korean Red Ginseng in patients with nonalcoholic fatty liver disease. Journal of ginseng research. 2016;40(3):203–210. doi: 10.1016/j.jgr.2015.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ki S.H., Yang J.H., Ku S.K., Kim S.C., Kim Y.W., Cho I.J. Red ginseng extract protects against carbon tetrachloride-induced liver fibrosis. Journal of ginseng research. 2013;37(1):45. doi: 10.5142/jgr.2013.37.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Tsuchida T., Friedman S.L. Mechanisms of hepatic stellate cell activation. Nat Rev Gastroenterol Hepatol. 2017;14(7):397–411. doi: 10.1038/nrgastro.2017.38. [DOI] [PubMed] [Google Scholar]
- 42.Inagaki Y., Okazaki I. Emerging insights into transforming growth factor β Smad signal in hepatic fibrogenesis. Gut. 2007;56(2):284–292. doi: 10.1136/gut.2005.088690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ding Q., Zhu W., Diao Y., Xu G., Wang L., Qu S., Shi Y. Elucidation of the mechanism of action of ginseng against acute lung injury/acute respiratory distress syndrome by a network pharmacology-based strategy. Front Pharmacol. 2021;11 doi: 10.3389/fphar.2020.611794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Li Q.-Y., Hou C.-Z., Yang L.-P., Chu X.-L., Wang Y., Zhang P., Zhao Y. Study on the mechanism of ginseng in the treatment of lung adenocarcinoma based on network pharmacology. Evid base Compl Alternative Med. 2020:2020. doi: 10.1155/2020/2658795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lee J., Jung E., Lee J., Huh S., Kim J., Park M., So J., Ham Y., Jung K., Hyun C.-G. Panax ginseng induces human Type I collagen synthesis through activation of Smad signaling. J Ethnopharmacol. 2007;109(1):29–34. doi: 10.1016/j.jep.2006.06.008. [DOI] [PubMed] [Google Scholar]
- 46.Li J., Du J., Liu D., Cheng B., Fang F., Weng L., Wang C., Ling C. Ginsenoside Rh1 potentiates dexamethasone's anti-inflammatory effects for chronic inflammatory disease by reversing dexamethasone-induced resistance. Arthritis Res Ther. 2014;16(3):1–11. doi: 10.1186/ar4556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Luo H., Rankin G.O., Li Z., DePriest L., Chen Y.C. Kaempferol induces apoptosis in ovarian cancer cells through activating p53 in the intrinsic pathway. Food Chem. 2011;128(2):513–519. doi: 10.1016/j.foodchem.2011.03.073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chu Y., Zhang W., Kanimozhi G., Brindha G., Tian D. Ginsenoside Rg1 induces apoptotic cell death in triple-negative breast cancer cell lines and prevents carcinogen-induced breast tumorigenesis in sprague dawley rats. Evidence-Based Complementary and Alternative Medicine. 2020;8886955:1–12. doi: 10.1155/2020/8886955. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding authors.