Abstract
Although recombinant human adenovirus (HAV)-based vectors offer several advantages for somatic gene therapy and vaccination over other viral vectors, it would be desirable to develop alternative vectors with prolonged expression and decreased toxicity. Toward this objective, a replication-defective bovine adenovirus type 3 (BAV-3) was developed as an expression vector. Bovine cell lines designated VIDO R2 (HAV-5 E1A/B-transformed fetal bovine retina cell [FBRC] line) and 6.93.9 (Madin-Darby bovine kidney [MDBK] cell line expressing E1 proteins) were developed and found to complement the E1A deletion in BAV-3. Replication-defective BAV-3 with a 1.7-kb deletion removing most of the E1A and E3 regions was constructed. This virus could be grown in VIDO R2 or 6.93.9 cells but not in FBRC or MDBK cells. The results demonstrated that the E1 region of HAV-5 has the capacity to transform bovine retina cells and that the E1A region of HAV-5 can complement that of BAV-3. A replication-defective BAV-3 vector expressing bovine herpesvirus type 1 glycoprotein D from the E1A region was made. A similar replication-defective vector expressing the hemagglutinin-esterase gene of bovine coronavirus from the E3 region was isolated. Although these viruses grew less efficiently than the replication-competent recombinant BAV-3 (E3 deleted), they are suitable for detailed studies with animals to evaluate the safety, duration of foreign gene expression, and ability to induce immune responses. In addition, a replication-competent recombinant BAV-3 expressing green fluorescent protein was constructed and used to evaluate the host range of BAV-3 under cell culture conditions. The development of bovine E1A-complementing cell lines and the generation of replication-defective BAV-3 vectors is a major technical advancement for defining the use of BAV-3 as vector for vaccination against diseases of cattle and somatic gene therapy in humans.
Replication-defective, recombinant adenoviral vectors are widely used for in vivo gene transfer because of the ability of these vectors to enter many different target cells efficiently and to express the transgene (6). Although these vectors are effective with respect to the entry and expression processes, expression of the transgene is transient (8). The transient expression is explained at least in part by the host immune responses, both innate and acquired, to cells transduced with adenoviral vectors (37). To overcome the problem associated with the preexisting immune responses, nonhuman adenoviruses have been proposed as vectors for gene therapy (21, 29, 30, 38, 39, 42). Animal adenoviruses are species specific, can enter human cells but do not replicate, and thus have great potential as gene transfer vectors. In addition, these vectors could be used as live viral vaccines in animals. Currently used recombinant adenovirus vectors are made replication defective by deletion of E1A and E1B sequences (4). The E1A region encodes the immediate-early proteins that transactivate all other viral early region genes. By deleting these sequences, the expression of all other early and the late region genes is reduced to a large extent, resulting in lower immune responses to adenoviral proteins and thus in long-term expression of a transgene. In addition, deletion of the E1 region removes the oncogenic potential of the viruses.
Bovine adenoviruses (BAVs) belong to the Mastadenovirus genus of Adenoviridae family. Currently, the accepted 10 serotypes of BAVs are divided into two subgroups on the basis of the differences in their biological and serological properties (3). Serotypes 1, 2, 3, and 9 belong to subgroup I and grow relatively well in established bovine cell lines. We have been studying BAV type 3 (BAV-3) with a goal of developing it as an expression vector. BAV-3 was chosen for this purpose based on the lack of virulence and the ability of the virus to grow to high titers in cell culture. In addition, experimental infection of calves with BAV-3 failed to produce either clinical signs or gross lesions, but all animals seroconverted (22). Molecular studies of the genome are essential for the development of BAV-3 as an expression vector. Determination of the complete nucleotide sequence and transcription map of BAV-3 was recently reported (30). Nucleotide sequence analysis of the E1 region of BAV-3 identified open reading frames for proteins that are homologous to the E1A and E1B proteins of human adenovirus type 5 (HAV-5) (11, 43). Transcription analysis of the E1 region in BAV-3 has indicated that the transcripts of the E1A, E1B, and pIX regions are 3′ coterminal (28). The proteins produced from the E1 and neighboring pIX region of BAV-3 were also identified and characterized by using specific antibodies raised in rabbits (28).
Among current methods for generating adenovirus-based vectors, the Escherichia coli BJ5183 recombination system is the most simple and efficient (5, 29, 42). We applied this method to clone the full-length BAV-3 genome as a stable infectious bacterial plasmid and to generate the replication-competent viral vector (42). In this paper, we describe generation of E1A-complementing cell lines, E1A deletion mutants, and replication-defective BAV-3 vectors. This is the first report, to our knowledge, of the development of E1A-complementing cell lines and construction of replication-defective BAV-3 expression vectors.
MATERIALS AND METHODS
Virus and viral DNA.
The WBR-1 strain of BAV-3 was cultivated in Madin-Darby bovine kidney (MDBK) and VIDO R2 cells. VIDO R2 is a transformed fetal bovine retina cell (FBRC) line expressing the E1 proteins of HAV-5. The cells were grown in Eagle’s minimum essential medium supplemented with 5% fetal bovine serum. The viral DNA was extracted from virus-infected cell monolayers by the method of Hirt (17). Replication-defective HAV-5 expressing β-galactosidase (Ad5d1E1AlacZ) was propagated as described previously (43).
Construction of recombinant plasmids.
The recombinant plasmid vectors were constructed by standard procedures (31) using restriction enzymes and other DNA-modifying enzymes as directed by the manufacturers.
(i) Construction of plasmid pFBAV500.
Plasmid pTG5435, containing the full-length BAV-3 genome in pPOLYIIsn14, was digested with HindIII, and the restriction enzyme fragment harboring the terminal fragments (nucleotides [nt] 1 to 1656 and 33232 to 34446) of BAV-3 was gel purified and self-ligated to produce plasmid pBAV-101. Nucleotide numbers of the BAV-3 genome referred in this report are according to GenBank accession no. AF030154. To delete a part of the E1A sequence between the nt 536 and 1077, pBAV-101 was digested with AccI and SpeI, and ends were blunt repaired and dephosphorylated. To facilitate the cloning of foreign genes, an XbaI linker was inserted into the deleted region, and the resulting plasmid was designated pBAV-102. The deletion of the E1A region was then rescued into the full-length infectious clone by homologous recombination in E. coli BJ5183 between the HindIII-linearized pBAV-102 and the genomic DNA from BAV3.E3d, a virus lacking the E3 region (42), creating plasmid pFBAV500.
(ii) Construction of recombinant plasmid pFBAV501.
The gene coding for glycoprotein D (gD) (33) of bovine herpesvirus type 1 (BHV-1) under the control of the simian virus 40 (SV40) immediate-early promoter, a 137-bp length chimeric intron, and SV40 late poly(A) signal was first cloned into the blunt-ended XbaI site of pBAV-102 to generate plasmid pBAV-102.gD. A full-length plasmid (E1A-E3 deletion) with the gD gene in the E1A region (pFBAV501) was generated by homologous recombination in E. coli BJ5183 between the HindIII-digested pBAV-102.gD and the genomic DNA of BAV3.E3d (42).
(iii) Construction of recombinant plasmid pFBAV502.
The genome of recombinant BAV333 (27) contains the hemagglutinin-esterase (HE) gene under the control of SV40 early promoter, a 137-bp chimeric intron, and SV40 poly(A) signal inserted in the E3 region of the BAV3.E3d genome (42). To construct a replication-defective recombinant BAV-3 expressing the HE gene of bovine coronavirus (BCV) from the E3 region of BAV-3, the genomic DNA of recombinant BAV333 was recombined with HindIII-digested plasmid pBAV-102 in E. coli BJ5183 to create plasmid pFBAV502.
(iv) Construction of plasmid pFBAV304.
The green fluorescent protein (GFP) gene under the control of the cytomegalovirus (CMV) immediate-early promoter and bovine growth hormone poly(A) signal was obtained from pQBI 25 (Quantum Biotechnologies) by BglII and DraIII digestions followed by blunt ending with T4 DNA polymerase. This fragment was first cloned into the SrfI site of the E3 transfer vector pBAV-301 (plasmid constructed by ligating the 7,635-bp KpnI-SspI fragment from pFBAV302 to KpnI-NotI [T4 treated]-digested plasmid PpolyIIsn14) in the same orientation as the E3 transcription unit to generate pBAV301.gfp. A KpnI-SwaI fragment encompassing the modified E3 region was isolated from pBAV301.gfp and recombined with SrfI-digested pFBAV.302 DNA (42) in E. coli BJ5183, creating plasmid pFBAV304.
Isolation of recombinant BAV-3.
VIDO R2 cell monolayers in 60-mm-diameter dishes were transfected with 5 to 10 μg of PacI-digested pFBAV500, pFBAV501, pFBAV502, and pFBAV304 recombinant plasmid DNAs by using Lipofectin (Gibco/BRL). After incubation at 37°C, the transfected cells showing cytopathic effects were collected and freeze-thawed twice, and the recombinant viruses were plaque purified on VIDO R2 cells.
Western blot analysis.
Extracts of mock- or virus-infected cells were resolved (5 μg per lane) by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) and transferred to a nitrocellulose membrane (Bio-Rad). Nonspecific binding sites on the membrane were blocked with 1% bovine serum albumin before incubation of the blots with protein-specific monoclonal or polyclonal antibodies. The membranes were washed and exposed to anti-mouse or anti-rat immunoglobulin G conjugated to horseradish peroxidase or alkaline phosphatase and developed by using a horseradish peroxidase or alkaline phosphatase color development kit (Bio-Rad).
Immunoprecipitation.
Confluent monolayers of VIDO R2 cells in six-well dishes were infected with the virus at a multiplicity of infection (MOI) of more than 5. The cells were preincubated for 2 h in minimal essential medium free of methionine and cystine prior to labeling with 50 μCi of [35S]methionine (Tran35S-label; phosphate-buffered saline, 1,000 Ci/mmol; ICN Radiochemicals Inc., Irvine, Calif.) for 4 h. The cells were washed once with phosphate-buffered saline, harvested by scraping, and then lysed with ice-cold modified radioimmunoprecipitation assay buffer. The radiolabeled proteins were immunoprecipitated with a pool of anti-BHV-1 gD monoclonal antibodies (MAbs) (18) or polyclonal anti-BCV rabbit antibodies (9, 10) and analyzed by SDS-PAGE. The gels were dried, and protein bands were visualized by autoradiography.
Infection of human and animal cells with BAV-3.
MDBK, VIDO R2 (bovine), STR (swine testicular cells expressing the E1 proteins of HAV-5), VIDO R1 (fetal porcine retina cell line expressing E1 proteins of HAV-5), 293, A549, HeLa, HepG2 (human), Cos, Vero (monkey), C3HA (mouse), sheep skin fibroblast, dog kidney, or cotton rat lung cells were infected with more than 5 PFU of wild-type BAV-3 or BAV304 per cell. The cells infected with wild-type BAV-3 were harvested at 2 h and 3 days postinfection, and the virus titers were determined in MDBK cells. Cells infected with BAV304 were examined for green fluorescence with the aid of a fluorescence microscope.
RESULTS
Development of MDBK cell line 6.93.9.
Our initial goal was to develop an MDBK cell line expressing E1 proteins of BAV-3, as these cells are commonly used for the propagation of BAV-3. MDBK cells are also permissive to wild-type HAV-5 infection, and the available E1 deletion mutants of HAV-5 would allow us the selection of the clones that are expressing E1 proteins of BAV-3 by cross-complementation (43). MDBK cells were transfected with plasmid pVD-Neo, containing a SalI B fragment of BAV-3 covering 0 to 23.4 map units on the viral genome and a Neor marker under the control of the long terminal repeat from Rous sarcoma virus. The SalI B fragment contains all of the E1 region, pIX, IVa2, and pol genes and part of the terminal protein (pTP) gene. The Neor system permits the selection of cells that take up the plasmid vector containing the Neor marker. Several G418-resistant clones were isolated, expanded, and tested for the ability to support the growth of Ad5dlE1AlacZ (43). Five clones supported the growth of an E1A deletion mutant, and a clone (6.93.9) which supported the growth of the virus to the highest titer was selected and subjected to single-cell cloning. However, even though these cells supported the growth of an E1A deletion mutant, we could not detect E1 protein expression in any of the clones including 6.93.9 by Western blotting using polyclonal antibodies specific to E1 proteins of BAV-3 (28). Clone 6.93.9 cells grew slowly, rapidly acidified the medium, and never formed a confluent monolayer.
Development of FBRC line VIDO R2.
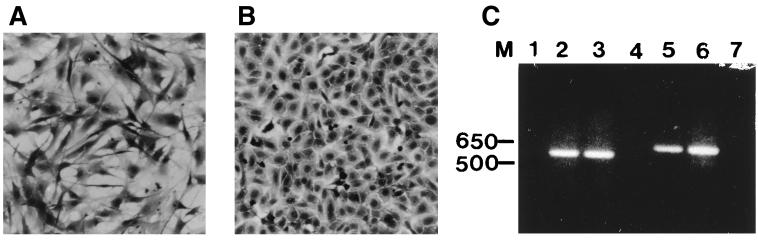
Since MDBK cells were not useful for the generation of replication-defective BAV-3 vectors, we attempted to develop an FBRC-derived cell line. Early-passage secondary FBRCs were prepared by standard techniques. Two attempts to transform FBRCs by the E1 region of BAV-3 (pVD-Neo) with or without G418 selection were not successful. As our experience with clone 6.93.9 indicated that the E1A region of BAV-3 could complement the E1A of HAV-5, we assumed that the FBRCs transformed with the E1 region of HAV-5 should also complement the E1 region of BAV-3. Human fetal retina cells are efficiently transformed with the E1 region of HAV-5 (13). To develop a bovine cell line expressing the E1 protein of HAV-5, subconfluent monolayers of FBRCs were transfected with plasmid pTG4671(pTG6559 [20] without the E1blarge mutation) by the calcium phosphate technique. Plasmid pTG4671 contains the entire E1A and E1B coding sequences of HAV-5 (nt 505 to 4034). The transcription of E1A is under the control of the constitutive mouse phosphoglycerate kinase gene (PGK) promoter, and transcription of the E1B region is under the control of its natural promoter and a globin poly(A) signal. This plasmid also has a selection marker, the puromycin acetyltransferase gene, under the control of the SV40 early promoter and SV40 poly(A) signal. Several morphologically transformed colonies were observed 4 weeks after transfection without selection for puromycin resistance. The cells in transformed foci showed altered morphology, smaller and rounded, whereas the untransformed FBRCs were long and slender (Fig. 1). The transformed cells expressed Vimentin, not cytokeratin, indicating that they are mesenchymal in origin (data not shown). Transformed cell line VIDO R2 was established from a separated foci and then subjected to single-cell cloning. Initially, the expression of E1 in VIDO R2 cell line was analyzed by reverse transcription (RT)-PCR using primers specific to E1A and E1B regions of HAV-5. The PCR products generated using the E1 region DNA as templates were used as size controls (Fig. 1C, lanes 3 and 6). As seen in Fig. 1C, HAV-5 E1A (lane 2) and 19-kDa E1B (lane 5) genes of expected lengths were transcribed in VIDO R2 cells. The RNA samples without reverse transcriptase added did not show any bands (lanes 1 and 4), indicating that genes were amplified from E1 mRNAs and not from residual DNA.
FIG. 1.
VIDO R2 cell line. Morphological features of untransformed FBRCs (A) and transformed VIDO R2 cells (B). Cells were stained with crystal violet and photographed at a magnification of ×100. (C) Analysis of ethidium bromide-stained RT-PCR products. Products of RT-PCR using DNase-treated RNA (lanes 1, 2, 4, 5, and 7) or of PCR using DNA (lanes 3 and 6) as a template were synthesized by using primer pairs for E1A (sense, 5′-ATGAGACATATTATCTGCCA-3′; antisense, 5′-CTTACTGTAGACAAACATGC) (lanes 1 to 3) and 19-kDa E1B (sense, 5′-ATGTTTAACTTGCATGGCGT-3′; antisense, 5′-ATTCCCGAGGGTCCAGGCCG-3′) (lanes 4 to 6). Lanes 1 (E1A primers) and 4 (19-kDa E1B primers) shows RT-PCR of RNA without reverse transcriptase; lane 7 shows RT-PCR using RNA from FBRCs. Sizes of marker (M) DNA are shown in base pairs.
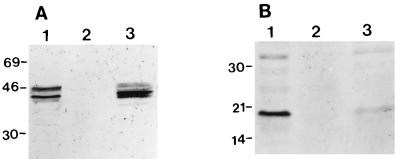
For complementation of an essential viral protein by a cell line, the protein of interest must be synthesized in sufficient quantities by the host cells. To determine the level of E1A proteins produced by VIDO R2, Western blot analysis was carried out. The mouse monoclonal antibody M73, which recognizes all HAV-5 E1A proteins, and antibody 3D11 (Calbiochem), directed against the 19-kDa HAV-5 E1B protein, were used. In VIDO R2, the levels of E1A proteins produced were similar to or little higher than those produced in 293 cells (Fig. 2A). However, the amount of 19-kDa E1B protein produced was slightly less in VIDO R2 cells (Fig. 2B). Western blot analysis using the same antibodies did not recognize any proteins from extracts from FBRCs (Fig. 2, lanes 2). Attempts to detect the HAV-5 E1B 55-kDa protein in 293 and VIDO R2 cells in immunoprecipitation assays using a rat MAb (DP 08; Cedarlane) specific to the protein were not successful.
FIG. 2.
Western blot analysis of E1 proteins. Proteins from 293 cells (lane 1), FBRCs (lane 2), and VIDO R2 cells (lane 3) were separated by SDS-PAGE (10% gel) under reducing conditions and transferred to nitrocellulose. The separated proteins were probed in Western blots by MAbs against HAV-5 E1A (A) and 19-kDa E1B (B). Sizes are indicated in kilodaltons.
To investigate the complementing properties of the VIDO R2, the cells were infected with the E1A deletion mutant of HAV-5 (Ad5dlE1AlacZ [43]). This cell line supported the growth of the deletion mutant to 107 PFU/ml. To determine whether the VIDO R2 cell line could support plaque formation, cells cultured in 35-mm-diameter dishes were infected with BAV-3 or HAV-5 and incubated in a CO2 incubator. Clear plaque formation was evident on days 5 and 7 postinfection with HAV-5 and BAV-3, respectively. We also observed a substantially more rapid onset of viral cytopathic effect in E1-expressing cell lines than in MDBK cells and FBRCs. In addition, VIDO R2 supported the formation of clear plaques by recombinant BAV-3.
Transfectability of E1-expressing cell lines.
To test the ability of the cells to take up large DNA, MDBK, 6.93.9, and VIDO R2 cells in 35-mm-diameter dishes were transfected with 1 to 3 μg of PacI-restricted plasmid pFBAV304 DNA (described below) by using Lipofectin (GIBCO/BRL). This plasmid contains the entire BAV-3 genome with the E3 region replaced by a GFP gene under the control of a CMV immediate-early promoter. When observed under a fluorescence microscope 24 h posttransfection, more than 3% of VIDO R2 cells showed fluorescence, as opposed to less than 0.1% cells in MDBK and 6.93.9 cultures. Further incubation of the transfected VIDO R2 cells for 10 to 14 days resulted in the production of a recombinant virus (named BAV304) expressing GFP. These observation suggest that VIDO R2 is a better cell line for the generation of recombinant BAV-3 due to its greater transfection efficiency and/or the presence of HAV-5 E1A/B sequences.
Construction of E1A deletion mutant of BAV-3.
To construct a replication-defective recombinant BAV-3, full-length BAV-3 genomic DNA containing deletions in the E1A (nt 536 to 1077) and E3 (nt 26456 to 27701) regions was cloned in a plasmid named pFBAV500 (Fig. 3a). Plasmid pFBAV500 DNA digested with PacI (to release the recombinant viral genome from the plasmid) was used to transfect VIDO R2 cells. A recombinant virus (named BAV500) with the E1A and E3 regions deleted was obtained 10 days following transfection. The virus was amplified in VIDO R2 cells, and the viral DNA was extracted from infected cells. The DNA was analyzed after digestion with restriction enzyme ClaI. The wild-type BAV-3 had ClaI fragments of 3.1, 6, and 7.5 kb (Fig. 4, lane 1), which were missing in the recombinant BAV500 genome, which instead had fragments of 2.5 and 12.2 kb (lane 2). This is consistent with the expected ClaI fragments of the wild type and the deletion mutant.
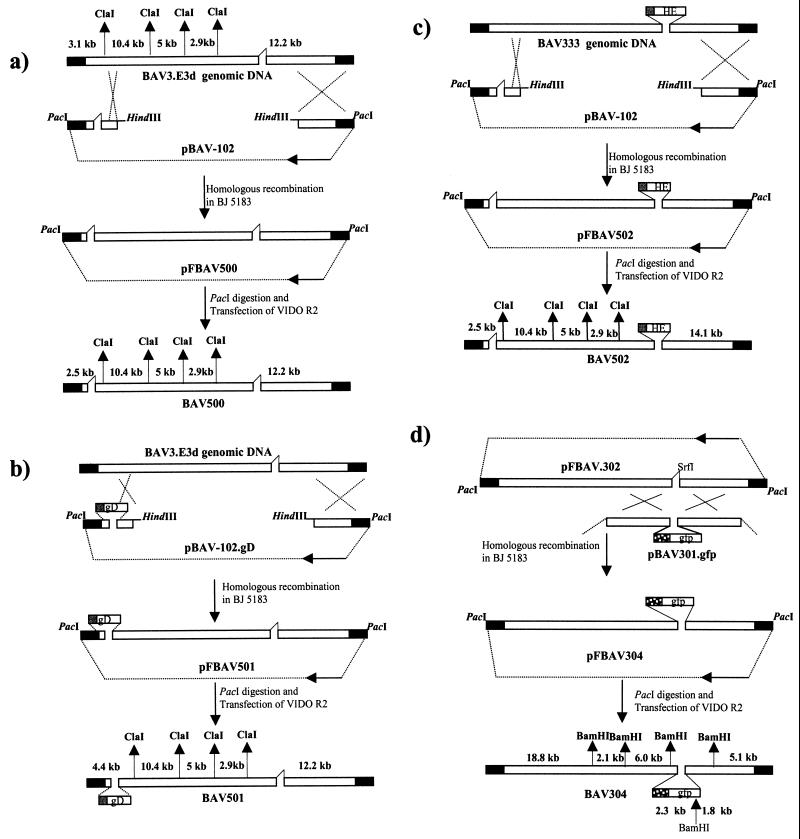
FIG. 3.
Strategy used for the generation of recombinant BAV-3. Different plasmids were constructed from different genomic clones as described in the text. Origins of DNA sequences: plasmid DNA, thin line; BAV-3 genomic DNA, hollow box; inverted terminal repeats, filled box. The plasmid maps are not drawn to scale.
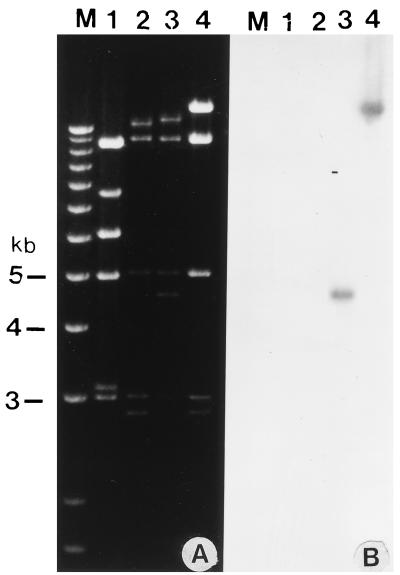
FIG. 4.
Restriction enzyme analysis of the recombinant BAV-3 genome. (A) The DNAs were extracted from VIDO R2 cells infected with BAV-3 (lane 1), BAV500 (lane 2), BAV501 (lane 3), or BAV502 (lane 4) as described previously (17) and digested with ClaI. (B) The fragments shown in panel A were transferred to Nytran membranes and probed with α-32P-labeled gD and HE probes. Lane M, 1 Kb Plus DNA Ladder (Gibco/BRL) used for sizing the viral DNA fragments.
Construction of E1A deletion-based recombinants.
To determine the usefulness of E1A-deleted (replication-defective) BAV-3 recombinants as gene delivery vehicles, we constructed recombinant BAV-3 expressing BHV-1 gD (32) or BCV HE (26). The full-length gD gene [flanked by the SV40 early promoter, chimeric intron, and SV40 poly(A) signal] was inserted into the E1A region of the BAV500 genome in the same transcriptional orientation as E1 (using the homologous recombination machinery of E. coli), (5), creating plasmid pFBAV501 (Fig. 3b). Earlier, we constructed a replication-competent recombinant BAV-3 (BAV333) expressing BCV HE (27). To construct a replication-defective recombinant BAV-3 expressing BCV HE, the full-length HE gene [flanked by the SV40 promoter, chimeric intron, and SV40 poly(A) signal] was inserted into the E3 region of the BAV500 genome in the same transcriptional orientation as E3 (using the homologous recombination machinery of E. coli, (5), creating plasmid pFBAV502 (Fig. 3c). The PacI-digested pFBAV501 or pFBAV502 DNA was transfected into VIDO R2 cells to isolate a recombinant virus named BAV501 or BAV502, respectively. BAV501 and BAV502 were amplified in VIDO R2 cells, and viral DNAs were extracted from the infected cells. The presence of foreign genes in viral genomes (gD in BAV501 and HE in BAV502) was confirmed by ClaI restriction enzyme analysis. The BAV500 genome had a ClaI fragment of 2.5 kb (Fig. 4A, lane 2) that was missing in the recombinant BAV501 genome, which instead had a fragment of 4.4 kb (Fig. 4A, lane 3). This suggested that recombinant BAV501 contained a gD gene, a possibility confirmed by Southern blot analysis (Fig. 4B, lane 3). As expected, the 12.2-kb BAV500 DNA fragment (Fig. 4A, lane 2) was replaced with a 14.1-kb DNA fragment in BAV502 (Fig. 4A, lane 4). This suggested that recombinant BAV502 contains an HE gene in the E3 region, a possibility confirmed by Southern blot analysis (Fig. 4B, lane 4).
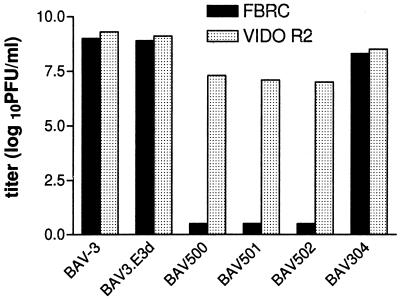
To demonstrate that infection with the E1 mutant viruses is abortive in noncomplementing cell lines, MDBK, FBRC, 6.93.9, and VIDO R2 cultures were used for infection studies. Cells were infected at an MOI of less than 1, grown for a week, subjected to two freeze-thaw cycles, and titrated on the VIDO R2 cells. Wild-type BAV-3 and BAV-3.E3d (E3 deletion [42]) grew to high titers (109 PFU/ml) in all cells tested, whereas the replication-defective recombinant viruses (E1A-E3 deletion) grew only in the VIDO R2 (Fig. 5) and 6.93.9 (106 to 107 PFU/ml) cells. This reduction in titer of replication-defective BAV-3s could be due to lower than optimum level of E1A expression in cell lines or differences in the transcriptional activation property of different E1As in different cell lines, which may affect the efficiency of virus production. Alternatively, it is possible that certain unspecified replication functions of bovine E1A cannot be efficiently replaced by HAV-5 E1A.
FIG. 5.
Production of recombinant BAV-3 by VIDO R2 and FBRC cells. Near-confluent monolayers were infected with less than 1 MOI of wild-type or recombinant BAV-3. After a week, the cells were freeze-thawed and the virus was titrated on VIDO R2 cells as described in the text.
Kinetics of expression of gD and HE.
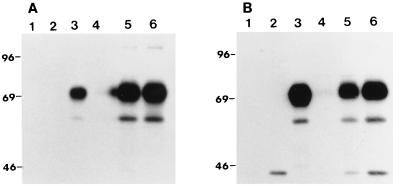
The kinetics of gD expression by recombinant BAV501 was determined at three different time points postinfection in VIDO R2 and MDBK cells by immunoprecipitation assay. Electrophoretic analysis of metabolically radiolabeled immunoprecipitates from recombinant BAV501-infected VIDO R2 cells lysates revealed immunoreactive proteins with approximate molecular masses of 63 kDa (unglycosylated) and 71 kDa (glycosylated) (Fig. 6A, lanes 5 and 6). No corresponding proteins could be detected in mock (lane 1) or wild-type BAV-3 (lane 2)-infected cells. The molecular weight of the unglycosylated and glycosylated gD corresponds to that of the authentic gD immunoprecipitated from BHV-1-infected cell extracts (lane 3). In BAV501-infected cells, expression of gD was first detected at 24 h postinfection (lane 5) and continued to be produced up to 36 h postinfection (lane 6), the last time point used in the study. Similar kinetics of gD expression was observed in MDBK cells (Fig. 6B, lanes 5 and 6).
FIG. 6.
Kinetics of gD expression by recombinant BAV-3. Proteins from lysates of [35S]methionine-labeled mock-infected (lane 1), BAV-3-infected (lane 2), BHV-1-infected (lane 3), or BAV501-infected cells harvested at 12 (lane 4), 24 (lane 5), and 36 (lane 6) h postinfection were immunoprecipitated with anti-gD MAbs (18) and separated by SDS-PAGE 10% gel under reducing conditions. (A) VIDO R2 cells; (B) MDBK cells. Positions of size markers (in kilodaltons) are shown to the left of each panel.
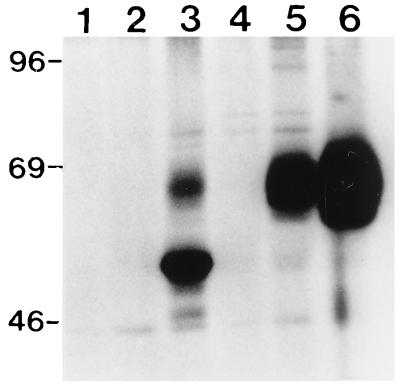
The kinetics of HE expression by recombinant BAV502 was determined in VIDO R2 cells. Anti-BCV polyclonal rabbit serum immunoprecipitated a 65-kDa polypeptide from the cells infected with BAV502 (Fig. 7, lanes 5 and 6). This polypeptide comigrated with the authentic HE protein produced from BCV-infected cells (lane 3). No such protein was immunoprecipitated from mock (lane 1)- or wild-type BAV-3 (lane 2)-infected cells. The kinetics of HE expression (lanes 5 and 6) was similar to that observed for gD in VIDO R2 cells (Fig. 6A, lanes 5 and 6). Glycosylation of the recombinant gD and HE proteins was examined by labeling the virus-infected cells with [3H]glucosamine and then performing immunoprecipitation assays. The results confirmed that the proteins produced by recombinant adenoviruses are glycosylated and are indistinguishable in migration rate from the authentic proteins produced from virus-infected cells (data not shown).
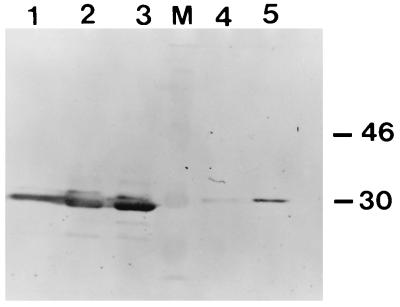
FIG. 7.
Kinetics of HE expression by recombinant BAV-3 in VIDO R2 cells. Proteins from lysates of [35S]methionine-labeled mock-infected (lane 1), BAV-3-infected (lane 2), BCV-infected (lane 3), or BAV502-infected cells harvested at 12 (lane 4), 24 (lane 5), and 36 (lane 6) h postinfection were immunoprecipitated with anti-BCV polyclonal antibodies (9, 10) and separated by SDS-PAGE (10% gel) under reducing conditions. Positions of size markers (in kilodaltons) are shown on the left.
Construction of replication-competent BAV-3 expressing GFP.
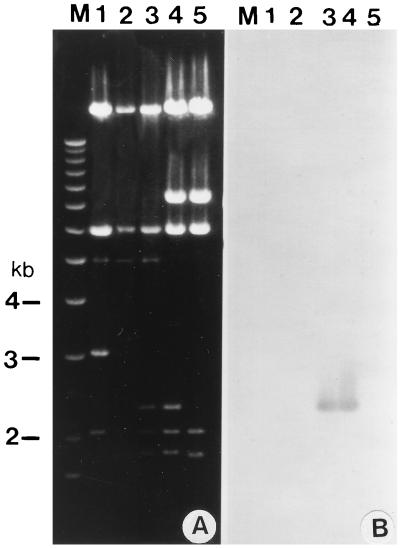
Plasmid pFBAV304 DNA (Fig. 3d) was digested with PacI and transfected into VIDO R2 cells to isolate recombinant virus BAV304. BAV304 was amplified in MDBK cells, and viral DNA was extracted from the infected cells. The viral DNA was analyzed by agarose gel electrophoresis after digestion with restriction enzyme BamHI. We observed a 2.3-kb DNA fragment in the BAV304 genome (Fig. 8A, lane 3) which was absent in the BAV3.E3d genome (Fig. 8B, lane 2). This suggested that the BAV304 genome contained a GFP gene, a possibility confirmed by Southern blot analysis (Fig. 8B, lane 3). A similar 2.3-kb DNA fragment was also observed in the pFBAV304 genome (Fig. 8A, lane 4) but not in pFBAV302 (lane 5). This suggests that pFBAV304 contained a GFP gene, a possibility confirmed by Southern blot analysis (Fig. 8B, lane 5).
FIG. 8.
Analysis of recombinant BAV304 genome. (A) The DNAs were extracted from BAV-3 (lane 1), BAV3.E3d (lane 2), BAV304 (lane 3), pFBAV304 (lane 4), and pFBAV302 (lane 5) digested with BamHI and analyzed by ethidium bromide staining of an agarose gel. (B) The fragments shown in panel A were transferred to a Nytran membrane and probed with a α-32P-labeled GFP gene. Lane M, 1 Kb Plus DNA Ladder (Gibco/BRL) used for sizing the DNA fragments.
To examine the expression of GFP, recombinant BAV304-infected cell lysates were analyzed by Western blotting using GFP-specific polyclonal antibodies (Clontech). The anti-GFP serum identified a band of 28 kDa in recombinant BAV304-infected cells (Fig. 9, lanes 1 to 3). No such band was observed in mock (lane 4)- or wild-type BAV-3 (lane 5)-infected cells. In BAV304-infected cells, GFP was detected from 12 to 36 h postinfection (lanes 1 to 3).
FIG. 9.
Western blot analysis of GFP expression. Proteins from mock (lane 4)-infected, BAV-3-infected (lane 5), or BAV304-infected MDBK cells harvested at 12 (lane 1), 24 (lane 2), and 36 (lane 3) h postinfection were separated by SDS-PAGE (10% gel) under reducing conditions and transferred to nitrocellulose. The separated proteins were probed in Western blots by GFP-specific polyclonal antiserum. Molecular masses (lane M) are indicated in kilodaltons.
Replication of BAV-3 and expression of GFP in different cell types.
To determine the host species restriction of BAV-3, 13 different cell types obtained from six different mammalian species were infected with wild-type BAV-3. BAV-3 replicated in cells of bovine origin and cotton rat lung fibroblasts. In contrast to the earlier observations (23), BAV-3 was able to replicate in cotton rat lung cells as well as or better than in bovine cells. No virus replication was apparent in cells derived from all other species, since titers were lower than those of input virus. To further characterize the host species restriction of BAV-3, we examined GFP expression in cells infected with BAV304. The greatest number of GFP-expressing cells were obtained in MDBK, VIDO R2, STR, and cotton rat lung fibroblast cultures, where as very few sheep skin fibroblasts and VIDO R1 cells showed fluorescence. Human cell lines 293 and HeLa infected with BAV304 showed mild fluorescence, while A549 and HepG2 cells showed no fluorescence (data not shown). All other virus-infected cells had low-level or no fluorescence (data not shown).
DISCUSSION
Human 293 (15), 911 (12), HAV-5 E1-expressing A549 (20), and PER (13) cells have been used to generate replication-defective HAVs. These cell lines are not suitable for BAV-3 infection, as they are nonpermissive for BAVs. To develop a bovine cell line for the generation of replication-defective BAV-3 vectors, initially MDBK cells were transfected with a plasmid, pVD-Neo, containing E1 sequences of BAV-3. A number of G418-resistant clones which supported the growth of an E1A deletion mutant of HAV-5 were selected. However, attempts to detect the E1 proteins of BAV-3 by Western blot analysis were unsuccessful. We have also not been able to generate E1 deletion mutants of BAV-3 by using this cell line. We hypothesized that this failure is due to the low transfection efficiency of MDBK cells. Consequently, we developed a second series of E1-complementing cell lines derived from FBRCs. Retina cells were chosen for the purpose because they are considered easy to transfect and transform with the E1 region of adenoviruses (13, 14). Two attempts to transform FBRCs with plasmid pVD-Neo containing the left end of the BAV-3 genome were not successful. Reasons for our failure to transform FBRCs with the E1 region of BAV-3 are unknown. However, several transformed foci were evident following the transfection of FBRCs with a plasmid, pTG4671, containing the E1 region of HAV-5 under the control of the mouse PGK promoter. The resulting transformed cells contained the integrated E1 sequences of HAV-5 in their genomic DNA as determined by PCR. These cells also produced E1A and at least one of the E1B proteins in amounts roughly as high as in 293 cells. The results of this study indicate that the diploid cultures of FBRCs could be immortalized with the E1 region of HAV-5. The resultant cell line, called VIDO R2, has been passaged more than 50 times and shown to be stable with respect to retention of the E1 region, morphology, and the ability to support the replication of E1A deletion viruses.
Functional homologies between the E1A proteins of the mouse adenovirus type 1 and HAV-5 were demonstrated in transient transfection assays (2). Similarly, Zheng et al. (43) showed transactivation activity of BAV-3 E1A proteins on the E2 and E3 promoters of HAV-5 in MDBK cells coinfected with E1A-deleted HAV-5 and BAV-3. In the present study, we demonstrated that the E1A deletion mutants of HAV-5 could be grown on 6.93.9 cells, suggesting that E1A proteins of BAV-3 could complement those of HAV-5. Using the VIDO R2 cell line and E1A deletion mutants of BAV-3, we further demonstrated that the E1A proteins of HAV-5 could complement those of BAV-3 during viral replication. This may be explained by strong homologies noticed among the E1A proteins of adenoviruses, especially in conserved region 3 (2, 28, 43). The amino-terminal part of conserved region 3 contains a zinc finger motif required for the transactivation function, and the carboxy terminus contains a domain required for association with various transcription factors (7). Once it was established that the E1A proteins of HAV-5 could complement for those of BAV-3, two BAV-3 recombinants expressing gD of BHV-1 and HE of BCV were generated from the same cell line. These viruses grow exclusively in bovine cell lines expressing E1 proteins of either BAV-3 (6.93.9) or HAV-5 (VIDO R2) but not in MDBK cells and FBRCs.
The presence of replication-competent adenoviruses (RCAs) in batches of replication-defective adenoviruses is a major problem for the use of these vectors in gene therapy and is potentially a safety risk. RCAs are generated by recombination between sequences in the adenovirus vector and the matching sequences in complementing cell lines such as 293 and 911, resulting in acquisition of the E1 region by the vector (16). The RCAs, in addition to providing helper function to replication-defective vectors, will aggravate the host immune response and enhance tissue damage (19). Although problems with RCAs can be overcome by carefully designing the vectors, an advantage of the VIDO R2 cells is that they contain the E1 region of HAV-5 and do not contain sequences that are present in replication-defective BAV-based vectors. Thus, the absence of homologous sequences should completely eliminate the problem of RCA generation by homologous recombination and allow the production of safe recombinant BAV-3 vectors.
The demonstration of a mucosal immune response following mucosal delivery of recombinant HAV-5 expressing BHV-1 gD or BCV HE in cotton rats (1, 24, 25) has stimulated interest in the development of BAV-3, vector-based vaccines in cattle. Recently, we constructed replication-competent BAV-3 recombinants expressing BHV-1 gD (42) and demonstrated that intranasal immunization of calves with these recombinants induced protective immune responses against BHV-1 (41). However, these vectors have the potential to shed recombinant virus into the environment.
Ideally, a recombinant vaccine antigen should be produced in the target animal without virus replication, thus making the live vectors safe by eliminating transmission of the virus to other farm animals and humans. Cell lines complementing E1A function allowed us to construct replication-defective BAV-3 recombinants that may serve as safe expression systems or nonreplicating vaccine vectors for cattle. For our initial experiments, we chose genes coding for protective antigens from BHV-1 (a virus involved in the development of a complex respiratory disease syndrome called shipping fever [33–35, 40]) and BCV (a virus involved in causing neonatal diarrhea [9, 10, 26]). The availability of these recombinant viruses expressing vaccine antigens should help in determining the potential of the replication-defective recombinant BAV-3 for delivery of vaccine antigens to mucosal surfaces of cattle.
Recombinant BAV-3 also has potential as a viral vector for somatic gene therapy in humans. In this respect, a recombinant BAV-3 expressing GFP represents an important tool for characterizing and potentially improving its properties as a vector. The GFP gene was chosen as a marker because it provides a simple and reliable method for the detection of transgene expression in cells. The observation of low levels of GFP expression in human cells even though the gene was placed under the control of a strong CMV immediate-early promoter indicates that the entry of BAV-3 to human cells is restricted. For successful somatic gene therapy, we need to develop gene transfer vectors with very high transduction efficiency. There are several ways to improve the transduction efficiency of BAV-3 into human cells, by utilizing high doses of the virus or by combining the virus with cationic liposomes. An alternative strategy is to replace the knob region of the fiber of BAV-3 with that of an HAV. This seems to be possible, as the entry of an ovine adenovirus into human cells was enhanced when the knob region of the fiber was replaced with that of HAV-5 (38). The RGD motif, which interacts with surface integrins (36) and facilitates the entry of virus into cells, is present in the penton base protein of HAV-5 but absent in BAV-3 (30). Introduction of such a motif into BAV-3 may facilitate the entry of BAV-3 into human cells.
ACKNOWLEDGMENTS
We thank members of the laboratory, including A. N. Zakhartchouk, Mohit Baxi, and Caron Pyne for help.
This work was supported by grants from National Science and Engineering Research Council of Canada, Medical Research Council of Canada, Saskatchewan Agriculture Development Fund, Saskatchewan Beef Development Fund, Western Economic Diversification, and Alberta Agriculture Research Institute. P.S.R. was the recipient of a postdoctoral research fellowship from the Health Services Utilization and Research Commission, Saskatoon, Saskatchewan, Canada.
Footnotes
Published with the permission of the Director of VIDO as journal series no. 258.
REFERENCES
- 1.Baca-Estrada M E, Liang X, Babiuk L A, Yoo D. Induction of mucosal immunity in cotton rats to hamagglutinin-esterase glycoprotein of bovine coronavirus by recombinant adenovirus. Immunology. 1995;86:134–140. [PMC free article] [PubMed] [Google Scholar]
- 2.Ball A O, Williams M E, Spindler K R. Identification of mouse adenovirus type 1 early region 1: DNA sequence and a conserved transactivating function. J Virol. 1988;62:3947–3957. doi: 10.1128/jvi.62.11.3947-3957.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bartha A. Proposal for subgrouping of bovine adenoviruses. Acta Vet. 1969;19:319–321. [PubMed] [Google Scholar]
- 4.Bett A J, Haddara W, Prevec L, Graham F L. An efficient and flexible system for construction of adenovirus vectors with insertions or deletions in early regions 1 and 3. Proc Natl Acad Sci USA. 1994;91:8802–8806. doi: 10.1073/pnas.91.19.8802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chartier C, Degryse E, Gantzer M, Dieterle A, Pavirani A, Mehtali M. Efficient generation of recombinant adenovirus vectors by homologous recombination in Escherichia coli. J Virol. 1996;70:4805–4810. doi: 10.1128/jvi.70.7.4805-4810.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Crystal R G. Transfer of genes to humans: early lessons and obstacles to success. Science. 1995;270:404–410. doi: 10.1126/science.270.5235.404. [DOI] [PubMed] [Google Scholar]
- 7.Culp J S, Webster L C, Friedman D J, Smith C L, Huang W J, Wu F Y, Rosenberg M, Ricciardi R P. The 289-amino acid E1A protein of adenovirus binds zinc in a region that is important for trans-activation. Proc Natl Acad Sci USA. 1988;85:6450–6454. doi: 10.1073/pnas.85.17.6450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dai Y, Schwarz E M, Gu D, Zhang W, Sarvetnick N, Verma I M. Cellular and humoral responses to adenoviral vectors containing factor IX gene: tolerization of factor IX and vector antigens allows for long term expression. Proc Natl Acad Sci USA. 1995;92:1401–1405. doi: 10.1073/pnas.92.5.1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Deregt D, Babiuk L A. Monoclonal antibodies to bovine coronavirus: characteristics and topographical mapping of neutralizing epitopes on the E2 and E3 glycoproteins. Virology. 1987;161:410–420. doi: 10.1016/0042-6822(87)90134-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Deregt D, Gilford G A, Ijaz M K, Watts T C, Gilchrist J E, Haines D M, Babiuk L A. Monoclonal antibodies to bovine coronavirus glycoproteins E2 and E3: demonstration of in vivo virus-neutralizing activity. J Gen Virol. 1989;70:993–998. doi: 10.1099/0022-1317-70-4-993. [DOI] [PubMed] [Google Scholar]
- 11.Elgadi M M, Rghei N, Haj-Ahmad Y. Sequence and sequence analysis of E1 and pIX regions of the BAV3 genome. Intervirology. 1993;36:113–120. doi: 10.1159/000150329. [DOI] [PubMed] [Google Scholar]
- 12.Fallaux F J, Kraneburg O, Cramer S J, Houweling A, van Ormondt H, Hoeben R C, van der Eb A J. Characterization of 911: a new helper cell line for the titration and propagation of early region 1-deleted adenoviral vectors. Hum Gene Ther. 1996;7:215–222. doi: 10.1089/hum.1996.7.2-215. [DOI] [PubMed] [Google Scholar]
- 13.Fallaux F J, Bout A, van der Velde I, van den Wollenberg D J M, Hehir K M, Keegan J, Auger C, Cramer S J, van Ormondt H, van der Eb A J, Valerio D, Hoeben R C. New helper cells and matched early region 1-deleted adenovirus vectors prevent generation of replication-competent adenoviruses. Hum Gene Ther. 1998;9:1909–1917. doi: 10.1089/hum.1998.9.13-1909. [DOI] [PubMed] [Google Scholar]
- 14.Gallimore P H, Grand R J, Byrd P J. Transformation of human embryo retinablasts with simian virus 40, adenovirus and ras oncogenes. Anticancer Res. 1986;6:499–508. [PubMed] [Google Scholar]
- 15.Graham F L, Smiley J, Russell W C, Nairn R. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J Gen Virol. 1977;36:59–74. doi: 10.1099/0022-1317-36-1-59. [DOI] [PubMed] [Google Scholar]
- 16.Hehir K M, Armentano D, Cardoza L M, Choquette T L, Berthelette P B, White G A, Couture L A, Everton M B, Keegan J, Martin J M, Pratt D A, Smith M P, Smith A E, Wadsworth S C. Molecular characterization of replication-competent variants of adenovirus vectors and genome modifications to prevent their occurrence. J Virol. 1996;70:8459–8467. doi: 10.1128/jvi.70.12.8459-8467.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967;26:365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- 18.Hughes G, Babiuk L A, van Drunen Littel-van den Hurk S. Functional and topographical analysis of epitopes on bovine herpesvirus-1 glycoprotein IV. Arch Virol. 1988;103:47–60. doi: 10.1007/BF01319808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Imler J L, Bout A, Dreyer D, Dieterle A, Schultz H, Valerio D, Mehtali M, Pavirani A. Trans-complementation of E1-deleted adenovirus: a new vector to reduce the possibility of codissemination of wild-type and recombinant adenoviruses. Hum Gene Ther. 1995;6:711–721. doi: 10.1089/hum.1995.6.6-711. [DOI] [PubMed] [Google Scholar]
- 20.Imler J L, Chartier C, Dreyer D, Dieterle A, Sainte-Marie M, Faure T, Pavirani A, Mehtali M. Novel complementation cell lines derived from human lung carcinoma A549 cells support the growth of E1-deleted adenovirus vectors. Gene Ther. 1996;3:75–84. [PubMed] [Google Scholar]
- 21.Klonjkowski B, Gillardi-Hebenstreit P, Hadchouel J, Randrianarison V, Boutin S, Yeh P, Perricaudet M, Kremer E J. A recombinant E1-deleted canine adenoviral vector capable of transduction and expression of a transgene in human derived cells in vivo. Hum Gene Ther. 1997;8:2103–2115. doi: 10.1089/hum.1997.8.17-2103. [DOI] [PubMed] [Google Scholar]
- 22.Lehmkuhl H D, Smith M H, Dierks R E. A bovine adenovirus type 3: isolation, characterization and experimental infection in calves. Arch Virol. 1975;48:39–46. doi: 10.1007/BF01320564. [DOI] [PubMed] [Google Scholar]
- 23.Mittal S K, Middleton D M, Tikoo S K, Prevec L, Graham F L, Babiuk L A. Pathology and immunogenicity in the cotton rat (Sigmodon hispidus) model after infection with a bovine adenovirus type 3 recombinant virus expressing the firefly luciferase gene. J Gen Virol. 1996;77:1–9. doi: 10.1099/0022-1317-77-1-1. [DOI] [PubMed] [Google Scholar]
- 24.Mittal S K, Papp Z, Tikoo S K, Baca-Esrada M E, Yoo D, Benko M, Babiuk L A. Induction of systemic and mucosal immune responses in cotton rats immunized with human adenovirus type 5 recombinants expressing the full length and truncated forms of bovine herpesvirus type 1 glycoprotein gD. Virology. 1996;222:299–309. doi: 10.1006/viro.1996.0427. [DOI] [PubMed] [Google Scholar]
- 25.Papp Z, Middleton D, Mittal S K, Babiuk L A, Baca-Estrada M E. Mucosal immunization with recombinant adenoviruses:induction of immunity and protection of cotton rats against respiratory bovine herpesvirus type 1 infection. J Gen Virol. 1997;78:2933–2943. doi: 10.1099/0022-1317-78-11-2933. [DOI] [PubMed] [Google Scholar]
- 26.Parker M D, Cox G J, Deregt D, Fitzpatrick D R, Babiuk L A. Cloning and in vitro expression of the gene for the E3 haeagglutinin glycoprotein of bovine coronavirus. J Gen Virol. 1989;70:155–164. doi: 10.1099/0022-1317-70-1-155. [DOI] [PubMed] [Google Scholar]
- 27.Reddy, P. S., N. Idamakanti, L. A. Babiuk, and S. K. Tikoo. 1999. Unpublished data.
- 28.Reddy P S, Chen Y, Idamakanti N, Pyne C, Babiuk L A, Tikoo S K. Characterization of early region 1 and pIX of bovine adenovirus-3. Virology. 1999;253:299–308. doi: 10.1006/viro.1998.9510. [DOI] [PubMed] [Google Scholar]
- 29.Reddy P S, Idamakanti N, Hyun B, Tikoo S K, Babiuk L A. Development of porcine adenovirus-3 as an expression vector. J Gen Virol. 1999;80:563–570. doi: 10.1099/0022-1317-80-3-563. [DOI] [PubMed] [Google Scholar]
- 30.Reddy P S, Idamakanti N, Zakhartchouk A N, Baxi M K, Lee J B, Pyne C, Babiuk L A, Tikoo S K. Nucleotide sequence, genome organization and transcription map of bovine adenovirus type 3. J Virol. 1998;72:1394–1402. doi: 10.1128/jvi.72.2.1394-1402.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 32.Tikoo S K, Fitzpatrick D R, Babiuk L A, Zamb T J. Molecular cloning, sequencing and expression of functional bovine herpesvirus 1 glycoprotein gIV in transfected bovine cells. J Virol. 1990;64:5132–5142. doi: 10.1128/jvi.64.10.5132-5142.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Tikoo S K, Campos M, Babiuk L A. Bovine herpesvirus 1(BHV-1): biology, pathogenesis and control. Adv Virus Res. 1995;45:191–223. doi: 10.1016/s0065-3527(08)60061-5. [DOI] [PubMed] [Google Scholar]
- 34.van Drunen Littel-van den Hurk S, Parker M D, Massie B, van den Hurk J V, Harland R, Babiuk L A, Zamb T J. Protection of cattle from BHV-1 infection by immunization with recombinant glycoprotein gIV. Vaccine. 1993;11:25–35. doi: 10.1016/0264-410x(93)90336-v. [DOI] [PubMed] [Google Scholar]
- 35.van Drunen Littel-van den Hurk S, Tikoo S K, Liang X, Babiuk L A. Bovine herpesvirus vaccines. Immunol Cell Biol. 1993;71:405–420. doi: 10.1038/icb.1993.47. [DOI] [PubMed] [Google Scholar]
- 36.Wicham T J, Filardo E J, Cheresh D A, Nemerow G R. Integrin-alpha-v-beta and integrin-alpha-v-beta-5 promote adenovirus internalization but not virus attachment. Cell. 1993;73:309–319. doi: 10.1016/0092-8674(93)90231-e. [DOI] [PubMed] [Google Scholar]
- 37.Worgall S, Wolf G, Falck-Pedersen E, Crystal R G. Innate immune mechanisms dominate elimination of adenoviral vectors following in vivo administration. Hum Gene Ther. 1997;8:37–44. doi: 10.1089/hum.1997.8.1-37. [DOI] [PubMed] [Google Scholar]
- 38.Xu Z Z, Both G W. Altered tropism of an ovine adenovirus carrying the fiber protein cell binding domain of human adenovirus type 5. Virology. 1998;248:156–163. doi: 10.1006/viro.1998.9261. [DOI] [PubMed] [Google Scholar]
- 39.Xu Z Z, Hyatt A, Boyle D B, Both G W. Construction of ovine adenovirus recombinants by gene insertion or deletion of related terminal region sequences. Virology. 1997;230:62–71. doi: 10.1006/viro.1997.8452. [DOI] [PubMed] [Google Scholar]
- 40.Yates W D G. A review of infectious bovine rhinotracheitis, shipping fever pneumonia and viral-bacterial synergism in respiratory diseases of cattle. Can J Comp Med. 1982;46:225–263. [PMC free article] [PubMed] [Google Scholar]
- 41.Zakhartchouk A N, Pyne C, Mutwiri G K, Papp Z, Baca-Estrada M, Griebel P, Babiuk L A, Tikoo S K. Mucosal immunization of calves with recombinant bovine adenovirus-3: induction of protective immunity to bovine herpesvirus-1. J Gen Virol. 1999;80:1263–1269. doi: 10.1099/0022-1317-80-5-1263. [DOI] [PubMed] [Google Scholar]
- 42.Zakhartchouk A N, Reddy P S, Baxi M, Baca-Estrada M E, Mehtali M, Babiuk L A, Tikoo S K. Construction and characterization of E3 deleted bovine adenovirus type 3 expressing full-length and truncated form of bovine herpesvirus type 1 glycoprotein gD. Virology. 1998;250:220–229. doi: 10.1006/viro.1998.9351. [DOI] [PubMed] [Google Scholar]
- 43.Zheng B, Mittal S K, Graham F L, Prevec L. The E1 sequence of bovine adenovirus type 3 and complementation of human adenovirus type 5 E1A function in bovine cells. Virus Res. 1994;31:163–186. doi: 10.1016/0168-1702(94)90002-7. [DOI] [PubMed] [Google Scholar]