Abstract
The distribution of the immune system throughout the body complicates in vitro assessments of coronavirus disease 2019 (COVID-19) immunobiology, often resulting in a lack of reproducibility when extrapolated to the whole organism. Consequently, developing animal models is imperative for a comprehensive understanding of the pathology and immunology of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection. This review summarizes current progress related to COVID-19 animal models, including non-human primates (NHPs), mice, and hamsters, with a focus on their roles in exploring the mechanisms of immunopathology, immune protection, and long-term effects of SARS-CoV-2 infection, as well as their application in immunoprevention and immunotherapy of SARS-CoV-2 infection. Differences among these animal models and their specific applications are also highlighted, as no single model can fully encapsulate all aspects of COVID-19. To effectively address the challenges posed by COVID-19, it is essential to select appropriate animal models that can accurately replicate both fatal and non-fatal infections with varying courses and severities. Optimizing animal model libraries and associated research tools is key to resolving the global COVID-19 pandemic, serving as a robust resource for future emerging infectious diseases.
Keywords: SARS-CoV-2, COVID-19, Animal models, Infection immunology, Immunotherapy
INTRODUCTION
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the pathogen responsible for coronavirus disease 2019 (COVID-19), which can manifest as mild to severe respiratory illness and can cause death in humans (Riggioni et al., 2020). Since its emergence in late 2019, the highly transmissible and variable nature of this virus has posed a significant challenge to the global medical community. As of February 2024, the pandemic has led to over 774 million confirmed cases and more than 7 million deaths worldwide (https://covid19.who.int/). Despite the continuous evolution of the virus, the current prevalent variants are not associated with increased disease severity (Statement from the Fifteenth Meeting of the International Health Regulations (IHR) (2005) Emergency Committee on the COVID-19 Pandemic). Consequently, on 4 May 2023, the World Health Organization (WHO) declared that COVID-19 no longer constituted a “public health emergency of international concern” (PHEIC). Nonetheless, considerable research challenges remain, particularly in our understanding of the role of the host immune system in disease progression, as well as the development of broad-spectrum vaccines capable of targeting rapidly mutating SARS-CoV-2 strains and the implementation of different immunotherapies. Therefore, animal model research remains a priority for advancing our knowledge of COVID-19.
Studying the immunobiology of COVID-19 using only in vitro tools is difficult due to the complexity of the human immune system. Animal models offer significant advantages as they enable the simulation and control of various factors, such as host characteristics, viral dosage, and antiviral interventions. These models facilitate procedures such as transgene expression, cell selection, and tissue sampling, which are challenging in humans, thereby providing valuable experimental data. Hence, animal models are essential for the advancement of immunological research and immunotherapy for COVID-19, particularly those involving higher mammals. During the early stages of the pandemic, various animal models were established to support vaccine and drug development, and to investigate SARS-CoV-2 pathogenesis (Muñoz-Fontela et al., 2020). To date, naturally susceptible animals such as non-human primates (NHPs), Syrian hamsters (Mesocricetus auratus), ferrets (Mustela furo), and mice (Mus musculus) — either sensitized through transgenic techniques or infected with SARS-CoV-2-adapted strains — have been widely utilized. Various studies have documented the creation, pathology, and application of animal models for COVID-19 (Bi et al., 2021; Chu et al., 2022; Kane et al., 2023; Muñoz-Fontela et al., 2020; Shou et al., 2021; Zhang et al., 2023; Zhao et al., 2023), which will not be repeated here. Other reviews have also summarized the immune responses to SARS-CoV-2 infection in both patients and animals, highlighting the unique immune characteristics of diverse animal models (Chen et al., 2024; Saravanan et al., 2022). This review focuses on current advancements in our understanding of the immunological characteristics and mechanisms of animal models for COVID-19, as well as their applications in studying immune prevention and treatment strategies, emphasizing the effective use of each model to address specific problems (Figure 1).
Figure 1.
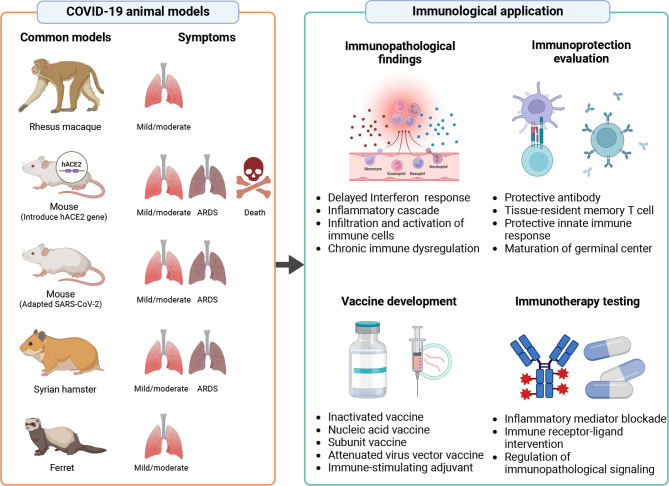
Immunological applications of common COVID-19 animal models
Rhesus macaques, mice transduced with human ACE2 gene, mice infected with adaptive SARS-CoV-2, Syrian hamsters, and ferrets are widely used to study immune pathogenesis and immune protection mechanisms of COVID-19, and also play important roles in COVID-19 vaccine and immunotherapeutic drug development. Figure created using BioRender (https://biorender.com/).
ANIMAL MODELS OF COVID-19
In response to the rapid global spread of SARS-CoV-2, various animal species, including NHPs, mice, ferrets, tree shrews, and Syrian hamsters, have been established as models to facilitate COVID-19 research. The suitability of these animals as models predominantly depends on the molecular structure of the primary binding receptor (ACE2) for the virus. Catarrhine primates, including apes and African/Asian monkeys, have ACE2 amino acid residues identical to those in human ACE2 (hACE2), while New World monkeys exhibit over 90% similarity, making both groups highly susceptible to SARS-CoV-2 (Gao & Zhang, 2020; Karki et al., 2021; Melin et al., 2020). However, the high costs associated with using large animal models often limit their utility in research, prompting a shift towards smaller animals. Tree shrews, which possess certain primate-like characteristics, show a divergence of 10 amino acids in their ACE2 key domain compared to hACE2, resulting in lower susceptibility to SARS-CoV-2 infection, characterized by mild viral shedding and lung lesions (Xu et al., 2020a; Zhao et al., 2020). Furthermore, mice and ferrets have ACE2 sequences that differ from hACE2 by eight and seven amino acids, respectively. Although mice are naturally resistant to SARS-CoV-2, humanized mouse models are widely utilized in COVID-19 research due to advancements in gene-editing techniques, including transgenic expression, clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) editing, and transient overexpression of hACE2 (Sefik et al., 2022; Zhang et al., 2023; Zhou et al., 2023). Conversely, ferrets exhibit a natural susceptibility to SARS-CoV-2, manifesting with mild clinical symptoms and relatively low lung viral titers (Zhao et al., 2023). Syrian hamsters, with only two amino acid differences in ACE2 compared to hACE2, are a preferred model for in vivo preclinical assessment of SARS-CoV-2 virulence and for testing vaccines, antivirals, and therapeutics (Handley et al., 2023). Despite these advancements, however, no single animal model can fully replicate all aspects of COVID-19, highlighting the need for tailored research approaches that utilize the unique features of each animal model (Table 1).
Table 1. Common animal models for COVID-19 immunobiology research.
| Animal | Immunopathology | Immune response | Advantage | Disadvantages |
| NHPs | Diffuse alveolar damage Lung consolidation Infiltration of inflammatory cells Thoracic adhesion Glassy opacity Hepatosplenopathy Age-related pathological changes |
Neutralizing antibody response Virus-specific T cell response Up-regulation of IL-1, IL-6, IL-10, and other inflammatory factors Transient neutropenia and lymphocytopenia |
Closer to human immune response Essential models for preclinical trials |
Lack of severe clinical symptoms High technical requirements for biosecurity High experimental costs |
| hACE2 transgenic mice | Severe interstitial pneumonia Infiltration of inflammatory cells Thickening of alveolar septum Unique vascular system damage |
Lymphopenia Pulmonary immune cell recruitment Up-regulation of IFN-γ, IL-6, MCP-1 and other inflammatory factors Diffuse microglia activation |
Excellent simulation of severe COVID-19 Dose-dependent respiratory symptoms and mortality Central nervous system infection |
Vastly different ACE2 expression pattern from humans Complex and expensive transgenic operations Lack of genetic diversity and disease models |
| AdV/AAV-hACE2 mice | Infiltration of inflammatory cells from perivascular to mesenchymal Diffuse alveolar intraepithelial infection Alveolar edema Increased vascular congestion and bleeding |
Virus-specific T cell response Up-regulation of TNF-α, IFN-γ, IL-10 and other inflammatory factors Reduced activation of CD4+, CD8+, or NK cells |
Fast and simple model construction Diverse genetic backgrounds or disease models Suitable for large-scale drug and vaccine research |
Interference from anti-Adv/AAV immune response Time and tissue limitations of viral infection |
| Mouse adapted SARS-COV-2 | Interstitial pneumonia Edema Diffuse alveolar damage Mild to moderate lung inflammation |
Proinflammatory and monocyte chemokine responses Up-regulation of IL-6, IL-1α, IL-1β and other inflammatory factors |
Easy to simulate multiple COVID-19 symptoms Diverse genetic backgrounds or disease models |
Clinical SARS-CoV-2 strains cannot be used |
| Syrian hamsters | Alveolar destruction Monocyte infiltration Alveolar collapse Lung consolidation Pulmonary hemorrhage |
Neutralizing antibody response Infiltration of macrophages and T lymphocytes Up-regulation of IFN-γ, IL-4, IL-6 and other inflammatory factors |
Naturally susceptible to SARS-CoV-2 Similar to human COVID-19 symptoms Suitable for large-scale drug and vaccine research |
Lack of reagents and tools for immunology research Lack of genetic diversity and disease models |
NHPs
Compared to other laboratory animals, NHPs possess immune and respiratory systems, as well as tissue structures, that closely resemble those of humans (Estes et al., 2018). Old World monkeys, such as rhesus macaques (Macaca mulatta), cynomolgus macaques (Macaca fascicularis), African green monkeys (Chlorocebus aethiops), and baboons (Papio hamadryas), and New World monkeys, such as common marmosets (Callithrix jacchus), are frequently employed as models for SARS-CoV-2 infection, displaying mild to moderate respiratory symptoms. In rhesus macaques, SARS-CoV-2 infection causes acute localized to widespread pneumonia, but typically without pronounced clinical respiratory symptoms (Shan et al., 2020). Notably, age significantly impacts the severity of SARS-CoV-2 in rhesus macaques, with older animals demonstrating more severe interstitial pneumonia and cytokine storm, despite a delayed immune response, compared to their younger counterparts (Song et al., 2020; Yu et al., 2020b). Cynomolgus macaques, which are smaller and less expensive to maintain, are predominant in China’s experimental primate population, with crab-eating monkeys (211 171) far outnumbering rhesus macaques (28 806). Post-infection, cynomolgus macaques show symptoms such as abnormal chest radiographs, elevated body temperature, and weight loss, although their cytokine response and lung disease are less severe than in rhesus macaques (Lu et al., 2020). Despite the abundance of cynomolgus macaques, rhesus macaques are preferred in COVID-19 research due to their higher suitability. Northern pig-tailed macaques (Macaca leonina), lacking the retroviral restriction factor tripartite motif-containing protein 5α (TRIM5α), exhibit more severe inflammation and physical impairment from SARS-CoV-2 than rhesus macaques (Kuang et al., 2009; Song et al., 2021). However, limited research on these macaques due to their small number hinders a full understanding of their potential in studies. Common marmosets exhibit lower susceptibility to SARS-CoV-2, with less pronounced and replicable manifestations of COVID-19 (Ireland et al., 2022; Lu et al., 2020). African green monkeys and baboons, though utilized less frequently, can also serve as SARS-CoV-2 models. Compared to macaques, baboons experience prolonged viral RNA shedding and more severe lung inflammation (Singh et al., 2020), while African green monkeys display higher SARS-CoV-2 replication and marked respiratory issues, including pronounced inflammation and clotting disorders (Woolsey et al., 2021). The correlation between age and COVID-19 severity is consistent across these NHP species, highlighting their utility in exploring the effects of aging and metabolic diseases on SARS-CoV-2 infection (Singh et al., 2020; Song et al., 2020; Yu et al., 2020b). Despite requiring biosafety level 3 (BSL-3) facilities and specialized personnel to handle SARS-CoV-2-infected NHPs, these models are essential for evaluating COVID-19 treatments and vaccines (Guebre-Xabier et al., 2020; Rabdano et al., 2023; Sui et al., 2021b; Vogel et al., 2021).
Mice
Due to their small size, large populations, and low feeding costs, mice are extensively used as model organisms for a variety of human diseases. However, significant structural differences between hACE2 and mouse ACE2 (mACE2) inhibit the natural replication of SARS-CoV-2 in wild-type mice (Mahdy et al., 2020). Consequently, current strategies for developing COVID-19-susceptible mouse models involve either introducing the hACE2 gene or adapting mACE2 through viral passage (Knight et al., 2021). The diverse array of genetic and immunological tools available for mice allows for a more effective exploration of COVID-19 pathogenesis, offering advantages not achievable with other animal models and contributing to a more profound understanding of this complex disease.
hACE2 transgenic mice
Animal susceptibility to SARS-CoV-2 is influenced by the structural characteristics of ACE2, while COVID-19 severity is closely associated with its expression (Ni et al., 2020). As a result, the transgenic introduction of hACE2 into mice under the control of efficient promoters has become a highly effective strategy for constructing COVID-19 models. In 2007, hACE2 transgenic mice were generated to evaluate SARS-CoV in vivo using cytomegalovirus (CMV) immediate enhancer/β-actin (CAG) and keratin-18 (K18) promoters, with the former enabling robust hACE2 expression across various mouse cell types and the latter restricting high hACE2 expression to epithelial cells (McCray et al., 2007; Tseng et al., 2007). The K18-hACE2 mouse model has also been extensively used in COVID-19 research, replicating dose-dependent characteristics of SARS-CoV-2 infection, including loss of olfaction, severe lung pathology, thrombosis, vasculitis, nerve damage, and mortality, similar to those observed in human COVID-19 patients (Zheng et al., 2021b). However, K18-initiated hACE2 is overexpressed in the brain, exceeding ACE2 expression levels in humans (Chen et al., 2021b). Consequently, K18-hACE2 transgenic mice exhibit a disparity in brain infection levels compared to COVID-19 patients (Winkler et al., 2020). To address this issue, hACE2 transgenic mice were developed using the mACE2 promoter to ensure equivalent hACE2 expression in mACE2+ cells (Yang et al., 2007), resulting in a self-limiting and non-lethal course of infection, which is invaluable for assessing the efficacy of antiviral therapeutics and vaccines (Bao et al., 2020). Additionally, CRISPR/Cas9 knock-in technology has been employed to generate transgenic mouse models expressing hACE2, which display detectable SARS-CoV-2 in the lung, trachea, and brain, accompanied by interstitial pneumonia and elevated cytokine levels (Sun et al., 2020b).
Viral vectors for hACE2 transduction in mice
Employing adenoviral vectors (AdV) or adeno-associated virus vectors (AAV) to transiently express hACE2 in mouse lungs is considered highly effective for modeling SARS-CoV-2 infection in strains with distinct genetic characteristics or diseases. Notably, intranasal administration of AdV-encoded hACE2 in wild-type BALB/c or C57BL/6 mice, with subsequent SARS-CoV-2 infection at the peak of hACE2 expression at 5 days post-transduction, leads to weight loss, severe lung lesions, and high viral replication in the lungs (Sun et al., 2020a). In general, AAVs are preferred over AdVs due to their lower immunogenicity, prolonged expression of target genes, and enhanced suitability for in vivo gene delivery. Intranasal administration of hACE2 encoded by AAV9 and AAV-DJ results in robust hACE2 gene expression in the lungs of mice, while AAV6 yields relatively low expression (Gary et al., 2021; Glazkova et al., 2022; Israelow et al., 2020). Mice humanized with AAV-hACE2, specifically the MSTRG6 strain, exhibit chronic COVID-19 features, including weight loss, persistent viral RNA presence, fibrotic lung pathology, human-like macrophage responses, sustained interferon (IFN)-stimulated gene (ISG) expression, and T cell lymphopenia (Glazkova et al., 2022; Sefik et al., 2022). These AdV/AAV-hACE2 mouse models provide a readily available and genetically varied framework for studying SARS-CoV-2 infection, crucial for rapid preclinical assessments.
Mouse-adapted viruses
For research involving COVID-19 animal models where SARS-CoV-2 mutation is not a factor, using a mouse-adapted strain of SARS-CoV-2 may be a preferable option. Dinnon et al. (2020) engineered a mouse-adapted recombinant SARS-CoV-2 variant by modifying the Q498Y/P499T amino acid residue in the receptor-binding domain (RBD) of the spike (S) protein, enabling viral replication in both the upper and lower respiratory tracts of wild-type BALB/c mice, although young mice rapidly cleared the infection within 4 days. Leist et al. (2020) generated the MA10 variant by passaging mouse-adapted SARS-CoV-2 through mice 10 times, inducing pulmonary disease and potential acute respiratory distress syndrome (ARDS) in BALB/c mice, but only mild lung inflammation and weight loss in C57BL/6 mice. Another variant carrying the N501Y substitution, generated by serial passage of SARS-CoV-2 through the respiratory tract of elderly BALB/c mice, induces interstitial pneumonia without visible clinical symptoms or weight loss (Gu et al., 2020). The SARS2-N501Y MA30 strain, produced through successive passages of SARS2-N501Y in mouse lungs, shares spike protein mutations with the Omicron variant and can cause fatal disease in young BALB/c mice (Kibler et al., 2022; Wong et al., 2022). The generation of highly virulent mouse-adapted SARS-CoV-2 through in vivo passage alone mirrors unrestricted SARS-CoV-2 transmission in the population without preventative measures, providing a vital model for studying viral transmission dynamics.
Hamsters
Hamsters, particularly Syrian hamsters (Mesocricetus auratus), are highly susceptible to SARS-CoV-2 infection, making them a preferred model for viral study. SARS-CoV-2 effectively replicates in the pulmonary system of Syrian hamsters, targeting the lower respiratory tract within 12 hours post-inoculation and causing severe pathological alterations and olfactory dysfunction similar to symptoms observed in COVID-19 patients (Clancy et al., 2023; Imai et al., 2020; Merle-Nguyen et al., 2024). SARS-CoV-2 infection in Syrian hamsters is characterized by rapid viral clearance by day 7, resembling infection dynamics in mildly infected humans, possibly attributed to varying viral infection titers (Sia et al., 2020). Few animal models of COVID-19 exhibit clinical features of ARDS, limiting our understanding of the molecular mechanisms underlying the host immune response to SARS-CoV-2 infection. However, SARS-CoV-2-infected Syrian hamsters display key ARDS characteristics, such as lung injury, increased permeability, acute inflammation, and hypoxemia, particularly when inflammatory mediators are up-regulated and persist after viral clearance (Bednash et al., 2022). Thus, Syrian hamster models of SARS-CoV-2 infection reflect several aspects observed in moderate, self-limiting cases of COVID-19, including respiratory and vascular inflammation and age-related patterns (Bednash et al., 2022; Gruber et al., 2022; Imai et al., 2020). In addition, Roborovski pygmy hamsters (Phodopus roborovskii) are highly susceptible to SARS-CoV-2, developing severe acute diffuse alveolar injury and pulmonary hyaline microthrombi similar to severe COVID-19 patients (Trimpert et al., 2020). Chinese hamsters (Cricetulus griseus) can also be infected with SARS-CoV-2, presenting with clinical symptoms akin to the Syrian hamster models but with prolonged symptom duration (Bertzbach et al., 2021). As valuable models for SARS-CoV-2 infection, hamsters, mice, and NHPs have been used in combination for the development of viral vaccines and drugs (Baum et al., 2020; Lu et al., 2023; Yu et al., 2023).
Other animal models
Various other rodents have been used in studies of SARS-CoV-2 infection. Guinea pigs (Cavia porcellus) do not appear to be susceptible to SARS-CoV-2 infection, as the virus cannot be detected despite early histological changes in the lungs by day 3 (Hewitt et al., 2020). In contrast, New Zealand white rabbits (Oryctolagus cuniculus) are sensitive to SARS-CoV-2 but remain asymptomatic (Mykytyn et al., 2021), making them unsuitable as a disease model. Interestingly, Yu et al. (2022a) established a SARS-CoV-2-susceptible hACE2-transgenic rat (Rattus norvegicus) model but found that wild-type Sprague-Dawley (SD) rats infected with the prototype strain of SARS-CoV-2 showed detectable viral loads in the upper respiratory tract and lung lesions. The increase in demand for COVID-19 research has expanded the range of animals used for testing and model development, beyond conventional primate and rodent models. Studies have indicated that SARS-CoV-2 exhibits limited replication in dogs, pigs, chickens, and ducks, but shows efficient replication in the upper respiratory tract of ferrets and in the respiratory and digestive systems of cats (Shi et al., 2020). The respiratory tract of ferrets, which is physiologically and anatomically similar to that of humans, is naturally sensitive to a variety of human respiratory viruses, including influenza virus, SARS-CoV, and SARS-CoV-2 (Belser et al., 2011; Kim et al., 2020; Shi & Hu, 2008). SARS-CoV-2 infection in ferrets results in fever, mild respiratory symptoms, acute bronchiolitis, and diffuse interstitial histiocytic pneumonia, but not death (Kim et al., 2020; Kreft et al., 2022). Minks (Neovison vison), also belonging to the weasel family, experience severe respiratory disease and death following SARS-CoV-2 infection, presenting with pulmonary edema, moderate vasculitis, and fibrinous interstitial pneumonia (Eckstrand et al., 2021). Domestic cats (Felis catus) also exhibit high susceptibility to SARS-CoV-2 infection, although their widespread use is impeded by the challenges of non-standard animal models (Shi et al., 2020). Additionally, in-depth study of infection immunology in these models is hindered by the scarcity of specialized immunological research tools.
INFECTION IMMUNOLOGY IN ANIMAL MODELS OF COVID-19
Immune pathogenesis of SARS-CoV-2 infection
The pathophysiology of COVID-19 primarily arises from host dysregulation in controlling SARS-CoV-2 replication and modulating the immune response, resulting in delayed viral clearance, inflammation, and tissue damage that extends beyond pulmonary involvement to systemic manifestations (Arish et al., 2023). The host innate immune system detects RNA viruses, like coronaviruses, through various pattern recognition receptors (PRRs). Toll-like receptor 3 (TLR3), TLR7, and TLR8 recognize viral genomic RNA, double-stranded RNA (dsRNA), and single-stranded RNA (ssRNA). Cytoplasmic RNA sensors, such as retinoic acid-inducible gene I (RIG-I) and melanoma differentiation-associated protein 5 (MDA5), detect dsRNA intermediates formed during viral replication. This detection triggers the activation of IFN regulatory factor 3 (IRF3)/IRF7-dependent type I and III IFN transcription, as well as nuclear factor kappa B (NF-κB)-dependent proinflammatory cytokines and chemokines, initiating the antiviral process (Merad et al., 2022).
SARS-CoV-2 disrupts the innate immune system by targeting viral sensors and blocking downstream antiviral signaling molecules, facilitating successful transmission and adaptation to human hosts (Kasuga et al., 2021). Patients with moderate to severe COVID-19 exhibit impaired responses to both IFN-I and IFN-III, associated with persistent viral load in the bloodstream, while patients with mild disease show early induction and higher levels of IFN (Galani et al., 2021; Hadjadj et al., 2020). As the virus evades immune surveillance and efficiently infects neighboring target cells, leading to acute lung injury, it continues to activate the innate immune response via PRRs and damage-associated molecular patterns (Chu et al., 2020). Uncontrolled immune responses trigger secondary systemic inflammation and a cytokine storm, marked by immune cell overactivation, abnormal blood counts, and elevated levels of circulating cytokines, such as tumor necrosis factor (TNF)-α, IFN-I and IFN-II, interleukin (IL)-1, IL-6, IL-12, IFN-γ-inducible protein 10 (IP-10), and monocyte chemoattractant protein 1 (MCP-1), culminating in septic shock, ARDS, and multiple organ failure (Fajgenbaum & June, 2020; Kasuga et al., 2021).
Initial insights into the pathological mechanisms of COVID-19 were derived from patient blood samples, biopsies, and autopsy specimens. With the advancement of animal models replicating SARS-CoV-2 infection, a more comprehensive understanding of COVID-19 pathogenesis has been achieved (Figure 2). An ideal animal model should capture the key aspects of SARS-CoV-2 pathogenesis and associated immune responses, allow detailed study of multiple anatomical sites of viral infection through invasive sampling, and support experimental approaches to manipulate the immune system. Currently, NHPs, mice, and golden hamsters are commonly used as models of SARS-CoV-2 infection due to their similarity to human patients regarding virus sensitivity, anatomical sites of viral infection, immune system components, and clinical symptoms. However, no single animal model can perfectly replicate all characteristics of COVID-19. By effectively and thoughtfully utilizing each animal model, we can gain a deeper understanding of the immunopathological mechanism underpinning SARS-CoV-2 infection.
Figure 2.
Infection immunology based on COVID-19 animal models
Application of animal models of SARS-CoV-2 infection has promoted systematic research on COVID-19 immunopathology. Animal experiments on vaccines and adjuvants have advanced our understanding of the immunoprotective mechanisms against SARS-CoV-2 infection. Figure created using BioRender (https://biorender.com/).
IFN-I response
The role of IFN-I in SARS-CoV-2 infection is complex and extensively studied. IFN-I is crucial for antiviral immunity, exerting various antiviral effects and promoting immune cell recruitment, thereby playing a protective role against SARS-CoV-2 infection (Zhang et al., 2020a). However, evidence also links a strong IFN-I response and increased ISG expression with the development of severe COVID-19 (Lee & Shin, 2020; Zhu et al., 2020). This abnormal IFN-I response in COVID-19 is further supported by animal models. Notably, C57BL/6J mice infected with the SARS-CoV-2 MA10 strain do not develop severe disease, while those lacking IFN-I and IFN-II receptors develop more pronounced and longer-lasting pulmonary dysfunction after infection, confirming the protective role of IFN-I signaling (Leist et al., 2020). Studies have shown that SARS-CoV-2 suppresses IFN-I expression, promoting infection in models involving rhesus macaques and Syrian hamsters (Francis et al., 2021; Sui et al., 2021a). However, most animal models have reported an association between IFN-I signal intensity and disease severity. For instance, AAV-hACE2-transfected IFN-α/β receptor (IFNAR)−/− and IRF3/7−/− mice have revealed that the IFN-I response post-SARS-CoV-2 infection is essential for the recruitment of proinflammatory monocytes and macrophages to lung lesions, which induces inflammation (Israelow et al., 2020). Prolonged IFN signaling in mice with respiratory tract infection disrupts lung epithelial cell repair, thereby increasing disease severity and susceptibility to bacterial infection (Broggi et al., 2020; Major et al., 2020). In baboons, SARS-CoV-2 infection leads to a higher production of inflammatory factors, including IFN-α, compared to rhesus macaques, resulting in more severe lung pathology (Singh et al., 2020). K18-hACE2 transgenic mice infected with SARS-CoV-2 exhibit high and persistent IFN-I, IFN-II, and IFN-III expression, highlighting the correlation between IFN expression and COVID-19 severity (Oladunni et al., 2020; Zheng et al., 2021b).
Inflammatory response
Activation of inflammasomes by SARS-CoV-2 infection initiates an inflammatory cascade, which can, in severe cases, lead to a lethal cytokine storm and systemic organ failure (Mehta et al., 2020). Animal models have been instrumental in elucidating the inflammation caused by SARS-CoV-2 infection and its role in disease pathogenesis. NHPs infected with SARS-CoV-2 exhibit typical clinical symptoms, such as fever, cough, dyspnea, and interstitial pneumonia. However, severe lung injury and diffuse alveolitis are primarily observed in rhesus macaques and African green monkeys (Azkur et al., 2020; Yuan et al., 2021). This severe condition is marked by thickening of the alveolar wall and infiltration of numerous monocytes and lymphocytes, as well as a small number of eosinophils (Shan et al., 2020). Beyond the respiratory tract, NHP models also show pathological changes in the heart, liver, kidneys, spleen, lymph nodes, and brain, although direct evidence of SARS-CoV-2 infection in these tissues is still lacking (Lu et al., 2020; Philippens et al., 2022). Research involving K18-hACE2 transgenic mice has shown that lung function impairment is associated with the infiltration of monocytes, neutrophils, and activated T cells, leading to a heightened innate immune response (Winkler et al., 2020). Similarly, Syrian hamsters infected with SARS-CoV-2 exhibit symptoms such as interstitial pneumonia, inflammatory cell infiltration, alveolar septum thickening, and vascular system injury, as well as a pulmonary immune response driven by macrophages (Bednash et al., 2022; Mulka et al., 2022). These findings suggest that inflammatory injury induced by SARS-CoV-2 is more critical than the multiorgan damage resulting from direct viral infection.
The cytokine storm associated with COVID-19 is a key pathogenic factor and potential therapeutic target in COVID-19 patients (Zanza et al., 2022). Clinical studies have highlighted the significant role of cytokines such as IL-6, IL-1, IL-17, and TNF-α in lung injury (Montazersaheb et al., 2022). In cases of severe COVID-19, SARS-CoV-2 induces a chronic immune response mediated by transforming growth factor-β (TGF-β), which promotes fibrosis (Ferreira-Gomes et al., 2021). SARS-CoV-2-infected NHPs also show elevated expression of inflammatory cytokines, including IL-10, IL-1A, IL-8, IL-15, MCP-1, IFN-β, and IP-10, but unlike in severe COVID-19 patients, IL-6 is not highly expressed (Lu et al., 2020; Song et al., 2020), underscoring its critical role in cytokine storms. Transcriptomic analysis of lung tissue from SARS-CoV-2-infected mice with severe disease has shown that antiviral responses in younger mice are dominated by IFN and IL-6 pathway activation, while fatal outcomes in older animals are associated with TNF and TGF-β signaling, highlighting the significant impact of age on cytokine response in COVID-19 (Bader et al., 2023). In the Syrian hamster model of SARS-CoV-2 infection, various cytokines, such as IFN-α, IL-6, IL-1β, and TNF, are associated with severe lung disease, consistent with clinical observations (Fomin et al., 2023; Francis et al., 2021).
Immune cell activation
Clinical studies have demonstrated that cytokine storms in COVID-19 are driven by the robust activation of monocytes and macrophages, leading to the development of SARS-CoV-2-related complications, including ARDS, disseminated intravascular coagulation syndrome (DICS), edema, and pneumonia (Kosyreva et al., 2021; Liao et al., 2020). Similarly, SARS-CoV-2-infected K18-hACE2 transgenic mice exhibit pronounced recruitment of pulmonary immune cells, including dendritic cells (DCs), innate myeloid cells (IMMs), and CD4+ and CD8+ T lymphocytes (Zheng et al., 2021b). In rhesus macaques, SARS-CoV-2 infection induces rapid recruitment of macrophage subsets, particularly CD163+ mannose receptor C-type 1 (MRC1)- and triggering receptor expressed on myeloid cells 2 (TREM2)+, which are primary sources of inflammatory cytokines (Upadhyay et al., 2023). In ferrets, SARS-CoV-2 infection also results in notable alterations in the composition of macrophage subsets in bronchoalveolar lavage fluid (BALF), with monocyte-derived M1 and M2 macrophages playing pivotal roles in both early viral clearance and late-phase excessive inflammation (Lee et al., 2021). Furthermore, in Syrian hamsters, SARS-CoV-2 infection induces rapid and extensive infiltration of monocyte-derived macrophages (MDM) in the lungs, contributing to tissue remodeling and fibrosis via the up-regulation of prothrombotic factors, tissue repair, and alveolar cell proliferation (Bagato et al., 2024). Neutrophils, the most abundant white blood cells, are the first to infiltrate infected lungs following viral infection. Chemokine (C-X-C motif) ligand 5 (CXCL5) plays an important role in the process of neutrophil recruitment, with its knockout in SARS-CoV-2-infected mice found to significantly reduce pulmonary inflammation (Liang et al., 2020).
Hospitalized COVID-19 patients, especially those with severe disease, frequently exhibit a notable decrease in blood lymphocyte counts (Liu et al., 2020a; Sekine et al., 2020). Autopsies have revealed T cell infiltration and migration in inflamed tissues, suggesting that the decrease in blood lymphocytes may be due to their relocation to infection sites (Adamo et al., 2021; Jafarzadeh et al., 2020). However, the percentage of CD8 T cells in the BALF is lower in severe patients compared to those with moderate disease, possibly due to apoptotic signals (André et al., 2022; Liao et al., 2020). Despite variations in T cell numbers, their activation in SARS-CoV-2 infection-induced pulmonary inflammation is significantly increased. This heightened activation, characterized by overactivation, depletion, or apoptosis sensitivity, is identified through various cell surface markers, including CD38, human leukocyte antigen-DR isotype (HLA-DR), programmed cell death 1 (PD-1), T cell immunoreceptor with Ig and ITIM domains (TIGIT), T cell immunoglobulin and mucin domain-containing protein 3 (Tim-3), and natural killer (NK) cell receptor A (NKG2A). These activated T cells are closely linked to the development of severe COVID-19 (Du et al., 2021; Georg et al., 2022; Zheng et al., 2020), as supported by various animal models. For example, Zheng et al. (2021a) observed a significant increase in T cell CXCR3 and CD38 expression in the lung tissues of elderly macaques infected with SARS-CoV-2, indicative of both activation and inflammation and potentially valuable for predicting severe COVID-19. Similarly, SARS-CoV-2-infected rhesus macaques exhibit a marked increase in T cell activation markers in their lung tissues and cerebrospinal fluid (Verma et al., 2021), while SARS-CoV-2-infected mice show a significant up-regulation in CD44+CD3+CD8+T cell activation in their lungs, accompanied by alveolar macrophage loss (Winkler et al., 2020). In K18-hACE2 mice, T cell activation is correlated with diffuse microglial activation and plays a critical role in SARS-CoV-2 brain infection (Seehusen et al., 2022). Wu et al. (2021) found that the SARS-CoV-2 membrane (M) protein activates STAT1 phosphorylation and T-bet transcription via TREM-2, contributing to T cell activation. Experiments using CD4-specific conditional TREM-2 knockout mice have demonstrated that TREM-2 enhances the TH1 response and aggravates lung injury.
Immune protection against SARS-CoV-2
Following the SARS-CoV-2 outbreak, the development of vaccines and drugs necessitated relevant animal studies to evaluate their safety and efficacy in vivo. Various NHP models, particularly rhesus macaques, have been employed to test the efficiency of different SARS-CoV-2 vaccines developed using different technologies, including inactivation (A'La et al., 2023; Chen et al., 2021a), spike protein or RBD subunit expression (Guebre-Xabier et al., 2020; Prenafeta et al., 2023), DNA (Yu et al., 2020a), mRNA (Oh et al., 2023; Vogel et al., 2021), and attenuated viral vectors (Feng et al., 2020; Jacob-Dolan et al., 2021). Mouse and hamster models have primarily been applied for testing new vaccines and adjuvants and studying immunoprotective mechanisms, such as immune-stimulating adjuvant chimeric vaccines (Ashhurst et al., 2022), multivalent vaccines (Afkhami et al., 2022), lipid nanoparticle vaccines (Elia et al., 2021), adenovirus vaccines (Port et al., 2023), DC vaccines (Tada et al., 2023), novel adjuvants (Machado et al., 2023; Vijayanand et al., 2023), and cross-protection mechanisms (Liu et al., 2023a). The effects of different vaccine delivery modes on immune efficacy have also been verified using animal models. In addition to traditional intramuscular injection, oral (Beddingfield et al., 2021; Langel et al., 2022; Yu et al., 2022b) and intranasal vaccines (Langel et al., 2022; Sui et al., 2021b; Tokunoh et al., 2023) have also been shown to provide protection. These animal models have significantly enhanced our understanding of the immunoprotective mechanisms against SARS-CoV-2 (Figure 2).
Protective adaptive immune response
Animal models have been instrumental in elucidating the importance of adaptive immune protection in SARS-CoV-2 infection. Speranza et al. (2022) discovered that older rhesus macaques exhibit a sustained inflammatory innate response to SARS-CoV-2 infection, while younger animals initiate an earlier local effector T-cell response, leading to more rapid recovery of immune homeostasis. A key immunological feature of SARS-CoV-2-infected NHPs and hamsters is their ability to develop protective immunity, which not only protects against secondary infection but may also cross-react with other coronaviruses (Chandrashekar et al., 2020; Horiuchi et al., 2021; Jacob-Dolan et al., 2021). McMahan et al. (2021) demonstrated that the transfer of purified IgG from convalescent rhesus macaques protects recipient macaques from SARS-CoV-2, highlighting that CD8+ T cell responses can confer protection even with low or weak antibody levels. The CD4+ T cell response in rhesus macaques is strongly correlated with the efficacy of SARS-CoV-2 spike protein mRNA vaccines and neutralizing antibody levels (Corbett et al., 2021). In mice, the IFN-I pathway-mediated virus-specific T cell immune response is critical for protecting against severe SARS-CoV-2 infection (Zhuang et al., 2021). CD8+ T cells are essential for early control and elimination of SARS-CoV-2, while CD4+ T cells play a crucial role in eliciting antibody responses (Israelow et al., 2021), underscoring the importance of adaptive immunity in mitigating disease progression and preventing secondary infections. Despite the significant role of T cells in recovery from acute SARS-CoV-2 infection in rhesus macaques, their depletion does not result in severe disease, suggesting that T cells alone do not fully account for the innate resistance of rhesus macaques to severe COVID-19 (Hasenkrug et al., 2021).
The respiratory mucosal surface is a crucial site for interactions between SARS-CoV-2 and the immune system, where mucosal antibodies and tissue-resident memory T (TRM) and B cells provide early antiviral immune responses (Lee & Oh, 2022; Zheng & Wakim, 2022). However, challenges in sampling from human mucosal sites, especially the lower respiratory tract, have hindered studies on local immunity to SARS-CoV-2. Animal model vaccination experiments have offered vital insights into mucosal immune memory following SARS-CoV-2 infection. Tang et al. (2022) demonstrated that mice immunized with a combination of systemic mRNA vaccine and mucosal AdV-S protein vaccine exhibit a strong neutralizing antibody response, providing better respiratory mucosal protection than the systemic mRNA vaccine alone. Intratracheal inoculation of rhesus macaques with a bivalent SARS-CoV-2 vaccine based on Ad26 significantly induces humoral and cellular immunity in mucosa, with almost complete protection against SARS-CoV-2 BQ.1.1 challenge (McMahan et al., 2024). Furthermore, intranasal delivery of the ChAdOx1 vaccine induces potent respiratory RBD-specific IgA antibody titers, antibody-dependent neutrophil and monocyte phagocytosis, complement activation, NK cell activation, and reduced viral shedding following SARS-CoV-2 infection (Oh et al., 2021). Intranasal immunization with trivalent COVID-19 AdV vaccine elicits a robust multifunctional respiratory tract mucosal TRM response in K18-hACE2 mice, enhancing mucosal protection and conferring complete protection against SARS-CoV-2 infection-induced disease or mortality (Afkhami et al., 2022). Research on SARS-CoV-2-infected K18-hACE2 mice has also shown that respiratory memory T cells, both CD4+ and CD8+, induced by a subunit-based adjuvant system, can protect against SARS-CoV-2 infection, even in the absence of virus-neutralizing antibodies (Van Doremalen et al., 2021).
Protective innate immune response
The efficacy of COVID-19 vaccines may be reduced in certain populations, such as the elderly, highlighting the need for more personalized vaccination regimens. Targeted activation of innate immune signals has gained attention as a strategy to elicit strong immune responses in vaccine development. Studies have shown that mRNA vaccines can enhance the innate immune response in obese mice with impaired adaptive immunity by enhancing IFN-I signaling and providing protection against SARS-CoV-2 infection (Chen et al., 2023). In rhesus macaques, mRNA-based vaccines can effectively stimulate all immune system components and improve the connection between innate and adaptive immunity after a second dose via key mediators, such as TNFα, IFNγ, IL-21, and chemokine (C-C motif) ligand 3 (CCL3) (Schramm et al., 2023). Marx et al. (2022) found that therapeutic RNA oligonucleotides, acting as RIG-I agonists, can trigger a robust NK cell response in K18-hACE2 mice, providing transient yet potent antiviral protection against SARS-CoV-2 infection. TLR2, an important innate immune receptor on respiratory epithelial and lung immune cells, is crucial for effective defense against infection (Beckett et al., 2012). The TLR2 ligand Pam 2 Cys, when used as a mucosal adjuvant in COVID-19 vaccines, enhances the recruitment of antigen-presenting cells (APCs), including DCs, alveolar macrophages, and monocytes, to the respiratory mucosa, leading to robust activation of the IL-17+CD4+ T cell response and significantly boosting the protective efficacy of COVID-19 vaccines in murine models (Ashhurst et al., 2022). Lymphoblastoid cells play a crucial role in shaping the microenvironment of lymph nodes and the structure and function of the germinal center (GC). Age-related decline in GC response impedes the establishment of durable humoral immunity post-vaccination (Masters et al., 2018). Denton et al. (2022) discovered that TLR4 agonists can promote the maturation and expansion of follicular DCs (FDCs) and the activation and proliferation of stromal cells, thus enhancing GC immune response initiation and COVID-19 vaccine efficacy in aged mice. Additionally, the RBD dimer fusion protein carrying IFN exhibits potential as a candidate for COVID-19 vaccination by targeting DCs in lymph nodes and stimulating T follicular helper (Tfh) cell differentiation and GC formation, providing complete protection against high-dose SARS-CoV-2 challenge in rhesus macaques (Sun et al., 2021).
Protective immune response against novel SARS-CoV-2 variants
In late 2020, after nearly a year of transmission in humans, SARS-CoV-2 underwent significant adaptive changes. These highly mutated variants, referred to as “variants of concern” (VOC), exhibit higher transmission rates than earlier strains. To date, the WHO has identified five SARS-CoV-2 variants as VOCs, including Alpha, Beta, Gamma, Delta, and Omicron, all of which exhibit significant changes in transmissibility or immune escape, warranting close surveillance (Carabelli et al., 2023). As of 9 February 2024, the primary circulating variants of interest include XBB.1.5, XBB.1.16, EG.5, BA.2.86, and JN.1 (https://www.who.int/activities/tracking-SARS-CoV-2-variants). These variants have numerous nonsynonymous mutations in the spike protein and unique phenotypic characteristics, leading to changes in transmissibility and antigenicity, which facilitate evasion of host immune responses. This evolution may reduce the effectiveness of current SARS-CoV-2 vaccines (Kim et al., 2022). Therefore, it is crucial to use appropriate animal models to further study the etiology, transmission, and pathogenesis of the virus and to evaluate the efficacy of vaccines against the latest SARS-CoV-2 variants.
Compared to the Delta variant, the Omicron BA.1 and BA.2 variants have been shown to exhibit weakened lower respiratory tract replication and lower pathogenicity in a variety of animal models, including rodents and NHPs (Boon et al., 2022; Halfmann et al., 2022; Van Doremalen et al., 2022). Halfmann et al. (2023) reported that the Omicron subvariant XBB.1.5 exhibited greater airborne transmissibility than its predecessor BA.2 and partial immune escape from previous infection with BA.1 in Syrian hamsters. Tamura et al. (2024) found that nonsense mutations in ORF8 of XBB.1.5 impaired MHC suppression and were associated with reduced virulence in Syrian hamsters infected with this subvariant. XBB.1.16, an independent variant from XBB.1.5, has two amino acid substitutions in its spike protein compared to XBB.1.5: E180V in the N-terminal domain and T478R in the receptor-binding domain (Yamasoba et al., 2023). Pseudovirus experiments have shown that XBB.1.16 has similar infectivity to XBB.1.5 and similar sensitivity to XBB.1 in convalescent serum as XBB.1 and XBB.1.5 (Tamura et al., 2023). Meehan et al. (2023) developed a digital pathology algorithm to quantitatively assess respiratory lesions caused by SARS-CoV-2, confirming that Omicron subvariants BA.2.75 and EG.5.1 have regained some virulence, although not to the levels previously observed in Omicron. Similar to XBB.1.5, EG.5.1 is more transmissible among hamsters, but exhibits significantly enhanced immune evasion characteristics, and shows no significant differences in replication ability or pathogenicity compared to BA.2 (Uraki et al., 2023). Herder et al. (2023) evaluated the virulence of BA.2.86 and BA.2.75, among the most virulent Omicron subvariants, and found that BA.2.86 exhibited an attenuated phenotype in hamsters, suggesting no greater risk to public health than its parental Omicron subvariants. JN.1, derived from BA.2.86, harbors a key mutation (S:L455S) in the spike protein, which enhances its immune evasion ability and resistance to monovalent XBB.1.5 vaccine serum (Altamimi et al., 2024). However, the lack of animal models associated with JN.1 has hindered pathology and vaccine development.
Several animal-based vaccine studies have been conducted against novel SARS-CoV-2 variants. An Omicron (BA.1)-specific mRNA vaccine has shown effective protection in hamsters against BA.1 at 253 days after immunization, and moderate neutralizing activity against Omicron subvariants in rhesus macaques at 9 months after immunization (Wu et al., 2023b). Immunization of mice and rhesus macaques with the XBB.1.5-based monovalent recombinant spike protein COVID-19 vaccine induces neutralizing antibodies against the XBB.1.5, XBB.1.16, XBB.2.3, EG.5.1, and XBB.1.16.6 subvariants and elicits a CD4+ Th1 cell response specific to the XBB subvariant (Patel et al., 2023). The Omicron XBB.1.5 RBD dimer-based vaccine completely protects mice from Omicron XBB.1.16 and significantly reduces respiratory viral infection in Syrian hamsters, demonstrating excellent protective efficacy against SARS-CoV-2 and its variants (Wu et al., 2023a). Wang et al. (2024) identified a highly conserved neutralizing epitope targeted by the broad-spectrum neutralizing antibody BA7535, which was highly neutralizing not only to previous variants (e.g. Alpha, Beta, Gamma, Delta and Omicron BA.1-BA.5) but also to recently emerging Omicron subvariants BF.7, CH.1.1, XBB.1, XBB.1.5, XBB.1.9.1, and EG.5. BA7535 effectively protects female mice from Omicron BA.5 and XBB.1 variants. The novel ChAd-SARS-CoV-2 vaccine, consisting of a bivalent Ad36 vector vaccine encoding SARS-CoV-2 WA1 and BA.5 spike proteins, induces persistent antibody and T cell responses against XBB.1.16 infection in rhesus macaques immunized by intranasal spray or inhalation aerosol (Gagne et al., 2023). The new generation of RBD nano-antibodies neutralizes Alpha, Delta and Omicron BA.2.75, BA.1, BA.2, BA.4/5 and XBB.1 variants, significantly reducing viral load, weight loss, and pathogenicity in infected hamsters (Aksu et al., 2024).
IMMUNOLOGICAL INSIGHTS OF POST-COVID-9 SYMPTOMS (LONG COVID-19) FROM ANIMAL MODELS
Development of long COVID-19
Although most individuals fully recover from SARS-CoV-2 infection within days or weeks, some experience persistent or new symptoms months or even years later, a condition known as long COVID-19 (Schmidt, 2021). This complex multisystem disorder is characterized by over 200 identified symptoms, including fatigue, dyspnea, arthralgia, myalgia, cardiac arrhythmias, and cognitive impairments, affecting up to 75% of those who have recovered from COVID-19 (Jansen et al., 2022). Huang et al. (2022) found that patients infected with the original strain of SARS-CoV-2 continued to experience long-term symptoms more than 2 years after hospital discharge, although most symptoms had improved. Significant risk factors for long COVID-19 include female sex, obesity, and severe COVID-19 disease; however, even outpatients with mild symptoms may experience prolonged COVID-19 manifestations (Frontera et al., 2022; Subramanian et al., 2022). The original SARS-CoV-2 strain is associated with a higher incidence of long COVID-19 compared to the Alpha or Delta variants (Fernández-de-las-Peñas et al., 2022). In contrast, the Omicron variant is associated with a lower risk of long-term pneumonia compared to the Delta variant (Antonelli et al., 2022; Wise, 2022). Older adults infected with SARS-CoV-2 are more likely to suffer from long COVID-19, presenting with fatigue, difficulty breathing, coughing, and joint pain, as well as abnormalities in chest imaging and lung function tests. The situation is more complex for children and adolescents (Daitch et al., 2022), with immunocompromised children showing a higher prevalence of gastrointestinal symptoms associated with long COVID-19, while immunocompetent children report increased levels of fatigue (Roessler et al., 2022).
The mechanisms behind long COVID-19 and reasons for differences in symptoms across viruses and populations are unclear. Key hypotheses include changes in the immune system, persistence of residual viral components driving chronic inflammation, endothelial dysfunction or activation, changes in the microbiome, mitochondrial dysfunction, abnormal metabolites, reactivation of pre-existing chronic viral infections, microbiota dysbiosis, and unrepaired tissue damage (Davis et al., 2023; Liu et al., 2023b; Santopaolo et al., 2023). From an immunological perspective, uncontrolled immune dysregulation following SARS-CoV-2 clearance is an important contributor to the development of long COVID-19. In-depth immunophenotyping analysis has revealed significant gene expression interference in innate immune cells (such as NK cells, low-density neutrophils, and CXCR3+ monocytes) and adaptive immune cells (such as T helper (Th) cells, Tfh cells, and regulatory T cells) in long-term COVID-19 patients (Ryan et al., 2022). Klein et al. (2023) demonstrated that long COVID-19 patients exhibit an excessive humoral response to SARS-CoV-2 and higher levels of antibody response to non-SARS-CoV-2 viral pathogens, especially Epstein-Barr virus. Single-cell omics and serologic studies have also revealed systemic inflammation and immune dysregulation in patients with long COVID-19, characterized by an increased frequency of inflammatory CD4+ T cells and an exhausted phenotype of virus-specific CD8+ T cells, indicating disharmony between humoral and cell-mediated responses (Yin et al., 2024). Despite these findings, clinical challenges remain in identifying potential therapeutic targets for COVID-19, necessitating urgent in vivo intervention studies using animal models.
Animal models of long COVID-19
Animal models for long COVID-19 are still in the early stages. Paidas et al. (2022) inoculated mice with mouse hepatitis virus 1 (MHV-1), a member of the same β-coronavirus family as SARS-CoV-2, for 12 months to simulate long COVID-19, finding severe pathological changes and inflammation in their brain, lungs, and heart. Although not a direct SARS-CoV-2 infection model, these results imply that coronaviruses can have long-term effects, even after viral clearance. Sefik et al. (2022) examined innate and adaptive immune responses to SARS-CoV-2 infection in an AAV-hACE2 mouse model over 28 days, revealing characteristics of chronic COVID-19, such as progressive weight loss, persistent viral RNA presence, lung pathology accompanied by fibrosis, inflammatory macrophage response, sustained expression of ISGs, and depletion of T lymphocytes. Mice infected with SARS-CoV-2 for up to 120 days exhibit persistent inflammation and fibrosis, important features of chronic COVID-19 lung injury, as well as an increase in interstitial macrophages (Dinnon et al., 2022). The neurological symptoms associated with SARS-CoV-2 infection may potentially arise from cytokine storm, neuroimmune stimulation, and systemic infection, rather than direct viral mechanisms of injury (Efstathiou et al., 2022). Evidence suggests that SARS-CoV-2 infection-induced CCL11 evokes a selective microglial response in white matter, leading to dysfunction and structural dysregulation of multiple neuronal lineages in the central nervous system, contributing to cognitive impairment during the acute phase following COVID-19 (Li et al., 2022). In Syrian hamsters, 31 days of SARS-CoV-2 infection results in permanent lung and kidney damage, along with unique effects on the olfactory bulb and olfactory epithelium, including myeloid and T cell activation, proinflammatory cytokine production, and IFN responses, leading to neurobehavioral disorders such as anosmia and depression (Frere et al., 2022). Due to the lack of animal models of long COVID-19 and the high cost of BSL-3 laboratories, it is difficult to summarize the immunological mechanisms underlying COVID-19 development in a small number of cases, hindering the advancement of appropriate treatments. Solving this issue requires the development of pseudovirus and transgenic mouse models suitable for establishing animal models for long COVID-19.
APPLICATION OF ANIMAL MODELS IN COVID-19 IMMUNOTHERAPY
Following the COVID-19 outbreak, various new and repurposed compounds exhibiting in vitro anti-coronavirus activity have rapidly advanced towards clinical application. Prior to the development of an effective SARS-CoV-2 vaccine, the US Food and Drug Administration (FDA) granted emergency use authorization (EUA) for a nucleoside analog and three monoclonal antibodies (MAbs) (Tao et al., 2021). Despite widespread vaccination, developing drug therapy for SARS-CoV-2 remains crucial, particularly for immunocompromised individuals and those who may not respond to the rapidly mutating virus (Boby et al., 2023). Current COVID-19 antiviral drugs are primarily divided into the following categories: ① Polymerase inhibitors, which act on RNA-dependent RNA polymerase (RdRps), mainly nucleoside analogs, including remdesivir, molnupiravir, and nelfinavir, and have been validated in rhesus monkeys (Johnson et al., 2023; Tao et al., 2021; Williamson et al., 2020) and Syrian hamsters (Abdelnabi et al., 2021); ② Protease inhibitors, which act on 3C-like proteinase protein (3CLpro), mainly nirmatrelvir, PF-07321332, S-217622, ensitrelvir, and have been validated in rhesus monkeys (Rosenke et al., 2023), Syrian hamsters (Sasaki et al., 2023; Tao et al., 2021), and mice (Jeong et al., 2022; Tao et al., 2021); ③ Entry inhibitors, including antibodies, fusion inhibitors, soluble recombinant hACE2, which have achieved good results in SARS-CoV-2 infected rhesus monkeys (Streblow et al., 2023), mice (Morgan et al., 2022; Tao et al., 2021), Syrian hamsters (Linsky et al., 2020; Uraki et al., 2022), and ferrets (De Vries et al., 2021). The rapid development of COVID-19 immunotherapy research, relying on in vivo testing, can be attributed to the establishment of diverse animal models for SARS-CoV-2 infection, leading to numerous potential therapeutic drugs and protocols. These findings will serve as a valuable resource for addressing future infections, even if COVID-19 ceases to be a major public health threat. This discussion will not delve into the efficacy and mechanisms of these antivirals but will instead focus on trial outcomes of immunomodulatory drugs in COVID-19 animal models (Figure 3).
Figure 3.
COVID-19 immunotherapy based on animal models
Validation of immune interventions targeting immune receptors, inflammatory mediators, and immunopathological signals in SARS-CoV-2-infected animal models is crucial for the development of COVID-19 immunotherapy. Figure created using BioRender (https://biorender.com/).
Targeting immune receptors
Since the early stages of the global pandemic, research has focused on targeting immune receptors for COVID-19. Early studies revealed that severe COVID-19 patients often show reduced T-cell counts, linked to poor clinical outcomes and indicating potential T-cell exhaustion (Liu et al., 2020b; Zheng et al., 2020). Lymphopenia in severe COVID-19 is linked to the overexpression of immune checkpoint receptors PD-1, cytotoxic T-lymphocyte-associated protein 4 (CTLA-4), TIGIT, and Tim-3 (Jin et al., 2023; Yin et al., 2024). With numerous inhibitory receptor antagonists already in clinical trials and available for use, their rapid implementation in COVID-19 trials is likely, minimizing the necessity for preclinical animal testing (Loretelli et al., 2021; Tan & Li, 2022). Given our advancing comprehension of the interactions between SARS-CoV-2 and immune receptors, the validation of novel immune receptor-targeted drugs will largely depend on animal models to confirm their efficacy.
SARS-CoV-2 enters host cells and is recognized by TLRs, NOD-like receptors (NLRs), and NLR protein 3 (NLRP3), initiating IFN-I production and innate immune reactions. Targeting these immune receptors therapeutically may help ameliorate infection-induced inflammation. TLRs, crucial for recognizing SARS-CoV-2 and initiating innate immunity, are expressed in various cell types, including immune cells, fibroblasts, epithelial cells, and ACE2-expressing type II lung cells (Jiang et al., 2022; Qi et al., 2020). TLRs, including those in human (TLR1–TLR10) and mouse cells (TLR1–TLR9 and TLR11–TLR13), are broadly categorized into cell surface TLRs, including TLR1, TLR2, TLR4–6, and TLR10, and intracellular somatic membrane TLRs, including TLR3 and TLR7–9 (O'Neill et al., 2013). Upon activation, TLRs recruit specific bridging molecules like myeloid differentiation primary response protein 88 (MyD88) and Toll/IL-1 receptor (TIR) domain-containing linker-induced interferon β (TRIF), which trigger the IRF and NF-κB signaling pathways to initiate IFN-I and inflammatory cytokine production (Jiang et al., 2022). TLR-mediated immune dysregulation plays a critical role in hyperinflammation and cytokine storm observed in COVID-19, making it a potential target for intervention (Liu et al., 2022). Zheng et al. (2021c) demonstrated that the SARS-CoV-2 E protein induces TLR2-related inflammatory responses in mice, while inhibition of TLR2 with oxPAPC significantly attenuates the release of proinflammatory cytokines and chemokines. Fontes-Dantas et al. (2023) reported that intracerebral administration of the spike protein in mice induces neuroinflammation and synaptic loss through TLR4-mediated microglial activation, with early intervention with TLR4 inhibitors shown to prevent late neuronal damage in mice, identifying TLR4 as a potential target for long-term cognitive dysfunction induced by COVID-19. The role of TLRs in COVID-19 is complex, extending beyond immunopathology. Pre-COVID-19 studies indicated that MyD88- and TRIF-deficient mice infected with SARS-CoV experience higher mortality rates, weight loss, and viral loads, suggesting that MyD88 and TRIF are essential for preventing fatal SARS-CoV infection (Sheahan et al., 2008; Totura et al., 2015). Furthermore, combining a TLR9 agonist with mRNA enhances both T cell and neutralizing antibody responses against RBD in mice, compared to mRNA alone (Haabeth et al., 2021). Treating K18-hACE2 transgenic mice with a lethal SARS-CoV-2 dose and the synthetic agonist Poly(I:C) for TLR3/MDA5 temporarily boosts the innate immune response in the lungs, subsequently reducing viral load and cytokine storm levels, and significantly improving survival (Tamir et al., 2022).
Inflammatory processes at the cellular level are typically mediated by inflammasomes, particularly NLRP3 inflammasomes, which are highly responsive to RNA viruses. NLRP3 inflammasomes are composed of the NLRP3 receptor, apoptosis-associated speck-like protein containing a C-terminal caspase recruitment domain (ASC), and caspase-1. Upon assembly, caspase-1 is activated by autoproteolytic cleavage, facilitating the maturation of proinflammatory cytokines such as IL-1β, IL-18, IL-6, TNF-α, and gasdermin-D (GSDMD), which all contribute to the development of COVID-19-induced cytokine storm (Broz & Dixit, 2016; Dutta et al., 2022). Increasing evidence suggests that hyperactivation of NLRP3 inflammasomes plays a role in the pathogenesis of SARS-CoV-2-induced multiorgan failure (Ratajczak & Kucia, 2020). Studies have indicated that Nlrp3−/− mice, which lack NLRP3 inflammasomes, exhibit reduced severity of SARS-CoV-2-induced lung pathology compared to wild-type C57BL/6 mice, while specific inhibition of MCC950 reduces excessive lung inflammation in SARS-CoV-2-infected hACE2 transgenic mice, confirming NLRP3 as a potential immunotherapeutic target for COVID-19 (Zeng et al., 2022). Albornoz et al. (2023) discovered that SARS-CoV-2 infection activates NLRP3 inflammasomes in microglia through the NF-κB and ACE2 signaling pathways, leading to neurodegeneration in hACE2 transgenic mice, with these effects significantly mitigated and survival rates improved with oral administration of an NLRP3 inhibitor. These findings suggest that targeting NLRP3 may be beneficial in alleviating neurocognitive disorders associated with long-term SARS-CoV-2 infection.
Targeting inflammatory mediators
In the context of acute lung injury in COVID-19, macrophage activation syndrome and ARDS arise from the release of proinflammatory cytokines, such as IL-1, IL-6, IL-18, and TNF-α. Consequently, targeting these cytokines presents significant therapeutic potential for reducing cytokine storm in COVID-19 patients (McGonagle et al., 2020). Rubsamen et al. (2020) noted that early administration of α-IL-6 mAbs in mice infected with Ebola more effectively combated cytokine release syndrome (CRS) than IL-6 receptor blockade alone, offering insights for potential SARS-CoV-2 treatments. Subsequently, Xu et al. (2020b) found that the recombinant human IL-6 mAb tocilizumab binds specifically to soluble and membrane-bound IL-6 receptors (sIL-6R and mIL-6R), inhibiting IL-6-mediated signal transduction and improving clinical outcomes in severe and critically ill COVID-19 patients. IL-18 is also implicated in SARS-CoV-2-induced inflammatory lung injury. Notably, IL-18 is associated with the excessive recruitment, infiltration, and activation of neutrophils observed in the lung tissue of COVID-19 patients, as well as progression of cardiovascular dysfunction (Liang et al., 2023). In hACE2 transgenic mice, exposure to the spike protein increases NF-κB activity and NLRP3-inflammasome-dependent IL-18 levels in heart and lung tissues, with subsequent improvement in cardiac function and reduced NF-κB activity following inhibition of IL-18 (Liang et al., 2023). In K18-hACE2 mice, SARS-CoV-2 infection activates the NLRP3 pyroptosis signaling pathway in the lungs, resulting in IL-1β release, pulmonary vascular injury, and associated protein-rich pulmonary edema, while the selective IL-1 receptor antagonist anastrozole blocks IL-1 receptor signaling and prevents VE-cadherin down-regulation and excessive pulmonary vascular permeability, significantly improving survival (Xiong et al., 2021). Clinical findings indicate that the bispecific IL-1β/IL-18 MAb MAS825 effectively inhibits biomarkers of clinical and inflammatory pathways associated with COVID-19 and promotes the rapid clearance of SARS-CoV-2 (Hakim et al., 2023). Dysregulation of TNF-α, an important proinflammatory cytokine in the innate immune response, can induce CRS (Ragab et al., 2020). Karki et al. (2021) demonstrated that inhibition of TNF-α and IFN-γ can effectively mitigate sepsis, hemophagocytic lymphohistiocytosis (HLH), and cytokine shock in K18-ACE-2 transgenic mice following SARS-CoV-2 infection. Several anti-TNF therapeutics, including Humira (adalimumab), Remicade (infliximab), Simponi (golimumab), Cimzia (certolizumab pegol), and Entyvio (vedolizumab), are currently undergoing clinical trials for COVID-19, with most studies confirming favorable impact on reducing or ameliorating disease progression (Hakim et al., 2023).
Targeting immunopathological signaling pathways
The Janus kinase (JAK)-signal transducer and activator of transcription (STAT) pathway is pivotal in the regulation of inflammatory factors and formation of cytokine storms in COVID-19. Consequently, several JAK/STAT signaling inhibitors, including baricitinib, ruxolitinib, and tofacitinib, have been investigated as potential treatments for COVID-19 (Hall et al., 2023; Rein et al., 2022). Despite this, clinical trials have indicated that JAK/STAT signaling inhibitors are less effective than expected in alleviating disease severity in severe COVID-19 cases. Baricitinib effectively inhibits JAK1/2 kinase by competitively blocking adenosine triphosphate (ATP), essential for JAK1/2 activation by inflammatory cytokines such as IL-6, and disrupts various proinflammatory signaling pathways, including IL-6, IL-1β, IL-12, granulocyte-macrophage colony-stimulating factor (GM-CSF), and IL-2 (Ravid et al., 2022; Zhang et al., 2020b). Additionally, baricitinib effectively attenuates pulmonary inflammation in SARS-CoV-2-infected rhesus macaques by suppressing cytokines and chemokines, which, in turn, reduce neutrophil recruitment and pulmonary macrophage inflammation. However, its effect on viral replication and the IFN-I response is minimal, resulting in incomplete resolution of immunopathology (Hoang et al., 2021).
Bruton’s tyrosine kinase (BTK) is essential for B cell development in the bone marrow and plays a key role in the proliferation and survival of leukemia B cells (Hendriks et al., 2014). Elevated BTK activation levels in blood monocytes from severe COVID-19 patients are associated with systemic inflammation (Roschewski et al., 2020). Ibrutinib, a BTK inhibitor, is widely used in the treatment of indolent B cell malignancies and chronic graft-versus-host disease (cGVHD) and shows potential in eliminating inflammatory cytokines in the lungs, minimizing lung injury, and lowering mortality risk in a lethal influenza mouse model (Florence et al., 2018). Ibrutinib effectively inhibits poly I:C and lipopolysaccharide (LPS)-induced activation of BTK, fms-like tyrosine kinase 3 (FLT3), and epidermal growth factor receptor (EGFR)-related pathways, significantly reducing acute lung injury in mice (Rao et al., 2022). These findings suggest that BTK inhibitors may serve as potential therapeutic agents for regulating SARS-CoV-2-induced acute lung injury. BTK inhibitors, including acalabrutinib, ibrutinib, and zanubrutinib, have been utilized in COVID-19 clinical trials, showing protection against lung injury and reduced inflammatory activity (Kifle, 2021; Treon et al., 2020).
Excessive up-regulation of p38 activity may drive the intense inflammatory response observed in COVID-19 infection through two potential mechanisms: first, the loss of ACE2 after viral entry may hinder the conversion of angiotensin II to angiotensin 1–7, thereby releasing the inhibition of p38 activity; second, SARS-CoV-2 may exploit p38 activity to facilitate its replication (Grimes & Grimes, 2020; Higgins et al., 2023). Therefore, p38 inhibition may be a promising therapeutic strategy for alleviating COVID-19. Jimenez-Guardeño et al. (2014) found that during SARS-CoV infection, syntenin binds to the E protein, leading to syntenin redistribution from the nucleus to the cytoplasm, activating the p38 mitogen-activated protein kinase (MAPK) pathway and promoting excessive inflammatory cytokine expression. Furthermore, the application of p38 MAPK inhibitors significantly improves survival in SARS-CoV-2-infected mice by reducing IL-1α and IL-6 levels in the BALF, decreasing neutrophil infiltration, and alleviating interstitial edema (Gu et al., 2021). Selective p38 signaling inhibitors, such as PH-797804 and VX-702, markedly reduce the expression of proinflammatory cytokines IL6, CXCL8, CXCL10, and TNF-α during SARS-CoV-2 infection, while concurrently reducing viral replication and modulating the IFN-mediated antiviral response (Faist et al., 2023).
The NF-κB pathway plays a central role as an immune regulator in COVID-19, contributing to excessive inflammation and cytokine storms. Inhibiting the NF-κB pathway can suppress the release of various proinflammatory cytokines, chemokines, and adhesion molecules, making it an important potential therapeutic target for COVID-19 (Jiang et al., 2022). During the early phase of infection, the SARS-CoV-2 nucleocapsid protein, present in body fluids, activates NF-κB p65 phosphorylation, promotes M1 macrophage polarization and proinflammatory cytokine expression, and induces acute lung injury. The NF-κB inhibitor pyrrolidine dithiocarbamate alleviates the effects of the N protein on acute lung injury in mouse models (Xia et al., 2021). Auranofin, another NF-κB inhibitor, also exerts inhibitory effects on SARS-CoV-2 replication and inflammatory cytokine expression, including IL-6, TNF-α, and IL-1β, in vitro (Cirri et al., 2021; Rothan et al., 2020). Oral administration of auranofin significantly attenuates lung tissue damage, cell infiltration, inflammatory response, and IL-6 production in Syrian hamsters infected with SARS-CoV-2 (Biji et al., 2021). Neufeldt et al. (2022) reported that in human epithelial cells, SARS-CoV-2 triggers an inflammatory immune response via cyclic GMP-AMP synthase (cGAS)-stimulator of IFN genes (STING)-mediated NF-κB activation, which can be attenuated by various STING-targeting drugs, suggesting that cGAS-STING may be a potential target for COVID-19 therapy. However, STING-deficient K18-hACE2 mice do not exhibit altered disease progression after SARS-CoV-2 infection, suggesting that STING deficiency does not impact viral replication or IFN and inflammatory cytokine production (Marino et al., 2023). In contrast, the small-molecule STING agonist diABZI limits SARS-CoV-2 replication in mice and effectively inhibits SARS-CoV-2 infection in multiple viral strains (Li et al., 2021). These findings emphasize the need to verify the effects of immunomodulatory drugs via in vitro and in vivo experiments and conduct comprehensive validations in animal models before advancing with COVID-19 treatment strategies.
CONCLUSIONS
The COVID-19 pandemic continues to pose unprecedented challenges to global health, particularly in terms of prevention and treatment efforts. While activation of the immune system is critical for defending against invading pathogens, it also plays a key role in COVID-19 pathogenesis. Understanding the interactions between SARS-CoV-2 infection and the host immune system is essential for developing effective prevention and treatment strategies. Animal models have been instrumental in studying COVID-19, uncovering the pathological and immunological mechanisms of SARS-CoV-2 infection and accelerating vaccine and drug development. Future research in COVID-19 immunobiology should focus on optimizing the existing animal model library, incorporating specialized animal strains to better represent diverse human populations, and developing relevant tools and methods to address the challenges posed by the high variability and continuous evolution of novel SARS-CoV-2 variants.
Acknowledgments
COMPETING INTERESTS
The authors declare that they have no competing interests.
AUTHORS’ CONTRIBUTIONS
Y.T.Z. conceived the project; H.Y.Z., T.Z.S., and Y.T.Z. wrote the manuscript; Y.T.Z. edited the manuscript. All authors read and approved the final version of the manuscript.
Funding Statement
The work was supported by the National Key Research and Development Program of China (2022YFC2303700, 2021YFC2301300), Yunnan Key Research and Development Program (202303AC100026), National Natural Science Foundation of China (82302002, 82341069), Yunnan Fundamental Research Project (202201AS070047), and Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0490000)
References
- Abdelnabi R, Foo CS, De Jonghe S, et al Molnupiravir inhibits replication of the emerging SARS-CoV-2 variants of concern in a hamster infection model. The Journal of Infectious Diseases. 2021;224(5):749–753. doi: 10.1093/infdis/jiab361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adamo S, Chevrier S, Cervia C, et al Profound dysregulation of T cell homeostasis and function in patients with severe COVID‐19. Allergy. 2021;76(9):2866–2881. doi: 10.1111/all.14866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Afkhami S, D’Agostino MR, Zhang AL, et al. 2022. Respiratory mucosal delivery of next-generation COVID-19 vaccine provides robust protection against both ancestral and variant strains of SARS-CoV-2. Cell, 185 (5): 896–915. e19.
- Aksu M, Kumar P, Güttler T, et al Nanobodies to multiple spike variants and inhalation of nanobody-containing aerosols neutralize SARS-CoV-2 in cell culture and hamsters. Antiviral Research. 2024;221:105778. doi: 10.1016/j.antiviral.2023.105778. [DOI] [PubMed] [Google Scholar]
- A'La R, Wijaya AY, Susilowati H, et al Inactivated SARS-CoV-2 vaccine candidate immunization on non-human primate animal model: B-cell and T-cell responses immune evaluation. Heliyon. 2023;9(7):e18039. doi: 10.1016/j.heliyon.2023.e18039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albornoz EA, Amarilla AA, Modhiran N, et al SARS-CoV-2 drives NLRP3 inflammasome activation in human microglia through spike protein. Molecular Psychiatry. 2023;28(7):2878–2893. doi: 10.1038/s41380-022-01831-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altamimi I, Alabdulkarim IM, Alhumimidi AS, et al Navigating novel uncertainties of COVID-19: the rise of the JN. 1 variant. Cureus. 2024;16(1):e51497. doi: 10.7759/cureus.51497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- André S, Picard M, Cezar R, et al T cell apoptosis characterizes severe Covid-19 disease. Cell Death & Differentiation. 2022;29(8):1486–1499. doi: 10.1038/s41418-022-00936-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antonelli M, Pujol JC, Spector TD, et al Risk of long COVID associated with delta versus omicron variants of SARS-CoV-2. The Lancet. 2022;399(10343):2263–2264. doi: 10.1016/S0140-6736(22)00941-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arish M, Qian W, Narasimhan H, et al COVID‐19 immunopathology: from acute diseases to chronic sequelae. Journal of Medical Virology. 2023;95(1):e28122. doi: 10.1002/jmv.28122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashhurst AS, Johansen MD, Maxwell JWC, et al Mucosal TLR2-activating protein-based vaccination induces potent pulmonary immunity and protection against SARS-CoV-2 in mice. Nature Communications. 2022;13(1):6972. doi: 10.1038/s41467-022-34297-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azkur AK, Akdis M, Azkur D, et al Immune response to SARS‐CoV‐2 and mechanisms of immunopathological changes in COVID‐19. Allergy. 2020;75(7):1564–1581. doi: 10.1111/all.14364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bader SM, Cooney JP, Sheerin D, et al SARS-CoV-2 mouse adaptation selects virulence mutations that cause TNF-driven age-dependent severe disease with human correlates. Proceedings of the National Academy of Sciences of the United States of America. 2023;120(32):e2301689120. doi: 10.1073/pnas.2301689120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bagato O, Balkema-Buschmann A, Todt D, et al Spatiotemporal analysis of SARS-CoV-2 infection reveals an expansive wave of monocyte-derived macrophages associated with vascular damage and virus clearance in hamster lungs. Microbiology Spectrum. 2024;12(1):e0246923. doi: 10.1128/spectrum.02469-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bao LL, Deng W, Huang BY, et al The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature. 2020;583(7818):830–833. doi: 10.1038/s41586-020-2312-y. [DOI] [PubMed] [Google Scholar]
- Baum A, Ajithdoss D, Copin R, et al REGN-COV2 antibodies prevent and treat SARS-CoV-2 infection in rhesus macaques and hamsters. Science. 2020;370(6520):1110–1115. doi: 10.1126/science.abe2402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beckett EL, Phipps S, Starkey MR, et al TLR2, but Not TLR4, is required for effective host defence against Chlamydia respiratory tract infection in early life. PLoS One. 2012;7(6):e39460. doi: 10.1371/journal.pone.0039460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beddingfield BJ, Maness NJ, Fears AC, et al Effective prophylaxis of COVID-19 in rhesus macaques using a combination of two parenterally-administered SARS-CoV-2 neutralizing antibodies. Frontiers in Cellular and Infection Microbiology. 2021;11:753444. doi: 10.3389/fcimb.2021.753444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bednash JS, Kagan VE, Englert JA, et al Syrian hamsters as a model of lung injury with SARS-CoV-2 infection: pathologic, physiologic, and detailed molecular profiling. Translational Research. 2022;240:1–16. doi: 10.1016/j.trsl.2021.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belser JA, Katz JM, Tumpey TM The ferret as a model organism to study influenza A virus infection. Disease Models & Mechanisms. 2011;4(5):575–579. doi: 10.1242/dmm.007823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertzbach LD, Vladimirova D, Dietert K, et al SARS‐CoV‐2 infection of Chinese hamsters (Cricetulus griseus) reproduces COVID‐19 pneumonia in a well‐established small animal model. Transboundary and Emerging Diseases. 2021;68(3):1075–1079. doi: 10.1111/tbed.13837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bi ZF, Hong WQ, Yang JY, et al Animal models for SARS‐CoV‐2 infection and pathology. MedComm. 2021;2(4):548–568. doi: 10.1002/mco2.98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biji A, Khatun O, Swaraj S, et al Identification of COVID-19 prognostic markers and therapeutic targets through meta-analysis and validation of Omics data from nasopharyngeal samples. EBioMedicine. 2021;70:103525. doi: 10.1016/j.ebiom.2021.103525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boby ML, Fearon D, Ferla M, et al Open science discovery of potent noncovalent SARS-CoV-2 main protease inhibitors. Science. 2023;382(6671):eabo7201. doi: 10.1126/science.abo7201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boon ACM, Darling TL, Halfmann PJ, et al Reduced airborne transmission of SARS-CoV-2 BA. 1 omicron virus in Syrian hamsters. PLoS Pathogens. 2022;18(12):e1010970. doi: 10.1371/journal.ppat.1010970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broggi A, Ghosh S, Sposito B, et al Type III interferons disrupt the lung epithelial barrier upon viral recognition. Science. 2020;369(6504):706–712. doi: 10.1126/science.abc3545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broz P, Dixit VM Inflammasomes: mechanism of assembly, regulation and signalling. Nature Reviews Immunology. 2016;16(7):407–420. doi: 10.1038/nri.2016.58. [DOI] [PubMed] [Google Scholar]
- Carabelli AM, Peacock TP, Thorne LG, et al SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nature Reviews Microbiology. 2023;21(3):162–177. doi: 10.1038/s41579-022-00841-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandrashekar A, Liu JY, Martinot AJ, et al SARS-CoV-2 infection protects against rechallenge in rhesus macaques. Science. 2020;369(6505):812–817. doi: 10.1126/science.abc4776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen HB, Xie ZP, Long RX, et al Immunological evaluation of an inactivated SARS-CoV-2 vaccine in rhesus macaques. Molecular Therapy - Methods & Clinical Development. 2021a;23:108–118. doi: 10.1016/j.omtm.2021.08.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen RR, Wang KE, Yu J, et al The spatial and cell-type distribution of SARS-CoV-2 receptor ACE2 in the human and mouse brains. Frontiers in Neurology. 2021b;11:573095. doi: 10.3389/fneur.2020.573095. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen YX, Song WC, Li C, et al COVID-19 mRNA vaccine protects against SARS-CoV-2 Omicron BA. 1 infection in diet-induced obese mice through boosting host innate antiviral responses. EBioMedicine. 2023;89:104485. doi: 10.1016/j.ebiom.2023.104485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z, Yuan YC, Hu QT, et al SARS-CoV-2 immunity in animal models. Cellular & Molecular Immunology. 2024;21(2):119–133. doi: 10.1038/s41423-023-01122-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu H, Chan JFW, Wang YX, et al Comparative replication and immune activation profiles of SARS-CoV-2 and SARS-CoV in human lungs: an ex vivo study with implications for the pathogenesis of COVID-19. Clinical Infectious Diseases. 2020;71(6):1400–1409. doi: 10.1093/cid/ciaa410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu H, Chan JFW, Yuen KY Animal models in SARS-CoV-2 research. Nature Methods. 2022;19(4):392–394. doi: 10.1038/s41592-022-01447-w. [DOI] [PubMed] [Google Scholar]
- Cirri D, Marzo T, Tolbatov I, et al In vitro anti-SARS-CoV-2 activity of selected metal compounds and potential molecular basis for their actions based on computational study. Biomolecules. 2021;11(12):1858. doi: 10.3390/biom11121858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clancy CS, Meade-White K, Shaia C, et al Histopathologic characterization of experimental peracute SARS-CoV-2 infection in the Syrian hamster. Veterinary Sciences. 2023;10(9):536. doi: 10.3390/vetsci10090536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corbett KS, Nason MC, Flach B, et al Immune correlates of protection by mRNA-1273 vaccine against SARS-CoV-2 in nonhuman primates. Science. 2021;373(6561):eabj0299. doi: 10.1126/science.abj0299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daitch V, Yelin D, Awwad M, et al Characteristics of long-COVID among older adults: a cross-sectional study. International Journal of Infectious Diseases. 2022;125:287–293. doi: 10.1016/j.ijid.2022.09.035. [DOI] [PubMed] [Google Scholar]
- Davis HE, Mccorkell L, Vogel JM, et al Long COVID: major findings, mechanisms and recommendations. Nature Reviews Microbiology. 2023;21(3):133–146. doi: 10.1038/s41579-022-00846-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Vries RD, Schmitz KS, Bovier FT, et al Intranasal fusion inhibitory lipopeptide prevents direct-contact SARS-CoV-2 transmission in ferrets. Science. 2021;371(6536):1379–1382. doi: 10.1126/science.abf4896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denton AE, Dooley J, Cinti I, et al Targeting TLR4 during vaccination boosts MAdCAM-1+ lymphoid stromal cell activation and promotes the aged germinal center response. Science Immunology. 2022;7(71):eabk0018. doi: 10.1126/sciimmunol.abk0018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dinnon KH, Leist SR, Okuda K, et al SARS-CoV-2 infection produces chronic pulmonary epithelial and immune cell dysfunction with fibrosis in mice. Science Translational Medicine. 2022;14(664):eabo5070. doi: 10.1126/scitranslmed.abo5070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dinnon KH, Leist SR, Schäfer A, et al A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature. 2020;586(7830):560–566. doi: 10.1038/s41586-020-2708-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du J, Wei LR, Li GL, et al Persistent high percentage of HLA-DR+CD38high CD8+ T cells associated with immune disorder and disease severity of COVID-19. Frontiers in Immunology. 2021;12:735125. doi: 10.3389/fimmu.2021.735125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dutta D, Liu JN, Xiong HG NLRP3 inflammasome activation and SARS-CoV-2-mediated hyperinflammation, cytokine storm and neurological syndromes. International Journal of Physiology, Pathophysiology and Pharmacology. 2022;14(3):138–160. [PMC free article] [PubMed] [Google Scholar]
- Eckstrand CD, Baldwin TJ, Rood KA, et al An outbreak of SARS-CoV-2 with high mortality in mink (Neovison vison) on multiple Utah farms. PLoS Pathogens. 2021;17(11):e1009952. doi: 10.1371/journal.ppat.1009952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efstathiou V, Stefanou MI, Demetriou M, et al Long COVID and neuropsychiatric manifestations (Review) Experimental and Therapeutic Medicine. 2022;23(5):363. doi: 10.3892/etm.2022.11290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elia U, Rotem S, Bar Haim E, et al Lipid nanoparticle RBD-hFc mRNA vaccine protects hACE2 transgenic mice against a lethal SARS-CoV-2 infection. Nano Letters. 2021;21(11):4774–4779. doi: 10.1021/acs.nanolett.1c01284. [DOI] [PubMed] [Google Scholar]
- Estes JD, Wong SW, Brenchley JM Nonhuman primate models of human viral infections. Nature Reviews Immunology. 2018;18(6):390–404. doi: 10.1038/s41577-018-0005-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faist A, Schloer S, Mecate-Zambrano A, et al Inhibition of p38 signaling curtails the SARS-CoV-2 induced inflammatory response but retains the IFN-dependent antiviral defense of the lung epithelial barrier. Antiviral Research. 2023;209:105475. doi: 10.1016/j.antiviral.2022.105475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fajgenbaum DC, June CH Cytokine storm. New England Journal of Medicine. 2020;383(23):2255–2273. doi: 10.1056/NEJMra2026131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng LQ, Wang Q, Shan C, et al An adenovirus-vectored COVID-19 vaccine confers protection from SARS-COV-2 challenge in rhesus macaques. Nature Communications. 2020;11(1):4207. doi: 10.1038/s41467-020-18077-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fernández-de-las-Peñas C, Cancela-Cilleruelo I, Rodríguez-Jiménez J, et al Associated-onset symptoms and Post-COVID-19 symptoms in hospitalized COVID-19 survivors infected with wuhan, alpha or delta SARS-CoV-2 variant. Pathogens. 2022;11(7):725. doi: 10.3390/pathogens11070725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferreira-Gomes M, Kruglov A, Durek P, et al SARS-CoV-2 in severe COVID-19 induces a TGF-β-dominated chronic immune response that does not target itself. Nature Communications. 2021;12(1):1961. doi: 10.1038/s41467-021-22210-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Florence JM, Krupa A, Booshehri LM, et al Inhibiting Bruton’s tyrosine kinase rescues mice from lethal influenza-induced acute lung injury. American Journal of Physiology-Lung Cellular and Molecular Physiology. 2018;315(1):L52–L58. doi: 10.1152/ajplung.00047.2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fomin G, Tabynov K, Islamov R, et al Cytokine response and damages in the lungs of aging Syrian hamsters on a high-fat diet infected with the SARS-CoV-2 virus. Frontiers in Immunology. 2023;14:1223086. doi: 10.3389/fimmu.2023.1223086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fontes-Dantas FL, Fernandes GG, Gutman EG, et al SARS-CoV-2 Spike protein induces TLR4-mediated long-term cognitive dysfunction recapitulating post-COVID-19 syndrome in mice. Cell Reports. 2023;42(3):112189. doi: 10.1016/j.celrep.2023.112189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Francis ME, Goncin U, Kroeker A, et al SARS-CoV-2 infection in the Syrian hamster model causes inflammation as well as type I interferon dysregulation in both respiratory and non-respiratory tissues including the heart and kidney. PLoS Pathogens. 2021;17(7):e1009705. doi: 10.1371/journal.ppat.1009705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frere JJ, Serafini RA, Pryce KD, et al SARS-CoV-2 infection in hamsters and humans results in lasting and unique systemic perturbations after recovery. Science Translational Medicine. 2022;14(664):eabq3059. doi: 10.1126/scitranslmed.abq3059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frontera JA, Sabadia S, Yang D, et al Life stressors significantly impact long-term outcomes and post-acute symptoms 12-months after COVID-19 hospitalization. Journal of the Neurological Sciences. 2022;443:120487. doi: 10.1016/j.jns.2022.120487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gagne M, Flynn BJ, Andrew SF, et al. 2023. Mucosal adenoviral-vectored vaccine boosting durably prevents XBB. 1.16 infection in nonhuman primates. BioRxiv.
- Galani IE, Rovina N, Lampropoulou V, et al Untuned antiviral immunity in COVID-19 revealed by temporal type I/III interferon patterns and flu comparison. Nature Immunology. 2021;22(1):32–40. doi: 10.1038/s41590-020-00840-x. [DOI] [PubMed] [Google Scholar]
- Gao S, Zhang LL ACE2 partially dictates the host range and tropism of SARS-CoV-2. Computational and Structural Biotechnology Journal. 2020;18:4040–4047. doi: 10.1016/j.csbj.2020.11.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gary EN, Warner BM, Parzych EM, et al A novel mouse AAV6 hACE2 transduction model of wild-type SARS-CoV-2 infection studied using synDNA immunogens. iScience. 2021;24(7):102699. doi: 10.1016/j.isci.2021.102699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georg P, Astaburuaga-García R, Bonaguro L, et al. 2022. Complement activation induces excessive T cell cytotoxicity in severe COVID-19. Cell, 185 (3): 493–512. e25.
- Glazkova DV, Bogoslovskaya EV, Urusov FA, et al Generation of SARS-CoV-2 mouse model by transient expression of the human ACE2 gene mediated by intranasal administration of AAV-hACE2. Molecular Biology. 2022;56(5):705–712. doi: 10.1134/S0026893322050065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grimes JM, Grimes KV p38 MAPK inhibition: a promising therapeutic approach for COVID-19. Journal of Molecular and Cellular Cardiology. 2020;144:63–65. doi: 10.1016/j.yjmcc.2020.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruber AD, Firsching TC, Trimpert J, et al Hamster models of COVID-19 pneumonia reviewed: how human can they be? Veterinary Pathology. 2022;59(4):528–545. doi: 10.1177/03009858211057197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu HJ, Chen Q, Yang G, et al Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science. 2020;369(6511):1603–1607. doi: 10.1126/science.abc4730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu TX, Zhao SM, Jin GG, et al Cytokine signature induced by SARS-CoV-2 spike protein in a mouse model. Frontiers in Immunology. 2021;11:621441. doi: 10.3389/fimmu.2020.621441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guebre-Xabier M, Patel N, Tian JH, et al NVX-CoV2373 vaccine protects cynomolgus macaque upper and lower airways against SARS-CoV-2 challenge. Vaccine. 2020;38(50):7892–7896. doi: 10.1016/j.vaccine.2020.10.064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haabeth OaW, Lohmeyer JJK, Sallets A, et al An mRNA SARS-CoV-2 vaccine employing charge-altering releasable transporters with a TLR-9 agonist induces neutralizing antibodies and T cell memory. ACS Central Science. 2021;7(7):1191–1204. doi: 10.1021/acscentsci.1c00361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hadjadj J, Yatim N, Barnabei L, et al Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science. 2020;369(6504):718–724. doi: 10.1126/science.abc6027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hakim AD, Awili M, O'Neal HR, et al Efficacy and safety of MAS825 (anti-IL-1β/IL-18) in COVID-19 patients with pneumonia and impaired respiratory function. Clinical and Experimental Immunology. 2023;213(3):265–275. doi: 10.1093/cei/uxad065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halfmann PJ, Iida S, Iwatsuki-Horimoto K, et al SARS-CoV-2 Omicron virus causes attenuated disease in mice and hamsters. Nature. 2022;603(7902):687–692. doi: 10.1038/s41586-022-04441-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halfmann PJ, Uraki R, Kuroda M, et al Transmission and re-infection of Omicron variant XBB. 1.5 in hamsters. eBioMedicine. 2023;93:104677. doi: 10.1016/j.ebiom.2023.104677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall FC, Cheriyan J, Cope AP, et al Efficacy and safety of baricitinib or ravulizumab in adult patients with severe COVID-19 (TACTIC-R): a randomised, parallel-arm, open-label, phase 4 trial. The Lancet Respiratory Medicine. 2023;11(12):1064–1074. doi: 10.1016/S2213-2600(23)00376-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Handley A, Ryan KA, Davies ER, et al SARS-CoV-2 disease severity in the golden Syrian hamster model of infection is related to the volume of intranasal inoculum. Viruses. 2023;15(3):748. doi: 10.3390/v15030748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hasenkrug KJ, Feldmann F, Myers L, et al Recovery from acute SARS-CoV-2 infection and development of anamnestic immune responses in T cell-depleted rhesus macaques. mBio. 2021;12(4):e0150321. doi: 10.1128/mBio.01503-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hendriks RW, Yuvaraj S, Kil LP Targeting Bruton's tyrosine kinase in B cell malignancies. Nature Reviews Cancer. 2014;14(4):219–232. doi: 10.1038/nrc3702. [DOI] [PubMed] [Google Scholar]
- Herder V, Mendonca DC, Upfold N, et al. 2023. The SARS-CoV-2 Omicron sub-variant BA. 2.86 is attenuated in hamsters. BioRxiv.
- Hewitt JA, Lutz C, Florence WC, et al ACTIVating resources for the COVID-19 pandemic: in vivo models for vaccines and therapeutics. Cell Host & Microbe. 2020;28(5):646–659. doi: 10.1016/j.chom.2020.09.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Higgins CA, Nilsson-Payant BE, Bonaventure B, et al SARS-CoV-2 hijacks p38β/MAPK11 to promote virus replication. mBio. 2023;14(4):e0100723. doi: 10.1128/mbio.01007-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoang TN, Pino M, Boddapati AK, et al. 2021. Baricitinib treatment resolves lower-airway macrophage inflammation and neutrophil recruitment in SARS-CoV-2-infected rhesus macaques. Cell, 184 (2): 460–475. e21.
- Horiuchi S, Oishi K, Carrau L, et al Immune memory from SARS-CoV-2 infection in hamsters provides variant-independent protection but still allows virus transmission. Science Immunology. 2021;6(66):eabm3131. doi: 10.1126/sciimmunol.abm3131. [DOI] [PubMed] [Google Scholar]
- Huang LX, Li X, Gu XY, et al Health outcomes in people 2 years after surviving hospitalisation with COVID-19: a longitudinal cohort study. The Lancet Respiratory Medicine. 2022;10(9):863–876. doi: 10.1016/S2213-2600(22)00126-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imai M, Iwatsuki-Horimoto K, Hatta M, et al Syrian hamsters as a small animal model for SARS-CoV-2 infection and countermeasure development. Proceedings of the National Academy of Sciences of the United States of America. 2020;117(28):16587–16595. doi: 10.1073/pnas.2009799117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ireland RE, Davies CD, Keyser E, et al Histopathological and immunological findings in the common marmoset following exposure to aerosolized SARS-CoV-2. Viruses. 2022;14(7):1580. doi: 10.3390/v14071580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Israelow B, Mao TY, Klein J, et al Adaptive immune determinants of viral clearance and protection in mouse models of SARS-CoV-2. Science Immunology. 2021;6(64):eabl4509. doi: 10.1126/sciimmunol.abl4509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Israelow B, Song E, Mao TY, et al Mouse model of SARS-CoV-2 reveals inflammatory role of type I interferon signaling. Journal of Experimental Medicine. 2020;217(12):e20201241. doi: 10.1084/jem.20201241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacob-Dolan C, Feldman J, McMahan K, et al Coronavirus-specific antibody cross reactivity in Rhesus Macaques following SARS-CoV-2 vaccination and infection. Journal of Virology. 2021;95(11):e00117–21. doi: 10.1128/JVI.00117-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jafarzadeh A, Jafarzadeh S, Nozari P, et al Lymphopenia an important immunological abnormality in patients with COVID‐19: possible mechanisms. Scandinavian Journal of Immunology. 2020;93(2):e12967. doi: 10.1111/sji.12967. [DOI] [PubMed] [Google Scholar]
- Jansen EB, Orvold SN, Swan CL, et al After the virus has cleared—Can preclinical models be employed for Long COVID research? PLoS Pathogens. 2022;18(9):e1010741. doi: 10.1371/journal.ppat.1010741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jeong JH, Chokkakula S, Min SC, et al Combination therapy with nirmatrelvir and molnupiravir improves the survival of SARS-CoV-2 infected mice. Antiviral Research. 2022;208:105430. doi: 10.1016/j.antiviral.2022.105430. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang YJ, Zhao TM, Zhou XY, et al Inflammatory pathways in COVID‐19: mechanism and therapeutic interventions. MedComm. 2022;3(3):e154. doi: 10.1002/mco2.154. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jimenez-Guardeño JM, Nieto-Torres JL, Dediego ML, et al The PDZ-binding motif of severe acute respiratory syndrome coronavirus envelope protein is a determinant of viral pathogenesis. PLoS Pathogens. 2014;10(8):e1004320. doi: 10.1371/journal.ppat.1004320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jin JY, Wang XW, Li YZ, et al Weak SARS-CoV-2-specific responses of TIGIT-expressing CD8+ T cells in people living with HIV after a third dose of a SARS-CoV-2 inactivated vaccine. Chinese Medical Journal. 2023;136(24):2938–2947. doi: 10.1097/CM9.0000000000002926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson DM, Brasel T, Massey S, et al Evaluation of molnupiravir (EIDD-2801) efficacy against SARS-CoV-2 in the rhesus macaque model. Antiviral Research. 2023;209:105492. doi: 10.1016/j.antiviral.2022.105492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kane Y, Wong G, Gao GF Animal models, zoonotic reservoirs, and cross-species transmission of emerging human-infecting coronaviruses. Annual Review of Animal Biosciences. 2023;11:1–31. doi: 10.1146/annurev-animal-020420-025011. [DOI] [PubMed] [Google Scholar]
- Karki R, Sharma BR, Tuladhar S, et al. 2021. Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell, 184 (1): 149–168. e17.
- Kasuga Y, Zhu BH, Jang KJ, et al Innate immune sensing of coronavirus and viral evasion strategies. Experimental & Molecular Medicine. 2021;53(5):723–736. doi: 10.1038/s12276-021-00602-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kibler KV, Szczerba M, Lake D, et al Intranasal immunization with a vaccinia virus vaccine vector expressing pre-fusion stabilized SARS-CoV-2 spike fully protected mice against lethal challenge with the heavily mutated mouse-adapted SARS2-N501YMA30 strain of SARS-CoV-2. Vaccines. 2022;10(8):1172. doi: 10.3390/vaccines10081172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kifle ZD Bruton tyrosine kinase inhibitors as potential therapeutic agents for COVID-19: a review. Metabolism Open. 2021;11:100116. doi: 10.1016/j.metop.2021.100116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim YI, Casel MaB, Choi YK Transmissibility and pathogenicity of SARS-CoV-2 variants in animal models. Journal of Microbiology. 2022;60(3):255–267. doi: 10.1007/s12275-022-2033-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim YI, Kim SG, Kim SM, et al. 2020. Infection and rapid transmission of SARS-CoV-2 in ferrets. Cell Host & Microbe, 27 (5): 704–709. e2.
- Klein J, Wood J, Jaycox JR, et al Distinguishing features of long COVID identified through immune profiling. Nature. 2023;623(7985):139–148. doi: 10.1038/s41586-023-06651-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knight AC, Montgomery SA, Fletcher CA, et al Mouse models for the study of SARS-CoV-2 infection. Comparative Medicine. 2021;71(5):383–397. doi: 10.30802/AALAS-CM-21-000031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kosyreva A, Dzhalilova D, Lokhonina A, et al The role of macrophages in the pathogenesis of SARS-CoV-2-associated acute respiratory distress syndrome. Frontiers in Immunology. 2021;12:682871. doi: 10.3389/fimmu.2021.682871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreft IC, Winiarczyk RRA, Tanis FJ, et al Absence of COVID-19-associated changes in plasma coagulation proteins and pulmonary thrombosis in the ferret model. Thrombosis Research. 2022;210:6–11. doi: 10.1016/j.thromres.2021.12.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuang YQ, Tang X, Liu FL, et al Genotyping of TRIM5 locus in northern pig-tailed macaques (Macaca leonina), a primate species susceptible to Human Immunodeficiency Virus type 1 infection. Retrovirology. 2009;6(1):58. doi: 10.1186/1742-4690-6-58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langel SN, Johnson S, Martinez CI, et al Adenovirus type 5 SARS-CoV-2 vaccines delivered orally or intranasally reduced disease severity and transmission in a hamster model. Science Translational Medicine. 2022;14(658):eabn6868. doi: 10.1126/scitranslmed.abn6868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CM, Oh JE Resident memory B cells in barrier tissues. Frontiers in Immunology. 2022;13:953088. doi: 10.3389/fimmu.2022.953088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JS, Koh JY, Yi K, et al Single-cell transcriptome of bronchoalveolar lavage fluid reveals sequential change of macrophages during SARS-CoV-2 infection in ferrets. Nature Communications. 2021;12(1):4567. doi: 10.1038/s41467-021-24807-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee JS, Shin EC The type I interferon response in COVID-19: implications for treatment. Nature Reviews Immunology. 2020;20(10):585–586. doi: 10.1038/s41577-020-00429-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leist SR, Dinnon KH, Schäfer A, et al. 2020. A mouse-adapted SARS-CoV-2 induces acute lung injury and mortality in standard laboratory mice. Cell, 183 (4): 1070–1085. e12.
- Li MH, Ferretti M, Ying BL, et al Pharmacological activation of STING blocks SARS-CoV-2 infection. Science Immunology. 2021;6(59):eabi9007. doi: 10.1126/sciimmunol.abi9007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Q, Dang C, Wang LH Neuroinflammation in mild respiratory COVID-19: insights into cognitive impairment in milder cases. Military Medical Research. 2022;9(1):72. doi: 10.1186/s40779-022-00431-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang SX, Bao CL, Yang Z, et al SARS-CoV-2 spike protein induces IL-18-mediated cardiopulmonary inflammation via reduced mitophagy. Signal Transduction and Targeted Therapy. 2023;8(1):108. doi: 10.1038/s41392-023-01368-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liang Y, Li H, Li J, et al Role of neutrophil chemoattractant CXCL5 in SARS-CoV-2 infection-induced lung inflammatory innate immune response in an in vivo hACE2 transfection mouse model. Zoological Research. 2020;41(6):621–631. doi: 10.24272/j.issn.2095-8137.2020.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao MF, Liu Y, Yuan J, et al Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nature Medicine. 2020;26(6):842–844. doi: 10.1038/s41591-020-0901-9. [DOI] [PubMed] [Google Scholar]
- Linsky TW, Vergara R, Codina N, et al De novo design of potent and resilient hACE2 decoys to neutralize SARS-CoV-2. Science. 2020;370(6521):1208–1214. doi: 10.1126/science.abe0075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu DL, Chen CK, Chen DB, et al Mouse models susceptible to HCoV-229E and HCoV-NL63 and cross protection from challenge with SARS-CoV-2. Proceedings of the National Academy of Sciences of the United States of America. 2023a;120(4):e2202820120. doi: 10.1073/pnas.2202820120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu R, Wang Y, Li J, et al Decreased T cell populations contribute to the increased severity of COVID-19. Clinica Chimica Acta. 2020a;508:110–114. doi: 10.1016/j.cca.2020.05.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Gu XY, Li HB, et al Mechanisms of long COVID: an updated review. Chinese Medical Journal Pulmonary and Critical Care Medicine. 2023b;1(4):231–240. doi: 10.1016/j.pccm.2023.10.003. [DOI] [Google Scholar]
- Liu ZM, Long W, Tu MQ, et al Lymphocyte subset (CD4+, CD8+) counts reflect the severity of infection and predict the clinical outcomes in patients with COVID-19. Journal of Infection. 2020b;81(2):318–356. doi: 10.1016/j.jinf.2020.03.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu ZM, Yang MH, Yu K, et al Toll-like receptor (TLRs) agonists and antagonists for COVID-19 treatments. Frontiers in Pharmacology. 2022;13:989664. doi: 10.3389/fphar.2022.989664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loretelli C, Abdelsalam A, D’addio F, et al PD-1 blockade counteracts post–COVID-19 immune abnormalities and stimulates the anti–SARS-CoV-2 immune response. JCI Insight. 2021;6(24):e146701. doi: 10.1172/jci.insight.146701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu SY, Zhao Y, Yu WH, et al Comparison of nonhuman primates identified the suitable model for COVID-19. Signal Transduction and Targeted Therapy. 2020;5(1):157. doi: 10.1038/s41392-020-00269-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu Y, Shen F, He WQ, et al HR121 targeting HR2 domain in S2 subunit of spike protein can serve as a broad-spectrum SARS-CoV-2 inhibitor via intranasal administration. Acta Pharmaceutica Sinica B. 2023;13(8):3339–3351. doi: 10.1016/j.apsb.2023.05.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Machado RRG, Walker JL, Scharton D, et al Immunogenicity and efficacy of vaccine boosters against SARS-CoV-2 Omicron subvariant BA. 5 in male Syrian hamsters. Nature Communications. 2023;14(1):4260. doi: 10.1038/s41467-023-40033-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahdy MaA, Younis W, Ewaida Z An overview of SARS-CoV-2 and animal infection. Frontiers in Veterinary Science. 2020;7:596391. doi: 10.3389/fvets.2020.596391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Major J, Crotta S, Llorian M, et al Type I and III interferons disrupt lung epithelial repair during recovery from viral infection. Science. 2020;369(6504):712–717. doi: 10.1126/science.abc2061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marino G, Zhang BC, Schmitz A, et al STING is redundant for host defense and pathology of COVID-19-like disease in mice. Life Science Alliance. 2023;6(8):e202301997. doi: 10.26508/lsa.202301997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marx S, Kümmerer BM, Grützner C, et al RIG-I-induced innate antiviral immunity protects mice from lethal SARS-CoV-2 infection. Molecular Therapy - Nucleic Acids. 2022;27:1225–1234. doi: 10.1016/j.omtn.2022.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masters AR, Jellison ER, Puddington L, et al Attrition of T cell zone fibroblastic reticular cell number and function in aged spleens. Immunohorizons. 2018;2(5):155–163. doi: 10.4049/immunohorizons.1700062. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCray Jr PB, Pewe L, Wohlford-Lenane C, et al Lethal infection of K18-hACE2 mice infected with severe acute respiratory syndrome coronavirus. Journal of Virology. 2007;81(2):813–821. doi: 10.1128/JVI.02012-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGonagle D, Sharif K, O'regan A, et al The role of cytokines including interleukin-6 in COVID-19 induced pneumonia and macrophage activation syndrome-like disease. Autoimmunity Reviews. 2020;19(6):102537. doi: 10.1016/j.autrev.2020.102537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMahan K, Wegmann F, Aid M, et al Mucosal boosting enhances vaccine protection against SARS-CoV-2 in macaques. Nature. 2024;626(7998):385–391. doi: 10.1038/s41586-023-06951-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMahan K, Yu JY, Mercado NB, et al Correlates of protection against SARS-CoV-2 in rhesus macaques. Nature. 2021;590(7847):630–634. doi: 10.1038/s41586-020-03041-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meehan GR, Herder V, Allan J, et al Phenotyping the virulence of SARS-CoV-2 variants in hamsters by digital pathology and machine learning. PLoS Pathogens. 2023;19(11):e1011589. doi: 10.1371/journal.ppat.1011589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mehta P, Mcauley DF, Brown M, et al COVID-19: consider cytokine storm syndromes and immunosuppression. The Lancet. 2020;395(10229):1033–1034. doi: 10.1016/S0140-6736(20)30628-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melin AD, Janiak MC, Marrone F, et al Comparative ACE2 variation and primate COVID-19 risk. Communications Biology. 2020;3(1):641. doi: 10.1038/s42003-020-01370-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merad M, Blish CA, Sallusto F, et al The immunology and immunopathology of COVID-19. Science. 2022;375(6585):1122–1127. doi: 10.1126/science.abm8108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merle-Nguyen L, Ando-Grard O, Bourgon C, et al Early corticosteroid treatment enhances recovery from SARS-CoV-2 induced loss of smell in hamster. Brain, Behavior, and Immunity. 2024;118:78–89. doi: 10.1016/j.bbi.2024.02.020. [DOI] [PubMed] [Google Scholar]
- Montazersaheb S, Hosseiniyan Khatibi SM, Hejazi MS, et al COVID-19 infection: an overview on cytokine storm and related interventions. Virology Journal. 2022;19(1):92. doi: 10.1186/s12985-022-01814-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morgan MS, Yan KX, Le TT, et al Monoclonal antibodies specific for SARS-CoV-2 spike protein suitable for multiple applications for current variants of concern. Viruses. 2022;15(1):139. doi: 10.3390/v15010139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mulka KR, Beck SE, Solis CV, et al Progression and resolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection in golden syrian hamsters. The American Journal of Pathology. 2022;192(2):195–207. doi: 10.1016/j.ajpath.2021.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muñoz-Fontela C, Dowling WE, Funnell SGP, et al Animal models for COVID-19. Nature. 2020;586(7830):509–515. doi: 10.1038/s41586-020-2787-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mykytyn AZ, Lamers MM, Okba NMA, et al Susceptibility of rabbits to SARS-CoV-2. Emerging Microbes & Infections. 2021;10(1):1–7. doi: 10.1080/22221751.2020.1868951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neufeldt CJ, Cerikan B, Cortese M, et al SARS-CoV-2 infection induces a pro-inflammatory cytokine response through cGAS-STING and NF-κB. Communications Biology. 2022;5(1):45. doi: 10.1038/s42003-021-02983-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni WT, Yang XW, Yang DQ, et al Role of angiotensin-converting enzyme 2 (ACE2) in COVID-19. Critical Care. 2020;24(1):422. doi: 10.1186/s13054-020-03120-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oh DQP, Grothe I, Lukaß H, et al Monitoring and immunogenicity of SARS-CoV-2 vaccination of laboratory rhesus monkeys (Macaca mulatta) Scientific Reports. 2023;13(1):3274. doi: 10.1038/s41598-023-30473-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oh JE, Song E, Moriyama M, et al Intranasal priming induces local lung-resident B cell populations that secrete protective mucosal antiviral IgA. Science Immunology. 2021;6(66):eabj5129. doi: 10.1126/sciimmunol.abj5129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oladunni FS, Park JG, Pino PA, et al Lethality of SARS-CoV-2 infection in K18 human angiotensin-converting enzyme 2 transgenic mice. Nature Communications. 2020;11(1):6122. doi: 10.1038/s41467-020-19891-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Neill LAJ, Golenbock D, Bowie AG The history of Toll-like receptors — redefining innate immunity. Nature Reviews Immunology. 2013;13(6):453–460. doi: 10.1038/nri3446. [DOI] [PubMed] [Google Scholar]
- Paidas MJ, Cosio DS, Ali S, et al Long-term sequelae of COVID-19 in experimental mice. Molecular Neurobiology. 2022;59(10):5970–5986. doi: 10.1007/s12035-022-02932-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel N, Trost JF, Guebre-Xabier M, et al XBB. 1.5 spike protein COVID-19 vaccine induces broadly neutralizing and cellular immune responses against EG. 5.1 and emerging XBB variants. Scientific Reports. 2023;13(1):19176. doi: 10.1038/s41598-023-46025-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Philippens IHCHM, Böszörményi KP, Wubben JAM, et al Brain inflammation and intracellular α-Synuclein aggregates in macaques after SARS-CoV-2 infection. Viruses. 2022;14(4):776. doi: 10.3390/v14040776. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Port JR, Yinda CK, Riopelle JC, et al Infection- or AZD1222 vaccine-mediated immunity reduces SARS-CoV-2 transmission but increases Omicron competitiveness in hamsters. Nature Communications. 2023;14(1):6592. doi: 10.1038/s41467-023-42346-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prenafeta A, Bech-Sàbat G, Moros A, et al Preclinical evaluation of PHH-1V vaccine candidate against SARS-CoV-2 in non-human primates. iScience. 2023;26(7):107224. doi: 10.1016/j.isci.2023.107224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi FR, Qian S, Zhang SY, et al Single cell RNA sequencing of 13 human tissues identify cell types and receptors of human coronaviruses. Biochemical and Biophysical Research Communications. 2020;526(1):135–140. doi: 10.1016/j.bbrc.2020.03.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabdano SO, Ruzanova EA, Pletyukhina IV, et al Immunogenicity and in vivo protective effects of recombinant nucleocapsid-based SARS-CoV-2 vaccine convacell®. Vaccines. 2023;11(4):874. doi: 10.3390/vaccines11040874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ragab D, Salah Eldin H, Taeimah M, et al The COVID-19 cytokine storm; What we know so far. Frontiers in Immunology. 2020;11:1446. doi: 10.3389/fimmu.2020.01446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rao HN, Song XMT, Lei JT, et al Ibrutinib prevents acute lung injury via multi-targeting BTK, FLT3 and EGFR in mice. International Journal of Molecular Sciences. 2022;23(21):13478. doi: 10.3390/ijms232113478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ratajczak MZ, Kucia M SARS-CoV-2 infection and overactivation of Nlrp3 inflammasome as a trigger of cytokine “storm” and risk factor for damage of hematopoietic stem cells. Leukemia. 2020;34(7):1726–1729. doi: 10.1038/s41375-020-0887-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravid JD, Leiva O, Chitalia VC Janus kinase signaling pathway and its role in COVID-19 inflammatory, vascular, and thrombotic manifestations. Cells. 2022;11(2):306. doi: 10.3390/cells11020306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rein L, Calero K, Shah R, et al Randomized phase 3 trial of ruxolitinib for COVID-19-associated acute respiratory distress syndrome. Critical Care Medicine. 2022;50(12):1701–1713. doi: 10.1097/CCM.0000000000005682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riggioni C, Comberiati P, Giovannini M, et al A compendium answering 150 questions on COVID‐19 and SARS‐CoV‐2. Allergy. 2020;75(10):2503–2541. doi: 10.1111/all.14449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roessler M, Tesch F, Batram M, et al Post-COVID-19-associated morbidity in children, adolescents, and adults: a matched cohort study including more than 157, 000 individuals with COVID-19 in Germany. PLoS Medicine. 2022;19(11):e1004122. doi: 10.1371/journal.pmed.1004122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roschewski M, Lionakis MS, Sharman JP, et al Inhibition of Bruton tyrosine kinase in patients with severe COVID-19. Science Immunology. 2020;5(48):eabd0110. doi: 10.1126/sciimmunol.abd0110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenke K, Lewis MC, Feldmann F, et al Combined molnupiravir-nirmatrelvir treatment improves the inhibitory effect on SARS-CoV-2 in macaques. JCI Insight. 2023;8(4):e166485. doi: 10.1172/jci.insight.166485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothan HA, Stone S, Natekar J, et al The FDA-approved gold drug auranofin inhibits novel coronavirus (SARS-COV-2) replication and attenuates inflammation in human cells. Virology. 2020;547:7–11. doi: 10.1016/j.virol.2020.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubsamen R, Burkholz S, Massey C, et al Anti-IL-6 Versus Anti-IL-6R blocking antibodies to treat acute ebola infection in BALB/c mice: potential implications for treating cytokine release syndrome. Frontiers in Pharmacology. 2020;11:574703. doi: 10.3389/fphar.2020.574703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ryan FJ, Hope CM, Masavuli MG, et al Long-term perturbation of the peripheral immune system months after SARS-CoV-2 infection. BMC Medicine. 2022;20(1):26. doi: 10.1186/s12916-021-02228-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santopaolo M, Gregorova M, Hamilton F, et al Prolonged T-cell activation and long COVID symptoms independently associate with severe COVID-19 at 3 months. eLife. 2023;12:e85009. doi: 10.7554/eLife.85009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saravanan UB, Namachivayam M, Jeewon R, et al Animal models for SARS-CoV-2 and SARS-CoV-1 pathogenesis, transmission and therapeutic evaluation. World Journal of Virology. 2022;11(1):40–56. doi: 10.5501/wjv.v11.i1.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sasaki M, Tabata K, Kishimoto M, et al S-217622, a SARS-CoV-2 main protease inhibitor, decreases viral load and ameliorates COVID-19 severity in hamsters. Science Translational Medicine. 2023;15(679):eabq4064. doi: 10.1126/scitranslmed.abq4064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt C COVID-19 long haulers. Nature Biotechnology. 2021;39(8):908–913. doi: 10.1038/s41587-021-00984-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schramm CA, Moon D, Peyton L, et al Interaction dynamics between innate and adaptive immune cells responding to SARS-CoV-2 vaccination in non-human primates. Nature Communications. 2023;14(1):7961. doi: 10.1038/s41467-023-43420-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seehusen F, Clark JJ, Sharma P, et al Neuroinvasion and neurotropism by SARS-CoV-2 variants in the K18-hACE2 mouse. Viruses. 2022;14(5):1020. doi: 10.3390/v14051020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sefik E, Israelow B, Mirza H, et al A humanized mouse model of chronic COVID-19. Nature Biotechnology. 2022;40(6):906–920. doi: 10.1038/s41587-021-01155-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekine T, Perez-Potti A, Rivera-Ballesteros O, et al. 2020. Robust T cell immunity in convalescent individuals with asymptomatic or mild COVID-19. Cell, 183 (1): 158–168. e14.
- Shan C, Yao YF, Yang XL, et al. 2020. Infection with novel coronavirus (SARS-CoV-2) causes pneumonia in Rhesus macaques. Cell Research, 30 (8): 670–677.
- Sheahan T, Morrison TE, Funkhouser W, et al MyD88 is required for protection from lethal infection with a mouse-adapted SARS-CoV. PLoS Pathogens. 2008;4(12):e1000240. doi: 10.1371/journal.ppat.1000240. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi JZ, Wen ZY, Zhong GX, et al Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2. Science. 2020;368(6494):1016–1020. doi: 10.1126/science.abb7015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi ZL, Hu ZH A review of studies on animal reservoirs of the SARS coronavirus. Virus Research. 2008;133(1):74–87. doi: 10.1016/j.virusres.2007.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shou SY, Liu MH, Yang Y, et al Animal models for COVID-19: hamsters, mouse, ferret, mink, tree shrew, and non-human primates. Frontiers in Microbiology. 2021;12:626553. doi: 10.3389/fmicb.2021.626553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sia SF, Yan LM, Chin AWH, et al Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature. 2020;583(7818):834–838. doi: 10.1038/s41586-020-2342-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singh DK, Singh B, Ganatra SR, et al Responses to acute infection with SARS-CoV-2 in the lungs of rhesus macaques, baboons and marmosets. Nature Microbiology. 2020;6(1):73–86. doi: 10.1038/s41564-020-00841-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song TZ, Zheng HY, Han JB, et al Delayed severe cytokine storm and immune cell infiltration in SARS-CoV-2-infected aged Chinese rhesus macaques. Zoological Research. 2020;41(5):503–516. doi: 10.24272/j.issn.2095-8137.2020.202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song TZ, Zheng HY, Han JB, et al Northern pig-tailed macaques (Macaca leonina) infected with SARS-CoV-2 show rapid viral clearance and persistent immune response. Zoological Research. 2021;42(3):350–353. doi: 10.24272/j.issn.2095-8137.2020.334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Speranza E, Purushotham JN, Port JR, et al Age-related differences in immune dynamics during SARS-CoV-2 infection in rhesus macaques. Life Science Alliance. 2022;5(4):e202101314. doi: 10.26508/lsa.202101314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streblow DN, Hirsch AJ, Stanton JJ, et al Aerosol delivery of SARS-CoV-2 human monoclonal antibodies in macaques limits viral replication and lung pathology. Nature Communications. 2023;14(1):7062. doi: 10.1038/s41467-023-42440-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Nirantharakumar K, Hughes S, et al Symptoms and risk factors for long COVID in non-hospitalized adults. Nature Medicine. 2022;28(8):1706–1714. doi: 10.1038/s41591-022-01909-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sui YJ, Li JP, Venzon DJ, et al SARS-CoV-2 spike protein suppresses ACE2 and type I interferon expression in primary cells from macaque lung bronchoalveolar lavage. Frontiers in Immunology. 2021a;12:658428. doi: 10.3389/fimmu.2021.658428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sui YJ, Li JP, Zhang RS, et al Protection against SARS-CoV-2 infection by a mucosal vaccine in rhesus macaques. JCI Insight. 2021b;6(10):e148494. doi: 10.1172/jci.insight.148494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J, Zhuang Z, Zheng J, et al. 2020a. Generation of a broadly useful model for COVID-19 pathogenesis, vaccination, and treatment. Cell, 182 (3): 734–743. e5.
- Sun SY, Cai YQ, Song TZ, et al Interferon-armed RBD dimer enhances the immunogenicity of RBD for sterilizing immunity against SARS-CoV-2. Cell Research. 2021;31(9):1011–1023. doi: 10.1038/s41422-021-00531-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun SH, Chen Q, Gu HJ, et al. 2020b. A mouse model of SARS-CoV-2 infection and pathogenesis. Cell Host & Microbe, 28 (1): 124–133. e4.
- Tada T, Peng JY, Dcosta BM, et al Single-epitope T cell–based vaccine protects against SARS-CoV-2 infection in a preclinical animal model. JCI Insight. 2023;8(7):e167306. doi: 10.1172/jci.insight.167306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamir H, Melamed S, Erez N, et al Induction of innate immune response by TLR3 agonist protects mice against SARS-CoV-2 infection. Viruses. 2022;14(2):189. doi: 10.3390/v14020189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura T, Irie T, Deguchi S, et al Virological characteristics of the SARS-CoV-2 Omicron XBB. 1.5 variant. Nature Communications. 2024;15(1):1176. doi: 10.1038/s41467-024-45274-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura T, Ito J, Uriu K, et al Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants. Nature Communications. 2023;14(1):2800. doi: 10.1038/s41467-023-38435-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tan JX, Li YQ Immune checkpoint alterations and their blockade in COVID-19 patients. Blood Science. 2022;4(4):192–198. doi: 10.1097/BS9.0000000000000132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang JY, Zeng C, Cox TM, et al Respiratory mucosal immunity against SARS-CoV-2 after mRNA vaccination. Science Immunology. 2022;7(76):eadd4853. doi: 10.1126/sciimmunol.add4853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tao KM, Tzou PL, Nouhin J, et al SARS-CoV-2 antiviral therapy. Clinical Microbiology Reviews. 2021;34(4):e0010921. doi: 10.1128/CMR.00109-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tokunoh N, Tamiya S, Watanabe M, et al A nasal vaccine with inactivated whole-virion elicits protective mucosal immunity against SARS-CoV-2 in mice. Frontiers in Immunology. 2023;14:1224634. doi: 10.3389/fimmu.2023.1224634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Totura AL, Whitmore A, Agnihothram S, et al Toll-like receptor 3 signaling via TRIF contributes to a protective innate immune response to severe acute respiratory syndrome coronavirus infection. mBio. 2015;6(3):e00638–15. doi: 10.1128/mBio.00638-15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treon SP, Castillo JJ, Skarbnik AP, et al The BTK inhibitor ibrutinib may protect against pulmonary injury in COVID-19–infected patients. Blood. 2020;135(21):1912–1915. doi: 10.1182/blood.2020006288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trimpert J, Vladimirova D, Dietert K, et al The roborovski dwarf hamster is a highly susceptible model for a rapid and fatal course of SARS-CoV-2 infection. Cell Reports. 2020;33(10):108488. doi: 10.1016/j.celrep.2020.108488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tseng CTK, Huang C, Newman P, et al Severe acute respiratory syndrome coronavirus infection of mice transgenic for the human angiotensin-converting enzyme 2 virus receptor. Journal of Virology. 2007;81(3):1162–1173. doi: 10.1128/JVI.01702-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Upadhyay AA, Viox EG, Hoang TN, et al TREM2+ and interstitial-like macrophages orchestrate airway inflammation in SARS-CoV-2 infection in rhesus macaques. Nature Communications. 2023;14(1):1914. doi: 10.1038/s41467-023-37425-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uraki R, Kiso M, Imai M, et al Therapeutic efficacy of monoclonal antibodies and antivirals against SARS-CoV-2 Omicron BA. 1 in Syrian hamsters. Nature Microbiology. 2022;7(8):1252–1258. doi: 10.1038/s41564-022-01170-4. [DOI] [PubMed] [Google Scholar]
- Uraki R, Kiso M, Iwatsuki Horimoto K, et al. 2023. Characterization of a SARS-CoV-2 EG. 5.1 clinical isolate in vitro and in vivo. Cell Reports, 42 (12): 113580.
- Van Doremalen N, Purushotham JN, Schulz JE, et al Intranasal ChAdOx1 nCoV-19/AZD1222 vaccination reduces viral shedding after SARS-CoV-2 D614G challenge in preclinical models. Science Translational Medicine. 2021;13(607):eabh0755. doi: 10.1126/scitranslmed.abh0755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Doremalen N, Singh M, Saturday TA, et al SARS-CoV-2 Omicron BA. 1 and BA. 2 are attenuated in rhesus macaques as compared to Delta. Science Advances. 2022;8(46):eade1860. doi: 10.1126/sciadv.ade1860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verma A, Hawes CE, Lakshmanappa YS, et al Monoclonal antibodies protect aged rhesus macaques from SARS-CoV-2-induced immune activation and neuroinflammation. Cell Reports. 2021;37(5):109942. doi: 10.1016/j.celrep.2021.109942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vijayanand S, Patil S, Menon I, et al An adjuvanted inactivated SARS-CoV-2 microparticulate vaccine delivered using microneedles induces a robust immune response in vaccinated mice. Pharmaceutics. 2023;15(3):895. doi: 10.3390/pharmaceutics15030895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogel AB, Kanevsky I, Che Y, et al BNT162b vaccines protect rhesus macaques from SARS-CoV-2. Nature. 2021;592(7853):283–289. doi: 10.1038/s41586-021-03275-y. [DOI] [PubMed] [Google Scholar]
- Wang YQ, Yan A, Song DY, et al Identification of a highly conserved neutralizing epitope within the RBD region of diverse SARS-CoV-2 variants. Nature Communications. 2024;15(1):842. doi: 10.1038/s41467-024-45050-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williamson BN, Feldmann F, Schwarz B, et al Clinical benefit of remdesivir in rhesus macaques infected with SARS-CoV-2. Nature. 2020;585(7824):273–276. doi: 10.1038/s41586-020-2423-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winkler ES, Bailey AL, Kafai NM, et al SARS-CoV-2 infection of human ACE2-transgenic mice causes severe lung inflammation and impaired function. Nature Immunology. 2020;21(11):1327–1335. doi: 10.1038/s41590-020-0778-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wise J Covid-19: long covid risk is lower with omicron than delta, researchers find. BMJ. 2022;377:o1500. doi: 10.1136/bmj.o1500. [DOI] [PubMed] [Google Scholar]
- Wong LYR, Zheng J, Wilhelmsen K, et al Eicosanoid signalling blockade protects middle-aged mice from severe COVID-19. Nature. 2022;605(7908):146–151. doi: 10.1038/s41586-022-04630-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woolsey C, Borisevich V, Prasad AN, et al Establishment of an African green monkey model for COVID-19 and protection against re-infection. Nature Immunology. 2021;22(1):86–98. doi: 10.1038/s41590-020-00835-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y, Shi J, He XX, et al Protection of the receptor binding domain (RBD) dimer against SARS-CoV-2 and its variants. Journal of Virology. 2023a;97(11):e0127923. doi: 10.1128/jvi.01279-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y, Wu NM, Jia XY, et al Long‐term immune response to Omicron‐specific mRNA vaccination in mice, hamsters, and nonhuman primates. MedComm. 2023b;4(6):e460. doi: 10.1002/mco2.460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu YJ, Wang MN, Yin H, et al TREM-2 is a sensor and activator of T cell response in SARS-CoV-2 infection. Science Advances. 2021;7(50):eabi6802. doi: 10.1126/sciadv.abi6802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia J, Tang WQ, Wang JM, et al SARS-CoV-2 N protein induces acute lung injury in mice via NF-ĸB activation. Frontiers in Immunology. 2021;12:791753. doi: 10.3389/fimmu.2021.791753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiong SQ, Zhang LH, Qadir AS, et al Interleukin-1RA mitigates SARS-CoV-2–induced inflammatory lung vascular leakage and mortality in humanized K18-hACE-2 mice. Arteriosclerosis, Thrombosis, and Vascular Biology. 2021;41(11):2773–2785. doi: 10.1161/ATVBAHA.121.316925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu L, Yu DD, Ma YH, et al COVID-19-like symptoms observed in Chinese tree shrews infected with SARS-CoV-2. Zoological Research. 2020a;41(5):517–526. doi: 10.24272/j.issn.2095-8137.2020.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu XL, Han MF, Li TT, et al Effective treatment of severe COVID-19 patients with tocilizumab. Proceedings of the National Academy of Sciences of the United States of America. 2020b;117(20):10970–10975. doi: 10.1073/pnas.2005615117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamasoba D, Uriu K, Plianchaisuk A, et al Virological characteristics of the SARS-CoV-2 omicron XBB. 1.16 variant. The Lancet Infectious Diseases. 2023;23(6):655–656. doi: 10.1016/S1473-3099(23)00278-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang XH, Deng W, Tong Z, et al Mice transgenic for human angiotensin-converting enzyme 2 provide a model for SARS coronavirus infection. Comparative Medicine. 2007;57(5):450–459. [PubMed] [Google Scholar]
- Yin KL, Peluso MJ, Luo XY, et al Long COVID manifests with T cell dysregulation, inflammation and an uncoordinated adaptive immune response to SARS-CoV-2. Nature Immunology. 2024;25(2):218–225. doi: 10.1038/s41590-023-01724-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu DD, Long YHP, Xu L, et al Infectivity of SARS-CoV-2 and protection against reinfection in rats. Zoological Research. 2022a;43(6):945–948. doi: 10.24272/j.issn.2095-8137.2022.339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu JY, Collins ND, Mercado NB, et al Protective efficacy of gastrointestinal SARS-CoV-2 delivery against intranasal and intratracheal SARS-CoV-2 challenge in rhesus macaques. Journal of Virology. 2022b;96(2):e0159921. doi: 10.1128/JVI.01599-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu JY, Tostanoski LH, Peter L, et al DNA vaccine protection against SARS-CoV-2 in rhesus macaques. Science. 2020a;369(6505):806–811. doi: 10.1126/science.abc6284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu P, Qi FF, Xu YF, et al Age‐related rhesus macaque models of COVID‐19. Animal Models and Experimental Medicine. 2020b;3(1):93–97. doi: 10.1002/ame2.12108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu WH, Wang JB, Yang Y, et al SARS‐CoV‐2 Omicron (B. 1.1. 529) infection in rhesus macaques, hamsters, and BALB/c mice with severe lung histopathological damage. Journal of Medical Virology. 2023;95(6):e28846. doi: 10.1002/jmv.28846. [DOI] [PubMed] [Google Scholar]
- Yuan LZ, Tang QY, Zhu HC, et al SARS-CoV-2 infection and disease outcomes in non-human primate models: advances and implications. Emerging Microbes & Infections. 2021;10(1):1881–1889. doi: 10.1080/22221751.2021.1976598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zanza C, Romenskaya T, Manetti AC, et al Cytokine storm in COVID-19: immunopathogenesis and therapy. Medicina. 2022;58(2):144. doi: 10.3390/medicina58020144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zeng JX, Xie XC, Feng XL, et al Specific inhibition of the NLRP3 inflammasome suppresses immune overactivation and alleviates COVID-19 like pathology in mice. eBioMedicine. 2022;75:103803. doi: 10.1016/j.ebiom.2021.103803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Q, Bastard P, Liu ZY, et al Inborn errors of type I IFN immunity in patients with life-threatening COVID-19. Science. 2020a;370(6515):eabd4570. doi: 10.1126/science.abd4570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang XH, Zhang Y, Qiao WZ, et al Baricitinib, a drug with potential effect to prevent SARS-COV-2 from entering target cells and control cytokine storm induced by COVID-19. International Immunopharmacology. 2020b;86:106749. doi: 10.1016/j.intimp.2020.106749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang ZY, Zhang N, Lu XC, et al Anti-infection effects of heparin on SARS-CoV-2 in a diabetic mouse model. Zoological Research. 2023;44(6):1003–1014. doi: 10.24272/j.issn.2095-8137.2023.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y, Wang CL, Gao ZY, et al Ferrets: a powerful model of SARS-CoV-2. Zoological Research. 2023;44(2):323–330. doi: 10.24272/j.issn.2095-8137.2022.351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y, Wang JB, Kuang DX, et al Susceptibility of tree shrew to SARS-CoV-2 infection. Scientific Reports. 2020;10(1):16007. doi: 10.1038/s41598-020-72563-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng HY, He XY, Li W, et al Pro-inflammatory microenvironment and systemic accumulation of CXCR3+ cell exacerbate lung pathology of old rhesus macaques infected with SARS-CoV-2. Signal Transduction and Targeted Therapy. 2021a;6(1):328. doi: 10.1038/s41392-021-00734-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng HY, Zhang M, Yang CX, et al Elevated exhaustion levels and reduced functional diversity of T cells in peripheral blood may predict severe progression in COVID-19 patients. Cellular & Molecular Immunology. 2020;17(5):541–543. doi: 10.1038/s41423-020-0401-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng J, Wong LYR, Li K, et al COVID-19 treatments and pathogenesis including anosmia in K18-hACE2 mice. Nature. 2021b;589(7843):603–607. doi: 10.1038/s41586-020-2943-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng M, Karki R, Williams EP, et al TLR2 senses the SARS-CoV-2 envelope protein to produce inflammatory cytokines. Nature Immunology. 2021c;22(7):829–838. doi: 10.1038/s41590-021-00937-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng MZM, Wakim LM Tissue resident memory T cells in the respiratory tract. Mucosal Immunology. 2022;15(3):379–388. doi: 10.1038/s41385-021-00461-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou XY, Sun WY, Zhang Y, et al A novel hACE2 knock-in mouse model recapitulates pulmonary and intestinal SARS-CoV-2 infection. Frontiers in Microbiology. 2023;14:1175188. doi: 10.3389/fmicb.2023.1175188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu LN, Yang PH, Zhao YZ, et al. 2020. Single-cell sequencing of peripheral mononuclear cells reveals distinct immune response landscapes of COVID-19 and influenza patients. Immunity, 53 (3): 685–696. e3.
- Zhuang Z, Lai XM, Sun J, et al Mapping and role of T cell response in SARS-CoV-2–infected mice. Journal of Experimental Medicine. 2021;218(4):e20202187. doi: 10.1084/jem.20202187. [DOI] [PMC free article] [PubMed] [Google Scholar]