Abstract
Prostate cancer with a cribriform pattern, including invasive cribriform carcinoma (ICC) and/or intraductal carcinoma (IDC) is associated with a poor prognosis, and the underlying mechanisms are unclear. Therefore, we aimed to identify biomarkers for this feature. Using a radical prostatectomy cohort, we performed within-patient differential expression analyses with RNA sequencing data to compare samples with a cribriform pattern to those with non-cribriform Gleason pattern 4 (NcGP4; n=13). ACSM1, GRIN3A, PCDHB2, and REG4 were identified as differentially expressed, and validation was performed using real-time reverse transcription polymerase chain reaction (n=99; 321 RNA samples) and RNA in situ hybridization on tissue microarrays (n=479; 2047 tissue cores). GRIN3A was significantly higher expressed in cribriform pattern vs. NcGP4, when assessed within the same patient (n=27; p=0.005) and between different patients (n=83; p=0.001). Tissue cores with IDC more often expressed GRIN3A compared to ICC, NcGP4, and benign tissue (52 % vs. ≤ 32 %). When IDC and NcGP4 was compared within the same patient (173 pairs of tissue cores; 54 patients), 38 (22 %) of the tissue microarray core pairs had GRIN3A expression in only IDC, 33 (19 %) had expression in both IDC and NcGP4, 14 (8 %) in only NcGP4 and 88 (51 %) were negative in both entities (p=0.001). GRIN3A was as well associated with biochemical recurrence (log-rank, p=0.002). In conclusion, ectopic GRIN3A expression is an RNA-based biomarker for the presence of cribriform prostate cancer, particularly for IDC.
Keywords: Biomarker, Intraductal carcinoma, Invasive cribriform carcinoma, Prostate cancer, RNA sequencing, RNA in situ hybridization
Abbreviations
- BCR
biochemical recurrence
- CR
clinical recurrence
- cT-stage
clinical tumor stage
- CT
cycle threshold
- DGEA
differential gene expression analysis
- FFPE
formalin-fixed paraffin-embedded
- GP4
Gleason pattern 4
- HGPIN
high-grade prostatic intraepithelial neoplasia
- ICC
invasive cribriform carcinoma
- IDC
intraductal carcinoma
- IQR
interquartile range
- ISUP
International Society of Urological Pathology
- MRI
magnetic resonance imaging
- PSA
prostate-specific antigen
- pT-stage
pathological tumor stage
- RP
radical prostatectomy
- RT-PCR
reverse transcription polymerase chain reaction
- TMA
tissue microarray
Introduction
Prostate cancer is characterized by substantial clinical, morphological, and molecular heterogeneity, reflected in the highly variable outcomes among patients. Despite the available diagnostic tools for risk stratification, including Gleason score/Grade Group, biomarkers are still needed to improve clinical decision-making. Throughout the past decade, the prognostic value of specific morphological features in prostate cancer has gained increasing attention. Two such features are intraductal carcinoma (IDC) and invasive cribriform carcinoma (ICC)—the latter being a subtype of Gleason pattern 4 (GP4)—where several studies have demonstrated their association with a poorer prognosis for prostate cancer patients [[1], [2], [3], [4], [5], [6], [7], [8], [9], [10]].
Previous studies have demonstrated DNA, RNA, and protein aberrations in tumors with a cribriform pattern (IDC and/or ICC) compared to tumors without this feature [11]. The aberrations include loss of tumor suppressor genes, e.g., PTEN and NKX3-1, amplification of MYC, and point mutations in SPOP ([12,13]). Mutations causing defects in DNA repair genes, such as BRCA2 and CDK12, have been reported to be associated specifically with IDC [14]. Tumors with a cribriform pattern more often have a high RNA expression of the long non-coding RNA SCHLAP1 ([15,16]), and higher so-called genomic risk score as determined by commercially available RNA-based tests performed on samples from needle biopsies [17] and radical prostatectomy (RP) specimens ([18,19]). One study discovered a relation between cribriform pattern and the molecular subtype luminal proliferating type B [19], which is a PAM50 classifier subtype [20] that has been linked to a poor prognosis in prostate cancer [21]. Additionally, tumors with a cribriform pattern have been found to have a downregulation of apoptotic pathways, with concurrent upregulation of proliferative and MYC pathway signaling [19]. Alterations in protein expression in tissue with a cribriform pattern include PTEN loss, which has been shown to occur more frequently in both IDC and ICC than in malignant tissue without a cribriform pattern ([22,23]).
Summarized, previous studies support the association of a cribriform pattern with more aggressive disease. With few exceptions, previous research has not differentiated between IDC and ICC. Therefore, there is a need for molecular biomarkers that distinguish between these entities. Here we applied a methodological approach of RNA differential gene expression analysis within different malignant phenotypes, which identified GRIN3A as a novel biomarker for a subtype of IDC and poor outcome in prostate cancer.
Materials and methods
Prospective cohort and prostate cancer biobank
The prospective cohort is comprised of 571 patients treated with RP at Oslo University Hospital–Radiumhospitalet between 2010 and 2012, as previously described ([16,[24], [25], [26]]). The prostate cancer biobank includes both formalin-fixed paraffin-embedded tissue (FFPE) slides and blocks, in addition to three to seven fresh-frozen tissue samples from each patient. From the fresh-frozen tissue samples, total RNA was isolated and complementary DNA (cDNA) generated, as previously described ([16,24]). In the present study, 515 treatment-naïve patients without distant metastases at time of surgery and with available tissue slides were included (Table S1). Among the 515 patients, different subseries were included in the various analyses (Fig. 1 and Table S2). The median follow-up time for all 515 patients was 8.74 years (interquartile range [IQR]: 7.68–9.82 years).
Fig. 1.
Patient inclusion in the different analyses. Schematic illustration showing overlap of patient inclusion for the different analyses. Abbreviations: RNA-seq, RNA sequencing; RT-PCR, reverse transcription polymerase chain reaction.
The study was undertaken with the understanding and written consent of each patient included in the study. The study methodologies conformed to the standards set by the Declaration of Helsinki and were approved by the regional Ethics Committee South-East Norway (2013/595).
Histopathological evaluation and tissue microarrays
For all 515 RP specimens, histopathological evaluation was performed by two investigators (MB and UA) according to the 2014 International Society of Urological Pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma [27], and evaluation of morphological features was performed as previously described [26]. We defined IDC as a proliferation of malignant epithelial cells within predefined acini/ducts and with at least a partially preserved basal cell layer based on the definition by Montironi et al. [28], and ICC as sheets of cells with multiple glandular lumina without intervening stroma based on the definition as stated by Kweldam et al. [6].
The areas surrounding the fresh-frozen tissue sampling (range 3–7 per patient) were evaluated on the whole-mount tissue sections for the presence of malignant and/or benign tissue and for the morphological features: cribriform IDC, ICC, and non-cribriform GP4 (ill-defined, fused, and glomeruloid). In cases with cribriform ICC or IDC and comedonecrosis, the cribriform gland was graded as Gleason pattern 5 (rather than Gleason pattern 4), and were not included in analyses. If no malignant tissue was present, the fresh-frozen sample was characterized as a benign sample. Benign samples also included high-grade prostatic intraepithelial neoplasia (HGPIN).
Regions of interest were selected and included on tissue microarrays (TMAs), as previously described [26]. Inter- and intrafocal heterogeneity were evaluated, following previously described criteria [26] on the TMAs for 152 and 306 patients, respectively.
Differential gene expression analysis from RNA sequencing data
Whole transcriptome short-read paired-end RNA sequencing data was available from 23 of the patients [29]. These patients were originally selected as they had fresh-frozen samples available from multiple malignant foci [24]. Briefly, for each patient, three to four fresh-frozen tissue samples were sequenced, which consisted of two to three malignant samples and one benign sample, rendering a total of 87 samples (64 malignant, 23 benign). The RNA sequencing data had an average of 44 million uniquely mapped read pairs per sample [29].
Differential gene expression analysis (DGEA) was performed on the RNA sequencing data using the “DESeq2” R package (version 1.24.0) [30]. A paired sampling design was used to compare samples with a cribriform pattern (IDC and/or ICC) to samples with non-cribriform GP4 from within the same patient (intrapatient pairwise comparison). In total, 13 patients with 32 samples (15 with cribriform pattern and 17 with non-cribriform GP4) were included. Raw read counts quantified by HTSeq (version 0.10.0) [31] were provided as input. P-values were adjusted for multiple testing by independent hypothesis weighting provided by the “IHW” R package (version 1.12.0) [32], weighted by the average expression of each gene across all samples. Genes with absolute log2 fold changes greater than 1 and adjusted p-values less than 0.05 were considered significantly higher expressed in cribriform pattern compared to non-cribriform GP4. Genes with a log2 fold change less than -1 were considered significantly lower expressed in cribriform pattern samples compared to non-cribriform GP4. Four genes that were significantly associated with the cribriform pattern samples were selected for further analysis.
Semi-quantitative gene expression analyses
Real-time reverse transcription polymerase chain reaction (RT-PCR) was performed to 1) corroborate the RNA abundance of the four candidate genes for the patients included in DGEA, 2) validate results using a paired sampling design (intrapatient pairwise comparison) in a larger cohort of patients not included in DGEA, and 3) assess findings from the DGEA in a larger cohort of patients using an unpaired sampling design (interpatient comparison). In total, real-time RT-PCR was performed on RNA from 321 prostatic tissue samples from 99 patients, of which 211 samples were from malignant and 110 from benign tissue.
TaqMan Gene expression assays were used to investigate five genes (Thermo Fisher Scientific, Waltham, MA, USA): Hs01048218_m1 (ACSM1), Hs01077968_m1 (GRIN3A), Hs00251764_s1 (PCDHB2), Hs00230746_m1 (REG4), and Hs01104728_m1 (ABL1). All samples were run in triplicates on an ABI 7900HT Fast Real-time PCR System (Thermo Fisher Scientific), with 10 ng cDNA input in each 10 μl reaction.
Cycle threshold (CT) values were recorded for all reactions, and the median of each sample triplicate was used for analyses. CT medians > 35 and “undetermined” were considered as “below threshold.” Expression levels for each of the four candidate genes were normalized to the reference gene ABL1 using the standard curve method. The ratio between the median relative quantity of each of the candidate genes and ABL1 in the benign tissue samples was used as a calibrator.
To investigate the correlation between RNA sequencing and real-time RT-PCR results, the Pearson correlation coefficient was calculated using non-transformed expression values for both methods. All samples that were RNA sequenced from the 13 patients in the DGEA were included, rendering a total of 48 samples (35 malignant, 13 benign). However, only samples with values above threshold (CT median < 35) were included in correlation analyses.
To further validate the results from the DGEA, a separate, larger confirmation cohort of 27 patients with 78 samples (43 cribriform pattern, 35 non-cribriform GP4) were analyzed using real-time RT-PCR. Among the 27 patients, 13 patients had more than one sample with the same morphological feature, either cribriform pattern or non-cribriform GP4. Therefore, if a patient had, e.g., two samples classified as non-cribriform GP4 and only one sample with a cribriform pattern, the same cribriform pattern sample was used twice to allow for the comparison with both non-cribriform GP4 samples.
In the analysis with an unpaired sampling design (interpatient comparison), samples with a cribriform pattern were compared to samples with a non-cribriform GP4. To ensure that the samples being compared were independent of each other, i.e., not from the same patient, samples with a cribriform pattern were compared to samples with a non-cribriform GP4 from patients without any cribriform pattern present in their prostates. In total, 83 patients (52 patients with a cribriform pattern, 31 patients with only a non-cribriform GP4) with a total of 121 samples (79 cribriform pattern, 42 non-cribriform GP4) were included. All patients included in the DGEA, in addition to the larger, confirmation cohort were included in this analysis.
The expression levels of the four candidate genes were also compared in malignant versus benign samples using real-time RT-PCR. In total, 93 patients with at least one malignant and one benign sample were included, with a total of 301 samples (191 malignant, 110 benign). Both patients from the DGEA and additional patients were included.
RNA in situ hybridization
RNA in situ hybridization was performed to investigate the expression of the candidate genes within the context of tissue morphology and to assess a larger cohort, including both patients from the DGEA and additional patients (Fig. 1). RNA in situ hybridization was performed on the TMAs using the RNAscope® VS Universal HRP assay from Advanced Cell Diagnostics (ACD, Hayward, CA, USA, catalog number: 323200) [33] on the Discovery Ultra Platform (Roche Tissue Diagnostics, Tucson, AZ, USA) according to the manufacturer's recommendations.
The housekeeping gene PPIB (RNAscope® 2.5 VS positive control probe_Hs-PPIB (catalog number: 313909) was used as a positive control on selected TMAs to assess RNA quality. Target probes included ACSM1 (RNAscope® 2.5 VS probe-Hs-ACSM1, catalog number: 867949), GRIN3A (RNAscope® 2.5 VS Probe-Hs-GRIN3A, catalog number: 534849), PCDHB2 (RNAscope® 2.5 VS Probe-Hs-PCDHB2, catalog number: 867959), and REG4 (RNAscope® 2.5 VS Probe-Hs-REG4, catalog number: 312079). The DISCOVERY mRNA Purple HRP Detection kit (RUO) (catalog number: 760-255, Roche Tissue Diagnostics) was used for visualization of target probes. The evaluation of RNA abundance was visually scored, and each tissue core was graded as either “negative” or “positive.” A tissue core was graded as “positive” if at least two glands showed any “positive” staining in malignant cells, indicated by purple “dots” or “clusters,” and an RP specimen was classified as “positive” if any malignant sample from that RP specimen was “positive.”
On the TMAs, 479 patients (93 %) had at least one malignant sample that could be evaluated by RNAscope for all four target probes. For these 479 patients, a total of 2047 tissue cores (1425 malignant, 622 benign) could be evaluated.
Immunohistochemistry
In addition to morphologic evaluation, immunohistochemistry was used to differentiate between IDC and ICC, with a cocktail of three antibodies: Cytokeratin, High Molecular Weight (34BE12; clone: 34βE12; dilution: 1:20; Agilent, Santa Clara, CA, USA), Tumor protein 63 (p63; clone: DAK-p63; dilution: 1:20; Agilent), and α-methylacyl coenzyme A racemase (AMACR)/p504s (clone: 13H4; dilution: 1:50; Agilent). Immunohistochemical staining for the cocktail of three antibodies was performed using the Ventana Benchmark Ultra system (Roche Tissue Diagnostics) with the Ventana UltraView Universal DAB Detection Kit (product number: 760-500, Roche Tissue Diagnostics) or UltraView Universal Alkaline Phosphatase Red (product number: 760-501, Roche Tissue Diagnostics).
Immunohistochemistry for PTEN was also used in cases when there was uncertainty as to whether IDC or HGPIN was present. The antibody used was anti-human PTEN (clone 6H2.1; dilution 1:100; Agilent).
PTEN staining was performed using DAKO Autostainer Link (Agilent) and Dako EnV FLEX+, High pH (Link) detection kit (product number: K8002; Agilent). Controls for PTEN included PTEN negative endometrioid adenocarcinoma, PTEN positive clear cell adenocarcinoma, and tonsillar tissue.
If a malignant gland with cribriform morphology had any positive (brown) basal cell staining, the cribriform structure was characterized as IDC, and not ICC. If PTEN expression was absent (indicated by lack of brown nuclear and cytoplasmic staining in malignant cells) and there was more than moderate atypia (or severe atypia) in the cribriform structure, the cribriform structure was characterized as IDC, and not HGPIN.
Statistical analysis
All statistical analyses were performed using R (version 4.1.1) and RStudio (version 1.4.1717), with the packages “ggplot2” (version 3.3.5), “ggpubr” (version 0.4.0), “survival” (version 3.2-11), and “survminer” (version 0.4.9). A p-value of 0.05 was used as the threshold for statistical significance. IDC and ICC were grouped together as “cribriform pattern” for all analyses, except when assessing results from RNA in situ hybridization on the TMA core level.
For statistical analyses of real-time RT-PCR data, the Wilcoxon signed-rank test was used to compare malignant to benign samples using a paired analysis, or to compare cribriform pattern samples to non-cribriform GP4 samples using either a paired or unpaired analysis. For each separate analysis, samples “below threshold” were given a value 0.5 below the lowest log2 fold change for each gene, so that they could be included in plots and statistical analyses.
For analyses of RNA in situ hybridization data, the χ2 test of independence or Fisher's exact test (categorical), or Wilcoxon rank-sum test (continuous) was used compare RP specimens with “positive” expression of a particular candidate gene to RP specimens with “negative” expression. For the paired analyses comparing ICC vs. IDC, ICC vs. non-cribriform GP4 and IDC vs. non-cribriform GP4 (intrapatient pairwise comparison), McNemar's test was used. Similar to the paired analyses performed using the real-time RT-PCR data, if a patient had e.g., two samples classified as ICC and only one sample classified as IDC, the same IDC sample was used twice to allow for comparisons with both ICC samples.
Time-to-event analyses
For time-to-event analyses using RNA in situ hybridization data, patients with persistently elevated prostate-specific antigen (PSA) levels and/or patients who received adjuvant treatment (n = 58) were excluded. In addition, one patient with unavailable information on the preoperative PSA was excluded, leaving a total of 420 patients included in time-to-event analyses. The Kaplan-Meier method and log-rank test was used to compare time to events, either biochemical recurrence (BCR) or clinical recurrence (CR). Univariable and multivariable Cox regression analyses were used to calculate hazard ratios with 95 % confidence intervals for BCR and CR. Only variables that were significant in univariable analyses were included in multivariable analyses. Time-to-event analyses using metastatic castration-resistant prostate cancer and prostate cancer-related death as endpoints were not performed due to few events.
BCR was defined as PSA ≥ 0.2 ng/ml in two blood samples collected at least one week apart. CR was defined as local recurrence in the prostatic bed or seminal vesicles, or metastasis to lymph nodes or distant sites. Time to BCR and CR was defined as the time from date of surgery to verified BCR or start of treatment for CR, respectively. Patients not experiencing BCR or CR were censored at the last date of follow-up. The proportional hazards assumption was assessed using the “cox.zph” function implemented in the “survival” R package.
Results
Differentially expressed genes between cribriform pattern and non-cribriform GP4 samples
To discover novel genes associated with a cribriform pattern (IDC and/or ICC), DGEA with a paired sampling design was performed using whole transcriptome short-read paired-end RNA sequencing data. RNA samples from tissue with a cribriform pattern were compared to non-cribriform GP4 tissue from the same patients. In total, 154 genes were significantly differentially expressed, whereof 10 genes were higher expressed in the cribriform pattern samples and 144 genes were higher in the non-cribriform GP4 samples (Tables S3 and S4). Among the cribriform-associated genes, ACSM1, GRIN3A, PCDHB2, and REG4 were selected as candidates for further investigation based on previous knowledge of their potential relevance in prostate cancer [[34], [35], [36], [37], [38], [39]], and/or intrapatient separation of the gene expression levels in cribriform pattern compared to non-cribriform GP4 samples.
Experimental corroboration of RNA levels of selected cribriform-associated genes
Experimental corroboration of gene expression estimates from RNA sequencing data was performed using real-time RT-PCR for the four cribriform-associated genes in the same sample set. There was a strong correlation between the two methods for all four genes (Fig. 2A and Fig. S1A–C; all Pearson correlation coefficients r ≥ 0.95). Furthermore, it appeared that the samples with cribriform pattern present clustered together for three of the four genes (ACSM1, GRIN3A, and PCDHB2), and were the samples with higher log2 fold changes for both methods.
Fig. 2.
Assessment of GRIN3A expression levels. (A) Plot showing the correlation between GRIN3A expression levels as determined by RNA sequencing and real-time RT-PCR. Each dot represents one sample from a patient that was included in the differential gene expression analysis with a paired sampling analysis of cribriform pattern versus non-cribriform Gleason pattern 4. (B) Paired analysis (intrapatient pairwise comparison) comparing log2 fold changes for GRIN3A expression in cribriform samples to non-cribriform Gleason pattern 4 samples. Samples from patients in an independent subcohort were included (n = 27). Each dot represents one sample, and gray lines between samples indicate the sample pair being compared. Box plots show median relative log2 expression, first and third quartile, and maximum and minimum log2 values. (C) Unpaired analysis (interpatient pairwise comparison) comparing log2 fold changes for GRIN3A expression in cribriform samples to non-cribriform Gleason pattern 4 samples (n = 83). Abbreviations: GP3, Gleason pattern 3; NcGP4, non-cribriform Gleason pattern 4; RT-PCR, reverse transcription polymerase chain reaction; * statistical significance.
Validation of gene expression levels using real-time RT-PCR
Validation of the results from the DGEA was performed using real-time RT-PCR in a larger patient cohort not included in the DGEA (Fig. 2B, Fig. S1D–F, and Table S5). Of the four candidate genes, only GRIN3A was significantly higher expressed in cribriform pattern samples compared to non-cribriform GP4 samples (Fig. 2B; p=0.005).
Interpatient comparison of gene expression levels in cribriform versus non-cribriform Gleason pattern 4 samples
To determine whether the candidate genes were higher expressed in samples with a cribriform pattern compared to samples from RP specimens without any cribriform pattern, an unpaired sampling analysis (interpatient comparison) was performed using expression levels determined by real-time RT-PCR (Figs. 2C and S1G–I). Only GRIN3A was significantly higher expressed in cribriform pattern samples (Fig. 2C; p=0.001).
Gene expression in the context of tissue morphology
The expression of the four candidate genes was then assessed in the context of tissue morphology using RNA in situ hybridization (RNAscope®) on TMAs, including a larger patient cohort (Table S6). Here, we identified which cells expressed the candidate genes, and specifically whether they were expressed in IDC and/or ICC. Expression of all four genes was confined to epithelial cells and was not detected in stromal cells (Fig. 3).
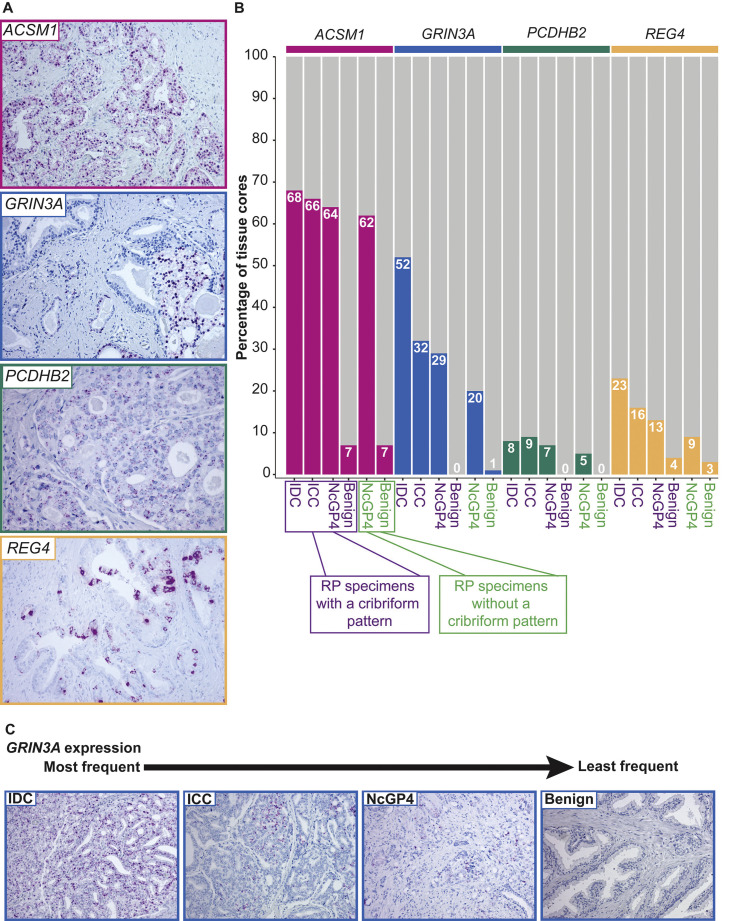
Fig. 3.
Assessment of ACSM1, GRIN3A, PCDHB2, and REG4 expression by RNA in situ hybridization. (A) Example images of “positive” staining for the four candidate genes (ACSM1, GRIN3A, PCDHB2, and REG4), at 10x magnification for ACSM1,GRIN3A and REG4, and 20x magnification for PCDHB2. (B) Stacked bar charts showing the percentage of samples with either “positive” or “negative” expression of the candidate genes. Classification of the tissue cores by morphological feature is shown on the x-axis, and colored according to whether they are from radical prostatectomy specimens with (purple) or without (green) a cribriform pattern. (C) Example images (10x magnification) of a selection of the different morphological features and their GRIN3A expression in descending order of frequency. Abbreviations: ICC, invasive cribriform carcinoma; IDC, intraductal carcinoma; NcGP4, non-cribriform Gleason pattern 4; RP, radical prostatectomy.
ACSM1 was most commonly identified in RP specimens (78 %), followed by GRIN3A (39 %), REG4 (21 %), and PCDHB2 (12 %). Interfocal heterogeneity was evaluated in patients with “positive” expression of the candidate genes in malignant tissue, and with at least one sample from two different malignant foci. Interfocal heterogeneity was identified for ACSM1 expression in 43 % (56/131) of RP specimens, in 85 % (55/65) for GRIN3A, 65 % (13/20) for PCDHB2, and 88 % (38/43) for REG4. For patients with multiple samples from a focus defined as “positive,” intrafocal heterogeneity was evaluated. Intrafocal heterogeneity was identified for ACSM1 expression in 51 % (129/252) of RP specimens, in 73 % (103/141) for GRIN3A, 84 % (42/50) for PCDHB2, and 85 % (62/73) for REG4.
We then compared tissue cores with IDC or ICC to non-cribriform GP4. ACSM1 and PCDHB2 expression were detected in a similar percentage of tissue cores with the different morphological features, while differences in the percentage of tissue cores depending on the morphological feature were identified for GRIN3A and REG4 expression (Fig. 3B). Tissue cores with IDC most commonly had GRIN3A expression, whereas ICC and non-cribriform GP4 demonstrated similar expression levels for this gene (Fig. 3B–C). However, we did observe a field effect as not only was IDC characterized as positive, but also neighboring malignant glands. Similar findings were identified for REG4, although the expression was often limited to a few cells within a particular malignant gland.
Paired analyses (intrapatient pairwise comparisons) comparing IDC to ICC, IDC to non-cribriform GP4, and ICC to non-cribriform GP4 were also performed specifically for GRIN3A. A statistically significant association was identified when comparing IDC to non-cribriform Gleason pattern 4 (p=0.001), but not for the other comparisons comparing IDC to ICC (p=1) nor ICC to non-cribriform GP4 (p=0.1). Of a total of 173 pairs of tissue cores from 54 patients that had tissue cores with IDC and also non-cribriform GP4, 38 (22 %) had GRIN3A expression only in IDC and not non-cribriform GP4, 33 (19 %) had GRIN3A expression in both IDC and non-cribriform GP4, 14 (8 %) had only GRIN3A expression in non-cribriform GP4 and 88 (51 %) did not have GRIN3A expression in IDC nor non-cribriform GP4.
Interestingly, all four candidate genes were less commonly expressed in benign compared to malignant samples (Fig. 3B). We explored this further using real-time RT-PCR and a paired sample analysis. We found that all four candidate genes were significantly lower expressed in benign samples compared to malignant samples (Table S7 and Fig. S2).
Associations with clinicopathological parameters
Associations between expression of the four candidate genes, as assessed by RNA in situ hybridization, and clinicopathological parameters are summarized in Table 1 and Tables S8–S10. For three of the four genes (ACSM1, GRIN3A, and REG4), RP specimens with “positive” expression more often had a cribriform pattern compared to RP specimens without a cribriform pattern. RP specimens with GRIN3A expression were more often classified as having higher Grade Groups in preoperative needle biopsies and RP specimens, advanced clinical tumor (cT) and pathological tumor (pT) stages, and reactive stroma. Moreover, a significant association between GRIN3A expression and the presence of a post-RP persistent PSA was identified (Table 1).
Table 1.
Clinicopathological characteristics for prostate cancer patients evaluated with RNA in situ hybridization, stratified by presence or absence of GRIN3A expression in malignant tissue. A patient was classified as GRIN3A-positive (+) if at least one malignant sample was GRIN3A-positive. The χ2 test of independence was used for categorical variables unless otherwise specified. The Wilcoxon rank-sum test was used for continuous variables. a Fisher's exact test used; b analysis only comparing pN0 versus pN1; c analysis only comparing patients with minor high-grade pattern 5 absent versus present. Patients with unavailable/missing data for a specific variable were excluded from analyses. Abbreviations: cT-stage, clinical tumor stage; IQR, interquartile range; mCRPC, metastatic castration-resistant prostate cancer; mpMRI, multiparametric magnetic resonance imaging; n, number of patients; Nbx, needle biopsy; PSA, prostate-specific antigen; pN-stage, pathological node stage; pT-stage, pathological tumor stage; RP, radical prostatectomy; rT, radiological tumor stage; * statistical significance.
| Characteristic | GRIN3A + (n = 188; 39 %) | GRIN3A – (n = 291; 61 %) | p |
|---|---|---|---|
| Age at time of RP, median (IQR) | 64 (60–69) | 64 (59–67) | 0.4 |
| Preoperative PSA group, n (%) | 0.7 | ||
| < 10 ng/ml | 91 (48) | 132 (45) | |
| 10–20 ng/ml | 67 (36) | 103 (35) | |
| > 20 ng/ml | 30 (16) | 55 (19) | |
| Missing | 0 (0) | 1 (0.3) | |
| Grade Group for preoperative Nbx, n (%) | 0.01* | ||
| 1 | 31 (16) | 73 (25) | |
| 2 | 65 (35) | 118 (41) | |
| 3 | 46 (24) | 46 (16) | |
| 4 | 31 (16) | 42 (14) | |
| 5 | 15 (8) | 12 (4) | |
| cT-stage, n (%) | < 0.001* | ||
| cT1 | 73 (39) | 157 (54) | |
| cT2 | 79 (42) | 101 (35) | |
| cT3 | 35 (19) | 27 (9) | |
| Missing | 1 (1) | 6 (2) | |
| Preoperative mpMRI-stage, n (%) | 0.001*,a | ||
| rT2 | 65 (35) | 124 (43) | |
| rT3 | 78 (41) | 72 (25) | |
| rT4 | 1 (0.5) | 0 (0) | |
| Not available | 44 (23) | 95 (33) | |
| Multifocal cancer, n (%) | 116 (62) | 206 (71) | 0.048* |
| Grade Group for RP specimen, n (%) | 0.002* | ||
| 1 | 4 (2) | 25 (9) | |
| 2 | 74 (39) | 138 (47) | |
| 3 | 61 (32) | 72 (25) | |
| 4 | 20 (11) | 31 (11) | |
| 5 | 29 (15) | 25 (9) | |
| pT-stage, n (%) | < 0.001* | ||
| pT2 | 47 (25) | 154 (53) | |
| pT3a | 110 (59) | 111 (38) | |
| pT3b | 31 (16) | 25 (9) | |
| Missing | 0 (0) | 1 (0.3) | |
| pN-stage, n (%) | 0.1 | ||
| pN0 | 47 (25) | 70 (24) | |
| pN1 | 13 (7) | 8 (3) | |
| pNX | 128 (68) | 213 (73) | |
| Positive surgical margins, n (%) | 37 (20) | 41 (14) | 0.1 |
| Minor high-grade pattern 5, n (%) | 0.5 | ||
| Present | 32 (17) | 42 (14) | |
| Absent | 103 (55) | 168 (58) | |
| Not applicable | 53 (28) | 81 (28) | |
| Reactive stroma, n (%) | 92 (49) | 93 (32) | < 0.001* |
| Cribriform pattern, n (%) | 140 (74) | 132 (45) | < 0.001* |
| Post-RP persistent PSA, n (%) | 23 (12) | 15 (5) | 0.008* |
| mCRPC, n (%) | 10 (5) | 5 (2) | 0.05,a |
| Death due to prostate cancer, n (%) | 4 (2) | 3 (1) | 0.4a |
Prognostic value of the candidate genes
Eighty-nine patients experienced BCR with a median time to BCR of 3.33 years (IQR: 1.94–6.00 years) and 47 patients experienced CR with a median time to CR of 5.19 years (IQR 3.40–7.40 years).
The prognostic value of the candidate genes was assessed using the patient classification from RNA in situ hybridization (Fig. 4, Table 2, Fig. S3, and Table S11). Of the candidate genes, only GRIN3A expression was significantly associated with BCR (log-rank, p=0.002); however, not with CR (log-rank, p=0.06). It was also assessed whether the prognostic impact of a cribriform pattern varied by GRIN3A expression; however, significant interactions were not identified for BCR nor CR (likelihood ratio test; p=0.1 and p=0.9). In multivariable analysis, GRIN3A expression was not identified as an independent prognostic factor for BCR (Table 2), although in multivariable analysis excluding a cribriform pattern as a variable, GRIN3A expression demonstrated a trend towards statistical significance (Table S12; p=0.06).
Fig. 4.
Biochemical recurrence-free and clinical-recurrence-free survival in patients treated with radical prostatectomy stratified by GRIN3A expression status in malignant samples. Kaplan-Meier curves for (A) biochemical recurrence-free survival and (B) clinical recurrence-free survival, stratified by GRIN3A expression status in malignant samples. A patient was classified as GRIN3A-positive (+) if at least one malignant tissue core from that patient was scored as “positive,” as determined by RNA in situ hybridization. Abbreviations: BCR, biochemical recurrence; CR, clinical recurrence.
Table 2.
Univariable and multivariable Cox regression analyses assessing the prognostic value of the candidate genes. Patients were classified as “positive” for a particular gene if at least one malignant sample was scored as “positive,” as determined by RNA in situ hybridization. a violated proportional hazards assumption. Abbreviations: CI, confidence interval; events, number of patients experiencing biochemical recurrence; HR, hazard ratio; pN-stage, pathological lymph node stage; PSA, prostate-specific antigen; pT-stage, pathological tumor stage; RP, radical prostatectomy; * statistical significance.
| Biochemical recurrence (420 patients; 89 events) |
||||
|---|---|---|---|---|
| Covariable | Univariable |
Multivariable |
||
| HR (95 % CI) | p | HR (95 % CI) | p | |
| Age at time of RP | ||||
| (continuous) | 1.02 (0.99–1.06) | 0.2 | – | |
| Preoperative PSA group | ||||
| < 10 ng/ml | 1.00 (reference) | 1.00 (reference) | ||
| 10–20 ng/ml | 1.78 (1.12–2.83) | 0.02* | 1.09 (0.66–1.77) | 0.7 |
| > 20 ng/ml | 2.47 (1.37–4.46) | 0.003* | 1.03 (0.53–1.97) | 0.9 |
| Grade Group for RP specimen | ||||
| 1–2 | 1.00 (reference) | 1.00 (reference) | ||
| 3 | 4.78 (2.72–8.42) | < 0.001* | 2.65 (1.43–4.89) | 0.002* |
| 4–5 | 8.00 (4.53–14.2) | < 0.001* | 4.27 (2.20–8.29) | < 0.001* |
| pT-stage | ||||
| pT2 | 1.00 (reference) | 1.00 (reference) | ||
| pT3a | 3.64 (2.08–6.37) | < 0.001* | 2.09 (1.16–3.77) | 0.01* |
| pT3b | 11.1 (5.76–21.3) | < 0.001* | 3.93 (2.10–9.17) | < 0.001* |
| pN-stage | ||||
| pN0 | 1.00 (reference) | 1.00 (reference) | ||
| pN1 | 6.11 (2.64–14.1) | < 0.001* | 4.53 (1.82–11.3) | 0.001* |
| pNX | 0.60 (0.38–0.95) | 0.03* | 1.28 (0.76–2.16) | 0.3 |
| Surgical margin status | ||||
| Negative | 1.00 (reference) | 1.00 (reference) | ||
| Positive | 3.58 (2.15–5.95) | < 0.001* | 3.33 (1.94–5.72) | < 0.001* |
| Cribriform pattern | ||||
| Absent | 1.00 (reference) | 1.00 (reference) | ||
| Present | 3.97 (2.37–6.67) | < 0.001*,a | 2.37 (1.33–4.23) | 0.004* |
| ACSM1 RNA expression status | ||||
| Negative | 1.00 (reference) | – | ||
| Positive | 1.10 (0.66–1.83) | 0.7 | – | |
| GRIN3A RNA expression status | ||||
| Negative | 1.00 (reference) | 1.00 (reference) | ||
| Positive | 1.90 (1.25–2.88) | 0.003* | 1.26 (0.80–1.98) | 0.3 |
| PCDHB2 RNA expression status | ||||
| Negative | 1.00 (reference) | – | ||
| Positive | 0.98 (0.53–1.87) | 1 | – | |
| REG4 RNA expression status | ||||
| Negative | 1.00 (reference) | – | ||
| Positive | 1.46 (0.92–2.34) | 0.1 | – | |
Discussion
In the present study, we nominated genes associated with a cribriform growth pattern by RNA sequencing and differential gene expression in paired samples within each prostate. Four candidates were selected for further analyses by real-time RT-PCR and RNA in situ hybridization, and assessed in relation to clinicopathological data. One of the candidate genes, GRIN3A, was shown to have potential as a diagnostic and prognostic biomarker for prostate cancer, particularly for patients with IDC.
A previous report has demonstrated that GRIN3A is more strongly expressed in prostate cancer compared to other cancer types [34]. Moreover, in line with our results, GRIN3A was more strongly expressed in malignant compared to benign prostate tissue [34]. We show that GRIN3A is higher expressed in cribriform pattern compared to non-cribriform pattern samples, and specifically in IDC when compared to both ICC and non-cribriform GP4, implying that GRIN3A may be particularly useful in suggesting the presence of IDC.
GRIN3A encodes for a subunit of the N-methyl-D-aspartate (NMDA) receptors. These receptors are involved in processes in the central nervous system ([40,41]). In prostate cancer, GRIN3A has been shown to promote proliferation and maintain viability of malignant cells in vitro [34]. Thus, it is not unexpected that GRIN3A is more highly expressed in malignant glands with a cribriform pattern, as both we [42] and others have demonstrated that malignant glands with a cribriform pattern more often have a high proliferation index (as determined by Ki-67 expression), compared to non-cribriform GP4 [43]. However, the difference in GRIN3A expression between IDC and ICC was rather surprising, as we would have expected GRIN3A expression to be similar in these two morphological features if GRIN3A is involved in cell proliferation [34]. Thus, further functional studies assessing the role of GRIN3A expression in prostate cancer ought to be explored.
The morphological features IDC and ICC are now established as markers of poor prognosis for patients treated with RP [[1], [2], [3], [4], [5], [6], [7], [8], [9], [10]]. The detection of cribriform morphology in the diagnostic setting is highly relevant as it has been suggested that it should be an exclusion criterion for active surveillance ([44,45]), and the failure to detect cribriform morphology at the time of diagnosis could lead to undertreatment of patients. Currently, a suspicion of prostate cancer is based on PSA, clinical examination, and magnetic resonance imaging (MRI), and verified if prostate cancer is identified in prostate biopsies [46]. Nevertheless, prostate biopsies may lead to underdiagnosis as only small areas of the tumor are sampled and, specifically, cribriform morphology may be unsampled. Although MRI has shown promise in the detection of cribriform morphology, the sensitivities vary ([47,48]), and as of yet there is no consensus as to whether cribriform morphology is readily detectable by mpMRI [49].We found that GRIN3A was more often expressed in tumors with a cribriform pattern, and not only in malignant glands with a cribriform pattern, suggesting a field effect. Field effects have been demonstrated for other molecular biomarkers found to be associated with a cribriform pattern in prostate cancer, such as SCHLAP1 [15], Ki-67 [43], and PTEN ([22,50]). We speculate that, in patients where a cribriform pattern is not identified on needle biopsy, GRIN3A expression could still be detected due to a field effect and strengthen the suspicion of an unsampled cribriform pattern area, and thus aid in its improved detection in a diagnostic setting.
The distinction between IDC and ICC may be difficult due to similar morphology, and although basal cell staining may be a useful adjunct tool, basal cell staining may be absent in the particular malignant gland with a cribriform pattern that is examined [51]. The differentiation between IDC and ICC by GRIN3A expression is therefore relevant, and despite the poor prognosis of both morphological features, differences between the two entities with regard to prognosis and molecular features have been shown [52]. Nevertheless, many studies have combined IDC and ICC, and this is supported by the latest ISUP guidelines which recommend incorporation of IDC into grading when invasive carcinoma is present [44].
The distinction between IDC and HGPIN may be challenging, but is critical in diagnostic biopsies—particularly in cases where invasive carcinoma is absent—as IDC, and not HGPIN, is an indication for immediate re-biopsy. Currently PTEN and ERG are two markers that may aid in distinguishing these entities. However, the interpretation of PTEN is not always straightforward [53] and ERG may also be identified in a subset of cases with HGPIN ([54,55]). GRIN3A expression was identified in slightly more than 50 % of IDC cases, which is comparable to the frequency of ERG in IDC [56]. Therefore, we speculate that GRIN3A could be used to identify IDC either in conjunction with PTEN and ERG or with PTEN alone. Although not the focus of the current study, it would be of interest to specifically compare GRIN3A expression in IDC and HGPIN.
We found that GRIN3A had prognostic value, although not independently of other prognostic factors, as expected due to its close association with cribriform pattern and other features of poor prognosis. Associations with features such as pT-stage, a post-RP persistent PSA, and metastatic castration-resistant prostate cancer, all support that GRIN3A is a marker of more aggressive prostate cancer.
We found that all four candidate genes were more highly expressed in malignant compared to benign tissue. In particular, ACSM1 and GRIN3A may be relevant as tumor markers, as they were identified in a relatively high percentage of patients and only rarely detected in benign tissue. This is in agreement with other studies, which also have demonstrated that ACSM1 ([35,37]) and GRIN3A [34] are more highly expressed in malignant tissue compared to benign tissue.
ACSM1 is a medium chain Acyl-CoA synthetase family member involved in fatty acid metabolism [57], and is a putative androgen-regulated gene in prostate cancer [36]. In functional studies, ACSM1 is thought to be involved in cell growth and protect against ferroptosis [36]. Furthermore, a study by Guo et al. demonstrated that ACSM1 may be involved in metabolic and extracellular matrix interaction signaling pathways, and may have value as a prognostic biomarker [37].
Since GRIN3A RNA expression was identified as a biomarker associated with a cribriform pattern in prostate cancer, we sought to determine whether GRIN3A protein expression could be detected by immunohistochemistry—as immunohistochemistry is still easier to implement in routine practice. We tested two different antibodies (data not shown); however, the results were inconclusive, warranting further research.
Conclusions
In conclusion, through differential gene expression analysis with a paired sampling design, we identified that ectopically expressed GRIN3A is a RNA-based biomarker for the more frequent presence of a cribriform pattern in prostate cancer, particularly for IDC. Moreover, we demonstrate that GRIN3A is a marker of poor prognosis in primary prostate cancer.
Data availability
The majority of the data that support the findings of this study are available from the supporting information published online, and any additional data is available from the corresponding author upon reasonable request.
CRediT authorship contribution statement
Mari Bogaard: Writing – review & editing, Writing – original draft, Methodology, Investigation, Formal analysis, Data curation. Jonas M. Strømme: Writing – review & editing, Methodology, Investigation, Formal analysis, Data curation. Susanne G. Kidd: Writing – review & editing, Investigation, Data curation. Bjarne Johannessen: Writing – review & editing, Methodology, Formal analysis. Anne C. Bakken: . Ragnhild A. Lothe: Writing – review & editing, Supervision, Resources, Project administration. Karol Axcrona: Writing – review & editing, Writing – original draft, Supervision, Project administration, Methodology, Investigation, Data curation, Conceptualization. Rolf I. Skotheim: Writing – review & editing, Writing – original draft, Supervision, Project administration, Investigation, Funding acquisition, Data curation, Conceptualization. Ulrika Axcrona: Writing – review & editing, Writing – original draft, Supervision, Resources, Project administration, Methodology, Investigation, Funding acquisition, Formal analysis, Data curation, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Funding sources and disclosure of conflicts of interests
The authors have no conflicts of interest to declare. This work was supported by the South-Eastern Norway Regional Health Authority (grant numbers 2016003, 2019016, and 2020063), the Research Council of Norway through its FRIPRO funding scheme (262529/F20 and Toppforsk-250993), and the Norwegian Cancer Society (grant number 208197). The study was granted secure storage and high-performance computation resources from NorStore and University of Oslo's Services for Sensitive Data (NS9013S and p19, respectively).
Acknowledgments
We would like to thank the individuals with prostate cancer and their families for contributing to this study. We would also like to thank Hogne Røed Nilsen for technical assistance with the RNAscope technology, and Mette S. Førsund and Don Trinh for technical assistance with the construction of TMAs.
Footnotes
Supplementary material associated with this article can be found, in the online version, at doi:10.1016/j.neo.2024.101023.
Appendix. Supplementary materials
References
- 1.Iczkowski K.A., Torkko K.C., Kotnis G.R., Wilson R.S., Huang W., Wheeler T.M., et al. Digital quantification of five high-grade prostate cancer patterns, including the cribriform pattern, and their association with adverse outcome. Am. J. Clin. Pathol. 2011;136(1):98–107. doi: 10.1309/AJCPZ7WBU9YXSJPE. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.van Leenders G., Kweldam C.F., Hollemans E., Kümmerlin I.P., Nieboer D., Verhoef E.I., et al. Improved prostate cancer biopsy grading by incorporation of invasive cribriform and intraductal carcinoma in the 2014 Grade Groups. Eur. Urol. 2020;77(2):191–198. doi: 10.1016/j.eururo.2019.07.051. [DOI] [PubMed] [Google Scholar]
- 3.Miura N., Mori K., Mostafaei H., Quhal F., Motlagh R.S., Pradere B., et al. The prognostic impact of intraductal carcinoma of the prostate: a systematic review and meta-analysis. J. Urol. 2020;204(5):909–917. doi: 10.1097/JU.0000000000001290. [DOI] [PubMed] [Google Scholar]
- 4.Hollemans E., Verhoef E.I., Bangma C.H., Rietbergen J., Helleman J., Roobol M.J., et al. Large cribriform growth pattern identifies ISUP grade 2 prostate cancer at high risk for recurrence and metastasis. Mod. Pathol. 2019;32(1):139–146. doi: 10.1038/s41379-018-0157-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hollemans E., Verhoef E.I., Bangma C.H., Rietbergen J., Osanto S., Pelger R.C.M., et al. Cribriform architecture in radical prostatectomies predicts oncological outcome in Gleason score 8 prostate cancer patients. Mod. Pathol. 2021;34(1):184–193. doi: 10.1038/s41379-020-0625-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kweldam C.F., Wildhagen M.F., Steyerberg E.W., Bangma C.H., van der Kwast T.H., van Leenders G.J. Cribriform growth is highly predictive for postoperative metastasis and disease-specific death in Gleason score 7 prostate cancer. Mod. Pathol. 2015;28(3):457–464. doi: 10.1038/modpathol.2014.116. [DOI] [PubMed] [Google Scholar]
- 7.Haffner M.C., Salles D.C., Gao G., Epstein J.I. Gleason pattern 4 with cribriform morphology on biopsy is associated with adverse clinicopathological findings in a prospective radical prostatectomy cohort. Hum. Pathol. 2020;98:74–80. doi: 10.1016/j.humpath.2020.02.004. [DOI] [PubMed] [Google Scholar]
- 8.Dong F., Yang P., Wang C., Wu S., Xiao Y., McDougal W.S., et al. Architectural heterogeneity and cribriform pattern predict adverse clinical outcome for Gleason grade 4 prostatic adenocarcinoma. Am. J. Surg. Pathol. 2013;37(12) doi: 10.1097/PAS.0b013e3182a02169. [DOI] [PubMed] [Google Scholar]
- 9.Sæter T., Vlatkovic L., Waaler G., Servoll E., Nesland J.M., Axcrona K., et al. Intraductal carcinoma of the prostate on diagnostic needle biopsy predicts prostate cancer mortality: a population-based study. Prostate. 2017;77(8):859–865. doi: 10.1002/pros.23326. [DOI] [PubMed] [Google Scholar]
- 10.Russo G.I., Soeterik T., Puche-Sanz I., Broggi G., Lo Giudice A., De Nunzio C., et al. Oncological outcomes of cribriform histology pattern in prostate cancer patients: a systematic review and meta-analysis. Prostate Cancer Prostatic. Dis. 2022 doi: 10.1038/s41391-022-00600-y. [DOI] [PubMed] [Google Scholar]
- 11.Hesterberg A.B., Gordetsky J.B., Hurley P.J. Cribriform prostate cancer: clinical pathologic and molecular considerations. Urology. 2021;155:47–54. doi: 10.1016/j.urology.2021.05.028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Böttcher R., Kweldam C.F., Livingstone J., Lalonde E., Yamaguchi T.N., Huang V., et al. Cribriform and intraductal prostate cancer are associated with increased genomic instability and distinct genomic alterations. BMC Cancer. 2018;18(1):8. doi: 10.1186/s12885-017-3976-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Elfandy H., Armenia J., Pederzoli F., Pullman E., Pertega-Gomes N., Schultz N., et al. Genetic and epigenetic determinants of aggressiveness in cribriform carcinoma of the prostate. Mol. Cancer Res. 2019;17(2):446–456. doi: 10.1158/1541-7786.MCR-18-0440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zhao J., Sun G., Zhu S., Dai J., Chen J., Zhang M., et al. Circulating tumour DNA reveals genetic traits of patients with intraductal carcinoma of the prostate. BJu Int. 2022;129(3):345–355. doi: 10.1111/bju.15530. [DOI] [PubMed] [Google Scholar]
- 15.Chua M.L.K., Lo W., Pintilie M., Murgic J., Lalonde E., Bhandari V., et al. A prostate cancer "Nimbosus": genomic instability and SChLAP1 dysregulation underpin aggression of intraductal and cribriform subpathologies. Eur. Urol. 2017;72(5):665–674. doi: 10.1016/j.eururo.2017.04.034. [DOI] [PubMed] [Google Scholar]
- 16.Kidd S.G., Carm K.T., Bogaard M., Olsen L.G., Bakken A.C., Løvf M., et al. High expression of SCHLAP1 in primary prostate cancer is an independent predictor of biochemical recurrence, despite substantial heterogeneity. Neoplasia. 2021;23(6):634–641. doi: 10.1016/j.neo.2021.05.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Greenland N.Y., Zhang L., Cowan J.E., Carroll P.R., Stohr B.A., Simko J.P. Correlation of a commercial genomic risk classifier with histological patterns in prostate cancer. J. Urol. 2019;202(1):90–95. doi: 10.1097/JU.0000000000000175. [DOI] [PubMed] [Google Scholar]
- 18.Taylor A.S., Morgan T.M., Wallington D.G., Chinnaiyan A.M., Spratt D.E., Mehra R. Correlation between cribriform/intraductal prostatic adenocarcinoma and percent Gleason pattern 4 to a 22-gene genomic classifier. Prostate. 2020;80(2):146–152. doi: 10.1002/pros.23926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Lone Z., Benidir T., Rainey M., Nair M., Davicioni E., Gibb E.A., et al. Transcriptomic features of cribriform and intraductal carcinoma of the prostate. Eur. Urol. Focus. 2022 doi: 10.1016/j.euf.2022.05.005. [DOI] [PubMed] [Google Scholar]
- 20.Parker J.S., Mullins M., Cheang M.C., Leung S., Voduc D., Vickery T., et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 2009;27(8):1160–1167. doi: 10.1200/JCO.2008.18.1370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhao S.G., Chang S.L., Erho N., Yu M., Lehrer J., Alshalalfa M., et al. Associations of luminal and basal subtyping of prostate cancer with prognosis and response to androgen deprivation therapy. JAMa Oncol. 2017;3(12):1663–1672. doi: 10.1001/jamaoncol.2017.0751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ronen S., Abbott D.W., Kravtsov O., Abdelkader A., Xu Y., Banerjee A., et al. PTEN loss and p27 loss differ among morphologic patterns of prostate cancer, including cribriform. Hum. Pathol. 2017;65:85–91. doi: 10.1016/j.humpath.2017.04.024. [DOI] [PubMed] [Google Scholar]
- 23.Shah R.B., Shore K.T., Yoon J., Mendrinos S., McKenney J.K., Tian W. PTEN loss in prostatic adenocarcinoma correlates with specific adverse histologic features (intraductal carcinoma, cribriform Gleason pattern 4 and stromogenic carcinoma) Prostate. 2019;79(11):1267–1273. doi: 10.1002/pros.23831. [DOI] [PubMed] [Google Scholar]
- 24.Løvf M., Zhao S., Axcrona U., Johannessen B., Bakken A.C., Carm K.T., et al. Multifocal primary prostate cancer exhibits high degree of genomic heterogeneity. Eur. Urol. 2019;75(3):498–505. doi: 10.1016/j.eururo.2018.08.009. [DOI] [PubMed] [Google Scholar]
- 25.Carm K.T., Hoff A.M., Bakken A.C., Axcrona U., Axcrona K., Lothe R.A., et al. Interfocal heterogeneity challenges the clinical usefulness of molecular classification of primary prostate cancer. Sci. Rep. 2019;9(1):13579. doi: 10.1038/s41598-019-49964-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kidd S.G., Bogaard M., Carm K.T., Bakken A.C., Maltau A.M.V., Løvf M., et al. In situ expression of ERG protein in the context of tumor heterogeneity identifies prostate cancer patients with inferior prognosis. Mol. Oncol. 2022;16(15):2810–2822. doi: 10.1002/1878-0261.13225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Epstein J.I., Egevad L., Amin M.B., Delahunt B., Srigley J.R., Humphrey PA. The 2014 international society of urological pathology (ISUP) consensus conference on Gleason grading of prostatic carcinoma: definition of grading patterns and proposal for a new grading system. Am. J. Surg. Pathol. 2016;40(2):244–252. doi: 10.1097/PAS.0000000000000530. [DOI] [PubMed] [Google Scholar]
- 28.Montironi R., Cimadamore A., Gasparrini S., Mazzucchelli R., Santoni M., Massari F., et al. Prostate cancer with cribriform morphology: diagnosis, aggressiveness, molecular pathology and possible relationships with intraductal carcinoma. Expert. Rev. AntiCancer Ther. 2018;18(7):685–693. doi: 10.1080/14737140.2018.1469406. [DOI] [PubMed] [Google Scholar]
- 29.Strømme J.M., Johannessen B., Kidd S.G., Bogaard M., Carm K.T., Zhang X., et al. Expressed prognostic biomarkers for primary prostate cancer independent of multifocality and transcriptome heterogeneity. Cancer Gene Ther. 2022;29:1276–1284. doi: 10.1038/s41417-022-00444-7. [DOI] [PubMed] [Google Scholar]
- 30.Love M.I., Huber W., Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550. doi: 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Anders S., Pyl P.T., Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31(2):166–169. doi: 10.1093/bioinformatics/btu638. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Ignatiadis N., Klaus B., Zaugg J.B., Huber W. Data-driven hypothesis weighting increases detection power in genome-scale multiple testing. Nat. Methods. 2016;13(7):577–580. doi: 10.1038/nmeth.3885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Wang F., Flanagan J., Su N., Wang L.C., Bui S., Nielson A., et al. RNAscope: a novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 2012;14(1):22–29. doi: 10.1016/j.jmoldx.2011.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lei H., Wang Z., Jiang D., Liu F., Liu M., Lei X., et al. CRISPR screening identifies CDK12 as a conservative vulnerability of prostate cancer. Cell Death Dis. 2021;12(8):740. doi: 10.1038/s41419-021-04027-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Alinezhad S., Väänänen R.M., Mattsson J., Li Y., Tallgrén T., Tong Ochoa N., et al. Validation of novel biomarkers for prostate cancer progression by the combination of bioinformatics, clinical and functional studies. PLoS One. 2016;11(5) doi: 10.1371/journal.pone.0155901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shrestha R.K., Townley S., Hanson A., Pickering M., Nassar Z.D., Mah C.Y., et al. Abstract PO-036: ACSM1 and ACSM3 regulate fatty acid oxidation in prostate cancer to promote growth and protect against oxidative stress. Cancer Res. 2020;80(23_Supplement) PO-036-PO. [Google Scholar]
- 37.Guo Y., Ren C., Huang W., Yang W., Bao Y. Oncogenic ACSM1 in prostate cancer is through metabolic and extracellular matrix-receptor interaction signaling pathways. Am. J. Cancer Res. 2022;12(4):1824–1842. [PMC free article] [PubMed] [Google Scholar]
- 38.Ohara S., Oue N., Matsubara A., Mita K., Hasegawa Y., Hayashi T., et al. Reg IV is an independent prognostic factor for relapse in patients with clinically localized prostate cancer. Cancer Sci. 2008;99(8):1570–1577. doi: 10.1111/j.1349-7006.2008.00846.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gu Z., Rubin M.A., Yang Y., Deprimo S.E., Zhao H., Horvath S., et al. Reg IV: a promising marker of hormone refractory metastatic prostate cancer. Clin. Cancer Res. 2005;11(6):2237–2243. doi: 10.1158/1078-0432.CCR-04-0356. [DOI] [PubMed] [Google Scholar]
- 40.Pérez-Otaño I., Larsen R.S., Wesseling JF. Emerging roles of GluN3-containing NMDA receptors in the CNS. Nat. Rev. Neurosci. 2016;17(10):623–635. doi: 10.1038/nrn.2016.92. [DOI] [PubMed] [Google Scholar]
- 41.Gene [Internet]. Bethesda, MD: National Library of Medicine (US), National Center for Biotechnology Information; 2004- [updated 2024 Jun 17; cited 2024 Jun 20]. GRIN3A glutamate ionotropic receptor NMDA type subunit 3A [Homo sapiens (human)] - Gene - NCBI. Available from: https://www.ncbi.nlm.nih.gov/gene/116443.
- 42.Bogaard M., Skotheim R.I., Maltau A.V., Kidd S.G., Lothe R.A., Axcrona K., et al. High proliferative cribriform prostate cancer' defines a patient subgroup with an inferior prognosis. Histopathology. 2023;83(6):853–869. doi: 10.1111/his.15012. [DOI] [PubMed] [Google Scholar]
- 43.Fu L., Hwang M., Adeniran A.J., Humphrey PA. Proliferation index of different Gleason pattern 4 histomorphologies and associated pattern 3 adenocarcinoma of the prostate. Hum. Pathol. 2017;70:1–5. doi: 10.1016/j.humpath.2017.06.007. [DOI] [PubMed] [Google Scholar]
- 44.van Leenders G., van der Kwast T.H., Grignon D.J., Evans A.J., Kristiansen G., Kweldam C.F., et al. The 2019 international society of urological pathology (ISUP) consensus conference on grading of prostatic carcinoma. Am. J. Surg. Pathol. 2020;44(8):e87–e99. doi: 10.1097/PAS.0000000000001497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Epstein J.I., Amin M.B., Fine S.W., Algaba F., Aron M., Baydar D.E., et al. The 2019 genitourinary pathology society (GUPS) white paper on contemporary grading of prostate cancer. Arch. Pathol. Lab. Med. 2021;145(4):461–493. doi: 10.5858/arpa.2020-0015-RA. [DOI] [PubMed] [Google Scholar]
- 46.Mottet N., Cornford P., van den Bergh R.C.N., Briers E., Expert Patient Advocate (European Prostate Cancer Coalition/Europa UOMO), De Santis M., et al. EAU–EANM–ESUR–SIOG guidelines on prostate cancer. Edn. presented at the EAU Annual Congress Amsterdam 2022 Arnhem, The Netherlands: EAU Guidelines Office; 2022 [updated 2022 Mar; cited 2022 Sep 9]. Available from: https://uroweb.org/guidelines/prostate-cancer.
- 47.Tonttila P.P., Ahtikoski A., Kuisma M., Pääkkö E., Hirvikoski P., Vaarala M.H. Multiparametric MRI prior to radical prostatectomy identifies intraductal and cribriform growth patterns in prostate cancer. BJu Int. 2019;124(6):992–998. doi: 10.1111/bju.14812. [DOI] [PubMed] [Google Scholar]
- 48.Truong M., Feng C., Hollenberg G., Weinberg E., Messing E.M., Miyamoto H., et al. A comprehensive analysis of cribriform morphology on magnetic resonance imaging/ultrasound fusion biopsy correlated with radical prostatectomy specimens. J. Urol. 2018;199(1):106–113. doi: 10.1016/j.juro.2017.07.037. [DOI] [PubMed] [Google Scholar]
- 49.Norris J.M., Carmona Echeverria L.M., Simpson B.S., Ball R., Freeman A., Kelly D., et al. Conspicuity of cribriform prostate cancer on multiparametric magnetic resonance imaging: the jury is still out. BJu Int. 2021;127(2):169–170. doi: 10.1111/bju.15295. [DOI] [PubMed] [Google Scholar]
- 50.Trock B.J., Fedor H., Gurel B., Jenkins R.B., Knudsen B.S., Fine S.W., et al. PTEN loss and chromosome 8 alterations in Gleason grade 3 prostate cancer cores predicts the presence of un-sampled grade 4 tumor: implications for active surveillance. Mod. Pathol. 2016;29(7):764–771. doi: 10.1038/modpathol.2016.63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Varma M. Intraductal carcinoma of the prostate: a guide for the practicing pathologist. Adv. Anat. Pathol. 2021;28(4):276–287. doi: 10.1097/PAP.0000000000000303. [DOI] [PubMed] [Google Scholar]
- 52.Gordetsky J.B., Schaffer K., Hurley P.J. Current conundrums with cribriform prostate cancer. Histopathology. 2022;80(7):1038–1040. doi: 10.1111/his.14665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Downes M.R., Satturwar S., Trudel D., van der Kwast T.H. Evaluation of ERG and PTEN protein expression in cribriform architecture prostate carcinomas. Pathol. Res. Pract. 2017;213(1):34–38. doi: 10.1016/j.prp.2016.10.007. [DOI] [PubMed] [Google Scholar]
- 54.Cerveira N., Ribeiro F.R., Peixoto A., Costa V., Henrique R., Jerónimo C., et al. TMPRSS2-ERG gene fusion causing ERG overexpression precedes chromosome copy number changes in prostate carcinomas and paired HGPIN lesions. Neoplasia. 2006;8(10):826–832. doi: 10.1593/neo.06427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Shah R.B., Li J., Dhanani N., Mendrinos S. ERG overexpression and multifocality predict prostate cancer in subsequent biopsy for patients with high-grade prostatic intraepithelial neoplasia. Urol. Oncol. 2016;34(3):120. doi: 10.1016/j.urolonc.2015.10.008. e1-7. [DOI] [PubMed] [Google Scholar]
- 56.Lotan T.L., Gumuskaya B., Rahimi H., Hicks J.L., Iwata T., Robinson B.D., et al. Cytoplasmic PTEN protein loss distinguishes intraductal carcinoma of the prostate from high-grade prostatic intraepithelial neoplasia. Mod. Pathol. 2013;26(4):587–603. doi: 10.1038/modpathol.2012.201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Gene [Internet]. Bethesda, MD: National Library of Medicine (US), National Center for Biotechnology Information; 2004, [updated 2024 Jun 17; cited 2024 Jun 20]. ACSM1 acyl-CoA synthetase medium chain family member 1 [Homo sapiens (human)] - Gene - NCBI. Available from: https://www.ncbi.nlm.nih.gov/gene/116285.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The majority of the data that support the findings of this study are available from the supporting information published online, and any additional data is available from the corresponding author upon reasonable request.