Abstract
Background
Diabetic foot ulcers (DFU) seriously threaten the health and quality of life of patients. The microbiota is the primary reason for the refractory and high recurrence of DFU. This study aimed to determine the wound microbiota at different DFU stages.
Methods
Wound samples were collected from 48 patients with DFU and divided into three phases: inflammatory (I, n = 49), proliferation (P, n = 22), and remodeling (R, n = 19). The wound samples obtained at different stages were then subjected to 16S rRNA gene sequencing. The number of operational taxonomic units (OTUs) in the different groups was calculated according to the criterion of 97 % sequence similarity. The diversity of the microbiota differentially presented bacterial taxa at the phylum and genus levels, and important phyla and genera in the different groups were further explored.
Results
After sequencing, 3351, 925, and 777 OTUs were observed in groups I, P, and R, respectively, and 175 OTUs overlapped. Compared with the inflammatory stage, the α-diversity of wound microbiota at proliferation and remodeling stages was significantly decreased (P < 0.05). At the phylum level, Firmicutes, Proteobacteria, Actinobacteriota, and Bacteroidota were the dominant phyla, accounting for more than 90 % of all the phyla. At the genus level, Random Forest and linear discriminant analysis effect size analyses showed that Peptoniphilus, Lactobacillus, Prevotella, Veillonella, Dialister, Streptococcus, and Ruminococcus were the signature wound microbiota for the inflammatory stage; Anaerococcus, Ralstonia, Actinomyces, and Akkermansia were important species for the proliferation stage; and the crucial genera for the remodeling stage were Enterobacter, Pseudomonas, Sondgrassella, Bifidobacterium, and Faecalibacterium.
Conclusions
There were significant differences in the composition and structure of the wound microbiota in patients with DFU at different stages, which may lay a foundation for effectively promoting wound healing in DFU.
Keywords: Diabetic foot ulcers, Wound microbiota, Inflammation stage, Proliferation stage, Remodeling stage
1. Introduction
Diabetes is a growing public health problem, with rising prevalence and mortality threatening the health of people worldwide [1]. In 2021, 537 million people worldwide were living with diabetes, mainly type 2 diabetes, and it is predicted that by 2045, the number of patients with diabetes will reach 783 million worldwide [2,3]. Diabetic foot ulcers (DFU) are among the most serious complications of chronic diabetes. It has been reported that one-third to one-fifth of individuals with diabetes may be affected by DFU during their lifetime. Currently, treatment strategies for DFU include local wound care and surgical debridement, regular wound assessment and dressing changes, active infection control, and blood glucose control [4]. The failure rate for patients with DFU to recover within one year of treatment is as high as 20 % [5]. The recurrence rate of DFU is high, at 40 % within one year and 65 % within five years, and the lifetime incidence of DFU can be as high as 19%–34 % [6,7]. Consequently, DFU is expensive and can lead to amputation and death [8]. Therefore, it is necessary to investigate the pathogenesis of DFU further and identify effective diagnostic and therapeutic targets to improve treatment efficacy.
Human skin, the largest organ of the body, not only acts as a physical protective barrier against environmental damage but also as a dynamic interface between the host and microbiota [9]. The microbiota that inhabit human skin is very complex and can be divided into symbiotic and pathogenic bacteria [10]. Pathogenic bacteria are harmful and can spread directly to host tissues, causing infections [11]. In contrast, symbiotic bacteria can provide essential nutrients to host tissues and are beneficial in fighting infections [12]. Therefore, skin microbiota play a paradoxical role in wound healing, and changing the microbiota composition of a wound directly affects its healing potential [13]. A previous study has demonstrated that the healing and recurrence of chronic skin wounds are associated with alterations in bacterial diversity and instability in the composition of the skin wound microbiota [14]. There were significant differences in skin microbiota composition between patients with and without diabetes. Grice et al. showed that the abundances of Lachnospiraceae, Streptococcus, and Acinetobacter decreased, while Aerococcus, Porphyromonadaceae, Staphylococcus, and Proteobacteria increased in the wounds of diabetic mice [15]. In addition, a study by Gardner et al. found that Staphylococcus, particularly Staphylococcus aureus, were most common in DFU skin as well as the depth of DFU was positively correlated with the abundance of anaerobic bacteria and negatively correlated with the abundance of Staphylococcus; and the duration of DFU was positively correlated with the bacterial diversity, richness, and Proteobacteria abundance [16]. Another study illustrated that patients with DFU displayed more pairs of pathogenic microorganisms, such as Pseudomonas and Candida [17]. These results suggest that dysbiosis of the skin microbiota may be the main factor in the development of DFU.
Skin wound healing is a complex and well-organized natural repair process that includes four stages: hemostasis, inflammation, proliferation, and remodeling [18]. Skin damage induces wounds, which make it easy for the microbiota to penetrate the skin tissues from the wound, thus resulting in an imbalance in the skin microbiota [19]. Changes in skin wound microbiota are closely related to changes in wound healing dynamics [20]. However, the specific microbiota of wounds at different stages in patients with DFUs remain unclear. Therefore, in this study, wound tissues at different stages were collected for 16S rRNA gene sequencing, and crucial wound microbiota were identified at different stages in patients with DFU. This study is expected to shed new light on the relationship between DFU and wound microbiota and promote DFU healing.
2. Results
2.1. The overall structure of microbiota in the wounds of DFU at different stages
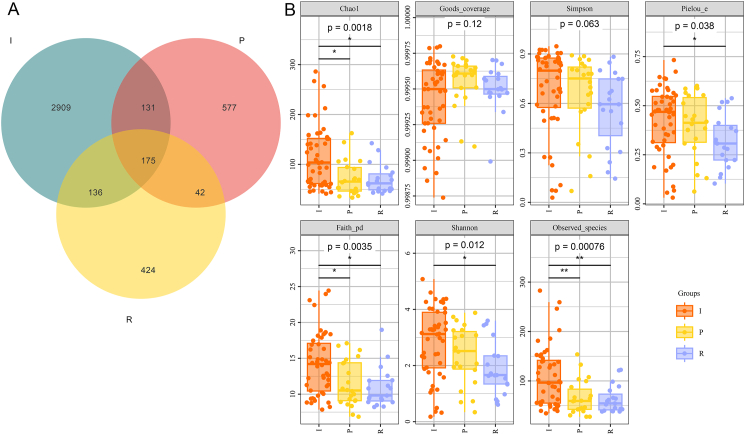
To investigate the roles of the microbiota in DFU wounds at different stages, wound samples at different stages were subjected to 16S rRNA gene sequencing. To verify that the sequencing depth was sufficient to cover all OTU for this species, rarefaction and rank-abundance curves were plotted. As shown in Fig. S1A, with an increase in the sample number, the rarefaction curves of the different groups gradually flattened, and the inflection points of all the sample curves were less than 5000, which was far lower than the average sequence of each sample in the experiment. Rank-abundance curves showed that the curves decreased rapidly before 500 OUT level ranks; that is, the species distribution changed greatly; however, after 500 OUT ranks, the curves tended to flatten out, indicating that the distribution of OTUs in each sample was relatively uniform (Fig. S1B). These results suggest that the sample size was reasonable and the sequencing depth was sufficient to reflect the species composition of the microbiota in the wound and to capture most of the diversity. In addition, the Venn diagram showed that there were 3351, 925, and 777 OTUs in groups I, P, and R, respectively, and that 175 OTUs overlapped in these three groups (Fig. 1A).
Fig. 1.
Overall structure of the microbiota in the wounds of diabetic foot ulcers (DFU) at different stages. (A) Venn diagram of annotated wound microbiota (OTUs) in different groups. (B) The α-diversity of the wound microbiota including Chao1, Good's coverage, Simpson, Pielou's evenness, Faith's PD, Shannon, and observed species in the different groups. I: inflammatory stage; P: proliferation stage; R: remodeling stage.
Subsequently, the diversity indices in the different groups were calculated, including Chao1, Good's coverage, Simpson, Pielou's evenness, Faith's PD, Shannon, and observed species. Chao1 and observed species indices reflected species richness, and the Shannon and Simpson indices were used to evaluate species diversity. Faith's PD index refers to species evolutionary diversity, and Pielou's evenness index represents species evenness. The Good's coverage in the I, P, and R groups were respectively 0.9994 ± 0.0003, 0.9996 ± 0.0002, and 0.9995 ± 0.0002, respectively (Fig. 1B), implying that the current sequencing essentially contained most species, and can be used for further analysis. Additionally, there was no significant difference in the Simpson index among groups I, P, and R (P = 0.063; Fig. 1B). Compared with the I group, the values of Chao1, Pielou's evenness, Faith's PD, Shannon, and observed species were significantly lower in the P and R groups (P < 0.05; Fig. 1B). It can be inferred that the diversity of the microbiota in DFU wounds at the proliferation and remodeling stages could be lower than that at the inflammatory stage.
2.2. The specific microbiota in the wounds of DFU at different stages at the phylum level
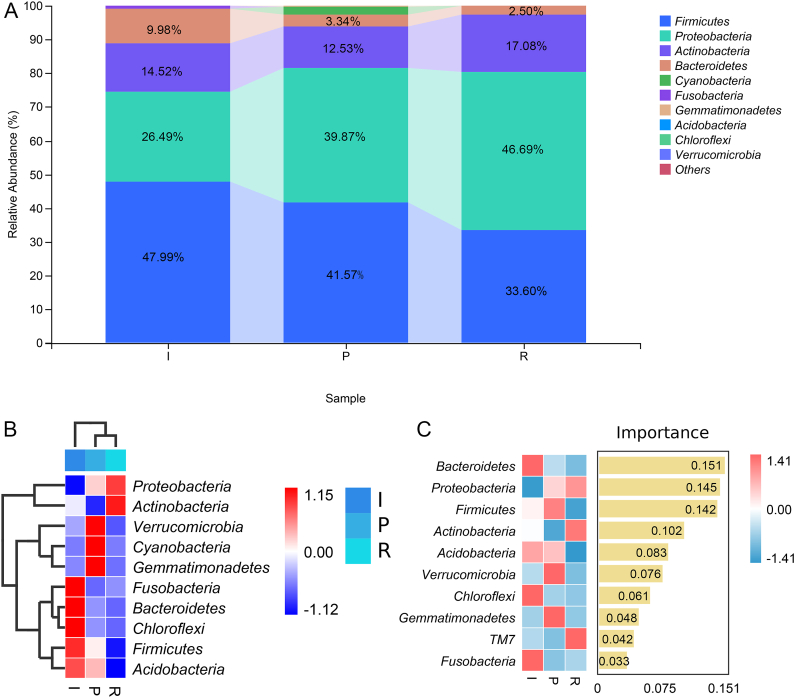
After analyzing the diversity of the microbiota in the wounds of the different groups, we further explored the changes in the composition of specific microbiota in DFU wounds at the phylum level. Based on Fig. 2A, Firmicutes, Proteobacteria, Actinobacteriota, and Bacteroidota were the dominant phyla in all the different groups. Among them, the relative abundances of Firmicutes (41.57 % in P and 33.60 % in R) and Bacteroidota (3.34 % in P and 2.50 % in R) were lower in the P and R groups than in the I group (47.99 % for Firmicutes and 9.98 % for Bacteroidota); as well as the Proteobacteria (39.87 % in P and 46.69 % in R) abundance was higher in the P and R groups than in the I group (26.49 %) (Fig. 2A). A clustering heatmap of the top 10 phyla in groups I, P, and R is displayed in Fig. 2B. Random Forest analysis at the phylum level observed that Bacteroidetes (importance = 0.151), Chloroflexi (importance = 0.061), and Fusobacteria (importance = 0.033) were important phyla for the wound at the inflammatory stage; and Firmicutes (importance = 0.142), Verrucomicrobia (importance = 0.076), and Gemmatimonadetes (importance = 0.048) were crucial for the wound at the proliferation stage; as well as the essential phyla for the wound in the remodeling stage were TM7 (importance = 0.042), Actinobacteria (importance = 0.102), and Proteobacteria (importance = 0.145) (Fig. 2C).
Fig. 2.
Specific microbiota in DFU wounds at different stages at the phylum level. (A) Distribution of the top 10 dominant phyla of the wound microbiota in different groups. (B) Clustering heatmap of the top 10 phyla in different groups. (C) Differences in the top 10 phyla among the different groups using Random Forest analysis. The heatmap shows the abundance distribution of the wound microbiota phyla in different samples, and the importance of species to the model decreases from top to bottom. I: inflammatory stage; P: proliferation stage; R: remodeling stage.
2.3. The specific microbiota in the wounds of DFU at different stages at the genus level
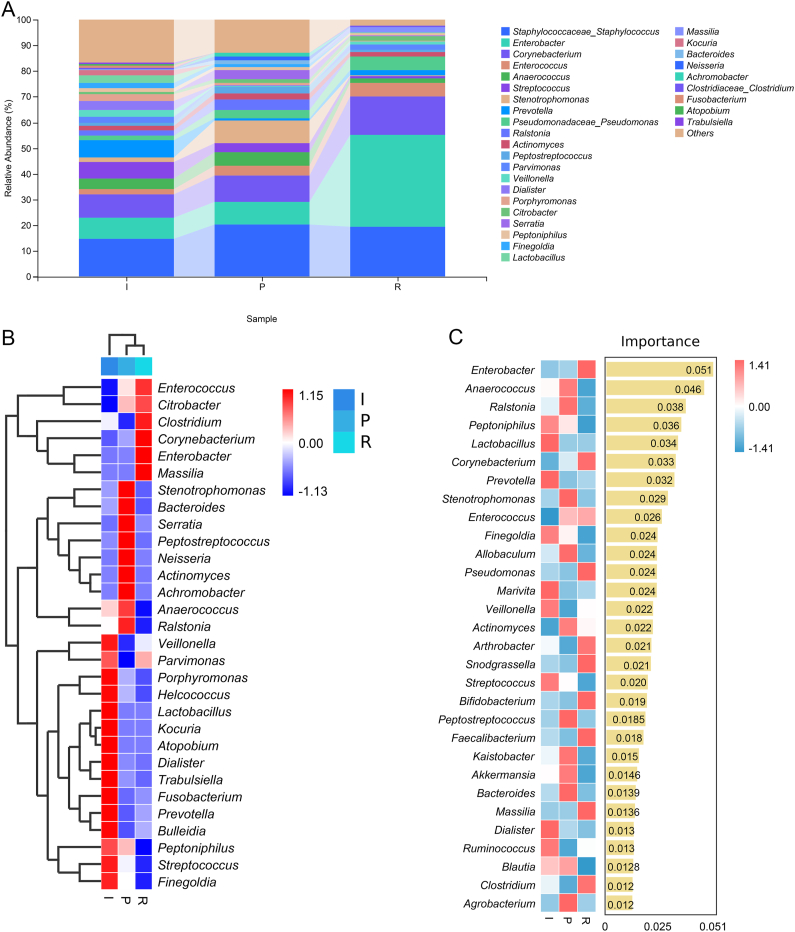
Furthermore, alterations in the composition of specific microbiota in DFU wounds at the genus level were studied. At the genus level, top 30 bacterial genera were analyzed, including Staphylococcaceae-Staphylococcus, Enterobacter, Corynebacterium, Enterococcus, Anaerococcus, Streptococcus, Stenotrophomonas, Prevotella, Pseudomonadaceae-Pseudomonas, Veillonella, Dialister, Serratia, Lactobacillus, Massillia, and Kocuria (Fig. 3A). Compared with the I and P groups, the relative abundance of Enterobacter (8.21 % in I, 8.96 % in P and 35.48 % in R), Corynebacterium (8.92 % in I, 10.02 % in P and 15.15 % in R), Enterococcus (2.05 % in I, 3.95 % in P and 5.23 % in R), Massilia (0.0057 % in I, 0.0024 % in P and 2.03 % in R), Citrobacter (0.75 % in I, 1.45 % in P and 1.70 % in R), and Clostridium (0.38 % in I, 0.17 % in P and 0.63 % in R) was significantly increased in the R group (Fig. 3A and B, Table S1). In the P group, the abundances of Stenotrophomonas (8.59 %), Serratia (3.38 %), Ralstonia (4.25 %), Anaerococcus (5.34 %), Peptostreptococcus (2.76 %), and Bacteroides (1.44 %) were higher; while the higher genus in the I group included Veillonella (2.57 %), Dialister (3.58 %), Lactobacillus (3.01 %), Kocuria (2.00 %), Streptococcus (6.64 %), Prevotella (6.82 %), Fusobacterium (0.88 %), and Trabulsiella (0.64 %) (Fig. 3A and B, Table S1). Random Forest analysis at the genus level showed that Enterobacter (importance = 0.051), Anaerococcus (importance = 0.046), Ralstonia (importance = 0.038), Lactobacillus (importance = 0.034), Corynebacterium (importance = 0.033), Prevotella (importance = 0.032), Stenotrophomonas (importance = 0.0.029), Pseudomonas (importance = 0.024), Bifidobacterium (importance = 0.019), Faecalibacterium (importance = 0.018), and Akkermansia (importance = 0.0146) were important genus to distinguish the I, P, and R groups (Fig. 3C).
Fig. 3.
Specific microbiota in DFU wounds at different stages at the genus level. (A) Distribution of the top 30 dominant genera of the wound microbiota in the different groups. (B) Cluster heatmap of the top 30 genera in the different groups. (C) Differences in the top 30 genera in the different groups using Random Forest analysis. The heatmap shows the abundance distribution of the genera of the wound microbiota in the different samples, and the importance of species in the model decreases from top to bottom. I: inflammatory stage; P: proliferation stage; R: remodeling stage.
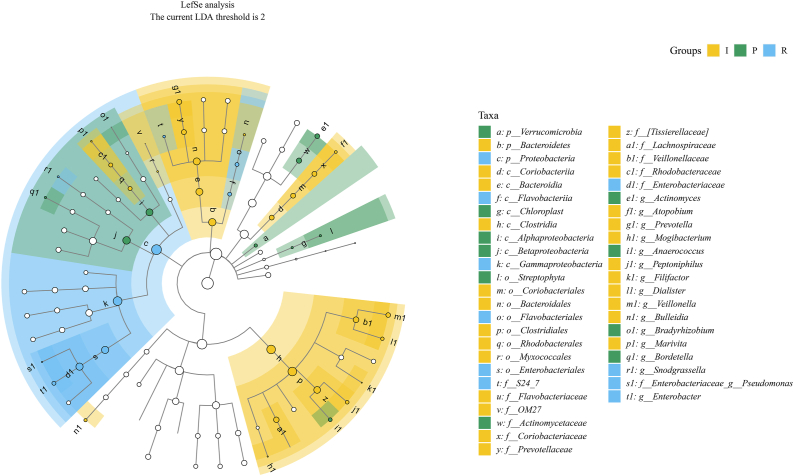
Linear discriminant analysis effect size (LEfSe) can be used to analyze the differences in the microbiota between groups and identify the major microbiota specific to each group, helping to uncover potential biomarkers. Our study found that Bacteroidetes was the primary phylum in group I, Verrucomicrobia was important for group P, and Proteobacteria were especially abundant in group R (Fig. 4). At the genus level, Atopobium, Prevotella, Mogibacterium, Peptoniphilus, Filifactor, Dialister, Veillonella, Bulleidia, and Marivita were the signature wound microbiota in group I; the crucial genera in the P group were Actinomyces, Anaerococcus, Bradyrhizobium, and Bordetella; and Sondgrassella and Enterobacter were important species in group R (Fig. 4).
Fig. 4.
Linear discriminant analysis (LDA) effect size (LEfSe) was employed to identify biomarkers of bacterial communities among different groups at different levels. The current low-density LDA threshold is 2. I: inflammatory stage; P: proliferation stage; R: remodeling stage.
2.4. Functional analyses of the annotated microbiota in the wounds of DFU
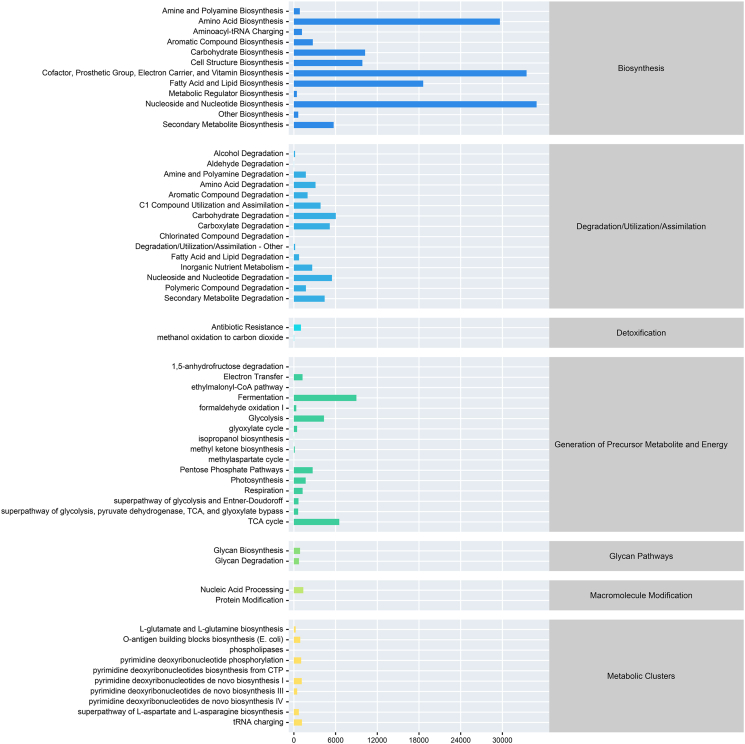
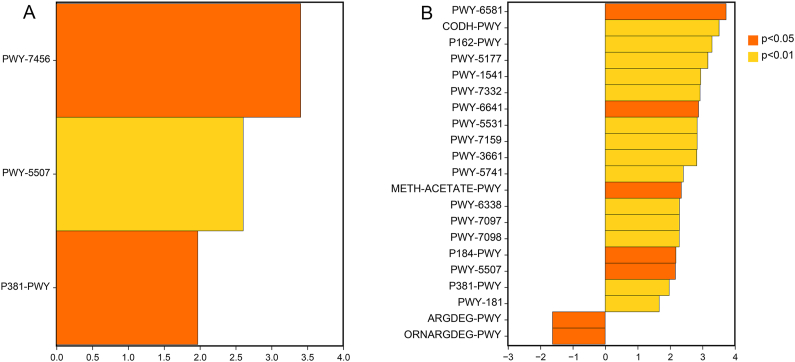
The annotated microbiota in DFU wounds were subjected to functional prediction, “fatty acid and lipid biosynthesis,” “amino acid biosynthesis,” “cell structure biosynthesis,” “carbohydrate biosynthesis,” “secondary metabolites biosynthesis,” “nucleoside and nucleotide biosynthesis,” “cofactor, prosthetic group, electron carrier, and vitamin biosynthesis,” “fermentation,” and “TCA cycle” (Fig. 5). The metagenomeSeq method was used to screen for significantly different metabolic pathways between the groups. Comparing groups I and P, the significantly different metabolic pathways were PWY-7456 (mannan degradation), PWY-5507 (adenosylcobalamin biosynthesis I (early cobalt insertion)), and P381-PWY (adenosylcobalamin biosynthesis II (late cobalt incorporation)) (Fig. 6A). Additionally, 21 differential functional pathways were observed between the I and R groups, such as PWY-6581 (spirilloxanthin and 2,2′-diketo-spirilloxanthin biosynthesis), CODH-PWY (reductive acetyl coenzyme A pathway), PWY-6641 (superpathway of sulfolactate degradation), P184-PWY (protocatechuate degradation I (meta-cleavage pathway)), PWY-5507 (adenosylcobalamin biosynthesis I (early cobalt insertion)), METH-ACETATE-PWY (methanogenesis from acetate), ARGDEG-PWY (superpathway of l-arginine, putrescine, and 4-aminobutanoate degradation), and ORNARGDEG-PWY (superpathway of l-arginine and l-ornithine degradation) (Fig. 6B).
Fig. 5.
Functional analysis of the annotated microbiota in the wounds of DFU using the MetaCyc database. The horizontal coordinate represents the abundance of functional pathways, the ordinate represents the item of the functional pathway of the second MetaCyc classification level, and the far right represents the first-level classification to which the pathway belongs.
Fig. 6.
Significantly different metabolic pathways enriched by bacterial communities among different groups. Significantly different metabolic pathways were enriched in bacterial communities between the inflammatory and proliferation stages (A) and between the inflammatory and remodeling stages (B). I: inflammatory stage; P: proliferation stage; R: remodeling stage.
2.5. Correlation analysis between the top 50 genera at different stages and the clinical indices in DFU
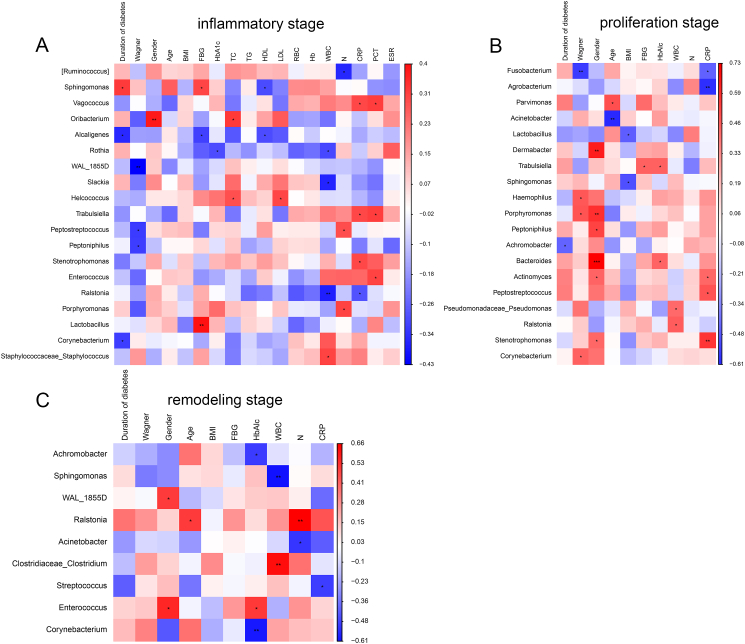
The correlation between the top 50 gut microbiota at the genus level in the inflammatory, proliferative, and remodeling stages and the clinical indices of DFU was further investigated. At the inflammation stage, the gut microbiota of Sphingomonas was significantly positively correlated with the duration of diabetes and fasting blood glucose but negatively correlated with high-density lipoprotein (Fig. 7A). Vagococcus positively correlated with C-reactive protein and procalcitonin, Enterococcus positively correlated with procalcitonin, and Ralstonia negatively correlated with white blood cells and C-reactive protein (Fig. 7A). For Lactobacillus, it significantly positively correlated with fasting blood glucose levels (P < 0.01; Fig. 7A). At the proliferation stage, Lactobacillus and Sphingomonas were negatively correlated with body mass index, while Haemophilus, Porphyromonas, and Corynebacterium were significantly positively correlated with the Wagner stage of DFU (P < 0.05, Fig. 7B). C-reactive protein was negatively associated with Fusobacterium and Agrobacterium and positively associated with Actinomyces, Peptrostreptococcus, and Stenotrophomonas (Fig. 7B). In addition, at the remodeling stage, the level of white blood cells was negatively correlated with Sphingomonas and positively correlated with Clostridiaceae-Clostridum (Fig. 7C).
Fig. 7.
Correlation analysis between the top 50 genus at different stages and clinical indices in DFU. Correlation between genus and clinical indices at the inflammatory (A), proliferation (B), and remodeling (C) stages. BMI: body mass index; FBG: fasting blood glucose; HbA1c: glycated hemoglobin; TC: total cholesterol; TG: triglyceride; HDL: high-density lipoprotein; LDL: low-density lipoprotein; RBC: red blood cell; Hb: hemoglobin; WBC: white blood cells; N: nitrogen; CRP: C-reactive protein; PCT: procalcitonin; ESR: erythrocyte sedimentation rate.
3. Discussion
DFU is a major chronic complication of diabetes and a primary cause of disability and death, seriously endangering the health and quality of life of patients [21]. The skin microbiota influence various processes in the skin, such as keratinocyte proliferation, epithelial differentiation, epidermal blood vessel growth, and cell signaling [22,23]. An imbalance in the skin microbiota can damage skin immune homeostasis, thus causing skin lesions, and the microbiota is the main reason for the refractory and high recurrence of DFU [24]. An earlier study compared the bacterial communities in the wounds of DFU, venous leg ulcers, decubitus ulcers, and non-healing surgical wounds in American populations (White, Black, and Hispanic races) using 16S rDNA pyrosequencing and found that neither patient demographics nor wound type could affect the bacterial composition of the microbiome in chronic wounds, and Staphylococcus and Pseudomonas were higher in wound samples, accounting for 63 % and 25 % of all wound samples, respectively [25]. Pang et al. [26] showed that the diversity and richness of the foot skin microbiome increased during the development of diabetes and that the main skin microbiota phyla were Firmicutes, Proteobacteria, Actinobacteria, and Bacteroides. Another study in China demonstrated that the microbial diversity in diabetic foot osteomyelitis samples was significantly higher than that in post-traumatic foot osteomyelitis; Prevotellaceae, Firmicutes, and Prevotella were the most abundant in diabetic foot osteomyelitis, and the most abundant microbiota in post-traumatic foot osteomyelitis were Prevotella jejuni, Proteobacteria, Prevotella fusca, and Halomonadaceae [27]. In this study, 16s rRNA gene sequencing was employed to identify the key wound microbiota at different stages in patients with DFU in China, and we found that the α-biodiversity of wound microbiota at the inflammatory stage could be higher than that at the proliferation and remodeling stages. The possible reason is that there are more pro-inflammatory cytokines (interleukin (IL)-1, and tumor necrosis factor-α) in the inflammatory period, and the presence of pro-inflammatory cytokines may make the bacteria persist in the wound, thereby further damaging the wound healing process [19,23]. However, at the proliferation and remodeling stages, the wound gradually heals, and some pathogenic microorganisms may be suppressed under treatment; thus, the diversity of the wound microbiota may be reduced [22,28]. Therefore, we speculate that the overall diversity and structure of the wound microbiota in DFU may be higher during the inflammatory period than during the proliferation and remodeling periods; however, specific reasons need to be investigated in the future.
Furthermore, crucial wound microbiota were identified at different stages at the phylum and genus levels. At the phylum level, Firmicutes, Proteobacteria, Actinobacteriota, and Bacteroidota were the dominant phyla and were important for distinguishing the three stages. Zhang et al. [29] showed that with the progression of diabetes complications, the abundance of gram-negative bacteria could increase in the plantar skin, accompanied by decreased Firmicutes and increased Proteobacteria and Bacteroidetes. Another study also reported that Firmicutes, Actinobacteria, Proteobacteria, Bacteroidetes, and Clostriobacteria were the dominant phyla in DFU [30]. At the genus level, Peptoniphilus, Lactobacillus, Prevotella, Veillonella, Dialister, Streptococcus, and Ruminococcus were the signature wound microbiota for the inflammatory stage; Anaerococcus, Ralstonia, Stenotrophomonas, Actinomyces, and Akkermansia were important species for the proliferation stage; and the crucial genera for the remodeling stage were Enterobacter, Pseudomonas, Sondgrassella, Bifidobacterium, and Faecalibacterium.
The regulation of inflammation is an important mechanism of wound healing. A blood clot forms one to three days after skin injury, which controls bleeding and induces neutrophils to release factors, such as platelet-derived growth factor, tumor necrosis factor-α, and IL-1, thus triggering inflammation [31]. Gram-positive anaerobes, which are resident bacteria in the human skin, can express various virulence factors and cause inflammation, thereby contributing to increased pathogenicity and impaired wound healing [32]. Peptoniphilus, a gram-positive anaerobe, is associated with impaired healing of DFU [33]. Although it is a symbiotic bacterium, Prevotella, together with other microbes from normal microbiota communities, has been implicated in various human infections [34]. Zou et al. [27] have demonstrated that Prevotella is the most abundant microbiota in diabetic foot osteomyelitis and is positively correlated with the duration of diabetic foot infections. Veillonella, a gram-negative obligate anaerobic bacterium, is significantly enriched in the intestines of patients with inflammatory bowel disease and can transition from fermentation to anaerobic respiration in an inflammatory environment, thus further promoting inflammation progression [35]. Park et al. [30] showed that compared with the mild DFU group, the abundance of Peptoniphilus, Prevotella, and Dialister was enhanced in severe DFU. Ruminococcus is a mucin-degrading gut bacterium that alleviates atopic dermatitis by enhancing the proportion of regulatory T cells and producing short-chain fatty acids [36]. A previous study indicated that Streptococcus may be closely related to the occurrence of DFU and participate in its progression through the insulin pathway [37]. Lactobacillus, a beneficial bacterium, can compete against skin pathogens by inhibiting adhesion, producing antimicrobial metabolites, and influencing pathogen metabolism [38]. In addition to typical skin symbiont bacteria, Lactobacillus may be beneficial for wound healing, with lysates that increase the migration and proliferation of keratinocytes or produce organic acids that fight pathogens and inhibit wound biofilm formation [39,40]. Our study found that Lactobacillus was enriched during the inflammatory period, which may be related to the location of sample collection [41]. Peptoniphilus, Lactobacillus, Prevotella, Veillonella, Dialister, Streptococcus, and Ruminococcus play important roles in DFU wounds during inflammation.
After the inflammatory stage, inflammatory cytokines can activate fibroblasts and release keratinocyte growth factor-1 and -2 and IL-6, inducing adjacent keratinocytes to migrate to the wound area, resulting in proliferation (proliferation stage) and subsequent epithelialization (remodeling stage) [20]. It has been reported that wounds following blunt or penetrating trauma may also alter the skin microbiome, as well as Anaerococcus, Pseudomonas, Streptococcus, Staphylococcus, Corynebacterium, and Acinetobacter were dominant [42]. Ralstonia is a gram-negative opportunistic bacterium that can survive in various solutions and cause various infections [43]. Stenotrophomonas is a gram-negative obligatory aerobic bacterium reported to be closely associated with major amputations in patients with limb-threatening DFU [44]. Actinomyces is closely associated with insulin resistance and inflammation [45]. Akkermansia is a probiotic known to relieve metabolic disorders and inflammatory responses, and its abundance is reduced in patients with diabetes [46]. Akkermansia muciniphila can strengthen the intestinal barrier and regulate the balance between pro-inflammatory and anti-inflammatory cytokines through various signaling pathways, thus relieving insulin resistance and metabolic syndrome [46]. Enterobacter, Staphylococcus aureus, Klebsiella pneumoniae, and Acinetobacter are important pathogens isolated from DFU that are involved in their progression [47]. Pseudomonas, a gram-negative bacterium, is a common pathogen that causes longer hospital stays and increases morbidity and mortality [48]. Sondgrassella is a core functional gut bacterium that has not been reported in DFU [49]. Faecalibacterium is a strictly anaerobic, extremely oxygen-sensitive, gram-positive bacterium, and its low levels are associated with inflammation [50]. Bifidobacterium, also a probiotic, can metabolize propionic acid and lactic acid to produce butyrate, which has anti-inflammatory, anti-glycemic, and barrier-enhancing activities, thereby helping maintain the balance of gut microbes and prevent invasion by pathogenic bacteria [51]. These reports, combined with our results, suggest that Anaerococcus, Ralstonia, Stenotrophomonas, Actinomyces, and Akkermansia may be candidate signatures for DFU wounds at the proliferation stage, and the essential biomarkers for the remodeling stage may be Enterobacter, Pseudomonas, Sondgrassella, Bifidobacterium, and Faecalibacterium. However, a recent study by Jaber et al. [52] comprehensively investigated the microbial ecology of diabetic foot infections using PCR and found that Staphylococcus epidermidis, Bacteroides fragilis, Candida albicans, Staphylococcus aureus, and Clostridium perfringens were the main pathogenic bacteria responsible for diabetic foot infections. Therefore, the underlying effects of the identified microbiota on DFU wound healing at different stages require further investigation through a series of experiments such as PCR.
In conclusion, 16s rRNA gene sequencing revealed characteristic changes in the wound microbiota of patients with DFU at the inflammatory, proliferative, and remodeling stages. Specifically, wounds in the inflammatory period of DFU may have a higher α-diversity of microbiota than those in the proliferation and remodeling periods. At the phylum level, Firmicutes, Proteobacteria, Actinobacteriota, and Bacteroidota were the dominant phyla that differentiated among the three stages. In addition, Peptoniphilus, Lactobacillus, Prevotella, Veillonella, Dialister, Streptococcus, and Ruminococcus may serve as crucial biomarkers of wounds at the inflammatory stage; and the candidate signatures for wounds at the proliferation stage may be Anaerococcus, Ralstonia, Actinomyces, and Akkermansia; as well as Enterobacter, Pseudomonas, Sondgrassella, Bifidobacterium, and Faecalibacterium may be key for wound at the remodeling stage in DFU. These findings broaden our understanding of how the microbiota affects wound healing in DFU and provide novel potential targets for therapeutic interventions to promote wound healing in DFU at different stages.
4. Materials and methods
4.1. Ethics statement
This study was approved by the Institutional Ethics Committee of Taiyuan Central Hospital (approval no. 2023046), and informed consent was obtained from all participants in accordance with national legislation and the Declaration of Helsinki.
4.2. Patients and sample collection
From November 1, 2022, to December 31, 2023, 48 patients diagnosed with DFU were recruited from Taiyuan Central Hospital. The inclusion criteria for DFU were the patients with DFU aged of 50–90 years, duration of DFU >6 weeks, and no antibiotic use for at least two weeks. The exclusion criteria for DFU were as follows: patients with malignant tumors, digestive system diseases, immune system diseases, or mental diseases; patients with a history of bowel diseases (inflammatory bowel disease, bowel cancer, constipation, diarrhea, etc.); pregnant or lactating women; patients with antibiotic use within the last month; and patients with serious illness or an inability to provide test samples. Wound samples were collected via full flushing during the first debridement after admission. Based on the Wagner and Infectious Disease Society of America-International Working Group of Diabetic Foot (IDSA-IWGDF) classification system, all wound samples were divided into three stages: inflammatory (I, n = 49), proliferation (P, n = 22), and remodeling (R, n = 19). Basic clinical information of the enrolled participants is presented in Table 1.
Table 1.
The basic clinical information of patients with diabetic foot ulcers.
| Characteristics | Mean | Standard deviation |
|---|---|---|
| Age (years) | 63.75 | 11.03 |
| BMI (kg/m2) | 23.78 | 2.62 |
| FBG (mmol/L) | 10.06 | 2.80 |
| FINS (uU/mL) | 5.29 | 6.90 |
| fasting C-peptide (ng/mL) | 2.04 | 3.45 |
| HbA1c (%) | 8.63 | 1.99 |
| TC (mmol/L) | 3.78 | 1.14 |
| TG (mmol/L) | 1.53 | 0.87 |
| HDL (mmol/L) | 0.90 | 0.24 |
| LDL (mmol/L) | 2.52 | 0.86 |
| ALT (lU/L) | 18.38 | 11.86 |
| AST (lU/L) | 18.10 | 6.63 |
| BUN (mmol/L) | 8.77 | 4.88 |
| Scr (μmol/L) | 114.29 | 136.50 |
| UA (μmol/L) | 319.71 | 114.97 |
| 24 h urine protein (g/24h) | 1.16 | 2.24 |
| RBC (*1012/L) | 4.12 | 0.90 |
| Hb (g/L) | 119.85 | 25.23 |
| WBC (*109/L) | 9.21 | 3.21 |
| N (%) | 76.40 | 8.34 |
| CRP (mg/L) | 67.17 | 72.34 |
| PCT | 0.23 | 0.37 |
| ESR (mm/h) | 57.27 | 34.24 |
| Course (years) | 16.79 | 8.04 |
| N | % | |
| Sex (male) | 36 | 75 |
| Smoking history | 27 | 56.25 |
| Drinking history | 12 | 25 |
| Hypertension | 20 | 41.67 |
| Coronary heart disease | 18 | 37.5 |
| Cerebrovascular disease | 12 | 25 |
| Insulinize | 31 | 64.58 |
| Amputation | 29 | 60.42 |
| Diabetic retinopathy | 25 | 52.08 |
| Diabetic nephropathy | 27 | 56.25 |
| Peripheral artery disease | 48 | 100 |
| Diabetic peripheral neuropathy | 48 | 100 |
BMI: body mass index; FBG: fasting blood glucose; FINS: fasting insulin; HbA1c: glycated hemoglobin; TC: total cholesterol; TG: triglyceride; HDL: high density lipoprotein; LDL: low density lipoprotein; ALT: glutamic-pyruvic transaminase; AST: aspartate amino transferase; BUN: blood urea nitrogen; Scr: serum creatinine; UA: uric acid; RBC: red blood cell; Hb: hemoglobin; WBC: white blood cells; N: nitrogen; CRP: C-reactive protein; PCT: procalcitonin; ESR: erythrocyte sedimentation rate.
4.3. 16S rRNA gene sequencing
Wound samples obtained at different stages were submitted to Personal Bio-Technology Co., Ltd. (Shanghai, China) for 16S rRNA gene sequencing. Briefly, total microbial genomic DNA was extracted from each wound sample using the QIAamp DNA isolation kit (Qiagen, Hilden, Germany) according to the manufacturer's instructions. DNA concentration and purity were assessed using an ultraviolet spectrophotometer and 1 % agarose gel electrophoresis, respectively. Subsequently, the DNA samples were used to amplify the V3–V4 region of the 16S rRNA gene with the following primers: F: 5′-ACTCCTACGGGAGGCAGCA-3′; and R: 5′-GGACTACHVGGGTWTCTAAT-3′ [53]. Subsequently, 1 % agarose gel electrophoresis was performed to evaluate the PCR products, and gel recovery reagents from AXYGEN were used to recover and purify the PCR products. The recovered products were quantified using the Quant-iT PicoGreen dsDNA Assay Kit and mixed at equal concentrations. Equal amounts of DNA were used to prepare sequencing libraries using the TruSeq Nano DNA LT Library Prep Kit. The quality of the sequencing libraries was evaluated using an Agilent High Sensitivity DNA Kit, and the constructed sequencing libraries were quantified using the Quant-iT PicoGreen dsDNA Assay Kit with the Promega QuantiFluor system in accordance with the manufacturer's protocols. Finally, 16S rRNA gene sequencing was performed using the Illumina HiSeq platform. The raw data of 16S rRNA gene sequencing were deposited in the NCBI SRA database, and the accession number is “PRJNA1105325.”
4.4. Sequencing data analysis
The PersonalBio cloud platform (https://www.genescloud.cn/home) was used for sequencing data analysis. The raw sequencing data were stored in the FASTQ format, and QIIME2 software (version 2019.4) was used for analysis. The raw data were quality-filtered, denoised, and assembled; chimeric sequences were removed using the DADA2 method, and amplicon sequence variants were obtained. Subsequently, the Greengenes reference database classifier (release 13.8) was used to annotate OTUs according to the criterion of 97 % sequence similarity. Afterward, the diversity of the microbiota was analyzed using the ggplot 2 package of R with the QIIME2 software (version 2019.4), as well as the differentially presented bacterial taxa at phylum and genus levels in the different groups, the important phyla/genera in the different groups were analyzed using the self-written perl script in the QIIME2 software (version 2019.4), and Random Forest and LefSe analyses. In addition, the identified microbiota among the different groups were predicted using PICRUSt2 (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) [54] based on the MetaCyc database (https://metacyc.org/), and the significantly different pathways between the two groups were further explored using the metagenomeSeq package in R with a threshold of P < 0.05. In addition, the correlation analysis of the top 50 genera at different stages and the clinical indices were further explored using the Spearman correlation coefficient with |R| > 0.3 and P < 0.05 as the significant differences.
Ethics approval and consent to participate
This study was approved by the Institutional Ethics Committee of Taiyuan Central Hospital (approval no. 2023046), as well as the informed consent was obtained from all the participants in accordance with the national legislation and the Declaration of Helsinki.
Consent for participate
The informed consent was obtained from all the participants in accordance with the national legislation and the Declaration of Helsinki.
Availability of data and materials
Raw 16S rRNA gene sequencing data are openly available in a public repository that issues datasets with accession number PRJNA1105325 in the NCBI SRA database (https://dataview.ncbi.nlm.nih.gov/object/PRJNA1105325?reviewer=6e2lh8k8b5f10kv2p2926bq6d2). Additionally, other data used and/or analyzed in the current study are available from the corresponding author upon reasonable request.
Funding
This work was supported by Taiyuan Bureau of Science and Technology, Science, Technology, and Innovation Program of National Regional Medical Center (Project No. 202210).
CRediT authorship contribution statement
Ying Li: Writing – original draft, Methodology, Investigation, Formal analysis, Conceptualization. Li Zhang: Validation, Resources, Methodology, Investigation, Data curation. Meifang He: Software, Resources, Methodology, Investigation, Formal analysis. Yuebin Zhao: Writing – review & editing, Visualization, Supervision, Project administration, Funding acquisition, Conceptualization.
Declaration of competing interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgements
Not applicable.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.heliyon.2024.e34368.
Appendix A. Supplementary data
The following is/are the supplementary data to this article.
References
- 1.Rehman Z.U., Khan J., Noordin S. Diabetic foot ulcers: contemporary assessment and management. JPMA The Journal of the Pakistan Medical Association. 2023;73(7):1480–1487. doi: 10.47391/jpma.6634. [DOI] [PubMed] [Google Scholar]
- 2.Zhang P., Lu J., Jing Y., Tang S., Zhu D., Bi Y. Global epidemiology of diabetic foot ulceration: a systematic review and meta-analysis (†) Ann. Med. 2017;49(2):106–116. doi: 10.1080/07853890.2016.1231932. [DOI] [PubMed] [Google Scholar]
- 3.Xu Y.F., Wu Y.X., Wang H.M., Gao C.H., Xu Y.Y., Yan Y. Bone marrow-derived mesenchymal stem cell-conditioned medium ameliorates diabetic foot ulcers in rats. Clinics. 2023;78 doi: 10.1016/j.clinsp.2023.100181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kim J., Nomkhondorj O., An C.Y., Choi Y.C., Cho J. Management of diabetic foot ulcers: a narrative review. Journal of Yeungnam medical science. 2023;40(4):335–342. doi: 10.12701/jyms.2023.00682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Deng H., Li B., Shen Q., Zhang C., Kuang L., Chen R., et al. Mechanisms of diabetic foot ulceration: a review. J. Diabetes. 2023;15(4):299–312. doi: 10.1111/1753-0407.13372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang X.N., Wu C.Y., Wu Z.W., Xu L.X., Jiang F.T., Chen H.W. Association between the diabetic foot ulcer and the bacterial colony of the skin based on 16S rRNA gene sequencing: an observational study. Clin. Cosmet. Invest. Dermatol. 2023;16:2801–2812. doi: 10.2147/ccid.S425922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Edmonds M., Lázaro-Martínez J.L., Alfayate-García J.M., Martini J., Petit J.M., Rayman G., et al. Sucrose octasulfate dressing versus control dressing in patients with neuroischaemic diabetic foot ulcers (Explorer): an international, multicentre, double-blind, randomised, controlled trial. Lancet Diabetes Endocrinol. 2018;6(3):186–196. doi: 10.1016/s2213-8587(17)30438-2. [DOI] [PubMed] [Google Scholar]
- 8.Gao D., Zhang Y., Bowers D.T., Liu W., Ma M. Functional hydrogels for diabetic wound management. APL Bioeng. 2021;5(3) doi: 10.1063/5.0046682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Ito Y., Amagai M. Dissecting skin microbiota and microenvironment for the development of therapeutic strategies. Curr. Opin. Microbiol. 2023;74 doi: 10.1016/j.mib.2023.102311. [DOI] [PubMed] [Google Scholar]
- 10.Chen Y., Knight R., Gallo R.L. Evolving approaches to profiling the microbiome in skin disease. Front. Immunol. 2023;14 doi: 10.3389/fimmu.2023.1151527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gao T., Wang X., Li Y., Ren F. The role of probiotics in skin health and related gut-skin Axis: a review. Nutrients. 2023;15(14) doi: 10.3390/nu15143123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Liu Q., Ranallo R., Rios C., Grice E.A., Moon K., Gallo R.L. Crosstalk between skin microbiota and immune system in health and disease. Nat. Immunol. 2023;24(6):895–898. doi: 10.1038/s41590-023-01500-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Luqman A., Götz F. The ambivalent role of skin microbiota and adrenaline in wound healing and the interplay between them. Int. J. Mol. Sci. 2021;22(9) doi: 10.3390/ijms22094996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Xu Z., Hsia H.C. The impact of microbial communities on wound healing: a review. Ann. Plast. Surg. 2018;81(1):113–123. doi: 10.1097/sap.0000000000001450. [DOI] [PubMed] [Google Scholar]
- 15.Grice E.A., Snitkin E.S., Yockey L.J., Bermudez D.M., Liechty K.W., Segre J.A. Longitudinal shift in diabetic wound microbiota correlates with prolonged skin defense response. Proc Natl Acad Sci U S A. 2010;107(33):14799–14804. doi: 10.1073/pnas.1004204107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gardner S.E., Hillis S.L., Heilmann K., Segre J.A., Grice E.A. The neuropathic diabetic foot ulcer microbiome is associated with clinical factors. Diabetes. 2013;62(3):923–930. doi: 10.2337/db12-0771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Oates A., Bowling F.L., Boulton A.J., McBain A.J. Molecular and culture-based assessment of the microbial diversity of diabetic chronic foot wounds and contralateral skin sites. J. Clin. Microbiol. 2012;50(7):2263–2271. doi: 10.1128/jcm.06599-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Guo S., Dipietro L.A. Factors affecting wound healing. J. Dent. Res. 2010;89(3):219–229. doi: 10.1177/0022034509359125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Ersanli C., Tzora A., Voidarou C.C., Skoufos S., Zeugolis D.I., Skoufos I. Biodiversity of skin microbiota as an important biomarker for wound healing. Biology. 2023;12(9) doi: 10.3390/biology12091187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Zielińska M., Pawłowska A., Orzeł A., Sulej L., Muzyka-Placzyńska K., Baran A., et al. Wound microbiota and its impact on wound healing. Int. J. Mol. Sci. 2023;24(24) doi: 10.3390/ijms242417318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wang Y., Shao T., Wang J., Huang X., Deng X., Cao Y., et al. An update on potential biomarkers for diagnosing diabetic foot ulcer at early stage. Biomed. Pharmacother. 2021;133 doi: 10.1016/j.biopha.2020.110991. [DOI] [PubMed] [Google Scholar]
- 22.Johnson T.R., Gómez B.I., McIntyre M.K., Dubick M.A., Christy R.J., Nicholson S.E., et al. The cutaneous microbiome and wounds: new molecular targets to promote wound healing. Int. J. Mol. Sci. 2018;19(9) doi: 10.3390/ijms19092699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wolcott R., Sanford N., Gabrilska R., Oates J.L., Wilkinson J.E., Rumbaugh K.P. Microbiota is a primary cause of pathogenesis of chronic wounds. J. Wound Care. 2016;25(Sup10):S33–s43. doi: 10.12968/jowc.2016.25.Sup10.S33. [DOI] [PubMed] [Google Scholar]
- 24.Liu C., Ponsero A.J., Armstrong D.G., Lipsky B.A., Hurwitz B.L. The dynamic wound microbiome. BMC Med. 2020;18(1):358. doi: 10.1186/s12916-020-01820-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wolcott R.D., Hanson J.D., Rees E.J., Koenig L.D., Phillips C.D., Wolcott R.A., et al. Analysis of the chronic wound microbiota of 2,963 patients by 16S rDNA pyrosequencing. Wound Repair Regen. : official publication of the Wound Healing Society [and] the European Tissue Repair Society. 2016;24(1):163–174. doi: 10.1111/wrr.12370. [DOI] [PubMed] [Google Scholar]
- 26.Pang M., Zhu M., Lei X., Chen C., Yao Z., Cheng B. Changes in foot skin microbiome of patients with diabetes mellitus using high-throughput 16S rRNA gene sequencing: a case control study from a single center. Med Sci Monit. 2020;26 doi: 10.12659/msm.921440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zou M., Cai Y., Hu P., Cao Y., Luo X., Fan X., et al. Analysis of the composition and functions of the microbiome in diabetic foot osteomyelitis based on 16S rRNA and metagenome sequencing Technology. Diabetes. 2020;69(11):2423–2439. doi: 10.2337/db20-0503. [DOI] [PubMed] [Google Scholar]
- 28.Patel B.K., Patel K.H., Huang R.Y., Lee C.N., Moochhala S.M. The gut-skin microbiota Axis and its role in diabetic wound healing-A review based on current literature. Int. J. Mol. Sci. 2022;23(4) doi: 10.3390/ijms23042375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Zhang S., Li S., Huang J., Ding X., Qiu Y., Luo X., et al. Gram-negative bacteria and lipopolysaccharides as risk factors for the occurrence of diabetic foot. J. Clin. Endocrinol. Metab. 2023;108(10):2604–2614. doi: 10.1210/clinem/dgad178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Park J.U., Oh B., Lee J.P., Choi M.H., Lee M.J., Kim B.S. Influence of microbiota on diabetic foot wound in comparison with adjacent normal skin based on the clinical features. BioMed Res. Int. 2019;2019 doi: 10.1155/2019/7459236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Witte M.B., Barbul A. General principles of wound healing. Surg. Clin. 1997;77(3):509–528. doi: 10.1016/s0039-6109(05)70566-1. [DOI] [PubMed] [Google Scholar]
- 32.Murphy E.C., Frick I.M. Gram-positive anaerobic cocci--commensals and opportunistic pathogens. FEMS Microbiol. Rev. 2013;37(4):520–553. doi: 10.1111/1574-6976.12005. [DOI] [PubMed] [Google Scholar]
- 33.Min K.R., Galvis A., Baquerizo Nole K.L., Sinha R., Clarke J., Kirsner R.S., et al. Association between baseline abundance of Peptoniphilus, a Gram-positive anaerobic coccus, and wound healing outcomes of DFUs. PLoS One. 2020;15(1) doi: 10.1371/journal.pone.0227006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sharma G., Garg N., Hasan S., Shirodkar S. Prevotella: an insight into its characteristics and associated virulence factors. Microb. Pathog. 2022;169 doi: 10.1016/j.micpath.2022.105673. [DOI] [PubMed] [Google Scholar]
- 35.Rojas-Tapias D.F., Brown E.M., Temple E.R., Onyekaba M.A., Mohamed A.M.T., Duncan K., et al. Inflammation-associated nitrate facilitates ectopic colonization of oral bacterium Veillonella parvula in the intestine. Nat Microbiol. 2022;7(10):1673–1685. doi: 10.1038/s41564-022-01224-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ahn J.R., Lee S.H., Kim B., Nam M.H., Ahn Y.K., Park Y.M., et al. Ruminococcus gnavus ameliorates atopic dermatitis by enhancing Treg cell and metabolites in BALB/c mice. Pediatr. Allergy Immunol. : official publication of the European Society of Pediatric Allergy and Immunology. 2022;33(1) doi: 10.1111/pai.13678. [DOI] [PubMed] [Google Scholar]
- 37.Wang Y., Zhang H., Ma G., Tian Z., Wang B. The contribution of intestinal Streptococcus to the pathogenesis of diabetic foot ulcers: an analysis based on 16S rRNA sequencing. Int. Wound J. 2022;19(7):1658–1668. doi: 10.1111/iwj.13766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Delanghe L., Spacova I., Van Malderen J., Oerlemans E., Claes I., Lebeer S. The role of lactobacilli in inhibiting skin pathogens. Biochem. Soc. Trans. 2021;49(2):617–627. doi: 10.1042/bst20200329. [DOI] [PubMed] [Google Scholar]
- 39.Fijan S., Frauwallner A., Langerholc T., Krebs B., Ter Haar Née Younes J.A., Heschl A., et al. Efficacy of using probiotics with antagonistic activity against pathogens of wound infections: an integrative review of literature. BioMed Res. Int. 2019;2019 doi: 10.1155/2019/7585486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mohammedsaeed W., Cruickshank S., McBain A.J., O'Neill C.A. Lactobacillus rhamnosus GG lysate increases Re-epithelialization of keratinocyte scratch assays by promoting migration. Sci. Rep. 2015;5 doi: 10.1038/srep16147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kim J.H., Son S.M., Park H., Kim B.K., Choi I.S., Kim H., et al. Taxonomic profiling of skin microbiome and correlation with clinical skin parameters in healthy Koreans. Sci. Rep. 2021;11(1) doi: 10.1038/s41598-021-95734-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Travis D.J., Bradbury J., Benkendorff K. Toward non-invasive collection methods for sampling the microbiome of diabetic foot ulcers. Int. Wound J. 2023;20(9):3731–3737. doi: 10.1111/iwj.14267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Liao L., Lin D., Liu Z., Gao Y., Hu K. A case of meningitis caused by Ralstonia insidiosa, a rare opportunistic pathogen. BMC Infect. Dis. 2023;23(1):548. doi: 10.1186/s12879-023-08506-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hung S.Y., Yeh Y.M., Chiu C.H., Armstrong D.G., Lin C.W., Yang H.M., et al. Microbiome of limb-threatening diabetic foot ulcers indicates the association of fastidious Stenotrophomonas and major amputation. Journal of microbiology, immunology, and infection = Wei mian yu gan ran za zhi. 2024;57(1):156–163. doi: 10.1016/j.jmii.2023.10.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Freitas R., Vasques A.C.J., da Rocha Fernandes G., Ribeiro F.B., Solar I., Shivappa N., et al. Gut bacterial markers involved in association of dietary inflammatory index with visceral adiposity. Nutrition. 2024;122 doi: 10.1016/j.nut.2024.112371. [DOI] [PubMed] [Google Scholar]
- 46.Li J., Yang G., Zhang Q., Liu Z., Jiang X., Xin Y. Function of Akkermansia muciniphila in type 2 diabetes and related diseases. Front. Microbiol. 2023;14 doi: 10.3389/fmicb.2023.1172400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Makeri D., Eilu E., Odoki M., Agwu E. A systematic review of the microbial landscape of diabetic foot ulcers in Uganda. Infect. Drug Resist. 2024;17:143–151. doi: 10.2147/idr.S446838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Alhussain F.A., Yenugadhati N., Al Eidan F.A., Al Johani S., Badri M. Risk factors, antimicrobial susceptibility pattern and patient outcomes of Pseudomonas aeruginosa infection: a matched case-control study. Journal of infection and public health. 2021;14(1):152–157. doi: 10.1016/j.jiph.2020.11.010. [DOI] [PubMed] [Google Scholar]
- 49.Zhu L., Wang K., Wu X., Zheng H., Liao X. Association of specific gut microbiota with polyethylene microplastics caused gut dysbiosis and increased susceptibility to opportunistic pathogens in honeybees. Sci. Total Environ. 2024;918 doi: 10.1016/j.scitotenv.2024.170642. [DOI] [PubMed] [Google Scholar]
- 50.Martín R., Rios-Covian D., Huillet E., Auger S., Khazaal S., Bermúdez-Humarán L.G., et al. Faecalibacterium: a bacterial genus with promising human health applications. FEMS Microbiol. Rev. 2023;47(4) doi: 10.1093/femsre/fuad039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Huang Y., Wang Z., Ye B., Ma J.H., Ji S., Sheng W., et al. Sodium butyrate ameliorates diabetic retinopathy in mice via the regulation of gut microbiota and related short-chain fatty acids. J. Transl. Med. 2023;21(1):451. doi: 10.1186/s12967-023-04259-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Jaber D., Younes N., Khalil E., Albsoul-Younes A., Zawiah M., Al-Bakri A.G. Studying microbial ecology of diabetic foot infections: significance of PCR analysis for prudent antimicrobial stewardship. Int. J. Low. Extrem. Wounds. 2024 doi: 10.1177/15347346241230288. [DOI] [PubMed] [Google Scholar]
- 53.Dong C., Yang Y., Wang Y., Hu X., Wang Q., Gao F., et al. Gut microbiota combined with metabolites reveals unique features of acute myocardial infarction patients different from stable coronary artery disease. J. Adv. Res. 2023;46:101–112. doi: 10.1016/j.jare.2022.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Yang C., Mai J., Cao X., Burberry A., Cominelli F., Zhang L. ggpicrust2: an R package for PICRUSt2 predicted functional profile analysis and visualization. Bioinformatics. 2023;39(8) doi: 10.1093/bioinformatics/btad470. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Raw 16S rRNA gene sequencing data are openly available in a public repository that issues datasets with accession number PRJNA1105325 in the NCBI SRA database (https://dataview.ncbi.nlm.nih.gov/object/PRJNA1105325?reviewer=6e2lh8k8b5f10kv2p2926bq6d2). Additionally, other data used and/or analyzed in the current study are available from the corresponding author upon reasonable request.