Abstract
In a bioeconomy that relies on synthetic DNA sequences, the ability to ensure their authenticity is critical. DNA watermarks can encode identifying data in short sequences and be combined with error correction and encryption protocols to ensure sequences are robust to errors and securely communicated. New digital signature techniques allow for public verification that a sequence has not been modified and can contain sufficient information for synthetic DNA to be self-documenting. In translating these techniques from bacteria to more complex genetically modified organisms (GMOs), special considerations must be made to allow for public verification of these products. We argue these approaches should be widely implemented to assert authorship, increase the traceability, and detect the unauthorized use of synthetic DNA.
Keywords: DNA cryptography, cyberbiosecurity, DNA watermark, digital signature, genetic engineering attribution
Attribution of synthetic DNA mitigates cyberbiosecurity risks
In the 45 years since recombinant human insulin was first derived from plasmids expressed in Escherichia coli, the ease and availability of DNA synthesis has grown significantly [1,2]. Products utilizing synthetic DNA sequences (i.e., biological drugs, gene and cell therapies, biofuels, and foods like the Impossible Burger) have become the foundation of the growing bioeconomy [3–9]. Yet, the cyber-physical nature of synthetic DNA has raised cyberbiosecurity concerns since the inception of recombinant DNA technology [5,9–13]. For example, stringent cyberbiosecurity protocols are needed to ensure DNA sequences encoding potentially dangerous products don’t pass the security checks of DNA synthesis companies [14]. One major outstanding challenge is the ability to easily and accurately confirm the integrity and original attribution of DNA sequences [15].
After synthesis, DNA sequences are often manipulated and/or shared between academic labs. Thus, over time, owners and recipients may lack exact reference sequences and/or knowledge of the sequence origins. In addition, proper attribution of DNA sequences is limited by complicated patent regulations. The benefits of improved traceability and attribution of DNA sequences range from better quality control and the protection of intellectual property (IP) rights to the mitigation of security risks stemming from misuse and improved microbial forensics.
Since DNA sequences are generally designed and documented electronically, attribution may be given by including the origin information in the documentation. However, this only allows for an indirect association between the (digital) description of a product and its actual (physical) realization [5]. Furthermore, in some cases, it can exacerbate security and IP risks, for example, if the full sequence of a pathogenic viral vector is freely available or if a company wants to keep proprietary sequence information confidential [5,7].
Recent research has moved beyond digital documentation to within-molecule documentation. DNA is a robust storage medium capable of long-term data storage: books, songs, even operating systems can be stored in DNA [16,17]. This review, intended for life science researchers interested in improving their knowledge of DNA cryptography, provides an overview of techniques for encoding encrypted information in DNA to ensure the traceability and attribution of synthetic DNA sequences. A discussion of early work on watermarking techniques, wherein short information-containing sequences are embedded into synthetic DNA sequences, provides a starting point for digital signature techniques, which provide both authenticity and integrity guarantees for an entire synthetic DNA sequence.
DNA watermarks can contain encrypted attribution information
One method for asserting ownership over digital media is watermarking, which involves embedding information in pictures, video, or audio. More recently, DNA watermarks have been developed, where a plaintext message (e.g., ownership information) is converted into a nucleotide sequence that can be extracted and verified, e.g., by PCR or sequencing. Jupiter and colleagues [18] recommended that all DNA watermarks be: (1) easily recoverable (stably integrated at a defined location), (2) biologically innocuous (have no function of their own nor alter the function of the original DNA), (3) error-tolerant (messages are recoverable in the event of unintentional errors), (4) highly available (all labs have many unique sequences at their disposal), and (5) resistant to attack (complex and secure). Thus, DNA watermarks must simultaneously be long enough to be unique, complex, and information-rich, but also short enough that they are innocuous and economical.
To achieve these goals, principles from both coding theory and cryptography are applied. Coding theory aims to ensure a message can be successfully encoded and transmitted over a noisy channel, and encompasses error detection and correction methods [19,20]. Cryptography, by contrast, aims to ensure secure, confidential communication only between specific parties (Box 1). Although watermarks do not inherently rely on encryption, here we primarily focus on instances where DNA watermarks are encrypted using a variety of cryptographic methods, from simple substitution ciphers to more complex public key cryptosystems, such as Rivest-Shamir-Adleman (RSA) encryption (see Glossary).
Box 1. Fundamentals of cryptography.
In information security, Confidentiality, Integrity, and Availability make up the CIA triad that forms the basis of data protection [77]. Confidentiality ensures that data is accessed only by authorized entities (the goal of cryptography). Integrity allows recipients to verify that data has not been tampered with during transmission (error tolerance, a feature of coding theory).
Historically, encryption schemes were developed to securely transfer secret military messages. Ciphers are the earliest encryption schemes where a message is transformed, or encrypted, to conceal its meaning. In cryptography, the original human-readable message is called plaintext, and the concealed message is called ciphertext. Ciphers are reversible: encryption produces ciphertext from plaintext, while decryption reveals the original message from the ciphertext. For example, the popular Caesar cipher encrypts a message by shifting each letter in the original message by an index, concealing the message. If the index is two and the message is “dna”, ‘d’ is shifted by two and becomes ‘f’, ‘n’ becomes ‘p’, and ‘a’ becomes ‘c’. The plaintext message ‘dna’ is thus transformed to the ciphertext ‘fpc’.
Encryption schemes use one or more keys and a message as parameters. The Caesar Cipher is a symmetric encryption scheme, wherein the same secret key (here, the index value of two) is used to decrypt the ciphertext in a simple reversal of the encryption process. Anyone who gains knowledge of the key can decrypt messages encrypted with the same key. Thus, it is difficult to share the key without compromising security.
To resolve this, asymmetric encryption, or public-key encryption, was developed using a pair of distinct keys that are mathematically linked: a public key and a private key. The public key is freely shared with anyone and is used for encrypting data, while the private key is kept secret and is used for decryption. When someone wants to send an encrypted message to the owner of the public key, they use that key to encrypt the data, and only the owner, who possesses the private key, can decrypt the original message, ensuring secure and confidential communication.
As messages became digitized for use in computers, the alphabet needed numerical representations, thus the American Standard Code for Information Interchange (ASCII) was introduced. ASCII maps every character and punctuation mark in the Latin alphabet to a number represented in binary code. The process of converting messages to ASCII and back is a type of encoding and decoding. With information now stored as numbers, encryption techniques shifted from human-friendly methods like the Caesar cipher to complex mathematical procedures that are computationally infeasible to break. There are a few mathematical concepts that are integral to cryptography, like the modulo and XOR operation. The modulo operation (mod, %) calculates the remainder when one integer is divided by another (example: 5%3 = 2). The XOR (exclusive or) operation is a binary logical operation that returns true (1) if an odd number of its inputs are true, and false (0) otherwise.
In a landmark study, Clelland and colleagues developed a substitution cipher to encode alphanumeric messages in DNA (Figure 1A). Each character is encoded by a unique codon (e.g., “M” is encoded by the nucleotides “TTC”), and the entire message is flanked by PCR primer sequences at either end. After hiding the secret message in a pool of DNA fragments, the secret message can be successfully retrieved only by a recipient who has been given (1) the PCR primer sequences, and (2) the sequence-to-character encryption key [21]. Wong and colleagues [22] expanded this system by transforming short (<100 base pairs [bp]) messages into bacteria to be replicated indefinitely. Importantly, the insert sequence is flanked by primer sequences which do not exist in the host genome, making it easily detectable. One must also note the landmark study by Gibson and colleagues [6] wherein they designed a synthetic Mycoplasma mycoides genome and differentiated it from the natural genome by embedding four unique watermark sequences. Although the watermarks weren’t designed to contain messages, the work provided a proof of principle for producing and tagging cells based on computer-designed genome sequences.
Figure 1. Watermarking techniques allow information storage in DNA.

(A, B, C) Substitution ciphers (yellow) encode messages in DNA by assigning each character to a short DNA sequence. Throughout, every other character of a message and its respective DNA/binary sequences are bolded for visibility. (A) DNA watermarks can be transformed into bacteria [21,22], (B) made more economical by decreasing the number of bases per character [23] and/or (C) more secure by splitting the message across multiple fragments [24]. (D, E, F) Methods using binary code (green) to encode messages in DNA add a layer of protection against interceptors. (D) DNA binary strings composed of START, STOP, 0-bit and 1-bit sequences can be visualized on a gel. The message X is mixed with dummy fragments Y1 and Y2 (collectively Y) and the message is decrypted by subtracting the image of the dummy strands from the mixed image (X+Y). Each method relies on the interpreter knowing the key primer sequences (half arrows) and/or the 0- and 1-bit primer sequences (D), which produce fragments from START to each 0 or 1, which are read from the bottom up [25]. (E) Binary messages can be encrypted with single-use keys. An “exclusive OR” calculation is performed wherein different bits produce 0 and matching bits produce 1 (bolded) [26]. (F) Characters encoded by 8-bit ASCII values can be encrypted with a secret key, converted into 6-bit binary sequences and embedded only into synonymous codons (black) wherein 0 and 1 represent how common the codon is [27]. (G) Discrete wavelet transform (DWT) coefficients can be calculated to find optimal synonymous codon subsequences (blue) [29]. Created with BioRender.com.
Once these early works showed that (hidden) information could be embedded into DNA and propagated in biological systems, methods that were more economical and/or optimized for use in living systems became desired. For example, Smith and colleagues adapted a Huffman code for use in DNA, wherein more frequently used characters are encoded by fewer nucleotides to shorten the message length (e.g., “E” is encoded by T, while “Z” is encoded by CCCTG), however, their method did not consider whether the message is biologically innocuous (Figure 1B) [23]. More recently, Zakeri and colleagues developed the iKey-64 system to map characters to codons while preventing homopolymeric stretches of more than 4 nucleotides (e.g., GGGGG): more frequently used characters like “E” are represented by 3 different nucleotides (e.g., TCA), while less common characters like “K” may have 2 nucleotides repeated (e.g., TTC) [24]. In addition, the message is split across multiple DNA fragments, which can be engineered to contain decoy messages so the true message can only be interpreted when a common primer is used to sequence a particular combination of fragments (Figure 1C).
Leier and colleagues [25] were the first to develop encrypted DNA watermarks using a binary (0 and 1) system (Figure 1D). Short (26 bp) oligonucleotides were engineered to represent either start, stop, 0-bit, or 1-bit, and upon ligation of these DNA fragments, the resulting sequence encoded a string of numbers. After being mixed with dummy fragments – DNA from other species or fragments encoding other messages – the binary message can be decrypted by a user who knows (1) the secret key forward primer sequence and (2) the reverse primer sequences corresponding to the 0-bit and 1-bit sequences. The PCR products from both reactions are visualized on the gel to reconstruct the order of the bits. The message can be further encrypted by utilizing the dummy fragments themselves as the key: the gel image obtained from the dummy pool is subtracted from the gel image obtained from the encrypted pool (dummy pool + message strand) to reveal the message. Although this system is both secure and simple, the technical aspect of decryption is laborious and the long length of each message makes this technique intractable on a large scale. Gehani and colleagues [26] describe a system for encrypting binary messages in DNA watermarks utilizing a one-time pad (Figure 1E). Briefly, an “exclusive or” (XOR or ⊕) calculation is performed using a single-use key which is the same length as the plaintext. A XOR calculation is performed on each pair of bits in the plaintext message and key: a pair of two different bits (0 and 1) result in 0, while two of the same bit result in 1 (Figure 1E). If the key is used only once, this method is more secure than substitution-based methods. However, the key length scales with the message length, therefore this method is not practical for encoding long messages.
Utilizing binary code as a conduit between a message and a DNA sequence presents a barrier to potential interceptors trying to read a DNA watermark. Embedding a watermark into a coding region of DNA may provide another level of concealment, however it is more difficult to achieve than with DNA fragments or non-coding regions, since the underlying amino acid sequence cannot change. Liss and colleagues [27] encoded innocuous binary messages by only modifying amino acids that have synonymous codons: the first, third, and fifth common synonymous codons represent 1 (odds), whereas the second, fourth, and sixth most common ones represent 0 (evens). Therefore, if the interpreter knows the optimized codon usage table for a particular species, they can determine the binary message, thus encrypting the message with a Vigenère cipher – a polyalphabetic substitution cipher – is critical. In brief, the American Standard Code for Information Interchange (ASCII) values of the plaintext message are added to the ASCII values of the secret key (Figure 1F). The encrypted ciphertext is generated through a few mathematical operations and made smaller by converting from typical 8-bit ASCII characters to a 6-bit binary code (Figure 1F). To decode the message, the process is followed in reverse, ending with the ASCII values of the key being subtracted from the watermark ciphertext. Since the receiver must know this secret key to interpret the watermark, decryption is more complex (secure) than with a simple substitution cipher.
Later, Haughton and Balado [28] developed BioCode, which facilitates watermarking in both non-coding and coding DNA regions using a binary to codon translation table. It ensures that no start codons are created by watermarks in non-coding regions, while in coding regions, it ensures that (1) the primary structure (translated protein) is conserved and (2) that the codon bias (usage) for a particular species is maintained. More recently, Lee [29] developed a discrete wavelet transform (DWT)-based method to optimize watermarks designed using the codon mapping table established by Cristea [30], wherein each codon is assigned to a number between 0 and 63 (Figure 1G). The codon sequence is separated into subsequences, then the DWT coefficients for each subsequence producing synonymous codons are calculated and used to choose the optimal subsequence for embedding the watermark. By choosing the optimal codon sequence, this approach was shown to be more robust to point mutations than BioCode [28] and the method by Liss [27], as well as DNA-Crypt [31] whose own merits are discussed below.
Error detection and correction in DNA watermarks
Innate to coding theory is the ability to successfully transmit a signal in the event of noise or errors. Given the inherent propensity of DNA to mutate, it is critical that DNA watermarking methods implement error detection, or better yet, correction techniques to ensure the attribution of DNA sequences in the long-term. Special attention has been paid to error correction within short (>20 bp) barcodes for use in sequencing experiments (Box 2), but methods for longer DNA watermarks must take alternative approaches.
Box 2. Error correction in sequencing barcodes.
Short (2–6 bp) DNA barcodes for use in sequencing experiments were first developed in the late 1990s, such that different complementary DNA (cDNA) libraries could be tagged with and identified by unique, randomly-generated barcodes [78]. Qiu and colleagues then developed 6 bp error correcting barcodes to tag individual cDNA molecules within multiplexed libraries [79].
This approach was based on the edit distance between two words (in this case, DNA sequences), which is the smallest number of changes needed to transform one string of characters into another (e.g., “GCG” and “GCA” have an edit distance of 1) [79,80]. Edit distance usually refers to Levenshtein distance, which is the most commonly used since it can correct not only substitutions, but insertions and deletions as well (e.g., “GCG” and “GC” have a Levenshtein distance of 1) [80,81]. Ashlock and Orth developed the Salmon algorithm which produces an optimized list of code words (barcodes) based on the Levenshtein distance and programmable biological constraints (e.g., GC content between 40 and 60%) [80,82]. Alternatively, Buschmann and Bystrykh [83] developed the Sequence-Levenshtein distance for designing short barcodes: since the words in a DNA sequence (the barcode and the tagged sequence) are not separate, this method considers how the presence of the adjacent sequence will affect the edit distance of the barcode in the case of insertions and deletions.
However, since the edit distance is directly related to the length of the message, it is difficult to scale up these error correction methods to longer DNA messages; they are better suited for designing short (~6 bp) barcodes that correct only one or two errors. Even when edit distance is not used, a balance between error correction capability and barcode length must be met: Hawkins and colleagues produced a large list of 1 bp-correcting filled/truncated right end edit (FREE) barcodes that are between 3 and 16 bp long, and 2 bp-correcting barcodes that are between 5 and 17 bp long [84]. Nevertheless, these FREE barcodes can successfully correct insertions and deletions without knowing the length of the altered barcode (as would be common in a sequencing or ligation error), an advantage over the Levenshtein distance.
Early on, Smith and colleagues [23] implemented a comma code to detect errors in DNA watermarks: one nucleotide acts as a “comma” that is consistently repeated throughout the watermark (Figure 2A). In this schema, the predictable pattern makes it easy to detect errors, and point mutations will introduce a stop codon 83% of the time. However, it greatly limits the information capacity of the message, since only 3 bases can be used for encoding. Arita and colleagues [32] instead included a parity bit in the binary sequence of each character for error detection (Figure 2D). To insert their watermark, they mutated only the 3rd nucleotide in each codon of a coding sequence, such that wobble base pairing can occur and the amino acid sequence is maintained. The wild-type codon encodes 0, whereas a mutated codon encodes 1. Each group of 6 bits (codons) encodes a character: the first 5 bits uniquely represent a character and the sixth bit is a parity bit for error detection (e.g., “M” must be 011001 to confirm the sequence has not changed). Thus, the integrity of the sequence can be verified upon sequencing if the interpreter knows the wild-type sequence.
Figure 2. Error detection and correction techniques in DNA watermarks.

(A) A predictable arrangement of one nucleotide makes it easy to confirm watermark integrity [23]. (B) Discrepancies between a watermark sequence and an obtained sequence can be visualized on a DNA dot plot [33]. (D) The presence of a parity bit (bold) in the binary code of each character verifies sequence identity [32]. While error detection can be rather simple (A, B, D; yellow), error correction techniques (C, E, F; green) are more involved. (C) Four information-containing bits are interspersed with four parity bits (underlined) in the construction of 1-base error-correcting 8/4 Hamming codes. The integrity of a resulting byte (blue) can be confirmed through a series of calculations on individual bits (h0-h7) [31]. (E) Hamming codes can be combined with additional encryption methods, such as cyclic permutation, to increase the security of the message [37]. (F) Groups of six characters can be arranged in 2D space, converted to base-four binary (0–3) and a block sum check for error correction performed. The sum of each column and row (after modulo 4) is converted to a nucleotide and appended to the linear DNA sequence, which is embedded into single nucleotide polymorphisms (SNPs) in the genome. Decryption follows the encryption process in reverse, and a single error can be corrected based on the expected parity nucleotides for a certain row and column [38]. Created with BioRender.com.
Yamamoto and colleagues [33] developed a simple approach to retrieve watermarks even in the face of mutations. Message characters are first translated into short binary strings (e.g., “M” is 100), then the binary strings are converted into nucleotides (i.e., 0 encodes a purine (A, G), while 1 encodes a pyrimidine (C, T)). This system allows for more flexibility in the sequence that can be embedded within coding regions. Given the necessary primer sequences, the decoder can visually assess the integrity and degree of similarity between the obtained sequence and the watermark sequence on a DNA dot plot (Figure 2B). However, this method is labor-intensive, and can only be applied when the watermark sequence is known, not when embedding secret messages or in the case of wide public dissemination.
Several studies have implemented error correction techniques alongside cryptographic methods. The DNA-Crypt algorithm by Heider and colleagues [31,34] is based on a simple substitution cipher, where each base is represented by 2 binary digits, i.e., T: 00, G: 01, C: 10, A: 11, and the watermark is placed within synonymous codons as in [32]. Each byte includes 4 bits of information (2 bases) plus 4 parity bits so they can implement 8/4 Hamming codes to detect errors up to 2 bits and correct a single bit (Figure 2C). However, Hamming codes are only able to correct substitutions, not deletions or insertions (unlike the Levenshtein distance, Box 2). Nevertheless, there are several algorithms to encrypt the message before embedding it. There are symmetric cryptosystems, in which the sender and receiver utilize the same (private) key for encryption and decryption, including one-time pad, advance encryption standard (AES), and Blowfish [35,36]. There are also asymmetric (public-key) cryptosystems, such as RSA, wherein the sender and receiver utilize different secret keys for encryption and decryption, increasing the security of the message [35,36].
A symmetric hybrid DNA encryption (HyDEn) approach by Tulpan and colleagues [37] also utilizes Hamming codes for error correction, but with an additional cyclic permutation to encrypt the message (Figure 2E). In brief, ASCII characters are randomly assigned to 8 bp binary DNA codewords, established based on the Hamming distance (e.g., “M” is “AACGGTGA”). After the message is converted to a DNA sequence, the message is encrypted by being permuted cyclically a certain number of positions based on a private key shared between the signer and recipient. This approach is less susceptible to a brute-force attack than simpler methods, such as a substitution approach.
A method by Na [38] encodes watermarks within single nucleotide polymorphism (SNP) hotspots, defined as more than 35 SNPs in a 1 kilobase region (Figure 2F). This approach is fairly secure against unwanted third parties, because there is no obvious insert in the genome and SNPs are naturally polymorphic. Following use of a character-to-codon encryption table (e.g., “M” is CAA), the codons are converted to a 4-bit binary string using a substitution system (A=0, T=1, G=2, C=3). A block sum check algorithm is then used for error correction. In brief, the resulting binary sequence is first arranged in 2D space. The values in each row and column are summed, divided by 4, and the remainder added to the end of the respective row or column (Figure 2F). The resultant parity values are converted back into nucleotides and inserted into the sequence. Upon decryption, the receiver can easily detect that the sequence has mutated if the obtained sequence doesn’t produce the same parity nucleotides. Furthermore, if the obtained parity nucleotide is different, then one original nucleotide can be easily deduced and corrected based on the difference in the remainder. Of course, with the advent of the CRISPR-Cas9 system, the flexibility in where watermarks are inserted has grown [39].
From watermarks to digital signatures
Even when the messages within DNA watermarks are encrypted, there are inherent security risks in the use of watermarks [12,40]. Ultimately, a watermark is independent of the DNA vector in which it is embedded. Once DNA watermark sequences are identified, they can be easily deleted, edited, or engineered into new DNA molecules to obscure their contents or origins and potentially encode unwanted or harmful products [12]. Furthermore, the watermarking techniques that are based on substitution ciphers [21–24,26,28–30,32,33] are particularly prone to interception and forgery. The idiosyncrasies of the language of the encrypted message can make it easier to crack the message [37,41]. The Vigenère cipher used by [27] is slightly more secure, but the Kasiski method to guess the secret key length and decrypt the message was first published in 1863 [42,43]. In addition, symmetric encryption, in which both the sender and receiver share a secret key (e.g., primer sequence or encryption table) [21,22,25–27,31,33,37], is not practical in all cases. Firstly, the watermark verification process is only possible within a pre-selected list of users who were given the key, and thus cannot be used in cases of further dissemination [40]. To share it with a new verifier, a new key must be generated and tracked. Secondly, an interceptor who gains access to the key can easily distort the encrypted message or generate new watermarked sequences. Furthermore, while the generation of many unique secret keys is trivial in the digital space, in the DNA domain, each unique watermark must be synthesized and shipped, imposing significant financial and time constraints on the process.
Digital signatures, developed to mimic handwritten signatures, are widely used to validate the overall authenticity (i.e., identity, integrity and origin) of digital documents, such as emails or financial transactions. They typically employ public-key (asymmetric) cryptographic methods, such as RSA, ensuring signed items can be publicly and widely verified [12,44]. As opposed to watermarks, only one public key is needed to generate an encrypted signature that can be verified by anyone. The first notion of a digital signature was described by [45] and over time, signatures have become equipped with stronger security and unforgeability guarantees [46]. Consequently, researchers have adapted digital signatures for synthetic DNA. In essence, the sender uses a private key to generate a unique signature for the message, which is then attached to the message itself. The recipient then uses the sender’s public key to verify the signature. If the signature is valid, it confirms that the message hasn’t been tampered with (integrity) and was sent by the claimed sender (authenticity). Digital signatures thus enforce not only the CIA triad, but also authenticate the data’s origin and ensure non-repudiation, wherein the validity of the signature cannot be denied.
One approach, first described by Kar and colleagues [7,47] and experimentally validated by Gallegos and colleagues [48], uses identity-based signatures [49] to verify plasmid sequences (Figure 3). With identity-based signatures, users’ secret signing keys are generated from their identifying information, such as an ORCID [50] or ID number, rather than being randomly generated. The public key is the identity (i.e., ORCID) itself. Anyone can derive their secret key from this public key using a trusted third party (TTP) server. Like with other public key systems, if the authority is compromised, numerous user signatures can be easily forged. However, knowing the expected number of genuine keys can appease this issue.
Figure 3. Digital signature techniques allow DNA sequences to be publicly verified.
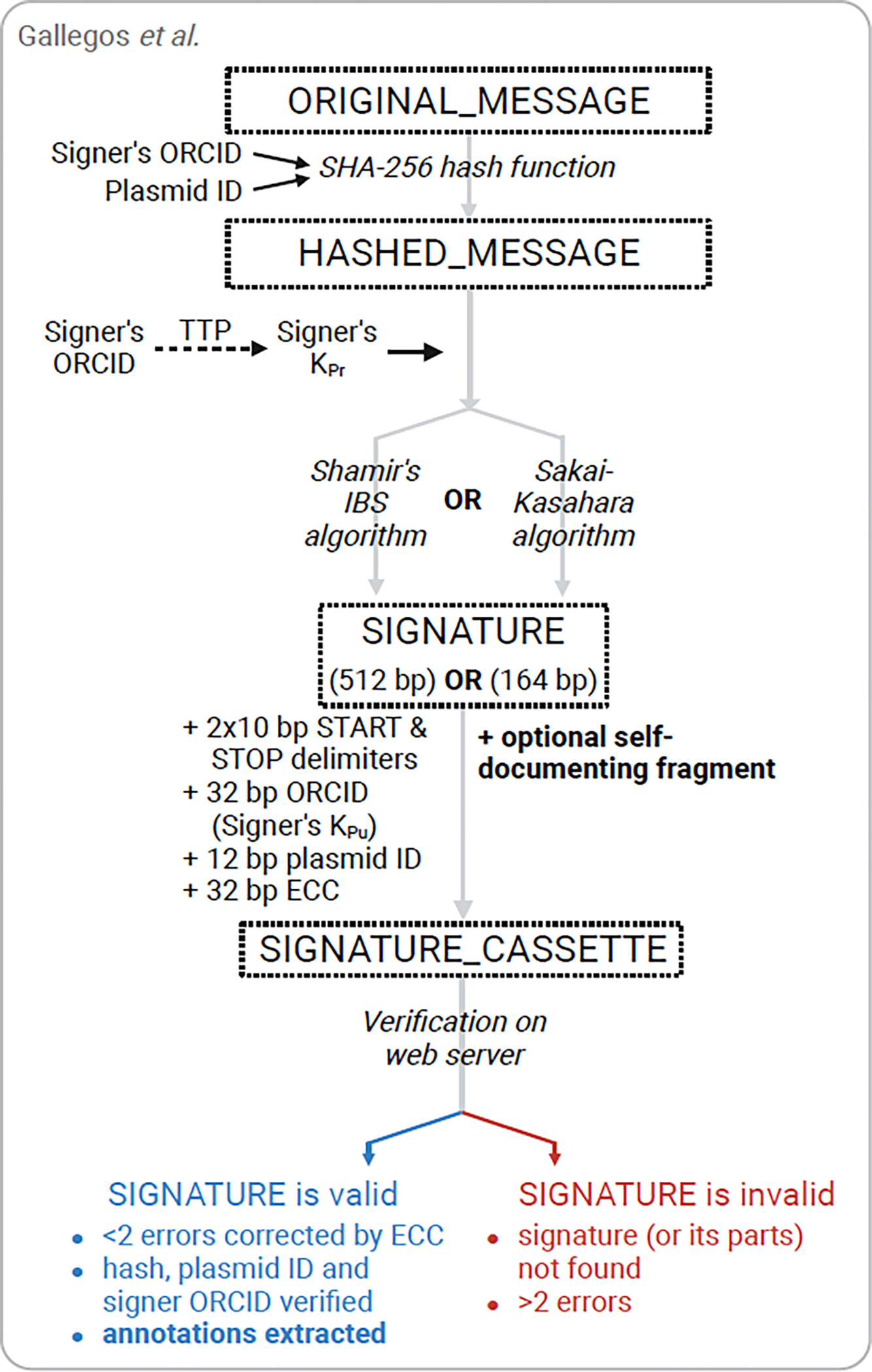
Signature techniques aim to securely encode more information into DNA than watermarks and are based around public-key cryptography methods that allow for wide, public verification of DNA sequences. An identity-based signature method developed for plasmids compresses the original message (the sequence itself) through one of two algorithms (resulting in two different sized signatures) which utilizes the signer’s private key, which is derived from their ORCID and issued by a trusted third party, to encrypt the hashed message. Additional information (e.g. plasmid ID, an error correction code (ECC), or a self-documenting fragment encoding a Genbank file with gene annotations) is added to the signature cassette. The signature and plasmid itself are verified through a simple process on a publicly available web server [48]. Created with BioRender.com.
The group initially used Shamir’s identity-based signature scheme, which is based on RSA [51], to generate a 512 bp signature representing the entire document (plasmid). This means that the accuracy of not just the signature, but the entire plasmid can be confirmed. The signature is produced from the message (a concatenation of the DNA sequence, plasmid ID and signer’s ORCID) using a SHA-256 hash function. Importantly, the signature was experimentally validated to not affect biological function (e.g. colony growth, antibiotic resistance, lacZ expression) [48]. The group then implemented the Sakai-Kasahara signature scheme [52] which generates a shorter 164 bp signature for any given input [48]. Both signature methods include a Reed-Solomon error-correction code (ECC) [53] in the signature cassette: it is 32 bp long which facilitates the correction of 2 errors. However, the ECC can be made longer to correct more errors.
With this method, users can generate a unique digital signature cassette to verify a plasmid and its signer, however the group also developed a method for inserting a fragment containing all the sequence annotations for the entire plasmid (Figure 3) [48]. Although increasing the number of annotations quickly makes the fragment quite large, a fragment containing minimal sequence annotations and a bibliographic reference to the source of the plasmid was successfully engineered into a plasmid using only a few thousand base pairs. This size could likely be decreased further with improved compression methods. Nevertheless, this approach allows a plasmid to be completely self-documenting: upon sequence assembly, the resulting FASTA file can be verified by the software and the resulting Genbank file with the annotations can be obtained [48]. If the signed plasmid is later edited beyond the 2 errors corrected by the ECC, the signature is thus invalidated. Because the software is openly available for users to sign and verify plasmids, the process of obtaining all the necessary documentation to validate the origin, contents, and terms of use of the plasmid is quick and easy after sequencing assembly is complete.
A simpler approach to self-documentation has been implemented in a novel strain version control system, CellRepo, which uses a DNA barcoding system to generate digital twins between DNA sequences in cell strains and their digital documentation [54,55]. Rather than the barcode itself containing the identifying information, it points to the digital documentation hosted online where the information such as the sequence and origin can be confirmed. The inserted barcode is comprised of a sequencing primer binding site, a 96 bp randomly generated universally unique identifier (containing the reference to the web documentation), a 9 bp synchronization sequence for aligning sequencing reads, and an 18 bp checksum sequence. Together, these components allow for error correction of the barcode, so it can be identified even with truncated or incorrect reads. Researchers can capture changes made to a cell strain in a new version, or “commit”, which prompts the insertion of a new barcode sequence into the strain. However, if the web documentation is not accessible for any reason, the barcode’s usefulness is lost, as no other identifying information is directly embedded into the DNA (such as the signer’s ID as in [48]).
Mueller [12,40] describes the considerations for implementing digital signature techniques for securing entire GMOs. One important feature is a zero-knowledge (ZK) proof: (1) a confirmer can convince a verifier that they have the secret (a valid signature) without revealing it, and (2) an interceptor cannot distinguish between a correct and incorrect secret nor learn the secret [56,57]. This could be achieved through designated confirmer signatures, first introduced by Chaum in [58], which are verified by engaging in a usually interactive protocol with a TTP. Another important feature is invisibility: one cannot tell whether a signature is valid or not just by looking at it [12,59]. In the case of GMOs, this can be accomplished by designing two sets of signatures – valid and dummy signatures – that are embedded into different clones [12,40]. The hashes of all signatures can then be made publicly verifiable without revealing which are valid. This also allows illegitimate GMOs to be identified based on not having the full collection of valid signatures or the presence of modifications in dummy clones [12].
To address the specific challenges in signing GMOs, different cryptographic primitives (e.g., hashes, digital signatures) can be combined like building blocks [12,40]. For example, an asymmetric ElGamal algorithm [60,61] can be applied [40]. In brief, the sender’s private key and the confirmer’s public key are used to encrypt a message, which is decrypted using the confirmer’s private key. If the verification process is successful, both the confirmer and TTP are assured that the GMO is authentic, even though the exact signature is not known. In case of a dispute, the TTP can confirm the signature by sequencing. Therefore, it is not the verification of the signature itself that confirms the authenticity, but rather the outcome of the verification process (pass/fail). In addition, the actual signature is never disclosed, since it can be hashed and hidden in the genome by matching the codon bias of the organism [12,40]. However, the choice of primitives depends on the application. For example, the ElGamal algorithm produces an encrypted message twice the length of the message which will impose limitations on the signed message. and the interactive protocol required by designated confirmer signatures may not always be feasible. Error detection and correction techniques can also be integrated [40,62]. Although all new schemes need to be experimentally validated, the prospect of implementing ZK proofs and invisibility in signature schemes that can verify increasingly longer sequences is critical for securing GMOs. Despite the daunting logistics of implementing such signatures in GMOs at a large scale, these efforts are necessary to protect consumers and designers of products utilizing synthetic DNA sequences.
Concluding Remarks
DNA watermarks and signatures allow information to be stably embedded in and retrieved from DNA sequences with varying levels of security and reliability (Figure 4, Key Figure). In general, they rely on the relative simplicity of isolating short sequences or plasmids, up to thousands of base pairs long, from cells to confirm specific DNA sequences. In many cases, such verification is sufficient to fully authenticate the product of interest: it is typically more appropriate to verify the gene or plasmid of interest than an entire bacterial genome with base-level precision. Not only it is more time- and cost-efficient, but it is still technically difficult to sequence entire genomes with current sequencing technologies, due to repetitive or GC-rich regions that inhibit sequencing, systemic errors introduced by PCR or sequencing, and errors introduced during the assembly of sequencing reads [63,64] (see Outstanding Questions). In addition, de novo assembly efforts are inherently more difficult than reference-based assembly, in the event a signature sequence is unknown before sequencing [65–67].
Figure 4, Key Figure. Watermark and digital signature methods vary in terms of security and reliability.

Security includes features like the encryption and secrecy of the message, whereas reliability represents the ease with which the message can be retrieved (i.e., mutation resistance, error correction, availability of tools). Watermarks using substitution ciphers are generally insecure due to their simple nature [21–24,32,33], while more complex systems like binary code [25–28,31,37,38,54] or discrete wavelet transform [29] increase the security of the message. Digital signatures are more secure than watermarks due to the use of more complex encryption algorithms and public key cryptography but vary in terms of their reliability [48]. The presence of error correcting mechanisms is critical for long-term information storage in DNA.
Outstanding Questions.
How can DNA sequencing and assembly techniques be optimized to ensure robust retrieval of information encoded in DNA watermarks and signatures, given that the effectiveness of these methods ultimately depends on reliable data retrieval?
How can error correction methods for DNA signatures be improved, given the limitations of current techniques? In other words, how do we ensure error correction capabilities are appropriately scaled with the size of the DNA to correct the entire sequence, while also ensuring their size doesn’t interfere with genetic functionality?
To what extent can cryptographic schemes for DNA signatures be standardized in the face of different biological requirements and applications, e.g., DNA data storage, bacterial plasmids used in research labs, viral vector vaccines, GMO crops and animals?
How can the diverse documentation practices for genetic constructs, found in various repositories with unique structures and requirements, be standardized to facilitate the creation of formal data structures for programs generating embedded documentation? What specific information should these standards encompass: origin, purpose, gene annotations, experimental data?
The idea that signatures are limited to verifying finite DNA sequences is especially pertinent as one moves into genetically modified plants and animals, as changes in unsigned regions of the genome could be missed [12,63]. However, it is unclear whether such long messages as genomic sequences can be signed in a way that biologically innocuous and cost-effective. As sequences become longer, error detection and correction methods become even more crucial to the authentication of DNA messages. The implementation of HEDGES (Hash Encoded, Decoded by Greedy Exhaustive Search) error-correcting codes, which correct substitutions, insertions and deletions, into current DNA signature schemes could prove fruitful [76].
Implementing signatures to verify GMOs is an important goal for the field, but further improvements are needed in both data encryption and sequencing methods to meet their specific challenges [12,40]. While data storage and compression in DNA encoding continue to improve [68], one feasible approach may be to have multiple signatures each verifying a particular DNA sequence. The ability to verify even short sequences with signatures would be an improvement over current GMO verification methods that primarily rely on experimental quantification and characterization of the target sequence or protein [69,70]. Moving forward, cryptographic schemes must be adapted for a wide variety of biological applications, such as plasmids used for research, viral vector vaccines, and GMO crops, to protect creators and consumers alike. Ideally, these schemes could be standardized to an extent that would facilitate wide usage and streamlined signing and verification protocols.
Implementing DNA signatures in non-GMO higher living organisms could also be of benefit. Currently, DNA barcodes composed of random synthetic sequences are used to track and identify organisms. For example, recent work has engineered DNA-loaded protein microcrystals that can persist throughout a mosquito’s lifespan [71]. Such DNA barcodes can be used to understand the circulation of vector-borne diseases or track endangered species, but these efforts would be strengthened by applying the cryptographic methods discussed here to embed meaningful identifying information within these barcodes.
In addition to watermarks and signatures, which aim to explicitly label DNA with identifying information, machine learning techniques are now emerging to predict the source lab or designer of non-labeled synthetic DNA sequences (Box 3). The premise is that the unique design choices made (consciously or not) by research groups (e.g., preferences for certain genetic parts, codon optimization, and the propagation of silent mutations) collectively leave a “methodological signature” which may be used to infer attribution of the DNA construct [15,72]. Since these approaches don’t rely on sequences being embedded in the DNA sequence, there are no concerns about modifying or losing the properties of the DNA as with watermarking or signature methods. However, for the same reason, these approaches do not provide authenticity guarantees. There is little to stop third parties from mimicking the methodological signature of a research group to conceal a construct’s origin [15]. Conversely, the presence of a feature that is commonly associated with one lab does not mean that lab created that construct. Although these models will likely never achieve 100% accuracy, they will be useful in providing a starting ground for investigations into the origins of DNA sequences [72,73].
Box 3. Machine learning for genetic engineering attribution.
Nielsen and colleagues [72] trained a convolutional neural network (CNN) on a dataset of over 30,000 plasmid sequences from over 800 labs. The depositing lab was predicted correctly 48% of the time and placed in the top-10 predictions 70% of the time. Importantly, accurate predictions could still be made even if, on average, 316 point mutations were made (about 10% of the sequence). Nevertheless, the effect of mutations greatly depends on the plasmid or genetic part: in one case, a 12 bp disruption in a region containing two restriction sites completely changed the predicted source lab.
Whereas CNNs are primarily used for pattern recognition in spatial data (i.e., usually images, sometimes text), recurrent neural networks (RNNs) are better suited for detecting and predicting patterns in sequential data (i.e., text, as in natural language processing models) [72,73]. Thus, the deteRNNt model established by Alley and colleagues achieved an accuracy of about 70% and the source lab was in the top-10 predictions nearly 85% of the time, using a larger dataset of over 80,000 plasmid sequences from over 3,500 labs [73].
A recent community-led approach led to even higher accuracy predictions [85]. Using a similar dataset as Alley and colleagues [73], 75 teams surpassed the top-10 accuracy of both of the existing models [72,73]. The top-ranked team achieved nearly 95% top-10 accuracy using a CNN-based approach, and the misclassification rate for the top-1 prediction was less than 20% (compared to 30% in [73] and 50% in [72]). In addition, one of the teams did not use neural networks, but rather a naïve Bayes classifier, resulting in a nearly 1000-fold increase in speed.
Machine learning models require high-quality, representative training datasets, are computationally expensive to run, and may have low explainability (how and why the model makes a decision). In response to these limitations, Wang and colleagues [86] developed an alternative alignment-based approach. Using a similar dataset as Nielsen and colleagues [72], they construct a pan-genome comprised of all the identified genomic regions. To identify a plasmid’s origin, its subsequences are aligned against the pan-genome and a lab score, favoring more unique sequences, is calculated. This approach led to an accuracy of 76% and the source lab was in the top-10 predictions 85% of the time. Although this method is still limited by the contents of the training dataset, it is easier to incorporate new sequences into the pan-genome than to retrain a machine learning model.
DNA signatures are a promising way of embedding identifying information in synthetic DNA sequences, such as their source, designer, terms of use, or even a compressed copy (hash) of the sequence itself, and then allowing public verification of the entire sequence. In line with efforts to make synthetic DNA design more transparent and standardized, the contents of DNA signatures must be standardized [73–76]. For example, the use of the ORCID as a standardized identification number is appealing due to its prevalence among scientific researchers. Though neural networks have shown some potential for identifying the originating lab of DNA sequences, they sorely lack the precision necessary for reliable authentication and ultimately fulfill a different need than DNA signatures. Rather than trying to computationally predict who designed a plasmid, researchers in the field must move towards signing synthetic DNA sequences themselves. As the use of DNA signatures becomes more widespread, the lack of a signature could be an indication in and of itself that the sequence is dubious.
Highlights.
The ability to quickly and accurately verify the authenticity of synthetic DNA sequences is critical for cyberbiosecurity efforts
Watermarks and digital signatures are the primary techniques for embedding attribution information into DNA sequences, and often implement error detection and correction to ensure robust communication
Digital signatures provide integrity, authenticity, and non-repudiation assurances to facilitate the secure public verification of DNA sequences, and can even make DNA sequences self-documenting
Features like invisibility and zero-knowledge proofs may allow DNA signatures to be used to combat counterfeit GMOs
Machine learning approaches are being implemented to predict the source of unsigned DNA sequences
Acknowledgements
J.P. and C.-T.B. are supported by the National Science Foundation (award #2123367) and the National Institutes of Health (R01GM147816).
Glossary
- Convolutional neural network (CNN):
A type of feed-forward neural network that leverages kernels or filters to process information locality. CNNs are well known for learning how to engineer features
- Designated confirmer signature:
This cryptographic technique is a type of digital signature that specifies a single confirmer or verifier when signed. The designated confirmer is then the only entity that can verify the signature, preventing others from doing so. This adds a level of privacy to digital signatures
- Discrete wavelet transform (DWT):
A mathematical method that decomposes a signal or data series into its different frequencies. An advancement of the Fourier Transform, this method provides information into frequency and temporal locality. Common applications are image compression, removing noise, and feature extraction
- ElGamal algorithm:
This public-key encryption algorithm leverages properties of modular exponentiation and the discrete logarithm problem to transmit data securely over insecure channels. It is used similarly to RSA
- Hamming code:
A type of binary error detecting and correcting code. Hamming codes can be used to detect one- and two-bit errors and correct only one-bit errors. The detection and correction techniques only work for substitutions, not for insertions or deletions
- Hash function:
A mathematical function extensively used in cryptography to calculate a fixed-size digest from input data. A key property is the computational infeasibility of reverse-engineering or deducing the original input from its digest, thereby guaranteeing data integrity and enhancing security
- Huffman coding:
A lossless data compression technique where a prefix Huffman code is inserted in the data to map common symbols to smaller symbols. This can be used in DNA by mapping frequently appearing characters in the data to fewer nucleotides
- Levenshtein distance:
A type of edit distance metric that allows for flexible edit operations including substitutions, insertions, and deletions. This flexibility makes it very applicable for genetic code
- Recurrent neural network (RNN):
A type of artificial neural network that maintains internal states as a form of memory making the model effective for sequential datasets
- Reed-Solomon code:
A type of binary erasure code that can detect and correct substitutions, insertions, and deletions
- Rivest-Shamir-Adleman (RSA):
A widely used cryptographic system that enables secure data transmission through asymmetric public-key encryption. The security of the system relies on the mathematical properties of large prime numbers. An alternative scheme to RSA would be the El-Gamal algorithm
- Sakai-Kasahara scheme:
This identity-based scheme encrypts a message for a given recipient with a known identity (like email or ORCID) so that it can only be decrypted by the recipient. This allows anyone to encrypt a message using only public information about the recipient
- Shamir’s identity-based signature scheme:
A digital signature that uses cryptographic techniques from RSA to secure a message and tie it to an identity. This scheme creates fixed sized signatures that do not scale well for use in DNA signatures
- Vigenère cipher:
This cipher encrypts an ASCII plaintext message using a changing Caesar cipher where each character has a different offset based on the key’s character. The offset for the Caesar cipher is calculated from the key’s value from the first value in the alphabet. The key’s size is arbitrary and can be repeated to encrypt a message of any size
- Zero-knowledge (ZK) proof:
A method that allows two parties, a verify and a prover, to assert the truth of some given information without revealing any of the information
Footnotes
Conflict of Interest
J. P. and S. P. have financial interests in GenoFAB, Inc. This company may benefit or be perceived as benefiting from this publication.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Hughes RA and Ellington AD (2017) Synthetic DNA Synthesis and Assembly: Putting the Synthetic in Synthetic Biology. Cold Spring Harbor Perspectives in Biology 9, a023812. 10.1101/cshperspect.a023812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Goeddel DV et al. (1979) Expression in Escherichia coli of chemically synthesized genes for human insulin. Proceedings of the National Academy of Sciences 76, 106–110. 10.1073/pnas.76.1.106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Voigt CA (2020) Synthetic biology 2020–2030: six commercially-available products that are changing our world. Nature Communications 11, 6379. 10.1038/s41467-020-20122-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Chiarabelli C et al. (2013) Chemical synthetic biology: a mini-review. Frontiers in microbiology 4, 285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Peccoud J et al. (2018) Cyberbiosecurity: from naive trust to risk awareness. Trends in biotechnology 36, 4–7 [DOI] [PubMed] [Google Scholar]
- 6.Gibson DG et al. (2010) Creation of a bacterial cell controlled by a chemically synthesized genome. science 329, 52–56 [DOI] [PubMed] [Google Scholar]
- 7.Kar DM et al. (2018). Digital signatures to ensure the authenticity and integrity of synthetic DNA Molecules. Proceedings of the New Security Paradigms Workshop. [Google Scholar]
- 8.Shankar S and Hoyt MA (2018). Expression constructs and methods of genetically engineering methylotrophic yeast. Google Patents [Google Scholar]
- 9.Walsh PF (2022) Securing the Bioeconomy: Exploring the Role of Cyberbiosecurity. In, pp. 335–355, Springer International Publishing [Google Scholar]
- 10.Murch RS et al. (2018) Cyberbiosecurity: an emerging new discipline to help safeguard the bioeconomy. Frontiers in bioengineering and biotechnology 6, 39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Mueller S (2021) Facing the 2020 pandemic: What does cyberbiosecurity want us to know to safeguard the future? Biosafety and health 3, 11–21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mueller S (2019) On DNA Signatures, Their Dual-Use Potential for GMO Counterfeiting, and a Cyber-Based Security Solution. Frontiers in bioengineering and biotechnology 7, 189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Berg P et al. (1974) Potential Biohazards of Recombinant DNA Molecules. Science 185, 303–303. doi: 10.1126/science.185.4148.303 [DOI] [PubMed] [Google Scholar]
- 14.Puzis R et al. (2020) Increased cyber-biosecurity for DNA synthesis. Nature Biotechnology 38, 1379–1381. 10.1038/s41587-020-00761-y [DOI] [PubMed] [Google Scholar]
- 15.Lewis G et al. (2020) The biosecurity benefits of genetic engineering attribution. Nature Communications 11. 10.1038/s41467-020-19149-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Church GM et al. (2012) Next-generation digital information storage in DNA. Science 337, 1628–1628 [DOI] [PubMed] [Google Scholar]
- 17.Erlich Y and Zielinski D (2017) DNA Fountain enables a robust and efficient storage architecture. Science 355, 950–954 [DOI] [PubMed] [Google Scholar]
- 18.Jupiter DC et al. (2010) DNA watermarking of infectious agents: Progress and prospects. PLoS Pathog 6, e1000950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Aydin N (2009) What is Coding Theory and What is Cryptography? Introduction to Coding Theory and Cryptography [Google Scholar]
- 20.Calderbank AR (1998) The art of signaling: fifty years of coding theory. IEEE Transactions on Information Theory 44, 2561–2595. 10.1109/18.720549 [DOI] [Google Scholar]
- 21.Clelland CT et al. (1999) Hiding messages in DNA microdots. Nature 399, 533–534. 10.1038/21092 [DOI] [PubMed] [Google Scholar]
- 22.Wong PC et al. (2003) Organic data memory using the DNA approach. Communications of the ACM 46, 95–98 [Google Scholar]
- 23.Smith GC et al. (2003) Some possible codes for encrypting data in DNA. Biotechnology letters 25, 1125–1130 [DOI] [PubMed] [Google Scholar]
- 24.Zakeri B et al. (2016) Multiplexed Sequence Encoding: A Framework for DNA Communication. PLOS ONE 11, e0152774. 10.1371/journal.pone.0152774 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Leier A et al. (2000) Cryptography with DNA binary strands. Biosystems 57, 13–22 [DOI] [PubMed] [Google Scholar]
- 26.Gehani A et al. (2004) DNA-based cryptography. Aspects of molecular computing: essays dedicated to tom head, on the occasion of his 70th birthday, 167–188 [Google Scholar]
- 27.Liss M et al. (2012) Embedding permanent watermarks in synthetic genes. PloS one 7, e42465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Haughton D and Balado F (2013) BioCode: Two biologically compatible Algorithms for embedding data in non-coding and coding regions of DNA. BMC bioinformatics 14, 121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lee S-H (2014) DWT based coding DNA watermarking for DNA copyright protection. Information Sciences 273, 263–286 [Google Scholar]
- 30.Cristea PD (2002) Conversion of nucleotides sequences into genomic signals. Journal of Cellular and Molecular Medicine 6, 279–303. 10.1111/j.1582-4934.2002.tb00196.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Heider D and Barnekow A (2007) DNA-based watermarks using the DNA-Crypt algorithm. BMC bioinformatics 8, 176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Arita M and Ohashi Y (2004) Secret signatures inside genomic DNA. Biotechnology progress 20, 1605–1607 [DOI] [PubMed] [Google Scholar]
- 33.Yamamoto N et al. (2014) A watermarking system for labeling genomic DNA. Plant Biotechnology, 14.0609 b [Google Scholar]
- 34.Heider D and Barnekow A (2008) DNA watermarks: A proof of concept. BMC molecular biology 9, 40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Patil P et al. (2016) A comprehensive evaluation of cryptographic algorithms: DES, 3DES, AES, RSA and Blowfish. Procedia Computer Science 78, 617–624 [Google Scholar]
- 36.Simmons GJ (1979) Symmetric and asymmetric encryption. ACM Computing Surveys (CSUR) 11, 305–330 [Google Scholar]
- 37.Tulpan D et al. (2013) HyDEn: A Hybrid Steganocryptographic Approach for Data Encryption Using Randomized Error-Correcting DNA Codes. BioMed Research International 2013, 1–11. 10.1155/2013/634832 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Na D (2020) DNA steganography: hiding undetectable secret messages within the single nucleotide polymorphisms of a genome and detecting mutation-induced errors. Microbial Cell Factories 19. 10.1186/s12934-020-01387-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Velázquez E et al. (2021) Targetron-Assisted Delivery of Exogenous DNA Sequences into Pseudomonas putida through CRISPR-Aided Counterselection. ACS Synthetic Biology 10, 2552–2565. 10.1021/acssynbio.1c00199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Mueller S et al. (2016) A covert authentication and security solution for GMOs. BMC bioinformatics 17, 1–8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Shiu H-J et al. (2010) Data hiding methods based upon DNA sequences. Information Sciences 180, 2196–2208 [Google Scholar]
- 42.Bhateja AK et al. (2015) Cryptanalysis of vigenere cipher using cuckoo search. Applied Soft Computing 26, 315–324 [Google Scholar]
- 43.Kasiski FW (1863) Die Geheimschriften und die Dechiffrir-Kunst: Mit bes. Berücks. der deutschen u. der franz. Sprache ES Mittler [Google Scholar]
- 44.Salomaa A (2013) Public-key cryptography Springer Science & Business Media [Google Scholar]
- 45.Diffie W and Hellman M (1976) New directions in cryptography. IEEE transactions on Information Theory 22, 644–654 [Google Scholar]
- 46.Lysyanskaya A (2002). Signature schemes and applications to cryptographic protocol design. Massachusetts Institute of Technology [Google Scholar]
- 47.Kar DM et al. (2020) Synthesizing DNA molecules with identity-based digital signatures to prevent malicious tampering and enabling source attribution. Journal of Computer Security, 1–31 [Google Scholar]
- 48.Gallegos JE et al. (2020) Securing the Exchange of Synthetic Genetic Constructs Using Digital Signatures. ACS Synthetic Biology 9, 2656–2664. 10.1021/acssynbio.0c00401 [DOI] [PubMed] [Google Scholar]
- 49.Baek J et al. (2004). A survey of identity-based cryptography. Proc. of Australian Unix Users Group Annual Conference. [Google Scholar]
- 50.Haak LL et al. (2012) ORCID: a system to uniquely identify researchers. Learned Publishing 25, 259–264. 10.1087/20120404 [DOI] [Google Scholar]
- 51.Shamir A (1984). Identity-based cryptosystems and signature schemes. Workshop on the theory and application of cryptographic techniques. Springer [Google Scholar]
- 52.Sakai R and Kasahara M (2003) ID based Cryptosystems with Pairing on Elliptic Curve. IACR Cryptology ePrint Archive 2003, 54 [Google Scholar]
- 53.Reed IS and Solomon G (1960) Polynomial codes over certain finite fields. Journal of the society for industrial and applied mathematics 8, 300–304 [Google Scholar]
- 54.Tellechea-Luzardo J et al. (2020) Linking Engineered Cells to Their Digital Twins: A Version Control System for Strain Engineering. ACS Synthetic Biology 9, 536–545. 10.1021/acssynbio.9b00400 [DOI] [PubMed] [Google Scholar]
- 55.Tellechea-Luzardo J et al. (2022) Versioning biological cells for trustworthy cell engineering. Nature Communications 13, 765. 10.1038/s41467-022-28350-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Goldreich O and Oren Y (1994) Definitions and properties of zero-knowledge proof systems. Journal of Cryptology 7, 1–32 [Google Scholar]
- 57.Goldwasser S et al. (1989) The knowledge complexity of interactive proof systems. SIAM Journal on computing 18, 186–208 [Google Scholar]
- 58.Chaum D (1994). Designated confirmer signatures. Workshop on the Theory and Application of of Cryptographic Techniques. Springer [Google Scholar]
- 59.Galbraith SD and Mao W (2003). Invisibility and anonymity of undeniable and confirmer signatures. Topics in Cryptology—CT-RSA 2003: The Cryptographers’ Track at the RSA Conference 2003 San Francisco, CA, USA, April 13–17, 2003 Proceedings. Springer [Google Scholar]
- 60.ElGamal T (1985) A public key cryptosystem and a signature scheme based on discrete logarithms. IEEE transactions on information theory 31, 469–472 [Google Scholar]
- 61.Rivest RL et al. (1978) A method for obtaining digital signatures and public-key cryptosystems. Communications of the ACM 21, 120–126 [Google Scholar]
- 62.Mueller S et al. (2015) Improving dependability and precision of data encoding in DNA. Eur. J. Exp. Biol 10, [Google Scholar]
- 63.Mueller S (2019) Are Market GM Plants an Unrecognized Platform for Bioterrorism and Biocrime? Frontiers in Bioengineering and Biotechnology 7. 10.3389/fbioe.2019.00121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hu T et al. (2021) Next-generation sequencing technologies: An overview. Human Immunology 82, 801–811. 10.1016/j.humimm.2021.02.012 [DOI] [PubMed] [Google Scholar]
- 65.Liao X et al. (2019) Current challenges and solutions of de novo assembly. Quantitative Biology 7, 90–109 [Google Scholar]
- 66.Prjibelski A et al. (2020) Using SPAdes De Novo Assembler. Current Protocols in Bioinformatics 70, e102. 10.1002/cpbi.102 [DOI] [PubMed] [Google Scholar]
- 67.Wick RR et al. (2017) Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS computational biology 13, e1005595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wang C et al. (2022) Mainstream encoding–decoding methods of DNA data storage. CCF Transactions on High Performance Computing 4, 23–33. 10.1007/s42514-022-00094-z [DOI] [Google Scholar]
- 69.Fu W et al. (2020) A universal analytical approach for screening and monitoring of authorized and unauthorized GMOs. LWT 125, 109176. 10.1016/j.lwt.2020.109176 [DOI] [Google Scholar]
- 70.Qian C et al. (2018) Recent advances in emerging DNA-based methods for genetically modified organisms (GMOs) rapid detection. TrAC Trends in Analytical Chemistry 109, 19–31. 10.1016/j.trac.2018.09.021 [DOI] [Google Scholar]
- 71.Stuart JD et al. (2022) Mosquito tagging using DNA-barcoded nanoporous protein microcrystals. PNAS Nexus 1. 10.1093/pnasnexus/pgac190 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Nielsen AA and Voigt CA (2018) Deep learning to predict the lab-of-origin of engineered DNA. Nature communications 9, 1–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Alley EC et al. (2020) A machine learning toolkit for genetic engineering attribution to facilitate biosecurity. Nature Communications 11. 10.1038/s41467-020-19612-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Martínez-García E et al. (2022) SEVA 4.0: an update of the Standard European Vector Architecture database for advanced analysis and programming of bacterial phenotypes. Nucleic Acids Research 51, D1558–D1567. 10.1093/nar/gkac1059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Czar MJ et al. (2009) Writing DNA with GenoCAD™. Nucleic Acids Research 37, W40–W47. 10.1093/nar/gkp361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Peccoud J et al. (2011) Essential information for synthetic DNA sequences. Nature biotechnology 29, 22–22 [DOI] [PubMed] [Google Scholar]
- 77.Samonas S and Coss D (2014) The CIA strikes back: Redefining confidentiality, integrity and availability in security. Journal of Information System Security 10, [Google Scholar]
- 78.Bonaldo Md.F. et al. (1996) Normalization and subtraction: two approaches to facilitate gene discovery. Genome research 6, 791–806 [DOI] [PubMed] [Google Scholar]
- 79.Qiu F et al. (2003) DNA sequence-based “bar codes” for tracking the origins of expressed sequence tags from a maize cDNA library constructed using multiple mRNA sources. Plant Physiology 133, 475–481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Ashlock D et al. (2012) On the synthesis of DNA error correcting codes. Biosystems 110, 1–8. 10.1016/j.biosystems.2012.06.005 [DOI] [PubMed] [Google Scholar]
- 81.Levenshtein VI (1966). Binary codes capable of correcting deletions, insertions, and reversals. Soviet physics doklady. Soviet Union [Google Scholar]
- 82.Orth J and Houghten S (2011). Optimizing the Salmon Algorithm for the construction of DNA error-correcting codes. IEEE [Google Scholar]
- 83.Buschmann T and Bystrykh LV (2013) Levenshtein error-correcting barcodes for multiplexed DNA sequencing. BMC bioinformatics 14, 1–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Hawkins JA et al. (2018) Indel-correcting DNA barcodes for high-throughput sequencing. Proceedings of the National Academy of Sciences 115, E6217–E6226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Crook OM et al. (2022) Analysis of the first genetic engineering attribution challenge. Nature communications 13, 7374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Wang Q et al. (2021) PlasmidHawk improves lab of origin prediction of engineered plasmids using sequence alignment. Nature Communications 12. 10.1038/s41467-021-21180-w [DOI] [PMC free article] [PubMed] [Google Scholar]


