ABSTRACT
Epidermodysplasia verruciformis (EV) is a rare genetic skin disorder that is characterized by the development of papillomavirus-induced skin lesions that can progress to squamous cell carcinoma (SCC). Certain high-risk, cutaneous β-genus human papillomaviruses (β-HPVs), in particular HPV5 and HPV8, are associated with inducing EV in individuals who have a homozygous mutation in one of three genes tied to this disease: EVER1, EVER2, or CIB1. EVER1 and EVER2 are also known as TMC6 and TMC8, respectively. Little is known about the biochemical activities of EVER gene products or their roles in facilitating EV in conjunction with β-HPV infection. To investigate the potential effect of EVER genes on papillomavirus infection, we pursued in vivo infection studies by infecting Ever2-null mice with mouse papillomavirus (MmuPV1). MmuPV1 shares characteristics with β-HPVs including similar genome organization, shared molecular activities of their early, E6 and E7, oncoproteins, the lack of a viral E5 gene, and the capacity to cause skin lesions that can progress to SCC. MmuPV1 infections were conducted both in the presence and absence of UVB irradiation, which is known to increase the risk of MmuPV1-induced pathogenesis. Infection with MmuPV1 induced skin lesions in both wild-type and Ever2-null mice with and without UVB. Many lesions in both genotypes progressed to malignancy, and the disease severity did not differ between Ever2-null and wild-type mice. However, somewhat surprisingly, lesion growth and viral transcription was decreased, and lesion regression was increased in Ever2-null mice compared with wild-type mice. These studies demonstrate that Ever2-null mice infected with MmuPV1 do not exhibit the same phenotype as human EV patients infected with β-HPVs.
IMPORTANCE
Humans with homozygous mutations in the EVER2 gene develop epidermodysplasia verruciformis (EV), a disease characterized by predisposition to persistent β-genus human papillomavirus (β-HPV) skin infections, which can progress to skin cancer. To investigate how EVER2 confers protection from papillomaviruses, we infected the skin of homozygous Ever2-null mice with mouse papillomavirus MmuPV1. Like in humans with EV, infected Ever2-null mice developed skin lesions that could progress to cancer. Unlike in humans with EV, lesions in these Ever2-null mice grew more slowly and regressed more frequently than in wild-type mice. MmuPV1 transcription was higher in wild-type mice than in Ever2-null mice, indicating that mouse EVER2 does not confer protection from papillomaviruses. These findings suggest that there are functional differences between MmuPV1 and β-HPVs and/or between mouse and human EVER2.
KEYWORDS: epidermodysplasia verruciformis, papillomavirus, MmuPV1, TMC8, EVER2
INTRODUCTION
Epidermodysplasia verruciformis (EV) is a rare inherited skin disease caused by certain cutaneous papillomaviruses. Individuals with this disease develop skin lesions that resemble benign flat warts, distinct from the common warts, arising during early childhood (1, 2). EV-associated lesions typically remain for the lifetime of the patient and, in a subset of cases, can progress to invasive squamous cell carcinoma (SCC) at sun-exposed areas of the skin (3). Nearly 20 cutaneous high-risk β-genus human papillomavirus (β-HPV) genotypes are associated with EV, with two, HPV5 and HPV8, associated with the vast majority of cases of cutaneous SCCs in EV patients (3–6). Earlier studies have shown that specific genotypes of EV-associated HPV are directly related to the morphology and malignant progression of lesions arising in EV patients (7, 8). These “high-risk” β-HPVs are distinct from high-risk α-HPVs that infect mucosal tissue and cause anogenital and head and neck cancers (e.g., HPV16 and HPV18) (3, 9). Although the distribution of EV is global and these cutaneous β-HPVs are found to be widespread, EV-associated HPVs are generally innocuous in the general population (10–12). There are two types of EV: typical and atypical (13). Most typical EV patients contain mutations in both alleles of one of two genes, EVER1 and EVER2, while a subset of typical EV patients without EVER mutations contain mutations in CIB1 (calcium- and integrin-binding protein 1) (14). In people without EV, the EVER1, EVER2, and CIB1 proteins form a complex (14, 15), which we will refer to as the EVER complex. Typical EV patients are not more susceptible to infection by other pathogens, nor are they prone to developing other types of cancers, indicating that the immune system is at least partially intact (3, 16, 17). In contrast to typical EV, atypical EV patients present with primary immunodeficiencies due to T cell deficits caused by mutations in a variety of other genes (13). Patients with either typical or atypical EV develop skin lesions in response to persistent β-HPV infection.
EVER1 and EVER2 are located, head to head, in chromosomal region 17q25, 4.7 kb apart from each other (18, 19). These two genes are members of a larger gene family, the transmembrane channel-like (TMC) gene family (20, 21), composing eight genes. EVER1 is equivalent to TMC6, and EVER2 is equivalent to TMC8. We will refer to these genes as EVER1 and EVER2. EVER gene products are integral membrane proteins containing multiple transmembrane domains that are expressed in the cytoplasm and localized to the endoplasmic reticulum (3, 22). EVER1 and EVER2 are highly expressed in CD4+ and CD8+ T lymphocytes, B lymphocytes, and to a lesser extent in skin, of both humans and mice (14, 15, 23). Multiple mutations in EVER1 and EVER2 have been identified in human EV patients (22, 24). These mutations include nonsense mutations, single nucleotide deletions, splice site mutations, and deletion of exons, all of which are predicted to lead to loss of expression of wild-type gene products (3). Although one mutation is present in any one EV patient, that mutation must be homozygous, indicating it is a recessive mutation.
To determine why mutations in EVER2 increase the susceptibility of EV patients to high-risk β-HPV-induced disease, we conducted infection studies with mouse papillomavirus (MmuPV1) in Ever2 knockout mice. MmuPV1 is a recently discovered papillomavirus that generates benign papillomas, squamous cell dysplasia, and squamous cell carcinomas in the skin of laboratory mice (25–27). The ability of MmuPV1 to infect mice has been well described by multiple labs and is emerging as an important new papillomavirus-infection model (28–31). Our lab has shown that UVB increases susceptibility of immunocompetent mice (FVB/N) to MmuPV1 (27, 32–34). This correlated with the ability of UVB to induce systemic immunosuppression (27). We have also found that the incidence of disease correlates with the dose of virus applied. As the viral dose is decreased, MmuPV1 induces a lower penetrance of development of lesions demonstrating that a threshold level of virus is needed to induce disease (27). MmuPV1, like high-risk β-HPVs associated with EV, does not encode an E5 gene (35, 36). E5 has been argued to inhibit the function of the EVER complex (14, 37). E6 and E7 encoded by MmuPV1 possess some of the same biochemical properties as those encoded by EV-associated β-HPVs (38). These similarities between MmuPV1 and EV-associated β-HPVs provided us strong incentive to assess whether deficiency in Ever2 in mice increased susceptibility to MmuPV1-induced pathogenesis.
Ever2-null mice were generated by removing exons 6–9 of the Ever2 gene (15). We backcrossed these knockout alleles onto the inbred FVB/N genetic background because this is an immunocompetent strain of mice susceptible to MmuPV1-induced skin disease (27, 39). Our studies focused on assessing the consequences of knocking out only Ever2 because EV patients carry homozygous mutations in only one EVER gene. Infection with MmuPV1 generated skin lesions in both wild-type and Ever2-null FVB/N mice, with and without UVB irradiation. As expected, either an increased dose of MmuPV1 or the presence of UVB treatment increased disease penetrance in both genotypes. Surprisingly, we did not find that Ever2 deficiency increased disease penetrance as a result of MmuPV1 infection. In fact, we found faster lesion growth in wild-type mice than in Ever2-null mice infected with a low viral dose and treated with UVB, and found strong evidence that Ever2 deficiency inhibited lesion growth at high viral doses in the absence of UVB irradiation. These data demonstrate that Ever2 deficiency alone decreases the susceptibility of mice to MmuPV1-induced pathogenesis.
RESULTS
Comparing MmuPV1-induced cutaneous disease between Ever2 knockout mice and wild-type syngeneic mice exposed to UVB irradiation
To investigate the role Ever2 may be playing in restricting cutaneous papillomavirus infection in stratified squamous epithelia, ear and tail sites of Ever2-null mice and wild-type mice on the same genetic background (FVB/N) were scarified to induce wounding in the skin before they were topically treated with a solution containing 108 viral genome equivalent (VGE) of MmuPV1. Mice were then exposed to 300 mJ/cm2 of UVB whole body irradiation 24 hours postinfection. A subset of mice was scarified, topically treated with a phosphate-buffered saline (PBS) solution that did not contain MmuPV1, and irradiated 24 hours later. Mice were monitored for the presence of lesions at the sites of infection, and the size of lesions was measured every other week until the 6-month study endpoint.
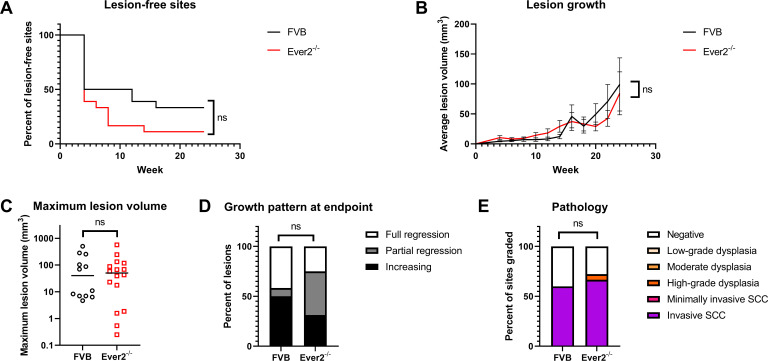
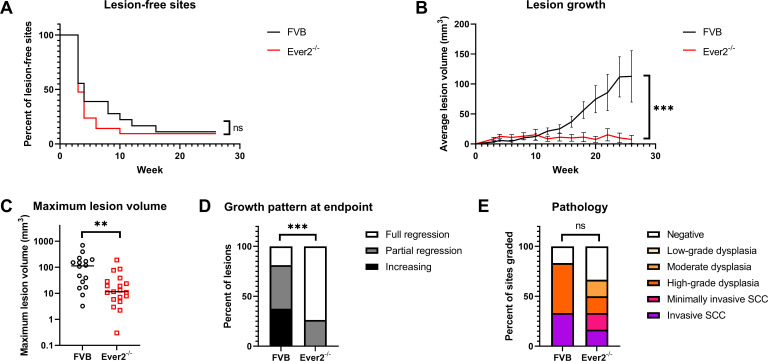
Lesions arose in both Ever2-null and wild-type mice beginning at 4 weeks postinfection (Fig. 1A). By study endpoint, 89% of infected sites had developed lesions in Ever2-null mice compared to 67% of sites in wild-type mice; however, lesion incidence did not differ statistically between genotypes (P = 0.90). There were no significant differences in lesion growth over time (P = 0.13, Fig. 1B) or maximum lesion volume (median 50 mm3 in Ever2-null mice and 40 mm3 in wild-type mice, P = 0.77, Fig. 1C). Differences in lesion regression, defined as declining lesion volume, were not significant (P = 0.84, Fig. 1D), although there was a trend toward greater regression of lesions in Ever2-null mice. At study endpoint, 31% of lesions were still increasing in size in Ever2-null mice, 44% had partially regressed, and 25% had fully regressed, while in wild-type mice, 50% of lesions were increasing, 8% had partially regressed, and 42% had fully regressed.
Fig 1.
Lesion characteristics in Ever2-null and wild-type mice infected with 108 VGE MmuPV1 and treated with UVB. Ear and tail sites were wounded to induce scarification and topically treated with MmuPV1 virions, then 24 hours later irradiated with UVB. Onset of overt lesions was monitored, and lesions were measured every other week over a period of 24 weeks postinfection. (A) Onset of lesions did not differ between Ever2-null and wild-type mice (P = 0.090, log-rank test, n = 18 infected sites per genotype). (B) Lesion volume did not differ between Ever2-null and wild-type mice (P = 0.13, type 3 test of interaction effect of Genotype × Week in a quadratic mixed model, n = 12–16 lesions per genotype). (C) Maximum lesion volume did not differ between Ever2-null and wild-type mice (P = 0.77, Wilcoxon rank-sum test, n = 12–16 lesions per genotype). (D) Lesion regression did not differ between Ever2-null and wild-type mice (P = 0.84, Wilcoxon rank-sum test, n = 12–16 lesions per genotype). (E) Disease severity did not differ between Ever2-null and wild-type mice (P = 0.60, Wilcoxon rank-sum test, n = 15–18 infected sites scored per genotype).
Tissues from infected sites on the ear and tail were harvested, fixed, paraffin embedded, hematoxylin and eosin (H&E) stained, and subjected to histopathological analysis to assess for the grade of dysplasia/carcinoma in situ and invasive carcinoma. There were no differences in pathology results between the groups (P = 0.60, Fig. 1E). Invasive cancer developed in 67% of infected sites in Ever2-null mice and in 60% of infected sites in wild-type mice. Consistent with previous findings from our lab (27, 39), wild-type FVB/N mice did not develop overt lesions or pathological disease after mock infection and UVB irradiation. Similarly, there was no overt or microscopic disease in mock-infected and irradiated Ever2-null mice (not shown).
To test whether the dose of viral infection used in the above study was so high that it could be masking differences in MmuPV1-induced disease penetrance in Ever2-null versus wild-type FVB/N mice, we infected mice on their ears and tails with 105, 106, or 107 VGE of MmuPV1 and irradiated the mice with 300 mJ/cm2 of UVB 24 hours postinfection. No lesions arose in any mice infected with 105 VGE MmuPV1 (Fig. S1A). Similar to our previous studies (27), there were no lesions in wild-type FVB/N mice infected with 106 VGE MmuPV1 and irradiated (Fig. S1B). At 106 VGE, Ever2-null mice developed lesions beginning at 4 weeks postinfection, with lesions arising at 11% of infected sites overall, but incidence did not differ from wild-type mice (P > 0.99). Lesions in Ever2-mice were small (median 2 mm3, Fig. S1C and D) and they all fully regressed by 10 weeks postinfection (Fig. S1E).
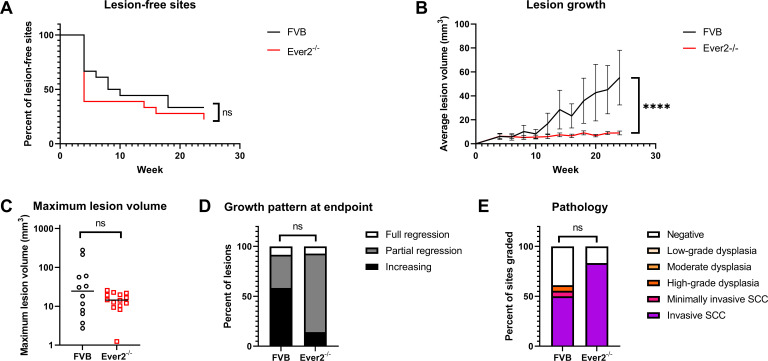
At 107 VGE MmuPV1, lesions arose by 4 weeks postinfection in both genotypes (Fig. 2A). Lesions developed in 78% of infected sites in Ever2-null mice and 67% of infected sites in wild-type mice by the 6-month study endpoint. Again, lesion incidence did not differ between genotypes (P = 0.39). Surprisingly, lesion growth was slower in Ever2-null mice than in wild-type mice (P < 0.0001, Fig. 2B). Maximum lesion volume and disease severity did not differ between Ever2-null mice and wild-type mice (P = 0.40 and 0.075, respectively, Fig. 2C and E). Because lesion growth rate differed between genotypes but maximum lesion volume over the course of longitudinal monitoring did not, we analyzed the timepoint at which lesions achieved their maximum volume. Lesions in FVB/N mice were largest at median 24 weeks postinfection, while lesions in Ever2-null mice were largest at median 14 weeks postinfection (P = 0.022, Fig. S2). Consistent with this finding, there was a trend toward greater lesion regression in Ever2-null mice (P = 0.059, Fig. 2D). Overall, we did not find that Ever2 knockout increased disease penetrance after MmuPV1 infection and UVB irradiation, and in fact we found some statistically significant evidence for faster lesion growth in wild-type mice than in Ever2-null mice.
Fig 2.
Lesion characteristics in Ever2-null and wild-type mice infected with 107 VGE MmuPV1 and treated with UVB. Ear and tail sites were wounded to induce scarification and topically treated with MmuPV1 virions, then 24 hours later irradiated with UVB. Onset of overt lesions was monitored, and lesions were measured every other week over a period of 24 weeks postinfection. (A) Onset of lesions did not differ between Ever2-null and wild-type mice (P = 0.39, log-rank test, n = 18 infected sites per genotype). (B) Lesion volume was lower in Ever2-null mice than in wild-type mice (P < 0.0001, type 3 test of interaction effect of Genotype × Week in a linear mixed model, n = 12–14 lesions per genotype). (C) Maximum lesion volume did not differ between Ever2-null and wild-type mice (P = 0.40, Wilcoxon rank-sum test, n = 12–14 lesions per genotype). (D) Lesion regression did not differ between Ever2-null and wild-type mice (P = 0.059, Wilcoxon rank-sum test, n = 12–14 lesions per genotype). (E) Disease severity did not differ between Ever2-null and wild-type mice (P = 0.075, Wilcoxon rank-sum test, n = 18 infected sites scored per genotype).
Comparing MmuPV1-induced cutaneous disease between Ever2 knockout mice and wild-type syngeneic mice in the absence of UVB irradiation
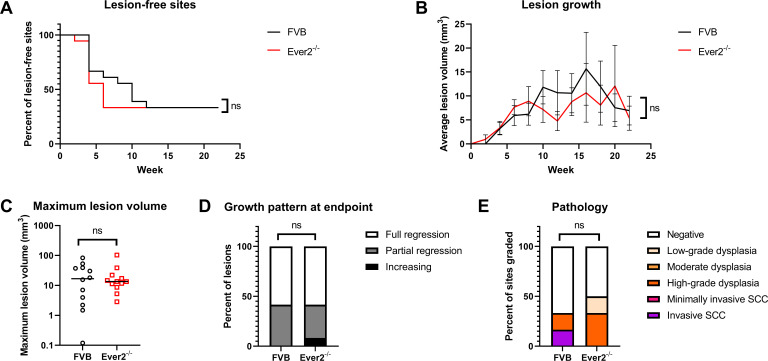
To determine whether the addition of UVB irradiation may be masking the potential effects of a knockdown of Ever2 in mice on susceptibility to MmuPV1-induced disease, we conducted MmuPV1 infection studies on Ever2-null and wild-type FVB/N mice in the absence of UVB exposure. Ear and tail sites were scarified and infected with 108 VGE of MmuPV1 virions. A subset of these mice was mock infected and showed no overt or microscopic lesions by the 6-month endpoint. Both Ever2-null and wild-type FVB/N mice developed lesions after MmuPV1 infection. Lesions arose by week 2 postinfection in Ever2-null mice and week 4 in wild-type mice, and developed at 67% of infected sites in both genotypes 6 months postinfection (Fig. 3A). Like the mice infected with 108 VGE MmuPV1 and UVB irradiated, there were no differences in lesion incidence (P = 0.64, Fig. 3A), growth rate (P = 0.53, Fig. 3B), maximum volume, regression, or severity (P = 0.98, >0.99, and >0.99, respectively, Fig. 3C through E).
Fig 3.
Lesion characteristics in Ever2-null and wild-type mice infected with 108 VGE MmuPV1 without irradiation. Ear and tail sites were wounded to induce scarification and topically treated with MmuPV1 virions. Onset of overt lesions was monitored, and lesions were measured every other week over a period of 22 weeks postinfection. (A) Onset of lesions did not differ between Ever2-null and wild-type mice (P = 0.64, log-rank test, n = 18 infected sites per genotype). (B) Lesion volume did not differ between Ever2-null and wild-type mice (P = 0.53, type 3 test of interaction effect of Genotype × Week in a quadratic mixed model, n = 12 lesions per genotype). (C) Maximum lesion volume did not differ between Ever2-null and wild-type mice (P = 0.98, Wilcoxon rank-sum test, n = 12 lesions per genotype). (D) Lesion regression did not differ between Ever2-null and wild-type mice (P > 0.99, Wilcoxon rank-sum test, n = 12 lesions per genotype). (E) Disease severity did not differ between Ever2-null and wild-type mice (P > 0.99, Wilcoxon rank-sum test, n = 6 infected sites scored per genotype).
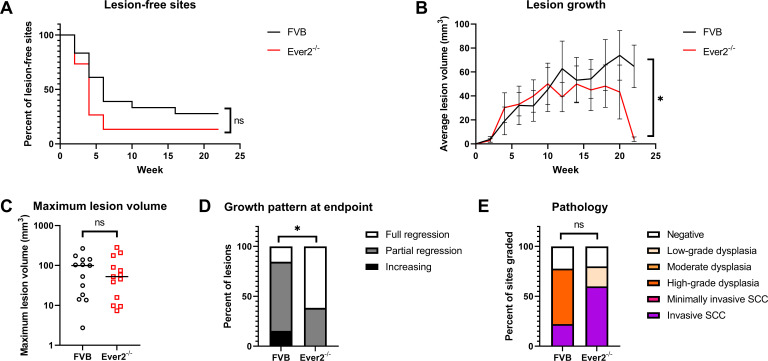
To test whether increasing viral dose in the absence of UVB irradiation would reveal differences in MmuPV1-induced disease penetrance in Ever2-null versus wild-type FVB/N mice, we infected mice on their ears and tails with 109 or 1010 VGE of MmuPV1 without UVB exposure. At 109 VGE, lesions arose in 87% of infected sites in Ever2-null mice and 72% of infected sites in wild-type mice by 6 months postinfection, but this difference again was not significant (P = 0.13, Fig. 4A). Similar to some of our experiments with UVB irradiation and lower viral doses, lesions grew more slowly in Ever2-null mice than in wild-type mice (P = 0.018, Fig. 4B). Here, we also found that more lesions regressed by the study endpoint in the Ever2-null mice (P = 0.021, Fig. 4D). There was no difference in maximum lesion volume (P = 0.69, Fig. 4C). At 1010 VGE, lesions arose at ~90% of infected sites in both Ever2-null and wild-type mice (P = 0.43, Fig. 5A). At this high viral dose, Ever2-null mice again displayed lower lesion growth rates (P = 0.0003, Fig. 5B) and greater lesion regression (P = 0.0003, Fig. 5D) than wild-type mice, as well as lower maximum lesion volumes (P = 0.0018, Fig. 5C). Lesions were largest at median 24 weeks in FVB/N mice and at 6 weeks in Ever2-null mice (P = 0.0014, Fig. S2). There were no differences in microscopic pathology evaluation at 109 or 1010 VGE (P = 0.57 and 0.58, respectively, Fig. 4E and 5E). Taken together, these results indicate that, contrary to our hypothesis, Ever2 knockout inhibited cutaneous MmuPV1-induced lesion growth in the absence of UVB irradiation.
Fig 4.
Lesion characteristics in Ever2-null and wild-type mice infected with 109 VGE MmuPV1 without irradiation. Ear and tail sites were wounded to induce scarification and topically treated with MmuPV1 virions. Onset of overt lesions was monitored, and lesions were measured every other week over a period of 22 weeks postinfection. (A) Onset of lesions did not differ between Ever2-null and wild-type mice (P = 0.13, log-rank test, n = 15–18 infected sites per genotype). (B) Lesions grew more slowly in Ever2-null mice than in wild-type mice (P = 0.018, type 3 test of interaction effect of Genotype × Week2 in a quadratic mixed model, n = 13 lesions per genotype). (C) Maximum lesion volume did not differ between Ever2-null and wild-type mice (P = 0.69, Wilcoxon rank-sum test, n = 13 lesions per genotype). (D) Lesion regression was higher in Ever2-null mice than in wild-type mice (P = 0.021, Wilcoxon rank-sum test, n = 13 lesions per genotype). (E) Disease severity did not differ between Ever2-null and wild-type mice (P = 0.57, Wilcoxon rank-sum test, n = 5–9 infected sites scored per genotype).
Fig 5.
Lesion characteristics in Ever2-null and wild-type mice infected with 1010 VGE MmuPV1 without irradiation. Ear and tail sites were wounded to induce scarification and topically treated with MmuPV1 virions. Onset of overt lesions was monitored, and lesions were measured every other week over a period of 26 weeks postinfection. (A) Onset of lesions did not differ between Ever2-null and wild-type mice (P = 0.43, log-rank test, n = 18–21 infected sites per genotype). (B) Lesions grew more slowly in Ever2-null mice than in wild-type mice (P = 0.0003, type 3 test of interaction effect of Genotype × Week2 in a quadratic mixed model, n = 16–19 lesions per genotype). (C) Maximum lesion volume was lower in Ever2-null mice than in wild-type mice (P = 0.0018, Wilcoxon rank-sum test, n = 16–19 lesions per genotype). (D) Lesion regression was higher in Ever2-null mice than in wild-type mice (P = 0.0003, Wilcoxon rank-sum test, n = 16–19 lesions per genotype). (E) Disease severity did not differ between Ever2-null and wild-type mice (P = 0.58, Wilcoxon rank-sum test, n = 6 infected sites scored per genotype).
Effect of either higher doses of MmuPV1 or UVB irradiation on disease penetrance in both Ever2-null and wild-type syngeneic FVB/N mice
In light of our unexpected findings that Ever2-null mice exhibited decreased susceptibility to MmuPV1-induced disease, we reanalyzed lesion data within both wild-type and Ever2-null genotypes to confirm our previous findings that disease penetrance increased with either higher doses of MmuPV1 or with UVB irradiation (27). To determine whether disease susceptibility increased with viral dose in the presence of UV, we compared lesion characteristics among mice infected with 105, 106, 107, or 108 VGE MmuPV1 and irradiated with UVB (Fig. S3). In both wild-type and Ever2-null FVB/N mice, lesion incidence (P < 0.0001 for both genotypes) and growth (P = 0.047 in wild-type mice and <0.0001 in Ever2-null mice) increased with higher viral doses. Ever2-null mice also demonstrated higher maximum lesion volumes at 108 VGE than at 107 VGE (P = 0.031). Maximum lesion volume trended higher in wild-type mice infected with the higher viral dose, but this difference was not significant (P = 0.55). Lesion regression and disease severity did not differ by dose of MmuPV1 in either genotype (P = 0.34 and 0.80 for wild-type mice, and >0.99 and 0.44 for Ever2-null mice). These results largely confirm increased disease penetrance with higher doses of MmuPV1 in the presence of UV.
To assess whether disease susceptibility increased with viral dose in the absence of UV in both wild-type and Ever2-null FVB/N mice, we compared lesion characteristics among treatment groups infected with 108, 109, or 1010 VGE MmuPV1 and not exposed to UVB within each genotype (Fig. S4). As expected, in wild-type FVB/N mice, lesion growth rates (P = 0.0004), maximum lesion volumes (P = 0.0098), and the proportion of lesions still increasing in size at the study endpoint (P = 0.013) all increased as viral dose increased. The number of invasive carcinomas also increased with viral dose, but differences in disease severity were not significant (P = 0.28). Lesion incidence did not differ by viral dose in wild-type mice (P = 0.11). In Ever2-null mice, lesion incidence (P = 0.031), growth rate (P < 0.0001), and size (P = 0.034) all differed by viral dose. Interestingly, lesions reached maximum volumes in Ever2-null mice infected with 109 VGE MmuPV1 and not the highest dose of 1010 (Fig. S4G and H). We also observed the largest proportion of invasive carcinomas at 109 VGE in Ever2-null mice, although differences in disease severity did not reach statistical significance (P = 0.28). There was no effect of viral dose on lesion regression in Ever2-null mice (P = 0.58). Overall, these results support increased disease penetrance with higher doses of MmuPV1 in the absence of UVB in wild-type FVB/N mice.
To confirm that UVB irradiation increased susceptibility to cutaneous MmuPV1-induced disease, we assessed differences between wild-type and Ever2-null FVB/N mice infected with 108 VGE MmuPV1 with UVB irradiation, and those infected with 108 VGE MmuPV1 without UVB irradiation (Fig. S5). Lesion incidence did not differ by UVB treatment in wild-type (P = 0.99) mice. Incidence was higher in Ever2-null mice with UVB irradiation than without, but this was not significant (P = 0.25). In both genotypes, lesions grew faster with UVB than without UVB (P = 0.0007 in wild-type mice and <0.0001 in Ever2-null mice). Both genotypes displayed trends toward higher maximum lesion volumes and lower rates of lesion regression that were not statistically significant (P = 0.16 and 0.092 in wild-type mice, and 0.090 and 0.073 in Ever2-null mice). Significantly higher disease severity was found in Ever2-null mice treated with UVB (P = 0.018), and wild-type mice treated with UVB showed higher rates of invasive carcinoma, but this was not statistically significant (P = 0.19). These results provide evidence for increased disease penetrance of MmuPV1 with UVB treatment in both wild-type and Ever2-null mice.
Comparing MmuPV1 viral gene expression between lesions arising in Ever2 knockout mice and wild-type syngeneic mice
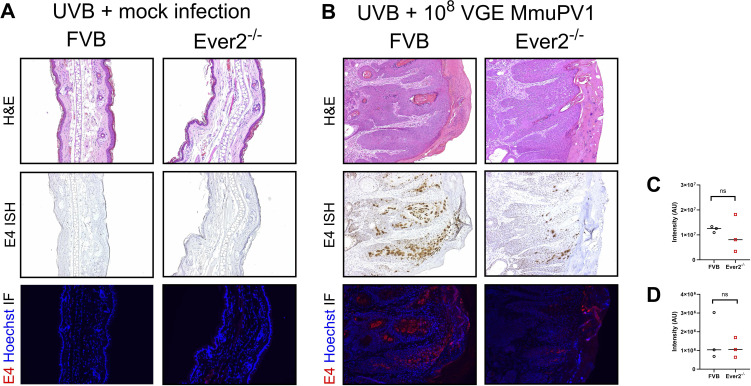
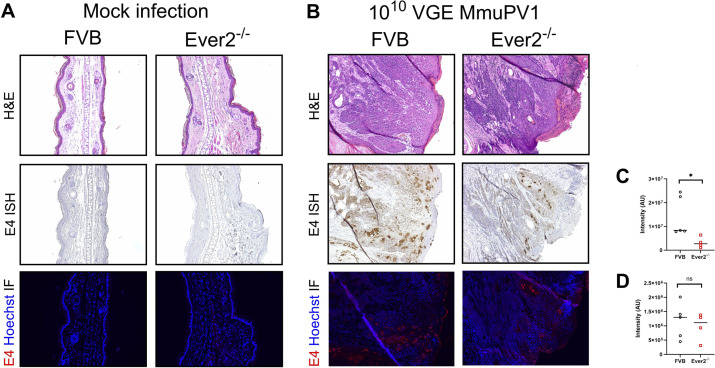
To determine whether sites that were infected with MmuPV1 display differences in late viral gene expression, we assessed tissue expression of the MmuPV1 gene E4 at transcript and protein level. Tissue sections from MmuPV1-infected Ever2-null and wild-type mice were subjected to E4-specific in situ hybridization (ISH) and immunofluorescent (IF) staining. Tissues from mock-infected mice were included as negative controls. Mock-infected tissues had all been graded negative for squamous dysplasia or carcinoma, and were found to be negative for MmuPV1 biomarkers (Fig. 6A and 7A). We compared levels of E4 transcript and protein in dysplasias and invasive carcinomas from Ever2-null and wild-type mice infected with 108 VGE MmuPV1 and treated with UVB irradiation, and in Ever2-null and wild-type mice infected with 1010 VGE MmuPV1 without UVB irradiation. By qualitative inspection, we observed decreased levels of E4 transcript in Ever2-null mice compared with wild-type mice (Fig. 6B and 7B). Quantitative analysis revealed that these differences reached significance in mice infected with 1010 VGE in the absence of UVB (P = 0.016, Fig. 7C), but not in mice infected with 108 VGE and treated with UVB (P = 0.70, Fig. 6C). E4 protein stains did not differ qualitatively (Fig. 6B and 7B) or quantitatively between genotypes (P > 0.99 and 0.56, respectively, Fig. 6D and 7D). We also completed ISH for MmuPV1 E6E7. As with E4, we observed decreased viral gene expression in skin lesions from Ever2-null mice, which was statistically significant in mice infected with 1010 VGE of MmuPV1 (Fig. S6). These findings indicate that Ever2-null mice display modest decreases in viral gene expression compared with wild-type mice 6 months after cutaneous infection with MmuPV1.
Fig 6.
MmuPV1 biomarkers in lesions from Ever2-null and wild-type mice infected with 108 VGE MmuPV1 and treated with UVB. (A) Normal tissues in mock-infected Ever2-null and wild-type mice irradiated with UVB. Serial sections of tissues stained with H&E (top row), MmuPV1 E4 RNAscope ISH (brown, middle row), and MmuPV1 E4 IF (red, bottom row). (B) Lesional tissue in MmuPV1-infected wild-type mice and Ever2-null mice irradiated with UVB. Depicted examples represent high-grade dysplasia with minimally invasive carcinoma seen at the base of the squamous epithelium in both genotypes. Serial sections stained with H&E (top row), MmuPV1 E4 RNAscope ISH (brown, middle row), and MmuPV1 E4 IF (red, bottom row). (C) E4 transcript staining intensity did not differ between Ever2-null and wild-type mice (P = 0.70, Wilcoxon rank-sum test, n = 3 lesions per genotype). (D) E4 protein staining intensity did not differ between Ever2-null and wild-type mice (P > 0.99, Wilcoxon rank-sum test, n = 3 lesions per genotype).
Fig 7.
MmuPV1 biomarkers in lesions from Ever2-null and wild-type mice infected with 1010 VGE MmuPV1 without irradiation. (A) Normal tissues in mock-infected Ever2-null and wild-type mice. Serial sections of tissues stained with H&E (top row), MmuPV1 E4 RNAscope ISH (brown, middle row), and MmuPV1 E4 IF (red, bottom row). (B) Minimally invasive squamous cell carcinoma in the background of high-grade dysplasia in MmuPV1-infected Ever2-null and wild-type mice. Serial sections stained with H&E (top row), MmuPV1 E4 RNAscope ISH (brown, middle row), and MmuPV1 E4 IF (red, bottom row). (C) E4 transcript staining intensity was lower in Ever2-null mice than in wild-type mice (P = 0.016, Wilcoxon rank-sum test, n = 4–5 lesions per genotype). (D) E4 protein staining intensity did not differ between Ever2-null and wild-type mice (P = 0.56, Wilcoxon rank-sum test, n = 4–5 lesions per genotype).
DISCUSSION
In this study, we found that Ever2-null mice infected with the mouse papillomavirus did not exhibit the same phenotype as humans with mutations in EVER2 who develop EV because of infection with a β-HPV. In contrast to human EV patients, in mice, Ever2 deficiency decreased the susceptibility to MmuPV1-induced squamous dysplasia and carcinoma.
Our findings contrast the proposed functions of the human EVER complex where mutations in either EVER1 or EVER2 contribute to HPV-induced disease. We started with the hypothesis that wild-type EVER proteins perform some anti-viral function in the context of immune responses to viral infection. Consistent with this hypothesis, high expression levels of EVER1 and EVER2 gene products have been found in multiple immune cells including, but not limited to, CD4+ and CD8+ T lymphocytes, B lymphocytes, and natural killer (NK) cells (3, 40). Therefore, we expected that knocking out EVER protein expression would allow viral infection to occur and persist by bypassing normal immune surveillance. We did observe a lower, albeit statistically insignificant, threshold for MmuPV1-induced lesions in Ever2-null mice than in wild-type mice (106 vs 107 VGE). However, somewhat unexpectedly, we found that lesion incidence was not significantly higher in Ever2-null mice than in wild-type mice across different viral doses and with and without potentiation by UVB treatment. Also, we did not observe an increase in the persistence of MmuPV1-induced lesions in Ever2-null mice compared to wild-type mice that could be indicative of a decrease in immune surveillance. An alternative hypothesis for the pro-oncogenic effect of EVER mutations in humans is that Ever genes perform anti-proliferative functions in epithelial cells, and knockdown of EVER proteins promotes increased cell proliferation or decreased cell death. EVER2 has been argued to play a role in inducing apoptosis (41). EVER proteins have also been argued to contribute to the regulation of transcription factors (via regulation of cellular zinc ion balance) that could contribute to cell growth (37). However, contrary to this hypothesis, MmuPV1-induced lesions in Ever2-null mice grew smaller than in wild-type mice. Another hypothesis is that knocking out Ever1 or Ever2 potentiates the productive phase of the viral life cycle. Supportive of this hypothesis, EVER2 has been found to impair the activity of AP-1 (activating protein 1), which is a transcription factor that plays a role in the papillomavirus life cycle (37). However, we found a decrease in viral gene expression in MmuPV1-infected Ever2-null mice compared with wild-type mice. Finally, it has been proposed that the E5 oncoprotein from high-risk mucosotropic α-HPVs, such as HPV16, is able to interfere with potential antiviral functions of the EVER complex. This could explain why humans both with and without EV are susceptible to high-risk α-HPVs, but only those with EV are susceptible to disease caused by cutaneous β-HPVs, while humans without EV are protected from β-HPVs because these viruses lack an E5 that could overcome the protection of an intact EVER complex. Consistent with this hypothesis, the HPV16 E5 oncoprotein has been shown to bind the EVER complex and upregulate viral transcription in vitro (14, 37). In addition, studies infecting HPV16 E5 transgenic mice with MmuPV1 showed that the transgenic mice were more susceptible to MmuPV1-induced disease, indicating that HPV16 E5 potentiates MmuPV1-induced pathogenesis (39). However, if the hypothesis were true that HPV16 E5 potentiates MmuPV1-induced disease by interfering with the function of the EVER complex, we would have expected that knocking out Ever genes in mice would produce a similar effect as seen with expressing E5. In contrast, we observed a decrease in lesion growth and viral transcription, and an increase in lesion regression in Ever2-null mice—a model with the same FVB/N genetic background as the HPV16 E5 mice that in our previous studies developed squamous dysplasia and invasive carcinoma with the same doses of MmuPV1 and UVB irradiation (39). Therefore, in the MmuPV1 cutaneous infection model, the effects of knocking out Ever2 contrast with the effects of adding HPV16 E5. Because we performed longitudinal monitoring of lesions over time, we were unable to assess reasons for lesion regression, such as immune infiltration or cell death. Future studies with multiple endpoints for tissue collection could address this important question.
One potential explanation for our findings is that MmuPV1 can overcome the protection from papillomaviruses conferred by an intact EVER complex in a way that β-HPVs cannot. MmuPV1 is a member of the papillomavirus π-genus, and has low overall sequence homology with HPVs (35). However, MmuPV1 does share important features with EV-associated β-HPVs including HPV5 and 8. Like these β-HPVs, MmuPV1 lacks an E5 open reading frame and contains separate promoters for the oncogenes E6 and E7 (36). The amino acid sequences and biochemical activities of E6 and E7 are similar between MmuPV1 and cutaneous β-HPVs (38). Specifically, the E6 proteins of MmuPV1, HPV5, and HPV8 bind MAML1 and SMAD2/3, which inhibits NOTCH signaling and TGFβ signaling, respectively (38). The E7 proteins of both MmuPV1 and β-HPVs target pRB; however, MmuPV1 E7 probably targets “non-canonical” functions of pRB because it lacks the LXCXE motif that β-HPV E7s use to interact with pRB and therefore MmuPV1 E7 does not inhibit pRB’s inhibition of the E2F transcription factor (38). MmuPV1, like HPV5 and 8, produces an E8^E2 spliced transcript (36), the function of which is dependent on the E8 part (42). The amino acid sequence of MmuPV1 E8 is similar to those of HPV5 and 8 E8s, and the function of E8^E2 is conserved in vitro among these papillomaviruses (42). MmuPV1 E8^E2 is required for cutaneous lesion formation in athymic nude mice (42), and abundant E8^E2 transcripts have been found in an HPV8+ lesion from a patient with EV (43), suggesting similar functions of E8^E2 in vivo. Finally, the papillomavirus E1^E4 transcript is abundant in both HPV-induced skin lesions in EV patients (43) and MmuPV1-induced skin lesions in mice (36, 44). Despite these similarities between MmuPV1 and the cutaneous β-HPVs associated with EV, biochemical interactions between MmuPV1 gene products and the EVER complex are unknown and could contribute to the differences we observed between Ever2-null mouse and EV patient susceptibility to papillomaviruses. Another factor to consider is that the FVB/N mice that we used are particularly susceptible to MmuPV1-induced lesions. These mice develop papillomavirus-associated SCC in weeks to months, rather than the years to decades it takes humans to develop the HPV-associated cancers that this model aims to mimic. Although quick disease development is a significant logistical advantage of MmuPV1 models, results should be interpreted in context of differences between host species.
Therefore, another hypothesis that could explain our findings is that mouse EVER2 does not confer protection from papillomaviruses the way that it does in humans. EVER2 shares 78% amino acid sequence identity between humans and mice (20). The EVER complex proteins, EVER1, EVER2, and CIB1, are found in multiple strains of mice, including FVB/N (15). In both humans and mice, EVER1 and 2 proteins are high in immune cells and low in keratinocytes, while CIB1 is strongly expressed in both immune cells and keratinocytes (14, 15, 23). These proteins regulate and stabilize each other in both species (14, 15). Like EV patients with EVER2 or EVER1 mutations (23, 45), Ever2- and Ever1-null mice are healthy, with a normal reproductive function, and do not have a T cell deficiency that increases their susceptibility to pathogens other than papillomavirus (15). In contrast, CIB1 function appears to differ between species. Cib1-null mice have vascular abnormalities and, in males, sterility (46). These phenotypes are not observed in human EV patients with CIB1 mutations (14), and CIB1 knockout in human keratinocytes results in minimal changes to gene expression (47), indicative of more limited functions of CIB1 in humans compared to mice. It is not known whether there may also be species-specific differences in the localization and function of the EVER complex, of which CIB1 is a member. There is evidence to suggest that the human EVER complex is present in keratinocytes and directly interacts with γ-HPV E8 and α-HPV E5 proteins (14). However, in mice, the EVER complex in keratinocytes is thought to be present at very low levels (15). Understanding EVER complex protein levels and localization in murine skin after UV treatment or MmuPV1 infection would begin to explain our findings. However, because of a lack of reliable antibodies for EVER1 and EVER2 that are suitable for immunofluorescence (14, 15), we were unable to characterize EVER complex proteins in our experimental tissues. The hypothesis that the mouse EVER complex differs from its human counterpart in protection from papillomaviruses may be supported by two recent studies that used Ever1- and Ever2-null mice on an FVB/N background to test HPV vaccines (48, 49). Although genotypes were not compared statistically to wild-type mice, neither Ever1- nor Ever2-null mice appeared to display increased susceptibility to HPV5 pseudovirus in the vagina (48, 49). Combined with our study, these findings provide evidence that the murine EVER complex is unable to prevent infection by EV-associated β-HPVs, or to completely prevent infection, persistence, or carcinogenesis associated with MmuPV1.
MATERIALS AND METHODS
Mice
Ever2-/- (Tmc8-/-) mice were generated on a 129 background, bred, and maintained on the C57BL/6 Cre inbred genetic background in the homozygous state, and backcrossed to an FVB/N inbred genetic background as previously described (15, 48, 49). FVB/N (Taconic) mice were purchased and bred to provide nontransgenic, syngeneic mice as wild-type controls for our studies.
All mice were housed in micro-isolator cages. Mice were fed with the 2019 Teklad rodent diet (Envigo) and were maintained on a 12-hour light/12-hour dark cycle. Mice were housed at the Medical Sciences Center Vivarium or the Clinical Sciences Center Vivarium at the University of Wisconsin School of Medicine and Public Health in strict accordance with guidelines approved by the Association for Assessment of Laboratory Animal Care.
MmuPV1 infections
In vivo infections were performed using purified MmuPV1 virus stocks that were generated by isolating MmuPV1 virions from papillomas arising on nude mice as described previously (27, 50). The concentration of virus in these stocks was quantified by determining the amount of encapsidated viral DNA to give VGEs. For any particular experiment, the same stock of virus was used for all cohorts in that experiment. Within any experiment, controls were included for direct comparison. Mock infections were performed with PBS solution. To conduct an infection, mice were anesthetized and the skin on the inner ear or tail was scarified using a 27-gauge syringe needle to scrape the skin to disrupt the epithelial tissue. Following scarification, the virus or mock solution was delivered to wounded sites by pipette delivery.
UV treatment
Mice that were irradiated were exposed to a single dose of UVB radiation using a Research Irradiation Unit (Daavlin, Bryan, OH, USA). Mice were exposed to 300 mJ/cm2 of UVB radiation 24 hours postinfection with MmuPV1 or mock infection.
Disease monitoring
Mice were examined every 2 weeks for the development of lesions at infected sites over the course of 6 months (22–26 weeks) postinfection. Lesion incidence was noted, and lesions were measured either by a micrometer or by calipers. For each lesion, four separate measurements were taken: (i) height of lesion on the ear/tail, (ii) height of ear/tail in nearby non-infected region, (iii) length of lesion, and (iv) width of lesion. For each lesion, the largest area was measured. To calculate lesion volume, we subtracted the height of the ear/tail in the non-infected nearby region (ii), from the height of the ear/tail at the site of the lesion (i), and multiplied this by the length and width of the lesion. The extent of lesion regression at the study endpoint was noted for each individual lesion. A lesion was “increasing” if its largest volume measurement was its last one, at the 6-month endpoint. A lesion that had “partially regressed” had reached its greatest volume at some point during the middle of the study, but a lesion still remained at the study endpoint. A lesion that had “fully regressed” had appeared during the study but was no longer overtly present at the study endpoint. Animals were euthanized at the 6-month endpoint. At the time of euthanasia, infected sites with and without lesions were isolated, fixed in 4% paraformaldehyde, processed, embedded in paraffin, and sectioned for histopathological analysis.
Histopathological analysis
Tissues were cut to generate 5-μm sections. Every 10th section was stained with H&E and subjected to histopathological analysis by a pathologist to determine disease severity ranging from normal epithelium to dysplasia to invasive SCC. Dysplasia was subcategorized into low, moderate, and severe (or low, intermediate, and high grade), and SCC was delineated by degree of invasiveness into minimally invasive vs invasive. The worst grade of disease for each site of infection was determined.
MmuPV1 biomarker staining and quantification
Serial sections were stained for MmuPV1 E4 transcript and protein. ISH for E4 transcript was completed using RNAscope (2.5 HD Reagent Kit-Brown, 322300, Advanced Cell Diagnostics, Newark, CA, USA) with probes specific for MmuPV1 E4 (473281) and E6E7 (409771) according to the manufacturer’s instructions. IF staining for E4 protein was completed following standard protocols. Briefly, sections were deparaffinized in xylene and rehydrated in a series of ethanols. Antigen retrieval was performed by boiling slides in a microwave oven for 20 minutes in 1X Tris-based antigen unmasking solution, pH 9.0 (H-3301, Vector Laboratories, Inc., Burlingame, CA, USA). Sections were permeabilized for 20 minutes in 0.1% PBS-Tween, blocked for 1 hour in 5% goat serum, then incubated in primary antibody (1:1,000 rabbit anti-E4, gift from John Doorbar, University of Cambridge, Cambridge, UK) overnight at 4°C. Slides were washed and incubated in secondary antibody (1:500 goat anti-rabbit Cy3, ab6939, Abcam, Cambridge, UK) for 1 hour at room temperature. Nuclei were counterstained with Hoechst dye for 10 minutes, and then slides were mounted and coverslipped with Prolong Glass mounting media (P36984, Fisher Scientific, Hampton, NH, USA), cured flat at room temperature in the dark for 24 hours, and stored at 4°C.
Images were captured using a Zeiss AxioImager M2 microscope using AxioVision software (Jena, Germany). For each stain, all slides were imaged during a single session using consistent microscope settings including exposure times. Staining intensity was quantified in one 10× image per lesion using ImageJ. For ISH, images were deconvoluted using the “H DAB” vector, and the 3, 3'-diaminobenzidine (DAB) channel (Color 2) was desaturated and used for analysis. For IF, the red channel was desaturated and used for analysis. Images were thresholded to remove background signal using consistent settings within each stain, then measured for density (RawIntDen).
Statistical analysis
Statistical analysis was carried out using GraphPad Prism 9.5.0 and SAS Studio 3.81. All tests were two-tailed with α = 0.05. The log-rank (Mantel-Cox) test was used to determine the significance of differences in lesion incidence. The significance of differences in lesion growth over time was tested by type 3 tests of fixed effects in mixed models that included group, linear and quadratic time postinfection, and interactions of group with linear and quadratic time as fixed effects. Insignificant quadratic terms were removed from final models. Significant interactions were interpreted as group differences in lesion growth rate. Lesion growth data are represented as mean ± standard error of the mean. The Wilcoxon rank-sum test (for two groups) or Kruskal-Wallis test (for more than two groups) was used to determine the significance of differences in maximum lesion volume, timepoint of achieving maximum lesion volume, intensity of MmuPV1 biomarkers, lesion growth pattern at endpoint, and disease severity. Maximum lesion volume data are represented as individual data points and group medians. Growth patterns were assigned ranks, with 0 = full regression, 1 = partial regression, and 2 = increasing. Disease severity was assigned ranks, with 0 = negative/no disease, 1 = low-grade dysplasia, 2 = moderate dysplasia, 3 = high-grade dysplasia, 4 = minimally invasive SCC, and 5 = invasive SCC. Results are represented as ns (not significant, P > 0.05); *P < 0.05, **P < 0.01, ***P < 0.001, or ****P < 0.0001.
ACKNOWLEDGMENTS
We thank Dr. Doug Lowy (NCI, USA) for connecting the Lambert and Griffith lab, which led to this study. We thank Dr. Andrew J. Griffith (NIH, USA) for the Tmc8-/- mice. We thank Dr. John Doorbar (University of Cambridge, UK) for MmuPV1 E4 antibody. We thank UWCCC Experimental Pathology Laboratory for processing harvested tissue.
This study was supported by NCI grants P01 CA022443 and R35 CA210807 to P.F.L. R.E.K. is supported by T32 CA090217.
Contributor Information
Paul F. Lambert, Email: plambert@wisc.edu.
Lawrence Banks, International Centre for Genetic Engineering and Biotechnology, Trieste, Italy.
DATA AVAILABILITY
Data supporting the findings of this study are available from the corresponding author upon request.
ETHICS APPROVAL
All protocols for animal work were approved by the University of Wisconsin School of Medicine and Public Health Institutional Animal Care and Use Committee (protocol number M005871).
SUPPLEMENTAL MATERIAL
The following material is available online at https://doi.org/10.1128/jvi.00174-24.
Figures S1 to S6.
ASM does not own the copyrights to Supplemental Material that may be linked to, or accessed through, an article. The authors have granted ASM a non-exclusive, world-wide license to publish the Supplemental Material files. Please contact the corresponding author directly for reuse.
REFERENCES
- 1. Jablonska S, Orth G. 1985. Epidermodysplasia verruciformis. Clin Dermatol 3:83–96. doi: 10.1016/0738-081x(85)90052-5 [DOI] [PubMed] [Google Scholar]
- 2. Majewski S, Jabłońska S. 1995. Epidermodysplasia verruciformis as a model of human papillomavirus-induced genetic cancer of the skin. Arch Dermatol 131:1312–1318. [PubMed] [Google Scholar]
- 3. Orth G. 2006. Genetics of epidermodysplasia verruciformis: insights into host defense against papillomaviruses. Semin Immunol 18:362–374. doi: 10.1016/j.smim.2006.07.008 [DOI] [PubMed] [Google Scholar]
- 4. McLaughlin-Drubin ME. 2015. Human papillomaviruses and non-melanoma skin cancer. Semin Oncol 42:284–290. doi: 10.1053/j.seminoncol.2014.12.032 [DOI] [PubMed] [Google Scholar]
- 5. Orth G. 1986. Epidermodysplasia verruciformis: a model for understanding the oncogenicity of human papillomaviruses, p 157–174. In Ciba foundation symposium 120 - papillomaviruses. John Wiley & Sons, Ltd. [DOI] [PubMed] [Google Scholar]
- 6. Majewski S, Jabłońska S, Orth G. 1997. Epidermodysplasia verruciformis. Immunological and nonimmunological surveillance mechanisms: role in tumor progression. Clin Dermatol 15:321–334. doi: 10.1016/s0738-081x(96)00169-1 [DOI] [PubMed] [Google Scholar]
- 7. Orth G, Jablonska S, Favre M, Croissant O, Jarzabek-Chorzelska M, Rzesa G. 1978. Characterization of two types of human papillomaviruses in lesions of epidermodysplasia verruciformis. Proc Natl Acad Sci U S A 75:1537–1541. doi: 10.1073/pnas.75.3.1537 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Orth G, Jablonska S, Jarzabek-Chorzelska M, Obalek S, Rzesa G, Favre M, Croissant O. 1979. Characteristics of the lesions and risk of malignant conversion associated with the type of human papillomavirus involved in epidermodysplasia verruciformis. Cancer Res 39:1074–1082. [PubMed] [Google Scholar]
- 9. de Villiers E-M, Fauquet C, Broker TR, Bernard H-U, zur Hausen H. 2004. Classification of papillomaviruses. Virology 324:17–27. doi: 10.1016/j.virol.2004.03.033 [DOI] [PubMed] [Google Scholar]
- 10. Astori G, Lavergne D, Benton C, Höckmayr B, Egawa K, Garbe C, de Villiers EM. 1998. Human papillomaviruses are commonly found in normal skin of immunocompetent hosts. J Invest Dermatol 110:752–755. doi: 10.1046/j.1523-1747.1998.00191.x [DOI] [PubMed] [Google Scholar]
- 11. Antonsson A, Forslund O, Ekberg H, Sterner G, Hansson BG. 2000. The ubiquity and impressive genomic diversity of human skin papillomaviruses suggest a commensalic nature of these viruses. J Virol 74:11636–11641. doi: 10.1128/jvi.74.24.11636-11641.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Boxman ILA, Berkhout RJM, Mulder LHC, Wolkers MC, Bavinck JNB, Vermeer BJ, ter Schegget J. 1997. Detection of human papillomavirus DNA in plucked hairs from renal transplant recipients and healthy volunteers. J Invest Dermatol 108:712–715. doi: 10.1111/1523-1747.ep12292090 [DOI] [PubMed] [Google Scholar]
- 13. de Jong SJ, Imahorn E, Itin P, Uitto J, Orth G, Jouanguy E, Casanova J-L, Burger B. 2018. Epidermodysplasia verruciformis: inborn errors of immunity to human beta-papillomaviruses. Front Microbiol 9:1222. doi: 10.3389/fmicb.2018.01222 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. de Jong SJ, Créquer A, Matos I, Hum D, Gunasekharan V, Lorenzo L, Jabot-Hanin F, Imahorn E, Arias AA, Vahidnezhad H, et al. 2018. The human CIB1–EVER1–EVER2 complex governs keratinocyte-intrinsic immunity to β-papillomaviruses. J Exp Med 215:2289–2310. doi: 10.1084/jem.20170308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Wu C-J, Li X, Sommers CL, Kurima K, Huh S, Bugos G, Dong L, Li W, Griffith AJ, Samelson LE. 2020. Expression of a TMC6-TMC8-CIB1 heterotrimeric complex in lymphocytes is regulated by each of the components. J Biol Chem 295:16086–16099. doi: 10.1074/jbc.RA120.013045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Jablonska S, Orth G, Jarzabek-Chorzelska M, Rzesa G, Obałek S, Glinski W, Favre M, Croissant O. 1979. Epidermodysplasia verruciformis versus disseminated verrucae planae: is epidermodysplasia verruciformis a generalized infection with wart virus? J Invest Dermatol 72:114–119. doi: 10.1111/1523-1747.ep12530383 [DOI] [PubMed] [Google Scholar]
- 17. Lutzner MA. 1978. Epidermodysplasia verruciformis. An autosomal recessive disease characterized by viral warts and skin cancer. A model for viral oncogenesis. Bull Cancer 65:169–182. [PubMed] [Google Scholar]
- 18. Ramoz N, Rueda LA, Bouadjar B, Favre M, Orth G. 1999. A susceptibility locus for epidermodysplasia verruciformis, an abnormal predisposition to infection with the oncogenic human papillomavirus type 5, maps to chromosome 17qter in a region containing a psoriasis locus. J Invest Dermatol 112:259–263. doi: 10.1046/j.1523-1747.1999.00536.x [DOI] [PubMed] [Google Scholar]
- 19. Ramoz N, Taïeb A, Rueda LA, Montoya LS, Bouadjar B, Favre M, Orth G. 2000. Evidence for a nonallelic heterogeneity of epidermodysplasia verruciformis with two susceptibility loci mapped to chromosome regions 2p21–p24 and 17q25. J Invest Dermatol 114:1148–1153. doi: 10.1046/j.1523-1747.2000.00996.x [DOI] [PubMed] [Google Scholar]
- 20. Kurima K, Yang Y, Sorber K, Griffith AJ. 2003. Characterization of the transmembrane channel-like (TMC) gene family: functional clues from hearing loss and epidermodysplasia verruciformis☆. Genomics 82:300–308. doi: 10.1016/s0888-7543(03)00154-x [DOI] [PubMed] [Google Scholar]
- 21. Keresztes G, Mutai H, Heller S. 2003. TMC and EVER genes belong to a larger novel family, the TMC gene family encoding transmembrane proteins. BMC Genomics 4:24. doi: 10.1186/1471-2164-4-24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Ramoz N, Rueda L-A, Bouadjar B, Montoya L-S, Orth G, Favre M. 2002. Mutations in two adjacent novel genes are associated with epidermodysplasia verruciformis. Nat Genet 32:579–581. doi: 10.1038/ng1044 [DOI] [PubMed] [Google Scholar]
- 23. Lazarczyk M, Dalard C, Hayder M, Dupre L, Pignolet B, Majewski S, Vuillier F, Favre M, Liblau RS. 2012. EVER proteins, key elements of the natural anti-human papillomavirus barrier, are regulated upon T-cell activation. PLoS One 7:e39995. doi: 10.1371/journal.pone.0039995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Kalińska-Bienias A, Kowalewski C, Majewski S. 2016. The EVER genes – the genetic etiology of carcinogenesis in epidermodysplasia verruciformis and a possible role in non-epidermodysplasia verruciformis patients. Postepy Dermatol Alergol 33:75–80. doi: 10.5114/ada.2016.59145 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Ingle A, Ghim S, Joh J, Chepkoech I, Bennett Jenson A, Sundberg JP. 2011. Novel laboratory mouse papillomavirus (MusPV) infection. Vet Pathol 48:500–505. doi: 10.1177/0300985810377186 [DOI] [PubMed] [Google Scholar]
- 26. Cladel NM, Budgeon LR, Cooper TK, Balogh KK, Christensen ND, Myers R, Majerciak V, Gotte D, Zheng Z-M, Hu J. 2017. Mouse papillomavirus infections spread to cutaneous sites with progression to malignancy. J Gen Virol 98:2520–2529. doi: 10.1099/jgv.0.000926 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Uberoi A, Yoshida S, Frazer IH, Pitot HC, Lambert PF. 2016. Role of ultraviolet radiation in papillomavirus-induced disease. PLoS Pathog 12:e1005664. doi: 10.1371/journal.ppat.1005664 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Hu J, Cladel NM, Budgeon LR, Balogh KK, Christensen ND. 2017. The mouse papillomavirus infection model. Viruses 9:246. doi: 10.3390/v9090246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Uberoi A, Lambert PF. 2017. Rodent papillomaviruses. Viruses 9:362. doi: 10.3390/v9120362 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Spurgeon ME, Lambert PF. 2020. Mus musculus papillomavirus 1: a new frontier in animal models of papillomavirus pathogenesis. J Virol 94:e00002-20. doi: 10.1128/JVI.00002-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Christensen ND, Budgeon LR, Cladel NM, Hu J. 2017. Recent advances in preclinical model systems for papillomaviruses. Virus Res 231:108–118. doi: 10.1016/j.virusres.2016.12.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Spurgeon ME, Uberoi A, McGregor SM, Wei T, Ward-Shaw E, Lambert PF. 2019. A novel in vivo infection model to study papillomavirus-mediated disease of the female reproductive tract. mBio 10:e00180-19. doi: 10.1128/mBio.00180-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Blaine-Sauer S, Shin M-K, Matkowskyj KA, Ward-Shaw E, Lambert PF. 2021. A novel model for papillomavirus-mediated anal disease and cancer using the mouse papillomavirus. mBio 12:e0161121. doi: 10.1128/mBio.01611-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Wei T, Buehler D, Ward-Shaw E, Lambert PF. 2020. An infection-based murine model for papillomavirus-associated head and neck cancer. mBio 11:e00908-20. doi: 10.1128/mBio.00908-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Joh J, Jenson AB, King W, Proctor M, Ingle A, Sundberg JP, Ghim S. 2011. Genomic analysis of the first laboratory-mouse papillomavirus. J Gen Virol 92:692–698. doi: 10.1099/vir.0.026138-0 [DOI] [PubMed] [Google Scholar]
- 36. Xue X-Y, Majerciak V, Uberoi A, Kim B-H, Gotte D, Chen X, Cam M, Lambert PF, Zheng Z-M. 2017. The full transcription map of mouse papillomavirus type 1 (MmuPV1) in mouse wart tissues. PLoS Pathog 13:e1006715. doi: 10.1371/journal.ppat.1006715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Lazarczyk M, Pons C, Mendoza J-A, Cassonnet P, Jacob Y, Favre M. 2008. Regulation of cellular zinc balance as a potential mechanism of EVER-mediated protection against pathogenesis by cutaneous oncogenic human papillomaviruses. J Exp Med 205:35–42. doi: 10.1084/jem.20071311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Romero-Masters JC, Lambert PF, Munger K. 2022. Molecular mechanisms of MmuPV1 E6 and E7 and implications for human disease. Viruses 14:2138. doi: 10.3390/v14102138 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Torres AD, Spurgeon ME, Bilger A, Blaine-Sauer S, Uberoi A, Buehler D, McGregor SM, Ward-Shaw E, Lambert PF. 2020. The human papillomavirus 16 E5 gene potentiates MmuPV1-dependent pathogenesis. Virology 541:1–12. doi: 10.1016/j.virol.2019.12.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Su AI, Wiltshire T, Batalov S, Lapp H, Ching KA, Block D, Zhang J, Soden R, Hayakawa M, Kreiman G, Cooke MP, Walker JR, Hogenesch JB. 2004. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc Natl Acad Sci U S A 101:6062–6067. doi: 10.1073/pnas.0400782101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Gaud G, Guillemot D, Jacob Y, Favre M, Vuillier F. 2013. EVER2 protein binds TRADD to promote TNF-α-induced apoptosis. Cell Death Dis 4:e499. doi: 10.1038/cddis.2013.27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Stubenrauch F, Straub E, Klein K, Kramer D, Iftner T, Wong M, Roden RBS. 2021. Expression of E8^E2 is required for wart formation by mouse papillomavirus 1 in vivo. J Virol 95:e01930-20. doi: 10.1128/JVI.01930-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Rehm TM, Straub E, Forchhammer S, Leiter U, Iftner T, Stubenrauch F. 2022. Transcription properties of beta-HPV8 and HPV38 genomes in human keratinocytes. J Virol 96:e0149822. doi: 10.1128/jvi.01498-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Wang W, Uberoi A, Spurgeon M, Gronski E, Majerciak V, Lobanov A, Hayes M, Loke A, Zheng Z-M, Lambert PF. 2020. Stress keratin 17 enhances papillomavirus infection-induced disease by downregulating T cell recruitment. PLoS Pathog 16:e1008206. doi: 10.1371/journal.ppat.1008206 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Crequer A, Picard C, Pedergnana V, Lim A, Zhang S-Y, Abel L, Majewski S, Casanova J-L, Jablonska S, Orth G, Jouanguy E. 2013. EVER2 deficiency is associated with mild T-cell abnormalities. J Clin Immunol 33:14–21. doi: 10.1007/s10875-012-9749-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Leisner TM, Freeman TC, Black JL, Parise LV. 2016. CIB1: a small protein with big ambitions. FASEB J 30:2640–2650. doi: 10.1096/fj.201500073R [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Imahorn E, Aushev M, Herms S, Hoffmann P, Cichon S, Reichelt J, Itin PH, Burger B. 2020. Gene expression is stable in a complete CIB1 knockout keratinocyte model. Sci Rep 10:14952. doi: 10.1038/s41598-020-71889-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Olczak P, Wong M, Tsai H-L, Wang H, Kirnbauer R, Griffith AJ, Lambert PF, Roden R. 2022. Vaccination with human alphapapillomavirus-derived L2 multimer protects against human betapapillomavirus challenge, including in epidermodysplasia verruciformis model mice. Virology 575:63–73. doi: 10.1016/j.virol.2022.08.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Olczak P, Matsui K, Wong M, Alvarez J, Lambert P, Christensen ND, Hu J, Huber B, Kirnbauer R, Wang JW, Roden RBS. 2022. RG2-VLP: a vaccine designed to broadly protect against anogenital and skin human papillomaviruses causing human cancer. J Virol 96:e0056622. doi: 10.1128/jvi.00566-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Bilger A, King RE, Schroeder JP, Piette JT, Hinshaw LA, Kurth AD, AlRamahi RW, Barthel MV, Ward-Shaw ET, Buehler D, Masters KS, Thibeault SL, Lambert PF. 2020. A mouse model of oropharyngeal papillomavirus-induced neoplasia using novel tools for infection and nasal anesthesia. Viruses 12:450. doi: 10.3390/v12040450 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figures S1 to S6.
Data Availability Statement
Data supporting the findings of this study are available from the corresponding author upon request.