Key Points
G3-p239-Nb55 and fenobody-55 can block p239 attachment and HEV infection of host cells.
Fenobody-55 can completely neutralize HEV in vivo with nonenveloped HEV.
The epitope was, to our knowledge, a novel conformational and a highly conserved one.
Visual Abstract
Abstract
Hepatitis E virus (HEV) is a worldwide zoonotic and public health concern. The study of HEV biology is helpful for designing viral vaccines and drugs. Nanobodies have recently been considered appealing materials for viral biological research. In this study, a Bactrian camel was immunized with capsid proteins from different genotypes (1, 3, 4, and avian) of HEV. Then, a phage library (6.3 × 108 individual clones) was constructed using peripheral blood lymphocytes from the immunized camel, and 12 nanobodies against the truncated capsid protein of genotype 3 HEV (g3-p239) were screened. g3-p239-Nb55 can cross-react with different genotypes of HEV and block Kernow-C1/P6 HEV from infecting HepG2/C3A cells. To our knowledge, the epitope recognized by g3-p239-Nb55 was determined to be a novel conformational epitope located on the surface of viral particles and highly conserved among different mammalian HEV isolates. Next, to increase the affinity and half-life of the nanobody, it was displayed on the surface of ferritin, which can self-assemble into a 24-subunit nanocage, namely, fenobody-55. The affinities of fenobody-55 to g3-p239 were ∼20 times greater than those of g3-p239-Nb55. In addition, the half-life of fenobody-55 was nine times greater than that of g3-p239-Nb55. G3-p239-Nb55 and fenobody-55 can block p239 attachment and Kernow-C1/P6 infection of HepG2/C3A cells. Fenobody-55 can completely neutralize HEV infection in rabbits when it is preincubated with nonenveloped HEV particles. Our study reported a case in which a nanobody neutralized HEV infection by preincubation, identified a (to our knowledge) novel and conserved conformational epitope of HEV, and provided new material for researching HEV biology.
Introduction
Hepatitis E virus (HEV) is the causative agent of hepatitis E, a usually self-limiting disease. However, it can cause an estimated 20 million human infections, leading to ∼52,100 deaths annually, as reported by the World Health Organization (https://www.who.int/publications/i/item/9789241565455). Notably, the mortality of HEV infection in pregnant women can reach up to 30% (1). HEV is mainly transmitted through the fecal-oral route through contaminated food, drinking water, or infected animal products (2). Although Hecolin is the exclusive HEV vaccine approved in China, it has not been applied on a large scale. The biology of HEV remains to be elucidated for the development of highly effective vaccines and drugs.
HEV, a quasi-enveloped, single-stranded, and positive-sense RNA virus, is classified as a member of the Hepeviridae family. It included two subfamilies, Orthohepevirinae (including four different genera: Avihepevirus, Chirohepevirus, Paslahepevirus, and Rocahepevirus) and Parahepevirinae (3, 4). Eight HEV genotypes (HEV-1 to HEV-8) belonging to Paslahepevirus balayani have been identified (4, 5). HEV-1 and HEV-2 exclusively infect humans and are common in developing countries (6). HEV-3, HEV-4, and HEV-7 are zoonotic pathogens with a broad host range and can cause sporadic cases of zoonotic hepatitis E in developed countries (7). The HEV genome is ∼7.2 kb long with at least three open reading frames (ORFs). Among the three ORFs, ORF2, encoding a viral capsid protein, also known as the ORF2 protein, plays an important role in viral assembly by entering host cells and inducing protective immune responses (8). Therefore, the ORF2 protein is regarded as a good candidate for the design of HEV subunit vaccines. The production of Abs against the HEV ORF2 protein and the mapping of epitopes in the protein is helpful for HEV biological research and the design of viral vaccines and drugs. The complete ORF2 protein is composed of 660 aa, with extensive homology (>85%) among the four major mammalian genotypes (genotyped 1–4) (9). Among the different regions of the ORF2 protein, the aa 368–606 region of the HEV ORF2 protein, named p239, is highly conserved among the different genotypes of HEV from mammals and can reserve immunogenic virion-like epitopes to induce an immune response (9). Currently, the HEV vaccine Hecolin, which is based on p239 of genotype 1 HEV expressed by Escherichia coli, has been shown to be well tolerated and to provide 100% protection from HEV infection (10, 11). Therefore, p239 can be used as a target for producing Abs against HEV and researching the antigenicity of the HEV ORF2 protein.
Nanobodies are an appealing alternative to traditional Abs. The VHHs are from the V domains of H chains of Abs (VHHs) of camelid animals, with a natural absence of the L chain and the constant H chain 1 domain of the H chain (12). Owing to its unique properties, such as its small size, solubility in an aqueous solvent, and ease of modification, it has been developed as a tool in biomedicine and biotechnology research. Nanobodies have been widely used for biological research, targeted drug delivery, bioimaging, and treatment (13, 14).
Considering the broad prospects of nanobodies and antigenicity of HEV p239, nanobodies against HEV p239 were screened and produced by phage display technology in this study. Then, an epitope recognized by one nanobody, which is common among the different HEV genotypes and located on the surface of nonenveloped HEV (neHEV) particles, was identified. Previously, it was reported that fenobody, a nanobody-derived product that can self-assemble into a ferritin structure to display 24 U of nanobodies, could improve the affinity of the nanobody for the Ag and its half-life (15). Based on this, a fenobody against p239 from genotype 3 of HEV was designed and produced. The neutralizing activities of the fenobody in vitro and in vivo were compared with those of the monovalent nanobody. Our study identified a (to our knowledge) novel common epitope among p239 proteins from different genotypes of HEV, revealed that a nanobody neutralizes the different genotypes of HEV, and investigated HEV biology.
Materials and Methods
Cells, viruses, and vectors
HepG2/C3A cells (ATCC CRL-10741) were cultured in MEM (Life Technologies, Gaithersburg, MD) supplemented with 10% FBS (Life Technologies, Gaithersburg, MD), penicillin (100 U/ml), and streptomycin (100 mg/ml) (Life Technologies, Invitrogen, Grand Island, NY) at 37°C in 5% CO2. The HEV strain Kernow-C1/P6 (genotype 3) can propagate in HepG2/C3A cells and was kindly provided by S.U. Emmerson (16). To construct the VHH library, we applied the pMECS vectors as previously described (17). The pCMV-N1-HRP vectors were designed and constructed as previously described to express nanobody-HRP fusion proteins (18).
Bactrian camel immunization and VHH library construction
Bactrian camel immunization and VHH library construction were performed as described in a previous study (17). To identify the nanobody recognizing common epitopes of HEV, we used five recombinant p239 proteins from different genotypes of HEV for immunization. In brief, a 4-y-old male Bactrian camel was immunized with a mixture of p239, including g3-p239 (HEV-3, Kernow-C1 strain, GenBank JQ679013), g1-p239 (HEV-1, Sar55 strain, GenBank AF444002), g3-rabbit p239 (rabbit HEV-3, CHN-SX-rHEV strain, GenBank KX227751), g4-p239 (HEV-4, CHN-SD-sHEV strain, GenBank KF176351), and ap237 (avian HEV, CaHEV strain, GenBank JN997392), via a s.c. route five times every 2 wk. The gene encoding p239 of different genotypes was ligated into the plasmid pET-21b (+) without fused tags. The expression and purification of p239 proteins were performed as previously described (Supplemental Fig. 1) (19). These p239 proteins were emulsified with CFA at the first immunization and with IFA at the latter four immunizations. After the camel was immunized five times, serum samples were collected to determine the titer of Abs against the HEV ORF2 protein with a commercial ELISA kit (Wantai Biological Pharmacy, Beijing, China). Rabbit anti-camel IgG was used to replace the secondary Ab in the kit. On the fifth day after the last immunization, PBMCs were isolated from the blood using Leucosep tubes (Greiner Bio-One, Frickenhausen, Germany) and divided into tubes containing 107 cells. Total RNA was extracted from PBMCs with an RNeasy Plus Mini Kit (QIAGEN, Dusseldorf, Germany) according to the manufacturer’s instructions. Then, cDNA was synthesized with total RNA as a template using the SuperScript III First-Strand Synthesis System for RT-PCR Kit (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. After that, nested PCR was conducted to amplify the VHH gene with the primers CALL001, CALL002, VHH-FOR, and VHH-REV, as previously described (20). The nested PCR products were then purified with an EasyPure Quick Gel Extraction Kit (TransGen Biotech, Beijing, China) and ligated into the phagemid vector pMECS by digestion with both the NotI and PstI enzymes. The ligated products were electrotransformed into E. coli TG1 cells and shaken at 220 rpm for 40 min. Finally, the library was harvested after the transformed TG1 cells were cultured at 37°C for 6 h on Luria-Bertani/ampicillin-glucose agar plates. Meanwhile, 100 μl of transformed TG1 cells was diluted to 10−2, 10−3, and 10−4 and placed on Luria-Bertani/ampicillin-glucose agar plates. The total number of colonies was counted to calculate the transformation efficiency. A total of 48 clones were randomly selected to determine the diversity of the library by PCR with the primers MP57 and GIII (Supplemental Table I).
Nanobody screening
Kernow-C1/P6 (genotype 3 HEV) is an HEV strain that can proliferate strongly in vitro. Then, after the confirmation of neutralizing activity with virus particles in vitro, g3-p239 from Kernow-C1/P6 was used as the Ag to screen nanobodies. After screening, all the screened nanobodies were determined to cross-react with other genotypes of HEV. To obtain specific nanobodies against g3-p239, we conducted three rounds of screening and phage rescue with indirect ELISA, as described previously (17). In brief, the ELISA plate was coated with purified g3-p239 (1 μg/well) in PBS. After the plates were blocked with 2.5% skim milk in PBS containing 0.5% Tween 20 (PBS′T), the recombinant phage particles were added and incubated for 2 h. After the plates were washed with PBS, the binding phage particles were eluted with trimethylamine (0.1 M, 100 μl). Then, the eluted particles were immediately neutralized with isopycnic 1 M Tris-HCl (pH 7.4). The titers of the recombinant phage particles were calculated during each screening round. After three rounds of screening, the specific phage particles binding to g3-p239 were enriched. In addition, 96 clones were randomly selected from the plates to determine the titers after three rounds of growth in liquid culture. Then, the periplasmic extracts of 96 colonies were extracted and tested by ELISA with purified g3-p239 as the coating Ag. The ELISA-positive colonies whose OD at 450 nm (OD450) was three times greater than that of the negative control were selected for sequencing by Tsingke Biotech Company (Beijing, China).
Expression and purification of nanobodies and fenobodies
The nanobodies were designed for monovalent and polyvalent forms and produced by a bacterial system to analyze neutralization. First, the VHH gene encoding the nanobody was amplified with the primer pair pET21b-Nb-F/R (Supplemental Table I) using recombinant pMECS vectors as templates (21). The VHH gene was directly cloned and inserted into the pET-21b vector with a His tag for the monovalent nanobody. For the polyvalent forms, as previously described, ferritin with a nanobody fusion protein can self-assemble into a nanocage, in which 24 nanobodies are displayed on the surface; this material was named fenobody (15). Ferritin from Pyrococcus furiosus (PDB: 2JD8_P), which is composed of 174 aa residues, was employed as the structural framework. The VHH gene was linked to the C terminus with the (GGGS)3 linker to substitute for the ε domain (aa 147–174) of ferritin (15). Then, the fused genes were cloned and inserted into the pET-28a vector for expression of the fenobody. The primers (Supplemental Table I) and operating procedures used were identical to those described previously (22).
After the recombinant plasmids were constructed, the positive ones were transformed into E. coli BL21 (DE3)-competent cells (TransGen Biotech, Beijing, China) for expression. When the OD600 of the positive bacteria reached 0.6–0.8, isopropyl-β-d-thiogalactoside (1 mM) was added, and the mixture was shaken at 16°C and 150 rpm for 24 h. Then, the bacterial cells were collected by centrifugation and resuspended in lysis buffer (50 mM NaH2PO4, 300 mM NaCl, 10 mM imidazole, pH 8.0) for purification with Ni-NTA Beads (Smart-Life Sciences, Changzhou, China) according to the manufacturer’s instructions.
Biophysical characterization of the monovalent nanobody and fenobody
The biophysical characteristics of the monovalent nanobody and fenobody were investigated as previously described (15). First, to determine whether the fenobody formed into a nanocage, we negatively stained the purified fenobody with uranyl acetate and examined with a JEM-1400-FLASH transmission electron microscope at 80 kV.
To determine the affinity between g3-p239 and monovalent nanobodies or fenobodies, we performed biolayer interferometry (BLI) analysis (Sino Biological, Beijing, China) based on the manufacturer’s instructions. In brief, purified g3-p239 (50 μg/ml) was coupled to the biosensor AR2G in sodium acetate solution (pH 4.0). Then, the monovalent nanobody and fenobody were diluted separately to 5 and 30 μg/ml, respectively, followed by 2-fold gradient dilutions. After incubation, the solidified complex was dissociated in 2× PBS. The equilibrium Kd was calculated by Data Analysis 12.0.
To determine the extent of the half-life of the monovalent nanobody and fenobody, we produced FITC-labeled monovalent nanobody and fenobody as previously described (15). In brief, 2 nmol FITC (MedChemExpress, Rahway, NJ) and 50 nM monovalent nanobody or fenobody were mixed in 1 ml of carbonate/bicarbonate buffer containing 100 mM carbonate (pH 9.0). Then, the mixtures were incubated in the dark for 2 h at room temperature. To remove the excess FITC, we filtered the mixtures through an ultrafiltration device (Sartorius, Gottingen, Germany). After the FITC-nanobody and FITC-fenobody were quantified with VICTOR X Series Multilabel Plate Readers (PerkinElmer), they (2 nmol FITC per mouse) were separately injected into the female BALB/c mice via the tail vein. At 15, 30, 45, 60, 120, 240, 360, 720, and 1440 min after the injection, blood samples were collected from the tail vein, and the fluorescence in the blood was determined (Ex 485/Em 535, 1.0 s).
To determine the binding of monovalent nanobodies and fenobodies to natural HEV particles, we designed and performed immunocapture RT-PCR. In brief, the ELISA plates were coated with the nanobody (2.1 μM = 3.2 μg) and fenobody (2.1 μM = 7.25 μg). After blocking, the plates were supplemented with HEV viral suspension (200 μl) and incubated at 4°C overnight. These particles were obtained from the supernatant of cells inoculated with Kernow-C1/P6 and pretreated or not pretreated with 0.1% sodium deoxycholate and 0.1% trypsin as previously described (23). After the plates were washed with PBS′T, RNAiso Plus (TAKARA, Beijing, China) was added to the wells to extract viral RNA. Then, the extracted RNA was reverse transcribed and amplified via nested RT-PCR using the PrimeScript One-Step RT-PCR Kit Ver.2 and Premix Taq (Ex Taq Version 2.0 plus dye) (TAKARA) as described previously (24) (Supplemental Table I). Finally, the PCR products with the expected size of 348 bp were analyzed by electrophoresis in a 1.0% agarose gel containing ethidium bromide. In addition, absolute quantitative real-time PCR (RT-qPCR) was performed to determine the number of HEV RNA copies in the captured particles (25).
Expression of the nanobody-HRP fusion protein
Nanobodies fused with HRP were designed and produced based on a previous study (18) to identify the cross-reaction between the nanobodies and different genotype epitopes. In brief, the VHH genes from the recombinant pMECS plasmids were obtained, cloned, and inserted into the pCMV-N1-HRP vector by digestion with PstI and NotI enzymes (New England Biolabs, Beverly, MA). After sequencing, the positive plasmids were transfected into HEK293T cells with Lipo8000 Transfection Reagent (Beyotime Biotechnology, Shanghai, China) according to the manufacturer’s instructions. At 72 h posttransfection, the medium containing the nanobody-HRP fusion proteins was harvested and filtered through 0.45-μm cellulose acetate membranes for further study. Then, these nanobody-HRP fusion proteins were used as the primary Abs for direct ELISA.
ELISA
Three kinds of ELISA, indirect ELISA, direct ELISA, and competitive ELISA (cELISA), were used in this study. Indirect ELISA was performed to detect the titers of Abs against the ORF2 protein in the immunized camel and to determine the binding of the monovalent nanobody and fenobody to g3-p239. In brief, 96-well ELISA plates (Thermo Fisher Scientific, Waltham, MA) were coated with purified g3-p239 (400 ng/well). For titrations, different dilutions of serum samples were added to the plates and incubated for 1 h at 37°C. Then, rabbit anti-camel Abs and HRP-labeled goat anti-rabbit Abs (Jackson ImmunoResearch Laboratories, West Grove, PA) were added. For specific binding, the purified monovalent nanobody (5 μM) and fenobody (5 μM) were added to the wells and incubated for 1 h at 37°C. The mAb 2C7 (5 μM), which recognizes the common epitope of different genotypes of HEV, was used as the positive control (19). Then, the anti-His tag mAb (1 μg/ml; Sino Biological, Beijing, China) and HRP-labeled goat-mouse Abs (1:5000; Jackson Immunoresearch Laboratories) were added. After incubation and washing, tetramethylbenzidine was added to the plates for color reactions, and the plates were incubated for 15 min at room temperature. After the reactions were stopped with 3 M H2SO4, the OD450 was read with an automatic ELISA plate reader.
Direct ELISA was used to analyze nanobody-HRP fusion protein cross-reactivity with HEV ORF2 proteins of other genotypes and for epitope mapping. For the cross-reaction, g1-p239, g3-rabbit p239, g4-p239, and ap237 were used to coat the ELISA plates. For epitope mapping, g3-p239 and mutated proteins were coated onto plates. Then, the nanobody-HRP fusion proteins were added to the wells. After washing, tetramethylbenzidine was added for the color reaction, and the reactions were stopped with 3 M H2SO4. Finally, the OD450 was read with an automatic ELISA plate reader.
cELISA was used to identify whether the epitope recognized by the nanobodies also existed in the natural viral particles (26). First, g3-p239 (200 ng/well) was coated on ELISA plates. After blocking, the positive and negative serum samples for Abs against HEV were mixed with the diluted nanobodies-HRP fusion protein (1:100). After 1 h of incubation at 37°C for 1 h and washing, the OD450 was read. The mean percent inhibition (PI) value was calculated using the following formula: PI (%) = (1 – [OD450 value of positive serum group/OD450 value of negative serum group]) × 100%.
Analysis of monovalent nanobodies and fenobodies blocking g3-p239 to attach to cells
Previously, it was shown that the p239 protein from genotype 1 HEV can mimic the natural viral particles attached to host cells (9). Therefore, the effects of monovalent nanobodies and fenobodies blocking p239 on cell attachment were analyzed. First, the cytotoxicity of the monovalent nanobody and fenobody to HepG2/C3A cells was evaluated with a Cell Counting Kit-8 (Biosharp, Guangzhou, China) according to the manufacturer’s instructions. In addition, the optimal amount of g3-p239 attached to the HepG2/C3A cells was also determined based on previous descriptions by IFA and Western blotting (19). In brief, a series of g3-p239 (1, 2, 4, 8, 16, or 32 μg) in 1 ml was incubated with HepG2/C3A cells at 4°C for 1 h and then at 37°C for 1 h (Supplemental Fig. 3). After these amounts were determined, g3-p239 was separately mixed with different concentrations (1, 2, 4, and 8 µM) of nanobody and fenobody in 500 μl of MEM, and the mixtures were incubated at 4°C overnight. Then, the mixtures were incubated with HepG2/C3A cells seeded onto 24-well cell plates for 2 h at 37°C and 5% CO2. The mAb 1B5, which recognizes a common epitope of different genotypes of mammalian HEV isolates and blocks g3-p239 attachment and swine HEV infection in HepG2 cells, was used as a positive control at 10 μg (10 μg/ml) (19). In addition, antisera collected from pigs that were naturally infected with genotype 4 swine HEV were used as a positive control (27). A monovalent nanobody and fenobody targeting Newcastle disease virus (NDV) were used as negative controls (22). After the plates were washed with PBS, the cells were prepared for IFA and Western blotting analysis to determine the amount of g3-p239. In addition, the attachment of p239 from other genotypes to HepG2/C3A cells was also verified to be blocked by monovalent nanobodies and fenobodies, as mentioned earlier.
In vitro neutralization analysis of the monovalent nanobody and fenobody
neHEV particles were obtained from the lysate of HepG2/C3A cells inoculated with Kernow-C1/P6. Similarly, 10 μg (10 μg/ml) of mAb 1B5 was used as a positive control. Then, the particles (2.48 × 106 copies) were incubated separately with different concentrations (0.1, 1, 2.5, and 5 µM) of nanobody and fenobody at 4°C overnight. The next day, the mixtures were inoculated into HepG2/C3A cells and cultured for 6 h at 37°C and 5% CO2. After that, the cells were washed three times with PBS and cultured for 6 d at 37°C and 5% CO2 in fresh medium supplemented with 2% FBS, 100 U/ml penicillin, and 0.1 mg/ml streptomycin. Then, IFA and relative RT-qPCR were performed to analyze the viral amounts.
Indirect immunofluorescence assay
After the HepG2/C3A cells were incubated with the mixtures described earlier, the cells were fixed with 4% paraformaldehyde. Then, the cells were permeabilized with 0.25% Triton X-100 in PBS at room temperature for 15 min. After being blocked with SuperBlock (TBS) blocking buffer (Thermo Scientific), the cells were incubated with an anti-HEV ORF2 protein Ab (1E6; Merck-Millipore, Billerica, MA) at 37°C for 1 h. After being washed with PBS, the cells were stained with Alexa Fluor 488–labeled goat anti-mouse IgG (Jackson ImmunoResearch Laboratories) for 1 h. Finally, the cell nuclei were stained with DAPI and then observed via fluorescence microscopy (Leica).
Western blotting
The treated HepG2/C3A cells described earlier were lysed with 2× Laemmli Sample Buffer (Bio-Rad, Hercules, CA). After the cell lysates were subjected to SDS-PAGE, they were transferred to polyvinylidene difluoride membranes with a Mini Trans-Blot apparatus (100 V, 1 h; Bio-Rad). After being blocked with blocking buffer (2.5% skim milk powder in PBS′T), the membranes were incubated with 1 μg/ml mAb 1B5 (19) and then with an HRP-conjugated goat anti-mouse Ab (1:5000). The membranes were visualized with an eECL Western Blot Kit (Cowin Bio, Beijing, China) and imaged.
Relative RT-qPCR
The expression levels of the GAPDH and HEV ORF1 genes were determined by RT-qPCR. The RNA of treated HepG2/C3A cells was extracted with RNAiso Plus (TRIzol) (TaKaRa, Tokyo, Japan) and then reverse transcribed into cDNA with PrimeScript RT Master Mix (TaKaRa). The primers used in this study were the same as those described in Supplemental Table I (28). The reaction system was composed of 7 μl of 2× Fast qPCR Master Mixture (Dining, Beijing, China), 0.375 μl of primers (10 μM), and then DNase/RNase-free water (Solarbio Life Science, Beijing, China) to 15 μl. The relative RT-qPCR was conducted on an Applied Biosystem StepOnePlus Real-Time PCR System (Applied Biosystems, Foster City, CA).
Neutralization analysis of the monovalent nanobody and fenobody in vivo
To identify the neutralization of monovalent nanobodies and fenobody in vivo, we used an animal model of rabbit HEV infection, as previously described, with some modifications (29). A total of 40 specific pathogen-free (SPF) 6-wk-old New Zealand White rabbits were purchased (Chengdu Dossy Experimental Animals, Chengdu, China) and then randomly divided into eight groups (n = 5/group). For the monovalent nanobody and fenobody, 133 μM nanobody and fenobody were preincubated with neHEV particles (1.1 × 104 genomic equivalents/ml) at 4°C overnight according to the quantity of nanobody previously described (29). The two groups were separately named the preincubated nanobody and fenobody groups. Similarly, for the blank control, negative control, and challenged groups, 1 ml of PBS, 133 μM NDV-Nb96, 133 μM NDV-fenobody-96, and rabbit HEV (1.1 × 104 genomic equivalents/rabbit) in PBS were preincubated. NDV-Nb96 and NDV-fenobody-96 were produced in our laboratory, and the same system used for the nanobody and fenobody in this study was used. Then, the mixtures were inoculated into the rabbits via the ear vein. To determine whether monovalent nanobodies and fenobodies have therapeutic effects on HEV infection, we infected SPF rabbits with neHEV. Then, g3-p239-Nb55 and fenobody-55 were injected via the ear vein after 24 h at accordant doses in the preincubated groups. The two groups were named the postinjection nanobody and fenobody groups.
Blood and fecal samples were collected before inoculation at 0 wk postinoculation (wpi) and then at 1, 2, 3, 4, and 5 wpi. The levels of HEV Abs in the serum samples were tested by indirect ELISA. Fecal samples were tested for HEV RNA with semiquantitative nested RT-PCR (30). Livers were also collected after the rabbits were euthanized at 5 wpi and fixed with 4% paraformaldehyde, after which three liver sections from each rabbit were stained with H&E to evaluate pathological and histopathological changes caused by HEV infection. In addition, to quantify the number of inflammatory cells in the liver, we counted three random 1-mm2 sections via ImageJ software.
Identification of the epitope recognized by the nanobody
To determine the epitope recognized by the nanobody, we first designed and expressed with the bacterial system a total of 14 truncated fragments of the HEV ORF2 protein, including aa 393–606, aa 380–606, aa 373–606, aa 415–606, aa 440–606, aa 453–606, aa 465–606, aa 523–606, aa 368–476, aa 393–476, aa 393–534, aa 393–558, aa 393–580, and aa 393–593. The primer pairs were designed and synthesized to amplify the eight genes encoding these fragments (Supplemental Table I). Then, these genes were separately ligated into pET-21b (+) and transformed into E. coli BL21 (DE3)-competent cells for expression based on the earlier methods. After purification with Ni-NTA beads, these proteins were used as Ags for direct ELISA of the nanobody-HRP fusion protein.
Determination of key motifs involved in the interaction between g3-p239 and the nanobody
To precisely define the motifs of the epitope, we separately predicted the secondary structures of g3-p239 and the nanobodies with PyMOL software to construct three-dimensional models according to the amino acid sequences. Then, the key residues involved in the interaction between g3-p239 and the nanobody were also simulated. A mutated nanobody with an HRP fusion protein was designed and expressed with the eukaryotic expression system mentioned earlier to further verify the possible binding regions. After harvesting, the mutated nanobody-HRP fusion protein was used as the primary Ab for ELISA detection.
In addition, the predicted key motif in g3-p239 was alanine (A), and the mutated genes were synthesized by Tsingke Biotech Company. Then, the mutated g3-p239 was expressed as described earlier and used as a coating protein for ELISA. The OD450 value was read and recorded. Finally, the conservation of the key amino acid for nanobody binding to g3-p239 among different genotypes of HEV was analyzed by sequence alignment with the MegAlign program.
Statistical analysis
Each experiment was independently repeated at least three times. Statistical significance was determined by Student t test when two groups were compared or by one-way ANOVA when more than two groups were compared. Asterisks indicate statistical significance as follows: *p < 0.05, **p < 0.01, ***p < 0.001; ns indicates no significant difference.
Ethics statement
The animal experiments were performed strictly per the recommendations in the Guide for the Care and Use of Laboratory Animals of Northwest A&F University. The protocols used for animal and serum collection were approved by the Institutional Animal Care and Use Committee of Northwest A&F University (approval no. 20221113/04).
Results
Construction of a VHH library containing HEV ORF2 proteins
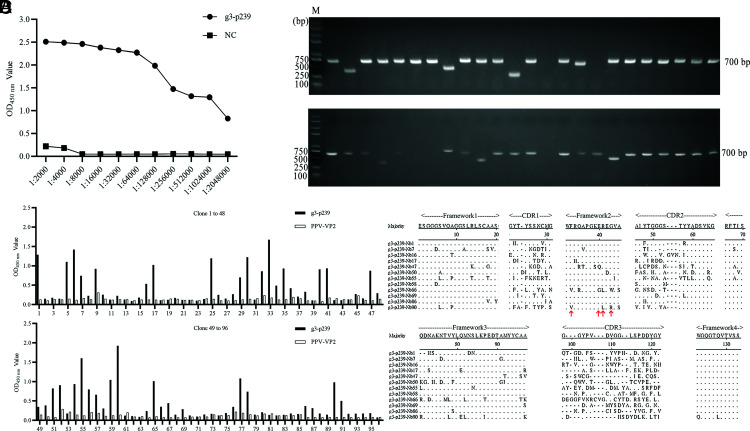
After the camel was immunized with p239 proteins from different genotypes of HEV five times, the ELISA results showed that the titers of Abs against HEV ORF2 proteins reached 1:2,048,000 (Fig. 1A). After the recombinant pMECS plasmid containing different VHH genes was electrotransformed into TG1 E. coli–competent cells, a phage-displayed VHH library consisting of ∼6.3 × 108 individual clones was successfully constructed. A total of 48 clones were randomly selected and tested by PCR to determine the insertion rate of the VHH genes, and the results showed that the insertion rate was ∼90% (Fig. 1B). Then, these positive clones were sequenced. The results showed that each clone contained a distinct VHH sequence, which confirmed the heterogeneity of the individual clones from the library (data not shown).
FIGURE 1.
Construction of the VHH library and screening of specific nanobodies against g3-p239 by phage display technology. (A) Titers of Abs against g3-p239 in the immunized camel after quintuple immunization with g1-p239, g3-p239, g4-p239, g3-rabbit p239, and ap237. (B) The percentage of positive clones among 48 randomly selected clones was determined by PCR. The expected size of the PCR products was ∼700 bp. (C) Identification of the periplasmic extracts from the 96 clones that specifically bind to g3-p239 via indirect ELISA. A total of 38 clones were identified as positive. (D) Alignments of amino acid sequences of 12 screened nanobodies. The sequences were grouped according to the variation in CDR3.
Screening of specific nanobodies against g3-p239
G3-p239 was the truncated HEV ORF2 protein (aa 368–606) from the genotype 3 HEV Kernow-C1/P6 strain. After three rounds of panning with g3-p239 as a coating Ag, phage particles carrying specific VHH genes encoding nanobodies against g3-p239 were strongly enriched (Table I). Then, the specific binding of the periplasmic extracts of 96 colonies randomly selected from the third-round panning plates to g3-p239 was tested via indirect ELISA. The results showed that a total of 38 colonies were positive, with the OD450 value of indirect ELISA being three times greater than that of the negative control (Fig. 1C). After these colonies were sequenced, sequence analysis revealed that 12 different sequences were screened according to variations within CDR3 regions (Fig. 1D), and the amino acid residues marked by red arrows were determined to be hydrophilic (Fig. 1D) (20).
Table I. Enrichment of phage particles carrying g3-p239–specific nanobodies during three rounds of panning.
| Round of Panning | Input Phage (PFUs/well) | P Output (PFUs/well) | N Output (PFUs/well) | Recovery | P/N |
|---|---|---|---|---|---|
| First round | 5 × 1010 | 8.3 × 106 | 2 × 105 | 1.66 × 10−4 | 41.5 |
| Second round | 5 × 1010 | 6.4 × 107 | 1.2 × 105 | 1.28 × 10−3 | 533 |
| Third round | 5 × 1010 | 4.2 × 106 | 7 × 102 | 8.4 × 10−5 | 6 × 103 |
N, negative; P, positive.
Expression and purification of 12 monovalent nanobodies against g3-p239 with a bacterial system
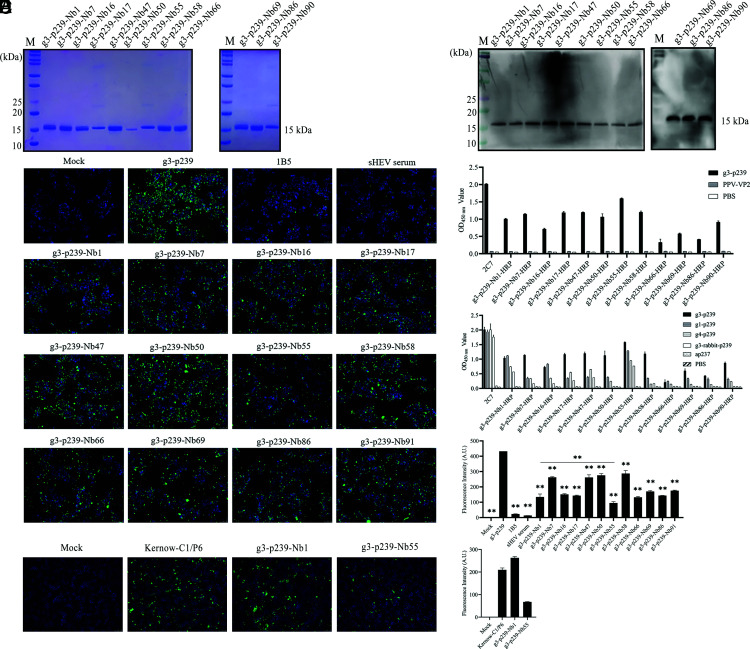
All 12 genes encoding nanobodies were successfully ligated into pET-21b. After the nanobodies were expressed using the E. coli system and purified with Ni-NTA columns, SDS-PAGE analysis showed that all 12 monovalent nanobodies were successfully expressed and purified with the expected size of 15 kDa (Fig. 2A). Using the anti-His mAb as the primary Ab, Western blot analysis further confirmed that the 12 His tags were successfully expressed (Fig. 2B). The 12 monovalent nanobodies were named g3-p239-Nb1, -Nb7, -Nb16, -Nb17, -Nb47, -Nb50, -Nb55, -Nb58, -Nb66, -Nb69, -Nb86, and -Nb90.
FIGURE 2.
Production and characterization of 12 nanobodies against different genotypes of HEV p239. (A) SDS-PAGE analysis of 12 monovalent nanobodies against g3-p239 expressed by the bacterial system. (B) Western blot analysis of the 12 nanobodies using an anti-His mAb as the primary Ab. (C) The binding capacity of 12 nanobody-HRP fusion proteins expressed in HEK293T cells with g3-p239 by direct ELISA. The PPV-VP2 protein was used as a negative control. (D) Cross-reaction of 12 nanobody-HRP fusion proteins with different genotypes of HEV p239s and ap237 by direct ELISA. (E) IFA analysis of the blocking effect of 12 nanobodies on the attachment of g3-p239 to HepG2/C3A cells. The nuclei were stained by DAPI, and the g3-p239 protein was stained by Alexa Fluor 488_labeled goat anti-mouse Ab. The fluorescence intensity was scanned with ImageJ software. The data were analyzed by one-way ANOVA using GraphPad software. (F) Analysis of g3-p239-Nb1 and g3-p239-Nb55 neutralizing HEV Kernow-C1/P6 infection in HepG2/C3A cells by IFA. The nuclei were stained by DAPI, and the virus was stained by Alexa Fluor 488_labeled goat anti-mouse Ab. The fluorescence intensity was scanned with ImageJ software. The experiment was independently repeated three times. The error bars indicate the mean ± SD. **p < 0.01.
Expression and characterization of 12 nanobody-HRP fusion proteins against g3-p239 in HEK293T cells
All 12 genes encoding nanobodies were successfully ligated into pCMV-N1-HRP. After the recombinant pCMV-N1-HRP plasmids were transfected into HEK293T cells, the supernatants containing the nanobody-HRP fusion proteins were collected at 72 h posttransfection. Direct ELISA using the supernatant as the primary Ab confirmed that the 12 nanobody-HRP fusion proteins were successfully expressed and specifically bound to g3-p239 (Fig. 2C). These fusion proteins were named g3-p239-Nb1-HRP, -Nb7-HRP, -Nb16-HRP, -Nb17-HRP, -Nb47-HRP, -Nb50-HRP, -Nb55-HRP, -Nb58-HRP, -Nb66-HRP, -Nb69-HRP, -Nb86-HRP, and -Nb90-HRP. Among these, g3-p239-Nb55-HRP had the highest affinity for g3-p239 according to the OD450 value of direct ELISA (Fig. 2C). In addition, the cross-reactivity between the 12 nanobody-HRP fusion proteins and ORF2 proteins from other HEV genotypes was also evaluated via direct ELISA. The results showed that g3-p239-Nb55-HRP simultaneously demonstrated exceptional affinity for p239 from genotype 1, 3, 4, and 3-rabbit HEVs (Fig. 2D). Then, using pig sera positive for Abs against g4 swine HEV as the competitive Ab, the results of cELISA showed that the binding of g3-p239 with g3-p239-Nb55-HRP can compete with that of serum Abs against swine HEV (Table II).
Table II. Detection of positive pig serum samples for anti-HEV Abs blocking 12 nanobody-HRP fusion proteins to bind to HEV capsid protein with cELISA.
| Sample | -Nb1 | -Nb7 | -Nb16 | -Nb17 | -Nb47 | -Nb50 | -Nb55 | -Nb58 | -Nb66 | -Nb69 | -Nb86 | -Nb91 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Positive (P) | 0.895 | 0.813 | 1.092 | 0.849 | 0.952 | 0.582 | 0.152 | 1.013 | 0.911 | 1.032 | 1.057 | 1.052 |
| Negative (N) | 1.097 | 1.023 | 1.131 | 1.067 | 1.060 | 1.055 | 0.988 | 1.024 | 1.084 | 1.117 | 1.076 | 1.134 |
| P/N | 0.816 | 0.795 | 0.965 | 0.795 | 0.898 | 0.552 | 0.154 | 0.990 | 0.841 | 0.923 | 0.982 | 0.927 |
| PI (%) | 18.38 | 20.46 | 3.47 | 20.50 | 10.20 | 44.84 | 84.57 | 1.05 | 15.94 | 7.66 | 1.78 | 7.25 |
g3-p239-Nb55 blocks the attachment of g3-p239 to cells and neutralizes infection caused by the HEV Kernow-C1/P6 strain
After 12 monovalent nanobodies were produced, they were first applied to verify whether they could block the attachment of g3-p239 to HepG2/C3A cells. According to the results in Supplemental Fig. 2, there was no cytotoxicity to the cells until 8 μM g3-p239-Nb55 was used. Then, 8 μM monovalent nanobodies was used, and 1B5 (10 µg) was used as the positive control. After mixtures of g3-p239 with different monovalent nanobodies were incubated with the cells, the IFA results and statistical analysis showed that all 12 nanobodies could block attachment (Fig. 2E). Notably, g3-p239-Nb1 and -Nb55 can block attachment to the greatest extent, and there was a significant difference between them (Fig. 2E).
Based on the earlier results of cross-reaction with nanobody-HRP fusion and blocking assays with monovalent nanobodies, g3-p239-Nb1 and -Nb55 were selected for evaluation of their neutralizing activity. After the mixtures of g3-p239-Nb1 and -Nb55 with Kernow-C1/P6 were separately inoculated into HepG2/C3A cells, the IFA results showed that the green fluorescence of g3-p239-Nb55 was lower than that of the negative control, indicating that g3-p239-Nb55 may have neutralized (Fig. 2F). However, the green fluorescence of the g3-p239-Nb1 group was the same as that of the group infected with only HEV (Fig. 2F). Therefore, g3-p239-Nb55 was selected for the following experiments.
Expression, purification, and identification of fenobody-55 against g3-p239
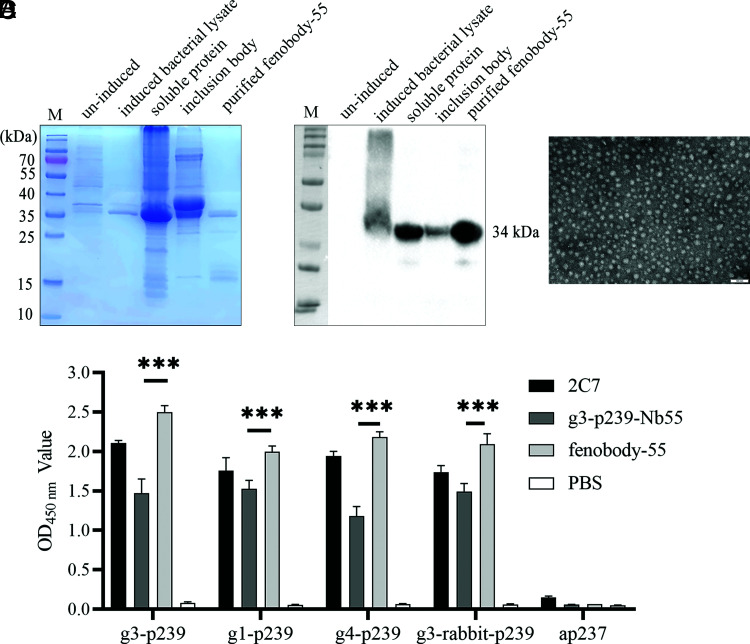
Previously, it was reported that the fenobody could form a nanocage, with 24 nanobodies located on the nanocage’s surface (15). In this study, ferritin fused with g3-p239-Nb55 (fenobody-55) was produced and compared with the neutralization of g3-p239-Nb55. First, SDS-PAGE and Western blot analysis showed that fenobody-55 was successfully expressed and purified with Ni-NTA columns (Fig. 3A, 3B). In addition, the results of transmission electron microscopy analysis showed that purified fenobody-55 can self-assemble into a 24-subunit nanocage (Fig. 3C). Using fenobody-55 as the primary Ab, the ELISA results showed that it still specifically reacted with g3-p239 and cross-reacted with g1-p239, g3-rabbit p239, and g4-p239, which was roughly consistent with that of g3-p239-Nb55 (Fig. 3D).
FIGURE 3.
Expression, purification, and characterization of fenobody-55. (A) Analysis of the expression and purification of fenobody-55 by SDA-PAGE. The expected size of fenobody-55 was ∼34 kDa. (B) Western blot analysis of the expression and purification of fenobody-55 using an anti-His mAb as the primary Ab. (C) Observation of purified fenobody-55 by transmission electron microscopy. Fenobody-55 self-assembled into a 24-subunit nanocage. (D) Cross-reactivity of fenobody-55 with different genotypes of HEV p239s compared with g3-p239-Nb55. G3-p239-Nb55 (1 μg/ml) and fenbody-55 (1 μg/ml) can bind to HEV p239 proteins from mammals but do not react with ap237 from avians. The experiment was independently repeated three times. The error bars indicate the mean ± SD. ***p < 0.001.
Comparisons of the affinities of g3-p239-Nb55 and fenobody-55 for binding to g3-p239
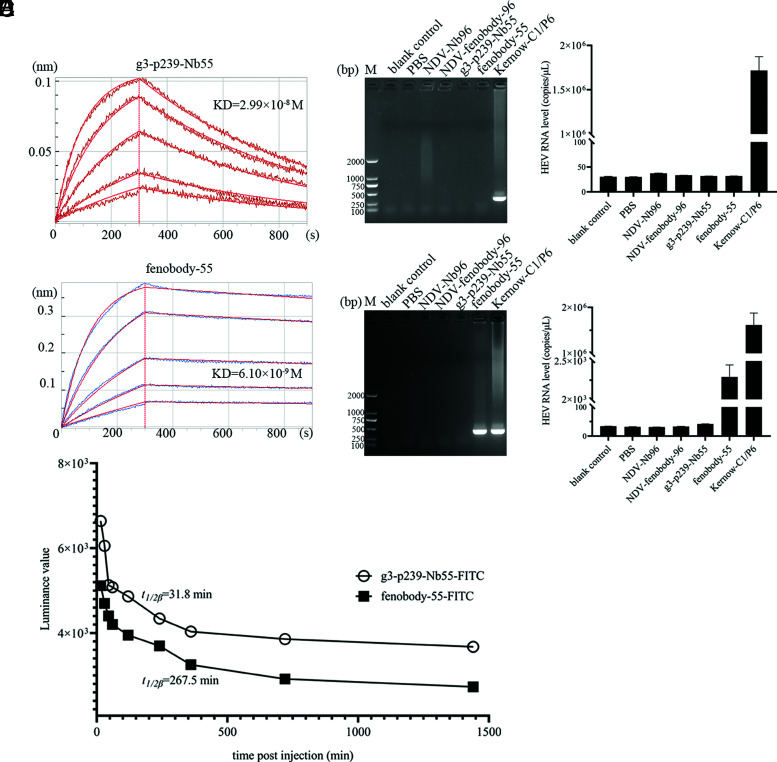
To determine the binding affinities of g3-p239-Nb55 and fenobody-55 for g3-p239, we performed a BLI assay. The results showed that the Kd values of g3-p239-Nb55 and fenobody-55 were 2.99 × 10−8 and 6.10 × 10−9 M, respectively, indicating that both g3-p239-Nb55 and fenobody-55 can react with g3-p239, and that the affinity of fenobody-55 for g3-p239 was ∼20 times greater than that of g3-p239-Nb55 (Fig. 4A, 4B). In addition, the results of immunocapture RT-PCR also showed that the pretreated neHEV could be captured by fenobody-55, but not by g3-p239-Nb55 (Fig. 4C, 4D). The RT-qPCR results also showed that the neHEV particles could be captured by fenobody-55, but not by g3-p239-Nb55 (Fig. 4C, 4D). These results indicated that the epitopes recognized by the nanobody were located on the surface of the viral particles and suggested that the affinity of fenobody-55 for viral particles was greater than that of g3-p239-Nb55.
FIGURE 4.
Characterization of the in vivo affinity, capture ability, and half-life of g3-p239-Nb55 and fenobody-55. Comparisons of the affinities of g3-p239-Nb55 (A) and fenobody-55 (B) for binding to g3-p239 with a BLI assay. The purified g3-p239 (50 μg/ml) was fixed on the solid biosensor AR2G. Then, the nanobody (5 μg/ml) and fenobody (30 μg/ml) were serially diluted for union and disunion. The binding affinity of fenbody-55 for g3-p239 was ∼20 times greater than that for g3-p239-Nb55. The binding capacity of g3-p239-Nb55 and fenobody-55 with no pretreatment (C) and pretreatment with deoxycholate (DOC-Na) and 0.1% trypsin. (D) HEV Kernow-C1/P6 particles by immunocapture PCR. Pretreated HEV particles could be captured by 2.1 μM (3.2 μg) fenobody-55, but not by 2.1 μM (7.25 μg) g3-p239-Nb55. Neither of the unpretreated HEV particles could be captured by either of them. In addition, absolute qRT-PCR was performed to determine the number of HEV RNA copies in the captured particles. The experiment was independently repeated three times. The error bars indicate the mean ± SD. (E) Plasma concentrations of g3-p239-Nb55 and fenobody-55 at different time points after i.v. injection into SPF BALB/C mice. The half-life was represented by t1/2β, the time required for the drug concentration in plasma to decrease by half. The half-life of fenobody-55 was longer than that of g3-p239-Nb55. M, 2000 marker.
Half-life extension of fenobody-55 in vivo
To evaluate the half-lives of g3-p239-Nb55 and fenobody-55, we labeled 50 nM g3-p239-Nb55 and fenobody-55, respectively, with 2 nmol FITC and i.v. injected it via the caudal vein. Serum samples were collected at different time points to analyze the half-lives of g3-p239-Nb55 and fenobody-55 by measuring fluorescence. As shown in Fig. 4E, the values gradually decreased over time, and the t1/2β (the time required for the plasma drug concentration to decrease by half) values of g3-p239-Nb55 and fenobody-55 were 31.8 and 267.5 min, respectively (a simulation performed with Kinetical software). Although equimolar amounts of g3-p239-Nb55 and fenobody-55 were used for labeling, the starting values were discordant, perhaps because of limitations in the FITC labeling efficiency. The half-life of fenobody-55 was approximately nine times greater than that of g3-p239-Nb55, demonstrating that fenobody-55 was metabolized more slowly in vivo.
Comparisons of the neutralizations of g3-p239-Nb55 and fenobody-55
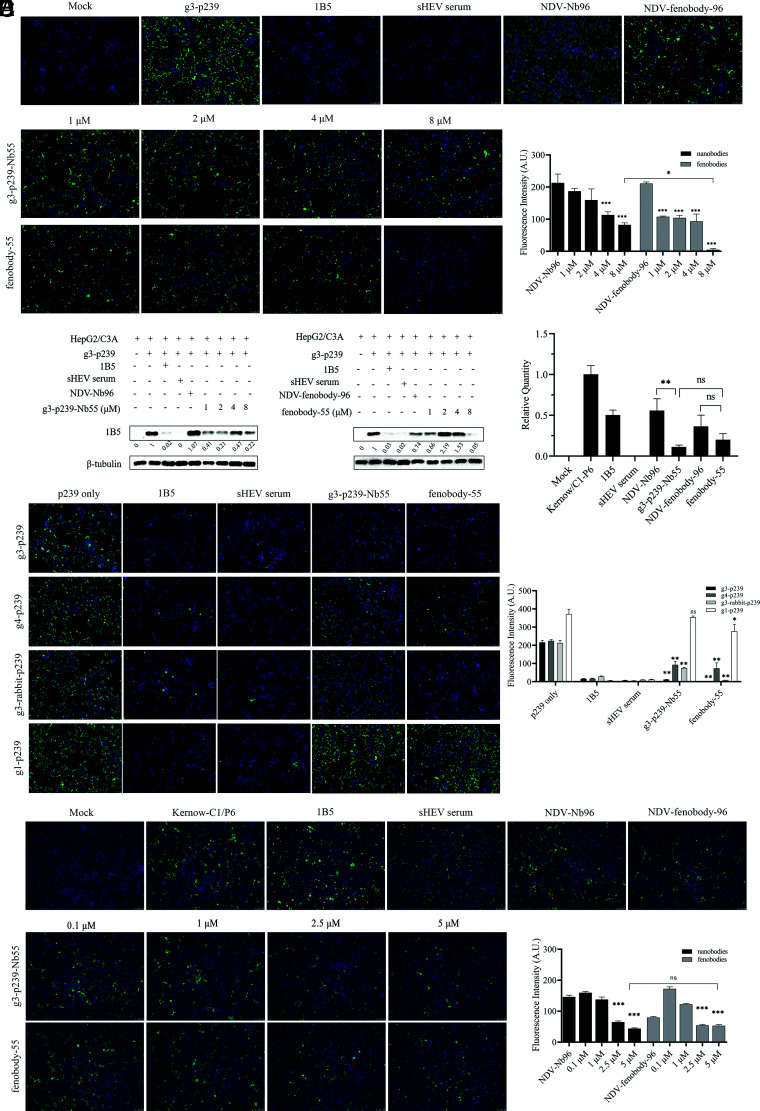
The analysis of cell viability showed that there was no cytotoxicity to HepG2/C3A cells when 8 µM g3-p239-Nb55 or fenobody-55 was added to the cells (Supplemental Fig. 2). Then, a concentration of 8 µM was used for subsequent neutralization assays. In addition, blocking assays of g3-p239 binding to HepG2/C3A by g3-p239-Nb55 and fenobody-55 were performed. As shown in Supplemental Fig. 3, 8 and 16 μg were used for the blocking assay with IFA and Western blot, respectively. After the mixtures of g3-p239-Nb55 or fenobody-55 and g3-p239 were incubated with HepG2/C3A, the IFA and Western blot results showed that both could block g3-p239 from attaching to the cells. At different nanobody concentrations, 8 μM fenobody-55 almost completely blocked the attachment of g3-p239 to HepG2/C3A cells (Fig. 5A, 5B). Compared with g3-p239-Nb55, fenobody-55 had a greater inhibitory effect (79.3 versus 97.8%, significant difference) (Fig. 5A, 5B). Because the same molar concentrations of g3-p239->Nb55 and fenobody-55 were employed in this assay, it was not unexpected that the blocking rate of fenobody-55 (24 U) was greater than that of g3-p239-Nb55 (1 U). In addition, using other HEV ORF2 protein genotypes, including g1-p239, g3-rabbit p239, and g4-p239, for the assay, the results also showed that both g3-p239-Nb55 and fenobody-55 could block attachment to HepG2/C3A cells. In addition, compared with the p239-only group, g3-p239-Nb55 and fenobody-55 had excellent blocking effects on g3-p239, g1-p239, g3-rabbit p239, and g4-p239, and the effect of g3-p239 was especially strong (Fig. 5C).
FIGURE 5.
Neutralizing activities of g3-p239-Nb55 and fenobody-55 against the HEV ORF2 proteins and Kernow-C1/P6 virus in vitro. G3-p239-Nb55 and fenobody-55 can block g3-p239 attachment to HepG2/C3A cells, as determined by IFA. The nuclei were stained by DAPI, and the g3-p239 protein was stained by Alexa Fluor 488_labeled goat anti-mouse Ab. (A) and Western blot (B). (C) G3-p239-Nb55 and fenobody-55 can block different genotypes of HEV ORF2 proteins (g1-p239, g3-rabbit-p239, and g4-p239) from attaching to HepG2/C3A cells. The nuclei were stained by DAPI, and the p239 proteins were stained by Alexa Fluor 488_labeled goat anti-mouse Ab. Compared with those in the p239-only group, the majority of HEV ORF2 proteins were strongly inhibited by g3-p239-Nb55 and fenobody-55, especially g3-p239. Neutralizing analysis of g3-p239-Nb55 and fenobody-55, which inhibited the infection of Kernow-C1/P6 into HepG2/C3A cells, by IFA. The nuclei were stained by DAPI, and the virus was stained by Alexa Fluor 488_labeled goat anti-mouse Ab. (D) and real-time RT-PCR (E). The experiment was independently repeated three times. The error bars indicate the mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001. ns, no significant difference.
The neutralizations of g3-p239-Nb55 and fenobody-55 for the natural particles were also evaluated. After mixtures of neHEV particles with different concentrations of g3-p239-Nb55 or fenobody-55 were inoculated into HepG2/C3A cells, IFA and RT-qPCR revealed that compared with NDV-Nb96 and NDV-fenobdoy-96, g3-p239-Nb55 and fenobody-55 inhibited HEV infection in the cells (79.9 versus 75.7%, respectively) (Fig. 5D, 5E). Unlike those of the g3-p239 protein, there were no significant differences between the antiviral effects of g3-p239-Nb55 and fenobody-55. It is speculated that the recognition or combination of neHEV particles in vitro was limited by the increasing m.w. of fenobody-55. In addition, the results also indicated that the neutralization activity of g3-p239-Nb55 and fenobody-55 against Kernow-C1/P6 infection in the cells was dose dependent (Fig. 5D).
Identification of g3-p239-Nb55 and fenobody-55 neutralizing HEV infection in vivo
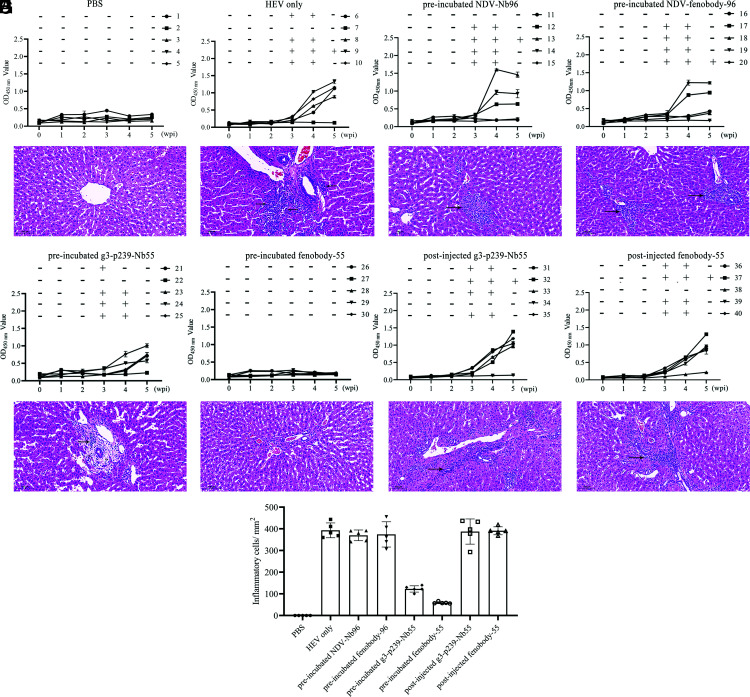
Before inoculation, all rabbits in this study were negative for HEV RNA in their fecal samples and for anti-HEV IgG Abs in their sera (Fig. 6). Fecal virus shedding and an increase in anti-HEV IgG Abs were originally observed from 3 wpi until 5 wpi in only the HEV-infected (except for No. 7), preincubated NDV-Nb96 (except for No. 11), preincubated NDV-fenobody-96 (except for No. 19), and preincubated g3-p239-Nb55 groups (except for No. 12) (Fig. 6B–E). Conversely, all rabbits in the PBS and preincubated fenobody-55 groups were negative for fecal virus shedding or anti-HEV IgG Abs (Fig. 6A, 6F). In addition, to study whether g3-p239-Nb55/fenobody-55 has therapeutic effects on HEV infection, we infected SPF rabbits with neHEV particles and then injected them with g3-p239-Nb55/fenobody-55. The results showed that fecal virus shedding was not reduced in the rabbits from the postinjection g3-p239-Nb55 (except for No. 34) and postinjection fenobody-55 groups (except for No. 38) (Fig. 6G, 6H). In addition, Abs were also induced by HEV infection in these rabbits (Fig. 6G, 6H). These results indicated that fenobody-55 entirely and g3-p239-Nb55 partially neutralized rabbit HEV infection in rabbits preincubated with neHEV particles. However, they cannot inhibit HEV replication when they are injected at 24 h after the rabbits are inoculated with neHEV.
FIGURE 6.
In vivo neutralization analysis of g3-p239-Nb55 and fenobody-55 and optical micrographs of liver lesions from necropsied SPF rabbits. Fecal virus shedding and the level of anti-HEV Abs in the serum of SPF rabbits were detected. The plus (+) and minus (−) symbols indicate positive and negative results, respectively, for HEV RNA in the fecal samples from the different rabbits at 1–5 wpi according to nested RT-PCR. The lines show the levels of Abs against HEV in the sera of different rabbits at 1–5 wpi determined by indirect ELISA with the ORF2 protein as the coating Ag. The liver tissues were stained by hematoxylin-eosin. The arrows show locally intense lymphocytic venous phlebitis, lymphocytic inflammatory cell infiltration, and periphlebitis. SPF rabbits were inoculated with PBS and labeled No. 1–5 (A), with only rabbit HEV and labeled No. 6–10 (B). SPF rabbits were inoculated with rabbit HEV mixed with 133 μM NDV-Nb96 and labeled No. 11–15 (C), mixed with the same dosage of NDV-fenobody-96 and labeled No. 16–20 (D), mixed with g3-p239-Nb55 and labeled No. 21–25 (E), mixed with fenobody-55 and labeled No. 26–30 (F). SPF rabbits were first inoculated with rabbit HEV, and after 24 h, they were injected with 133 μM g3-p239-Nb55 and labeled No. 31–35 (G), and injected with fenobody-55 and labeled No. 36–40 (H). Three liver sections from each rabbit were stained with H&E to evaluate pathological and histopathological changes caused by HEV infection. The error bars indicate the mean ± SD. (I) The average number of inflammatory cells in three random 1-mm2 sections of each slice was determined via ImageJ software.
Hepatic lesions were also observed in the liver tissue slices stained with H&E. Locally intense lymphocytic venous phlebitis, lymphocytic inflammatory cell infiltration, and periphlebitis occurred in the liver tissues of the HEV-challenged, preincubated NDV-Nb96, preincubated NDV-fenobody-96, postinjected g3-p239-Nb55, and postinjected fenobody-55 groups (Fig. 6B–D, 6G, 6H, black arrow). Preincubating g3-p239-Nb55 (Fig. 6E, black arrow) greatly reduced phlebitis, infiltration, and periphlebitis. In contrast, no significant liver lesions were observed in the PBS or preincubated fenobody-55 groups (Fig. 6A, 6F). These results showed that fenobody-55 could neutralize HEV infection in a rabbit animal model, and that g3-p239-Nb55 can provide partial protection. In addition, for the treatment groups, the rabbits in the postinjection g3-p239-Nb55 and postinjection fenobody-55 groups showed the same pathological lesions of the livers as the only virus infection group, indicating that neither had a therapeutic effect on HEV infection (Fig. 6G, 6H). As shown in Fig. 6I, the average number of inflammatory cells in the liver was counted in three randomly selected views of each slice.
Characterization of the epitope recognized by g3-p239-Nb55
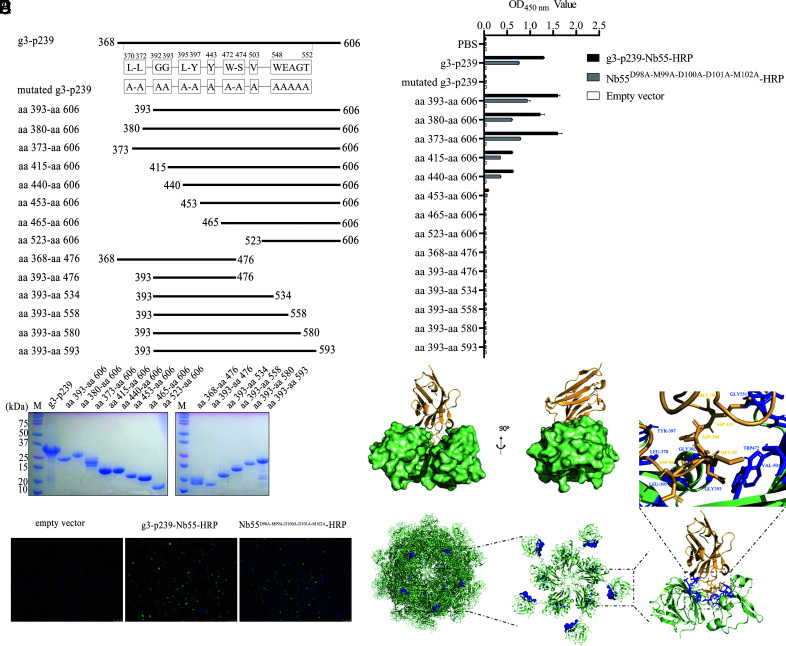
To determine the epitope recognized by g3-p239-Nb55, we designed (Fig. 7A) and produced with the bacterial system 14 different truncated fragments of g3-p239, including aa 393–606, aa 380–606, aa 373–606, aa 415–606, aa 440–606, aa 453–606, aa 465–606, aa 523–606, aa 368–476, aa 393–476, aa 393–534, aa 393–558, aa 393–580, and aa 393–593. SDS-PAGE analysis revealed that these fragments were successfully expressed and purified with the expected sizes (Fig. 7B). Using these fragments as coated Ags, the ELISA results indicated that the truncated fragments aa 393–606, aa 380–606, aa 373–606, aa 415–606, and aa 440–606 could react with g3-p239-Nb55-HRP, but the other truncated fragments could not react (Fig. 7C, 7D). Moreover, the truncated fragments lacking the C terminus of g3-p239 could not react with g3-p239-Nb55-HRP (Fig. 7D). These results suggested that both aa 440–453 (closing to the N terminus) and aa 593–606 (closing to the C terminus) of g3-p239 are indispensable for recognition by g3-p239-Nb55, and that the epitope is conformational. The deletion of amino acids upstream of aa 440 or the C-terminal residues should destroy the structure of the conformational epitope, preventing g3-p239 from being recognized by g3-p239-Nb55.
FIGURE 7.
Identification and characterization of the epitope recognized by g3-p239-Nb55. (A) Schematic diagram of g3-p239 and various truncated proteins. The mutated amino acid residues of g3-p239 are marked in detail. (B) SDS-PAGE analysis of purified g3-p239 and various truncated proteins. (C) Identification of g3-p239-Nb55-HRP and Nb55D98A-M99A-D100A-D101A-M102A-HRP fusion proteins expressed in HEK293T cells by IFA. The nuclei were stained by DAPI, and the fusion proteins were stained by Alexa Fluor 488_labeled goat anti-mouse Ab. (D) Detection of the binding capacity between g3-p239 or truncated and mutated proteins and Nb55-HRP or Nb55 D98A-M99A-D100A-D101A-M102A-HRP by ELISA. Spatial locations on g3-p239 of the epitope recognized by g3-p239-Nb55. Overall structures of g3-p239 monomers (E) and pentamers (F) and a detailed assessment of the epitope. The spatial locations of the g3-p239-Nb55 (yellow) binding epitope (blue) protruding from g3-p239 (green) are marked. The experiment was independently repeated three times. The error bars indicate the mean ± SD.
To determine the key motifs participating in the interaction between g3-p239 and g3-p239-Nb55, we predicted the candidate epitope with PyMOL software (Fig. 7E, 7F). The surface structures of g3-p239 in monomer form and g3-p239-Nb55 are shown on the front and side, respectively (Fig. 7F). Notably, the ribbon diagram revealed the structures of the g3-p239 and g3-p239-Nb55 complexes in pentamer form. The predicted motifs of g3-p239 were discretely located at L370, L372, G392, G393, L395, Y397, Y443, W472, S474, V503, W548, E549, A550, G551, and T552. These key motifs formed spatial structures that could be recognized by 98DMDDM102 of g3-p239-Nb55 (Fig. 7E).
To further verify the predicted epitope, we successfully designed and produced the mutated g3-p239-Nb55D98A-M99A-D100A-D101A-M102A-HRP fusion protein by transient transfection of HEK293T cells (Fig. 7C). The results showed that the binding titers of the g3-p239-Nb55D98A-M99A-D100A-D101A-M102A-HRP fusion protein binding to the aa 393–606, aa 380–606, aa 373–606, aa 415–606, and aa 440–606 fragments were lower than those of g3-p239-Nb55-HRP (significant differences) (Fig. 7D). In addition, when the key residues were entirely mutated to A, the g3-p239-Nb55-HRP or g3-p239-Nb55D98A-M99A-D100A-D101A-M102A-HRP fusion protein could not recognize the mutated g3-p239 protein (Fig. 7A, 7D). To this end, the conformational epitopes of g3-p239 at positions L370, L372, G392, G393, L395, Y397, Y443, W472, S474, V503, W548, E549, A550, G551, and T552 were likely recognized by 98DMDDM102 of g3-p239-Nb55. Based on the genome sequence alignment, these locations were highly conserved among different mammalian HEV genotypes (Table III), suggesting that a (to our knowledge) novel common conserved conformational epitope was present.
Table III. Conservation analysis of the epitope recognized by g3-p239-Nb55 among different genotypes HEV.
| Amino Acid Location of Epitope Recognized by g3-p239-Nb55 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| GenBank Accession No. | L370 | L372 | G392 | G393 | L395 | Y397 | Y443 | W472 | S474 | V503 | W548 | E549 | A550 | G551 | T552 |
| g1 AY230202 | |||||||||||||||
| g1 AF444002 (Sar55)a | |||||||||||||||
| g2 M74506 | |||||||||||||||
| g2 KX578717 | |||||||||||||||
| g3 AY115488 | |||||||||||||||
| g3 JQ679013 (Kernow-C1)a | |||||||||||||||
| g3 KX227751 (rabbit HEV)a | V | ||||||||||||||
| g4 KF176351 (swine HEV)a | |||||||||||||||
| g4 AB074915 | |||||||||||||||
| g4 AB108537 | |||||||||||||||
| g5 AB573435 | T | ||||||||||||||
| g5 BCD83331 | T | ||||||||||||||
| g6 AB602441 | T | ||||||||||||||
| g6 AB856243 | T | ||||||||||||||
| g7 KJ496143 | |||||||||||||||
| g7 KJ496144 | |||||||||||||||
| g8 KX387867 | M | ||||||||||||||
| g8 MH410174 | M | Q | |||||||||||||
| Ca JN997392 (CaHEV)a | G | S | Y | R | T | L | G | P | —b | —b | |||||
| Rat JX120573 | Q | F | T | Q | Q | ||||||||||
The amino acid residues were omitted when they are the same with that of the epitope.
Sequences used in this study.
Deletion mutation.
Discussion
HEV is classified into the subfamily Orthohepevirus, Hepeviridae (4). HEV infection is becoming a public health issue, with a high fatality rate of up to 30% among pregnant individuals (31). Moreover, HEV infection may develop into a chronic infection in immunocompromised individuals who have received organ transplantation (32). Therefore, developing neutralizing Abs to research HEV biology and designing viral vaccines and drugs are essential.
Nanobodies are characterized by small size, strict monomeric behavior, good solubility, thermostability, high affinity, and specificity for the Ag (33). Neutralizing nanobodies have been applied for antiviral infection, vaccine design, and viral biological research, the most striking example being SARS-CoV-2 (34). In this study, monovalent g3-p239-Nb55 and polyvalent fenobody-55 against the HEV ORF2 protein were first produced and were able to block the g3-p239 and truncated HEV ORF2 proteins from HEV genotype 3 (Kernow-C1 strain) from attaching to HepG2/C3A cells. In addition, they can also neutralize HEV infection by preincubation with neHEV particles both in vitro and in vivo. To our knowledge, epitope mapping revealed that the nanobody recognized a novel and common neutralizing epitope among p239 proteins from different genotypes of HEV, which is the first study to determine the novel epitope of HEV using a nanobody. However, when rabbits were injected with g3-p239-Nb55 and fenobody-55 as treatment agents after HEV infected them for 24 h, they could not inhibit HEV replication or treat the lesions caused by HEV infection. However, additional experiments are needed to clarify the causes of this phenomenon in the future.
Nanobodies are readily produced as highly soluble VHH domains, but their affinity is insufficient for many diagnostic or immunotherapy applications. Due to the strict monomeric behavior of nanobodies, it is possible to design multivalent nanobody formats. Multimerization is an efficient strategy for improving the affinity of nanobodies, and this method is named pentabody (35). Furthermore, ferritin was selected as the basic framework to display the nanobodies in 6 × 4 bundles (15). For the subunit structure, the ε domain of ferritin was replaced with a nanobody and then ligated with a GS linker. As a result, the affinity of the nanobody for the H5N1 virus dramatically increased (22). In this study, the affinity of fenobody-55 increased ∼20-fold, further confirming that fenobody can increase this affinity. In addition, the half-life of fenobody-55 was also extended, which was also in accordance with previous research (15, 22). Notably, because of the increased affinity and half-life of fenobody-55 in vivo, fenobody-55 exhibited a greater neutralizing effect on HEV infection in vivo than g3-p239-Nb55. Our results further suggested that nanobodies can be fused with ferritin to design multivalent nanobodies to improve their affinity, half-life, and application.
Interestingly, g3-p239-Nb55 did not completely capture the neHEV particles at a concentration of 2.1 μM, but it could still inhibit viral infection. We think there may be three reasons for this. On the one hand, the sensitivity of the capture ELISA was limited to detecting viruses at extremely low concentrations. On the other hand, when g3-p239-Nb55 was coated on plates and in the solid state, the locations recognizing the viral epitope may be covered. However, g3-p239-Nb55 in the liquid state more easily recognized the HEV particles and neutralized the HEV infection. In addition, as shown in Fig. 3D, both g3-p239-Nb55 and fenobody-55 can specifically bind to and cross-react with ORF2 proteins from different genotypes of HEV, and the OD450 values of ELISA also showed that fenobody-55 showed greater affinity than g3-p239-Nb55 when they were used at the same molar concentrations. The results showing the comparisons of affinity were the same as the results shown in Fig. 4A. Therefore, the other reason why g3-p239-Nb55 did not completely capture the neHEV particles may be that the affinity of g3-p239-Nb55 was lower than that of fenobody-55.
In this study, our results showed that g3-p239 could be blocked by 79.3 and 97.8% by g3-p239-Nb55 and fenobody-55, respectively, indicating that fenobody increased the blocking rate. When different dosages of g3-p239-Nb55 and fenobody-55 were used to block g3-p239 attachment to the cells, the results showed that there was no apparent titration with fenobody-55, as shown in Fig. 5A, with the exception of a decrease at 8 μM. There are two possible explanations. One was that the distances among the concentration gradients were insufficient, resulting in no obvious dose-dependent blocking effect of g3-p239-Nb55 or fenobody-55. Second, g3-p239 may also adsorb onto HepG2/C3A cells via other pathways. Therefore, when the initial dose is low, an attractive dose-dependent blocking effect cannot be demonstrated. However, when g3-p239-Nb55 or fenobody-55 is used at a high dosage, g3-p239 may be conformationally blocked. However, the inhibition of HEV infection by g3-p239-Nb55 and fenobody-55 in vitro was not significantly different (79.9 versus 75.7%). It is speculated that the increasing m.w. of fenobody-55 becomes an obstacle to recognizing or combining with HEV particles in vitro.
In addition, for the in vitro neutralization assay, the results showed that HEV infection could not be completely neutralized by g3-p239-Nb55 or fenobody-55. The possible reason was the dosage used in the assay. According to the results of the cytotoxicity test (Supplemental Fig. 2), 10 μM g3-p239-Nb55 or fenobody-55 showed significant cytotoxicity to HepG2/C3A, and only 8 μM was used, which was probably an insufficient dose for complete neutralization or even the formation of other conformational epitopes recognized by monovalent g3-p239-Nb55 and polyvalent fenobody-55. Moreover, unrelated NDV-Nb96 or NDV-fenobody-96 could slightly block the binding of g3-p239 to HepG2/C3A cells. The small m.w. of the proteins could explain the nonspecific binding between g3-p239 and NDV-Nb96 or NDV-fenobody-96.
In this study, we identified the common and conserved epitope within p239 proteins from different genotypes of HEV, indicating that the nanobody can neutralize the infection of different genotypes of HEV. However, except for the HEV Kernow-C1/P6 strain, no additional strains were chosen for verifying the neutralizing activity of g3-p239-Nb55 or fenobody-55 in this research. The self-assembly of p239 from various genotypes of HEV (genotyped 1, 3, and 4) into virus like particles was applied to mimic attachment to HepG2/C3A cells, followed by a blocking assay. A reasonable explanation is that only the Kernow-C1/P6 strain could replicate in the cell culture system. When it is possible to develop a cell culture system for other strains of HEV, the neutralizing activity of g3-p239-Nb55 or fenobody-55 will be immediately proven. For in vivo identification, rabbits were challenged with rabbit HEV to establish the animal model in this study because it was easier to inoculate native animals (36, 37) successfully. In addition, rabbits with small somatotypes are easier to manipulate than other animals with accessible somatotypes. Although nonhuman primates could not be used in this study, we look forward to evaluating the neutralizing activity of g3-p239-Nb55 or fenobody-55 in animals that are phylogenetically close to humans in the future.
The ORF2 protein comprises three functional domains, namely, the S (aa 129–319), M (aa 320–455), and P (aa 456–606) domains (38). To date, all the identified neutralizing epitopes have been proved to be mapped to the P domain (39). A previous study on the neutralized epitope revealed that the aa 368–393 region forms dimers and higher-order assemblies (40). Moreover, the discrete residues constituted the conformational epitope in the C terminus, including E479, S497, R512, K534, H577, and R578 for mAb 8C11 and E549, K554, G591, T553, G589, and P592 for mAb 8G12 (41, 42). Moreover, mAbs 2C7, 2G9, and 1B5 were shown to recognize the linear epitopes 458PSRPF462, 407EPTV410, and 410VKLYTS415, respectively (19). In this study, the discrete residues in the M and P domains played important roles in the formation of a conformational neutralizing epitope recognized by g3-p239-Nb55, in accordance with previous reports (39). In addition, these locations were highly common conserved regions among different genotypes of HEV, including g1-p239, g3-rabbit p239, and g4-p239. However, the amino acid sequences of genotype 1, 3, 4, and 3-rabbit HEVs were subtly different. As a result, there may be slight differences in the conformation and structure, and then they may show different affinities to the nanobody. Furthermore, the release of enveloped HEV (eHEV) particles with a lipid membrane decreased the ability of Kernow-C1/P6 to be captured by g3-p239-Nb55 or fenobody-55, as mentioned earlier.
Two forms of HEV, neHEV and eHEV virions, can coexist in hosts (43). Usually, neHEV virions are shed in the feces because of the presence of digestive enzymes in the gastrointestinal tract and bile, becoming a new starting point of the fecal-oral route. Nevertheless, eHEV particles with lipid membranes are formed in the cytoplasm and released by the cellular exosomal pathway (44). Research has indicated that the attachment of neHEV to cells is more efficient than that of eHEV (45). In addition, capsid protein was also detected in the blood and urine of solid organ transplant recipients (46). In this study, HEV virions from the cell culture supernatant (mostly eHEV) were pretreated before nanobody/fenobody immunocapture RT-PCR to harvest neHEV virions. As a result, not eHEV but neHEV virions could be captured by fenobody-55. The eHEV particles covering the ORF2 protein with lipid membranes circulating in the blood cannot be recognized by g3-p239-Nb55/fenobody-55. Based on HEV biology, we speculated that g3-p239-Nb55/fenobody-55 could not neutralize eHEV particles in the blood.
In addition, even though highly neutralizing g3-p239-Nb55 and fenbody-55 for neHEV were obtained in our study, they cannot inhibit HEV replication or treat lesions caused by HEV in rabbits 24 h after viral infection. We also speculated that eHEV was present in the blood of HEV-infected rabbits, and that g3-p239-Nb55 and fenbody-55 cannot neutralize eHEV. In the future, many experiments will be needed to confirm our speculations.
In summary, g3-p239-Nb55 and fenobody-55 exhibited neutralizing activities both in vitro and in vivo. Notably, the conformational epitope recognized by g3-p239-Nb55 was located on the surface of viral particles and highly conserved among genotypes 1–8 HEV. To our knowledge, this is the first study in which a nanobody-derived product (fenobody-55) completely blocked capsid proteins from different genotypes of HEV from attaching to host cells and neutralizing HEV infection in rabbits. Our study also provides a foundation for HEV diagnosis, treatment, and vaccine research.
Supplementary Material
Footnotes
This work was supported by the National Key R&D Program of China (2023YFD1800304 to Q.Z.), National Natural Science Foundation of China (31972676 to Q.Z.), and Natural Science Foundation of Shaanxi Province of China (2022JC-12 to Q.Z.).
The online version of this article contains supplemental material.
- BLI
- biolayer interferometry
- cELISA
- competitive ELISA
- eHEV
- enveloped HEV
- HEV
- hepatitis E virus
- NDV
- Newcastle disease virus
- neHEV
- nonenveloped HEV
- ORF
- open reading frame
- PBS′T
- PBS containing 0.5% Tween 20
- PI
- percent inhibition
- RT-qPCR
- quantitative real-time PCR
- SPF
- specific pathogen-free
- wpi
- weeks postinoculation
Disclosures
The authors have no financial conflicts of interest.
References
- 1. Pillot, J., Türkoglu S., Dubreuil P., Cosson A., Lemaigre G., Meng J., Lazizi Y.. 1995. Cross-reactive immunity against different strains of the hepatitis E virus transferable by simian and human sera. C. R. Acad. Sci III 318: 1059–1064. [PubMed] [Google Scholar]
- 2. Cuevas-Ferrando, E., Randazzo W., Pérez-Cataluña A., Sánchez G.. 2019. HEV occurrence in waste and drinking water treatment plants. Front. Microbiol. 10: 2937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Smith., D. B., Simmonds P.; Members of the International Committee on the Taxonomy of Viruses Study Group ; Jameel S., Emerson S. U., Harrison T. J., Meng X.-J., Okamoto H., Van der Poel W. H. M., and Purdy M. A.. 2014. Consensus proposals for classification of the family Hepeviridae. J. Gen. Virol. 95: 2223–2232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Purdy, M. A., Drexler J. F., Meng X.-J., Norder H., Okamoto H., Van der Poel W. H. M., Reuter G., de Souza W. M., Ulrich R. G., Smith D. B.. 2022. ICTV virus taxonomy profile: Hepeviridae 2022. J. Gen. Virol 103: jgv.0.001778. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Forni, D., Cagliani R., Clerici M., Sironi M.. 2018. Origin and dispersal of hepatitis E virus. Emerg. Microbes Infect. 7: 11–23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Zhou, Y. H., Purcell R. H., Emerson S. U.. 2005. A truncated ORF2 protein contains the most immunogenic site on ORF2: antibody responses to non-vaccine sequences following challenge of vaccinated and non-vaccinated macaques with hepatitis E virus. Vaccine 23: 3157–3165. [DOI] [PubMed] [Google Scholar]
- 7. Pavio, N., Meng X. J., Renou C.. 2010. Zoonotic hepatitis E: animal reservoirs and emerging risks. Vet. Res. 41: 46–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Montpellier, C., Wychowski C., Sayed I. M., Meunier J. C., Saliou J. M., Ankavay M., Bull A., Pillez A., Abravanel F., Helle F., et al. 2018. Hepatitis E virus lifecycle and identification of 3 forms of the ORF2 capsid protein. Gastroenterology 154: 211–223.e8. [DOI] [PubMed] [Google Scholar]
- 9. Mori, Y., Matsuura Y.. 2011. Structure of hepatitis E viral particle. Virus Res. 161: 59–64. [DOI] [PubMed] [Google Scholar]
- 10. Zhu, F. C., Zhang J., Zhang X. F., Zhou C., Wang Z. Z., Huang S. J., Wang H., Yang C. L., Jiang H. M., Cai J. P., et al. 2010. Efficacy and safety of a recombinant hepatitis E vaccine in healthy adults: a large-scale, randomised, double-blind placebo-controlled, phase 3 trial. Lancet 376: 895–902. [DOI] [PubMed] [Google Scholar]
- 11. Liu, P., Du R. J., Wang L., Han J., Liu L., Zhang Y. L., Xia J. K., Lu F. M., Zhuang H.. 2014. Management of hepatitis E virus (HEV) zoonotic transmission: protection of rabbits against HEV challenge following immunization with HEV 239 vaccine. PLoS One 9: e87600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Hamers-Casterman, C., Atarhouch T., Muyldermans S., Robinson G., Hamers C., Songa E. B., Bendahman N., Hamers R.. 1993. Naturally occurring antibodies devoid of light chains. Nature 363: 446–448. [DOI] [PubMed] [Google Scholar]
- 13. Sun, S., Ding Z., Yang X., Zhao X., Zhao M., Gao L., Chen Q., Xie S., Liu A., Yin S., et al. 2021. Nanobody: a small antibody with big implications for tumor therapeutic strategy. Int. J. Nanomedicine 16: 2337–2356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Schoof, M., Faust B., Saunders R. A., Sangwan S., Rezelj V., Hoppe N., Boone M., Billesbølle C. B., Puchades C., Azumaya C. M., et al. ; QCRG Structural Biology Consortium . 2020. An ultrapotent synthetic nanobody neutralizes SARS-CoV-2 by stabilizing inactive Spike. Science 370: 1473–1479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Fan, K., Jiang B., Guan Z., He J., Yang D., Xie N., Nie G., Xie C., Yan X.. 2018. Fenobody: a ferritin-displayed nanobody with high apparent affinity and half-life extension. Anal. Chem. 90: 5671–5677. [DOI] [PubMed] [Google Scholar]
- 16. Shukla, P., Nguyen H. T., Torian U., Engle R. E., Faulk K., Dalton H. R., Bendall R. P., Keane F. E., Purcell R. H., Emerson S. U.. 2011. Cross-species infections of cultured cells by hepatitis E virus and discovery of an infectious virus-host recombinant. Proc. Natl. Acad. Sci. USA 108: 2438–2443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Vincke, C., Gutierrez C., Wernery U., Devoogdt N., Hassanzadeh-Ghassabeh G., Muyldermans S.. 2012. Generation of single domain antibody fragments derived from camelids and generation of manifold constructs. Methods Mol. Biol. 907: 145–176. [DOI] [PubMed] [Google Scholar]
- 18. Sheng, Y., Wang K., Lu Q., Ji P., Liu B., Zhu J., Liu Q., Sun Y., Zhang J., Zhou E. M., Zhao Q.. 2019. Nanobody-horseradish peroxidase fusion protein as an ultrasensitive probe to detect antibodies against Newcastle disease virus in the immunoassay. J. Nanobiotechnology 17: 35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Chen, Y., Liu B., Sun Y., Li H., Du T., Nan Y., Hiscox J. A., Zhou E. M., Zhao Q.. 2018. Characterization of three novel linear neutralizing B-cell epitopes in the capsid protein of swine hepatitis E virus. J. Virol. 92: e00251-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Liu, H., Wang Y., Duan H., Zhang A., Liang C., Gao J., Zhang C., Huang B., Li Q., Li N., et al. 2015. An intracellularly expressed Nsp9-specific nanobody in MARC-145 cells inhibits porcine reproductive and respiratory syndrome virus replication. Vet. Microbiol. 181: 252–260. [DOI] [PubMed] [Google Scholar]
- 21. Liu, Z. H., Lei K. X., Han G. W., Xu H. L., He F.. 2020. Novel lentivirus-based method for rapid selection of inhibitory nanobody against PRRSV. Viruses 12: 229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Ji, P., Zhu J., Li X., Fan W., Liu Q., Wang K., Zhao J., Sun Y., Liu B., Zhou E. M., Zhao Q.. 2020. Fenobody and RANbody-based sandwich enzyme-linked immunosorbent assay to detect Newcastle disease virus. J. Nanobiotechnology 18: 44–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Nagashima., S., Takahashi M., Jirintai S., Tanggis, Kobayashi T., Nishizawa T., Okamoto H.. 2014. The membrane on the surface of hepatitis E virus particles is derived from the intracellular membrane and contains trans-Golgi network protein 2. Arch. Virol. 159: 979–991. [DOI] [PubMed] [Google Scholar]
- 24. Huang, F. F., Haqshenas G., Guenette D. K., Halbur P. G., Schommer S. K., Pierson F. W., Toth T. E., Meng X. J.. 2002. Detection by reverse transcription-PCR and genetic characterization of field isolates of swine hepatitis E virus from pigs in different geographic regions of the United States. J. Clin. Microbiol. 40: 1326–1332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Jothikumar, N., Cromeans T. L., Robertson B. H., Meng X. J., Hill V. R.. 2006. A broadly reactive one-step real-time RT–PCR assay for rapid and sensitive detection of hepatitis E virus. J. Virol. Methods 131: 65–71. [DOI] [PubMed] [Google Scholar]
- 26. Zhang, B., Fan J., Luo Y., Lv H., Zhao Q., Fan M., Duan H., Liu B., Nan Y., Sun Y., Zhou E. M.. 2021. Development of a competitive ELISA for detecting antibodies against genotype 1 hepatitis E virus. Appl. Microbiol. Biotechnol. 105: 8505–8516. [DOI] [PubMed] [Google Scholar]
- 27. Wang, X., Zhao Q., Dang L., Sun Y., Gao J., Liu B., Syed S. F., Tao H., Zhang G., Luo J., Zhou E. M.. 2015. Characterization of two novel linear B-cell epitopes in the capsid protein of avian hepatitis E virus (HEV) that are common to avian, swine, and human HEVs. J. Virol. 89: 5491–5501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Fan, M., Luo Y., Zhang B., Wang J., Chen T., Liu B., Sun Y., Nan Y., Hiscox J. A., Zhao Q., Zhou E. M.. 2021. Cell division control protein 42 interacts with hepatitis E virus capsid protein and participates in hepatitis E virus infection. Front. Microbiol. 12: 775083–775099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Liu, B., Chen Y., Sun Y., Nan Y., Li H., Du T., Hiscox J. A., Zhao Q., Zhou E. M.. 2019. Experimental infection of rabbit with swine-derived hepatitis E virus genotype 4. Vet. Microbiol. 229: 168–175. [DOI] [PubMed] [Google Scholar]
- 30. Geng, Y., Zhao C., Song A., Wang J., Zhang X., Harrison T. J., Zhou Y., Wang W., Wang Y.. 2011. The serological prevalence and genetic diversity of hepatitis E virus in farmed rabbits in China. Infect. Genet. Evol. 11: 476–482. [DOI] [PubMed] [Google Scholar]
- 31. Jameel, S. 1999. Molecular biology and pathogenesis of hepatitis E virus. Expert. Rev. Mol. Med. 1999: 1–16. [DOI] [PubMed] [Google Scholar]
- 32. Hering, T., Passos A. M., Perez R. M., Bilar J., Fragano D., Granato C., Medina-Pestana J. O., Ferraz M. L.. 2014. Past and current hepatitis E virus infection in renal transplant patients. J. Med. Virol. 86: 948–953. [DOI] [PubMed] [Google Scholar]
- 33. Hassanzadeh-Ghassabeh, G., Devoogdt N., De Pauw P., Vincke C., Muyldermans S.. 2013. Nanobodies and their potential applications. Nanomedicine (Lond.) 8: 1013–1026. [DOI] [PubMed] [Google Scholar]
- 34. Koenig, P. A., Das H., Liu H., Kümmerer B. M., Gohr F. N., Jenster L. M., Schiffelers L. D. J., Tesfamariam Y. M., Uchima M., Wuerth J. D., et al. 2021. Structure-guided multivalent nanobodies block SARS-CoV-2 infection and suppress mutational escape. Science 371: eabe6230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Zhang, J., Tanha J., Hirama T., Khieu N. H., To R., Tong-Sevinc H., Stone E., Brisson J. R., MacKenzie C. R.. 2004. Pentamerization of single-domain antibodies from phage libraries: a novel strategy for the rapid generation of high-avidity antibody reagents. J. Mol. Biol. 335: 49–56. [DOI] [PubMed] [Google Scholar]
- 36. Ma, H., Zheng L., Liu Y., Zhao C., Harrison T. J., Ma Y., Sun S., Zhang J., Wang Y.. 2010. Experimental infection of rabbits with rabbit and genotypes 1 and 4 hepatitis E viruses. PLoS One 5: e9160–e9168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Wang, L., Liu L., Wang L.. 2018. An overview: rabbit hepatitis E virus (HEV) and rabbit providing an animal model for HEV study. Rev. Med. Virol. 28: e1961–e1968. [DOI] [PubMed] [Google Scholar]
- 38. Yamashita, T., Mori Y., Miyazaki N., Cheng R. H., Yoshimura M., Unno H., Shima R., Moriishi K., Tsukihara T., Li T. C., et al. 2009. Biological and immunological characteristics of hepatitis E virus-like particles based on the crystal structure. Proc. Natl. Acad. Sci. USA 106: 12986–12991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Zhang, J., Zhao Q., Xia N.. 2016. Prophylactic hepatitis E vaccine. Adv. Exp. Med. Biol. 948: 223–246. [DOI] [PubMed] [Google Scholar]
- 40. Wei, M., Zhang X., Yu H., Tang Z. M., Wang K., Li Z., Zheng Z., Li S., Zhang J., Xia N., Zhao Q.. 2014. Bacteria expressed hepatitis E virus capsid proteins maintain virion-like epitopes. Vaccine 32: 2859–2865. [DOI] [PubMed] [Google Scholar]
- 41. Gu, Y., Tang X., Zhang X., Song C., Zheng M., Wang K., Zhang J., Ng M. H., Hew C. L., Li S., et al. 2015. Structural basis for the neutralization of hepatitis E virus by a cross-genotype antibody. Cell. Res. 25: 604–620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Tang, X., Yang C., Gu Y., Song C., Zhang X., Wang Y., Zhang J., Hew C. L., Li S., Xia N., Sivaraman J.. 2011. Structural basis for the neutralization and genotype specificity of hepatitis E virus. Proc. Natl. Acad. Sci. USA 108: 10266–10271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Nagashima, S., Takahashi M., Kobayashi T., Tanggis, Nishizawa T., Nishiyama T., Primadharsini P. P., Okamoto H.. 2017. Characterization of the quasi-enveloped hepatitis E virus particles released by the cellular exosomal pathway. J. Virol. 91: e00822-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Chapuy-Regaud, S., Dubois M., Plisson-Chastang C., Bonnefois T., Lhomme S., Bertrand-Michel J., You B., Simoneau S., Gleizes P. E., Flan B., et al. 2017. Characterization of the lipid envelope of exosome encapsulated HEV particles protected from the immune response. Biochimie 141: 70–79. [DOI] [PubMed] [Google Scholar]
- 45. Yin, X., Ambardekar C., Lu Y., Feng Z.. 2016. Distinct entry mechanisms for nonenveloped and quasi-enveloped hepatitis E viruses. J. Virol. 90: 4232–4242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Marion, O., Capelli N., Lhomme S., Dubois M., Pucelle M., Abravanel F., Kamar N., Izopet J.. 2019. Hepatitis E virus genotype 3 and capsid protein in the blood and urine of immunocompromised patients. J. Infect. 78: 232–240. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.