Abstract
This study investigates the cellular origin and tissue heterogeneity in bipolar disorder (BD) by integrating multiomics data. Four distinct datasets were employed, including single-cell RNA sequencing (scRNA-seq) data (embryonic and fetal brain, n = 8, 1,266 cells), BD Assay for Transposase-Accessible Chromatin using sequencing (ATAC-seq) data (adult brain, n = 210), BD bulk RNA-seq data (adult brain, n = 314), and BD genome-wide association study (GWAS) summary data (n = 413,466). The integration of scRNA-seq data with multiomics data relevant to BD was accomplished using the single-cell disease relevance score (scDRS) algorithm. We have identified a novel brain cell cluster named ADCY1, which exhibits distinct genetic characteristics. From a high-resolution genetic perspective, glial cells emerge as the primary cytopathology associated with BD. Specifically, astrocytes were significantly related to BD at the RNA-seq level, while microglia showed a strong association with BD across multiple panels, including the transcriptome-wide association study (TWAS), ATAC-seq, and RNA-seq. Additionally, oligodendrocyte precursor cells displayed a significant association with BD in both ATAC-seq and RNA-seq panel. Notably, our investigation of brain regions affected by BD revealed significant associations between BD and all three types of glial cells in the dorsolateral prefrontal cortex (DLPFC). Through comprehensive analyses, we identified several BD-associated genes, including CRMP1, SYT4, UCHL1, and ZBTB18. In conclusion, our findings suggest that glial cells, particularly in specific brain regions such as the DLPFC, may play a significant role in the pathogenesis of BD. The integration of multiomics data has provided valuable insights into the etiology of BD, shedding light on potential mechanisms underlying this complex psychiatric disorder.
Subject terms: Comparative genomics, Diagnostic markers, Bipolar disorder
Introduction
Bipolar disorder (BD) is a common psychiatric illness with a high disability rate [1]. The primary clinical features of BD are recurrent episodes of mania and hypomania, in which depressive syndromes is more prevalent [2]. These episodes are characterized by a distinct change in mood and behavior during discrete time periods [3]. BD affects 1 to 2 percent of the population, where patients have a life expectancy of 10 to 20 years shorter than that of the general population [2]. BD has widespread prevalence and serious impact, and its pathogenesis remains unclear.
BD is a highly heritable and polygenic genetic disorder [4], with estimated heritability ranging from 60 to 85 percent [5]. Recently, many multiomics data have been generated to annotate BD-associated SNPs and genomic regions, construct putative regulatory networks, and assess underlying causation or mediation links [6]. Evidence from genome-wide association studies (GWAS) suggested that at least three genes, namely TRANK1, LMAN2L and PTGFR, were significant genetic risk factors for BD development [7]. Recently, 33 novel BD-associated genomic loci were identified in the GWAS of 41,917 BD cases [8]. Numerous BD-dysregulated genes have been associated with diagnosis, symptom profile and prognosis [9–11]. Through multi-omics research, we can begin to explain the impact of BD-related variants or risk genes on brain anatomy, physiology and behavioural phenotypes in order to comprehensively understand etiology [6].
Compared to bulk omics data, which depend on average measurements of heterogeneous cell populations, single-cell RNA sequencing (scRNA-Seq) data offers a more precise, high-resolution transcriptome expression profile and captures cell type-specific features that may be missed by bulk tissue data [6]. The complex cellular nature of the brain makes it necessary to apply single-cell omics methods to comprehend genomic regulation at the single-cell level. For example, the Spatio-Temporal cell Atlas of the Human Brain defined 42 cell subtypes spanning 20 brain regions and 11 developmental periods by analyzing 13 available human brain scRNA-seq datasets [12]. A single-nucleus transcriptomics analysis identified major dysregulation of excitatory neurons and oligodendrocyte progenitor cells in major depressive disorder [13]. However, integrating single-cell data with multiomics data to understand the molecular function of brain cells and reveal its relationship to neuropsychiatric disorders is a new and challenging task.
scRNA-seq technology has evolved to fuse various multiomics information, which now includes not only the original quantitative data of each omics but also the widely available derived data like reference maps and quantitative trait loci [6]. A growing body of research has applied single-cell multiomics integrated analysis to extend insights into the biological mechanisms of neuropsychiatric disorders [14, 15]. One study integrating single-cell transcriptome data from mouse brain regions with 42 human GWAS results suggested that medium spiny neurons and pyramidal cells were associated with BD [16]. The authors employed predefined cell type annotations to aggregate cells of the same cell type, but this approach ignored the heterogeneity of individual cells within cell types. We introduce the single-cell disease relevance score (scDRS), which was developed to integrate the gene expression profile of scRNA-seq with the polygenic signatures of GWASs to correlate individual cells with diseases and complex traits [17]. The scDRS algorithm employs an appropriately matched empirical null distribution to assess whether a given cell exhibits increased expression across a set of putative disease genes derived from GWASs, thereby estimating well-calibrated P-values [17].
In this study, we aim to expand the application of scDRS to integrate scRNA-seq data with four BD-related omics datasets, encompassing diverse data types such as Transposase-Accessible Chromatin using sequencing (ATAC-seq), bulk RNA-seq, GWAS, and transcriptome-wide association studies (TWAS). Our comprehensive cell-type-level and gene-level analyses are designed to identify BD-associated cell types across multiple molecular layers, including open chromatin, RNA, and DNA-derived data, and to pinpoint specific brain regions associated with these BD-related cell types. This endeavor seeks to contribute to a deeper understanding of BD pathology at both cellular and regional levels, potentially unveiling new diagnostic markers and therapeutic targets to enhance current diagnosis and treatment strategies for BD.
Materials and methods
Processing and clustering of human brain single-cell RNA-seq data
The FASTQ files of scRNA-seq data were derived from PsychENCODE Consortium (Synapse ID: syn7067037) [18]. Samples of 1266 single cells were obtained from eight post-mortem embryonic and fetal brain tissue specimens, ranging from five to twenty weeks post-gestation. The sampled area corresponded to the dorsolateral prefrontal cortex (DLPFC), dorsal pallium, and cortical plate. Dissected tissue was dissociated into cell suspension. Captured cells were processed through lysis, reverse transcription, and PCR amplification to generate single-cell cDNA libraries, followed by Illumina sequencing [18].
CASAVA (Illumina) and FastQC were used for quality control measures and reporting. The FASTQ files were trimmed using Trim Galore (version 0.6.7) and aligned to the human transcriptome (GRCh37/hg19) by using HISAT2 (version 2.1.0) [19]. We used featureCounts (version 2.0.1) [20] to calculate the gene reads count values. We used the Scanpy python module (version 1.9.1) [21] based on Python (version 2.7.18) for clustering. We filtered out genes detected in fewer than three cells and excluded cells with more than eight percent of transcripts attributed to mitochondrial genes. A total of 978 cells with 7026 highly variable genes were left. We conducted principal component analysis (PCA) for cellular clustering and utilized Uniform Manifold Approximation and Projection (UMAP) [22] for visualization. Both PCA and UMAP functions are included in Scanpy. Wilcoxon rank-sum test was used to identify differentially expressed genes (DEGs) between clusters (false discovery rate (FDR) < 0.05). We retrieved markers for clusters to mark brain cell types with CellMarker 2.0 (http://bio-bigdata.hrbmu.edu.cn/CellMarker/) [23]. A cluster with significantly higher expression levels of a specific cell type marker gene was labelled as that cell type.
Processing and analysis of ATAC-seq data
The dataset for ATAC-seq consisted of 210 PEAKS files, including 25 samples from individuals with BD and 185 control samples, with BD not differentiated by symptom severity. The mean age of the samples was 74.40 (SE = 1.23) years, and out of the total samples, 83 (39.52%) were female. These files were obtained from the PsychENCODE consortium (Synapse ID: syn7349497) [24]. Samples from patients with BD and controls were obtained from bulk tissue collected from frozen left hemisphere coronal slabs excised from the DLPFC at autopsy. The libraries exhibiting periodic patterns were multiplexed and subjected to 50 bp sequencing on the HiSeq4000 (Illumina) platform, ensuring a minimum depth of 40 M paired-end reads per library. FASTQs were trimmed for adapter sequence and low-quality reads by cutadapt (version 1.2.0) [25]. We matched the clean reads to the GRCh37/hg19 reference genome by bowtie2 (version 2.1.0) [26]. All reads sequenced single- or paired-end in ATAC-seq libraries were treated as single-read sequences in the alignment. Aligned BAM files were sorted by Samtools (version 0.1.18) [27]. And, narrow open chromatin peaks were labelled with MACS2.
Diffbind (version 2.14.0) [28] in R (version 3.6.1) was implemented to identify differentially accessible regions (peaks) (FDR < 0.05). Reads were quantified for each region of overlapping open chromatin (ROCs) in each sample. Peaks were annotated using the HOMER (version 4.11) [29] and mapped to the GRCh37/hg19 human genome combination in the differential ROCs between BD and control samples. Various covariates were included for the differential chromatin analysis, such as GC (%), date submitted, ChrM aligned (%), nuclear respiratory factor (NRF), RNA-seq intronic aligned (%), RNA-seq astrocyte (%), RNA-seq oligodendrocyte (%), PC1 genotype, seizures, age of death, post-mortem interval (PMI), sex, and diagnosis [24]. Annotated genes with FDR < 0.05 were selected for subsequent analysis.
Processing and analysis of bulk RNA-seq data
The dataset for RNA-seq included 314 BAM files, comprising 69 samples from individuals with BD and 245 control samples, with BD not differentiated by symptom severity. The mean age of the samples was 66.72 (SE = 1.18) years, and out of the total samples, 121 (38.54%) were female. These BAM files were obtained from the PsychENCODE consortium (Synapse ID: syn3270015) [30]. Bipolar patients and controls were sampled from well-characterized post-mortem human brains at autopsy. All RNA was processed into rRNA-depleted stranded libraries. Libraries were sequenced on the HiSeq2000 on a high output flowcell for 100 bp paired-end sequencing and were 3-plexed per lane to reach 40 M paired-end reads per library. FASTQs were trimmed for adapter sequence and low base call quality (Phred score < 30 at ends) by cutadapt (v1.12). Trimmed reads were then matched to the GRCH37.p13/hg19 reference genome via STAR (2.4.2a).
For each BAM file generated by STAR, reads for each gene were summarized using featureCounts (version 2.0.1) [20] refer to the GRCh37/hg19 human genome. DESeq2 (version 1.6.3) [31] was used to assess DEGs between patients with BD and controls. The analysis considered various socio-demographic and clinical covariates. These covariates, including diagnosis, age, study/batch, sex, PMI, RNA integrity number (RIN), library preparation, sequencing platform, strand specificity, brain bank, brain region, ethnicity, and 29 sequence principal components (seqPCs), were incorporated into the analysis model. The model employed multivariate adaptive regression and a two-phase approach to maximize variance explanation by the covariates and eliminate redundant terms. Squared terms were included for continuous variables. Covariates present in at least half of the resulting pruned models, such as diagnosis, age, age2, study/batch, sex, PMI, RIN, RIN2, brain bank, brain region, seqPCs [1–3, 5–8, 10–14, 16, 18–25, 27–29], and seqPC32, were considered as known covariates [30]. The DEGs with FDR < 0.05 were selected for subsequent analysis.
GWGAS and TWAS analysis using GWAS summary data
Genome-wide association summary data for BD was obtained from a publicly available large-scale GWAS [8]. Briefly, the dataset comprised 57 collected cohorts, with a combined total of 41,917 individuals diagnosed with BD and 371,549 control subjects of European descent. Cases were required to meet internationally recognized consensus criteria (DSM-IV, ICD-9, or ICD-10) for a lifetime diagnosis of BD. In most cohorts, controls were randomly selected from the population to screen the absence of lifelong psychosis. To control test statistic inflation at SNPs with low minor allele frequency (MAF) in small cohorts, SNPs were retained only if cohort MAF was >1% and minor allele count was >10 in either cases or controls. Details of the sample collection, quality control, and correlation analysis of the dataset are described elsewhere [8].
The Genome-Wide Gene Association Analysis (GWGAS) for Bipolar Disorder (BD) was conducted using the MAGMA software (version 1.10) [32]. This analysis integrated Single Nucleotide Polymorphism (SNP)-based P-values derived from the Genome-Wide Association Study (GWAS) as input data. A total of 19,427 protein-coding genes, as defined by the NCBI 37.3 gene annotations, were included in the GWGAS. The gene-association tests accounted for the linkage disequilibrium (LD) between SNPs, ensuring accurate assessment of gene-level associations. Genes exhibiting a P-value below the threshold of 0.05 were selected for subsequent analysis. To establish a control panel, a GWGAS based on height GWAS was employed [33], enabling a reference for evaluating the specificity of the BD-associated genes.
Transcriptome-wide association study (TWAS) for BD was performed by FUSION (http://gusevlab.org/projects/fusion/) [34]. The reference weights of DLPFC were pre-calculated using the multiple prediction models in FUSION. For each gene, all SNPs within 1 Mb of the gene in the GWAS were selected. Imputation Z score of the cis genetic effect on BD was calculated as ZTWAS = W’Z/(W’SW)1/2. Z represents the scores of BD, while W represents the weights. S represents the SNP-correlation covariance matrix. We implemented 5000 permutations to control the potential effects of multiple testing. The genes with a permutated P-value < 0.05 were selected for subsequent analysis.
Multiomics analysis at single-cell level for bipolar disorder
Single-cell disease relevance score (scDRS) is a method for linking scRNA-seq to polygenic disease risk at single-cell resolution, independent of annotated cell type [17]. The scDRS method follows a three-step process. Firstly, it constructs a set of potential disease genes using the statistics derived from four BD-related omics panels. In this step, four putative gene sets G ⊂ {1, 2, …, ngene} with gene P values {P g}g ∈G were constructed, consisting of the top 1,000 BD-associated genes, such as differentially annotated peaks for ATAC-seq, DEGs for RNA-seq, significant genes for TWAS, and GWGAS respectively (Supplementary Table S1). Secondly, scDRS calculates a raw disease score and B Monte Carlo (MC) samples of raw control scores for each cell. Sample B sets of control genes matching the mean expression and expression variance of the BD-associated genes. define the estimate of gene-specific technical noise level. Thirdly, after performing gene set-wise and cell-wise normalization, scDRS determines an association P value for each cell c by comparing its normalized disease score to the empirical distribution of the pooled normalized control scores across all control gene sets and cells.
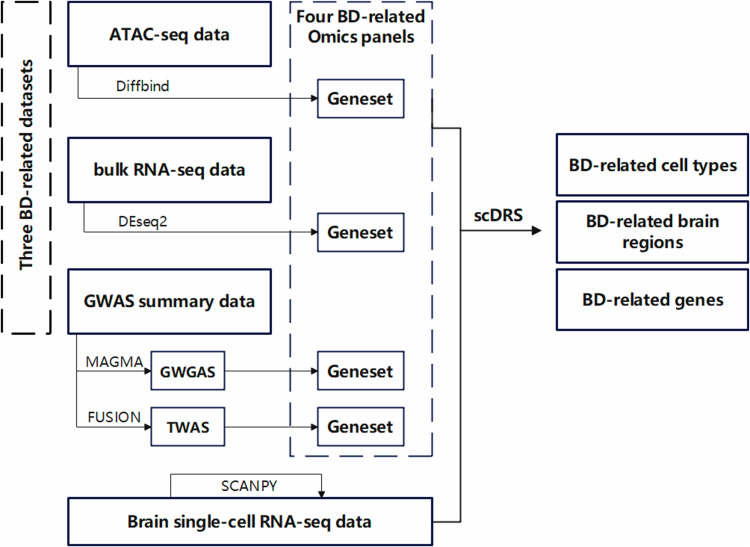
We performed two scDRS downstream analyses based on individual-cell-level disease-association P-values, raw omics disease scores, and raw control scores [17]. Initially, we conducted cell-type-level analyses to establish associations between predefined cell types and BD within four omics panels, including ATAC-seq, bulk RNA-seq, GWGAS, and TWAS. Within each predefined cell type, we assessed the heterogeneity in the association with BD across individual cells. We replicated these cell type-level analyses in various brain regions to identify the brain regions associated with specific cell types. Subsequently, we performed gene-level analyses in each omics panel to prioritize BD-related genes. The schematic diagram depicting the analyzed pipeline is presented in Fig. 1.
Fig. 1. Schematic overview of the analysis pipeline.
scRNA-seq single-cell RNA sequencing, ATAC-seq assay for transposase-accessible chromatin using sequencing. GWAS genome-wide association study, TWAS transcriptome-wide association study, GWGAS genome-wide gene association study, scDRS single-cell disease relevance score.
Gene ontology and kyoto encyclopedia of genes and genomes analyses of BD-associated genes
We utilized the Metascape tool (http://metascape.org/) to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) analyses on the genes associated with BD. Each omics panel, including ATAC-seq, RNA-seq, TWAS, and GWGAS, generated a distinct gene list. To perform enrichment analysis, we selected the top 300 ranked genes from each panel. In addition, we performed a combined enrichment analysis by integrating the multiple gene lists to identify shared biological processes associated with BD across different panels. P-values were computed using the cumulative hypergeometric distribution, and q-values were calculated using the Benjamini-Hochberg procedure to correct for multiple testing. Processes with q-value less than 0.05 were considered statistically significant.
Results
Discovery of a novel cell type in the human brain via scRNA-seq analysis
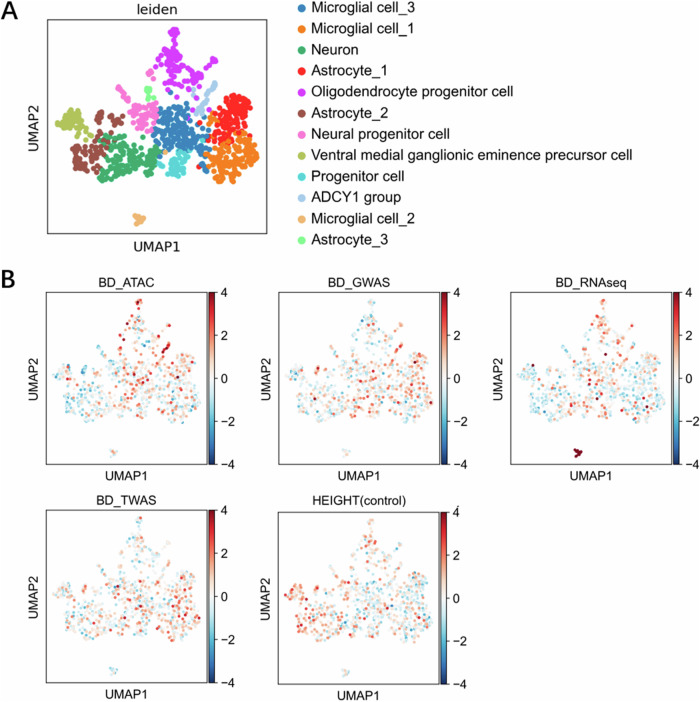
We employed dimensionality reduction techniques and categorized 978 cells into 12 distinct clusters using PCA, as visualized in the UMAP plot (Fig. 2A). Cell types were identified for each cluster based on characteristic markers and annotations from CellMarker 2.0 (Fig. S1) (Fig. 2A). The cells were primarily classified into eight types, including microglia (313 cells, 32.0%), astrocytes (224 cells, 22.9%), and neurons (130 cells, 13.3%). Interestingly, we identified a novel cluster, the ADCY1 group (30 cells, 3.1%) (Fig. 2A). To assess the reproducibility of this cell cluster, we examined the spatio-temporal cell atlas of the human brain (STAB) [12]. The STAB search results revealed consistently elevated levels of ADCY1 expression across the currently classified cell populations (Fig. S2). This observation raises the possibility of potentially misclassified ADCY1 cell populations within the existing classifications. However, there is currently no experimental evidence supporting ADCY1 as a brain cell marker according to Cell Marker 2.0 (Supplementary Table S2).
Fig. 2. scRNA-seq reveals cell clusters in the human brain.
The Uniform Manifold Approximation and Projection (UMAP) plot in Fig. 2A presents the results of an analysis of 978 cells. Principal Component Analysis (PCA) was employed for dimensionality reduction, and the cells were subsequently clustered into twelve distinct groups. Figure 2B shows BD multiomics panels in combination with predefined twelve brain cell subgroups. The color represents the disease score, where a darker color indicates a higher score, signifying greater disease enrichment for the specific cell cluster. Height was included as a negative control trait. ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study.
Single-cell multiomics analysis reveals specific subgroups of bipolar disorder
We conducted a multiomics analysis of BD in conjunction with the twelve predefined brain cell subgroups (Fig. 2B). Our results revealed eight significant correlations (P < 0.05) across four BD-related omics panels (Table 1). Five cell subgroups were associated with BD, including astrocyte_3 (PATAC = 0.040), microglial cell_1 (PTWAS = 0.015), microglial cell_2 (PRNA = 0.001), microglial cell_3 (PATAC = 0.024), oligodendrocyte progenitor cell (PATAC = 0.001, PRNA = 0.010), and the ADCY1 group (PATAC = 0.001, PGWAS = 0.041). Oligodendrocyte progenitor cells reflected cellular heterogeneity (PATAC = 0.039). Notably, none of the identified associations in the negative control group reached statistical significance.
Table 1.
Associations of multiomics panels with twelve brain cell subpopulations.
| GROUP | N_cell | ATAC | GWAS | RNA | TWAS | Height (control) | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| asso_p | heter_p | asso_p | heter_p | asso_p | heter_p | asso_p | heter_p | asso_p | heter_p | ||
| ADCY1 group | 30 | 0.001* | 0.136 | 0.041* | 0.435 | 0.176 | 0.094 | 0.268 | 0.058 | 0.717 | 0.779 |
| Astrocyte_1 | 123 | 0.231 | 0.006* | 0.185 | 0.545 | 0.998 | 0.362 | 0.184 | 0.984 | 0.976 | 0.979 |
| Astrocyte_2 | 91 | 0.990 | 0.007* | 0.615 | 0.742 | 1.000 | 0.128 | 0.879 | 0.375 | 0.016* | 0.166 |
| Astrocyte_3 | 10 | 0.040* | 0.767 | 0.511 | 0.034* | 0.134 | 0.776 | 0.286 | 0.886 | 0.532 | 0.636 |
| Microglial cell_1 | 138 | 0.463 | 0.234 | 0.412 | 0.067 | 0.644 | 0.158 | 0.015* | 0.077 | 0.909 | 0.553 |
| Microglial cell_2 | 15 | 0.415 | 0.880 | 0.786 | 0.948 | 0.001* | 0.296 | 0.876 | 0.839 | 0.363 | 0.103 |
| Microglial cell_3 | 160 | 0.024* | 0.298 | 0.329 | 0.430 | 0.541 | 0.204 | 0.208 | 0.753 | 0.900 | 0.787 |
| Neural progenitor cell | 71 | 0.418 | 0.013* | 0.963 | 0.832 | 0.826 | 0.26 | 0.689 | 0.974 | 0.416 | 0.566 |
| Neuron | 130 | 0.726 | 0.002* | 0.736 | 0.211 | 0.627 | 0.097 | 0.930 | 0.026* | 0.254 | 0.934 |
| Oligodendrocyte progenitor cell | 110 | 0.001* | 0.039* | 0.754 | 0.341 | 0.010* | 0.394 | 0.981 | 0.781 | 0.394 | 0.917 |
| Progenitor cell | 46 | 0.486 | 0.311 | 0.094 | 0.399 | 0.233 | 0.524 | 0.371 | 0.151 | 0.701 | 0.404 |
| Ventral medial ganglionic eminence precursor cell | 54 | 0.151 | 0.066 | 0.961 | 0.602 | 0.619 | 0.005* | 0.813 | 0.005* | 0.002* | 0.124 |
Height was included as a negative control trait.
ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study, N_cell the number of cells contained in the subgroup. * indicates P < 0.05.
Identification of specific cell subpopulations in brain tissues for bipolar disorder
Based on the aforementioned results, we focused on three types of glial cells: astrocytes, microglial cells, and oligodendrocyte progenitor cells. We investigated the performance of these glial cells in three brain tissues: the cortical plate, dorsal palm, and DLPFC. The Astrocyte_1 score showed a significant correlation with the cortical plate (PTWAS = 0.038), while the Astrocyte_3 score correlated with DLPFC (PATAC = 0.040). The Microglial cell_2 score was significantly correlated with the cortical plate (PRNA = 0.001), dorsal pallium (PRNA = 0.001), and DLPFC (PRNA = 0.001). The Microglial cell_3 score was significantly correlated with DLPFC (PATAC = 0.001). The Oligodendrocyte progenitor cell score was correlated with DLPFC (PATAC = 0.001, PRNA = 0.014). We additionally explored the relevance of the new cluster ADCY1 group with brain-related tissues. The results showed that the ADCY1 group score was associated with DLPFC (PATAC = 0.001, PGWAS = 0.041) (Table 2). Importantly, the absence of statistical significance associations in the negative control group indicates the specificity of these associations to BD.
Table 2.
Associations of specific cell subpopulations with brain tissues for bipolar disorder.
| Cell Type | Panel | Tissue | N_cell | asso_p | asso_z | heter_p | heter_z |
|---|---|---|---|---|---|---|---|
| Astrocyte_1 | ATAC | Cortical plate | 120 | 0.215 | 0.773 | 0.007* | 2.030 |
| ATAC | Dorsolateral prefrontal cortex | 3 | 0.529 | −0.153 | 0.326 | 0.003 | |
| GWAS | Cortical plate | 120 | 0.179 | 0.969 | 0.507 | 0.080 | |
| GWAS | Dorsolateral prefrontal cortex | 3 | 0.339 | 0.377 | 0.658 | −0.833 | |
| RNA | Cortical plate | 120 | 0.998 | −2.708 | 0.362 | 0.434 | |
| RNA | Dorsolateral prefrontal cortex | 3 | 0.462 | −0.031 | 0.379 | 0.484 | |
| TWAS | Cortical plate | 120 | 0.038* | 1.809 | 0.967 | −1.862 | |
| TWAS | Dorsolateral prefrontal cortex | 3 | 0.231 | 0.725 | 0.001* | 1.424 | |
| Height | Cortical plate | 120 | 0.975 | −1.795 | 0.991 | −2.374 | |
| Height | Dorsolateral prefrontal cortex | 3 | 0.195 | 0.878 | 0.001 | 1.371 | |
| Astrocyte_2 | ATAC | Dorsal pallium | 91 | 0.990 | −2.266 | 0.007* | 2.393 |
| GWAS | Dorsal pallium | 91 | 0.615 | −0.314 | 0.742 | −0.620 | |
| RNA | Dorsal pallium | 91 | 1.000 | −3.613 | 0.128 | 1.152 | |
| TWAS | Dorsal pallium | 91 | 0.623 | −0.347 | 0.916 | −1.384 | |
| Height | Dorsal pallium | 91 | 0.016* | 2.342 | 0.166 | 0.942 | |
| Astrocyte_3 | ATAC | Dorsolateral prefrontal cortex | 10 | 0.040* | 1.801 | 0.767 | −0.768 |
| GWAS | Dorsolateral prefrontal cortex | 10 | 0.511 | −0.085 | 0.034* | 1.912 | |
| RNA | Dorsolateral prefrontal cortex | 10 | 0.134 | 1.087 | 0.776 | −0.744 | |
| TWAS | Dorsolateral prefrontal cortex | 10 | 0.679 | −0.476 | 0.442 | 0.141 | |
| Height | Dorsolateral prefrontal cortex | 10 | 0.532 | −0.122 | 0.636 | −0.354 | |
| Microglial cell_1 | ATAC | Cortical plate | 138 | 0.463 | 0.085 | 0.234 | 0.736 |
| GWAS | Cortical plate | 138 | 0.412 | 0.177 | 0.067 | 1.430 | |
| RNA | Cortical plate | 138 | 0.644 | −0.394 | 0.158 | 0.935 | |
| TWAS | Cortical plate | 138 | 0.071 | 1.465 | 0.237 | 0.701 | |
| Height | Cortical plate | 138 | 0.909 | −1.344 | 0.553 | −0.094 | |
| Microglial cell_2 | ATAC | Cortical plate | 5 | 0.355 | 0.327 | 0.257 | 0.515 |
| ATAC | Dorsal pallium | 1 | 0.433 | 0.114 | / | / | |
| ATAC | Dorsolateral prefrontal cortex | 9 | 0.478 | 0.011 | 0.770 | −0.784 | |
| GWAS | Cortical plate | 5 | 0.843 | −1.064 | 0.857 | −1.061 | |
| GWAS | Dorsal pallium | 1 | 0.412 | 0.150 | / | / | |
| GWAS | Dorsolateral prefrontal cortex | 9 | 0.665 | −0.459 | 0.893 | −1.256 | |
| RNA | Cortical plate | 5 | 0.001* | 16.062 | 0.156 | 1.041 | |
| RNA | Dorsal pallium | 1 | 0.001* | 14.102 | / | / | |
| RNA | Dorsolateral prefrontal cortex | 9 | 0.001* | 16.235 | 0.410 | 0.135 | |
| TWAS | Cortical plate | 5 | 0.954 | −1.566 | 0.960 | −1.669 | |
| TWAS | Dorsal pallium | 1 | 0.580 | −0.189 | / | / | |
| TWAS | Dorsolateral prefrontal cortex | 9 | 0.952 | −1.633 | 0.871 | −1.061 | |
| Height | Cortical plate | 5 | 0.674 | −0.473 | 0.393 | 0.036 | |
| Height | Dorsal pallium | 1 | 0.701 | −0.546 | / | / | |
| Height | Dorsolateral prefrontal cortex | 9 | 0.110 | 1.229 | 0.100 | 1.623 | |
| Microglial cell_3 | ATAC | Cortical plate | 60 | 0.916 | −1.336 | 0.074 | 1.400 |
| ATAC | Dorsal pallium | 1 | 0.668 | −0.496 | / | / | |
| ATAC | Dorsolateral prefrontal cortex | 99 | 0.023* | 2.136 | 0.318 | 0.512 | |
| GWAS | Cortical plate | 60 | 0.276 | 0.581 | 0.184 | 0.902 | |
| GWAS | Dorsal pallium | 1 | 0.298 | 0.493 | / | / | |
| GWAS | Dorsolateral prefrontal cortex | 99 | 0.507 | −0.043 | 0.465 | 0.201 | |
| RNA | Cortical plate | 60 | 0.955 | −1.625 | 0.652 | −0.386 | |
| RNA | Dorsal pallium | 1 | 0.936 | −1.287 | / | / | |
| RNA | Dorsolateral prefrontal cortex | 99 | 0.350 | 0.341 | 0.255 | 0.703 | |
| TWAS | Cortical plate | 60 | 0.534 | −0.139 | 0.335 | 0.504 | |
| TWAS | Dorsal pallium | 1 | 0.325 | 0.450 | / | / | |
| TWAS | Dorsolateral prefrontal cortex | 99 | 0.793 | −0.793 | 0.804 | −0.862 | |
| Height | Cortical plate | 60 | 0.525 | −0.098 | 0.948 | −1.813 | |
| Height | Dorsal pallium | 1 | 0.739 | −0.646 | / | / | |
| Height | Dorsolateral prefrontal cortex | 99 | 0.999 | −2.942 | 0.508 | 0.015 | |
| Oligodendrocyte | ATAC | Cortical plate | 3 | 0.131 | 1.150 | 0.181 | 1.271 |
| progenitor cell | ATAC | Dorsolateral prefrontal cortex | 107 | 0.001* | 3.592 | 0.037* | 1.865 |
| GWAS | Cortical plate | 3 | 0.684 | −0.499 | 0.683 | −1.434 | |
| GWAS | Dorsolateral prefrontal cortex | 107 | 0.739 | −0.636 | 0.308 | 0.454 | |
| RNA | Cortical plate | 3 | 0.907 | −1.311 | 0.331 | −0.267 | |
| RNA | Dorsolateral prefrontal cortex | 107 | 0.014* | 2.428 | 0.429 | 0.183 | |
| TWAS | Cortical plate | 3 | 0.473 | 0.048 | 0.507 | −0.036 | |
| TWAS | Dorsolateral prefrontal cortex | 107 | 0.980 | −2.025 | 0.451 | 0.050 | |
| Height | Cortical plate | 3 | 0.205 | 0.820 | 0.843 | −1.350 | |
| Height | Dorsolateral prefrontal cortex | 107 | 0.379 | 0.310 | 0.910 | −1.369 | |
| ADCY1 group | ATAC | Dorsolateral prefrontal cortex | 30 | 0.001* | 4.472 | 0.136 | 1.174 |
| GWAS | Dorsolateral prefrontal cortex | 30 | 0.041* | 1.832 | 0.435 | 0.1 | |
| RNA | Dorsolateral prefrontal cortex | 30 | 0.176 | 0.961 | 0.094 | 1.313 | |
| TWAS | Dorsolateral prefrontal cortex | 30 | 0.491 | 0.053 | 0.507 | −0.05 | |
| HEIGHT | Dorsolateral prefrontal cortex | 30 | 0.717 | −0.589 | 0.779 | −0.743 |
Height was included as a negative control trait.
ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study. N_cell the number of cells contained in the subgroup. * indicates P < 0.05.
Pseudotime trajectory inference for glial cells
To elucidate the developmental trajectories of glial cell states, we utilized the Scanpy algorithm to organize three types of glial cells, namely astrocyte, microglial cell, and oligodendrocyte progenitor cell, into a pseudotime trajectory (Fig. 3). Pseudotime trajectory analysis involves ordering cells along a hypothetical timeline that represents their progressive stages of development or differentiation [35]. The neural precursor cell was manually designated as the root cell for diffusion pseudotime analysis. Our analysis revealed that the three glial cell types predominantly occupied the mid and late developmental states (Fig. 3A). Furthermore, we investigated the expression changes of marker genes along the trajectories of cell subpopulation using partition-based graph abstraction (PAGA) plots (Fig. 3B). The activation sequence of these marker genes demonstrated consistency with established cellular behaviors, with the markers becoming activated towards the terminal phase of the cell trajectory. Figure 3C delineates the developmental trajectories of glial cells across diverse brain regions. Our findings revealed that in the DLPFC, BD-associated subpopulations, such as Astrocyte_3, Microglial cell_3, and Oligodendrocyte progenitor cells, mainly exhibited early to middle stages of development. Conversely, in the cortical plate, BD-associated subpopulations like Astrocyte_1 and Microglial cell_2 primarily indicated advanced development.
Fig. 3. Pseudotime trajectory inference for glial cells.
The Scanpy algorithm was utilized to construct a pseudotime trajectory for three distinct glial cell types, including astrocytes, microglial cells, and oligodendrocyte progenitor cells. Figure A presents the pseudotime results and the cell subpopulation location annotations. The mid and late developmental stages were found to be the predominant stages across the three glial cell types. Figure B shows the expression patterns of marker genes for glial cell subpopulations along pseudotime. Figure C delineates the developmental trajectories of glial cells across diverse brain regions, including cortical plate and DLPFC. OPC oligodendrocyte progenitor cell, DLPFC dorsolateral prefrontal cortex.
Identification of BD-related genes across multiomics panels
We performed a Pearson correlation analysis of gene expression with scDRS disease scores in four BD-related omics panels. Figure 4 summarizes the top 20 genes with correlation coefficients in each panel and matches the correlation performance of these genes in the other three panels. We discovered that the same gene exhibits varying degrees of association with BD across different biological processes. Certain genes, such as CRMP1, SYT4, UCHL1, and ZBTB18, exhibited relatively strong correlations across panels. The most strongly correlated genes in each panel were GRIA2 (r ATAC-seq = 0.21), CRMP1 (r GWAS = 0.18), FOLR2 (r RNA-seq = 0.76), and VSTM2L (r TWAS = 0.18). Detailed results are listed in Supplementary Table S3–6.
Fig. 4. The top 20 genes with correlation coefficients in four omics-panels.
Pearson correlation analysis was conducted to examine the relationship between gene expression and scDRS disease scores in four different omics panels. The top 20 genes, as ranked by correlation coefficients in each panel, were compared to evaluate their performance across the remaining three panels. Value represents the Pearson correlation coefficient. ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study, scDRS single-cell disease relevance score.
Pathway enrichment analyses reveal BD-associated pathways
We conducted GO and KEGG pathway enrichment analyses to identify the most significantly enriched pathways associated with BD across four panels, as shown in Table 3. The results revealed several highly enriched terms, including salt transmembrane transporter activity (q ATAC-seq = 4.17 × 10−13), presynapse (q GWAS = 9.12 × 10−6), inflammatory response (q RNA-seq = 2.69 × 10−31), and cytoplasmic region (q ATAC-seq = 2.75 × 10−2). Furthermore, a combined enrichment analysis demonstrated commonalities observed across panels, such as cell–cell adhesion, cell leading edge, and locomotion, as depicted in Fig. 5.
Table 3.
Top 3 clusters of the top 300 genes associated with bipolar disorder in four omics panels.
| Panel | GO | Category | Description | Count (%) | P-value | q-value |
|---|---|---|---|---|---|---|
| BD_ATAC | GO:1901702 | GO Molecular Functions | Salt transmembrane transporter activity | 34 (12.06) | 1.86E-17 | 4.17E-13 |
| GO:0097447 | GO Cellular Components | Dendritic tree | 33 (11.7) | 8.13E-16 | 9.12E-12 | |
| GO:0098794 | GO Cellular Components | Postsynapse | 29 (10.28) | 5.50E-12 | 1.07E-08 | |
| BD_GWAS | GO:0098793 | GO Cellular Components | Presynapse | 24 (8.63) | 6.76E-10 | 9.12E-06 |
| GO:0097060 | GO Cellular Components | Synaptic membrane | 20 (7.19) | 8.13E-10 | 9.12E-06 | |
| GO:0030424 | GO Cellular Components | Axon | 24 (8.63) | 6.17E-09 | 4.27E-05 | |
| BD_RNASeq | GO:0006954 | GO Biological Processes | Inflammatory response | 52 (17.93) | 1.48E-35 | 2.69E-31 |
| GO:0001775 | GO Biological Processes | Cell activation | 57 (19.66) | 2.40E-35 | 2.69E-31 | |
| GO:0050865 | GO Biological Processes | Regulation of cell activation | 53 (18.28) | 3.72E-33 | 2.82E-29 | |
| BD_TWAS | GO:0099568 | GO Cellular Components | Cytoplasmic region | 13 (4.71) | 2.95E-06 | 2.75E-02 |
| GO:0099111 | GO Biological Processes | Microtubule-based transport | 11 (3.99) | 3.39E-06 | 2.75E-02 | |
| GO:0030424 | GO Cellular Components | Axon | 19 (6.88) | 6.92E-06 | 3.80E-02 |
GO gene ontology, ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study.
Fig. 5. Enrichment of characteristic genes in different omics panels.
ATAC-seq assay for transposase-accessible chromatin using sequencing, GWAS genome-wide association study, TWAS transcriptome-wide association study.
Discussion
In this study, we prioritized BD-associated cell types and revealed the etiology of BD from a genetic perspective by integrating multiomics characteristic gene sets with scRNA-seq data using the scDRS algorithm. Astrocytes were found to be significantly related to BD at the RNA-seq level. Microglia showed a strong association with BD at the TWAS, ATAC-seq, and RNA-seq levels. Oligodendrocyte precursor cells showed a significant association with BD at the ATAC-seq and RNA-seq levels. Notably, the BD-associated genes, as well as the identified cell types, exhibit distinct characteristics across multiple omics levels.
Our research identified eight principal clusters within 978 human brain cells, leading to a more detailed analysis of 12 sub-clusters of these primary cell types. Specifically, we discovered a novel cluster expressing adenylate cyclase 1 (ADCY1) in the human brain, which was correlated with DLPFC in ATAC-seq and GWAS panels. ADCY1 is an activity-dependent calcium-stimulated adenylate cyclase with mRNA transcripts expressed predominantly in nervous system tissues [36]. The expression of ADCY1 mRNA in the central nervous system (CNS) is regionally unique and developmentally controlled, mainly in neurons and to a lesser extent in oligodendrocytes, microglia, and astrocytes [37]. A GWAS has suggested that ADCY1 polymorphisms are associated with BD, and alterations in the ADCY1 gene may affect the therapeutic effect of the first-line drug lithium for BD [38]. ADCY1 controls brain development and synaptic plasticity by favorably regulating the cAMP signaling cascade response [39]. Studies have shown that ADCY1 expression in the hippocampus decreased with age and increased during the acquisition of spatial learning [40], highlighting a possible role of ADCY1 in learning and memory. Nevertheless, it is crucial to investigate whether the ADCY1 cell population in the brains of BD patients plays a unique role and contributes to disease progression. This necessitates further exploration.
Zhang et al. explored the spatial correlations between hippocampus coordinates and brain traits, revealing a link between BD and the proximal region of the hippocampus [17]. Their study focused on seven brain-related diseases/traits: schizophrenia, major depressive disorder, bipolar disorder, neuroticism, college education, body mass index, and smoking habits [17]. Diverging from this approach, our investigation broadens the scope by examining BD-associated cell types in various brain regions through multiomics approaches. Importantly, our findings precisely identify the brain regions where glial cells associated with BD are prevalent, offering specific insights into the neurobiological underpinnings of the disorder.
Our study revealed a significant association between BD and three glial cell types, including astrocytes, oligodendrocyte precursor cells, and microglia. The pathophysiology of BD is intricately linked with structural and functional abnormalities in these glial cells, as evidenced consistently across various research fields, such as histopathology, biochemistry, and neuroimaging [41]. A decreased density and number of these three glial cell types have been consistently reported in postmortem examinations of BD patients [42]. Interestingly, these glial cell families appear to possess regulatory mechanisms that interplay in the context of BD. For instance, activated microglia release reactive oxygen species, reactive nitrogen species, and chemokines [43]. This action induces an environment of oxidative stress and inflammation, subsequently affecting astrocyte functionality [44]. Additionally, microglia generate essential survival and proliferative factors such as insulin-like growth factor 1 (IGF-1), brain-derived neurotrophic factor (BDNF), transforming growth factor-beta (TGF-β), and nerve growth factor (NGF), which support oligodendrocyte precursor cells [45]. Activated astrocytes release cytokines such as interleukin 1 (IL-1), IL-6, and IL-10, which play multifaceted roles in modulating microglial activation, inducing phenotypic shifts in microglia, and influencing the development and neuroinflammatory response of oligodendrocytes [37]. Further research into the intricate interplay among these cells and their contribution to oxidative stress and inflammation may provide potential intervention strategies for BD. Our findings are corroborated by the team that conceived the scDRS algorithm, which similarly identified astrocytes and oligodendrocyte precursor cells as BD-associated using BD GWAS data [17]. Additionally, they observed a correlation between Bergmann glial cells and BD, which further strengthens our conclusion that BD is associated with specific glial cell types.
Our study demonstrated that astrocytes in the cortical plate were associated with BD in the TWAS panel, while astrocytes in the DLPFC were linked to BD in the ATAC panel. This suggests that gene expression in astrocytes during development may subtly influence susceptibility to BD. These multi-omics results consistently support the association between astrocytes with BD. Studies have confirmed the characteristics of astrocyte abnormalities [46] and neuroinflammation [47] in BD patients. Astrocytes are key mediators of the downstream immune response to inflammatory signals in the CNS [48]. Activated astrocytes generate cytokines that impact their own functionality and that of other brain cells, thereby inducing toxicity and inflammation [49].
The significance of epigenetic regulation in BD should not be underestimated. Chromatin accessibility, primarily influenced by histone acetylation, plays a crucial role in glial cell differentiation [50]. Research has indicated that valproic acid treatment enhances histone acetylation, leading to the upregulation of mood-regulating genes. This process aids in mitigating manic symptoms in BD patients [51]. While traditional histone deacetylase inhibitors like valproic acid offer some therapeutic benefits [52], emerging technologies such as site-directed histone acetylation using dCas9-HAT present promising avenues for more precise epigenetic interventions in psychiatric disorders [53]. Future research should explore these novel approaches to better target BD-related genetic pathways, potentially enhancing symptom management or decelerate the progression of BD.
Our analysis of the RNA-seq panel revealed that microglia were associated with BD in the cortical plate, dorsal pallium, and DLPFC. The ATAC panel, on the other hand, identified a correlation between BD and microglia in the DLPFC. These findings suggest that alterations in the typical developmental pathways of these cells might contribute to BD’s etiology. Numerous studies have reported elevated microglial biomarker levels in BD patients [47]. Research demonstrated that the expression of genes associated with microglia activation and phagocytosis, such as CD68, TREM2, and P2RY12, was downregulated in the BD patients’ DLPFC [54]. This downregulation could potentially impair microglia-related functions, including compensatory responses to myelin damage and regeneration [55]. Activated microglial cells are proposed as potential BD biomarkers [37]. As resident macrophages of the CNS [56], microglia can polarize into M1 and M2 phenotypes. This M1/M2 polarization of microglia likely regulates the equilibrium between the induction and resolution of neuroinflammation and immune response in BD [57]. Our findings further signified an association between BD and oligodendrocyte precursor cells in the DLPFC, as revealed by both ATAC and RNA-seq panels. Oligodendrocyte precursor cells are distributed throughout the CNS and are precursors of mature oligodendrocytes [58]. It’s been suggested that a dysregulation of myelination by these oligodendrocytes might be linked with BD [59]. Additionally, oligodendrocytes are susceptible to oxidative stress [60] and neuroinflammation [61], both known to be correlated with BD. Upon examining the specific brain regions associated with BD, we discovered significant associations between BD and all three types of glial cells in the DLPFC. This observation aligns with existing research suggesting that a reduced glial density in the DLPFC is a typical characteristic of individuals diagnosed with BD [62]. A holistic strategy covering molecular, functional, and cognitive areas is essential to better understand the DLPFC’s role in BD and uncover novel treatments.
Our Pearson association analysis of genes echoed the findings of many studies highlighting the multifaceted roles of brain genes in various biological processes [63]. Our research suggested that the BD-related genes exhibited variations across multiomics panels. Several genes, specifically CRMP1, SYT4, UCHL1, and ZBTB18, showed relatively robust correlations across these panels. Notably, the gene encoding collapsin response mediator protein 1 (CRMP1) showed significant association with BD status and the severity of mood symptoms [64]. In BD subjects, downregulation of CRMP1 expression following treatment best predicts a reduction in manic and depressive symptoms [64]. Synaptotagmin-4 (SYT4), predominantly expressed in the brain and neuroendocrine system, is essential for optimal neuronal function [65]. Perturbations in SYT4 levels have been linked to memory loss in mice and may contribute to psychiatric disorders in humans [66]. ZBTB18, also known as RP58 or ZNF238, is a KRAB C2H2 zinc-finger protein that has been characterized as a transcriptional repressor with crucial roles in brain development and neuronal differentiation [67]. This gene has been identified as a risk gene for neurodevelopmental disorders [68], highlighting its significance in neurodevelopmental processes. Additionally, alterations or reductions in ubiquitin carboxyl-terminal hydrolase L1 (UCHL1) have been associated with various neurodegenerative diseases. Research has demonstrated that altered knockdown of UCHL1 homologs in Drosophila results in abnormal mitochondrial function and oxidative stress in neurons [69]. These findings support the robustness of our methodology and provide valuable insights into the potential roles of ZBTB18 and UCHL1 in neurodevelopmental disorders and neurodegenerative diseases. Further research is warranted to explore the distinct functions of these identified genes within their respective omics panels and to unravel their precise mechanisms of action in the context of brain development and disease pathogenesis.
In summary, our single-cell multiomics integration analysis identified microglia, astrocytes, and oligodendrocyte precursor cells as possible cytopathology of BD at the genome-wide, chromatin accessible gene, and transcription levels. We also pinpointed the brain regions corresponding to BD-associated glial cells. Gene correlation analysis in multiomics panels highlighted BD-associated genes, including CRMP1, SYT4, UCHL1, and ZBTB18. These findings broaden the application of multiomics integrated analysis in the pathogenesis of BD, and provide reference for brain histopathology and cell targeted therapy of BD.
Strengths and limitations
This research highlights a key strength in employing single-cell multiomics integration analysis to thoroughly investigate specific cell types associated with BD and their respective brain regions. By integrating various omics data, this methodological approach provides a robust framework for uncovering the complex genetic mechanisms underlying BD. Furthermore, this progress in genetic and cellular understanding of BD hints at the potential for developing personalized future treatments, which might include genetic profiling, targeted therapies, predictive treatment responses, and integrated strategies to enhance patient care. However, there are certain limitations to consider. Despite the innovative methodology employed, the study lacks experimental validation for the newly identified cell cluster expressing ADCY1 and the implicated roles of the identified genes in BD. While the data-driven approach generates hypotheses, experimental studies are essential to validate these suppositions. The study utilized certain predefined markers to classify cell types, which may pose challenges in identifying new cell types using non-specific markers. Moreover, the multiomics analysis involves a multistep technical process, which introduces potential technical noise. Furthermore, while the datasets underwent stringent quality control, they exhibit variations in data quality, resolution, and coverage, thus requiring careful interpretation of the results. Lastly, the mismatch in developmental stages between single-cell data from fetal tissues and bulk omics data from adults may slightly compromise the precise identification of disease-specific cell types and their functions in the developmental stages of BD. Future research may utilize adult brain samples to more thoroughly explore the cellular and regional pathology characteristic of BD.
Supplementary information
Author contributions
Material preparation, data collection and analysis were performed by Wenming Wei and Bolun Cheng. The first draft of the manuscript was written by Wenming Wei. The figures and tables made by Xuena Yang, Xiaoge Chu, Dan He and Xiaoyue Qin. The literature searches were performed by Na Zhang, Yijing Zhao, Qingqing Cai, Sirong Shi, Yan Wen, Jingni Hui, Huan Liu, and Yumeng Jia. The study design was performed by Feng Zhang.
Funding
This research has received financial support from the Natural Science Basic Research Plan in Shaanxi Province of China [2021JCW-08]. Data were generated as part of the PsychENCODE Consortium, supported by: U01DA048279, U01MH103339, U01MH103340, U01MH103346, U01MH103365, U01MH103392, U01MH116438, U01MH116441, U01MH116442, U01MH116488, U01MH116489, U01MH116492, U01MH122590, U01MH122591, U01MH122592, U01MH122849, U01MH122678, U01MH122681, U01MH116487, U01MH122509, R01MH094714, R01MH105472, R01MH105898, R01MH109677, R01MH109715, R01MH110905, R01MH110920, R01MH110921, R01MH110926, R01MH110927, R01MH110928, R01MH111721, R01MH117291, R01MH117292, R01MH117293, R21MH102791, R21MH103877, R21MH105853, R21MH105881, R21MH109956, R56MH114899, R56MH114901, R56MH114911, R01MH125516, and P50MH106934 awarded to: Alexej Abyzov, Nadav Ahituv, Schahram Akbarian, Alexander Arguello, Lora Bingaman, Kristin Brennand, Andrew Chess, Gregory Cooper, Gregory Crawford, Stella Dracheva, Peggy Farnham, Mark Gerstein, Daniel Geschwind, Fernando Goes, Vahram Haroutunian, Thomas M. Hyde, Andrew Jaffe, Peng Jin, Manolis Kellis, Joel Kleinman, James A. Knowles, Arnold Kriegstein, Chunyu Liu, Keri Martinowich, Eran Mukamel, Richard Myers, Charles Nemeroff, Mette Peters, Dalila Pinto, Katherine Pollard, Kerry Ressler, Panos Roussos, Stephan Sanders, Nenad Sestan, Pamela Sklar, Nick Sokol, Matthew State, Jason Stein, Patrick Sullivan, Flora Vaccarino, Stephen Warren, Daniel Weinberger, Sherman Weissman, Zhiping Weng, Kevin White, A. Jeremy Willsey, Hyejung Won, and Peter Zandi.
Data availability
The BD GWAS data can be freely accessed via the PGC website at https://www.med.unc.edu/pgc/results-and-downloads. Data for Single-cell RNA-Seq (accession no. syn7067037), BD Bulk RNA-seq (accession no. syn3270015), and BD ATAC-Seq (accession no. syn7349497) were procured from the PsychENCODE Consortium, available on the Synapse platform. Interested researchers can request access to the PsychENCODE Consortium data by submitting an application form on the Synapse website at www.synapse.org/#!Synapse:syn4921369/wiki/235539. The datasets generated during and/or analyzed during the current study are available at https://github.com/weihuan12138/scmultiomics.git.
Code availability
All software utilized in this research is openly accessible and can be located through the respective URLs or references provided. The code developed to generate all results presented in this study can be accessed at https://github.com/weihuan12138/scmultiomics.git.
Competing interests
The authors declare no competing interests.
Ethics approval
The individual-level scRNA-seq, ATAC-seq, and bulk RNA-seq data, were sourced from the PsychENCODE Consortium (https://psychencode.synapse.org/) under application account fzhxjtu@synapse.org(SynID:syn5553626). In detail, the scRNA-seq data were obtained from SingleCellRNAseq study of the PsychENCODE Consortium. Neurotypical control tissue samples used in this study were obtained from various sources. Tissue was collected after obtaining parental or next of kin consent and with approval by the institutional review boards at the Yale University School of Medicine, and at each institution from which tissue specimens were obtained. Tissue was handled in accordance with ethical guidelines and regulations for the research use of human brain tissue set forth by the NIH and the WMA Declaration of Helsinki. Both ATAC-seq and bulk RNA-seq data were derived from the BrainGVEX study of the PsychENCODE Consortium. The BrainGVEX study utilizes well-characterized postmortem human brain samples from the Stanley Medical Research Institute (SMRI) and Banner Sun Health Research Institute (BSHRI). Ethical approval for the SMRI samples was granted based on Swedish regulations (Ethical Research Committee Ups dnr 99277) and from the University of New South Wales Human Research Ethics Committee (HREC #17826). For the BSHRI samples, ethical approval was secured for all subjects via Institutional Review Board-approved protocols and informed consent documents (Western IRB, Puyallup, WA; protocols 1132516 and 20201852).
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Wenming Wei, Bolun Cheng.
Supplementary information
The online version contains supplementary material available at 10.1038/s41398-024-03044-1.
References
- 1.Hu X, Yu C, Dong T, Yang Z, Fang Y, Jiang Z. Biomarkers and detection methods of bipolar disorder. Biosens Bioelectron. 2023;220:114842. 10.1016/j.bios.2022.114842 [DOI] [PubMed] [Google Scholar]
- 2.Nikolitch K, Saraf G, Solmi M, Kroenke K, Fiedorowicz JG. Fire and darkness: on the assessment and management of bipolar disorder. Med Clin North Am. 2023;107:31–60. 10.1016/j.mcna.2022.04.002 [DOI] [PubMed] [Google Scholar]
- 3.Nierenberg AA, Agustini B, Kohler-Forsberg O, Cusin C, Katz D, Sylvia LG, et al. Diagnosis and treatment of bipolar disorder: a review. JAMA. 2023;330:1370–80. 10.1001/jama.2023.18588 [DOI] [PubMed] [Google Scholar]
- 4.Craddock N, Sklar P. Genetics of bipolar disorder. Lancet. 2013;381:1654–62. 10.1016/S0140-6736(13)60855-7 [DOI] [PubMed] [Google Scholar]
- 5.Bienvenu OJ, Davydow DS, Kendler KS. Psychiatric ‘diseases’ versus behavioral disorders and degree of genetic influence. Psychol Med. 2011;41:33–40. 10.1017/S003329171000084X [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dong X, Liu C, Dozmorov M. Review of multi-omics data resources and integrative analysis for human brain disorders. Brief Funct Genomics. 2021;20:223–34. 10.1093/bfgp/elab024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chen DT, Jiang X, Akula N, Shugart YY, Wendland JR, Steele CJ, et al. Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol Psychiatry. 2013;18:195–205. 10.1038/mp.2011.157 [DOI] [PubMed] [Google Scholar]
- 8.Mullins N, Forstner AJ, O’Connell KS, Coombes B, Coleman JRI, Qiao Z, et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet. 2021;53:817–29. 10.1038/s41588-021-00857-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Weissleder C, Webster MJ, Barry G, Shannon Weickert C. Reduced insulin-like growth factor family member expression predicts neurogenesis marker expression in the subependymal zone in schizophrenia and bipolar disorder. Schizophr Bull. 2021;47:1168–78. 10.1093/schbul/sbaa159 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cruceanu C, Tan PPC, Rogic S, Lopez JP, Torres-Platas SG, Gigek CO, et al. Transcriptome sequencing of the anterior cingulate in bipolar disorder: dysregulation of G protein-coupled receptors. Am J Psychiat. 2015;172:1131–40. 10.1176/appi.ajp.2015.14101279 [DOI] [PubMed] [Google Scholar]
- 11.Akula N, Barb J, Jiang X, Wendland JR, Choi KH, Sen SK, et al. RNA-sequencing of the brain transcriptome implicates dysregulation of neuroplasticity, circadian rhythms and GTPase binding in bipolar disorder. Mol Psychiatr. 2014;19:1179–85. 10.1038/mp.2013.170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Song L, Pan S, Zhang Z, Jia L, Chen WH, Zhao XM. STAB: a spatio-temporal cell atlas of the human brain. Nucleic Acids Res. 2021;49:D1029–D37. 10.1093/nar/gkaa762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nagy C, Maitra M, Tanti A, Suderman M, Theroux JF, Davoli MA, et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat Neurosci. 2020;23:771–81. 10.1038/s41593-020-0621-y [DOI] [PubMed] [Google Scholar]
- 14.Zhu T, Liu J, Beck S, Pan S, Capper D, Lechner M, et al. A pan-tissue DNA methylation atlas enables in silico decomposition of human tissue methylomes at cell-type resolution. Nat Methods. 2022;19:296–306. 10.1038/s41592-022-01412-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Luquez T, Gaur P, Kosater IM, Lam M, Lee DI, Mares J, et al. Cell type-specific changes identified by single-cell transcriptomics in Alzheimer’s disease. Genome Med. 2022;14:136. 10.1186/s13073-022-01136-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Olislagers M, Rademaker K, Adan RAH, Lin BD, Luykx JJ. Comprehensive analyses of RNA-seq and genome-wide data point to enrichment of neuronal cell type subsets in neuropsychiatric disorders. Mol Psychiatry. 2022;27:947–55. 10.1038/s41380-021-01324-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zhang MJ, Hou K, Dey KK, Sakaue S, Jagadeesh KA, Weinand K, et al. Polygenic enrichment distinguishes disease associations of individual cells in single-cell RNA-seq data. Nat Genet. 2022;54:1572–80. 10.1038/s41588-022-01167-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wang D, Liu S, Warrell J, Won H, Shi X, Navarro FCP, et al. Comprehensive functional genomic resource and integrative model for the human brain. Science. 2018;362:eaat8464. 10.1126/science.aat8464 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37:907–15. 10.1038/s41587-019-0201-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30. 10.1093/bioinformatics/btt656 [DOI] [PubMed] [Google Scholar]
- 21.Wolf FA, Angerer P, Theis FJ. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 2018;19:15. 10.1186/s13059-017-1382-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Becht E, McInnes L, Healy J, Dutertre C-A, Kwok IWH, Ng LG, et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat Biotechnol. 2019;37:38–44. [DOI] [PubMed]
- 23.Zhang X, Lan Y, Xu J, Quan F, Zhao E, Deng C, et al. CellMarker: a manually curated resource of cell markers in human and mouse. Nucleic Acids Res. 2019;47:D721–D8. 10.1093/nar/gky900 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Bryois J, Garrett ME, Song L, Safi A, Giusti-Rodriguez P, Johnson GD, et al. Evaluation of chromatin accessibility in prefrontal cortex of individuals with schizophrenia. Nat Commun. 2018;9:3121. 10.1038/s41467-018-05379-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetjournal. 2011;17:10–2. [Google Scholar]
- 26.Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–9. 10.1038/nmeth.1923 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence alignment/map format and SAMtools. Bioinformatics. 2009;25:2078–9. 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ross-Innes CS, Stark R, Teschendorff AE, Holmes KA, Ali HR, Dunning MJ, et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature. 2012;481:389–93. 10.1038/nature10730 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38:576–89. 10.1016/j.molcel.2010.05.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gandal MJ, Zhang P, Hadjimichael E, Walker RL, Chen C, Liu S, et al. Transcriptome-wide isoform-level dysregulation in ASD, schizophrenia, and bipolar disorder. Science. 2018;362:eaat8127. 10.1126/science.aat8127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550. 10.1186/s13059-014-0550-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.de Leeuw CA, Mooij JM, Heskes T, Posthuma D. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput Biol. 2015;11:e1004219. 10.1371/journal.pcbi.1004219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yengo L, Sidorenko J, Kemper KE, Zheng Z, Wood AR, Weedon MN, et al. Meta-analysis of genome-wide association studies for height and body mass index in approximately 700,000 individuals of European ancestry. Hum Mol Genet. 2018;27:3641–9. 10.1093/hmg/ddy271 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Gusev A, Ko A, Shi H, Bhatia G, Chung W, Penninx BW, et al. Integrative approaches for large-scale transcriptome-wide association studies. Nat Genet. 2016;48:245–52. 10.1038/ng.3506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Bendall SC, Davis KL, Amir el AD, Tadmor MD, Simonds EF, Chen TJ, et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell. 2014;157:714–25. 10.1016/j.cell.2014.04.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chen J, Ding Q, An L, Wang H. Ca2+-stimulated adenylyl cyclases as therapeutic targets for psychiatric and neurodevelopmental disorders. Front Pharmacol. 2022;13:949384. 10.3389/fphar.2022.949384 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pinto JV, Passos IC, Librenza-Garcia D, Marcon G, Schneider MA, Conte JH, et al. Neuron-glia interaction as a possible pathophysiological mechanism of bipolar disorder. Curr Neuropharmacol. 2018;16:519–32. 10.2174/1570159X15666170828170921 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.International Consortium on Lithium G, Amare AT, Schubert KO, Hou L, Clark SR, Papiol S, et al. Association of polygenic score for schizophrenia and HLA antigen and inflammation genes with response to lithium in bipolar affective disorder: a genome-wide association study. JAMA Psychiatry. 2018;75:65–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yang M, Ding Q, Zhang M, Moon C, Wang H. Forebrain overexpression of type 1 adenylyl cyclase promotes molecular stability and behavioral resilience to physical stress. Neurobiol Stress. 2020;13:100237. 10.1016/j.ynstr.2020.100237 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mons N, Segu L, Nogues X, Buhot MC. Effects of age and spatial learning on adenylyl cyclase mRNA expression in the mouse hippocampus. Neurobiol Aging. 2004;25:1095–106. 10.1016/j.neurobiolaging.2003.10.014 [DOI] [PubMed] [Google Scholar]
- 41.Schroeter ML, Abdul-Khaliq H, Sacher J, Steiner J, Blasig IE, Mueller K. Mood disorders are glial disorders: evidence from in vivo studies. Cardiovasc Psychiatry Neurol. 2010;2010:780645. 10.1155/2010/780645 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bowley MP, Drevets WC, Ongur D, Price JL. Low glial numbers in the amygdala in major depressive disorder. Biol Psychiatry. 2002;52:404–12. 10.1016/S0006-3223(02)01404-X [DOI] [PubMed] [Google Scholar]
- 43.Maletic V, Raison C. Integrated neurobiology of bipolar disorder. Front Psychiatry. 2014;5:98. 10.3389/fpsyt.2014.00098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Myint AM. Kynurenines: from the perspective of major psychiatric disorders. FEBS J. 2012;279:1375–85. 10.1111/j.1742-4658.2012.08551.x [DOI] [PubMed] [Google Scholar]
- 45.Frost JL, Schafer DP. Microglia: architects of the developing nervous system. Trends Cell Biol. 2016;26:587–97. 10.1016/j.tcb.2016.02.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Dai N, Jones BDM, Husain MI. Astrocytes in the neuropathology of bipolar disorder: review of current evidence. Brain Sci. 2022;12:1513. 10.3390/brainsci12111513 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Jakobsson J, Bjerke M, Sahebi S, Isgren A, Ekman CJ, Sellgren C, et al. Monocyte and microglial activation in patients with mood-stabilized bipolar disorder. J Psychiatry Neurosci. 2015;40:250–8. 10.1503/jpn.140183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Vadodaria KC, Mendes APD, Mei A, Racha V, Erikson G, Shokhirev MN, et al. Altered neuronal support and inflammatory response in bipolar disorder patient-derived astrocytes. Stem Cell Rep. 2021;16:825–35. 10.1016/j.stemcr.2021.02.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Colombo E, Farina C. Astrocytes: key regulators of neuroinflammation. Trends Immunol. 2016;37:608–20. 10.1016/j.it.2016.06.006 [DOI] [PubMed] [Google Scholar]
- 50.Hsieh J, Gage FH. Chromatin remodeling in neural development and plasticity. Curr Opin Cell Biol. 2005;17:664–71. 10.1016/j.ceb.2005.09.002 [DOI] [PubMed] [Google Scholar]
- 51.Gavin DP, Kartan S, Chase K, Jayaraman S, Sharma RP. Histone deacetylase inhibitors and candidate gene expression: an in vivo and in vitro approach to studying chromatin remodeling in a clinical population. J Psychiatr Res. 2009;43:870–6. 10.1016/j.jpsychires.2008.12.006 [DOI] [PubMed] [Google Scholar]
- 52.Monneret C. Histone deacetylase inhibitors. Eur J Med Chem. 2005;40:1–13. 10.1016/j.ejmech.2004.10.001 [DOI] [PubMed] [Google Scholar]
- 53.Hilton IB, D’Ippolito AM, Vockley CM, Thakore PI, Crawford GE, Reddy TE, et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat Biotechnol. 2015;33:510–7. 10.1038/nbt.3199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zhang L, Verwer RWH, Lucassen PJ, Huitinga I, Swaab DF. Sex difference in glia gene expression in the dorsolateral prefrontal cortex in bipolar disorder: relation to psychotic features. J Psychiatr Res. 2020;125:66–74. 10.1016/j.jpsychires.2020.03.003 [DOI] [PubMed] [Google Scholar]
- 55.Xu Y, Gao W, Sun Y, Wu M. New insight on microglia activation in neurodegenerative diseases and therapeutics. Front Neurosci. 2023;17:1308345. 10.3389/fnins.2023.1308345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Saijo K, Glass CK. Microglial cell origin and phenotypes in health and disease. Nat Rev Immunol. 2011;11:775–87. 10.1038/nri3086 [DOI] [PubMed] [Google Scholar]
- 57.Ascoli BM, Gea LP, Colombo R, Barbe-Tuana FM, Kapczinski F, Rosa AR. The role of macrophage polarization on bipolar disorder: identifying new therapeutic targets. Aust N Z J Psychiatry. 2016;50:618–30. 10.1177/0004867416642846 [DOI] [PubMed] [Google Scholar]
- 58.Rosa PM, Martins LAM, Souza DO, Quincozes-Santos A. Glioprotective effect of resveratrol: an emerging therapeutic role for oligodendroglial cells. Mol Neurobiol. 2018;55:2967–78. 10.1007/s12035-017-0510-x [DOI] [PubMed] [Google Scholar]
- 59.Lloyd AJ, Ali HE, Nesbitt D, Moore PB, Young AH, Ferrier IN. Corpus callosum changes in euthymic bipolar affective disorder. Br J Psychiatry. 2014;204:129–36. 10.1192/bjp.bp.112.123687 [DOI] [PubMed] [Google Scholar]
- 60.Ibarretxe G, Sanchez-Gomez MV, Campos-Esparza MR, Alberdi E, Matute C. Differential oxidative stress in oligodendrocytes and neurons after excitotoxic insults and protection by natural polyphenols. Glia. 2006;53:201–11. 10.1002/glia.20267 [DOI] [PubMed] [Google Scholar]
- 61.Olympiou M, Sargiannidou I, Markoullis K, Karaiskos C, Kagiava A, Kyriakoudi S, et al. Systemic inflammation disrupts oligodendrocyte gap junctions and induces ER stress in a model of CNS manifestations of X-linked Charcot-Marie-Tooth disease. Acta Neuropathol Commun. 2016;4:95. 10.1186/s40478-016-0369-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Rajkowska G, Halaris A, Selemon LD. Reductions in neuronal and glial density characterize the dorsolateral prefrontal cortex in bipolar disorder. Biol Psychiatry. 2001;49:741–52. 10.1016/S0006-3223(01)01080-0 [DOI] [PubMed] [Google Scholar]
- 63.Escamez T, Bahamonde O, Tabares-Seisdedos R, Vieta E, Martinez S, Echevarria D. Developmental dynamics of PAFAH1B subunits during mouse brain development. J Comp Neurol. 2012;520:3877–94. 10.1002/cne.23128 [DOI] [PubMed] [Google Scholar]
- 64.McLean CK, Narayan S, Lin SY, Rai N, Chung Y, Hipolito MS, et al. Lithium-associated transcriptional regulation of CRMP1 in patient-derived olfactory neurons and symptom changes in bipolar disorder. Transl Psychiatry. 2018;8:81. 10.1038/s41398-018-0126-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhang G, Bai H, Zhang H, Dean C, Wu Q, Li J, et al. Neuropeptide exocytosis involving synaptotagmin-4 and oxytocin in hypothalamic programming of body weight and energy balance. Neuron. 2011;69:523–35. 10.1016/j.neuron.2010.12.036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ferguson GD, Vician L, Herschman HR. Synaptotagmin IV: biochemistry, genetics, behavior, and possible links to human psychiatric disease. Mol Neurobiol. 2001;23:173–85. 10.1385/MN:23:2-3:173 [DOI] [PubMed] [Google Scholar]
- 67.Xiang C, Baubet V, Pal S, Holderbaum L, Tatard V, Jiang P, et al. RP58/ZNF238 directly modulates proneurogenic gene levels and is required for neuronal differentiation and brain expansion. Cell Death Differ. 2012;19:692–702. 10.1038/cdd.2011.144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wang T, Hoekzema K, Vecchio D, Wu H, Sulovari A, Coe BP, et al. Large-scale targeted sequencing identifies risk genes for neurodevelopmental disorders. Nat Commun. 2020;11:4932. 10.1038/s41467-020-18723-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Huynh TKT, Mai TTT, Huynh MA, Yoshida H, Yamaguchi M, Dang TTP. Crucial roles of ubiquitin carboxy-terminal hydrolase L1 in motor neuronal health by Drosophila model. Antioxid Redox Signal. 2022;37:257–73. 10.1089/ars.2021.0057 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The BD GWAS data can be freely accessed via the PGC website at https://www.med.unc.edu/pgc/results-and-downloads. Data for Single-cell RNA-Seq (accession no. syn7067037), BD Bulk RNA-seq (accession no. syn3270015), and BD ATAC-Seq (accession no. syn7349497) were procured from the PsychENCODE Consortium, available on the Synapse platform. Interested researchers can request access to the PsychENCODE Consortium data by submitting an application form on the Synapse website at www.synapse.org/#!Synapse:syn4921369/wiki/235539. The datasets generated during and/or analyzed during the current study are available at https://github.com/weihuan12138/scmultiomics.git.
All software utilized in this research is openly accessible and can be located through the respective URLs or references provided. The code developed to generate all results presented in this study can be accessed at https://github.com/weihuan12138/scmultiomics.git.