Abstract
Managing renal fibrosis is challenging owing to the complex cell signaling redundancy in diseased kidneys. Renal fibrosis involves an immune response dominated by macrophages, which activates myofibroblasts in fibrotic niches. However, macrophages exhibit high heterogeneity, hindering their potential as therapeutic cell targets. Herein, we aimed to eliminate specific macrophage subsets that drive the profibrotic immune response in the kidney both temporally and spatially. We identified the major profibrotic macrophage subset (Fn1+Spp1+Arg1+) in the kidney and then constructed a 12-mer glycopeptide that was designated as bioactivated in vivo assembly PK (BIVA-PK) to deplete these cells. BIVA-PK specifically binds to and is internalized by profibrotic macrophages. By inducing macrophage cell death, BIVA-PK reshaped the renal microenvironment and suppressed profibrotic immune responses. The robust efficacy of BIVA-PK in ameliorating renal fibrosis and preserving kidney function highlights the value of targeting macrophage subsets as a potential therapy for patients with CKD.
Keywords: Renal fibrosis, immune microenvironment, pro-fibrotic macrophage, Bioactivated in vivo assembly-PK (BIVA-PK), cell death
Subject terms: Drug delivery, Mechanisms of disease, Monocytes and macrophages
Introduction
Chronic kidney disease (CKD) is characterized by long-term impairment of kidney structure and a progressive loss of renal function [1]. CKD affects 10% of adults globally [2, 3] and is responsible for 1.2 million deaths annually [4]. Renal fibrosis is the pathological manifestation of CKD and is characterized by myofibroblast activation and extracellular matrix (ECM) accumulation in the renal interstitium [5]. CKD arises from various renal injuries caused by ischemia, infection, toxins, obstruction, and nephritis. The pathological repair of injured renal tubule cells and the formation of scars in the renal interstitium reduce the number of functional nephrons and the glomerular filtration rate and ultimately cause renal failure [6, 7]. Managing CKD is challenging owing to the lack of specific treatments for renal tubule injury and consequent fibrosis [8]. One possible approach to prevention is to provide sufficient blood perfusion to the kidney and avoid infection, obstruction and renal toxins [1].
Monoclonal antibodies targeting key signaling pathways mediating renal fibrosis, including the tumor growth factor-β1 (TGF-β1)/SMAD family member 3 (SMAD3) pathway, have achieved minimal efficacy in clinical trials [5, 8, 9]. Moreover, small molecules targeting single pathways associated with inflammation or chemotaxis, including the nuclear 1 factor-erythroid-2-related factor 2 (NRF2) activator bardoxolone methyl [10], Janus kinase (JAK)1/2 inhibitor baricitinib [11], and C-C motif chemokine receptor (CCR)-2/5 inhibitor PF-04634817 [12], failed in basic research or clinical trials, highlighting the complex compensation of redundant signaling pathways in diseased kidneys. Therefore, novel therapeutic strategies that are effective in reshaping the profibrotic microenvironment of the kidney are needed to halt CKD progression.
Targeting the major cellular components of the fibrotic niche is a strategy for achieving robust efficacy. However, targeting myofibroblasts—the primary drivers of fibrosis—is challenging because their markers are only partially characterized [6, 7, 13–15], which could increase the risk of off-target adverse events. Previous studies have shown that macrophages infiltrate the injured kidney, initiating a profound fibrosis process [16–19]. However, the high heterogeneity of macrophages during the inflammation-to-fibrosis transition [20, 21] may significantly impact the efficacy of targeted therapeutics intended to regulate tissue inflammation and repair [22, 23]. Despite the extensive studies on acute proinflammatory kidney macrophage biology [21], the potential therapeutic value of targeting macrophage subpopulations involved in renal repair and fibrosis remains unknown.
In fact, macrophages characterized by Spp1, Fn1, Gpnmb, and Ctsk expression are considered fibrogenic macrophages [24–26]. These fibrogenic macrophage subsets produce cytokines and growth factors, such as TGF-β, platelet-derived growth factor (PDGF), and galactin-3, to activate myofibroblasts and promote fibrosis [7, 23, 27]. In addition to initiating myofibroblast activation, macrophages undergoing macrophage-myofibroblast transformation (MMT) serve as cellular sources of myofibroblasts [28, 29]. Hence, targeting fibrogenic macrophages in the chronic inflammatory niche might represent an effective therapeutic strategy for attenuating renal fibrosis. However, the challenge lies in temporally and spatially depleting these macrophages to inhibit their profibrotic effect.
In this study, we aimed to suppress the profibrotic immune response by temporally and spatially eliminating specific macrophage subsets that drive renal fibrosis. To this end, we identified the surface markers of profibrotic macrophages in the kidney and constructed the bioactivated in vivo assembly (BIVA) peptide BIVA-PK to specifically recognize and kill these problematic cells. BIVA nanotechnology is an enhanced targeting system that generates an in situ nanofiber-like assembly upon specific enzyme cleavage to improve retention within target cells [30–32]. We modified the BIVA peptide at the N-terminus to target a specific macrophage subset and then conjugated it to LTX-315 [33] (PK), a functional tumor-killing peptide, to trigger macrophage cell death. The efficacy of the resulting BIVA-PK peptide was validated both in vitro and in vivo, and the peptide showed high selectivity for the profibrotic macrophage subset and potent anti-fibrotic activity in a murine model of ischemia/reperfusion-injury (IRI)-induced renal fibrosis.
Results
CD206+ M2-like macrophages are major contributors to renal fibrosis
We established a unilateral ischemia/reperfusion injury (uIRI) mouse model to demonstrate the progression of IRI-induced renal fibrosis and evaluated the transition from injury/inflammation to fibrosis. As evidenced by the Masson’s trichrome staining and fibrosis score (Fig. 1A, B), renal fibrosis was initiated on day 3 (D3), increased on D7, and peaked on D14 post injury. Periodic acid-Schiff (PAS) staining (Fig. 1A) revealed that tubular epithelial cells exhibited signs of injury and apoptosis at D1 post injury, such as vacuolar degeneration, shedding from the basement membrane and the formation of casts in the tubule lumens. At D3, injured tubule cells initiated the regeneration program, and the unhealed tubule cells were gradually replaced by components of the ECM from D7 to D14. Immunohistochemical staining of CD45 in kidney tissues (Fig. 1A, C) revealed that CD45+ immune cells infiltrated on D1 after injury, and they peaked at D7, accompanied by the formation of an interstitial fibrotic niche, while immune cell infiltration decreased at D14 as fibrosis was established.
Fig. 1.
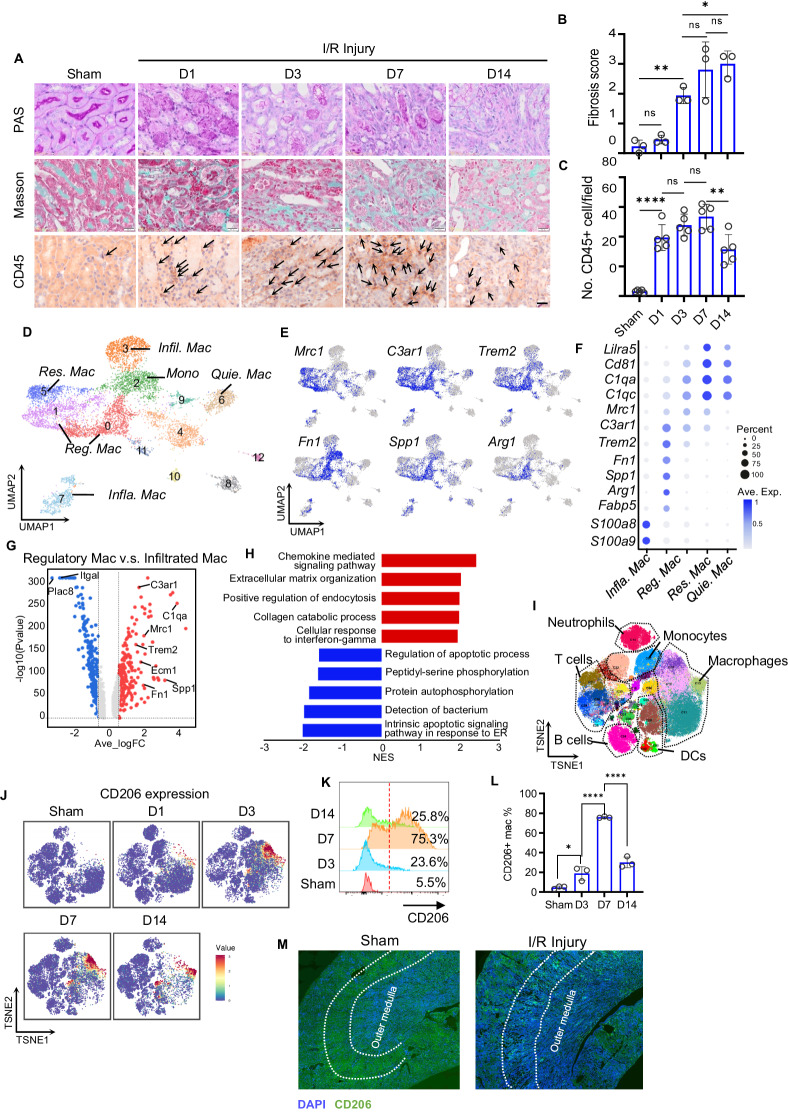
CD206+ macrophages are therapeutic targets for ischemia‒reperfusion injury (IRI)-induced renal fibrosis. A Periodic acid-Schiff staining, Masson’s trichrome staining and CD45 staining of kidney samples from unilateral IRI (uIRI) model mice at different time points. B Quantification of the fibrotic score determined by Masson’s trichrome staining in (A). C Quantification of the number of immune cells by CD45 staining in (A). D UMAP plot showing mononuclear phagocytic cell (MPC) clusters by single-cell transcriptome sequencing of CD45+ cells from IRI kidneys. E Feature plots of profibrotic gene expression in MPCs from IRI kidneys. F Typical marker genes of different macrophage clusters in kidneys with IRI. G Volcano plot displaying the genes differentially expressed between CD206+ and CD206- macrophages in IRI kidneys. H Enriched GOBP terms between CD206+ and CD206- macrophages. I TSNE plot of the clusters of CD45+ cells from IRI kidneys using cytometry by time of flight (CyTOF). J TSNE plot of the distribution of CD206+ macrophages in kidney samples from uIRI model mice at different time points determined by CyTOF. K, L The percentage of CD206+ macrophages in the kidneys of uIRI mice at the indicated times was determined via flow cytometry. M Fluorescence images of CD206+ macrophages in the kidneys of sham and uIRI mice. Scale bar, 20 μm. The data are shown as the means ± standard deviation (SD). *p < 0.05; **p < 0.01; and ***p < 0.001. n = 3–5. For all figures, the data are representative of three independent experiments. uIRI unilateral renal ischemia-reperfusion injury. MPCs mononuclear phagocytic cells
As previously reported [21, 34], mononuclear phagocytic cells (MPCs), including macrophages, monocytes, and other myeloid cells, are the predominant infiltrating immune cells involved in ischemia-reperfusion (I/R)-induced kidney injury. To understand the landscape of MPCs in the inflammation-to-fibrosis process and characterize the profibrotic populations, we performed scRNA-seq on CD45+ immune cells sorted from mouse kidneys on D1, D3, D7, and D14 post-IRI. Based on typical MPC gene expression, 8,838 cells were defined as MPCs (Fig. S1A, Fig. S2). These cells were further clustered into 12 subgroups and annotated as monocytes (C2; Ccr2, and Ly6c), macrophages (C0, C1, C3, C5–C7; Adgre1, and Cx3cr1), and dendritic cells (C4, C8, C12; Itgax, and Cd209a) (Fig. 1D; Fig. S1B; S2; S3). Homeostasis of kidneys without injury was represented by quiescent resident macrophages (C6; C1qa, Cd81, and Cd74), whereas acute inflammation after IRI was characterized by Ly6c+ monocyte expansion (C2; Plac8, and Ly6c), infiltrated macrophages (Infil. Mac; C3; Itgal, and Ace) and proinflammatory macrophages (Infla. Mac; C7; S100a8, S100a9, and Irg1) (Fig. 1D–F; Fig. S4A, B). IRG1 mRNA was not detected in either healthy or fibrotic kidneys but was markedly increased on D1 after injury, indicating that IRG1 is involved in acute inflammation (Fig. S4C). In the repair/fibrosis phase, Arg1+ regulatory macrophages (Reg. Mac; C0 and C1; Arg1, and Trem2) and C1qc+ resident-like macrophages (Res. Mac; C5; C1qa, and Cd81) predominated in the kidneys (Fig. 1D–F). Notably, both Reg. Mac and Res. Mac expressed Mrc1 and highly expressed profibrotic genes, such as Fn1, Spp1, Tgfb1, and C3ar1 (Fig. 1E, F). To understand the function and impact of the Mrc1+ macrophage population on kidney fibrosis, we assessed the DEGs between Mrc1+ (C0 and C1) and Mrc1- (C2) macrophages. Mrc1+ macrophages exhibited significantly upregulated expression of profibrotic genes (Fn1, Spp1, Ecm1, and C3ar1) in the kidneys of IRI model rats (Fig. 1G). These genes were enriched in pathways involved in lymphocyte chemotaxis and extracellular matrix organization after uIRI (Fig. 1H), supporting the correlation between CD206+ macrophages and renal fibrosis.
To confirm CD206 expression on macrophages, we performed mass cytometry by time of flight (CyTOF) on tissue samples from uIRI mice. Thirty-six clusters were identified among the CD45+ immune cells (Fig. 1I) according to the expression of specific cell surface markers (Fig. S5), among which four clusters comprised macrophages (Fig. 1I). Cluster 11 did not express CD206, were present in the homeostatic kidney, and disappeared 1 day after IRI (Fig. S6), indicating that this cluster comprised kidney-resident macrophages. Notably, compared with those in healthy kidneys, C10 and C12 were macrophage subsets that newly emerged on D3 and were retained until D7 in the kidneys of uIRI mice (Fig. S6). However, C10 and C12 disappeared and were replaced by C13 on D14 in the fibrotic kidney (Fig. S6). Hence, C10 and C12 actively participate in the chronic inflammation-to-fibrosis transition, and C13 may play a role in the maintenance of the renal fibrotic microenvironment. CD206 expression was highly enriched in C10 and C13, as well as in a portion of C12, compared with all other analyzed myeloid populations (Fig. 1J), suggesting the potential value of using CD206 to target these fibrogenic macrophages. Next, we detected CD206+ macrophages using immunofluorescence staining and flow cytometry to extend our CyTOF analyses. As shown by the flow cytometry data, the number of CD206+F4/80+ macrophages increased on D3 and peaked on D7 after injury when acute inflammation progressed to chronic inflammation and fibrosis (Fig. 1K, L). On D14 after injury, when fibrosis had developed, the number of CD206+F4/80+ macrophages decreased (Fig. 1K, L). Immunofluorescence staining revealed that CD206+ macrophages were primarily located in the renal outer medullary region (Fig. 1M; Fig. S7), which is prone to IRI and is the primary location of fibrotic lesions. Hence, CD206+ macrophages could be a major contributor to shaping the fibrotic microenvironment in the kidney.
Design and construction of BIVA-PK
To reduce the number of problematic M2-like macrophages and attenuate renal fibrosis, we designed a CD206+ macrophage-targeting peptide based on BIVA nanotechnology. BIVA-based peptides offer the advantages of ligand‒receptor-based targeting and specific enzyme cleavage-induced self-assembly and retention in situ. Here, the peptide was modified with mannose at the N-terminus to ensure specific binding to CD206 on the M2 macrophage surface. Additionally, a cleavable linker sequence was required for cell-specific tailoring and self-assembly of the C-terminal functional peptide. To identify the appropriate sequence, we assessed the expression of the macrophage enzymes cathepsin B, cathepsin S, and legumain in the uIRI kidneys. Western blotting revealed that cathepsin B expression was upregulated on D3–D7 as renal inflammation and fibrosis progressed (Fig. 2A, B), and immunofluorescence staining revealed cathepsin B expression in the renal interstitium on D7 (Fig. 2C). However, cathepsin S expression was not detected in the kidney (Fig. 2A, B; Fig. S8A, B), and legumain expression was primarily found in tubular epithelial cells, with minimal expression in the renal interstitium (Fig. S8C, D). Thus, the cleavable sequence of cathepsin B was selected as the peptide linker to trigger macrophage-specific cleavage and self-assembly in the renal fibrosis milieu. Ruxotemitide (LTX 315) is a peptide derived from bovine lactoferrin that triggers mitochondrial membrane permeabilization regulated by BCL2-associated X (BAX)/BCL2 antagonist/killer 1 (BAK1), inducing cell death [35]. This peptide has previously been used to kill tumor cells in preclinical studies [33, 36, 37]. Here, we leveraged LTX 315 as an effector peptide of BIVA-PK to perturb mitochondrial homeostasis and induce macrophage death.
Fig. 2.
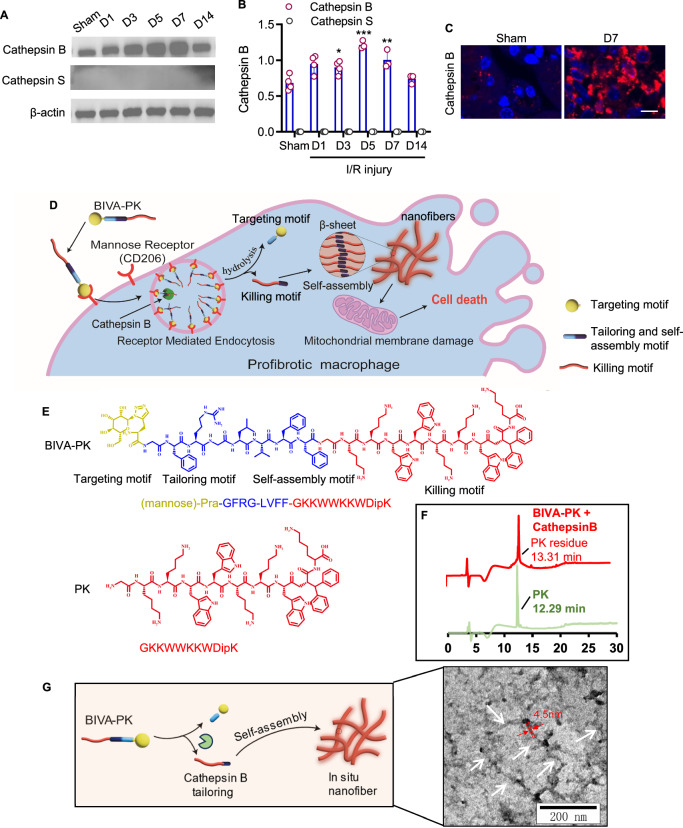
Illustration and chemical structures of bioactivated in vivo assembly PK (BIVA-PK). A, B Western blotting of cathepsin B and cathepsin S expression in the kidneys of uIRI mice at the indicated time points. C Immunofluorescence staining showing cathepsin B expression in the kidneys of uIRI mice. D Schematic illustration showing the modular design of BIVA-PK and the mechanisms for recognizing and killing M2 macrophages. E Chemical structures of BIVA-PK and control PK peptides. F High-performance liquid chromatography profiles of BIVA-PK and PK. G Schematic illustration of cathepsin B-dependent tailoring and in situ assembly of BIVA-PK and the morphology of assembled nanofibers via transmission electron microscopy. Scale bar, 2 μm. The data are shown as the means ± standard deviation (SD). *p < 0.05; **p < 0.01; and ***p < 0.001. n = 3–4. The data are representative of three independent experiments
Herein, the glycopeptide BIVA-PK, which we used to deplete CD206+ macrophages, had a modular design comprising four functional motifs (Fig. 2D): a targeting motif with mannose modification, a tailoring motif with a substrate of cathepsin B, a self-assembly motif with the sequence LVFF, and a killing motif with the LTX-315 peptide. BIVA-PK specifically targeted to CD206 on the profibrotic macrophage surface, triggering CD206 dependent endocytosis (Fig. 2D). In the lysosome, the glycopeptide was cleaved by cathepsin B at the peptide linkage site between F and R. Subsequently, the transmembrane residue RGLVFF-PK translocated to the cytosol and assembled in situ into β-sheet nanofibers. When PK was arranged outside the nanofibers, it caused an increased pertubation of mitochondrial plasma and cell death compared with free PK. The chemical structures of the peptides are presented in Fig. 2E, and the molecular weight of BIVA-PK was 2721 Da (Supplementary Fig. S9). The peptides were characterized as shown in Fig. 2F, G and Supplementary Fig. S9. By co-incubating peptide substrate (100 μM) with cathepsin B (1 μg/mL) in MES/NaOH buffer (pH 5.5) overnight, the BIVA-PK molecules were completely cleaved by the enzyme, resulting in the formation of truncated residues with two different retention times (Fig. 2F). By comparing the residue peaks with those of the PK control (12.29 min), the primary sharp peak at 13.31 min could be identified as the assembly and functional residues (Fig. 2F). Moreover, transmission electron microscopy (TEM) images showing the morphology of the activated assembled nanofibers were obtained (Fig. 2G). After hydrolysis by cathepsin B, the residues of BIVA-PK in situ assembled into nanofibers with a diameter of 4–5 nm (Fig. 2G).
Selective targeting and killing of CD206+ macrophages by BIVA-PK
To detect the specificity of BIVA-PK for polarized macrophages, bone marrow-derived macrophages (BMDMs) were isolated and treated with lipopolysaccharide (LPS) or interleukin-4 (IL-4) to induce M1 or M2 polarization, respectively; untreated BMDMs represented M0 macrophages (Fig. 3A). As shown by the flow cytometry data, BMDMs were polarized to the M1 phenotype (CD86highCD206low) or the M2 phenotype (CD86lowCD206high) (Fig. 3B). Polarized macrophages were incubated with AF647-conjugated BIVA-PK, and binding and internalization were detected using confocal microscopy. BIVA-PK negligibly bound to M1 macrophages, while significant binding and internalization were observed in M2 macrophages (Fig. 3C, D). These results indicate that the specific binding of BIVA-PK to M2 macrophages depends on CD206 expression; the use of mannose as a ligand allows BIVA-PK to target CD206+ macrophages actively. To investigate the binding specificity of BIVA-PK, macrophages and renal tubule epithelial cells (TCMK-1) were incubated with AF647-conjugated BIVA-PK. After incubation for 30 min, AF647-BIVA-PK was detected in the cytosol of M2 macrophages, whereas no BIVA-PK was detected in the TCMK-1 cells, indicating that BIVA-PK selectively recognized M2 macrophages and did not bind to the renal epithelial cells (Fig. 3E, F).
Fig. 3.
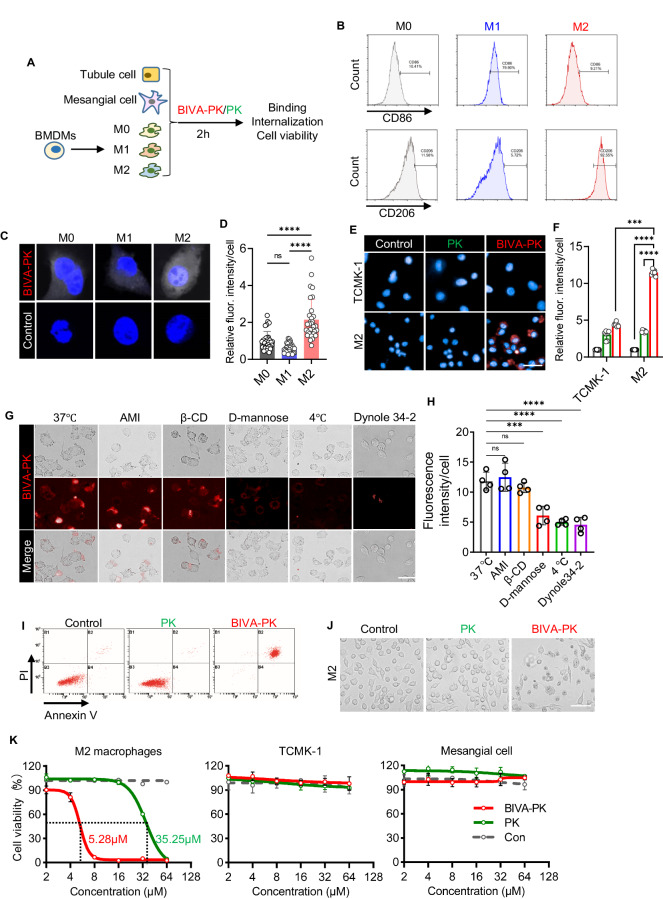
BIVA-PK induces specific internalization and cell death in M2 macrophages. A Bone marrow-derived macrophages (BMDMs) were polarized into M1/M2 macrophages and treated with PK or BIVA-PK at the indicated concentrations to analyze internalization and cell death. B Flow cytometry analysis of BMDMs polarized into M0, M1, and M2 macrophages. C Immunofluorescence images of BMDMs polarized into M0, M1, and M2 macrophages incubated with AF647-conjugated BIVA-PK. D Quantification of the fluorescence in (C). E Confocal images of renal tubule epithelial cells and BMDMs polarized into M2 macrophages incubated with AF647-conjugated PK and BIVA-PK at 37 °C for 2 h and nuclei stained with 4′,6-diamidino-2-phenylindole (DAPI). F Quantification of the induced fluorescence in (E). G BMDMs polarized into M2 macrophages were incubated with AF647-conjugated BIVA-PK in the presence of amiloride (AMI), β-cyclodextrin (β-CD), D-mannose, or dynole 34-2, or at 4 °C. H Quantification of the intracellular fluorescence signals in (G). I Flow cytometry of M2 macrophages co-stained with Annexin V and propidium iodide (PI) after BIVA-PK or PK treatment. J Images of M2 macrophages treated with BIVA-PK or PK. K Dose-dependent cell viability of M2 macrophages, TCMK-1 cells, and mesangial cells after BIVA-PK or PK treatment. Phosphate-buffered saline (PBS) served as a negative control. Scale bar, 20 μm. The data are shown as the means ± standard deviation (SD). *p < 0.05; **p < 0.01; ***p < 0.001; and ****p < 0.0001. n = 3–5. The data are representative of three independent experiments
To investigate whether BIVA-PK endocytosis relies on CD206 internalization, we treated M2 macrophages with mannose, a CD206 ligand, and various endocytosis pathway inhibitors. Mannose competed with BIVA-PK for binding sites and the cellular uptake of BIVA-PK was attenuated at 4 °C, indicating that the internalization of BIVA-PK was an energy-dependent pathway that relied on surface CD206-mediated endocytosis (Fig. 3G, H). Furthermore, treating macrophages with amiloride (AMI, a macropinocytosis inhibitor) or β-cyclodextrin (β-CD, a lipid raft inhibitor) failed to block BIVA-PK uptake, but the uptake of BIVA-PK was dramatically inhibited by the endocytosis inhibitor dynole 34-2 (Fig. 3G, H), consistent with the clathrin-dependent endocytosis pathway of CD206, further validating its significance in BIVA-PK endocytosis.
To confirm that BIVA-PK induces cell death, we examined the viability of M2 macrophages treated with different concentrations of BIVA-PK. BMDMs were isolated and polarized to M2 macrophages via IL-4 treatment for 24 h, followed by exposure to BIVA-PK or the control peptide PK. The staining of Annexin V and PI increased 1 h after BIVA-PK treatment (20 μM) (Fig. 3I), indicating that BIVA-PK induced late-phase cell death in M2 macrophages. The morphological alterations of CD206+ macrophages after BIVA-PK treatment are shown in Fig. 3J, and the expression of cleaved Caspase 3 in CD206+ macrophages was increased by BIVA-PK treatment (Fig. S10). Furthermore, the cell counting kit-8 (CCK-8) assay showed that BIVA-PK induced M2 macrophage-specific cell death in a concentration-dependent manner, with an IC50 of 5.28 μM (Fig. 3K), which was 6.7-fold lower than that of PK (IC50: 35.25 μM). Hence, the in situ nanofiber assembly amplified cytotoxicity and increased the drug use rate of BIVA-PK. These results support the conclusion that the specific targeting of CD206 and the AIR effect of BIVA-PK enhance its macrophage-killing ability. Moreover, TCMK-1 and mesangial cell viability were unaffected by BIVA-PK, further validating the specificity and safety of BIVA-PK (Fig. 3K).
BIVA-PK enhances phagocytosis and mitochondrial injury in M2 macrophages
To investigate the additional biological functions of BIVA-PK in M2 macrophages, we examined the phagocytic and mitochondrial functions of M2 macrophages after low-dose BIVA-PK treatment. To exclude the direct induction of endocytosis by BIVA-PK binding, M2 macrophages were treated with 100 nM BIVA-PK for 2 h and cultured in complete Dulbecco’s modified Eagle’s medium (DMEM) for 4 h before phagocytosis was evaluated. The expression levels of early- and late-phase endosome markers, such as the Ras-related protein Rab-5A (RAB-5A), the Ras-related protein Rab-7a, and lysosomal-associated membrane protein 1 (LAMP1), increased in BIVA-PK-treated M2 macrophages, indicating an increase in macrophage phagocytosis following BIVA-PK treatment (Fig. 4A, B). We then evaluated the phagocytic function of M2 macrophages after BIVA-PK treatment using beads assay (Fig. 4C; Fig. S11). The M2 macrophages showed increased phagocytosis of beads after BIVA-PK treatment; however, there was no significant difference in beads phagocytosis between mannose-treated M2 macrophages and untreated cells. This result suggested that BIVA-PK enhanced the phagocytosis of M2 macrophages in a mannose receptor signaling-independent manner.
Fig. 4.
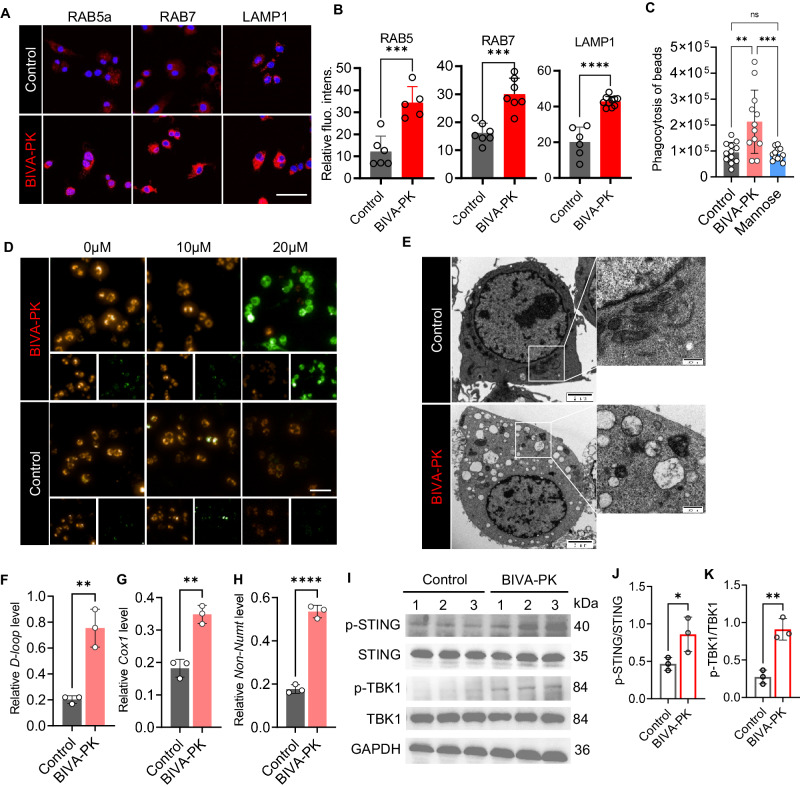
BIVA-PK enhances phagocytosis and mitochondrial injury in M2 macrophages. A Immunofluorescence images of BMDMs stained with anti-Ras-related protein Rab-5A (RAB5A), Ras-related protein Rab-7A (RAB7A), and lysosomal-associated membrane protein 1 (LAMP1) antibodies after BIVA-PK treatment. The intracellular fluorescence is quantified in (B). C After BIVA-PK (100 nM) or mannose (100 nM) treatment, BMDMs were incubated with fluorescence-labeled beads for 2 h, and the phagocytosis of the beads was evaluated using confocal imaging. D Confocal images of M2 macrophages stained with a JC-1 probe to determine the mitochondrial membrane potential after BIVA-PK or vehicle treatment. Scale bar, 20 μm. E Electron microscopy images of M2 macrophages treated with BIVA-PK or vehicle for 6 h. Scale bar, 2 μm. F–H The released mtDNA in the cell cytosol was quantified using qPCR with the primers for mtDNA, such as D-loop, Cox1, and Non-Numt. I–K Western blotting of STING, p-STING, TBK1, and p-TNK1 in BMDMs treated with or without BIVA-PK. The data are shown as the means ± standard deviation (SD). *p < 0.05; **p < 0.01; and ***p < 0.001. n = 3–5. The data are representative of three independent experiments
To investigate the impact of BIVA-PK on the mitochondria, we assessed the mitochondrial membrane potential of M2 macrophages after BIVA-PK treatment using JC-1 probe. In BMDMs, healthy mitochondria showed red aggregate fluorescence (590 nm); after exposure to BIVA-PK for 20 min, there was a concentration-dependent shift from red aggregated fluorescence to green monomer fluorescence (529 nm), indicating that mitochondrial depolarization was induced by BIVA-PK (Fig. 4D). A TEM analysis of the mitochondrial morphology revealed a significant loss of mitochondrial cristae and the accumulation of round fragments in BMDMs treated with 1 μM BIVA-PK, indicating severe mitochondrial damage in macrophages induced by BIVA-PK (Fig. 4E). To investigate whether BIVA-PK treatment enhanced the release of mitochondrial DNA (mtDNA) into the cell cytosol and thus activated the stimulator of interferon genes (STING) pathway, we treated polarized M2 macrophages with BIVA-PK (1 μM) and extracted DNA from the cell cytosol fraction. The released mtDNA in the cell cytosol was quantified using qRT-PCR with the primers for mtDNA (D-loop, Cox1, and Non-Numt; Supplementary Table S3). mtDNA from the whole cell lysate was also extracted and quantified for normalization. As shown in Fig. 4F–H, the release of mtDNA into the cytosol was increased by BIVA-PK; moreover, the phosphorylation of STING and downstream TANK binding kinase 1 (TBK1) were also enhanced after BIVA-PK treatment (Fig. 4I–K), accompanied by a slight increase in type I interferon production (Fig. S12), indicating that the STING pathway was slightly activated by the mtDNA released due to low-dose BIVA-PK treatment.
BIVA-PK is enriched in fibrotic kidneys and safe for major organs
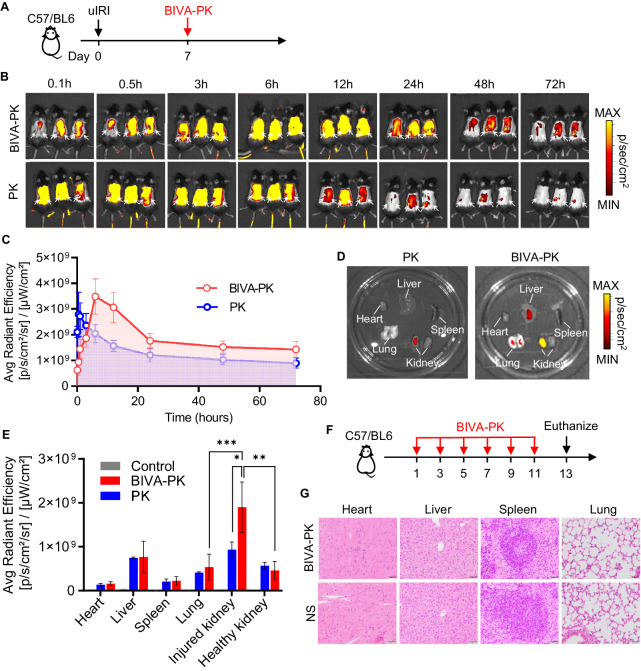
To investigate the biodistribution and organ selectivity of BIVA-PK in vivo, BIVA-PK and PK were conjugated with sulfo-cyanine 7.5 NHS ester. IRI was subsequently induced in the left kidney of C57/BL6 mice with intact contralateral kidneys. The mice were intravenously administered 10 mg/kg of conjugated BIVA-PK or PK on D7 after IRI (Fig. 5A). The distribution of the labeled molecules in the uIRI mice at different time points was monitored via in vivo fluorescence imaging. Although BIVA-PK and PK exhibited systemic distribution within the initial hours postinjection, BIVA-PK demonstrated preferential accumulation in injured kidneys after 3 h (Fig. 5B, C). Furthermore, BIVA-PK persisted in the injured kidney for up to 72 h post-injection, whereas PK was eliminated within 48 h (Fig. 5B, C). The time-dependent area under the curve indicated that BIVA-PK exhibited a higher accumulation dose and a more prolonged elimination half-life than PK (Fig. 5C). This characteristic can be attributed to the significant increase in accumulation and reduction in elimination resulting from in situ nanofiber assembly.
Fig. 5.
In vivo fluorescence imaging of the BIVA-PK distribution. A, B Fluorescence images of uIRI mice at the indicated time points after administration of cyanine 7.5-labeled BIVA-PK or PK. uIRI mice were injected with 10 mg/kg cyanine 7.5-conjugated BIVA-PK or PK at D7. C Quantification of the fluorescence intensity in the injured kidneys of the uIRI mice in (B). D, E Biodistribution of cyanine 7.5-labeled BIVA-PK or PK in the major organs of uIRI mice. uIRI mice were euthanized on day 3 (D3) after BIVA-PK or PK administration, and major organs were harvested for biodistribution evaluation and quantification. F, G Hematoxylin and eosin (HE) staining of the major organs of C57/BL6 mice treated with 50 mg/kg BIVA-PK six times. The data are shown as the means ± standard deviation (SD). *p < 0.05; **p < 0.01; and ***p < 0.001. n = 3–5. The data are representative of three independent experiments
To validate the kidney specificity of BIVA-PK, the mice were euthanized 72 h post-injection, and the major organs were collected for fluorescence analysis. Consistent with the in vivo fluorescence imaging data, the injured kidney (left) of the BIVA-PK-treated group exhibited the strongest fluorescence signals, whereas negligible signals were observed in the heart, liver, spleen, lungs, and healthy kidney of the same mouse (Fig. 5D, E). In the PK group, fluorescence was undetectable in all major organs 72 h after injection, with only weak accumulation observed in the injured kidney (Fig. 5D, E). Moreover, the quantitative analysis of the organ distribution demonstrated the competitive advantage of BIVA-PK over PK in injured kidneys, with a greater than 2-fold increase in accumulation (Fig. 5E).
To evaluate the toxicity and adverse effects of BIVA-PK treatment, healthy C57/BL6 mice were intravenously administered BIVA-PK (50 mg/kg) every other day for a period of 10 days (Fig. 5F). Mice receiving saline injections served as controls. After treatment, the mice exhibited no signs of distress, weight loss, or death. The acute toxicity of BIVA-PK to major organs was further tested using hematoxylin and eosin (H&E) staining (Fig. 5G). No pathological abnormalities were observed in the liver, heart, lungs, or spleen of the BIVA-PK-treated mice compared with those of the control mice. The toxicity of BIVA-PK was further investigated using complete blood count (CBC) and blood chemistry tests (Fig. S13). Compared to those in the healthy group (PBS), there were no obvious changes in the representative biomarkers of liver and kidney function (Fig. S13). In addition, an assessment of white blood cells, red blood cells, and platelets and the percentages of neutrophils, lymphocytes, and monocytes showed no noticeable differences between the BIVA-PK-treated and untreated mice (Fig. S13). These results indicated that BIVA-PK exerted mild toxicity on the liver, kidney, and general lymphocytes of healthy mice.
BIVA-PK improves antifibrotic activity and renal repair after IRI
To determine the optimal dosing window and maximum therapeutic benefit, the uIRI model was used to investigate the efficacy of BIVA-PK in ameliorating the progression of renal fibrosis. In brief, IRI was induced in the left kidney of C57BL/6 mice, and the contralateral kidney was removed 1 day before the mice were euthanized. To investigate whether the treatment time frame could influence the efficacy of BIVA-PK, we intravenously treated uIRI mice with BIVA-PK and the control peptide PK every other day after IR injury. Mice treated with BIVA-PK from D3 to D7 post-IRI were defined as the early-phase treatment group, those treated from D9 to D13 post-IRI were defined as the late-phase treatment group, and those treated from D3 to D13 were defined as the whole-phase treatment group (Fig. 6A).
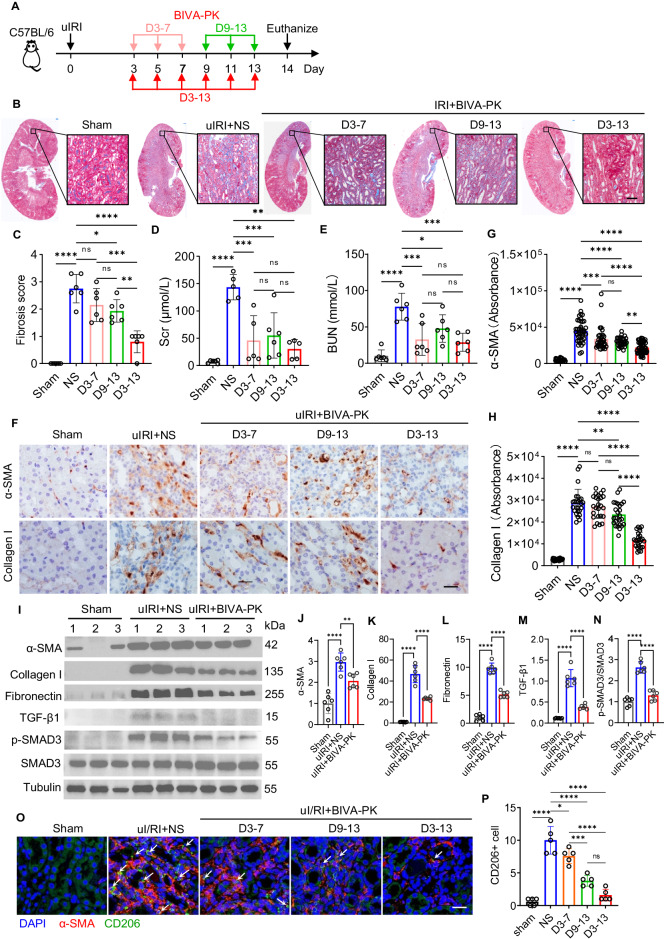
Fig. 6.
In vivo antifibrotic activity of BIVA-PK in renal fibrosis induced by IRI. A uIRI mice were injected with BIVA-PK (10 mg/kg) or normal saline (NS) at the indicated time points. B Masson’s trichrome staining of kidney samples from uIRI mice treated with BIVA-PK or NS. C Fibrosis score calculated from Masson’s trichrome staining of kidney samples from (B). D, E Serum creatinine (Scr) and BUN levels in uIRI mice treated with BIVA-PK or NS were measured by enzyme-linked immunosorbent assay (ELISA). F Immunohistochemical staining of α-smooth muscle actin (α-SMA) and collagen I in the kidneys of uIRI mice treated with BIVA-PK or NS. G, H Quantification of α-SMA and collagen I expression in (F). I Western blotting of the fibrotic markers α-SMA, collagen I, fibronectin, TGF-β1, SMAD3, and p-SMAD3. J–N Quantification of the expression of the fibrotic markers α-SMA, collagen I, fibronectin, TGF-β1, SMAD3, and p-SMAD3 in (I). O Immunofluorescence staining of CD206 and α-SMA in kidneys from uIRI mice treated with BIVA-PK or NS. P Quantification of CD206+ macrophages in (O). The data are shown as the means ± standard deviation (SD), *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. Scale bar = 20 μm. n = 5–6. The data are representative of three independent experiments. BUN blood urea nitrogen, Scr serum creatinine
On D14 post-IR injury, blood samples and IR kidneys were harvested to evaluate renal function and renal fibrosis progression. As shown by Masson’s trichrome staining, ECM deposition was ameliorated after BIVA-PK treatment (Fig. 6B). Compared with the early- and late-phase treatments, the whole-phase treatment exhibited superior inhibition of ECM deposition; however, there was no significant difference between the early- and late-phase treatments according to a quantitative analysis (Fig. 6C). Moreover, the levels of serum creatinine (Scr) and blood urea nitrogen (BUN), two crucial factors in identifying kidney function, were decreased in all three groups of BIVA-PK-treated mice compared with those in the untreated group (Fig. 6D, E). Interestingly, the renal function was similarly improved in the three treatment groups, with the whole-phase treatment group showing slightly better renal function, but the difference was not statistically significant (Fig. 6D, E). This result could be attributed to the robust compensation capacity of the tubule cells. The repaired tubule cells compensate for the function of the damaged tubule cells to a large extent, lowering the Scr and BUN levels in the blood. However, the compensating effect is fragile and susceptible to recurrent injuries; therefore, in the case of a second shock, the renal function of early-phase treated uIRI mice would deteriorate more severely than that of whole-phase-treated mice.
To validate the antifibrotic efficacy of BIVA-PK, we performed immunohistochemical staining of several fibrosis markers in the kidney (Fig. 6F). α-Smooth muscle actin (α-SMA) is a hallmark of myofibroblasts, which are the principal effector cells that promote renal fibrosis [6], and collagen I is one of the major ECM components in the interstitium, impairing normal kidney structure and function [38]. As shown by the immunohistochemical staining, α-SMA was expressed at low levels in healthy kidneys, whereas abundant α-SMA expression was evident in the kidneys of uIRI mice (Fig. 6F, G), indicating the prevalent activation of myofibroblasts. However, the increase in α-SMA expression was abolished by BIVA-PK, and the whole-phase treatment had the greatest effect (Fig. 6F, G). Consistent with the Masson staining results, uIRI triggered the deposition of collagen I in the kidney interstitium (Fig. 6F, H), impairing normal kidney structure and function. BIVA-PK treatment greatly reduced collagen I accumulation, with the most significant effect achieved via whole-phase treatment (Fig. 6F, H). These results indicate that whole-phase BIVA-PK treatment is significantly more effective at mitigating renal fibrosis than early- or late-phase treatment.
Western blotting of the fibrotic markers (α-SMA, collagen I, and fibronectin) further confirmed the suppression of renal fibrosis by BIVA-PK treatment (Fig. 6I–L). We also measured the levels of active TGF-β1 and the downstream phosphorylated SMAD3 (p-SMAD3) molecule using Western blotting (Fig. 6I, M, N). The expression of active TGF-β1 and p-SMAD3 in the BIVA-PK-treated group was clearly decreased compared with that in the untreated group, which is consistent with the reduction in TGF-β1 mRNA expression after BIVA-PK treatment (Fig. 7Q). These results suggested that the TGF-β-SMAD3 signaling pathway may participate in the anti-fibrotic role of BIVA-PK. Several studies have demonstrated the profibrotic effect of IL-11 on toxin- or unilateral ureteral obstruction-induced renal injury and fibrosis [39, 40] and renal ischemia-reperfusion injury, but its effect on I/R injury-induced fibrosis is undetermined. We then measured the expression level of IL-11 in the kidneys of NS-treated or BIVA-PK-treated uIRI mice using Western blotting (Fig. S14). Compared with those in the sham kidneys, the expression levels of IL-11 in the uIRI kidneys were marginally upregulated, and the IL-11 expression levels in the BIVA-PK-treated kidneys were similar to those in the NS-treated kidneys, indicating that the mechanisms underlying the antifibrotic effect of BIVA-PK might be independent of IL-11. Overall, the activation of fibroblasts and deposition of ECM were effectively attenuated by BIVA-PK treatment, with a superior effect achieved via whole-phase treatment, resulting in therapeutic benefits for renal fibrosis.
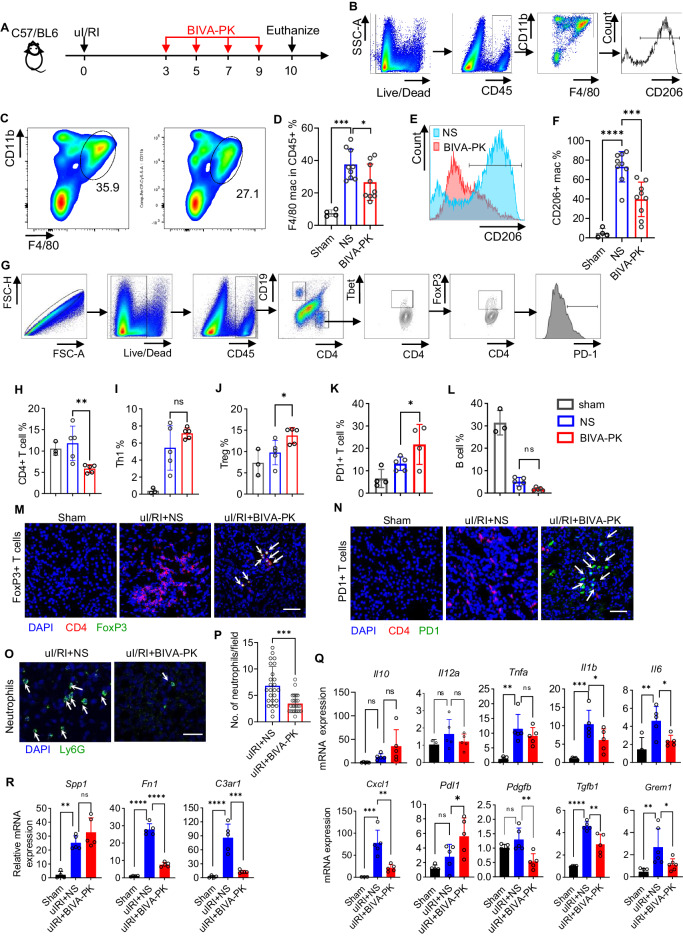
Fig. 7.
The immune microenvironment was reshaped by BIVA-PK in IRI kidneys. A uIRI mice were injected with 10 mg/kg BIVA-PK or NS on D3, D5, D7, and D9 and euthanized on D10 to harvest kidneys for flow cytometry. B Flow cytometry gating strategy for CD206+ macrophages. C, D Percentage of F4/80+ macrophages and (E, F) CD206+ macrophages in the parental population, in the kidneys of uIRI mice. G Gating strategy for T and B-cell populations. H–L Percentage of the indicated immune cells in the parental population from the kidneys of uIRI mice. M Immunofluorescence images of FoxP3+ T cells and (N) PD1+ T cells in the kidneys of uIRI mice treated with BIVA-PK or NS. O Immunofluorescence images of Ly6G+ neutrophils in the kidneys of uIRI mice treated with BIVA-PK or NS. P Calculation of Ly6G+ neutrophils in (O). Q Il10, Il12a, Tnfa, Il1b, Il6, Cxcl1, Pdcd1, Pdl1, Pdgfb, and Tgfb1 transcript expression (from whole kidney) on day 10. R Spp1, Fn1, and C3ar1 transcript expression (from isolated F4/80+ macrophages in the kidney) on D10. The data are shown as the means ± standard deviation (SD). *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001. Scale bar = 20 μm. n = 3–9. The data are representative of three independent experiments
To confirm that the antifibrotic effect of BIVA-PK was due to the specific elimination of CD206+ macrophages, we assessed tissue infiltration by CD206+ macrophages in the fibrotic kidney. Immunofluorescence staining showed that CD206+ macrophage accumulation in fibrotic kidneys was decreased by BIVA-PK, with whole-phase treatment demonstrating the greatest reduction (Fig. 6O, P), suggesting that the suppression of CD206+ macrophages is a potential mechanism of BIVA-PK efficacy.
BIVA-PK remodels the immune microenvironment of the AKI-to-CKD transition
To confirm that the excellent antifibrotic effect of BIVA-PK was due to the specific elimination of CD206+ macrophages, we treated uIRI mice with BIVA-PK or NS on D3, D5, D7, and D9 after injury and harvested the injured kidneys on D10 to quantify CD206+ macrophages in the fibrotic kidney (Fig. 7A, B). After BIVA-PK treatment, the proportion of F4/80+ macrophages among CD45+ immune cells decreased from 37.56% to 26.57% (p = 0.0246) (Fig. 7C, D), and the proportion of CD206+ macrophages among F4/80+ macrophages decreased from 73.1% to 39.6% (p = 0.0006) (Fig. 7E, F), indicating the effective targeting and elimination of M2-like macrophages by BIVA-PK in the fibrotic kidneys.
T cells and B cells are important cellular components of the kidney fibrogenic niche and are essential players in inflammation, wound healing and scar formation after I/R injury [7, 38]. To examine how BIVA-PK affects the immune microenvironment of IR-injured kidneys, we evaluated the major T-cell and B-cell populations using flow cytometry (Fig. 7G). The proportion of CD4+ T cells in the CD45+ cell population was significantly lower in BIVA-PK-treated mice than in NS-treated control mice (11.80% vs. 5.85%, p = 0.0059; Fig. 7H). The proportion of Type 1 T helper cells (Th1, Tbet+) among the CD4+ T cells was unchanged by BIVA-PK treatment, although it increased in the uIRI model (0.15% vs. 5.42%, p = 0.0059; Fig. 7I). Hence, BIVA-PK did not exert anti-inflammatory effects by downregulating proinflammatory Th1 cells. Notably, the percentages of regulatory T cells (Tregs, FoxP3+) and PD1+ T cells were significantly increased by BIVA-PK (9.73% vs. 13.74%, p = 0.0331, and 13.14% vs. 21.73%, p = 0.0485, respectively; Fig. 7J, K). This observation suggested that BIVA-PK significantly activated the immunosuppressive response and aided in attenuating chronic inflammation, thus ameliorating renal fibrosis progression. Similarly, the proportion of B cells in CD45+ cells in the kidney was lower in the uIRI group than in the healthy group (31.4% vs. 5.08%, p < 0.0001) and remained unchanged by BIVA-PK treatment (Fig. 7L). This result may have occurred because B-cell infiltration and activation are not markedly involved in the immune response after IR injury, consistent with previous evidence showing that myeloid cells are the predominant immune cells involved in fibrosis after IR injury.
Immunofluorescence staining confirmed increased FoxP3+ Tregs and PD1+ T cells in the fibrotic lesions of the outer medullary region (Fig. 7M, N). We then investigated whether BIVA-PK affects neutrophils—another major proinflammatory population of myeloid cells. We examined Ly6G+ neutrophils in the diseased kidneys in the absence or presence of BIVA-PK and found that Ly6G+ neutrophils were relatively absent in healthy kidneys but strongly infiltrated fibrotic kidneys (Fig. 7O, P). However, BIVA-PK significantly reduced their infiltration into the injured kidney (Fig. 7O, P), suggesting that BIVA-PK exerts protective effects in part by mitigating neutrophil-mediated chronic inflammation.
To determine whether BIVA-PK injection during the early phase is sufficient for treatment, we treated uIRI mice with BIVA-PK at D3, D5 and D7 and harvested the kidneys at D8 and D10 to determine the percentages of CD206+ macrophages and other immune cell populations (Fig. S15A). The percentage of CD206+ macrophages among F4/80+ cells decreased from 58.17% in NS-treated mice to 23.5% in BIVA-PK-treated mice at D8 (p = 0.0119); however, the percentage of CD206+ macrophages increased to 40.7% at D10 (p = 0.0489) (Fig. S15B), indicating the continuous infiltration and differentiation of CD206+ macrophages in the kidney. The proportions of either CD4+ T cells or B cells among CD45+ cells remained unchanged in early-phase BIVA-PK-treated mice compared with those in NS-treated mice (Fig. S15C, D). Moreover, the proportions of Th1 cells and Treg cells among CD4+ T cells were also unchanged after BIVA-PK treatment (Fig. S15E, F). Notably, the percentage of PD1+ T cells among CD4+ T cells significantly increased at D8 in early-phase BIVA-PK-treated mice compared with that in untreated mice (20.17% vs. 11.12%, p = 0.0139), but the percentage decreased to 13.77% at D10 (p = 0.0256) (Fig. S15G), indicating the loss of the immunosuppressive microenvironment after early-phase treatment was completed. These results demonstrated the continuous infiltration of CD206+ macrophages into the kidney, suggesting that early-phase BIVA-PK treatment might not be sufficient to maintain the antifibrotic immune microenvironment in this organ.
We also examined the renal expression of inflammatory and fibrotic transcripts to assess whether the renal microenvironment was reshaped by BIVA-PK treatment. The mRNA expression of inflammatory cytokines, such as Il1b and Il6, and the neutrophil chemotactic chemokine Cxcl1 was inhibited in the BIVA-PK-treated kidney; moreover, the mRNA level of Pdl1 increased, indicating a shift to an immunosuppressive environment by BIVA-PK (Fig. 7Q). BIVA-PK also reduced the mRNA expression of profibrotic factors, such as Pdgfb, Tgfb1 and Grem1, confirming its suppressive effect on renal fibrosis in uIRI kidneys (Fig. 7Q). We then sorted F4/80+ macrophages from uIRI kidneys and assessed the mRNA expression of the profibrotic genes Spp1, Fn1, and C3ar1 (Fig. 7R). Fn1 and C3ar1 mRNA expression in macrophages increased in uIRI mice but decreased in BIVA-PK-treated mice. Notably, the Spp1 mRNA levels in the BIVA-PK-treated mice were greater than those in the uIRI mice, indicating the complexity and plasticity of macrophages. These results suggest that BIVA-PK reshapes the local microenvironment of the kidney by promoting an immunosuppressive response, inhibiting inflammatory and fibrotic factors, disrupting the fibrotic niche, and shifting the environment from profibrotic to anti-fibrotic.
Discussion
Anti-renal fibrosis treatment has attracted increasing attention in recent years; however, until recently, most therapies have remained in the experimental research stage, and no specific antifibrotic drug has been approved for the treatment of kidney fibrosis. Although significant progress has been made in treating CKD patients using renin‐angiotensin‐aldosterone system inhibitors, novel sodium glucose cotransporter 2 inhibitors, and non-steroidal mineralocorticoid receptor antagonists, all these treatments have indirect antifibrotic effects by affecting blood pressure, glomerular perfusion, tubule epithelial metabolism or inflammation; none of these medications are specific antifibrotic drugs [41]. To date, most clinical trials targeting the TGF-β signaling pathway have shown unsatisfactory results [42]. Several clinical trials using other antifibrotic strategies in kidney diseases, such as modulating Nrf2-mediated renal oxidative stress, endothelin receptor or connective tissue growth factor, have not produced highly promising results that would justify clinical approval until now [41]. The possible reasons for the unsatisfactory results of anti-fibrotic drugs in clinical trials could be the complex microenvironment composed of various cell types and redundant signaling pathways in the fibrotic kidney and the heterogeneity of CKD patients with recurrent injuries and comorbidities. Therefore, drugs targeting a single signaling pathway may be insufficient to halt fibrosis; nevertheless, therapies targeting immune cells are more effective at reshaping the profibrotic microenvironment and preventing redundant molecular mechanisms. In this study, we demonstrated that Fn1+Spp1+Mrc1+ macrophages are major cellular contributors to the renal profibrotic microenvironment. Then, we leveraged BIVA technology to target these profibrotic macrophages and used functional peptides to induce macrophage cell death (Fig. 8). We showed that the selective depletion of Fn1+Spp1+Mrc1+ macrophages by the BIVA-PK peptide shifted the profibrotic microenvironment and ameliorated IRI-induced renal fibrosis, suggesting that macrophage-targeted BIVA-PK might show superiority and benefit CKD patients in clinical practice. More importantly, renal fibrosis arising from many heterogeneous diseases, including diabetes, transplant and nephritis, commonly manifests as increased CD206+ macrophages in the fibrotic kidney [29, 43], indicating that BIVA-PK could be used in various disease conditions to halt fibrosis progression.
Fig. 8.
Schematic diagram showing the mechanisms by which BIVA-PK ameliorates kidney fibrosis by inducing profibrotic macrophage cell death. Fn1+Spp1+Mrc1+ macrophages are major cellular contributors to the renal profibrotic microenvironment. The bioactivated in vivo assembly (BIVA) peptide BIVA-PK was constructed to recognize and kill these problematic cells specifically. Mechanistically, BIVA-PK targets CD206 expressed on the profibrotic macrophage surface, triggering CD206-dependent endocytosis. In the lysosome, BIVA-PK is cleaved by cathepsin B at the peptide linkage site. Subsequently, the functional residue RGLVFF-PK translocates to the cytosol and assembles in situ into β-sheet nanofibers, causing enhanced perturbation of mitochondrial plasma and cell death. The selective depletion of Fn1+Spp1+Mrc1+ macrophages by the BIVA-PK peptide reshaped the profibrotic microenvironment and ameliorated IRI-induced renal fibrosis
According to our study, late-phase treatment had no effect on renal fibrosis or kidney function, indicating that BIVA-PK treatment should be initiated before inflammation is resolved and fibrosis is fully established. Given that CKD patients usually undergo multiple injuries as the disease progresses in comparison to a single injury in uIRI mice, further clinical studies are required to determine the optimal time point to initiate BIVA-PK treatment in CKD patients. However, biomarkers identified in patients with fibrotic kidney injury [44], such as SPP1, which is produced by profibrotic macrophages [45], and MCP-1, which drives macrophage chemotaxis to the injured kidney [46], can serve as biomarkers for identifying potential populations of patients who may benefit from macrophage depletion therapy. As shown in a previous study [29], patients who underwent kidney transplantation also exhibited profibrotic macrophage infiltration, indicating that the BIVA-PK strategy could also be used for the treatment of transplanted kidney fibrosis.
Unlike myofibroblasts, pericytes, and other fibrosis contributors, macrophages have distinct surface markers, facilitating the development of targeting approaches [47]. We showed that the mannose receptor (CD206) is specifically expressed in macrophages in the fibrotic kidney and that their capacity for endocytosis is an optimal target for delivering drugs to profibrotic macrophages [48]. In this regard, we modified BIVA-PK with mannose at the N-terminus, allowing its specific recognition and internalization by CD206+ macrophages without binding to other renal cells. Notably, 72 h after intravenous injection, BIVA-PK primarily accumulated in the injured kidney rather than in other major organs or healthy kidneys. The primary explanation for this finding is that CD206+ macrophages specifically infiltrate injured kidneys, and when BIVA-PK binds to these macrophages, it is trapped in the local microenvironment. To enhance the cellular selectivity of the M2 macrophage depletion approach, we used BIVA nanotechnology to achieve a specific intracellular assembly-induced retention (AIR) effect [30, 49]. Herein, a cleavable sequence of cathepsin B was incorporated into the BIVA-PK peptide as a cleavable linker to self-assemble the functional peptide. Cathepsin B is a highly conserved lysosomal cysteine protease that is essential for cleaving intracellular proteins [50, 51]. Because of the upregulated expression of cathepsin B in the interstitium of fibrotic kidneys, the retention of BIVA-PK was greater in fibrotic kidneys than in healthy kidneys, confirming the microenvironment-specific retention of BIVA-PK.
In addition to the depletion of CD206+ macrophages, BIVA-PK altered macrophage behavior at a low-dose, promoting macrophage phagocytosis and inducing mitochondrial injury. The observed decrease in renal fibrosis in the early- and whole-phase-treated mice might be partially explained by the increased phagocytic activity of CD206+ macrophages, which facilitated the clearance of apoptotic tubule cells during acute injury and inhibited the second wave of inflammation [52, 53]. Moreover, BIVA-PK-enhanced macrophage phagocytosis helped reduce the ECM in chronic fibrotic lesions and remodel the tissue structure, contributing to the efficacy of late- and whole-phase treatments. Our findings are consistent with a previous study on liver fibrosis in which cytotherapy, comprising the transfer of M1-like macrophages, enhanced the clearance of the matrix, thereby ameliorating fibrogenesis [54]. By administering BIVA-PK at various stages of the AKI-to-CKD transition, the dynamic function of CD206+ macrophages in promoting inflammation and fibrosis after kidney injury was observed, providing insight into the immune cell regulatory mechanisms involved in fibrosis progression.
Peptide-based antifibrotic therapeutics have considerable advantages over antibody-based therapies. For example, their small molecular size before self-assembly facilitates rapid penetration from the circulation into the target tissue, and BIVA-PK enters fibrotic kidneys within 30 min. The half-life of BIVA-PK is significantly enhanced by cathepsin B-induced self-assembly, facilitating sustained treatment efficacy. However, owing to the prevalence of CD206+ macrophages in other organs, BIVA-PK, which has a high affinity for the mannose receptor, may cause concerns regarding non-specific toxicity. In vivo data showed that systemic administration of BIVA-PK caused only mild retention and morphological changes in other major organs and healthy kidneys, suggesting that it is a safe remedy for treating renal fibrosis.
This study had several limitations. First, the specific mechanism by which macrophage death influences the immune response and kidney fibrosis is unclear and requires further elucidation. Second, in addition to inducing cell death, new functional modules in the macrophage targeting system might be exploited to fine-tune macrophage phenotype and function, facilitating inflammation resolution and fibrosis blockade [49]. Considering the species differences between mice and humans, mice with humanized immune systems or large animal models (e.g., nonhuman primates) should be used to reproduce human CKD progression in future preclinical studies. Moreover, because CKD patients usually have complex conditions with recurrent renal injuries and multiple comorbidities, it is necessary to stratify CKD patients who might benefit from this therapy based on proper biomarker profiles of profibrotic macrophages when clinical trials are designed in the future.
In summary, in this study, we demonstrated that Fn1+Spp1+Mrc1+ macrophages are major contributors to the renal profibrotic microenvironment, and we identified a 12-mer peptide, BIVA-PK, that perturbs the macrophage membrane and mitochondrial homeostasis and induces cell death. We showed the robust efficacy of BIVA-PK in blocking renal fibrosis and preserving renal function by reshaping the immune microenvironment. In addition, BIVA-PK offers advantages, including specific kidney distribution, prolonged in situ retention and reliable safety, providing a potential therapeutic approach for treating CKD patients with renal fibrosis.
Materials and methods
Macrophage culture and polarization
BMDMs were isolated from C57/BL6 mice as described previously [34], and the cells were cultured in vitro in DMEM supplemented with 10% FBS and 20 ng/mL macrophage colony-stimulating factor (M-CSF, PeproTech, #315-02) for 7 days.
To polarize the macrophages into M0, M1, or M2 cells, the cells were treated with phosphate-buffered saline (PBS), lipopolysaccharide (LPS, 10 ng/mL, Sigma Aldrich, # L2630), interferon-gamma (IFN-γ, 20 ng/mL, PeproTech, # 315-05) or interleukin-4 (IL-4, 20 ng/mL, PeproTech, #214-14) for 12 h. Macrophage polarization was assessed through the expression of CD86 and CD206 using flow cytometry.
Internalization assay
The BMDMs were seeded in a 96-well black plate at 5,000 cells per well and cultured in complete DMEM supplemented with IL-4 for 12 h to induce M2 polarization. BIVA-PK was labeled with fluorescent Alexa Fluor 647 (AF647, Thermo Fisher Scientific, # A20006). Subsequently, the cells were incubated with AF647-labeled BIVA-PK at 37 °C or 4 °C for 40 min. Amiloride hydrochloride (2 mM, MedChemExpress, # HY-B0285A), β-cyclodextrin (5 mM, MedChemExpress, # HY-107201), sucrose (450 nM, MedChemExpress, # HY-107687), D (+)-mannose (450 mM, Sigma Aldrich, # M6020) or dynole 34-2 (5 μM, MedChemExpress, # HY-107545) were added to the culture medium to treat the cells. The macrophages were then fixed using 4% paraformaldehyde and stained with 4’,6-diamidino-2-phenylindole (DAPI, Abcam, #ab228549). Lastly, the cells were imaged under a confocal microscope (FV3000, Olympus, Japan).
Cytotoxicity assay
The mouse kidney epithelial cell line TCMK-1 (# CCL-139) and the mesangial cell line SV40 MES 13 (# CRL-1927) were purchased from ATCC (American Type Culture Collection, https://www.atcc.org). The cytotoxicity of BIVA-PK to mouse tubule epithelial cells, mesangial cells, and M0, M1, or M2 macrophages was evaluated using a Cell Counting Kit-8 (CCK-8, Sigma Aldrich, #91963). Cells were seeded into 96-well plates at a density of 2 × 104 cells per well and cultured for 14 h. The cells were then treated with serially diluted BIVA-PK (0, 2, 4, 8, 16, 32, and 64 μM) for 6 h. Then, CCK-8 solution (10 μL per well) was added and incubated for 2 h. The absorbance of each well was measured at 450 nm using a Spark Multimode Microplate Reader (Tecan). The cell viability (%) was calculated as follows: (A450 of BIVA-PK treatment/A450 of PBS treatment). The PK treatment served as controls.
uIRI model
The animal experiments complied with the Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines and were conducted according to the Guide for the Care and Use of Laboratory Animals approved by the Institutional Animal Care and Use Committee of the Chinese PLA General Hospital. Male C57BL/6J mice aged 6–8 weeks were obtained from Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China) and housed under specific pathogen-free (SPF) conditions. The mice were used to establish uIRI models, which allow renal fibrosis to progress after ischemia-reperfusion injury. In brief, the mice were anesthetized, and their left renal pedicles were clamped for 35 min, followed by clamp release to allow reperfusion. The contralateral kidney was left intact and functional. The mice were kept on a heating pad at 37 °C during the procedures and were provided with free access to food and water.
In vivo imaging
BIVA-PK was labeled with sulfo-cyanine 7.5 NHS ester (Lumiprobe, #25320) according to the manufacturer’s instructions. In vivo fluorescence imaging of uIRI mice was performed to measure the biodistribution of BIVA-PK and PK. In brief, uIRI mice were intravenously injected with 10 mg/kg sulfo-cyanine 7.5-BIVA-PK in one dose, anesthetized with isoflurane and imaged using an IVIS Lumina III imager (PerkinElmer Chemagen Technologie GmbH) with identical settings (2-s exposure, f/stop ¼ 2) at the indicated time points.
In vivo treatment of uIRI mice
The mice in each group were treated with BIVA-PK every other day, and NS served as the control. All the animal treatments started at a final administered volume of 200 μL. Blood samples (600–700 μL per mouse) and kidney samples were harvested for further analysis on D14. For the Scr and BUN measurement in the blood, the uIRI mice were anesthetized, and the right kidneys were removed 1 day before blood samples were collected.
scRNA-Seq by 10× genomics and data analysis
Single cells isolated from the IRI kidneys at the indicated time points were stained with Zombie NIR viability dye (BioLegend, # 423106); CD45- and CD 45+ live cells were sorted using a BD FACS AriaTM cell sorter. FACS-sorted cells were resuspended to a final cell concentration of 700–1200 cells/μL with >80% viability as determined by a LUNA-FL™ Automated Fluorescence Cell Counter (Logos Biosystems). The 3′ sequencing library was constructed using the 10× Genomics Chromium Next GEM Single Cell 3ʹ Reagent Kit v3.1 (Dual Index) according to the manufacturer’s protocols. In brief, the isolated single cells were encapsulated and barcoded using a Chromium Controller (10× Genomics, Pleasanton, CA). After reverse transcription and emulsion breaking, the product cDNA was purified and amplified, followed by sample index attachment. Libraries were sequenced on an Illumina NovaSeq 6000 platform using 150 paired-end reads.
We demultiplexed cell barcodes using the Cell Ranger software pipeline (version 5.0.0) provided by 10× Genomics, mapped reads to genomes and transcriptomes using the STAR aligner, and down-sampled reads as necessary to generate normalized aggregate data across samples to generate a matrix of gene counts versus cells. The unique molecular identifier (UMI) count matrix was processed using the R package Seurat (version 3.1.1) [55]. We filtered out cells with gene counts of less than 200 and greater than 10,000 and with a UMI of less than 1000 to remove low-quality cells and possible multiple captures. We also eliminated low-quality cells with greater than 20% mitochondrial gene expression. A total of 36,353 single cells were included in the downstream analysis after applying the quality control criteria. The library size was normalized using the NormalizeData function in Seurat to obtain normalized counts. Specifically, the global scaling normalization method “LogNormalize” normalizes gene expression measurements for each cell to total expression, multiplies by a scaling factor (default 10,000), and then log-transforms the result.
The top variable genes in single cells were determined using the method described by Macosko et al. [56]. The variable genes were screened using the FindVariableGenes function in Seurat (mean.function = FastExpMean, dispersion.function = FastLogVMR). Principal component analysis (PCA) was used to reduce dimensionality by using the RunPCA function in Seurat. Graph-based clustering was performed using the FindClusters function in Seurat based on the gene expression profiles of the cells. MPC populations were clustered at a resolution of 0.3. The cells were visualized using the two-dimensional Unified Surface Approximation and Projection (UMAP) algorithm with the RunUMAP function in Seurat. We used the FindAllMarkers function in Seurat (test.use = presto) to identify marker genes for each cluster and manually annotated a given cluster based on the lists of marker genes in Table S1 and previously published articles. Gene expression was visualized as scatter plots, dot plots and volcano plots using the FeaturePlot() and VlnPlot() functions in the Seurat package. Differentially expressed genes (DEGs) were identified using the FindMarkers function in Seurat (test.use = presto). P < 0.01 and |log2foldchange| > 0.58 were used as thresholds for significant differential expression. GO enrichment and KEGG pathway enrichment analyses of DEGs were performed using hypergeometric distribution-based R software.
Mass cytometry (CyTOF) and data analysis
Kidney single-cell solutions were prepared as described previously. For each sample, a total of 1–3 × 106 cells were incubated with cisplatin (250 nM, Fluidigm) for 5 min on ice to identify dead cells and then stained with cell-surface antibodies for 30 min at 4 °C followed by Fc blocking for 20 min on ice. The cells were washed twice in CSM (1 × PBS + 0.5% BSA) and fixed and stained overnight at 4 °C in a Cell ID Intercalator Ir (Fluidigm). A Foxp3/Transcription Factor Staining Buffer Set (eBioscience, # 00-5523-00) was used to permeabilize the cells, followed by intracellular antibody staining for 30 min at 4 °C. After being washed in deionized water, the cells were diluted in an EQ™ Four Element Calibration Beads solution (Fluidigm). The data were acquired using a Helios mass cytometer. The antibodies used for these experiments are listed in Table S2.
Mass cytometry data were normalized using EQ Four Element Calibration Beads. The FCS data were de-barcoded using a doublet filtering scheme with mass-tagged barcodes, which allowed for the differentiation of individual samples. The following gating strategy was used: cells, live cells, singlets, and CD45+ immune cells. An X-shift algorithm was used to obtain accurate immune cell clusters and annotations for all the samples. To characterize the immune cell clusters specifically, separate t-distributed stochastic neighbor embedding (t-SNE) plots and heat maps were generated for all the CD45+ cells.
Flow cytometry
Single-cell suspensions of kidney cells were stained with surface marker antibodies at 4 °C for 30 min. Then, the Foxp3/Transcription Factor Staining Buffer Set (eBioscience, # 00-5523-00) was used to fix and permeabilize the cells, followed by intracellular antibody staining for 30 min at 4 °C. The cell suspension was washed twice after each staining step, and the data were acquired using a BD LSRFortessaTM cytometer. The data were analyzed using BD FlowJo TM v10.6.2 software.
The following reagents and antibodies were used: live-dead-Zombie (BioLegend, #423106), CD45-PE (BioLegend, #147712), CD11b-Alexa Fluor 488 (BioLegend, #101217), F4/80-BV421 (BioLegend, #123132), CD206-APC (BioLegend, #141708), CD4-APC (BioLegend, #100515), CD19-Brilliant Violet 421 (BioLegend, #115537), FoxP3-PE (eBioscience, #12-5773-82), PD-1-APC (BioLegend, #135210), T-bet-PE-Cy7 (BioLegend, #644823) and isotype controls.
Statistics
All the data are presented as the means ± standard deviation, unless otherwise stated. Statistical analyses were performed using Prism GraphPad 9.0. Differences in means were analyzed using Student’s t test and one-way analysis of variance (ANOVA); p < 0.05 was considered to indicate statistical significance. The data are representative of three independent experiments.
Supplementary information
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant numbers 82000657 to QO, 82030025, 32141005 to XC and E3041101 to LL) and the National Key R&D Program of China (grant number 2021YFC3002203 to LZ). We thank Dr. Jianwen Chen and Dr. Tiantian Wang for their help in establishing the uIRI mouse models. We thank Dr. Jiaqi Liu and Shaoyuan Cui for their help with confocal imaging.
Author contributions
QO and LL conceived and designed the study. QO and YT performed the scRNA-seq data analysis. LLL contributed to the synthesis of the glycol peptide. QO, CW, TS, YT, JZ, YC, and XW performed the in vitro and in vivo experiments. XW and RL established the mouse models and cultured the cells. PC and JL conducted the pathology analysis. WS performed the transmission electron microscopy analysis. QO, LW, ZF, XS, and LZ analyzed and interpreted the data. QO drafted the manuscript. LLL, GC, and XC supervised the study and edited the manuscript. All the authors have approved the submitted version of the manuscript.
Data availability
All the data are available in the main text or the supplementary materials. The sc-RNA seq data that support the findings of this study are openly available in the Gene Expression Omnibus at https://www.ncbi.nlm.nih.gov/geo/, reference number GSE240885. All unique materials generated in this study, such as plasmids and cell lines, will be made available to researchers with a completed Materials Transfer Agreement upon request to the lead contact.
Competing interests
QO, TS, LLL, and XC are co-inventors on China Invention Patent Application no. ZL 2022 1 0056384.3 filed by the Chinese PLA General Hospital related to the subject matter of this manuscript.
Footnotes
These authors contributed equally: Qing Ouyang, Chao Wang, Tian Sang, Yan Tong.
Contributor Information
Qing Ouyang, Email: nolimithyer3169@outlook.com.
Guangyan Cai, Email: caiguangyan@sina.com.
Li-Li Li, Email: lill@nanoctr.cn.
Xiangmei Chen, Email: xmchen301@126.com.
Supplementary information
The online version contains supplementary material available at 10.1038/s41423-024-01190-6.
References
- 1.Kalantar-Zadeh K, Jafar TH, Nitsch D, Neuen BL, Perkovic V. Chronic kidney disease. Lancet. 2021;398:786–802. [DOI] [PubMed] [Google Scholar]
- 2.GBD Chronic Kidney Disease Collaboration. Global, regional, and national burden of chronic kidney disease, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet. 2020;395:709–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Xie Y, Bowe B, Mokdad AH, Xian H, Yan Y, Li T, et al. Analysis of the Global Burden of Disease study highlights the global, regional, and national trends of chronic kidney disease epidemiology from 1990 to 2016. Kidney Int. 2018;94:567–81. [DOI] [PubMed] [Google Scholar]
- 4.Foreman KJ, Marquez N, Dolgert A, Fukutaki K, Fullman N, McGaughey M, et al. Forecasting life expectancy, years of life lost, and all-cause and cause-specific mortality for 250 causes of death: reference and alternative scenarios for 2016-40 for 195 countries and territories. Lancet. 2018;392:2052–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zhao M, Wang L, Wang M, Zhou S, Lu Y, Cui H, et al. Targeting fibrosis, mechanisms and cilinical trials. Signal Transduct Target Ther. 2022;7:206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Humphreys BD. Mechanisms of renal fibrosis. Annu Rev Physiol. 2018;80:309–26. [DOI] [PubMed] [Google Scholar]
- 7.Li L, Fu H, Liu Y. The fibrogenic niche in kidney fibrosis: components and mechanisms. Nat Rev Nephrol. 2022;18:545–57. [DOI] [PubMed] [Google Scholar]
- 8.Ruiz-Ortega M, Rayego-Mateos S, Lamas S, Ortiz A, Rodrigues-Diez RR. Targeting the progression of chronic kidney disease. Nat Rev Nephrol. 2020;16:269–88. [DOI] [PubMed] [Google Scholar]
- 9.Hodson EM, Sinha A, Cooper TE. Interventions for focal segmental glomerulosclerosis in adults. Cochrane Database Syst Rev. 2022;2:CD003233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.de Zeeuw D, Akizawa T, Audhya P, Bakris GL, Chin M, Christ-Schmidt H, et al. Bardoxolone methyl in type 2 diabetes and stage 4 chronic kidney disease. N Engl J Med. 2013;369:2492–503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chen D, Liu Y, Chen J, Lin H, Guo H, Wu Y, et al. JAK/STAT pathway promotes the progression of diabetic kidney disease via autophagy in podocytes. Eur J Pharm. 2021;902:174121. [DOI] [PubMed] [Google Scholar]
- 12.Gale JD, Gilbert S, Blumenthal S, Elliott T, Pergola PE, Goteti K, et al. Effect of PF-04634817, an oral CCR2/5 chemokine receptor antagonist, on albuminuria in adults with overt diabetic nephropathy. Kidney Int Rep. 2018;3:1316–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kuppe C, Ibrahim MM, Kranz J, Zhang X, Ziegler S, Perales-Patón J, et al. Decoding myofibroblast origins in human kidney fibrosis. Nature. 2021;589:281–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tai Y, Woods EL, Dally J, Kong D, Steadman R, Moseley R, et al. Myofibroblasts: function, formation, and scope of molecular therapies for skin fibrosis. Biomolecules. 2021;11:1095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tarbit E, Singh I, Peart JN, Rose’Meyer RB. Biomarkers for the identification of cardiac fibroblast and myofibroblast cells. Heart Fail Rev. 2019;24:1–15. [DOI] [PubMed] [Google Scholar]
- 16.Chen T, Cao Q, Wang Y, Harris DCH. M2 macrophages in kidney disease: biology, therapies, and perspectives. Kidney Int. 2019;95:760–73. [DOI] [PubMed] [Google Scholar]
- 17.Sato Y, Yanagita M. Immune cells and inflammation in AKI to CKD progression. Am J Physiol Ren Physiol. 2018;315:F1501–F1512. [DOI] [PubMed] [Google Scholar]
- 18.Jang HR, Rabb H. Immune cells in experimental acute kidney injury. Nat Rev Nephrol. 2015;11:88–101. [DOI] [PubMed] [Google Scholar]
- 19.Horsburgh S, Todryk S, Ramming A, Distler JHW, O’Reilly S. Innate lymphoid cells and fibrotic regulation. Immunol Lett. 2018;195:38–44. [DOI] [PubMed] [Google Scholar]
- 20.Conway BR, O’Sullivan ED, Cairns C, O’Sullivan J, Simpson DJ, Salzano A, et al. Kidney single-cell atlas reveals myeloid heterogeneity in progression and regression of kidney disease. J Am Soc Nephrol. 2020;31:2833–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Yao W, Chen Y, Li Z, Ji J, You A, Jin S, et al. Single cell RNA sequencing identifies a unique inflammatory macrophage subset as a druggable target for alleviating acute kidney injury. Adv Sci. 2022;9:e2103675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Huen SC, Cantley LG. Macrophages in renal injury and repair. Annu Rev Physiol. 2017;79:449–69. [DOI] [PubMed] [Google Scholar]
- 23.Tang PM-K, Nikolic-Paterson DJ, Lan H-Y. Macrophages: versatile players in renal inflammation and fibrosis. Nat Rev Nephrol. 2019;15:144–58. [DOI] [PubMed] [Google Scholar]
- 24.Hoeft K, Schaefer G, Kim H, Schumacher D, Bleckwehl T, Long Q, et al. Platelet-instructed SPP1+ macrophages drive myofibroblast activation in fibrosis in a CXCL4-dependent manner. Cell Rep. 2023;42:112131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Qi J, Sun H, Zhang Y, Wang Z, Xun Z, Li Z, et al. Single-cell and spatial analysis reveal interaction of FAP+ fibroblasts and SPP1+ macrophages in colorectal cancer. Nat Commun. 2022;13:1742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fabre T, Barron A, Christensen SM, Asano S, Bound K, Lech MP, et al. Identification of a broadly fibrogenic macrophage subset induced by type 3 inflammation. Sci Immunol. 2023;8:eadd8945. [DOI] [PubMed] [Google Scholar]
- 27.Distler J, Györfi AH, Ramanujam M, Whitfield ML, Königshoff M, Lafyatis R. Shared and distinct mechanisms of fibrosis. Nat Rev Rheumatol. 2019;15:705–30. [DOI] [PubMed] [Google Scholar]
- 28.Tang PC-T, Chung JYF, Xue VW, Xiao J, Meng XM, Huang XR, et al. Smad3 promotes cancer-associated fibroblasts generation via macrophage-myofibroblast transition. Adv Sci. 2022;9:e2101235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Wang Y-Y, Jiang H, Pan J, Huang XR, Wang YC, Huang HF, et al. Macrophage-to-myofibroblast transition contributes to interstitial fibrosis in chronic renal allograft injury. J Am Soc Nephrol. 2017;28:2053–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ren H, Zeng XZ, Zhao XX, Hou DY, Yao H, Yaseen M, et al. A bioactivated in vivo assembly nanotechnology fabricated NIR probe for small pancreatic tumor intraoperative imaging. Nat Commun. 2022;13:418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.An H-W, Mamuti M, Wang X, Yao H, Wang MD, Zhao L, Li LL. Rationally designed modular drug delivery platform based on intracellular peptide self-assembly. Exploration. 2021;1:20210153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang D, Qi GB, Zhao YX, Qiao SL, Yang C, Wang H. In situ formation of nanofibers from purpurin18-peptide conjugates and the assembly induced retention effect in tumor sites. Adv Mater. 2015;27:6125–30. [DOI] [PubMed] [Google Scholar]
- 33.Camilio KA, Wang MY, Mauseth B, Waagene S, Kvalheim G, Rekdal Ø, et al. Combining the oncolytic peptide LTX-315 with doxorubicin demonstrates therapeutic potential in a triple-negative breast cancer model. Breast Cancer Res. 2019;21:9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Li D, Zhang J, Yuan S, Wang C, Chang J, Tong Y, et al. TGF-β1 peptide-based inhibitor P144 ameliorates renal fibrosis after ischemia-reperfusion injury by modulating alternatively activated macrophages. Cell Prolif. 2022;55:e13299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Vitale I, Yamazaki T, Wennerberg E, Sveinbjørnsson B, Rekdal Ø, Demaria S, Galluzzi L. Targeting cancer heterogeneity with immune responses driven by oncolytic peptides. Trends Cancer. 2021;7:557–72. [DOI] [PubMed] [Google Scholar]
- 36.Yamazaki T, Wennerberg E, Hensler M, Buqué A, Kraynak J, Fucikova J, et al. LTX-315-enabled, radiotherapy-boosted immunotherapeutic control of breast cancer by NK cells. Oncoimmunology. 2021;10:1962592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tang T, Huang X, Zhang G, Lu M, Hong Z, Wang M, et al. Oncolytic peptide LTX-315 induces anti-pancreatic cancer immunity by targeting the ATP11B-PD-L1 axis. J Immunother Cancer. 2022;10:e004129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Huang R, Fu P, Ma L. Kidney fibrosis: from mechanisms to therapeutic medicines. Signal Transduct Target Ther. 2023;8:129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Li Y, Luo C, Zeng Y, Zheng Z, Tao D, Liu Q, et al. Renal fibrosis is alleviated through targeted inhibition of IL-11-induced renal tubular epithelial-to-mesenchymal transition. Am J Pathol. 2023;193:1936–52. [DOI] [PubMed] [Google Scholar]
- 40.Widjaja AA, Viswanathan S, Shekeran SG, Adami E, Lim WW, Chothani S, et al. Targeting endogenous kidney regeneration using anti-IL11 therapy in acute and chronic models of kidney disease. Nat Commun. 2022;13:7497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Yamashita N, Kramann R. Mechanisms of kidney fibrosis and routes towards therapy. Trends Endocrinol Metab. 2024;35:31–48. [DOI] [PubMed] [Google Scholar]
- 42.Voelker J, Berg PH, Sheetz M, Duffin K, Shen T, Moser B, et al. Anti-TGF-β1 antibody therapy in patients with diabetic nephropathy. J Am Soc Nephrol. 2017;28:953–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Fu J, Sun Z, Wang X, Zhang T, Yuan W, Salem F, et al. The single-cell landscape of kidney immune cells reveals transcriptional heterogeneity in early diabetic kidney disease. Kidney Int. 2022;102:1291–304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mansour SG, Puthumana J, Coca SG, Gentry M, Parikh CR. Biomarkers for the detection of renal fibrosis and prediction of renal outcomes: a systematic review. BMC Nephrol. 2017;18:72. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Cheng Y, Li Y, Scherer N, Grundner-Culemann F, Lehtimäki T, Mishra BH, et al. Genetics of osteopontin in patients with chronic kidney disease: The German Chronic Kidney Disease study. PLoS Genet. 2022;18:e1010139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Klinkhammer BM, Boor P. Kidney fibrosis: emerging diagnostic and therapeutic strategies. Mol Asp Med. 2023;93:101206. [DOI] [PubMed] [Google Scholar]
- 47.Hashimoto D, Miller J, Merad M. Dendritic cell and macrophage heterogeneity in vivo. Immunity. 2011;35:323–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jaynes JM, Sable R, Ronzetti M, Bautista W, Knotts Z, Abisoye-Ogunniyan A, et al. Mannose receptor (CD206) activation in tumor-associated macrophages enhances adaptive and innate antitumor immune responses. Sci Transl Med. 2020;12:eaax6337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Li L-L, Qiao SL, Liu WJ, Ma Y, Wan D, Pan J, Wang H. Intracellular construction of topology-controlled polypeptide nanostructures with diverse biological functions. Nat Commun. 2017;8:1276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Bargh JD, Isidro-Llobet A, Parker JS, Spring DR. Cleavable linkers in antibody-drug conjugates. Chem Soc Rev. 2019;48:4361–74. [DOI] [PubMed] [Google Scholar]
- 51.Dheer D, Nicolas J, Shankar R. Cathepsin-sensitive nanoscale drug delivery systems for cancer therapy and other diseases. Adv Drug Deliv Rev. 2019;151–152:130–51. [DOI] [PubMed] [Google Scholar]
- 52.Xu L, Sharkey D, Cantley LG. Tubular GM-CSF promotes late MCP-1/CCR2-mediated fibrosis and inflammation after ischemia/reperfusion injury. J Am Soc Nephrol. 2019;30:1825–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Xu L, Guo J, Moledina DG, Cantley LG. Immune-mediated tubule atrophy promotes acute kidney injury to chronic kidney disease transition. Nat Commun. 2022;13:4892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ma P-F, Gao CC, Yi J, Zhao JL, Liang SQ, Zhao Y, et al. Cytotherapy with M1-polarized macrophages ameliorates liver fibrosis by modulating immune microenvironment in mice. J Hepatol. 2017;67:770–9. [DOI] [PubMed] [Google Scholar]
- 55.Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36:411–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell. 2015;161:1202–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All the data are available in the main text or the supplementary materials. The sc-RNA seq data that support the findings of this study are openly available in the Gene Expression Omnibus at https://www.ncbi.nlm.nih.gov/geo/, reference number GSE240885. All unique materials generated in this study, such as plasmids and cell lines, will be made available to researchers with a completed Materials Transfer Agreement upon request to the lead contact.