Abstract
The RNA modification m6A controls many aspects of RNA function that impact biological processes, including viral infection. In this review, we highlight recent work that shapes our current understanding of the diverse mechanisms by which m6A can regulate viral infection by acting on viral or cellular mRNA molecules. We focus on emerging concepts and understanding, including how viral infection alters the localization and function of m6A-machinery proteins, how m6A regulates antiviral innate immunity, and the multiple roles of m6A in regulating specific viral infections. We also summarize the recent studies on m6A during SARS-CoV-2 infection, focusing on points of convergence and divergence. Ultimately, this review provides a snapshot of the latest research on m6A during viral infection.
Introduction
The RNA modification N6-methyladenosine (m6A) controls many aspects of RNA function that impact biological processes, including viral infection. The addition and removal of m6A to mRNA is controlled by cellular proteins, which include a large m6A-methyltransferase complex and demethylase enzymes [1,2]. The key m6A-methyltransferase proteins METTL3 and METTL14, predominantly localized in the nucleus, catalyze RNA methylation on nascent RNA molecules as part of a large complex of proteins that assist in RNA targeting for m6A deposition [2]. In particular, the METTL3/METTL14-interacting protein WTAP plays a key role in RNA targeting of this complex as it mediates the localization and RNA binding activities of METTL3 and METTL14 in the nucleus [3]. The function of m6A on mRNA is regulated by interactions with RNA binding proteins, referred to as “readers”, that recognize and bind to m6A-modified mRNA. These reader proteins include those that contain YTH domains that have high affinity toward m6A-modified mRNAs, as well as other RNA binding proteins that bind through indirect mechanisms [4]. Through interactions with m6A-reader proteins, m6A can influence many aspects of RNA metabolism, including decreasing RNA stability and increasing mRNA translation [5]. Importantly, m6A can also be removed by the cellular demethylase proteins FTO and ALKBH5 (reviewed in [4,5]). In the last ten years, m6A and its associated cellular machinery have been shown to regulate many different aspects of viral infection.
Here, we highlight newly published studies that reveal diverse mechanisms by which m6A regulates infection by acting on viral or cellular mRNA molecules. We describe studies that reveal that viral infection can lead to changes in the subcellular localization of m6A-machinery proteins. We will expand upon recent studies that reveal multiple roles for m6A on viral RNA in specific viral infections, including how m6A on viral RNA can regulate aspects of the immune response. We will highlight novel strategies by which m6A-reader proteins regulate or are regulated during infection. Finally, we will summarize recent studies on m6A during SARS-CoV-2 infection, focusing on points of convergence and divergence. Ultimately, this review will provide a snapshot of the latest research describing how m6A can regulate viral infection. We note that this regulation could occur from viral or cellular RNA methylation changes. For comprehensive reviews on how m6A regulates viral infection, we point the readers to several recently published reviews by us and others [6,7], including one focusing on a framework for mapping m6A on viral RNA [8].
Viral infection alters the subcellular localization of m6A-machinery proteins.
As viral infection often leads to changes in normal cellular biological processes, it is unsurprising that viral infection alters the localization of several members of the m6A machinery. For viruses with lifecycle stages that take place in the nucleus, the nuclear m6A-methyltransferase proteins are in the appropriate subcellular compartment for viral and cellular RNA methylation; however, two recent examples illustrate novel strategies used by viruses to alter the subcellular localization of these proteins. First, hepatitis B virus (HBV) uses a viral protein to redirect METTL3 within the nucleus to the viral minichromosome and new cellular transcripts for methylation [9]. In a second example, the viral ICP27 nuclear export protein from the DNA virus herpes simplex virus-1 (HSV-1) facilitates the export of METTL3 and METTL14 out of the nucleus to the cytoplasm [10]. This leads to a decrease in both viral and cellular m6A, ultimately impacting both viral and host gene expression. These two examples highlight two different strategies used by nuclear-replicating viruses to redirect the function of the m6A-machinery, in one case towards viral RNA and specific cellular transcripts and the other case away from cellular RNA.
Viruses with lifecycle stages that take place in the cytoplasm must recruit the m6A-machinery to the cytoplasm to target their RNA for methylation. Recent studies show that the negative-strand RNA viruses Ebola virus and vesicular stomatitis virus (VSV), whose RNA molecules contain m6A, induce partial relocalization of METTL3 to the cytoplasm [11,12]. For Ebola virus, the relocalization is mediated by the interaction of METTL3 with the viral VP30 RNA binding nucleoprotein [11]. Similarly, the plant virus, wheat yellow mosaic virus, which has a positive-strand RNA genome and also contains m6A, uses the viral Nib protein to induce a partial relocalization of the wheat METTL14 homolog TaMtb to cytoplasmic aggregates [13]. This results in methylation of the viral RNA to stabilize it and ultimately promotes infection. Two positive-strand RNA viruses in the Flaviviridae family, hepatitis C virus (HCV) and Zika virus (ZIKV), also change the localization of proteins required for m6A modification of the viral RNA. HCV infection leads to increased WTAP localization in the cytoplasm for targeting METTL3 to the viral RNA [14]. ZIKV infection alters the localization of S-adenosylhomocysteine hydrolase AHCY to viral replication organelles [15]. As AHCY converts S-adenosylhomocysteine (SAH) to adenosine and L-homocysteine, this enzyme ultimately increases the levels of these metabolites for the generation of S-adenosylmethionine, which is then used by METTL3 for m6A-modification of the viral RNA. For ZIKV, this increase in m6A on the viral RNA inhibits innate immune sensing of the viral RNA, allowing for more viral replication [15]. Several independent studies have also recently shown that SARS-CoV-2, a positive-strand RNA virus that replicates in the cytoplasm, also induces partial localization of m6A-machinery proteins to the cytoplasm for regulation of viral m6A levels [16–18]. Similarly, the seasonal human beta-coronavirus HCoV-OC43 induces a partial relocalization of METTL3 [19]. In summary, recent work reveals that viral infection can alter the localization of the m6A-machinery (Figure 1). These relocalization strategies include the use of viral proteins to target the m6A-machinery to or away from viral or cellular RNA to regulate infection. Interestingly, in HIV-1 the viral Gag protein recruits the m6A demethylase FTO to the viral RNA for demethylation to enhance RNA packaging into the virion [20]. Future studies will undoubtedly define more viral proteins that interact with members of the m6A-machinery, including the m6A demethylases, to regulate RNA methylation during infection.
Figure 1. m6A-methyltransferase complex localization and function can be regulated by viral infection.
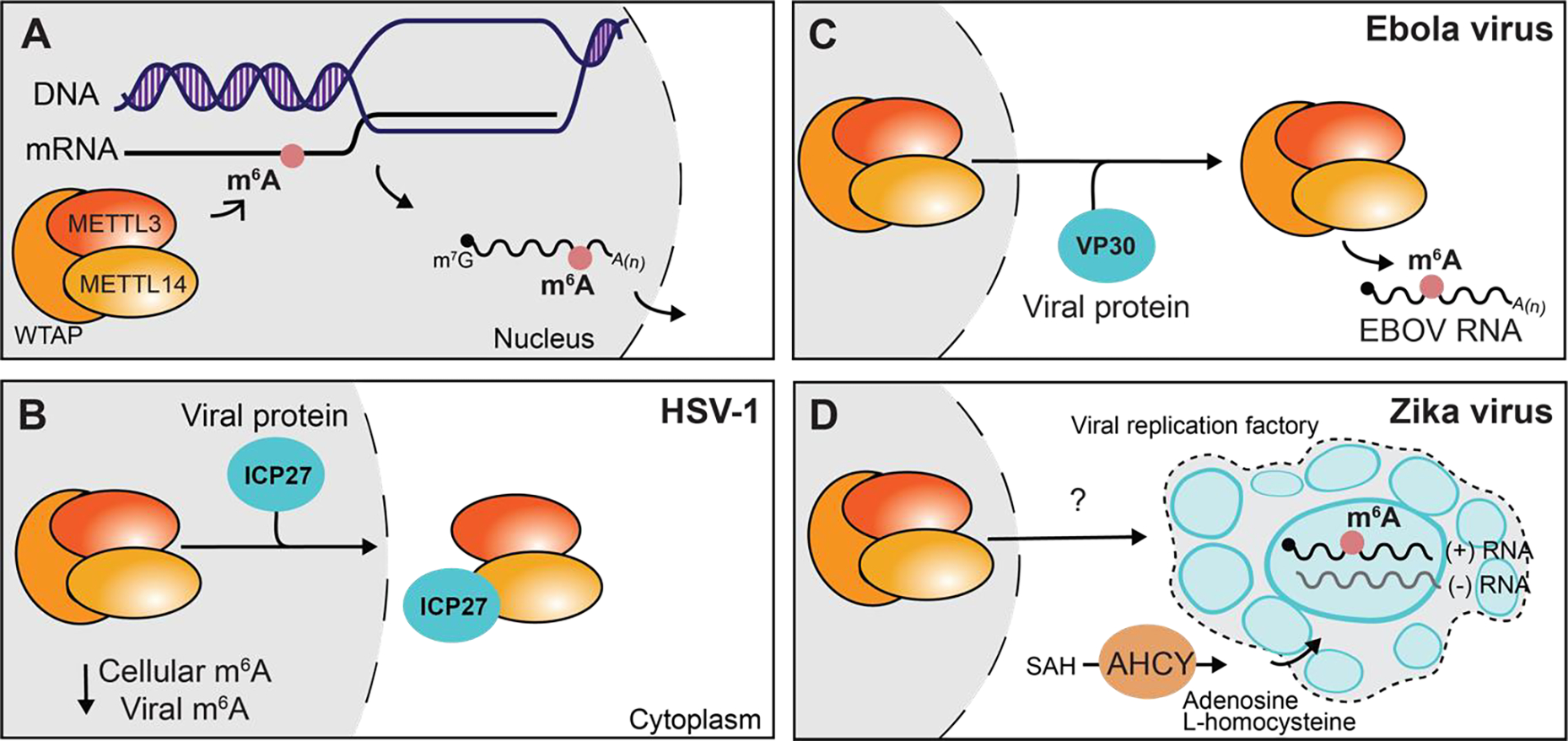
A) m6A is added to nascent mRNA in the nucleus by the METTL3-METTL14-WTAP m6A-methyltransferase complex. B) The HSV-1 ICP27 protein exports the m6A-methyltransferase complex proteins METTL3 and METTL14 from the nucleus, decreasing both viral and cellular RNA m6A-modification. C) The Ebola virus VP30 protein alters the localization of the m6A-methyltransferase complex to modify viral RNA in the cytoplasm. D) Zika virus recruits the metabolic enzyme AHCY to viral replication factories indirectly leading to m6A-modification of the viral RNA by the m6A-methyltransferase complex. See text for more details.
m6A and antiviral innate immunity
One of the described functions for m6A on viral RNA is to prevent recognition by the antiviral innate immune system, including the pattern recognition receptors RIG-I and MDA5 [6]. Indeed, this is generally the case for SARS-CoV-2 [21], described more below. Similarly, m6A on viral RNA limits RIG-I activation during infection by non-segmented, negative-strand RNA viruses, including those from the Pneumoviridae (respiratory syncytial virus (RSV)), Paramyxoviridae, and Rhabdoviridae families [22,23]. Experiments performed during infection with VSV, a virus in the Rhabdoviridae family, suggest that m6A can alter the dsRNA content of the viral RNA to prevent sensing by RIG-I and MDA5, thereby limiting induction of type I IFN [12]. As mentioned above, m6A-modification of ZIKV RNA also inhibits innate immune sensing of the viral RNA [15]. While these examples suggest that m6A is proviral, there are examples of m6A having antiviral functions. For example, in the plant species Arabidopsis thaliana, m6A on the RNA of alfalfa mosaic virus recruits the antiviral YTH domain-containing ECT proteins to the viral RNA to sequester it away from viral replication machinery [24]. Similarly, YTHDF proteins bind to m6A on HCV RNA to sequester the viral RNA from virion assembly machinery [25]. A complete understanding of how m6A interacts with the antiviral innate immune defenses will become clearer when the full complement of RNA binding proteins that differentially bind to viral RNA during infection are identified.
Regulation of m6A-reader proteins during viral infection
Recent studies have highlighted how a diverse set of m6A-reader proteins (e.g. YTHDF1, YTHDC1, YTHDC2, FMRP, and FXR1) can regulate specific steps of viral lifecycles (Figure 2). For example, they can increase viral RNA translation, promote nuclear export of viral transcripts, retain viral RNA in cells, or promote viral RNA stability [26–29]. Two recent examples in Kaposi’s sarcoma herpesvirus infection (KSHV) highlight how m6A-reader proteins alter the function of both viral and cellular RNA during infection. For example, the novel m6A-reader protein called SND1 stabilizes the m6A-modified viral ORF50 RNA [30]. On the other hand, during KSHV infection the m6A-reader and RNA helicase YTHDC2 prevents the degradation of the m6A-modified IL6 mRNA by the viral endoribonuclease SOX protein [31]. Thus, as illustrated here, both traditional and novel m6A-reader proteins can regulate the function of either viral or cellular RNA during infection.
Figure 2. m6A regulates RNA binding protein interaction with viral or cellular RNA to impact viral infection.

These panels indicate diverse roles of m6A during viral infection that can act on viral or cellular RNA to either promote (A) or inhibit (B) viral infection. See the text for more details.
The m6A-reader proteins can also be regulated by novel post-transcriptional and post-translational mechanisms during viral infection. For example, the enterovirus 2A protease cleaves the YTHDF proteins, which limits type I IFN signaling [32]. YTHDF2 can also be cleaved by cellular caspases during the latent cycle of infection with Epstein-Barr virus (EBV) [33]. This cleavage and subsequent inactivation of YTHDF2 enables sustained expression of the mRNAs encoding host factors required for lytic reactivation [33]. YTHDF2 can also become SUMOylated during EBV infection, and this maintains the latent cycle of EBV infection, as SUMOylated YTHDF2 has increased affinity to m6A-modified viral transcripts that would typically induce lytic replication. As such, SUMOylation of YTHDF2 promotes the decay of these viral transcripts, resulting in sustained viral latency [34]. It may be that other YTHDF proteins are similarly regulated during EBV infection such as YTHDF1, which also suppresses EBV lytic replication [35]. The expression of the YTHDF genes can also be regulated at the post-transcriptional level. For example, the translation of the mRNA molecules encoding the YTHDF genes is blocked by the viral host-shutoff program that occurs during vaccinia virus and HSV-1 infection, indirectly inducing antiviral interferon-stimulated genes [36]. Thus, the m6A-reader proteins can regulate viral infection in diverse ways, by acting on viral or host m6A-modified transcripts, and they can also be regulated during viral infection to affect viral replication directly or indirectly by altering the host response to infection.
m6A can have more than one function in a virus.
m6A on viral RNA can have more than one role in any viral infection, likely depending on how proviral or antiviral RNA binding proteins interact with individual m6A sites. These multiple roles have previously been described in the context of HIV-1 infection (reviewed in [37]); however, a newly published direct RNA sequencing study of HIV-1 RNA instead suggests that there are three highly abundant m6A sites redundant in function [38]. Other recent studies during infection with RSV, EBV, and HBV do suggest that m6A can have more than one function in any particular virus. While we describe those roles here, we note that these analyses are only as strong as the methods used to map m6A in each viruses described. As mentioned above, m6A can promote infection for RSV by limiting antiviral innate immune sensing of the viral RNA [23]. However, in some cell types, m6A on RSV can negatively regulate infection by promoting interactions with the YTHDF proteins that limit viral gene expression [39]. For EBV, m6A can similarly promote interactions with YTHDF proteins leading to decreased viral RNA stability resulting in viral latency [34,35]; it can also promote cell survival during lytic replication, as well as oncogenesis [40,41]. For HBV, m6A aids in nuclear export of the viral RNA [28], promotes viral RNA packaging [42], decreases viral RNA stability [43–45], and regulates HBV pregenomic RNA splicing [44]. The function of m6A in each of these cases seems to be different depending on the specific viral RNA that is modified and differential RNA binding protein interactions with the modified RNA molecule. As a general principle, for any given virus, m6A will likely have multiple regulatory functions that control the outcome of infection.
Studies on m6A during SARS-Cov-2 infection
Several groups have now studied m6A in the context of this SARS-CoV-2. Here, we will highlight many of the similar core findings of these studies. The published work on SARS-CoV-2 reveals that the viral RNA molecules contain m6A, although there are notable differences in its relative abundance and nucleotide position. These studies typically used meRIP-seq (m6A-RNA immunoprecipitation followed by sequencing) [46,47] to map the modification on the viral RNA, and several studies coupled this technique with a control experiment to remove m6A, or used higher resolution mapping methods, such as miCLIP (m6A individual-nucleotide-resolution cross-linking and immunoprecipitation) [48]. The results of all these studies reveal that m6A on SARS-CoV-2 is typically reproducibly detected in the nucleocapsid gene near the 3’ end of the genome, that there are ~5–14 m6A sites within the viral RNA, and that the negative strand replication intermediate RNA may have an m6A site [16–19,21]. Recent studies have also performed direct RNA sequencing of the SARS-CoV-2 genome, but they have not uniformly used m6A-negative controls for analysis of the data. For example, one direct RNA sequencing study does use this control and predicts a low-confidence METTL3-dependent m6A site that is not within a classical m6A consensus sequence motif (DRACH) [19]. In contrast, another direct RNA sequencing study does not use this control and instead predicts several m6A sites within these motifs [16]. This study also suggests that while a core set of m6A sites are conserved among cell types in SARS-CoV-2 viral RNA molecules, cell type-specific m6A sites exist [16]. Overall, the mapping results of m6A on SARS-CoV-2 do not indicate if the m6A profiles of individual viral RNA molecules are unique or if they co-occur. Further, they do not reveal if m6A occurs in full-length viral RNA, subgenomic viral RNA molecules, or shorter viral RNA molecules, such as the defective viral genomes that can form in SARS-CoV-2 RNA [49]. Further work will be required to understand the discrepancies between direct RNA sequencing and the other m6A mapping studies on SARS-CoV-2 viral RNA. m6A antibody-based mapping approaches are challenging to perform and can have significant limitations as they do not prove nucleotide resolution or stoichiometry, and as such, a combination of m6A mapping strategies, including quantitative approaches, likely coupled with direct RNA sequencing, may be required to define m6A sites accurately [8,50]. Based on the described data, it seems likely that the profile and abundance of some of these m6A sites on SARS-CoV-2 RNA may differ between cell types.
As SARS-CoV-2 RNA molecules contain m6A and the virus replicates exclusively in the cytoplasm, mechanisms to ensure cytoplasmic localization of m6A-methyltransferase complex proteins must exist. In fact, three studies all report that m6A-machinery proteins have altered localization to the cytoplasm during infection. In Vero E6 cells, SARS-CoV-2 infection results in cytoplasmic accumulation of m6A-methyltransferase complex proteins METTL3, METTL14, WTAP, as well as the m6A-demethylases ALKBH5 and FTO [16]. Others find that METTL3, but not METTL14, partially localizes to the cytoplasm after infection in both Vero cells and bronchial epithelial cell line, BEAS-2B [17]. In Huh7 cells, SARS-CoV-2 infection leads to increased expression of METTL14 and ALKBH5 in the cytoplasm [18]. While there are discrepancies on which m6A-machinery proteins have increased localization in the cytoplasm during SARS-CoV-2, there is cumulative evidence for altered localization of the m6A machinery. Intriguingly, this virally-induced relocalization of the m6A machinery to the cytoplasm can lower the cellular mRNA m6A levels and alter cellular gene expression, which can be reversed by treatment of cells with an inhibitor of the nuclear export protein XPO1 [17]. We do note that others have shown that instead of reducing cellular m6A levels, SARS-CoV-2 infection led to increased cellular m6A, although as this study also showed increased cytoplasmic localization of METTL14, it seems unclear how this could happen [18].
When studying how m6A impacts viral replication, most of the SARS-CoV-2 publications have found that m6A is proviral, meaning that depletion of the m6A-methyltransferase METTL3 or its inhibition by treatment with the chemical inhibitor STM2457 reduces viral infection [16,17,19,21]. However, one study found that m6A is antiviral to SARS-CoV-2, such that depletion of METTL3 or METTL14, or the m6A-binding protein YTHDF2, results in increased levels of viral RNA and infectious virus [18]. The reasons for these discrepancies are unclear. Similarly, it is not clear if the phenotypic differences observed between studies result from m6A changes to cellular or viral RNA. As each study used a different human cell line for these experiments, such as colorectal adenocarcinoma Caco2 cells, bronchial epithelial BEAS-2B cells, hepatocellular carcinoma Huh7 cells, and lung adenocarcinoma A549 or Calu3 cells, it could be that the complement of m6A regulators and functions are different between these cells. Previous studies on m6A in KSHV have similarly identified differential roles for m6A depending on the cell line [51]. None of the published SARS-CoV-2 studies definitively linked the relative contribution of m6A on viral RNA or cellular RNA to infection. One study has suggested that m6A on SARS-CoV-2 RNA prevents RIG-I sensing [21], but as this study used in vitro transcribed viral RNA, whose 5’ end structures are known to trigger RIG-I activation [52], and MDA5 is typically thought to be the primary RNA sensor for SARS-CoV-2 [53], the interpretation of these results is unclear. It is possible that m6A on infection-generated defective viral genomes, which are potent RIG-I ligands [54], does, in fact, inhibit their sensing by RIG-I. As described earlier, m6A levels on cellular mRNA during SARS-CoV-2 infection can change, with both increased and decreased m6A in cellular mRNAs reported, including in antiviral ISGs or proviral host factors, respectively [17,21]. If changes in the relative m6A levels of such key host mRNA molecules alter their expression, it seems likely that those changes could also impact SARS-CoV-2 infection, although the reasons for the discrepancy between these and the other studies described in this section are unclear.
Summary and Conclusions
This review highlights several recent studies that shape our understanding of the novel ways in which m6A can regulate viral infection by acting on either viral or cellular RNA molecules. We have described mechanisms that viruses use to alter the localization of the m6A-machinery to different subcellular compartments for novel viral or cellular RNA targeting. Interestingly, these mechanisms often use viral protein-mediated targeting of the m6A-machinery. Future studies will undoubtedly define more viral proteins that interact with members of the m6A-machinery, including the m6A demethylases FTO and ALKBH5, to regulate viral or cellular m6A-modification during infection. We have also highlighted how established and newly identified m6A-reader proteins can act on viral or cellular RNA to regulate their translation, localization, and stability during infection thereby impacting viral infection in diverse ways. Similarly, we have highlighted recent examples of how the expression of m6A-readers can be regulated at the post-transcriptional and post-translational levels by viral infection. Additionally, we have described several roles for m6A in regulating the host antiviral response to viral infection and illustrated that for any given virus, including SARS-CoV-2, m6A will likely have multiple regulatory functions that control the outcome of infection. Finally, we have delved into the latest studies on the function of m6A during SARS-CoV-2 infection, comparing the literature to highlight key themes.
Overall, numerous studies have implicated m6A as having either proviral or antiviral roles during viral infection. This regulation undoubtedly comes from the actions of m6A on both cellular and viral RNA molecules, as we have shown previously for HCV [14]. For some viruses, there are conflicting reports on the position and abundance of m6A on viral RNA (e.g. SARS-CoV-2 or HIV-1 noted above, but also others [25,55–57]). These differences are likely due to variances in cell lines used and m6A mapping strategies. In addition, the limited ability of many m6A mapping strategies to measure m6A stoichiometry in a nucleotide-specific manner prevents a real understanding of how individual m6A sites contribute to function in a viral RNA. New m6A mapping approaches may overcome some of these barriers, and a combination of methods may be useful for mapping viral m6A, as we outline in a recent perspective [8]. Ideally, high-resolution approaches to map m6A at single-nucleotide resolution on viral RNA molecules should focus on determining stoichiometry and the m6A profile of full-length viral RNA molecules in relevant cell types. Depending on the virus, this may include mapping m6A in viral RNA molecules unique to individual life cycle stages. Beyond higher-resolution m6A mapping, future research on viral m6A will also benefit from a more nuanced understanding of how m6A-RNA binding proteins interactions are regulated in the target cells of individual viruses. Undoubtedly, just as studies on viruses have revealed novel aspects of RNA biology, they will reveal novel regulation of m6A that could be important during viral infection or for cellular and organismal function.
Acknowledgments
We thank current members of the Horner Lab, especially Dr. Matthew Thompson, for valuable feedback and discussion. Research on m6A in the Horner lab has been supported by Burroughs Wellcome Fund and National Institutes of Health grant R01AI125416.
Footnotes
Declarations of interest:
none.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and recommended reading
Papers of particular interest, published within the period of review have been highlighted as:
* of special interest
** of outstanding interest
- 1.He PC, He C: m(6) A RNA methylation: from mechanisms to therapeutic potential. EMBO J 2021, 40:e105977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Garcias Morales D, Reyes JL: A birds’-eye view of the activity and specificity of the mRNA m(6) A methyltransferase complex. Wiley Interdiscip Rev RNA 2021, 12:e1618. [DOI] [PubMed] [Google Scholar]
- 3.Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS, et al. : Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res 2014, 24:177–189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Zaccara S, Ries RJ, Jaffrey SR: Reading, writing and erasing mRNA methylation. Nat Rev Mol Cell Biol 2019, 20:608–624. [DOI] [PubMed] [Google Scholar]
- 5.Flamand MN, Tegowski M, Meyer KD: The Proteins of mRNA Modification: Writers, Readers, and Erasers. Annu Rev Biochem 2023, 92:145–173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Aufgebauer CJ, Bland KM, Horner SM: Modifying the antiviral innate immune response by selective writing, erasing, and reading of m(6)A on viral and cellular RNA. Cell Chem Biol 2024, 31:100–109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Baquero-Perez B, Geers D, Diez J: From A to m(6)A: The Emerging Viral Epitranscriptome. Viruses 2021, 13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Horner SM, Thompson MG: Challenges to mapping and defining m(6)A function in viral RNA. RNA 2024, 30:482–490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kim GW, Siddiqui A: Hepatitis B virus X protein recruits methyltransferases to affect cotranscriptional N6-methyladenosine modification of viral/host RNAs. Proc Natl Acad Sci U S A 2021, 118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Srinivas KP, Depledge DP, Abebe JS, Rice SA, Mohr I, Wilson AC: Widespread remodeling of the m(6)A RNA-modification landscape by a viral regulator of RNA processing and export. Proc Natl Acad Sci U S A 2021, 118. **The HSV-1 viral protein ICP27 exports the m6A-methyltransferase proteins METTL3 and METTL14 from the nucleus to the cytoplasm over the course of infection, resulting in decreases viral and cellular RNA methylation. While HSV-1 mRNA molecules contain m6A at early times of infection, this redistribution of m6A at later times of infection suggests that the ongoing viral RNA methylation does not need to be sustained during infection. Importantly, it also suggests that demethylation of cellular mRNA molecules may benefit the viral gene expression program.
- 11.Wendt L, Pickin MJ, Bodmer BS, Reiche S, Feneant L, Holper JE, Fuchs W, Groseth A, Hoenen T: N(6)-methyladenosine is required for efficient RNA synthesis of Ebola virus and other haemorrhagic fever viruses. Emerg Microbes Infect 2023, 12:2223732. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Qiu W, Zhang Q, Zhang R, Lu Y, Wang X, Tian H, Yang Y, Gu Z, Gao Y, Yang X, et al. : N(6)-methyladenosine RNA modification suppresses antiviral innate sensing pathways via reshaping double-stranded RNA. Nat Commun 2021, 12:1582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Zhang T, Shi C, Hu H, Zhang Z, Wang Z, Chen Z, Feng H, Liu P, Guo J, Lu Q, et al. : N6-methyladenosine RNA modification promotes viral genomic RNA stability and infection. Nat Commun 2022, 13:6576. **A wheat homolog for METTL14, TaMTB, was identified analyzing GWAS and QTL for resistance to wheat yellow mosaic virus (WYMV) infection. TaMTB was found to have m6A methyltransferase activity and promote WYMV m6A-modification. Intriguingly, the viral Nib protein relocalized TaMTB from the nucleus to cytoplasmic aggerates during infection. This protein is a potential target for genetic engineering for pathogen resistance in wheat.
- 14.Sacco MT, Bland KM, Horner SM: WTAP Targets the METTL3 m(6)A-Methyltransferase Complex to Cytoplasmic Hepatitis C Virus RNA to Regulate Infection. J Virol 2022, 96:e0099722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Denolly S, Stukalov A, Barayeu U, Rosinski AN, Kritsiligkou P, Joecks S, Dick TP, Pichlmair A, Bartenschlager R: Zika virus remodelled ER membranes contain proviral factors involved in redox and methylation pathways. Nat Commun 2023, 14:8045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhang X, Hao H, Ma L, Zhang Y, Hu X, Chen Z, Liu D, Yuan J, Hu Z, Guan W: Methyltransferase-like 3 Modulates Severe Acute Respiratory Syndrome Coronavirus-2 RNA N6-Methyladenosine Modification and Replication. mBio 2021, 12:e0106721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Vaid R, Mendez A, Thombare K, Burgos-Panadero R, Robinot R, Fonseca BF, Gandasi NR, Ringlander J, Hassan Baig M, Dong JJ, et al. : Global loss of cellular m(6)A RNA methylation following infection with different SARS-CoV-2 variants. Genome Res 2023, 33:299–313. **This study reveals that upon SARS-CoV-2 infection, cellular mRNA molecules lose m6A, as METTL3 relocalizes to the cytoplasm to add m6A to viral RNA. Importantly, upon inhibition of the cellular nuclear export protein XPO1, expression of METTL3 in the nucleus is restored, which leads to recovery of m6A on cellular transcripts and decreases viral infection. This suggests that viral and cellular m6A likely have opposing effects on infection.
- 18.Liu J, Xu YP, Li K, Ye Q, Zhou HY, Sun H, Li X, Yu L, Deng YQ, Li RT, et al. : The m(6)A methylome of SARS-CoV-2 in host cells. Cell Res 2021, 31:404–414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Burgess HM, Depledge DP, Thompson L, Srinivas KP, Grande RC, Vink EI, Abebe JS, Blackaby WP, Hendrick A, Albertella MR, et al. : Targeting the m(6)A RNA modification pathway blocks SARS-CoV-2 and HCoV-OC43 replication. Genes Dev 2021, 35:1005–1019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pereira-Montecinos C, Toro-Ascuy D, Ananias-Saez C, Gaete-Argel A, Rojas-Fuentes C, Riquelme-Barrios S, Rojas-Araya B, Garcia-de-Gracia F, Aguilera-Cortes P, Chnaiderman J, et al. : Epitranscriptomic regulation of HIV-1 full-length RNA packaging. Nucleic Acids Res 2022, 50:2302–2318. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Li N, Hui H, Bray B, Gonzalez GM, Zeller M, Anderson KG, Knight R, Smith D, Wang Y, Carlin AF, et al. : METTL3 regulates viral m6A RNA modification and host cell innate immune responses during SARS-CoV-2 infection. Cell Rep 2021, 35:109091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Lu M, Xue M, Wang HT, Kairis EL, Ahmad S, Wei J, Zhang Z, Liu Q, Zhang Y, Gao Y, et al. : Nonsegmented Negative-Sense RNA Viruses Utilize N(6)-Methyladenosine (m(6)A) as a Common Strategy To Evade Host Innate Immunity. J Virol 2021, 95. *This study shows that the Arabidopsis YTH-containing m6A binding protein ECT2 binds viral m6A on alfalfa mosaic virus to limit infection. This suggests that m6A and/or the ECT2 protein is part of the plant antiviral defense system, broadly suggesting that m6A does not always inhibit antiviral innate immunity.
- 23.Xue M, Zhang Y, Wang H, Kairis EL, Lu M, Ahmad S, Attia Z, Harder O, Zhang Z, Wei J, et al. : Viral RNA N6-methyladenosine modification modulates both innate and adaptive immune responses of human respiratory syncytial virus. PLoS Pathog 2021, 17:e1010142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Martinez-Perez M, Aparicio F, Arribas-Hernandez L, Tankmar MD, Rennie S, von Bulow S, Lindorff-Larsen K, Brodersen P, Pallas V: Plant YTHDF proteins are direct effectors of antiviral immunity against an N6-methyladenosine-containing RNA virus. EMBO J 2023, 42:e113378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gokhale NS, McIntyre ABR, McFadden MJ, Roder AE, Kennedy EM, Gandara JA, Hopcraft SE, Quicke KM, Vazquez C, Willer J, et al. : N6-Methyladenosine in Flaviviridae Viral RNA Genomes Regulates Infection. Cell Host Microbe 2016, 20:654–665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kim GW, Siddiqui A: N6-methyladenosine modification of HCV RNA genome regulates cap-independent IRES-mediated translation via YTHDC2 recognition. Proc Natl Acad Sci U S A 2021, 118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kim GW, Moon JS, Gudima SO, Siddiqui A: N(6)-Methyladenine Modification of Hepatitis Delta Virus Regulates Its Virion Assembly by Recruiting YTHDF1. J Virol 2022, 96:e0112422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kim GW, Imam H, Siddiqui A: The RNA Binding Proteins YTHDC1 and FMRP Regulate the Nuclear Export of N(6)-Methyladenosine-Modified Hepatitis B Virus Transcripts and Affect the Viral Life Cycle. J Virol 2021, 95:e0009721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Hajikhezri Z, Kaira Y, Schubert E, Darweesh M, Svensson C, Akusjarvi G, Punga T: Fragile X-Related Protein FXR1 Controls Human Adenovirus Capsid mRNA Metabolism. J Virol 2023, 97:e0153922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Baquero-Perez B, Antanaviciute A, Yonchev ID, Carr IM, Wilson SA, Whitehouse A: The Tudor SND1 protein is an m(6)A RNA reader essential for replication of Kaposi’s sarcoma-associated herpesvirus. Elife 2019, 8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Macveigh-Fierro D, Cicerchia A, Cadorette A, Sharma V, Muller M: The m(6)A reader YTHDC2 is essential for escape from KSHV SOX-induced RNA decay. Proc Natl Acad Sci U S A 2022, 119. *This study reveals an example of a host mRNA molecule protecting its expression via m6A. Specifically, while most mRNA molecules are degraded by the KSHV SOX protein during infection, the presence of m6A and associated YTHDC2-binding on the interleukin-6 transcript, prevent this transcript from being degraded by SOX during infection.
- 32.Kastan JP, Tremblay MW, Brown MC, Trimarco JD, Dobrikova EY, Dobrikov MI, Gromeier M: Enterovirus 2A(pro) Cleavage of the YTHDF m(6)A Readers Implicates YTHDF3 as a Mediator of Type I Interferon-Driven JAK/STAT Signaling. mBio 2021, 12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zhang K, Zhang Y, Maharjan Y, Sugiokto FG, Wan J, Li R: Caspases Switch off the m(6)A RNA Modification Pathway to Foster the Replication of a Ubiquitous Human Tumor Virus. mBio 2021, 12:e0170621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Sugiokto FG, Saiada F, Zhang K, Li R: SUMOylation of the m6A reader YTHDF2 by PIAS1 promotes viral RNA decay to restrict EBV replication. mBio 2024, 15:e0316823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Xia TL, Li X, Wang X, Zhu YJ, Zhang H, Cheng W, Chen ML, Ye Y, Li Y, Zhang A, et al. : N(6)-methyladenosine-binding protein YTHDF1 suppresses EBV replication and promotes EBV RNA decay. EMBO Rep 2021, 22:e50128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hesser CR, Walsh D: YTHDF2 Is Downregulated in Response to Host Shutoff Induced by DNA Virus Infection and Regulates Interferon-Stimulated Gene Expression. J Virol 2023, 97:e0175822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Riquelme-Barrios S, Pereira-Montecinos C, Valiente-Echeverria F, Soto-Rifo R: Emerging Roles of N(6)-Methyladenosine on HIV-1 RNA Metabolism and Viral Replication. Front Microbiol 2018, 9:576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Baek A, Lee GE, Golconda S, Rayhan A, Manganaris AA, Chen S, Tirumuru N, Yu H, Kim S, Kimmel C, et al. : Single-molecule epitranscriptomic analysis of full-length HIV-1 RNAs reveals functional roles of site-specific m(6)As. Nat Microbiol 2024, 9:1340–1355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Figueroa F, Vega-Gibson A, Catrileo J, Gaete-Argel A, Riquelme-Barrios S, Alonso-Palomares LA, Tapia LI, Valiente-Echeverria F, Soto-Rifo R, Acevedo ML: N(6) - Methyladenosine Negatively Regulates Human Respiratory Syncytial Virus Replication. Front Cell Dev Biol 2021, 9:739445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yanagi Y, Watanabe T, Hara Y, Sato Y, Kimura H, Murata T: EBV Exploits RNA m(6)A Modification to Promote Cell Survival and Progeny Virus Production During Lytic Cycle. Front Microbiol 2022, 13:870816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lang F, Singh RK, Pei Y, Zhang S, Sun K, Robertson ES: EBV epitranscriptome reprogramming by METTL14 is critical for viral-associated tumorigenesis. PLoS Pathog 2019, 15:e1007796. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kim GW, Moon JS, Siddiqui A: N6-methyladenosine modification of the 5’ epsilon structure of the HBV pregenome RNA regulates its encapsidation by the viral core protein. Proc Natl Acad Sci U S A 2022, 119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Kim GW, Siddiqui A: Hepatitis B Virus X Protein Expression Is Tightly Regulated by N6-Methyladenosine Modification of Its mRNA. J Virol 2022, 96:e0165521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tsukuda S, Harris JM, Magri A, Balfe P, Siddiqui A, Wing PAC, McKeating JA: The N6-methyladenosine demethylase ALKBH5 regulates the hypoxic HBV transcriptome. PLoS Pathog 2024, 20:e1011917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Murata T, Iwahori S, Okuno Y, Nishitsuji H, Yanagi Y, Watashi K, Wakita T, Kimura H, Shimotohno K: N6-methyladenosine Modification of Hepatitis B Virus RNA in the Coding Region of HBx. Int J Mol Sci 2023, 24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR: Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 2012, 149:1635–1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, et al. : Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 2012, 485:201–206. [DOI] [PubMed] [Google Scholar]
- 48.Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR: Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 2015, 12:767–772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Girgis S, Xu Z, Oikonomopoulos S, Fedorova AD, Tchesnokov EP, Gordon CJ, Schmeing TM, Gotte M, Sonenberg N, Baranov PV, et al. : Evolution of naturally arising SARS-CoV-2 defective interfering particles. Commun Biol 2022, 5:1140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.McIntyre ABR, Gokhale NS, Cerchietti L, Jaffrey SR, Horner SM, Mason CE: Limits in the detection of m(6)A changes using MeRIP/m(6)A-seq. Sci Rep 2020, 10:6590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hesser CR, Karijolich J, Dominissini D, He C, Glaunsinger BA: N6-methyladenosine modification and the YTHDF2 reader protein play cell type specific roles in lytic viral gene expression during Kaposi’s sarcoma-associated herpesvirus infection. PLoS Pathog 2018, 14:e1006995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Schlee M, Hartmann E, Coch C, Wimmenauer V, Janke M, Barchet W, Hartmann G: Approaching the RNA ligand for RIG-I? Immunol Rev 2009, 227:66–74. [DOI] [PubMed] [Google Scholar]
- 53.Diamond MS, Kanneganti TD: Innate immunity: the first line of defense against SARS-CoV-2. Nat Immunol 2022, 23:165–176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Vignuzzi M, Lopez CB: Defective viral genomes are key drivers of the virus-host interaction. Nat Microbiol 2019, 4:1075–1087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Baquero-Perez B, Yonchev ID, Delgado-Tejedor A, Medina R, Puig-Torrents M, Sudbery I, Begik O, Wilson SA, Novoa EM, Diez J: N(6)-methyladenosine modification is not a general trait of viral RNA genomes. Nat Commun 2024, 15:1964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kim B, Arcos S, Rothamel K, Jian J, Rose KL, McDonald WH, Bian Y, Reasoner S, Barrows NJ, Bradrick S, et al. : Discovery of Widespread Host Protein Interactions with the Pre-replicated Genome of CHIKV Using VIR-CLASP. Mol Cell 2020, 78:624–640 e627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Bhattacharya T, Yan L, Crawford JM, Zaher H, Newton ILG, Hardy RW: Differential viral RNA methylation contributes to pathogen blocking in Wolbachia-colonized arthropods. PLoS Pathog 2022, 18:e1010393. [DOI] [PMC free article] [PubMed] [Google Scholar]


