Abstract
Background and Aims
The observed increase in the incidence of early-onset colorectal cancer (EOCRC) is being driven by sporadic cases, but the molecular characteristics of these tumors are not fully understood. Our objective was to investigate the prevalence of microsatellite instability (MSI) and selected mutations in sporadic EOCRC, and their association with survival.
Methods
Firstly, we compared the prevalence of molecular characteristics and survival within a population-based cohort study of 652 stage II and III colon cancer patients in Northern Ireland, comparing sporadic early-onset (<50 years, n = 35) with older (60–69 years, n = 179) patients. Secondly, a systematic review for studies reporting the prevalence of MSI, mismatch repair deficiency (dMMR), or BRAF, KRAS, NRAS, PIK3CA, and TP53 mutations in sporadic EOCRC was conducted. A meta-analysis was performed to calculate pooled estimates of the prevalence of molecular features in sporadic EOCRC.
Results
Firstly, within the cohort study, EOCRC patients did not have a significantly increased risk of colorectal cancer–specific death (adjusted hazard ratio 1.20; 95% confidence interval [CI] 0.61–2.39) compared with 60- to 69-year-olds. Second, 32 studies were included in the systematic review. The pooled analysis estimated a prevalence of 10% (95% CI 7%–14%) for MSI high/dMMR in sporadic EOCRC. BRAF and KRAS mutations had a prevalence of 1% (95% CI 0%–3%) and 32% (95% CI 23%–40%), respectively.
Conclusion
The molecular characteristics of sporadic EOCRC differ from those of cancers in older adults, particularly regarding reduced prevalence of BRAF mutations. Ten percent of sporadic EOCRC display MSI high/dMMR. Further studies are needed to address survival in sporadic EOCRC cases and whether molecular profiles influence EOCRC outcomes in this patient group.
Keywords: Microsatellite Instability, Mismatch Repair, Mutations
See editorial on page 301.
Introduction
An increase in the incidence of colorectal cancer (CRC) in adults younger than 50 years, known as early-onset CRC (EOCRC), has been observed in high-income countries.1, 2, 3, 4 Studies have suggested that the majority of EOCRC is sporadic in nature, with cases associated with identified germline mutations accounting for up to 35%.5,6 However, not all previous studies have separated sporadic from hereditary cases of EOCRC, and while the molecular pathogenesis of CRC due to inherited conditions such as Lynch syndrome is well defined, the pathways that lead to the development of sporadic EOCRC remain incompletely understood.
CRC is a molecularly heterogeneous disease resulting from stepwise accumulation of mutations in key oncogenes and tumor suppressor genes leading to the development of malignancy via a number of pathways, namely the chromosomal instability pathway (CIN), microsatellite instability (MSI) pathway, and the serrated pathway. Each pathway displays multiple characteristic gene mutations and epigenetic changes. CRCs developing via the CIN pathway are associated with mutations in APC as an early event, with subsequent mutations in RAS, RAF, PIK3CA, SMAD4, and/or TP53 genes, among others.7 KRAS and NRAS mutation status is used in the clinical setting to inform systemic treatment options.8 CRCs arising through the MSI pathway display deficient mismatch repair (dMMR), which is synonymous with MSI,9 resulting from uncorrected errors during DNA replication.10 Lynch syndrome, a hereditary condition predisposing to the development of several cancers, results in microsatellite-instability high (MSI-H) CRCs. However, due to the inclusion of both hereditary and sporadic cases of EOCRC in many studies to date, it is unclear how frequently sporadic MSI-H tumors occur in EOCRC cases. CRCs arising via the serrated pathway are MSI-H, associated with an increased prevalence of BRAF mutations (which is a distinguishing feature), and have high levels of CpG island methylation, known as the CpG island methylator phenotype (CIMP).11
A 2019 report reviewed 37 studies with regard to prognosis of EOCRC compared to late-onset CRC (LOCRC) and found conflicting results for a poorer, similar, or better prognosis in younger patients.12 It is possible that survival differences between EOCRC patients and older CRC patients may reflect the different molecular profiles of tumors occurring in these patients. However, to our knowledge, the evidence for molecular profiles of sporadic EOCRC tumors has not been systematically collated.
In the present study, we analyzed a population-based cohort of patients with stage II and III colon cancer to investigate molecular characteristics in sporadic CRCs and survival outcomes according to age categories. We also undertook a systematic review and meta-analysis of the prevalence of MSI status and selected tumor mutations in sporadic EOCRCs.
Methods
Population-Based Cohort Study
Patient Population
The study cohort (known as Epi700) was established as previously described.13, 14, 15 In summary, 661 stage II and III colon cancer patients diagnosed in 2 healthcare trusts in Northern Ireland from 2004 to 2008, for whom resection specimens were available to be retrieved from the Northern Ireland Biobank, were identified using the Northern Ireland Cancer Registry. Patients were followed up for recurrence and cause of death to December 31, 2013.
Tumor Pathology Characteristics and Clinical Data Collection
When tumor pathology characteristics were not readily available from routinely extracted cancer registry information, further pathology details, for example, tumor differentiation, were retrieved by manual review of pathology reports.
Clinical variables used in this study including family history of CRC, oncological treatments, Eastern Cooperative Oncology Group performance status, lifestyle information (including smoking and alcohol), and comorbidities were extracted from the Northern Ireland Clinical Oncology Information System, a prospective electronic record of patient management.
Tumor Molecular Analysis
Following tumor annotation and macrodissection, DNA was extracted according to the manufacturer’s instructions from 5-μm sections of representative whole-tumor blocks using the Maxwell 16 instrument (Promega, Southampton, UK) and Promega DNA extraction kit (Promega, Southampton, UK).
MSI analysis was performed within the Northern Ireland Molecular Pathology Laboratory, using the MSI Analysis System, version 1.2, kit (Promega, Southampton, UK) for 5 mononucleotide repeat markers (BAT-25, BAT-26, NR-21, NR-24, and MONO-27). PCR products were separated by capillary electrophoresis using an ABI 3500 Genetic Analyzer (Fisher Scientific, UK Ltd, Loughborough, UK). The output data were analyzed using GeneMapper v4.1 (Fisher Scientific, UK Ltd, Loughborough, UK) to determine MSI status.9
Tumor samples were analyzed for mutational status of established CRC markers. This included a ColoCarta panel of KRAS, NRAS, BRAF, CMET, and PIK3CA using a validated mass spectrometry–based targeted screening panel of 32 somatic mutations in 6 genes (Agena Bioscience, Hamburg, Germany). Samples were shipped via the Genomics Core Technology Unit (Queen’s University Belfast) and the assays performed by Agena Custom Services Laboratory (Hamburg, Germany).
Statistical Analysis
A statistical analysis was performed using Stata 16 (StataCorp, 2019. Stata Statistical Software: Release 16. StataCorp LLC, College Station, TX). Chi-squared tests were used to compare descriptive and molecular characteristics across age categories. A survival analysis was performed using the Cox proportional hazards model to calculate hazard ratios (HRs) and 95% confidence intervals (CIs). The multivariable model for CRC-specific survival included sex, family history of CRC, stage (II/III), grade/differentiation, adjuvant chemotherapy receipt, Eastern Cooperative Oncology Group performance status, alcohol, smoking, inflammatory bowel disease, and emergency surgery. Results from this study were included in the subsequent systematic review and meta-analysis.
Systematic Review and Meta-Analysis
This study was reported according to the Meta-analysis of Observation Studies in Epidemiology (MOOSE) checklist.16 The review protocol was registered on PROSPERO (CRD42021232567).
Study Population
The population of interest was patients with sporadic EOCRC, defined as adults younger than 50 years at their incident CRC diagnosis who had no identified inherited genetic syndrome that predisposes to CRC. Studies were included if sporadic cases were separated from hereditary cases by the authors or if the article contained information to enable distinction of hereditary from sporadic cases, such as results of genetic testing or family history.
Outcome
The primary outcome was to estimate the prevalence of MSI-H/dMMR status and KRAS, NRAS, BRAF, PIK3CA, and TP53 mutations in EOCRC. The protocol specified a secondary outcome investigating the influence of molecular profile on survival in EOCRC, but insufficient data were available in potentially eligible articles and so we restricted the reporting of the review to the prevalence of molecular features as outlined.
Search Strategy
The electronic databases Ovid Medline, Embase, and Web of Science were systematically searched from 2000 to April 12, 2021. The full search terms are available in Supplementary Appendix 1. Observational studies, descriptive studies, case series, and interventional studies were eligible for inclusion. All stages of CRC were included, with a focus on colorectal adenocarcinoma as the primary histology, and no language restrictions were imposed. Review articles, editorials, comments, abstract or conference proceedings, individual case studies, and case series with less than 10 patients were excluded.
Articles from the search were imported into Covidence, and duplicates were removed. Titles and abstracts were reviewed by 2 authors independently (A.C.H. and H.G.C.). The full text of all selected articles was read by 2 authors (A.C.H. reviewed all articles; H.G.C. and M.B.L. reviewed independent subsets). Any discrepancies were resolved by discussion among the 2 reviewers of the text, with the third reviewer involved if required.
Data Extraction
Data were extracted by A.C.H. and verified by H.G.C. Data extracted were related to study location, number of sporadic EOCRC cases, definition of sporadic cases, mutation testing, MSI or dMMR testing methods, and the prevalence of each molecular characteristic in the study population, along with sex and anatomical tumor location if available. Where mutation testing had resulted in an unknown or ambiguous result, our approach was to exclude these cases from the analysis. Where a study contained data for both MSI status and MMR proteins, the MSI status results were used. Details of molecular testing undertaken in each study are shown in Table A4. Immunohistochemistry for the p53 protein has been used as a surrogate for TP53 mutation testing,17 where reported in studies. In a change to the study protocol, the Joanna Briggs Institute Checklist for Prevalence Studies was used for quality assessment.
Statistical Analysis
Stata 16 was used to perform meta-analysis to produce pooled estimates of prevalence and 95% CI. The Freeman-Tukey Arcsine Transformation method was used to calculate estimates and standard errors, which were back-transformed to calculate a pooled prevalence.18 The logistic-normal random-effects model was also carried out to ensure the pooled estimates and 95% CI were similar. A sensitivity analysis was performed, and subgroup analysis was carried out by sex and anatomical tumor location. The heterogeneity between studies was determined using the I2 statistic.19 Publication bias was assessed using funnel plots.
Results
Population-Based Cohort Study
The number of patients in the cohort for whom a surgical resection specimen was retrieved for molecular analyses was 661. Nine patients had a known hereditary cancer syndrome: Lynch syndrome (n = 6), familial adenomatous polyposis (n = 1), and other familial syndromes (n = 2). These patients were excluded from the analyses, leaving a cohort of 652 patients with presumed sporadic CRCs.
Patients younger than 50 years (n = 35) comprised 5.4% of our cohort of all stage II and III sporadic colon adenocarcinoma patients within the jurisdiction of the Northern Ireland Biobank. The demographic and clinical characteristics of the overall cohort are summarized in Table A1.
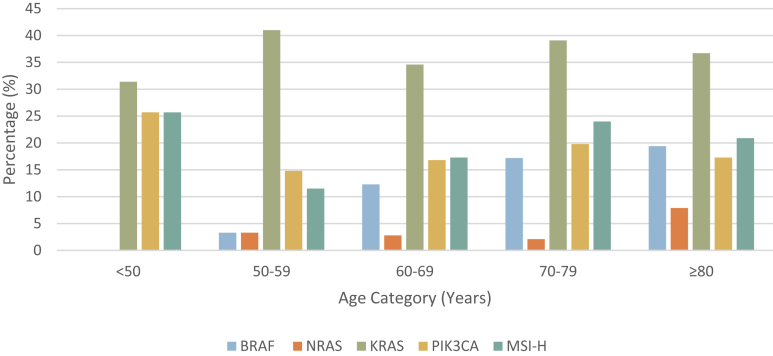
The distribution of mutations and molecular features by age category is shown in Figure 1. EOCRC patients did not have any BRAF or NRAS-mutant tumors, and the proportions of these features across age categories were significant (P < .01 and P = .01, respectively). EOCRCs had the highest proportion of MSI-H tumors (25.7%) and PIK3CA mutations (25.7%) of all the age groups, but this did not reach statistical significance.
Figure 1.
Distribution of molecular characteristics by age category in sporadic stage II and III colon cancer.
The results of survival analyses are shown in Table 1. Compared with 60 to 69-year-old patients, EOCRC patients did not have a significantly increased risk of CRC death in stage II/III disease (adjusted HR 1.20, 95% CI 0.61–2.39). Compared to patients with microsatellite-stable (MSS) tumors, patients with MSI-H tumors had a significantly reduced risk of CRC death (unadjusted HR 0.66, 95% CI 0.45–0.97) in stage II/III disease. In multivariable analysis, patients with MSI-H tumors had a reduced risk of CRC death, but this was not statistically significant (HR 0.71, 95% CI 0.47–1.09). Subgroup analyses of survival by MSI status are shown in Figure 2. EOCRC patients with MSI-H tumors did not have a significantly decreased risk of CRC death (adjusted HR 0.65, 95% CI 0.07–6.32). EOCRC patients with MSS tumors did not have a significantly increased risk of CRC death compared to 60 to 69-year-olds (adjusted HR 1.58, 95% CI 0.71–3.51).
Table 1.
Survival Analysis by Age Category in Sporadic Stage II and III Colon Cancer
| Age (y) | CRC death |
Overall survival |
||||
|---|---|---|---|---|---|---|
| No of CRC deaths/patients | Unadjusted HR (95% CI) | Adjusted HRa (95% CI) | No of deaths/CRC patients | Unadjusted HR (95% CI) | Adjusted HRb (95% CI) | |
| <50 | 11/34 | 1.14 (0.59–2.18) | 1.20 (0.61–2.39) | 12/35 | 1.14 (0.79–1.64) | 1.19 (0.81–1.75) |
| 50–59 | 16/58 | 0.83 (0.47–1.45) | 0.83 (0.47–1.46) | 19/61 | 0.75 (0.56–1.01) | 0.76 (0.57–1.03) |
| 60–69 | 52/170 | 1 (reference) | 1 (reference) | 61/179 | 1 (reference) | 1 (reference) |
| 70–79 | 70/193 | 1.04 (0.73–1.49) | 0.88 (0.60–1.28) | 115/238 | 0.98 (0.81–1.19) | 0.95 (0.77–1.17) |
| ≥80 | 61/104 | 1.96 (1.35–2.84) | 1.38 (0.91–2.10) | 96/139 | 1.66 (1.33–2.08) | 1.49 (1.16–1.91) |
ECOG, Eastern Cooperative Oncology Group.
Multivariable model adjusted HR, adjusted for sex, adjuvant chemotherapy receipt, stage, tumor differentiation, family history of CRC, ECOG, performance status, alcohol, smoking, inflammatory bowel disease, and emergency surgery.
Multivariable model adjusted for all variables in footnote “a” and Charlson comorbidity score.
Figure 2.
Risk of CRC–specific death according to age categories in stage II and III colon cancer cases, by microsatellite instability status. ∗∗Adjusted for sex, adjuvant chemotherapy receipt, stage, tumor differentiation, family history of CRC, ECOG performance status, smoking, alcohol, inflammatory bowel disease and emergency surgery. CI, confidence interval; CRC, colorectal cancer; ECOG, Eastern Cooperative Oncology Group; HR, hazard ratio; MSI, microsatellite instability.
Systematic Review and Meta-Analysis
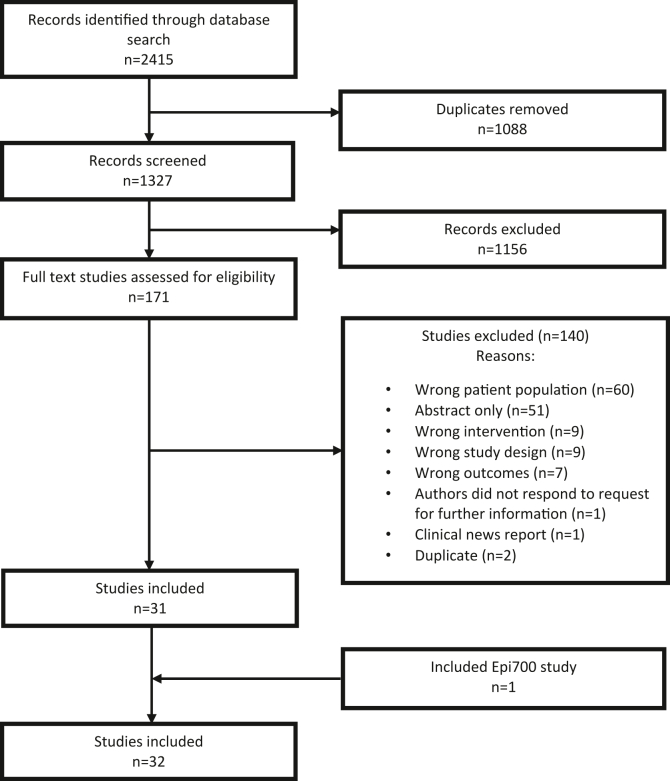
The search strategy identified 2415 studies, and 1088 duplicates were removed by Covidence, leaving 1327 articles for screening. Following title and abstract screening, 170 studies were eligible for full-text review. Following the full-text review, 140 studies were excluded for the reasons outlined in Figure 3. Thirty-two articles were included in the review (31 resulting from our search strategy and 1 resulting from our population-based cohort study described above, referred to herein as Hamilton et al, 2022). The characteristics of the included studies are summarized in Table A2. The rationale for determination of sporadic cases in each study is shown in Table A3.
Figure 3.
Flow chart of the selection of articles included in the review.
Microsatellite Instability/Deficient Mismatch Repair
Twenty-six studies reported data on tumor MSI status,20, 21, 22, 23, 24, 25, 26, 27, 28, 29, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44 including our population-based cohort study. Fifteen studies reported data on expression of MMR proteins, indicating MMR status.21,23,24,26,27,33,37,41,43,45, 46, 47, 48, 49, 50 A combined meta-analysis of studies that had MSI and/or MMR results was performed, and this is shown in Figure 4A. Pooled analysis revealed a prevalence of 10% (95% CI 7%–14%) of MSI-H/dMMR in presumed sporadic EOCRCs. Observed heterogeneity was high (I2 85.73%, P < .01). Separate meta-analyses were carried out for MSI-H and dMMR tumors (Figures A1 and A2).
Figure 4.
(A) Forest plot illustrating meta-analysis of the prevalence of microsatellite instability-high/deficient mismatch repair tumors in sporadic early-onset colorectal cancer. (B) Forest plot illustrating meta-analysis of the prevalence of BRAF mutations in sporadic early-onset colorectal cancer. (C) Forest plot illustrating meta-analysis of the prevalence of KRAS mutations in sporadic early-onset colorectal cancer. (D) Forest plot illustrating meta-analysis of the prevalence of NRAS mutations in sporadic early-onset colorectal cancer. (E) Forest plot illustrating meta-analysis of the prevalence of PIK3CA mutations in sporadic early-onset colorectal cancer. (F) Forest plot illustrating meta-analysis of the prevalence of TP53 mutations in sporadic early-onset colorectal cancer. Studies of Ak et al through Hamilton et al have presented microsatellite instability-high data, determined by PCR; studies of Aitchison et al through Suzuki et al have used mismatch repair immunohistochemistry. Details of molecular testing for each study are found in Table A4. CI, confidence interval; dMMR, deficient mismatch repair; ES, effect size (equivalent to proportion); MSI-H, microsatellite instability-high; PCR, polymerase chain reaction.
BRAF Mutations
Nine studies reported data on BRAF mutations,22,27,31,33,37,43,44,48 including our population-based cohort study. Pooled analysis revealed a prevalence of 1% (95% CI 0%–3%) for BRAF mutations in sporadic EOCRC (Figure 4B). The observed heterogeneity was moderate (I2 36.78%, P = .12).
KRAS Mutations
Thirteen studies reported data on KRAS mutations,22,24,27,31, 32, 33,37,38,40,43,44,48 including our population-based cohort study. Pooled analysis revealed a prevalence of 32% (95% CI 23%–40%) for KRAS mutations in sporadic EOCRC (Figure 4C). The observed heterogeneity was high (I2 84.52%, P < .01).
NRAS Mutations
Four studies reported data on NRAS mutations,43,44,48 including our population-based cohort study. Pooled analysis revealed a prevalence of 3% (95% CI 1%–4%) for NRAS mutations in sporadic EOCRCs (Figure 4D). The observed heterogeneity was low (I2 0.00%, P = .42).
PIK3CA Mutations
Five studies reported data on PIK3CA mutations,22,31,44,48 including our population-based cohort study. Pooled analysis revealed a prevalence of 14% (95% CI 5%–25%) for PIK3CA mutations in sporadic EOCRC (Figure 4E). The observed heterogeneity was high (I2 83.31%, P < .01).
TP53 Mutations
Seven studies reported data on TP53 mutations.22, 23, 24,31,38,40,44 Pooled analysis revealed a prevalence of 64% (95% CI 56%–71%) for TP53 mutations in sporadic EOCRC (Figure 4F). The observed heterogeneity was moderate (I2 57.80%, P = .03).
Sensitivity and Subgroup Analyses
Sensitivity analyses were performed by excluding one study at a time for each of the mutations and MSI/MMR status, to assess the robustness of the results. Results are shown in Table A5. Excluding some studies had a marginal effect on heterogeneity for BRAF, PIK3CA, TP53, and NRAS mutations, but the prevalence of these mutations remained similar.
Subgroup analyses were undertaken by sex for MSI/MMR status, KRAS, BRAF, and PIK3CA mutations (Table 2). There were insufficient data to undertake subgroup analyses for TP53 and NRAS mutations. The prevalence of MSI-H/dMMR and BRAF and PIK3CA mutations was similar between male and female patients. The prevalence of KRAS mutations was slightly higher in female patients (35%) than that in male patients (27%).
Table 2.
Subgroup Analysis by Sex and Tumor Location
| Mutation | Studies included | Pooled analysis (95% CI) | I2 (%) | P value |
|---|---|---|---|---|
| Males | ||||
| MSI-high/dMMR | 9 | 0.16 (0.09–0.24) | 55.53 | .02 |
| BRAF | 3 | 0.02 (0.00–0.09) | 27.43 | .25 |
| KRAS | 4 | 0.27 (0.16–0.38) | 0.00 | .78 |
| PIK3CA | 3 | 0.11 (0.00–0.35) | 79.80 | .01 |
| Females | ||||
| MSI-high/dMMR | 9 | 0.14 (0.07–0.21) | 46.86 | .06 |
| BRAF | 3 | 0.02 (0.00–0.13) | 55.04 | .11 |
| KRAS | 4 | 0.35 (0.23–0.48) | 0.00 | .54 |
| PIK3CA | 3 | 0.13 (0.00–0.40) | 81.83 | .00 |
| All colon | ||||
| MSI-high/dMMR | 7 | 0.16 (0.06–0.29) | 74.74 | .00 |
| BRAF | 3 | 0.04 (0.00–0.14) | 58.32 | .09 |
| KRAS | 3 | 0.34 (0.24–0.45) | 0.00 | .72 |
| PIK3CA | 3 | 0.12 (0.00–0.37) | 85.80 | .00 |
| Right colon | ||||
| MSI-high/dMMR | 6 | 0.32 (0.19–0.46) | 43.95 | .11 |
| BRAF | 3 | 0.02 (0.00–0.14) | 54.22 | .11 |
| KRAS | 3 | 0.35 (0.22–0.50) | 0.00 | .84 |
| PIK3CA | 3 | 0.18 (0.01–0.46) | 77.75 | .01 |
| Left colon | ||||
| MSI-high/dMMR | 6 | 0.03 (0.00–0.12) | 47.82 | .09 |
| BRAF | 3 | 0.04 (0.00–0.17) | 17.24 | .30 |
| KRAS | 3 | 0.33 (0.17–0.51) | 0.00 | .76 |
| PIK3CA | 3 | 0.01 (0.00–0.10) | 0.00 | .50 |
| Rectum | ||||
| MSI-high/dMMR | 6 | 0.06 (0.01–0.13) | 12.59 | .33 |
Subgroup analyses were undertaken by tumor location (Table 2). Analyses for colon were performed for MSI/MMR status; KRAS, BRAF, and PIK3CA mutations; and for rectum by MSI/MMR status. There were insufficient data for TP53 and NRAS mutations for any subgroup analysis by tumor location or for KRAS, BRAF, and PIK3CA mutations for rectal cancer. Results showed that the prevalence of MSI-H/dMMR was higher in the colon than in the rectum (16% vs 6%) and higher in the right colon than in the left colon (32% vs 3%). The prevalence of KRAS and BRAF mutations was similar in the right and left colon, while PIK3CA mutations showed a higher prevalence in the right colon than in the left colon (18% vs 1%).
Publication Bias
Publication bias was assessed using funnel plots where we plotted the proportion against the study size (Figure A3). No evidence of publication bias was detected.
Quality Assessment
Quality assessment was done using the Joanna Briggs Institute Checklist for prevalence studies, and this is shown in Table A6. No studies were excluded based on quality assessment.
Discussion
To our knowledge, this is the first systematic review investigating the prevalence of the MSI-H/dMMR status and somatic mutations in sporadic EOCRC. The importance of distinguishing sporadic from hereditary EOCRC is becoming increasingly recognized, with the molecular pathogenesis, treatment response, and outcomes of sporadic EOCRC less understood than those of hereditary cases.
MSI/MMR Status
Our systematic review shows that MSI-H/dMMR has a prevalence of 10% in sporadic EOCRC. These results were consistent across analyses for MSI-H tumors and dMMR tumors, suggesting that MSI and MMR statuses are highly correlated and that our findings are robust. Other studies have shown near-perfect concordance between immunohistochemistry testing for MMR proteins and MSI testing.9
MSI-H/dMMR CRCs are encountered in 2 clinical settings, representing the phenotypic convergence of 2 clinically distinct pathogeneses. Firstly, such tumors are the hallmark of CRCs arising in the context of Lynch syndrome, resulting from a germline mutation in one of the MMR genes, most commonly MLH1 or MSH2, and young age at cancer diagnosis is regarded as an indicator for a possible hereditary cause of the disease. Our results show that 10% of MSI-H/dMMR tumors in EOCRCs do not arise from Lynch syndrome. However, given the historical case series in reported studies, it is possible that some cases of seemingly sporadic EOCRC in our review may have undiagnosed Lynch syndrome or Lynch-like syndrome.51, 52, 53
Secondly, MSI-H tumors comprise a proportion of sporadic CRCs, and these are considerably more common than Lynch syndrome-related MSI-H CRCs. Sporadic MSI-H CRCs arise via the serrated neoplasia pathway and are strongly associated with BRAF mutations, older age, right-sided tumor location, and high levels of CIMP.54 Given the extremely low prevalence of BRAF mutations in EOCRC, it is unlikely the serrated pathway is the mechanism by which MSI-H tumors develop in younger patients. Similar to Lynch syndrome, results from the subgroup analysis suggest that sporadic MSI-H EOCRCs also have a predilection for the right colon.
CIMP-high CRCs are associated with older-age patients, female sex, proximal tumor location, MSI-H status, and somatic BRAF mutation.55 Evidence regarding CIMP in EOCRCs is sparse, but CIMP-high tumors appear to be less prevalent in younger patients with CRC.56 However, future studies are required to elucidate the role of CIMP in sporadic EOCRC.
BRAF Mutations
BRAF mutations occur in approximately 8% of all CRCs, the vast majority being V600E mutations.57 Our results show that the prevalence of BRAF mutations in sporadic EOCRC is 1%, which is lower than that in LOCRC and indicates this is a rare mutation in younger adults. Given the rarity of BRAF mutations in EOCRC, our results also suggest the association of BRAF mutations with MSI-H tumors seen in CRC in older-age groups58 does not apply to these younger patients.
BRAF mutations are a negative prognostic marker, with worse survival outcomes being reported in a metastatic disease.59 However, our results suggest a low prevalence of BRAF mutations in EOCRC does not necessarily translate into better survival in this group, and the reasons for this are unclear. In 2021, the National Institute for Health and Care Excellence approved the use of encorafenib, a BRAF inhibitor, in BRAF V600E-mutation-positive metastatic CRC in the United Kingdom.60 However, given the rarity of BRAF mutations in EOCRC, it is likely only a small proportion of young patients will be able to avail this treatment, and optimal treatment strategies for EOCRC remain to be determined.
RAS Mutations
Our results show that the prevalence of KRAS mutations in sporadic EOCRC is 32%. A large Memorial Sloan Kettering Cancer Centre study published after our literature search reported a prevalence of 42.5% for KRAS mutations in sporadic EOCRCs.61 Together with our meta-analysis, these results are comparable to a systematic review investigating KRAS mutations in metastatic CRCs, which reported a pooled prevalence of 35.9%.62 This suggests that the prevalence of KRAS mutations is broadly similar in EOCRC and LOCRC.
Knowledge regarding NRAS mutations in CRC is limited due to its low frequency. A systematic review found a prevalence of 4.1% (95% CI 3.5%–4.8%) of NRAS mutations in metastatic CRC in all ages.62 Our results show that the prevalence of NRAS mutations in tumors in sporadic EOCRCs is 3%, suggesting the prevalence of NRAS mutations is similar in young and older patients.
PIK3CA Mutations
A 2020 systematic review reported PIK3CA mutations had a prevalence of 12.9% in CRC in patients of all ages.63 Our results show that the prevalence of PIK3CA mutations in tumors in sporadic EOCRC is 14%, suggesting that the proportion of PIK3CA mutations is similar in EOCRC and LOCRC. However, knowledge regarding PIK3CA mutations in EOCRC remains limited, as shown by the small number of studies in our meta-analysis. PIK3CA mutations currently have no clinical role as predictive or prognostic biomarkers, with a previous systematic review and meta-analysis finding no significant association between PIK3CA mutation status and survival outcomes.64
TP53 Mutations
Mutation of TP53 is a late event in the stepwise development of CRC, most commonly via the CIN pathway.7 TP53 mutations have been shown to be present in up to 60% of CRCs.65 Our results show that the prevalence of TP53 mutations in sporadic EOCRC is 64%, the highest prevalence of any mutation in this study. Similar findings were observed in a whole-exome sequencing study which found TP53 was the most common mutation in EOCRC,66 with subsequent targeted deep sequencing (n = 833) showing a higher frequency of TP53 mutation in EOCRC than in LOCRC (80% vs 72%, Fisher’s exact P = 0.03). TP53 is currently not used as a prognostic or predictive biomarker in clinical practice, and more research is required into its clinical implications.
Survival
Results from our population-based cohort study indicate that stage II and III sporadic EOCRC does not have a significantly worse survival compared with those in patients aged 60–69 years but indicate there may be an aggressive subset within this young age group, driven by MSS tumors. CRCs displaying MSI have a better prognosis in an early-stage disease, with improved 5-year overall survival67 and 5-year recurrence-free survival,68 but less expected benefit from adjuvant chemotherapy.67,69 Results from our population-based cohort study support the conclusion that non-metastatic MSI-H tumors in sporadic EOCRC patients also carry a better prognosis than MSS tumors, although sample sizes were limited and so results were not statistically significant. We are unable to draw any conclusions about survival in metastatic sporadic EOCRC from this study.
Studies have shown that EOCRC patients have a more advanced stage at presentation than older patients, 70 which could be due to more aggressive biology or a delay in diagnosis.
Delayed diagnosis may be caused by a number of factors, including failure of younger patients to seek healthcare, a delay in referral by healthcare professionals, or the exclusion of younger individuals from bowel cancer screening programs.
Further research is urgently needed on outcomes for patients with EOCRC and more specifically on the impact of tumor molecular profile on survival, particularly how this varies by stage of disease.
Strengths and Limitations
Our study has a number of strengths. To our knowledge, this the first systematic review that determines the prevalence of key mutations and MSI-H/dMMR in sporadic EOCRC. The quality assessment was undertaken using the Joanna Briggs Institute Checklist for Prevalence Studies, which was felt to be rigorous in a recent systematic review.71 Sensitivity analyses demonstrated largely stable heterogeneity, particularly for MSI-H/dMMR tumors.
One weakness of the systematic review is that despite attempts to ensure that only sporadic EOCRC cases were included, there may be undiagnosed hereditary cases in our review. However, our methodology was rigorous, and studies were included if the information provided allowed us to confidently separate, as far as possible, sporadic from hereditary cases. For meta-analyses, we have used the Freeman-Tukey Arcsine Transformation method. A weakness of this method is that it breaks down with extremely sparse data.72 To ensure the accuracy of our results, we also carried out meta-analyses using the logistic-normal random-effects model, which showed similar pooled proportions and 95% CI. In addition, we were unable to undertake subgroup analysis by stage or race/ethnicity.
The studied molecular characteristics of CRC vary with stage of disease. For example, MSI-H occurs in approximately 15%–20% of stage II and III CRC but is less common in metastatic CRC, occurring in approximately 4% of cases.73,74 BRAF mutations are associated with an advanced stage of disease.75 Insufficient information was available to undertake subgroup analyses by stage in our study, but variation in molecular profile by stage may account for some of the observed differences between studies. This is an important issue to address in future studies.
Studies investigating the molecular profile of rectal cancer are lacking, and within this subgroup, we were only able to undertake an analysis for MSI/MMR status as the number of studies describing the selected mutations in rectal cancers was insufficient. Further research is needed into how rectal cancer differs from colon cancer in terms of mutational profile. In addition, newly discovered germline mutations in genes such as POLE and POLD1 will be contributing to a small percentage of EOCRC.76 However, while POLE/POLD1-mutated CRCs share some features with MSI-H CRCs (such as a high tumor mutation burden), they are typically MSS tumors and are unlikely to account for any of the sporadic MSI-H EOCRC cases in our meta-analysis.77
Conclusion
This systematic review addresses a research gap regarding sporadic EOCRC and provides evidence of differing molecular profiles in younger patients with CRC compared to older patients. Approximately 10% of seemingly sporadic EOCRCs are MSI-H, and BRAF mutations are a rare event in these tumors, having a much lower prevalence than in LOCRC. KRAS, NRAS, PIK3CA, and TP53 mutations have a similar prevalence to LOCRC. The molecular pathogenesis of sporadic EOCRC remains unclear, with the serrated neoplasia pathway unlikely to play a major role. EOCRC patients were not at increased risk of cancer-specific death compared with older patients in our population-based cohort, but further studies are needed to address whether molecular profiles differentially influence EOCRC patient outcomes in this patient group.
Acknowledgments
The authors would like to acknowledge the contributions of staff members who assisted with the creation of the Epi700 cohort data. The samples used in this research were received from the Northern Ireland Biobank13 which is funded by the Health and Social Care Research and Development Division of the Public Health Agency in Northern Ireland and Cancer Research UK (via the former Belfast CRUK Centre and the Northern Ireland Experimental Cancer Medicine Centre); additional support was received from the Friends of the Cancer Centre. The Northern Ireland Molecular Pathology Laboratory, which was responsible for creating resources for the Northern Ireland Biobank, has received funding from Cancer Research UK, Friends of the Cancer Centre, and the Sean Crummey Foundation. Clinical data collected and analyzed were facilitated by the Northern Ireland Cancer Registry, which is funded by the Public Health Agency, Northern Ireland.
Footnotes
Authors’ Contributions: Ashleigh C. Hamilton: Conceptualization, data curation, formal analysis, funding acquisition, investigation, methodology, project administration, software, visualization, writing—original draft. Finian J. Bannon: Conceptualization, methodology, writing—review and editing. Philip D. Dunne: Conceptualization, methodology, visualization, writing—review and editing. Chris R. Cardwell: Data curation, formal analysis, methodology, software, writing—review and editing. Maurice B. Loughrey: Conceptualization, methodology, investigation, validation, supervision, visualization, writing—original draft, writing—review and editing. Helen G. Coleman: Conceptualization, methodology, formal analysis, funding acquisition, investigation, project administration, software, supervision, validation, visualization, writing—original draft, writing—review and editing.
Conflicts of Interest: The authors disclose no conflicts.
Funding: Including departmental or institutional funding when no extramural funding was received; Ashleigh Hamilton is funded by a 2019 Health and Social Care (HSC) Research and Development Doctoral Fellowship (EAT/15494/18) from the R&D Division of the Public Health Agency in Northern Ireland. Helen Coleman is funded by a Cancer Research UK Career Establishment Award (reference: C37703/A25820). The resource creation and data analyzed for the Epi700 cohort analysis were supported by a HSC Research and Development Division of the Public Health Agency Doctoral Fellowship (EAT/4905/13—Ronan Gray), a Cancer Research UK Research Bursary (C50104/A17592—Ronan Gray), and a CRUK Population Health Postdoctoral Fellowship (C37703/A15333—Helen Coleman).
Ethical Statement: The corresponding author, on behalf of all authors, jointly and severally, certifies that their institution has approved the protocol for any investigation involving humans or animals and that all experiments were conducted in conformity with ethical and humane principles of research.
Data Transparency Statement: Applications for secondary use of data generated and presented in this manuscript, or materials described, as part of the Epi700 cohort creation, can be made by contacting the Northern Ireland Biobank, email: nibiobank@qub.ac.uk.
Reporting Guidelines: PRISMA, MOOSE, SAGER.
Material associated with this article can be found in the online version at https://doi.org/10.1016/j.gastha.2022.11.005.
Supplementary Materials
References
- 1.Lui R.N., Tsoi K.K.F., Ho J.M.W., et al. Global increasing incidence of young-onset colorectal cancer across 5 continents: a joinpoint regression analysis of 1,922,167 cases. Cancer Epidemiol Biomarkers Prev. 2019;28:1275–1282. doi: 10.1158/1055-9965.EPI-18-1111. [DOI] [PubMed] [Google Scholar]
- 2.Siegel R.L., Fedewa S.A., Anderson W.F., et al. Colorectal cancer incidence patterns in the United States, 1974-2013. J Natl Cancer Inst. 2017;109 doi: 10.1093/jnci/djw322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Vuik F.E., Nieuwenburg S.A., Bardou M., et al. Increasing incidence of colorectal cancer in young adults in Europe over the last 25 years. Gut. 2019;68:1820–1826. doi: 10.1136/gutjnl-2018-317592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sung J.J.Y., Chiu H.M., Jung K.W., et al. Increasing trend in young-onset colorectal cancer in Asia: more cancers in men and more rectal cancers. Am J Gastroenterol. 2019;114:322–329. doi: 10.14309/ajg.0000000000000133. [DOI] [PubMed] [Google Scholar]
- 5.Mork M.E., You Y.N., Ying J., et al. High prevalence of hereditary cancer syndromes in adolescents and young adults with colorectal cancer. J Clin Oncol. 2015;33:3544–3549. doi: 10.1200/JCO.2015.61.4503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Stoffel E.M., Koeppe E., Everett J., et al. Germline genetic features of young individuals with colorectal cancer. Gastroenterology. 2018;154:897–905.e1. doi: 10.1053/j.gastro.2017.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Nguyen L.H., Goel A., Chung D.C. Pathways of colorectal carcinogenesis. Gastroenterology. 2020;158:291–302. doi: 10.1053/j.gastro.2019.08.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Van Cutsem E., Cervantes A., Adam R., et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann Oncol. 2016;27:1386–1422. doi: 10.1093/annonc/mdw235. [DOI] [PubMed] [Google Scholar]
- 9.Loughrey M.B., McGrath J., Coleman H.G., et al. Identifying mismatch repair-deficient colon cancer: near-perfect concordance between immunohistochemistry and microsatellite instability testing in a large, population-based series. Histopathology. 2021;78:401–413. doi: 10.1111/his.14233. [DOI] [PubMed] [Google Scholar]
- 10.Shia J. The diversity of tumours with microsatellite instability: molecular mechanisms and impact upon microsatellite instability testing and mismatch repair protein immunohistochemistry. Histopathology. 2021;78:485–497. doi: 10.1111/his.14271. [DOI] [PubMed] [Google Scholar]
- 11.Weisenberger D.J., Siegmund K.D., Campan M., et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006;38:787–793. doi: 10.1038/ng1834. [DOI] [PubMed] [Google Scholar]
- 12.Mauri G., Sartore-Bianchi A., Russo A.G., et al. Early-onset colorectal cancer in young individuals. Mol Oncol. 2019;13:109–131. doi: 10.1002/1878-0261.12417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lewis C., McQuaid S., Clark P., et al. The Northern Ireland Biobank: a cancer focused repository of science. Open J Bioresour. 2018;5:9. [Google Scholar]
- 14.Gray R.T., Loughrey M.B., Bankhead P., et al. Statin use, candidate mevalonate pathway biomarkers, and colon cancer survival in a population-based cohort study. Br J Cancer. 2017;116:1652–1659. doi: 10.1038/bjc.2017.139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gray R.T., Cantwell M.M., Coleman H.G., et al. Evaluation of PTGS2 expression, PIK3CA mutation, aspirin use and colon cancer survival in a population-based cohort study. Clin Transl Gastroenterol. 2017;8:e91. doi: 10.1038/ctg.2017.18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Stroup D.F., Berlin J.A., Morton S.C., et al. Meta-analysis of observational studies in epidemiology: a proposal for reporting. JAMA. 2000;283:2008–2012. doi: 10.1001/jama.283.15.2008. [DOI] [PubMed] [Google Scholar]
- 17.Hall P.A., McCluggage W.G. Assessing p53 in clinical contexts: unlearned lessons and new perspectives. J Pathol. 2006;208:1–6. doi: 10.1002/path.1913. [DOI] [PubMed] [Google Scholar]
- 18.Miller J.J. The inverse of the Freeman – Tukey Double Arcsine Transformation. Am Stat. 1978;32:138. [Google Scholar]
- 19.Higgins J.P., Thompson S.G. Quantifying heterogeneity in a meta-analysis. Stat Med. 2002;21:1539–1558. doi: 10.1002/sim.1186. [DOI] [PubMed] [Google Scholar]
- 20.Ak S., Tunca B., Yilmazlar T., et al. Microsatellite instability status affects gene expression profiles in early onset colorectal cancer patients. J Surg Res. 2013;185:626–637. doi: 10.1016/j.jss.2013.07.014. [DOI] [PubMed] [Google Scholar]
- 21.Antelo M., Golubicki M., Roca E., et al. Lynch-like syndrome is as frequent as Lynch syndrome in early-onset nonfamilial nonpolyposis colorectal cancer. Int J Cancer. 2019;145:705–713. doi: 10.1002/ijc.32160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Berg M., Danielsen S.A., Ahlquist T., et al. DNA sequence profiles of the colorectal cancer critical gene set KRAS-BRAF-PIK3CA-PTEN-TP53 related to age at disease onset. PLoS One. 2010;5 doi: 10.1371/journal.pone.0013978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dieumegard B., Grandjouan S., Sabourin J.C., et al. Extensive molecular screening for hereditary non-polyposis colorectal cancer. Br J Cancer. 2000;82:871–880. doi: 10.1054/bjoc.1999.1014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fernebro E., Halvarsson B., Baldetorp B., et al. Predominance of CIN versus MSI in the development of rectal cancer at young age. BMC Cancer. 2002;2:25. doi: 10.1186/1471-2407-2-25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Fornasarig M., Viel A., Valentini M., et al. Microsatellite instability and MLH1 and MSH2 germline defects are related to clinicopathological features in sporadic colorectal cancer. Oncol Rep. 2000;7:39–43. doi: 10.3892/or.7.1.39. [DOI] [PubMed] [Google Scholar]
- 26.Giraldez M.D., Balaguer F., Bujanda L., et al. MSH6 and MUTYH deficiency is a frequent event in early-onset colorectal cancer. Clin Cancer Res. 2010;16:5402–5413. doi: 10.1158/1078-0432.CCR-10-1491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Goel A., Nagasaka T., Spiegel J., et al. Low frequency of Lynch syndrome among young patients with non-familial colorectal cancer. Clin Gastroenterol Hepatol. 2010;8:966–971. doi: 10.1016/j.cgh.2010.06.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kessels K., Fidder H.H., de Groot N.L., et al. Adherence to microsatellite instability testing in young-onset colorectal cancer patients. Dis Colon Rectum. 2013;56:825–833. doi: 10.1097/DCR.0b013e31828b6617. [DOI] [PubMed] [Google Scholar]
- 29.Kim H.C., Kim C.N., Yu C.S., et al. Methylation of the hMLH1 and hMSH2 promoter in early-onset sporadic colorectal carcinomas with microsatellite instability. Int J Colorectal Dis. 2003;18:196–202. doi: 10.1007/s00384-002-0445-0. [DOI] [PubMed] [Google Scholar]
- 30.Kim T.J., Kim E.R., Hong S.N., et al. Long-term outcome and prognostic factors of sporadic colorectal cancer in young patients: a large institutional-based retrospective study. Medicine (Baltimore) 2016;95 doi: 10.1097/MD.0000000000003641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kirzin S., Marisa L., Guimbaud R., et al. Sporadic early-onset colorectal cancer is a specific sub-type of cancer: a morphological, molecular and genetics study. PLoS One. 2014;9 doi: 10.1371/journal.pone.0103159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Liang J., Kalady M.F., Church J. Young age of onset colorectal cancers. Int J Colorectal Dis. 2015;30:1653–1657. doi: 10.1007/s00384-015-2341-4. [DOI] [PubMed] [Google Scholar]
- 33.Magnani G., Furlan D., Sahnane N., et al. Molecular features and methylation status in early onset (</=40 years) colorectal cancer: a population based, case-control study. Gastroenterol Res Pract. 2015;2015:132190. doi: 10.1155/2015/132190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Farrington S.M., McKinley A.J., Carothers A.D., et al. Evidence for an age-related influence of microsatellite instability on colorectal cancer survival. Int J Cancer. 2002;98:844–850. doi: 10.1002/ijc.10264. [DOI] [PubMed] [Google Scholar]
- 35.Perea J., Alvaro E., Rodriguez Y., et al. Approach to early-onset colorectal cancer: clinicopathological, familial, molecular and immunohistochemical characteristics. World J Gastroenterol. 2010;16:3697–3703. doi: 10.3748/wjg.v16.i29.3697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Perea J., Rueda D., Canal A., et al. Age at onset should be a major criterion for subclassification of colorectal cancer. J Mol Diagn. 2014;16:116–126. doi: 10.1016/j.jmoldx.2013.07.010. [DOI] [PubMed] [Google Scholar]
- 37.Pilozzi E., Maresca C., Duranti E., et al. Left-sided early-onset vs late-onset colorectal carcinoma histologic, clinical, and molecular differences. Am J Clin Pathol. 2015;143:374–384. doi: 10.1309/AJCPNOC55IOLXFUD. [DOI] [PubMed] [Google Scholar]
- 38.Raman R., Kotapalli V., Adduri R., et al. Evidence for possible non-canonical pathway(s) driven early-onset colorectal cancer in India. Mol Carcinog. 2014;53:E181–E186. doi: 10.1002/mc.21976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Siddique S., Tariq K., Rafiq S., et al. Sporadic early onset colorectal cancer in Pakistan: a case- control analysis of microsatellite instability. Asian Pac J Cancer Prev. 2016;17:2587–2592. [PubMed] [Google Scholar]
- 40.Soliman A.S., Bondy M.L., El-Badawy S.A., et al. Contrasting molecular pathology of colorectal carcinoma in Egyptian and Western patients. Br J Cancer. 2001;85:1037–1046. doi: 10.1054/bjoc.2001.1838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Stigliano V., Sanchez-Mete L., Martayan A., et al. Early-onset colorectal cancer patients without family history are “at very low risk” for lynch syndrome. J Exp Clin Cancer Res. 2014;33:1. doi: 10.1186/1756-9966-33-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tanskanen T., Gylfe A.E., Katainen R., et al. Exome sequencing in diagnostic evaluation of colorectal cancer predisposition in young patients. Scand J Gastroenterol. 2013;48:672–678. doi: 10.3109/00365521.2013.783102. [DOI] [PubMed] [Google Scholar]
- 43.Watson R., Liu T.C., Ruzinova M.B. High frequency of KRAS mutation in early onset colorectal adenocarcinoma: implications for pathogenesis. Hum Pathol. 2016;56:163–170. doi: 10.1016/j.humpath.2016.06.010. [DOI] [PubMed] [Google Scholar]
- 44.Willauer A.N., Liu Y.S., Pereira A.A.L., et al. Clinical and molecular characterization of early-onset colorectal cancer. Cancer. 2019;125:2002–2010. doi: 10.1002/cncr.31994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Aitchison A., Hakkaart C., Day R.C., et al. APC mutations are not confined to hotspot regions in early-onset colorectal cancer. Cancers. 2020;12:3829. doi: 10.3390/cancers12123829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Atef N., Alieldin N., Sherif G., et al. Microsatellite instability and life style factors in sporadic colorectal cancer. Asian Pac J Cancer Prev. 2020;21:1471–1480. doi: 10.31557/APJCP.2020.21.5.1471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Benmoussa A., Badre W., Pedroni M., et al. Clinical and molecular characterization of colorectal cancer in young Moroccan patients. Turk J Gastroenterol. 2012;23:686–690. doi: 10.4318/tjg.2012.0474. [DOI] [PubMed] [Google Scholar]
- 48.Jiang D., Shu C., Lei C.F., et al. Early-onset colorectal cancer: a distinct entity with unique genetic features. Oncol Lett. 2020;20:33. doi: 10.3892/ol.2020.11894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Sacdalan D.L., Garcia R.L., Diwa M.H., et al. Clinicopathologic factors associated with mismatch repair status among Filipino patients with young-onset colorectal cancer. Cancer Manag Res. 2021;13:2105–2115. doi: 10.2147/CMAR.S286618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Suzuki O., Eguchi H., Chika N., et al. Prevalence and clinicopathologic/molecular characteristics of mismatch repair-deficient colorectal cancer in the under-50-year-old Japanese population. Surg Today. 2017;47:1135–1146. doi: 10.1007/s00595-017-1486-x. [DOI] [PubMed] [Google Scholar]
- 51.Bateman A.C. DNA mismatch repair proteins: scientific update and practical guide. J Clin Pathol. 2021;74:264–268. doi: 10.1136/jclinpath-2020-207281. [DOI] [PubMed] [Google Scholar]
- 52.Valle L. Recent discoveries in the genetics of familial colorectal cancer and polyposis. Clin Gastroenterol Hepatol. 2017;15:809–819. doi: 10.1016/j.cgh.2016.09.148. [DOI] [PubMed] [Google Scholar]
- 53.Carethers J.M., Stoffel E.M. Lynch syndrome and Lynch syndrome mimics: the growing complex landscape of hereditary colon cancer. World J Gastroenterol. 2015;21:9253–9261. doi: 10.3748/wjg.v21.i31.9253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Weisenberger D.J., Levine A.J., Long T.I., et al. Association of the colorectal CpG island methylator phenotype with molecular features, risk factors, and family history. Cancer Epidemiol Biomarkers Prev. 2015;24:512–519. doi: 10.1158/1055-9965.EPI-14-1161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Advani S.M., Advani P., DeSantis S.M., et al. Clinical, pathological, and molecular characteristics of CpG island methylator phenotype in colorectal cancer: a systematic review and meta-analysis. Transl Oncol. 2018;11:1188–1201. doi: 10.1016/j.tranon.2018.07.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Ballester V., Rashtak S., Boardman L. Clinical and molecular features of young-onset colorectal cancer. World J Gastroenterol. 2016;22:1736–1744. doi: 10.3748/wjg.v22.i5.1736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Van Cutsem E., Cervantes A., Nordlinger B., et al. Metastatic colorectal cancer: ESMO clinical practice guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2014;25 Suppl 3:iii1–iii9. doi: 10.1093/annonc/mdu260. [DOI] [PubMed] [Google Scholar]
- 58.Lieu C.H., Golemis E.A., Serebriiskii I.G., et al. Comprehensive genomic landscapes in early and later onset colorectal cancer. Clin Cancer Res. 2019;25:5852–5858. doi: 10.1158/1078-0432.CCR-19-0899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tran B., Kopetz S., Tie J., et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer. 2011;117:4623–4632. doi: 10.1002/cncr.26086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.NICE Encorafenib plus cetuximab for previously treated BRAF V600E mutation-positive metastatic colorectal cancer 2021. https://www.nice.org.uk/guidance/ta668 Available from:
- 61.Cercek A., Chatila W.K., Yaeger R., et al. A comprehensive comparison of early-onset and average-onset colorectal cancers. J Natl Cancer Inst. 2021;113:1683–1692. doi: 10.1093/jnci/djab124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Levin-Sparenberg E., Bylsma L.C., Lowe K., et al. A systematic literature review and meta-analysis describing the prevalence of KRAS, NRAS, and BRAF gene mutations in metastatic colorectal cancer. Gastroenterol Res. 2020;13:184–198. doi: 10.14740/gr1167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Jin J., Shi Y., Zhang S., et al. PIK3CA mutation and clinicopathological features of colorectal cancer: a systematic review and meta-analysis. Acta Oncol. 2020;59:66–74. doi: 10.1080/0284186X.2019.1664764. [DOI] [PubMed] [Google Scholar]
- 64.Mei Z.B., Duan C.Y., Li C.B., et al. Prognostic role of tumor PIK3CA mutation in colorectal cancer: a systematic review and meta-analysis. Ann Oncol. 2016;27:1836–1848. doi: 10.1093/annonc/mdw264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Cancer Genome Atlas N Comprehensive molecular characterization of human colon and rectal cancer. Nature. 2012;487:330–337. doi: 10.1038/nature11252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Kim J.E., Choi J., Sung C.O., et al. High prevalence of TP53 loss and whole-genome doubling in early-onset colorectal cancer. Exp Mol Med. 2021;53:446–456. doi: 10.1038/s12276-021-00583-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Ribic C.M., Sargent D.J., Moore M.J., et al. Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med. 2003;349:247–257. doi: 10.1056/NEJMoa022289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Sinicrope F.A., Foster N.R., Thibodeau S.N., et al. DNA mismatch repair status and colon cancer recurrence and survival in clinical trials of 5-fluorouracil-based adjuvant therapy. J Natl Cancer Inst. 2011;103:863–875. doi: 10.1093/jnci/djr153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Sargent D.J., Marsoni S., Monges G., et al. Defective mismatch repair as a predictive marker for lack of efficacy of fluorouracil-based adjuvant therapy in colon cancer. J Clin Oncol. 2010;28:3219–3226. doi: 10.1200/JCO.2009.27.1825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Rho Y.S., Gilabert M., Polom K., et al. Comparing clinical characteristics and outcomes of young-onset and late-onset colorectal cancer: an international collaborative study. Clin Colorectal Cancer. 2017;16:334–342. doi: 10.1016/j.clcc.2017.03.008. [DOI] [PubMed] [Google Scholar]
- 71.Migliavaca C.B., Stein C., Colpani V., et al. Quality assessment of prevalence studies: a systematic review. J Clin Epidemiol. 2020;127:59–68. doi: 10.1016/j.jclinepi.2020.06.039. [DOI] [PubMed] [Google Scholar]
- 72.Nyaga V.N., Arbyn M., Aerts M. Metaprop: a Stata command to perform meta-analysis of binomial data. Arch Public Health. 2014;72:39. doi: 10.1186/2049-3258-72-39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Baretti M., Le D.T. DNA mismatch repair in cancer. Pharmacol Ther. 2018;189:45–62. doi: 10.1016/j.pharmthera.2018.04.004. [DOI] [PubMed] [Google Scholar]
- 74.Koopman M., Kortman G.A.M., Mekenkamp L., et al. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br J Cancer. 2009;100:266–273. doi: 10.1038/sj.bjc.6604867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Chen D., Huang J.F., Liu K., et al. BRAFV600E mutation and its association with clinicopathological features of colorectal cancer: a systematic review and meta-analysis. PLoS One. 2014;9 doi: 10.1371/journal.pone.0090607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Magrin L., Fanale D., Brando C., et al. POLE, POLD1, and NTHL1: the last but not the least hereditary cancer-predisposing genes. Oncogene. 2021;40:5893–5901. doi: 10.1038/s41388-021-01984-2. [DOI] [PubMed] [Google Scholar]
- 77.Briggs S., Tomlinson I. Germline and somatic polymerase epsilon and delta mutations define a new class of hypermutated colorectal and endometrial cancers. J Pathol. 2013;230:148–153. doi: 10.1002/path.4185. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.