Abstract

Titanium dioxide (TiO2) shows significant potential as a self-cleaning material to inactivate severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and prevent virus transmission. This study provides insights into the impact of UV-A light on the photocatalytic inactivation of adsorbed SARS-CoV-2 virus-like particles (VLPs) on a TiO2 surface at the molecular and atomic levels. X-ray photoelectron spectroscopy, combined with density functional theory calculations, reveals that spike proteins can adsorb on TiO2 predominantly via their amine and amide functional groups in their amino acids blocks. We employ atomic force microscopy and grazing-incidence small-angle X-ray scattering (GISAXS) to investigate the molecular-scale morphological changes during the inactivation of VLPs on TiO2 under light irradiation. Notably, in situ measurements reveal photoinduced morphological changes of VLPs, resulting in increased particle diameters. These results suggest that the denaturation of structural proteins induced by UV irradiation and oxidation of the virus structure through photocatalytic reactions can take place on the TiO2 surface. The in situ GISAXS measurements under an N2 atmosphere reveal that the virus morphology remains intact under UV light. This provides evidence that the presence of both oxygen and UV light is necessary to initiate photocatalytic reactions on the surface and subsequently inactivate the adsorbed viruses. The chemical insights into the virus inactivation process obtained in this study contribute significantly to the development of solid materials for the inactivation of enveloped viruses.
Keywords: SARS-CoV-2 virus-like particles (VLPs), titanium dioxide, photocatalytic oxidation, GISAXS, AFM, XPS
Introduction
The COVID-19 pandemic 2019, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has raised numerous questions for the scientific community.1 It has shown that interdisciplinary and multidisciplinary approaches encompassing medicine, material science, technology, biomedical science, chemistry, and applied physics are necessary to comprehend the mechanism behind the worldwide spread of viruses and to control them.1,2 One of the potential routes for SARS-CoV-2 transmission is through indirect contact with contaminated surfaces. The virus can remain active on various surfaces for minutes up to hours, depending on environmental conditions.3,4 Therefore, effectively controlling the transmission of the virus from surfaces is a key strategy for slowing its dissemination in both indoor and outdoor public environments. Various surface disinfection methods have been reported for virus inactivation, such as alcohol,5 heat, and ultraviolet (UV-C) (wavelength: 200–280 nm) irradiation,6−8 which inactivate the virus via denaturation, damage, or destruction of the ribonucleic acids (RNA) and proteins.8 However, several significant limitations are associated with the aforementioned disinfection methods, concerning their sustainability and safety. For example, the alcohol in antiviral products is not stable over time due to evaporation or dissipation.9 Instead, heat (>65 °C) and UV irradiation consume energy and may pose a threat to human health.8 Antiviral surfaces are promising technologies with practical applications in public places, such as hospitals, schools, and nursing homes, aimed at preventing the spread and inactivation of transmission viruses without using chemicals or requiring energy.10,11 Physical and chemical properties of the surface, such as porosity and hydrophobicity, are the key factors influencing their interaction with viruses. Therefore, there is a high demand to develop and design highly efficient antiviral surfaces using cost-effective and environmentally friendly materials.12 Photocatalytic materials represent promising antiviral coatings capable of preventing virus transmission and reducing indoor infections.13−16
Titanium dioxide (TiO2) stands out as one of the best photocatalytic materials, well-known for its high activity for the degradation of organic compounds, the inactivation of bacteria and viruses, and its self-cleaning properties.17 It is nontoxic, commercially available and chemically and thermally stable.8 By absorbing light with energy higher than the band gap (∼3.2 eV),8 photogenerated electron/hole (e–/h+) pairs form in TiO2 and can react with oxygen and water molecules, producing reactive oxygen species (ROS), such as superoxide (O2•–) and hydroxyl (HO•) radicals.8,18 These generated ROS possess high oxidative potential, effectively oxidizing organic components of microorganisms, resulting in cell wall and membrane rupture, RNA destruction, and protein alteration.19 Several metal oxide-based photocatalysts, such as CuO20 and TiO2,8 and nanoparticle decorated metal oxides (e.g., Ag/TiO2,21 Au/ZnO,22 and Pd/TiO28) have been used for virus inactivation. In most work reported so far, the reaction is performed in the liquid phase in the presence of a catalyst and virus suspension,9,23,24 and there is a lack of knowledge concerning the inactivation mechanism.
In our previous study, we demonstrated that the TiO2(101) photocatalyst surface can adsorb SARS-CoV-2 and further inactivate it through a combination action of light and thermal treatments.8 However, due to biosafety restrictions, we were unable to investigate the adsorption and interaction of the virus with the TiO2(101) surface before exposing it to UV light, as we had to inactivate the virus for further characterization outside of the biosafety level 3 (BSL-3) laboratory. Shifting all instrumentation to a BSL-3 laboratory was also unfeasible due to many technical and economic constraints. In order to overcome these limitations, in this work, we focus on the newly developed technology of SARS-CoV-2 virus-like particles (VLPs), which resemble infectious viruses in morphology but lack the viral genome.25 VLPs are also promising candidates for virus and vaccine studies (see Figure S1 in the Supporting Information).25−27 Therefore, with the present study, we report, for the first time, a combined experimental and theoretical investigation of the adsorption and photocatalytic inactivation of SARS-CoV-2 VLPs on TiO2. Furthermore, we reveal the role of light in the denaturation of viral proteins and changes in the membrane morphology of the adsorbed viruses. Such insights into interactions of VLPs with solid material surfaces pave the way for the further development and design of effective antiviral surfaces.
Results and Discussion
Adsorption of SARS-CoV-2 Virus-Like Particles (VLPs) on TiO2(101) Surface
It was reported in the literature that the SARS-CoV-2 virus can attach to some surfaces through various physical interactions, such as polar, nonpolar, and van der Waals.8,28 Moreover, it was suggested that the antiviral properties of a surface are correlated with its ability to adsorb viruses.12 As the first step of our investigation, we confirm the interaction and binding of VLPs to the TiO2(101) surface by combining fluorescence microscopy (FM), atomic force microscopy (AFM), nanoinfrared (Nano-IR), and X-ray photoelectron (XPS) spectroscopic analysis with density functional theory (DFT) calculations simulating XPS binding energies (BEs). SARS-CoV-2 VLPs, which are sketched in Figure S1, and high-quality TiO2(101) surface (single crystal 8 mm × 8 mm × 2 mm, Surface Net Ltd.) have been prepared by repeated sputtering and annealing cycles, as described in our previous study8 (see the Supporting Information for more information). In order to confirm the presence of VLPs on the surface, we carried out FM measurements. For this technique, we incubated the samples using rabbit-derived anti-SARS-CoV-2 nucleocapsid primary antibodies and antirabbit secondary antibodies conjugated with the fluorescent-dye Alexa568 (see the Experimental Section for additional details). FM intensity can indicate specific staining arising from the interaction of antibodies with VLPs, while residual nonspecific signals can result from the interaction of antibodies with other proteins adsorbed on the surface.29 To prevent antibody binding directly to the surface of TiO2, a blocking solution (a mixture of phosphate-buffered saline and bovine serum albumin, BSA) was applied to the sample with the VLPs adsorbed and the respective control samples (see Figure S3, Supporting Information). Mean fluorescence intensities for these measurements were reported in Table S1 (Supporting Information). According to the obtained values, specific staining resulted in a 1.9 times higher fluorescence intensity than the respective controls without primary antibodies or VLPs, confirming the presence of VLPs on the surface of TiO2(101).
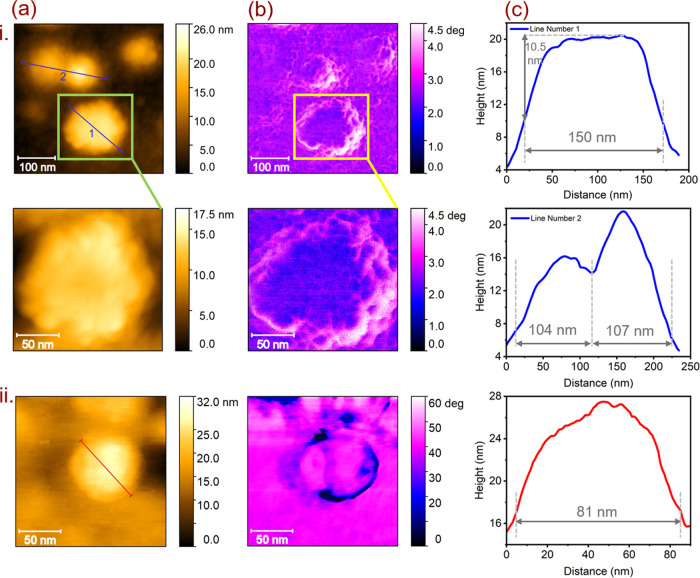
To visualize the shape and morphology of the adsorbed VLPs, the sample was characterized using AFM at DESY NanoLab.30Figure 1a,b display the AFM and phase images, respectively. The AFM images reveal spherical particles with a particle diameter in the range of 100–150 nm, corresponding to the VLPs (see line scan profiles in Figure 1c). SARS-CoV-2 VLPs have been reported in the size of 60–150 nm by Roessler et al.25 The common nonuniform size distribution is due to the engineered nature of VLPs.25 Nevertheless, these VLPs exhibit structural and visual similarities to living SARS-CoV-2 viruses,8 despite the lack of a complete virus genome. The height of the adsorbed VLPs (Figure 1c) appears to be smaller than the expected diameter (60–150 nm). A similar behavior was observed in the case of adsorbed SARS-CoV-2 on the solid surface of TiO2(101).8 The height of adsorbed SARS-CoV-2 particles on glass and tissue culture polystyrene substrates was reported in the range of 9–15 and 14–38 nm, respectively.5 This height reduction might occur due to two main factors: (i) the drying of virus particles on the surface, causing the VLPs become flatter due to the denaturation of spike and membrane proteins, and (ii) the accumulation of cell culture media (CCM, virus growth media) and other proteins smaller than VLPs on the area surrounding the virus.5,8 Additionally, virus proteins often undergo conformational changes upon surface adsorption, resulting in a lower height profile.31 This phenomenon is known as the “Vroman effect,” in which the adsorbed proteins are displaced by proteins with higher affinities for the surface.32 As proteins interact with a surface, they can bind to the surface. Over time, these adsorbed proteins undergo molecular relaxation or spread on the surface, resulting in enhanced affinity interaction with the surface and consequently shrinking in height.33 The spike proteins in SARS-CoV-2 play a crucial role in the virus’s life cycle and its interaction with host cells.8 Responsible for mediating viral entry into host cells, the spike protein becomes a key target for antiviral strategies. On average, SARS-CoV-2 is covered by approximately 24–40 spike proteins,34 each having a club-like shape measuring approximately 20 nm in length.35 The spike protein acts as a major mediator of cellular infection and consists of two distinct subunits, S1 and S2 (see Figure S1, Supporting Information).8 The size of the spike protein subunit 1, which contains the receptor binding domain to attach to the angiotensin-converting enzyme 2 receptor on host cell surfaces, has been reported to range between 10 and 15 nm.31,36
Figure 1.
(a) AFM topography (0.4 × 0.4 μm2 (i) and 0.15 × 0.15 μm2 (ii)) and (b) phase images of adsorbed SARS-CoV-2 VLPs (i) and ΔSVLP (ii) on the surface of TiO2(101). (c) Line scan profile of selected particles in AFM topography images (a, (i) and (ii)).
High-resolution AFM phase images of the VLPs corresponding to the topography images on TiO2(101) in Figure 1a are shown in Figure 1b. We observed spiky features along the edges of the virus particles in the phase image, potentially related to spike proteins. To confirm this interpretation, we further characterized engineered subviral particles without spike proteins (ΔSVLP) as a reference sample on TiO2(101). Figure 1a,b (ii) displays the AFM and phase images of ΔSVLPs/TiO2(101). The assembled ΔSVLP has a spherical shape with a size of 81 nm. Compared to the adsorbed VLPs on the surface of TiO2(101), the AFM and phase images of the ΔSVLP/TiO2(101) sample showed bald particles, providing proof of the presence of spike proteins in VLPs. We can clearly see in the AFM images that the shape and size of VLPs are similar to the SARS-CoV-2.8,37
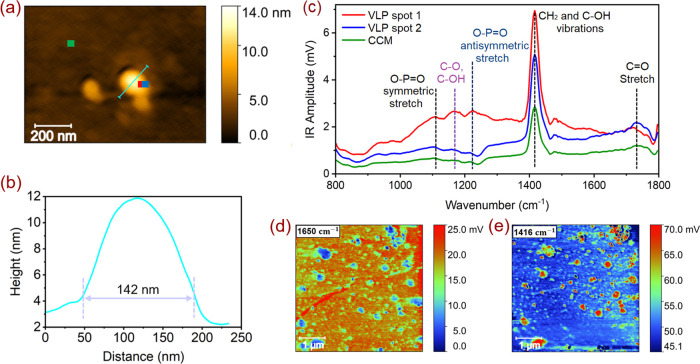
To obtain more information about the chemical binding of the adsorbed VLPs, AFM-based nanoinfrared (Nano-IR) spectroscopy was employed. Nano-IR enables the combination of AFM and IR techniques for the local visual-chemical composition mapping at the surface.38 In the case of the VLPs, the advantage of Nano-IR over bulk IR is the ability to separately identify virus signals from cell culture media (CCM) by the intrinsic spatial resolution, permitting two-dimensional (2D) chemical mapping with nanoscale resolution. In Figure S4, a broad view (5.0 × 5.0 μm2) of the surface displays many VLPs and a few self-assembled nanocrystals formed from the buffer medium during the sample drying after loading the VLPs. The AFM topography of a single virus particle and its measured line scan profile is shown in Figure 2a,b. The size of the selected VLP in this measurement is 142 nm in length and 8 nm in height. The spectra in Figure 2c were taken at two different locations inside the VLP (see the red and blue spots in the AFM image). To distinguish the information about the chemical binding of VLPs from that of the CCM, we also characterized a spot at the surface that is VLP-free as a reference containing only CCM (see the green spot in the AFM image). The strong band at 1416 cm–1 in the VLP spectrum, which exhibits lower intensity in the CCM spectrum, can be assigned to the CH239 and C–OH40 vibrations originating from the protein amino acids structure within the viral particle. Since the CCM also contains amino acids and vitamins,8 this vibration band was also observed at VLP-free spots as well. The IR band centered around 1734 cm–1 in the spectra is associated with the C=O stretching vibrations.41 The main parts of the virus such as lipids, RNA, and carbohydrates exhibit adsorption bands at 1290–1050 cm–1 region in the IR spectra.41,42 Contrary to the CCM spectrum, the observed broad bands at 1232, 1168, and 1105 cm–1 are attributed to the O–P=O antisymmetric stretching vibration,41 the functional groups of proteins (C–O stretching mode of the C–OH group),43 and O–P=O symmetric stretching vibration and C–N,41,44,45 respectively (Figure 2c). The main source of the P–O vibration is the nucleocapside protein in the VLP structure. AFM-IR nanoimaging was then carried out at two different IR frequencies: at the amide peak associated with protein at 1650 cm–1 (Figure 2d, protein map) and at the vibration of the CH2 groups at 1416 cm–1 (Figure 2e, CH2 map). The protein cargo of an individual virus particle was identified by local nano-IR measurement on the surface of solid TiO2. The nano-IR results confirm the presence of viral proteins by observing the different bands of functional groups in lipids and proteins of VLPs, compared to those in CCM.
Figure 2.
(a) AFM (1.0 × 1.0 μm2) topography image of the selected area in Figure S4. (b) Line scan profile of selected VLP for nano-IR measurement. (c) AFM-IR spectra of adsorbed SARS-CoV-2 VLPs on the surface of TiO2(101) at three different locations in the AFM image. The red, blue, and green spectra are acquired at positions corresponding to the red, blue, and green spots in the AFM topography image (a). AFM-IR maps were measured at two different frequencies: 1650 cm–1 (d) and 1416 cm–1 (e).
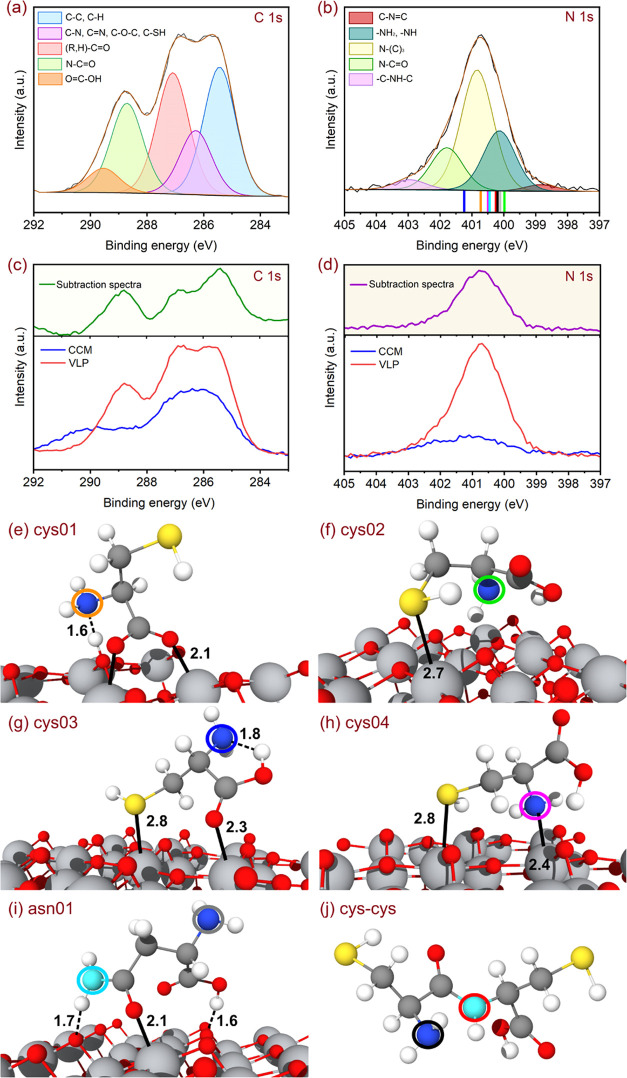
It was reported that the binding and interaction of SARS-CoV-2 with nonbiological surfaces are correlated with the virus stability and its potential for surface transmission in indoor environments.46 Therefore, gaining a deeper understanding of the interaction between various components of the functional groups in SARS-CoV-2 proteins and surfaces is crucial for developing effective strategies to mitigate virus transmission and functionalized surfaces. In addition to Nano-IR, the adsorbed VLPs on the surface of TiO2(101) are characterized using both experimental and simulated XPS (see characterization techniques section). Figure 3a,b reports high-resolution XP spectra of the C 1s and N 1s core levels. The deconvoluted XP spectrum of C 1s, which shows the contribution of lipid and proteins, is fitted with five components (Figure 3a) at binding energies (BEs) centered at 285.4 eV (C–C and C–H bonds8), 286.3 eV (C–N and C=N,8,47 C–O–C,48 and C–SH8), 287.1 eV ((R, H)–C=O bond48), 288.7 eV (N–C=O49,50), and 289.5 eV (O=C–OH8). These species are present in different amino acids and proteins within the structure of VLPs, such as spike proteins, nucleocapsid proteins, and lipid envelopes. However, since the VLPs sample contains CCM, the adsorbed CCM on the surface of TiO2(101) was separately characterized using XPS to distinguish the contributions from the VLPs and those from the CCM. Figure 3c,d shows the C 1s and N 1s core-level spectra for the VLPs and CCM samples as well as the corresponding difference spectra. These results can be compared with those we reported in our previous study on the adsorption of the SARS-CoV-2 virus on the surface of anatase TiO2(101).8 In the case of the core levels of C atoms, the spectrum of the VLPs agrees with that measured for the inactivated virus, with slightly different BEs 285.4 vs 284.5, 287.1 vs 286.5 eV and 289.5 vs 287.9 eV but similar energy separation 1.7 vs 1.6 eV and 1.4 vs 2.4 eV. The minor discrepancies in the intensity and the position of the peaks can be ascribed to a different folding of the proteins since in the case of inactivated virus, they were partly denatured under the thermal treatment. In our previous study, we concluded, based on the results of the DFT calculation, that the XP C 1s core level carries the fingerprints of the SARS-CoV-2 adsorbed on TiO2(101). Therefore, we assume these results to be valid for the VLP, which has the same shell structure as SARS-CoV-2. In the case of the N 1s contributions, the deconvolution of the XP spectrum reveals five components, centered at 398.8, 400.1, 400.8, 401.8, and 402.9 eV, characteristic of C–N=C,51 –NH2/–NH,51 N-(C)3,51 N–C=O,52 and –C–N–C–,8 respectively. After the removal of the contribution of the CCM, among the features related to the VLPs only (Figure 3d at 400.1, 400.8, and 401.8 eV), the first and the last peaks have an energy separation similar to that measured for the inactivated virus (1.7 vs 1.9 eV). On the contrary, the intermediate peak is observed exclusively for the VLP, with a BE 0.7 eV higher than the feature with the lowest BE. We model the adsorbed VLP with two of the most abundant amino acids in the spike proteins, cysteine (cys), and asparagine (asn), similarly to what some of the authors did for the inactivated virus.8 These model structures are shown in Figure 3e–i. In addition, we calculated the XPS signature of a cys–cys dipeptide, and its optimized structure is displayed in Figure 3j, in order to include the prototypical contribution of the secondary amide group involved in the peptide bonds forming the amino-acid chains in the spike protein. In this case, we did not investigate any interaction between such a functional group and the surface, since we assumed that the N amide was not exposed to the environment by the folded protein. The positions of the simulated XPS features are reported in Figure 3b through colored sticks. Our numerical approach only allows us to calculate the BE differences or core-level shift (CLS); therefore, we have aligned the lowest contributions measured experimentally with the fit component with the lowest BE calculated by DFT, while the numerical values of the CLS are collected in Table S2. Additional information about the CLS calculation can be found in the Supporting Information.
Figure 3.
C 1s (a) and N 1s (b) spectra of adsorbed CCM and VLPs on the surface of TiO2(101) and on top difference spectra of VLPs and CCM are included for the C 1s and N 1s core-level scans. Deconvoluted core-level photoelectron spectra of C 1s (c) and N 1s (d) for adsorbed VLPs on the surface of TiO2(101). In panel (b) the position of the peaks calculated for the different N species and adsorption modes is marked by colored sticks. The structure of the optimized DFT models of the simulated adsorbed cys or asn molecules as well as that of the isolated dipeptide are shown in panels (e–j). N, O, C, S, Ti, and H atoms are shown by blue, red, gray, yellow, (larger) gray, and white spheres, respectively. Each N atom is highlighted through circles, whose colors correspond to that of the sticks in panel (b). On all of the structures, chemical or hydrogen bonds are highlighted by solid or dashed black lines, respectively. The corresponding bond length is reported as well in Å.
The analysis of the correlation between the CLS of the different N atoms and their interactions with the surroundings allows us to effectively separate the calculated XP peak contributions into three groups, which qualitatively agree with those peaks deconvoluted from the experimental spectrum. In the first group, which we assign to the feature centered at 400.1 eV, we include the secondary amide group of the peptide bonds (as in the cys–cys model) and the amino groups with no interaction with the surface or other groups of the molecule itself (as in the cys02 and asn01 models). This attribution is supported by the fact that these CLSs are in agreement with the amino group of the isolated cys–cys dipeptide. In the second group, which we assign to the feature at the highest BE at 401.8 eV, we include the N groups, both amino and amide, close to a carbonyl group chemically bonded to a surface Ti atom (as in the cys03 model). Finally, we define a last group at intermediate BE, which we assign to the feature centered at 400.8 eV, where we include the amino and the amide groups that interact directly with the surface or other groups and also are located close to a carbonyl group bound to the surface as in cys01, cys04, and asn01 models. These atoms would experience two opposite effects, namely an increase of BE, triggered by the adsorbed carbonyl group nearby, and a red shift in BE, due to the formation of H-bonds as previously was observed in H-bonded amino groups in carbon nitride polymers.53 Therefore, these results support the adsorption of VLPs with spike proteins and that our synthesized VLPs can be considered reliable for studying the adsorption of the SARS-CoV-2 virus on the surface of TiO2(101).
UV-Induced Transformation of SARS-CoV-2 VLPs/TiO2(101) System
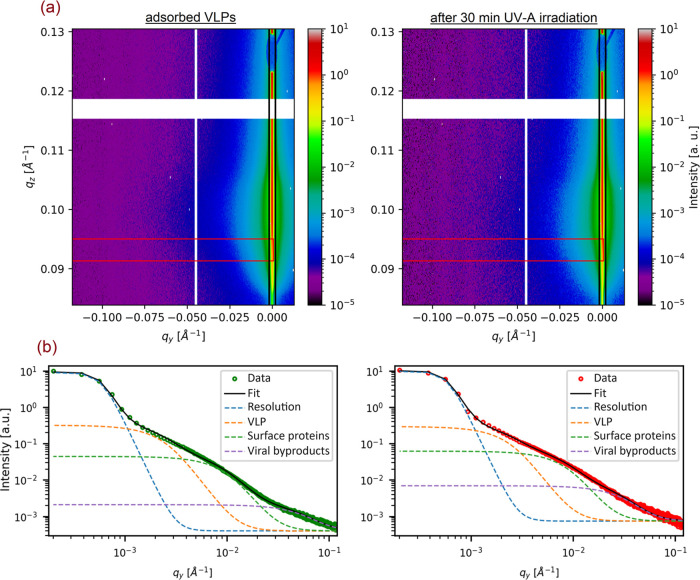
Recently, we discovered that UV-C (wavelength: 265 nm) can effectively inactivate SARS-CoV-2 adsorbed on a TiO2(101) surface through viral genome damage and photocatalytic oxidation.8 Since we characterized the sample after applying the inactivation method, we were unable to distinguish the photocatalytic inactivation mechanisms from the denaturation processes. Understanding the role of surface chemistry in virus inactivation is crucial, especially for the development of efficient self-cleaning materials for surface cleaning. To understand the role of TiO2(101) surface in the inactivation of the virus under UV light, we performed AFM and grazing-incidence small-angle X-ray scattering (GISAXS) experiments before and after UV treatment. These microscopic and scattering techniques provide us with more insights into the effect of light on the virus at the oxide interface, helping to understand the possible catalytic inactivation mechanisms of the virus. The GISAXS experiments were performed at beamline P03 of the synchrotron facility PETRA III (DESY, Hamburg) (see characterization techniques section).54,55 After loading the VLPs onto the surface of TiO2(101) (single crystal 8 mm × 8 mm × 2 mm), a GISAXS pattern was collected at room temperature in the air before exposing the sample to UV-A light (wavelength: 365 nm, intensity: 850 μW/cm2) and again after 30 min of UV exposure. To reduce the effects of temperature and light damage during sample transfer from the biosafety level 2 (BSL-2) laboratory to the beamline, the samples were transported in a closed dry ice bath to avoid viral protein denaturation. For studying the interaction of X-rays with the VLPs, one spot on the sample surface was continuously illuminated by the high-flux X-ray beam for 20 s (see Figure S7 in Supporting Information). During this time, a detector image (Figure 4a) was taken every 0.1 s. From each image, a horizontal (red box in Figure 4a) and vertical cut (black box in Figure 4a) was extracted. The position and width of the horizontal and vertical cuts were chosen to visualize X-ray-induced changes in the VLP morphology and sample composition, respectively. It is evident that continuous illumination for 20 s (see Figure S7), results in no X-ray-induced changes.
Figure 4.
(a) 2D GISAXS data of the adsorbed VLPs on the surface of TiO2(101) and after 30 min of UV irradiation in air. The black and red boxes mark the vertical and horizontal cut position, respectively. (b) Horizontal line cuts at the region of interest (red box in (a)) with the corresponding fit.
In order to not limit the study of the VLP morphology to one spot (perpendicular × parallel to the beam direction: 0.03 × 5 mm2) but over the whole sample surface, a total of 50 scattering images (illumination time: 0.1 s) were collected with a spacing of 100 μm between them over the whole VLP-loaded surfaces before and after the UV treatment. Consequently, all images were summed to provide averaged information regarding the VLP morphology across the whole sample surfaces. Since the main focus was on the in-plane structural changes of the VLP, only the horizontal cuts (red box in Figure 4a) were analyzed. Here, the proteins and VLPs sizes and size distribution were determined using the so-called effective interface approximation of the distorted wave Born approximation.56−58 For modeling, a combination of a Lorentzian resolution function (centered at qy = 0 Å–1) and three particle populations were assumed, which were described using spherical form factors with a Gaussian size distribution known from general small-angle X-ray scattering theory.59
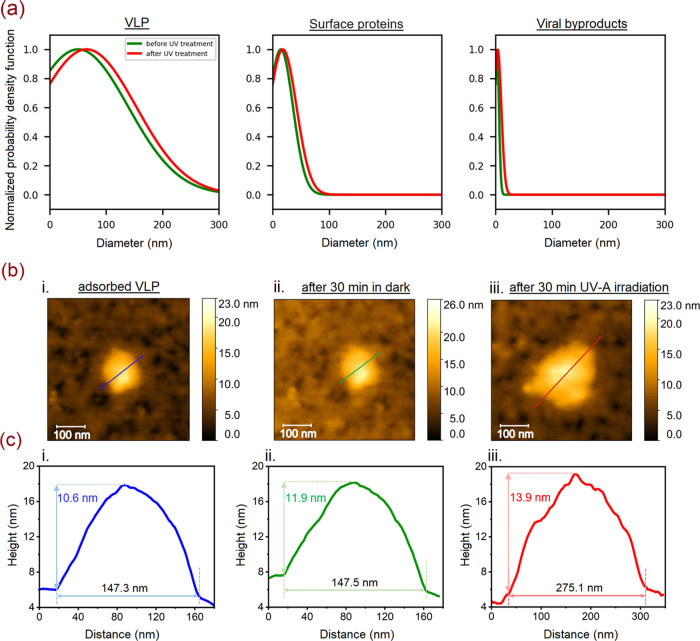
Considering the denaturation of spike proteins and other viral proteins in the virus structure during the adsorption process on the surface of TiO2(101),8 we examined three different models to fit data to cover all particles adsorbed on the surface, (i) VLPs, (ii) surface proteins (the spike, membrane, and envelope proteins), and (iii) viral byproducts. In contrast to the clean surface of TiO2(101), the intensity of the GISAXS curve for adsorbed VLPs is high, confirming the adsorption of VLPs (see Figure S9 in the Supporting Information). Based on the GISAXS curves, the intensity changed after UV treatment. Since the bulk beam does not affect the virus morphology (according to the beam damage scan results), the change in intensity is attributed to the effect of UV irradiation on the virus structure. By fitting the GISAXS curve (Figure 4b), we determined the average diameters of VLPs, surface proteins, and viral byproducts as 50.2, 14.4, and 2.5 nm, respectively (Figure 5a, for more information, see Table S3). After exposing the system to light, we observed an increase in the average diameter of both VLPs and surface proteins, suggesting that UV light has an effect on the size of the adsorbed particles. To delve deeper into the effect of UV irradiation, we performed a similar experiment using AFM and measured one adsorbed VLP before and after exposing the sample to light (Figure 5b). After VLPs were loaded onto the surface of TiO2(101), one adsorbed virus particle was measured with AFM, which exhibited a spherical particle with a size of 147.5 nm (see the line scan in Figure 5c). A control experiment was carried out by keeping this sample exposed to air and in dark conditions for 30 min, followed by characterizing the same particle using AFM. During the control experiment, the target particle exhibited no changes in shape and size and stayed intact (Figure 5b,c-ii). However, after 30 min of exposure to UV-A (Wavelength: 365 nm), the diameter of the particles increased and its spherical structure disappeared (Figure 5b,c-iii). The UV-A exposed VLP/TiO2(101) sample showed a particle with a size of 275.1 nm. Based on the line scan profile, the observed island on the line depicted in Figure 5c-iii exhibits a size ranging from 15 to 25 nm. This range of particle size may correspond to the surface proteins that dissociated from the virus during the photocatalytic reaction and/or as a result of the viral protein denaturation process at the surface of the oxide photocatalyst. According to the AFM results, the height of the selected particle decreased from 11.9 to 10.6 nm after the sample was kept in the dark due to the Vroman effect. However, after exposing the particle to UV-A light for 30 min, its height increased and reached 13.9 nm. This increase could be related to the polymerization of the dissociated surface proteins or membrane rupture during UV irradiation. The in situ AFM results, in line with the GISAXS findings, elucidate the photoinduced changes in the VLPs morphology on the TiO2(101) surface. When we conducted experiments both in dark conditions and under UV irradiation in air, no changes were observed in VLP morphology without UV light. This suggests that changes in VLP morphology occur only under oxygen pressure (air) combined with UV irradiation. There are two reactions that can take place on the surface of TiO2/VLPs sample under light and air exposure: (i) the denaturation of structural proteins induced by UV irradiation, leading to readsorption and cross-linking interactions between amino acids and (ii) the oxidation of the virus structure through photocatalytic reactions by producing ROS in the interaction of air/VLP/TiO2. In both cases, TiO2 acts as a photocatalyst providing an active surface for the photoinduced changes in VLPs. Protein denaturation refers to the alteration of a protein’s native structure, leading to the loss of its functional conformation. This can manifest as changes in size, including unfolding or aggregation of the protein. Additionally, the presence of ROS at the surface of the catalyst can induce modifications in the morphology of microorganisms.19 ROS have been reported to induce oxidative damage in viruses and bacteria, affecting various components such as proteins, RNA, DNA, and lipid membranes.60 ROS can impact the lipids, RNA, and cell membrane by selectively attacking nucleotides and sulfhydryl bonds. The result will be the breakdown of the cell wall, which subsequently leads to changes in the size and shape of the virus structure. For instance, it has been reported that the interaction between HO• radicals and Bacillus spore cells can disrupt their structure, changing their spherical shape and causing cell death.19 Additionally, the photogenerated holes in the valence band of TiO2 exhibit potent oxidizing properties capable of oxidizing the surface proteins, subsequently deactivating them.61 Air humidity can significantly affect the generation of hydroxyl radicals. In the presence of humidity, water molecules adsorb onto the surface of TiO2 and increase the surface hydroxyl groups after dissociation.62 These hydroxyl groups can then be transformed into radicals under UV irradiation, which enhances the photocatalytic reaction. The air humidity during the measurements was 65% in Hamburg, Germany.
Figure 5.
(a) Particle size distribution of VLP, surface proteins, and viral byproducts obtained from fitted GISAXS scattering curves in Figure 4b. (b) AFM topography images (0.5 × 0.5 μm2) of a single selected adsorbed VLP on the surface of TiO2(101) (i), after 30 min in dark and air (ii), and after UV irradiation in air for 30 min (iii). (c) Line scan profile of the particles in the corresponding AFM topography images in (b).
To confirm that the generated ROS are responsible for morphological changes through oxidation reactions, the effect of UV-A light on adsorbed VLPs was studied using the GISAXS method under an N2 atmosphere. After measuring the adsorbed VLPs in a N2 atmosphere, the sample was exposed to UV-A light while still under an N2 atmosphere to observe the effects of UV-A without oxygen on VLP morphology. GISAXS curves demonstrated no changes in the structure of VLPs under UV-A in an N2 atmosphere (see Figure S9 in the Supporting Information). This provides evidence that UV-A in the absence of oxygen cannot affect the morphology of adsorbed VLP on the surface of TiO2. Therefore, the presence of both light and oxygen is necessary to activate the surface of TiO2 and initiate photocatalytic oxidation. Our results suggest that oxygen and consequently the generated ROS on the surface of TiO2(101) play a critical role in inducing morphological changes and subsequently inactivating the virus through membrane oxidation.
Conclusions
We demonstrated the adsorption of SARS-CoV-2 VLPs on the surface of TiO2 as a photo active catalytic surface. XPS analysis revealed that the virus can interact with the surface by binding between amine (–NH2) and amide (–O=C–NH–) groups in the protein block structure, which was also confirmed by theoretical DFT simulations. We then used the TiO2(101)/VLP system under ambient conditions to observe the effects of UV-A light on VLP morphology. Our in situ GISAXS and AFM measurements confirmed the photoinduced changes in the morphology of the adsorbed VLPs. This is primarily due to the generation of ROS, which we propose contributes to the oxidation of the envelope protein in the virus structure. Our study elucidates the role of oxygen and subsequent ROS generation in inducing morphological changes in adsorbed VLPs at the surface of TiO2, leading to the effective inactivation of the virus. To the best of our knowledge, this study represents the first report providing visual confirmation of the structural damage and morphological alterations of SARS-CoV-2 VLPs at the surface of a solid photocatalyst, using AFM and GISAXS. These results emphasize the potential of such approaches in developing strategies for mitigating viral transmission in various environments.
Experimental Section
Preparation of SARS-CoV-2 Virus-Like Particles (VLPs)
To generate S+ VLPs containing appropriate levels of SARS-CoV-2 B.1 spike (D614G) and SVLPs controls, HEK293T cells were transfected using TransIT-293 (MIR2700, Mirus) according to the manufacturer’s protocol with carefully adjusted ratios of codon-optimized expression vectors coding for spike (S), membrane (M), nucleocapsid (N), and envelope (E) proteins. The expression plasmids are termed p7413.1 (S:D614G), p5025 (mock), p7395.LA3 (M), p7396.NA9 (E), and p7391.MA5 (N), respectively (Figure S1, see the Supporting Information). Media exchange was performed 6 h after the addition of the transfection mix to reduce a potential carryover of plasmid DNA. After 72 h, S+ VLPs, S VLPs, or control EVs were harvested from cell culture supernatants (DMEM, 8% FBS, supplemented with Pen/Strep, Gibco, Thermo Fisher Scientific) after low-speed centrifugation at 7 °C for 10 min at 300g and 20 min at 4200g and generally used without further processing. Particle preparations were flash frozen in liquid nitrogen for storage at −80 °C and subsequently characterized as described by Roessler et al.25
Characterization Techniques
Fluorescence Microscopy (FM)
To understand the specific and nonspecific binding of VLPs on the surface of TiO2, the sample was characterized by FM. In order to reduce nonspecific antibody binding, the sample including SARS-CoV-2 VLPs was covered with blocking solution phosphate-buffered saline (PBS) supplemented with saponin (0.25%) and bovine serum albumin (BSA, 3.0%). After 30 min, the sample was washed three times with PBS. Then, 20 μL of blocking solution containing 1:200 diluted primary antibody (anti-SARS-CoV-2 nucleocapsid antibody produced in rabbit, Thermo Fisher Scientific) was added to the sample. After incubation time (60 min), the sample was washed with PBS to remove unbound antibody, and then, blocking solution containing 1:500 fluorescent-dye conjugated secondary antibody (antirabbit antibody conjugated with Alexa568, Thermo Fisher Scientific) was added. After incubation (30 min) and washing (three times with PBS), the samples were examined with a fluorescence microscope (Leica DMi8 equipped with a 20× NA 0.4 air objective, a Leica DFC9000 GT sCMOS camera, and a Lumencore Sola SE FISH 365 LED light source). Figure S3 shows the sample preparation process for FM measurements (see the Supporting Information file).
Atomic Force Microscopy (AFM)
The morphology of adsorbed SARS-CoV-2 VLPs on the surface of TiO2(101) was investigated by AFM at DESY NanoLab.30 The AFM and phase images were recorded with different scan sizes (1.5 × 1.5 μm2 and 0.5 × 0.5 μm2) and 0.996 Hz rate under tapping mode. A RTESPA-300 silicon cantilever from Bruker with a nominal tip radius of 8 nm was used for the measurements. The visualization of measurements were done by the Gwyddion software package.63
AFM-Based Nanoinfrared (Nano-IR) Spectroscopy
Nano-IR spectroscopy (Anasys Instruments-Bruker) was used to probe the structural composition of adsorbed VLPs on the surface of TiO2(101) with nanoscale resolution. AFM-IR spectra were obtained with gold-coated dielectric probes (nominal tip radius: 20 nm) in tapping mode at a scan rate of 0–5 Hz. Analysis Studio and Gwyddion software63 were used to export the IR spectra and AFM image, respectively.
X-Ray Photoelectron Spectroscopy (XPS)
The XPS measurements of adsorbed VLPs on the surface of TiO2(101) was carried out using the XPS system at the DESY Nanolab, at the Centre for X-ray and Nano Science.30 The X-ray source employed was Al Kα at 1486.6 eV and a Phoibos 150 hemispherical energy analyzer with a base pressure of 1.2 × 10–10 mbar.
Computational Methods
We performed all calculations within the density functional theory (DFT) framework, using the QuantumESPRESSO suite.64,65 We modeled the interaction of a SARS-CoV-2 spike protein with the TiO2 anatase (101) considering different adsorption configurations of two of its most abundant amino acids, i.e., cysteine (cys) and asparagine (asn), similarly to those included in a previous work of some of the authors.8 In addition, we included in this work the model of an isolated prototypical molecule including the amide bond, i.e., the cys–cys dimer. A sketch of the three isolated molecules is shown in Figure S6 (see the Supporting Information file). For the ground state of all our models, we employed a plane-waves basis set with a cutoff of 52 and 575 Ry for the wave functions and the charge density, respectively, and the PBE exchange-correlation functional. We included the dispersion interactions through the Grimme-D3 pairwise correction.66 We modeled the TiO2 anatase (101) substrate as a slab including three triatomic layers, with lattice parameters fixed to those of the optimized bulk unit cell (a = 3.790 Å and b = 10.325 Å). We separated one slab from its periodic replicas along the z direction perpendicular to the surface by inserting a vacuum region 11 Å thick, and we optimized the two topmost triatomic layers, keeping the lowest layer of atoms fixed. The reciprocal space was sampled through a shifted Monkhorst–Pack 2 × 2 × 1 k-points mesh.
The simulation of the XPS features at the N K edge was carried out by calculating the core-level shifts (CLSs) using the ΔSCF method.67 We employed an additional N pseudopotential including a full core hole at its 1s state, requiring an extended basis set cutoff of 74 Ry for the wave functions. Since absolute values of the binding energy are forbidden with this methodology but only relative energy differences (the CLSs), we provided a common energy reference for all the models by including a N2 molecule far from the surface. Therefore, all supercells were enlarged along one (in the case of the adsorbed molecules) or more (in the case of isolated molecules) lattice directions in order to obtain a separation from the added reference molecule of at least 15 Å. Such an approach already gave satisfactory results in previous works.68,69 The CLS calculated for the different N species in our models are reported in Table S2.
Grazing-Incidence Small-Angle X-Ray Scattering (GISAXS)
For the GISAXS measurements, the X-ray wavelength was fixed at 1.05 Å (E = 11.8 keV) and the X-ray beam was focused to a 25 × 30 μm2 (vertical × horizontal direction) spot. The scattered photons were collected by using a Pilatus 2 M detector (DECTRIS, Baden, Switzerland), which was positioned roughly 5.5 m away from the samples. The incident angle was fixed at 0.4 deg for all measurements.
Acknowledgments
This study was supported by the initiative and networking fund of the Helmholtz Association of German Research Centers under the CORAERO Project (Grant KA1-Co-06). We acknowledge DESY (Hamburg, Germany), a member of the Helmholtz Association HGF, for the provision of experimental facilities. Parts of this research were carried out at PETRA III, and we would like to thank Andrei Chumakov and Jan Rubeck for assistance in using beamline P03. Beamtimes were allocated for proposals I-20211020 and I-20230640. The conductance of the Nano-IR spectroscopy analysis by Hartmut Stadler, Bruker Nano GmbH, Karlsruhe, Germany, is greatly acknowledged. C.D.V. acknowledges funding from the European Union—NextGenerationEU through the Italian Ministry of University and Research under PNRR—M4C2I1.4 ICSC—Centro Nazionale di Ricerca in High Performance Computing, Big Data and Quantum Computing (Grant No. CN00000013).
Glossary
Abbreviations
- SARS-CoV-2
severe acute respiratory syndrome coronavirus 2
- UV
ultraviolet
- RNA
ribonucleic acid
- TiO2
titanium dioxide
- ROS
reactive oxygen species
- O2•–
superoxide radical
- HO•
hydroxyl radical
- BSL-3
biosafety level 3
- VLPs
SARS-CoV-2 virus-like particles
- FM
fluorescence microscopy
- AFM
atomic force microscopy
- XPS
X-ray photoelectron spectroscopy
- UHV
ultra high vacuum
- CCM
cell culture media
- DFT
density functional theory
- cys
cysteine
- asn
asparagine
- CLS
core-level shift
- GISAXS
grazing-incidence small-angle X-ray scattering
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/acsami.4c07151.
Schematic structure of SARS-CoV-2 and SARS-CoV-2 virus-like particles (VLPs). LEED and XP spectra of the clean anatase TiO2(101) surface; sample preparation process for fluorescence microscopy (FM) measurements and FM signal for different experiments; AFM topographic image of a selected area on the surface for nano-IR measurement; deconvoluted O 1s and Ti 2p core-level XP spectrum of adsorbed VLPs on TiO2(101) surface showing the presence of different functional groups of virus proteins, which interact with the surface of the catalyst. Sketch of isolated cysteine and asparagine monopeptides and cysteine dimer with corresponding calculated core-level shifts (CLS) for different N species in these models; beam damage scan showing the bulk beam cannot affect on virus morphology during the GISAXS measurements; GISAXS detector image showing the good sample alignment at the beamline during the GISAXS measurements; GISAXS data fitting parameters for adsorbed VLPs on TiO2 surface before and after UV-A irradiation are reported in Table S3; horizontal line cuts at the region of interest in 2D GISAXS for the adsorbed VLPs and after 30 min UV-A irradiation under air and N2 showing that the presence of O2 and UV light are necessary to make the surface of TiO2 active (PDF)
Author Contributions
M.K.: sample preparation, FM, AFM, XPS, GISAXS, analysis of all data, and manuscript writing; A.U.: DFT and manuscript writing (DFT section); B.S.: GISAXS and analysis of GISAXS data; J.R.: VLP preparation; M.W.: XPS and GISAXS; A.M.: Nano-IR and GISAXS; E.E.B.: GISAXS; D.S.D.: GISAXS; M.B.G.: GISAXS; M.C.: GISAXS; T.F.K.: AFM; M.S.: GISAXS; S.K.V.: GISAXS; R.T.: FM; G.G.: sample preparation; C.L.: sample preparation; G.E.: virus loading; U.P.: virus loading; W.H.: VLP preparation; R.Z.: VLP preparation; S.V.R.: GISAXS and supervising analysis of GISAXS data; C.D.V.: DFT; A.S.: GISAXS; and H.N.: designed and performed experiments and supervised the project, analysis of data, manuscript writing, and funding acquisition. All the authors commented on the manuscript and have given approval to the final version of the manuscript.
The authors declare no competing financial interest.
Supplementary Material
References
- Prakash J.; Cho J. D.; Mishra Y. Photocatalytic TiO2 nanomaterials as potential antimicrobial and antiviral agents: Scope against blocking the SARS-COV-2 spread. Micro Nano Eng. 2022, 14, 100100 10.1016/j.mne.2021.100100. [DOI] [Google Scholar]
- Corman V. M.; Guggemos W.; Seilmaier M.; Zange S.; Müller M. A.; Niemeyer D.; Jones T. S.; Rothe C.; Hoelscher M.; Bleicker T.; Brünink S.; Schneider J.; Ehmann R.; Zwirglmaier K.; Drosten C.; Wendtner C.; et al. Virological assessment of hospitalized patients with COVID-2019. Nature 2020, 581, 465–469. 10.1038/s41586-020-2196-x. [DOI] [PubMed] [Google Scholar]
- Kampf G.; Todt D.; Pfaender S.; Steinmann E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. 10.1016/j.jhin.2020.01.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hasan J.; Pyke A.; Nair N.; Yarlagadda T.; Will G.; Spann K.; Yarlagadda P. Antiviral Nanostructured Surfaces Reduce the Viability of SARS-CoV-2. ACS Biomater. Sci. Eng. 2020, 6 (9), 4858–4861. 10.1021/acsbiomaterials.0c01091. [DOI] [PubMed] [Google Scholar]
- Celik U.; Celik K.; Celik S.; Abayli H.; Sahna K. C.; Tonbak S.; Toraman Y. A.; Oral A. Interpretation of SARS-CoV-2 behaviour on different substrates and denaturation of virions using ethanol: an atomic force microscopy study. RSC Adv. 2020, 10, 44079–44086. 10.1039/D0RA09083B. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Storm N. G. A.; McKay L.; Downs S. N.; Johnson R. I.; Birru D.; de Samber M.; Willaert W.; Cennini G.; Griffiths A. Rapid and complete inactivation of SARS-CoV-2 by ultraviolet-C irradiation. Sci. Rep. 2020, 10, 22421 10.1038/s41598-020-79600-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo C. W.; Matsuura R.; Iimura K.; Wada S.; Shinjo A.; Benno Y.; Nakagawa M.; Takei M.; Aida Y. UVC disinfects SARS-CoV-2 by induction of viral genome damage without apparent effects on viral morphology and proteins. Sci. Rep. 2021, 11, 13804 10.1038/s41598-021-93231-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohantorabi M.; Wagstaffe M.; Creutzburg M.; Ugolotti A.; Kulkarni S.; Jeromin A.; Krekeler T.; Feuerherd M.; Herrmann A.; Ebert G.; Protzer U.; Guédez G.; Löw C.; Thuenauer R.; Schlueter C.; Gloskovskii A.; Keller T. F.; Di Valentin C.; Stierle A.; Noei H. Adsorption and Inactivation of SARS-CoV-2 on the Surface of Anatase TiO2(101). ACS Appl. Mater. Interfaces 2023, 15 (6), 8770–8782. 10.1021/acsami.2c22078. [DOI] [PubMed] [Google Scholar]
- Nakano R.; Yamaguchi A.; Sunada K.; Nagai T.; Nakano A.; Suzuki Y.; Yano H.; Ishiguro H.; Miyauchi M. Inactivation of various variant types of SARS-CoV-2 by indoor-light-sensitive TiO2-based photocatalyst. Sci. Rep. 2022, 12, 5804 10.1038/s41598-022-09402-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun Z.; Ostrikov K. Future antiviral surfaces: Lessons from COVID-19 pandemic. Sustainable Mater. Technol. 2020, 25, e00203. 10.1016/j.susmat.2020.e00203. [DOI] [Google Scholar]
- Vasickova P.; Pavlik I.; Verani M.; Carducci A. Issues concerning survival of viruses on surfaces. Food Environ. Virol. 2010, 2, 24–34. 10.1007/s12560-010-9025-6. [DOI] [Google Scholar]
- Rakowska P. D.; Tiddia M.; Faruqui N.; Bankier C.; Pei Y.; Pollard A. J.; Zhang J.; Gilmore I. S. Antiviral surfaces and coatings and their mechanisms of action. Commun. Mater. 2021, 2, 53. 10.1038/s43246-021-00153-y. [DOI] [Google Scholar]
- Yoshizawa N.; Ishihara R.; Omiya D.; Ishitsuka M.; Hirano S.; Suzuki I. Application of a Photocatalyst as an Inactivator of Bovine Coronavirus. Viruses 2020, 12 (12), 1372. 10.3390/v12121372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abou Saoud W.; Kane A.; Le Cann P.; Gerard A.; Lamaa L.; Peruchon L.; Brochier C.; Bouzaza A.; Wolbert D.; Assadi A. A. Innovative photocatalytic reactor for the degradation of VOCs and microorganism under simulated indoor air conditions: Cu-Ag/TiO2-based optical fibers at a pilot scale. Chem. Eng. J. 2021, 411, 128622. 10.1016/j.cej.2021.128622. [DOI] [Google Scholar]
- Elgohary E. A.; A Mohammad Y. M.; ElNazar H. A.; Baaloudj O. S. S.; Alyami M.; El Jery A.; Assadi A. A.; Amrane A. A Review of the Use of Semiconductors as Catalysts in the Photocatalytic Inactivation of Microorganisms. Catalysts 2021, 11 (12), 1498. 10.3390/catal11121498. [DOI] [Google Scholar]
- Assadi I.; Guesmi A.; Baaloudj O.; Baaloudj O.; Zeghioud H.; Elfalleh W.; Benhammadi N.; Khezami L.; Assadi A. A. Review on inactivation of airborne viruses using non-thermal plasma technologies: from MS2 to coronavirus. Environ. Sci. Pollut. Res. 2022, 29, 4880–4892. 10.1007/s11356-021-17486-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diebold U. The surface science of titanium dioxide. Surf. Sci. Rep. 2003, 48 (5–8), 53–229. 10.1016/S0167-5729(02)00100-0. [DOI] [Google Scholar]
- Wagstaffe M.; Dominguez-Castro A.; Wenthaus L.; Palutke S.; Kutnyakhov D.; Heber M.; Pressacco F.; Dziarzhytski S.; Gleißner H.; Gupta V. K.; Redlin H.; Dominguez A.; Frauenheim T.; Rubio A.; Stierle A.; Noei H. Photoinduced Dynamics at the water/TiO2(101) Interface. Phys. Rev. Lett. 2023, 130, 108001. 10.1103/PhysRevLett.130.108001. [DOI] [PubMed] [Google Scholar]
- Kumar A.; Soni V.; Singh P.; Khan A. A. P.; Nazim M.; Mohapatra S.; Saini V.; Raizada P.; Hussain C. M.; Shaban M.; Marwani H. M.; Asiri A. M. Green aspects of photocatalysts during corona pandemic: a promising role for the deactivation of COVID-19 virus. RSC Adv. 2022, 12, 13609–13627. 10.1039/D1RA08981A. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hosseini M.; Chin A. W. H.; Behzadinasab S.; Poon L. L. M.; Ducker W. A. Cupric Oxide Coating That Rapidly Reduces Infection by SARS-CoV-2 via Solids. ACS Appl. Mater. Interfaces 2021, 13 (5), 5919–5928. 10.1021/acsami.0c19465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva D. J.; Duran A.; Cabral A. D.; Fonseca F. A. L.; Bueno R. F.; Wang S. H.; Rosa D. S. Delta SARS-CoV-2 inactivation and bactericidal performance of cotton wipes decorated with TiO2/Ag nanoparticles like Brazilian heavy-fruited Myrciaria cauliflora. Mater. Today Commun. 2022, 33, 104288. 10.1016/j.mtcomm.2022.104288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva D. J.; Duran A.; Cabral A. D.; Fonseca F. A. L.; Bueno R. F.; Rosa D. S. Questioning ZnO, Ag, and Ag/ZnO nanoparticles as antimicrobial agents for textiles: Do they guarantee total protection against bacteria and SARS-CoV-2?. J. Photochem. Photobiol., B 2022, 234, 112538. 10.1016/j.jphotobiol.2022.112538. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuura R.; Lo C. W.; Wada S.; Somei J.; Ochiai H.; Murakami T.; Saito N.; Ogawa T.; Shinjo A.; Benno Y.; Nakagawa M.; Takei M.; Aida Y. SARS-CoV-2 Disinfection of Air and Surface Contamination by TiO2 Photocatalyst-Mediated Damage to Viral Morphology, RNA, and Protein. Viruses 2021, 13 (5), 942. 10.3390/v13050942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatsuma T.; Nakakido M.; Ichinohe T.; Kuroiwa Y.; Tomioka K.; Liu C.; Miyamae N.; Onuki T.; Tsumoto K.; Hashimoto K.; Wakihara T. Inactivation and spike protein denaturation of novel coronavirus variants by CuxO/TiO2 nano-photocatalysts. Sci. Rep. 2023, 13, 4033 10.1038/s41598-023-30690-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roessler J.; Pich D.; Albanese M. R.; Wratil P.; Krähling V.; Hellmuth J.; Scherer C.; von Bergwelt-Baildon M.; Becker S. T.; Keppler O.; Brisson A.; Zeidler R.; Hammerschmidt W. Quantitation of SARS-CoV-2 neutralizing antibodies with a virus-free, authentic test. PNAS Nexus 2022, 1, pgac045. 10.1093/pnasnexus/pgac045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Syed A. M.; Taha T. Y.; Tabata T.; Chen I. P.; Ciling A.; Khalid M. M.; Sreekumar B.; Chen P. Y.; Hayashi J. M.; Soczek K. M.; Ott M.; Doudna J. A. Rapid assessment of SARS-CoV-2-evolved variants using virus-like particles. Science 2021, 374 (6575), 1626–1632. 10.1126/science.abl6184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roessler J.; Pich D.; Krähling V.; Becker S. T.; Keppler O.; Zeidler R.; Hammerschmidt W. SARS-CoV-2 and Epstein-Barr Virus-like Particles Associate and Fuse with Extracellular Vesicles in Virus Neutralization Tests. Biomedicines 2023, 11 (11), 2892. 10.3390/biomedicines11112892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller D. J.; Amrein M.; Engel A. Adsorption of Biological Molecules to a Solid Support for Scanning Probe Microscopy. J. Struct. Biol. 1997, 119 (21), 172–188. 10.1006/jsbi.1997.3875. [DOI] [PubMed] [Google Scholar]
- Putlyaeva L. V.; Lukyanov A. K. Studying SARS-CoV-2 with Fluorescence Microscopy. Int. J. Mol. Sci. 2021, 22 (12), 6558. 10.3390/ijms22126558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stierle A.; Keller T. F.; Noei H.; Vonk V.; Roehlsberger R. DESY NanoLab. J. Large-Scale Res. Facil. 2016, 2, 76. 10.17815/jlsrf-2-140. [DOI] [Google Scholar]
- Xin Y.; Grundmeier G.; Keller A. Adsorption of SARS-CoV-2 Spike Protein S1 at Oxide Surfaces Studied by High-Speed Atomic Force Microscopy. Adv. Nanobiomed. Res. 2021, 1 (2), 2000024 10.1002/anbr.202000024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slack S. M.; Horbett T. A.. The Vroman Effect: A Critical Review; ACS Publications, 1995; Vol. 8, pp 112–128. [Google Scholar]
- Lundström I. Models of protein adsorption on solid surfacesw. Prog. Colloid Polym. Sci. 1985, 70, 76–82. 10.1007/BFb0114308. [DOI] [Google Scholar]
- Akkız H. The Biological Functions and Clinical Significance of SARS-CoV-2 Variants of Corcern. Front. Med. 2022, 9, 849217. 10.3389/fmed.2022.849217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turoňová B.; Sikora M.; Schürmann C. J. H.; Hagen W.; Welsch S. E. C.; Blanc F.; von Bülow S.; Gecht M.; Bagola K.; Hörner C.; Zandbergen G.; Landry J.; Trevisan Doimo de Azevedo N.; Mosalaganti S.; Schwarz A.; Covino R.; Mühlebach M.; Hummer G.; Krijnse Locker J.; Beck M. In situ structural analysis of SARS-CoV-2 spike reveals flexibility mediated by three hinges. Science 2020, 370 (6513), 203–208. 10.1126/science.abd5223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asandei A.; Mereuta L.; Schiopu I.; Park J.; Chang H. S.; Park Y.; Luchian T. Non-Receptor-Mediated Lipid Membrane Permeabilization by the SARS-CoV-2 Spike Protein S1 Subunit. ACS Appl. Mater. Interfaces 2020, 12 (50), 55649–55658. 10.1021/acsami.0c17044. [DOI] [PubMed] [Google Scholar]
- Lyonnais S.; Hénaut M.; Neyret A.; Merida P.; Cazevieille C.; Gros N.; Chable-Bessia C.; Muriaux D. Atomic force microscopy analysis of native infectious and inactivated SARS-CoV-2 virions. Sci. Rep. 2021, 11, 11885 10.1038/s41598-021-91371-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mathurin J.; Deniset-Besseau A.; Bazin D.; Dartois E.; Wagner M.; Dazzi A. Photothermal AFM-IR spectroscopy and imaging: Status, challenges, and trends. J. Appl. Phys. 2022, 131, 010901 10.1063/5.0063902. [DOI] [Google Scholar]
- Farber C.; Wang R.; Chemelewski R.; Mullet J.; Kurouski D. Nanoscale Structural Organization of Plant Epicuticular Wax Probed by Atomic Force Microscope Infrared Spectroscopy. Anal. Chem. 2019, 91 (3), 2472–2479. 10.1021/acs.analchem.8b05294. [DOI] [PubMed] [Google Scholar]
- Chang T.; Babu R. P.; Zhao W.; Johnson C. M.; Hedström P.; Odnevall I.; Leygraf C. High-Resolution Microscopical Studies of Contact Killing Mechanisms on Copper-Based Surfaces. ACS Appl. Mater. Interfaces 2021, 13 (41), 49402–49413. 10.1021/acsami.1c11236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hull M. C.; Cambrea L. R.; Hovis J. S. Infrared Spectroscopy of Fluid Lipid Bilayers. Anal. Chem. 2005, 77 (18), 6096–6099. 10.1021/ac050990c. [DOI] [PubMed] [Google Scholar]
- Gamage S.; Howard M.; Makita H.; Cross B.; Hastings G.; Luo M.; Abate Y. Probing structural changes in single enveloped virus particles using nano-infrared spectroscopic imaging. PLoS One 2018, 13 (6), e0199112 10.1371/journal.pone.0199112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Movasaghi Z.; Rehman S.; ur Rehman I. Fourier Transform Infrared (FTIR) Spectroscopy of Biological Tissues. Appl. Spectrosc. Rev. 2008, 43 (2), 134–179. 10.1080/05704920701829043. [DOI] [Google Scholar]
- Dovbeshko G. I.; Chegel V. I.; Gridina N. Y.; Repnytska O. P.; Shirshov Y. M.; Tryndiak V. P.; Todor I. M.; Solyanik J. I. Surface enhanced IR absorption of nucleic acids from tumor cells: FTIR reflectance study. Biopolymers 2002, 67, 470–486. 10.1002/bip.10165. [DOI] [PubMed] [Google Scholar]
- Dou T.; Li Z.; Zhang J.; Evilevitch A.; Kurouski D. Nanoscale Structural Characterization of Individual Viral Particles Using Atomic Force Microscopy Infrared Spectroscopy (AFM-IR) and Tip-Enhanced Raman Spectroscopy (TERS). Anal. Chem. 2020, 92 (16), 11297–11304. 10.1021/acs.analchem.0c01971. [DOI] [PubMed] [Google Scholar]
- OßCallahan B.; Qafoku O.; Balema V.; Negrete O. A.; Passian A.; Engelhard M. H.; Waters K. M. Atomic Force Microscopy and Infrared Nanospectroscopy of COVID-19 Spike Protein for the Quantification of Adhesion to Common Surfaces. Langmuir 2021, 37 (41), 12089–12097. 10.1021/acs.langmuir.1c01910. [DOI] [PubMed] [Google Scholar]
- Kolanowska A.; Dzido G.; Krzywiecki M. M.; Tomczyk M.; Łukowiec D.; Ruczka S.; Boncel S. Carbon Quantum Dots from Amino Acids Revisited: Survey of Renewable Precursors toward High Quantum-Yield Blue and Green Fluorescence. ACS Omega 2022, 7 (45), 41165–41176. 10.1021/acsomega.2c04751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiu L.; Zhang R.; Zhang Y.; Li C.; Zhang Q.; Zhou Y. Superhydrophobic, mechanically flexible and recyclable reduced graphene oxide wrapped sponge for highly efficient oil/water separation. Front. Chem. Sci. Eng. 2018, 12 (3), 390–399. 10.1007/s11705-018-1751-6. [DOI] [Google Scholar]
- Sousa S. R.; Moradas-Ferreira P.; Barbosa M. A. TiO2 type influences fibronectin adsorption. J. Mater. Sci.: Mater. Med. 2005, 16, 1173–1178. 10.1007/s10856-005-4725-4. [DOI] [PubMed] [Google Scholar]
- Vanea E.; Simon V. XPS study of protein adsorption onto nanocrystalline aluminosilicate microparticles. Appl. Surf. Sci. 2011, 257 (6), 2346–2352. 10.1016/j.apsusc.2010.09.101. [DOI] [Google Scholar]
- Wang R.; Cao X.; Huang H.; Ji X.; Chen X.; Liu J.; Yan P.; Wei S.; Chen L.; Wang Y. Facile Chemical Vapor Modification Strategy to Construct Surface Cyano-Rich Polymer Carbon Nitrides for Highly Efficient Photocatalytic H2 Evolution. ChemSusChem 2022, 15, e202201575 10.1002/cssc.202201575. [DOI] [PubMed] [Google Scholar]
- Duderija B.; González-Orive A.; Ebbert C.; Neßlinger V.; Keller A.; Grundmeier G. Electrode Potential-Dependent Studies of Protein Adsorption on Ti6Al4V Alloy. Molecules 2023, 28, 5109. 10.3390/molecules28135109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ugolotti A.; Lanzilotto V.; Grazioli C.; Schio L.; Zamalloa-Serrano J. M.; Stredansky M.; Zhang T.; de Simone M.; Ferraro L.; Floreano L.; Coreno M.; Puglia C.; DiValentin C. In-Plane Hydrogen Bonds and Out-of-Plane Dipolar Interactions in Self-Assembled Melem Networks. J. Phys. Chem. C 2023, 127, 11307–11316. 10.1021/acs.jpcc.3c01990. [DOI] [Google Scholar]
- Roth S. V.; Herzog G.; Körstgens V.; Buffet A.; Schwartzkopf M.; Perlich J.; Abul Kashem M. M.; Döhrmann R.; Gehrke R.; Rothkirch A. In situ observation of cluster formation during nanoparticle solution casting on a colloidal film. J. Phys.: Condens. Matter 2011, 23, 254208 10.1088/0953-8984/23/25/25420. [DOI] [PubMed] [Google Scholar]
- Buffet A.; Rothkirch A.; Döhrmann R.; Körstgens V.; Abul Kashem M. M.; Perlich J.; Herzog G.; Schwartzkopf M.; Gehrke R.; Müller-Buschbaum P.; Roth S. V. P03, the microfocus and nanofocus X-ray scattering (MiNaXS) beamline of the PETRA III storage ring: the microfocus endstation. J. Synchrotron Radiat. 2012, 19, 647–653. 10.1107/S0909049512016895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller-Buschbaum P.; Metwalli E.; Moulin J. F.; Kudryashov V.; Haese-Seiller M.; Kampmann R. Time of flight grazing incidence small angle neutron scattering. Eur. Phys. J.: Spec. Top 2009, 167, 107–112. 10.1140/epjst/e2009-00944-5. [DOI] [Google Scholar]
- Frontini G. L.; Otero F.; Messineo M. G.; Eliçabe G. E. Estimation of size distribution in concentrated particle systems from light scattering measurements. Inverse Probl. Sci. Eng. 2008, 16 (8), 995–1004. 10.1080/00467600802082951. [DOI] [Google Scholar]
- Gazzillo D.; Giacometti A.; Guido Della Valle R.; Venuti E.; Carsughi F. A scaling approximation for structure factors in the integral equation theory of polydisperse nonionic colloidal fluids. J. Chem. Phys. 1999, 111 (16), 7636–7645. 10.1063/1.480089. [DOI] [Google Scholar]
- Pedersen J. S.; Gerstenberg M. C. Scattering Form Factor of Block Copolymer Micelles. Macromolecules 1996, 29 (4), 1363–1365. 10.1021/ma9512115. [DOI] [Google Scholar]
- Parra-Ortiz E.; Malmsten M. Photocatalytic nanoparticles—From membrane interactions to antimicrobial and antiviral effects. Adv. Colloid Interface Sci. 2022, 299, 102526. 10.1016/j.cis.2021.102526. [DOI] [PubMed] [Google Scholar]
- Bagheri S.; Julkapli N. M.; Hamid M. R. Y.; Ziaei R.; Sagadevan S. Nanomaterials Aspects for Photocatalysis as Potential for the Inactivation of COVID-19 Virus. Catalysts 2023, 13 (3), 620. 10.3390/catal13030620. [DOI] [Google Scholar]
- Zhang G.; Peyravi A.; Hashisho Z.; Sun Z.; Liu Y.; Zheng S.; Zhong L. Integrated adsorption and photocatalytic degradation of VOCs using a TiO2/diatomic composite: effects of relative humidity and reaction atmosphere. Catal. Sci. Technol. 2020, 10, 2378–2388. 10.1039/D0CY00168F. [DOI] [Google Scholar]
- Nečas D.; Klapetek P. Gwyddion: an open-source software for SPM data analysis. Open Phys. 2012, 10, 181–188. 10.2478/s11534-011-0096-2. [DOI] [Google Scholar]
- Giannozzi P.; Andreussi O.; Brumme T.; Bunau O.; Buongiorno Nardelli M.; Calandra; M Car R.; Cavazzoni C.; Ceresoli D.; Cococcioni M.; et al. Advanced capabilities for materials modelling with Quantum ESPRESSO. J. Phys.: Condens. Matter 2017, 29, 7465901 10.1088/1361-648X/aa8f79. [DOI] [PubMed] [Google Scholar]
- Giannozzi P.; Baseggio O.; Bonfà P.; Brunato D.; Car R.; Carnimeo I.; Cavazzoni C.; de Gironcoli S.; Delugas P.; Ferrari Ruffino F.; Ferretti A.; Marzari N.; Timrov I.; Urru; A Baroni S. Quantum ESPRESSO toward the exascale. J. Chem. Phys. 2020, 152, 154105. 10.1063/5.0005082. [DOI] [PubMed] [Google Scholar]
- Grimme S.; Antony J.; Ehrlich S.; Krieg H. A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J. Chem. Phys. 2010, 132, 154104. 10.1063/1.3382344. [DOI] [PubMed] [Google Scholar]
- García-Gil S.; García A.; Ordejón P. Calculation of core level shifts within DFT using pseudopotentials and localized basis sets. Eur. Phys. J. B 2012, 85, 239. 10.1140/epjb/e2012-30334-5. [DOI] [Google Scholar]
- Castrovilli M. C.; Bolognesi P.; Bodo E.; Mattioli G.; Cartoni A.; Avaldi L. An experimental and theoretical investigation of XPS and NEXAFS of 5-halouracils. Phys. Chem. Chem. Phys. 2018, 20, 6657–6667. 10.1039/C8CP00026C. [DOI] [PubMed] [Google Scholar]
- Ugolotti A.; di Valentin C. Ab-Initio Spectroscopic Characterization of Melem-Based Graphitic Carbon Nitride Polymorphs. Nanomaterials 2021, 11, 1863. 10.3390/nano11071863. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.