ABSTRACT
Despite their low quantity and abundance, the cellulolytic bacteria that inhabit the equine large intestine are vital to their host, as they enable the crucial use of forage-based diets. Fibrobacter succinogenes is one of the most important intestinal cellulolytic bacteria. In this study, Fibrobacter sp. HC4, one cellulolytic strain newly isolated from the horse cecum, was characterized for its ability to utilize plant cell wall fibers. Fibrobacter sp. HC4 consumed only cellulose, cellobiose, and glucose and produced succinate and acetate in equal amounts. Among genes coding for CAZymes, 26% of the detected glycoside hydrolases (GHs) were involved in cellulolysis. These cellulases belong to the GH5, GH8, GH9, GH44, GH45, and GH51 families. Both carboxymethyl cellulase and xylanase activities of Fibrobacter sp. HC4 were detected using the Congo red method and were higher than those of F. succinogenes S85, the type strain. The in vitro addition of Fibrobacter sp. HC4 to a fecal microbial ecosystem of horses with large intestinal acidosis significantly enhanced fibrolytic activity as measured by the increase in gas and volatile fatty acids production during the first 48 h. According to this, the pH decreased and the disappearance of dry matter increased at a faster rate with Fibrobacter sp. HC4. Our data suggest a high specialization of the new strain in cellulose degradation. Such a strain could be of interest for future exploitation of its probiotic potential, which needs to be further determined by in vivo studies.
IMPORTANCE
Cellulose is the most abundant of plant cell wall fiber and can only be degraded by the large intestine microbiota, resulting in the production of volatile fatty acids that are essential for the host nutrition and health. Consequently, cellulolytic bacteria are of major importance to herbivores. However, these bacteria are challenged by various factors, such as high starch diets, which acidify the ecosystem and reduce their numbers and activity. This can lead to an imbalance in the gut microbiota and digestive problems such as colic, a major cause of mortality in horses. In this work, we characterized a newly isolated cellulolytic strain, Fibrobacter sp. HC4, from the equine intestinal microbiota. Due to its high cellulolytic capacity, reintroduction of this strain into an equine fecal ecosystem stimulates hay fermentation in vitro. Isolating and describing cellulolytic bacteria is a prerequisite for using them as probiotics to restore intestinal balance.
KEYWORDS: culture, cellulose-degrading strain, xylan-degrading strain, CAZymes, large intestinal microbiota, in vitro, acidosis, probiotics, horses
INTRODUCTION
While cellulose is the major compound of dietary plant complex carbohydrates, only a small number of microorganisms, including cellulolytic bacteria, have the capacity to initiate its degradation in the gastrointestinal tract (1, 2). Cellulolytic bacteria possess specific carbohydrate-active enzymes (CAZymes) involved in the process of breaking down cellulose, named cellulases, that were found in at least 16 glycoside hydrolase (GHs) families (GH5-GH10, GH12, GH16, GH26, GH39, GH44-45, GH48, GH51, GH74, and GH124) according to CAZy database (http://www.cazy.org/). Cellulases are enzymes that hydrolyze the β−1,4 links of cellulose. A distinction is made between endoglucanases and exoglucanases. Endoglucanases “endohydrolyze” cellulose, releasing smaller cellulose chains known as cellodextrins, while exoglucanases, also known as cellobiosidases, hydrolyze the β−1,4 links at the ends of cellulose, releasing cellobiose. Some cellulolytic bacteria, such as Fibrobacter succinogenes, can break down hemicellulose only to gain access to cellulose (3). Xylan is the major hemicellulose, and the enzymes involved in its degradation were found in at least eight GHs families (GH3, GH5, GH8, GH10, GH11, GH16, GH30, and GH43) according to CAZy database (http://www.cazy.org/).
In the large intestine of herbivores, whose diet is very rich in fiber, cellulose degradation allows releasing nutrients on which a large part of the microbial community depends, designating cellulolytic bacteria as “keystone” species (4). Despite their crucial role, cellulolytic bacteria are present in very low abundance in the large intestine. In equine, these bacteria represent 1%–9% of the total bacteria and their concentration average 106 cells/mL of horse cecal or colonic contents (5). These bacteria are very sensitive to stressful environmental factors, such as vermifuge, antibiotics, or diet, and can decline rapidly under stressful conditions (6–9). Yet, studying the metabolic and physiological functions of such key species is critical not only to help optimizing fiber utilization but also to contribute to limiting microbial dysbiosis that plays a role in the pathogenesis of diseases such as laminitis or colitis in horses (8). Three cellulolytic bacterial species were detected in the large intestine of horse using specific nucleotide probes, i.e., F. succinogenes, Ruminococcus flavefaciens, and Ruminococcus albus (10–14). Four isolates resembled ruminal strains of R. flavefaciens morphologically but differed metabolically (11), and one isolate of F. succinogenes was identified as subspecies II of F. succinogenes and eight as a new F. succinogenes subspecies V (15). Today, none of these isolates are available in the collection.
Cellulolytic bacteria require specific nutrients, and growth factors, some of which are still unidentified. Thus, culture media use horse cecal or fecal fluid whose composition is variable from one sample to another and uncertain (16). Cellulolytic bacteria are highly sensitive to acidity. Maintaining a constant acidic intracellular pH creates a proton gradient that causes an excessive accumulation of toxic anions (17). Furthermore, when pH is lower than 6.0, the attachment of cellulolytic bacteria to fiber (rice straw) is inhibited (18). Such acidic environmental condition occurs when horses are fed a high starch diet, provoking the decrease of cellulolytic bacteria in the large intestine (8). Though specific tests for measuring sensitivity to a percentage of O2 have not been performed, cellulolytic bacteria from the large intestine have always been recognized as strict anaerobes (19–21) and require specific facilities for their growth under laboratory conditions. Obtaining cultured representatives of cellulolytic bacteria is thus difficult, and few cellulolytic bacteria have been isolated from the large intestine of equine using culture-dependent methods (21). Recently, our group isolated a novel strain of F. succinogenes from equine cecal content, named HC4, selected for its high efficiency to degrade cellulose (22). This strain is deposited at the German Collection of Microorganisms and Cell Cultures (DSMZ), under number DSM 33656.22
In the present study, we further characterized strain HC4 using phenotypic tests and carbon substrate utilization tests. Enzyme capacity was evaluated based on genomic characterization for carbon and nitrogen metabolism, and cellulases and xylanases were assessed by functional assays demonstrating fibrolytic activities. Finally, we conducted in vitro tests to investigate the functionality of this newly isolated cellulolytic strain in case of acidosis of the large intestinal ecosystem induced by dietary stress with high starch content.
RESULTS
Rapid growth on cellulose and large production of succinate and acetate
Fibrobacter sp. HC4 appeared as Gram-negative rods or pleomorphic ovoid cells under the light microscope. It grew on filter paper cellulose (Whatman No.1) or on microcrystalline cellulose powder (Sigmacell 50 cellulose -Sigma Chemicals) as the sole carbon source in the media. Observations by scanning electron microscope (SEM) showed rod cells of 1.5 µm in length and 0.63 µm in diameter attached on cellulose fiber of the filter paper used as a substrate (Fig. 1). Fibrobacter sp. HC4 consumed glucose and cellobiose. No growth was observed on raffinose, maltose, lactose, xylose, mannose, rhamnose, arabinose, sucrose, galactose, melezitose, mannitol, sorbitol, and on beechwood xylan, although xylanase was detected.
Fig 1.

Observation of Fibrobacter sp. HC4 cells on filter paper cellulose by SEM after a cryo-protection treatment (nitrogen gel at −140°C) by DImaCell after 48 h of culture (left: scale of 10.0 µm; right: scale of 5.00 µm).
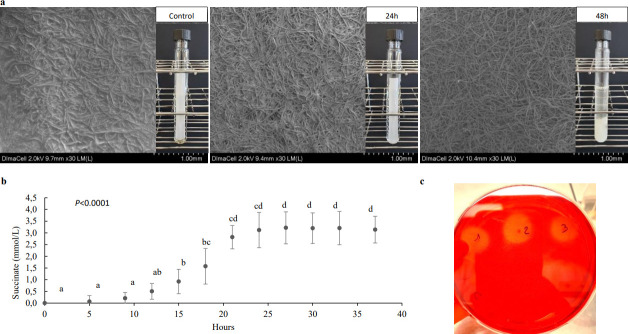
After 48 h cultures on filter paper cellulose at 38°C, Fibrobacter sp. HC4 produced equally succinate (47.02 ± 0.15 µmol/mg of protein) and acetate (45.69 ± 1.8 µmol/mg of protein). Propionate, butyrate, and formate were not detected. Using SEM, we observed that cellulose degradation occurred as early as 24 h, while nothing was yet discernable by naked-eye examination at the same time. The filter paper was completely degraded after 48 h (Fig. 2a). The stationary growth phase of Fibrobacter sp. HC4 was reached after 22 h on filter paper cellulose, as measured by the production of succinate (Fig. 2b). Concomitantly, the concentration estimated by the most probable number (MPN) reached 108 viable cells per mL. Fibrobacter sp. HC4 was also able to grow on washed dried ground hay and degraded 33.26% ± 2.05% of hay dry matter (DM) after 150 h of culture. Using Congo red, Fibrobacter sp. HC4 demonstrated a fibrolytic activity on carboxymethyl cellulose (CMC; Fig. 2c), beechwood xylan, and dried ground hay as the sole substrate.
Fig 2.
Fibrobacter sp. HC4 growth on cellulose. (a) SEM after a cryo-protection treatment (nitrogen gel at −140°C) by DImaCell of filter paper cellulose without bacteria (control), with Fibrobacter sp. HC4 growing on after 24 h and after 48 h on a scale of 1.00 mm. (b) Production of succinate by Fibrobacter sp. HC4 growing on filter paper cellulose (means and SD). Different letters indicate a significant difference in succinate concentration as a function of culture time. (c) Activity of CMCase by the strain HC4 detected by Congo Red method.
Anaerobic behavior
We tested O2 sensitivity of Fibrobacter sp. HC4 and F. succinogenes S85, the type strain, in Hungate tubes using adapted liquid media that were not saturated with CO2. Similar observations were made with both strains of Fibrobacter. Strains HC4 and S85 did not grow on cellulose or glucose in an aerated environment. After exposure to O2 for 2 min, both strains resumed growth when returned to strict anaerobic conditions with filter paper cellulose. However, exposure to O2 for 1, 5, or 24 h inhibited subsequent growth on filter paper cellulose, even when anaerobic conditions were restored.
Member of Fibrobacter sp. clade C
Using near-full-length 16S ribosomal RNA sequence, we generated the phylogenetic tree of Fibrobacter sp., with Bacteroides fragilis NCTC 9343 as an out group (Fig. 3). The strain Fibrobacter sp. HC4 isolated from the equine cecum was comparable to the eight ones previously isolated from horse feces and belonged to phenotype V of Fibrobacter (15). This phenotype is part of clade C, which is constituted only by strains of Fibrobacter isolated from feces of monogastric herbivores (horses, tapir, rhinoceros, and elephant), while other identified clades (A and B) contain strains of Fibrobacter originating from both rumen of ruminants and feces of monogastric animals (23). Phenotype V is mostly composed of strains originating from horses (6 out of 7). In the Genome Taxonomy Database (GTDB), strain HC4 belongs to the genus Fibrobacter, and it is classified in the same cluster (named Fibrobacter sp900142465 by GTDB) as the genomes coming from equine strains, including those isolated by Neuman (15). The representative strain of this cluster is Fibrobacter sp. UWH6.
Fig 3.

Molecular phylogenetic analysis by maximum likelihood method algorithm (1,000 bootstrap trial) method. The numbers at each branch represent bootstrap values. The phylogeny tree was constructed using a near-full-length 16S ribosomal RNA sequence (1,269 sites). The B. fragilis type strain NCTC 9343 was included as an outgroup. The red box shows bacteria isolated from horses, except from strain UW T1 which was isolated from tapir feces.
Multiple genes coding for fibrolysis as other Fibrobacter of herbivores
To investigate the putative capacity of Fibrobacter sp. HC4 to utilize dietary plant complex carbohydrates, the genome was screened for the presence of CAZymes. Among the predicted proteins, 222 types of CAZy domains (Table 1) could be detected, such as GHs, glycosyl transferases (GTs), polysaccharide lyases (PLs), carbohydrate esterases (CEs), and carbohydrate-binding modules (CBMs). Fibrobacter sp. HC4 genome had 91 GHs, distributed in 29 families, and including a significant number of GH5, GH43, and GH9 (Table 2). Fibrobacter strains used for comparison were isolated from mammals, either herbivores (ruminants: cattle and sheep; monogastrics: horse, elephant, colobus monkey tapir, and rhinoceros) or omnivores (rat, pig, and rhesus monkey), and from birds (ostriches). The total number of GHs was comparable between the genome of strain HC4 and that of strains originated from other herbivore mammals, especially horse strains, whereas there were twice less GHs in strains from omnivore mammals and herbivore birds. Like other strains of herbivores, Fibrobacter sp. HC4 demonstrated the most GH5 and GH43. The strains of monogastric herbivore animals have more GH5 in their genome than strains of ruminants and omnivores, whereas the number of GH43 is comparable to those of ruminant strains. Strains of omnivores have few GH43 in their genome. Fibrobacter strains of herbivores also have an important number of GH9. However, horse strains, include strain HC4, have the least. Based on CAZymes comparison, strain HC4 was closer to the strains originating from the equine digestive ecosystem than to strains of Fibrobacter originating from other species.
TABLE 1.
CAZyme families identified of Fibrobacter sp. HC4
| Families | |||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 18 | 19 | 23 | 26 | 27 | 28 | 30 | 32 | 35 | 39 | 43 | 44 | 45 | 48 | 50 | 51 | 53 | 57 | 74 | 77 | 88 | 94 | 95 | 116 | 141 | 146 | 171 | Total | |
| GH | 0 | 2 | 2 | 0 | 16 | 0 | 5 | 8 | 4 | 1 | 0 | 5 | 0 | 0 | 4 | 1 | 0 | 4 | 4 | 1 | 0 | 3 | 0 | 0 | 1 | 11 | 1 | 3 | 0 | 0 | 2 | 1 | 3 | 2 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | 91 |
| GT | 1 | 22 | 0 | 17 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 2 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 50 |
| PL | 6 | 0 | 0 | 0 | 0 | 0 | 1 | 2 | 1 | 2 | 1 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 14 |
| CE | 2 | 1 | 0 | 0 | 0 | 2 | 1 | 0 | 0 | 0 | 2 | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 10 |
| CBM | 0 | 0 | 0 | 5 | 0 | 22 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 0 | 11 | 0 | 0 | 0 | 0 | 1 | 8 | 3 | 0 | 0 | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 57 |
| CAZymes total | 222 | ||||||||||||||||||||||||||||||||||||||||||
TABLE 2.
Genes encoding for the different families of GH identified in the Fibrobacter sp. HC4 and in different Fibrobacter strains
| GH | 2 | 3 | 5 | 8 | 9 | 10 | 11 | 13 | 16 | 18 | 23 | 26 | 27 | 30 | 39 | 43 | 44 | 45 | 51 | 53 | 54 | 57 | 74 | 77 | 94 | 95 | 116 | 141 | 146 | 171 | Total | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cow rumen | F. succinogenes subsp. succinogenes S85 | 2 | 2 | 11 | 5 | 7 | 6 | 4 | 3 | 4 | 2 | 2 | 5 | 1 | 4 | 1 | 14 | 1 | 4 | 2 | 2 | 0 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 93 |
| Fibrobacter sp. UWB1 | 2 | 1 | 14 | 7 | 11 | 6 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 13 | 1 | 5 | 2 | 2 | 1 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 104 | |
| Fibrobacter sp. UWB3 | 2 | 3 | 12 | 7 | 10 | 8 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 11 | 1 | 4 | 2 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 100 | |
| Fibrobacter sp. UWB4 | 2 | 1 | 10 | 6 | 9 | 7 | 2 | 3 | 4 | 2 | 3 | 6 | 1 | 4 | 1 | 10 | 1 | 4 | 2 | 1 | 0 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 90 | |
| Fibrobacter sp. UWB5 | 2 | 3 | 13 | 6 | 12 | 9 | 5 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 13 | 1 | 5 | 2 | 2 | 1 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 109 | |
| Fibrobacter sp. UWB6 | 2 | 3 | 13 | 5 | 11 | 6 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 5 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 103 | |
| Fibrobacter sp. UWB7 | 2 | 3 | 12 | 7 | 9 | 8 | 4 | 3 | 4 | 2 | 3 | 6 | 1 | 4 | 1 | 11 | 1 | 3 | 2 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 2 | 100 | |
| Fibrobacter sp UWB8 | 2 | 3 | 13 | 5 | 11 | 6 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 5 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 103 | |
| Fibrobacter sp. UWB11 | 2 | 2 | 12 | 7 | 9 | 7 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 0 | 10 | 0 | 4 | 2 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 94 | |
| Fibrobacter sp. UWB12 | 2 | 1 | 12 | 7 | 9 | 8 | 4 | 3 | 4 | 2 | 3 | 6 | 1 | 4 | 0 | 10 | 0 | 4 | 2 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 95 | |
| Fibrobacter sp. UWB13 | 2 | 3 | 12 | 7 | 9 | 8 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 10 | 1 | 3 | 2 | 1 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 97 | |
| Fibrobacter sp. UWB15 | 2 | 3 | 13 | 5 | 11 | 6 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 5 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 103 | |
| Fibrobacter sp. UWB16 | 2 | 3 | 12 | 7 | 10 | 8 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 11 | 1 | 4 | 2 | 2 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 101 | |
| Sheep rumen | F. succinogenes subsp. elongatus HM2 | 2 | 2 | 12 | 6 | 9 | 9 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 4 | 2 | 2 | 1 | 3 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 102 |
| Fibrobacter sp. UWOV1 | 2 | 2 | 13 | 7 | 12 | 7 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 13 | 1 | 5 | 2 | 2 | 1 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 106 | |
| Horse cecum | Fibrobacter sp. HC4 | 2 | 2 | 16 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 4 | 1 | 3 | 1 | 11 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 91 |
| Horse feces | Fibrobacter sp. UWH1 | 2 | 2 | 16 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 4 | 1 | 3 | 1 | 10 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 90 |
| Fibrobacter sp. UWH3 | 2 | 2 | 16 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 4 | 1 | 3 | 1 | 10 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 90 | |
| Fibrobacter sp. UWH4 | 2 | 2 | 13 | 3 | 8 | 5 | 1 | 3 | 4 | 2 | 3 | 5 | 1 | 3 | 1 | 12 | 0 | 3 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 87 | |
| Fibrobacter sp. UWH5 | 2 | 2 | 17 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 4 | 1 | 3 | 1 | 10 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 91 | |
| Fibrobacter sp. UWH6 | 2 | 2 | 16 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 4 | 1 | 3 | 1 | 10 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 90 | |
| Fibrobacter sp. UWH8 | 2 | 2 | 16 | 5 | 9 | 4 | 1 | 5 | 4 | 1 | 4 | 3 | 1 | 3 | 1 | 11 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 91 | |
| Fibrobacter sp. UWH9 | 2 | 2 | 17 | 5 | 8 | 4 | 1 | 5 | 4 | 1 | 4 | 5 | 1 | 3 | 1 | 11 | 1 | 3 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 93 | |
| Rhinoceros feces | Fibrobacter sp. UWR4 | 2 | 2 | 17 | 6 | 8 | 4 | 3 | 5 | 4 | 2 | 4 | 6 | 1 | 3 | 1 | 12 | 1 | 4 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 100 |
| Fibrobacter sp. UWR2 | 2 | 1 | 13 | 6 | 9 | 6 | 5 | 3 | 4 | 2 | 3 | 6 | 1 | 4 | 1 | 11 | 1 | 4 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 2 | 1 | 1 | 1 | 98 | |
| Fibrobacter sp. UWR1 | 2 | 2 | 17 | 6 | 8 | 4 | 3 | 5 | 4 | 2 | 4 | 6 | 1 | 3 | 1 | 12 | 1 | 4 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 100 | |
| Fibrobacter sp. UWR3 | 2 | 1 | 13 | 5 | 10 | 5 | 4 | 3 | 3 | 2 | 3 | 4 | 1 | 4 | 1 | 12 | 1 | 4 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 94 | |
| Tapri feces | Fibrobacter sp. UWT2 | 2 | 3 | 14 | 6 | 11 | 5 | 5 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 6 | 2 | 2 | 1 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 107 |
| Fibrobacter sp. UWT3 | 2 | 2 | 16 | 4 | 10 | 4 | 5 | 3 | 4 | 2 | 3 | 5 | 1 | 3 | 0 | 13 | 1 | 4 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 98 | |
| Colobus monkey feces | Fibrobacter sp UWCM | 2 | 1 | 14 | 5 | 10 | 5 | 4 | 3 | 4 | 2 | 3 | 5 | 1 | 4 | 1 | 14 | 1 | 4 | 2 | 2 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 99 |
| Elephant feces | Fibrobacter sp. UWEL | 2 | 2 | 18 | 5 | 9 | 4 | 2 | 5 | 4 | 2 | 4 | 5 | 1 | 3 | 1 | 9 | 0 | 4 | 2 | 1 | 0 | 3 | 2 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 95 |
| Rhesus monkey feces | Fibrobacter sp UWRM | 1 | 0 | 11 | 3 | 5 | 2 | 1 | 4 | 2 | 1 | 3 | 2 | 1 | 3 | 0 | 4 | 1 | 2 | 1 | 1 | 0 | 3 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 57 |
| Rat cecum | F. intestinalis ATCC 43854 | 1 | 0 | 9 | 3 | 6 | 2 | 1 | 4 | 1 | 1 | 3 | 2 | 0 | 2 | 0 | 3 | 1 | 2 | 2 | 0 | 0 | 3 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 52 |
| Pig cecum | Fibrobacter sp. UWS2 | 1 | 0 | 11 | 5 | 5 | 3 | 1 | 4 | 1 | 2 | 3 | 2 | 1 | 2 | 1 | 4 | 1 | 2 | 1 | 0 | 0 | 3 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 59 |
| Pig feces | Fibrobacter sp. UWS1 | 1 | 0 | 9 | 3 | 6 | 2 | 1 | 4 | 1 | 1 | 3 | 2 | 0 | 2 | 0 | 4 | 1 | 2 | 2 | 0 | 0 | 3 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 53 |
| Ostrich feces | F. intestinalis UWOS | 1 | 0 | 9 | 3 | 5 | 2 | 1 | 4 | 1 | 1 | 3 | 1 | 0 | 2 | 0 | 3 | 1 | 2 | 2 | 0 | 0 | 3 | 2 | 1 | 1 | 0 | 1 | 0 | 0 | 1 | 50 |
Higher putative specialization in cellulose than xylan degradation
In the genome of Fibrobacter sp. HC4, we identified 50 GHs in families known to play a role in cellulose degradation over the 91 GHs identified. These families belonged to GH5, GH8, GH9, GH10, GH16, GH26, GH39, GH44, GH45, GH51, and GH74. At least 24 putative cellulases were predicted by BLAST (Table 3). The protein sequences of 13 GH5, 2 GH8, 6 GH9, 1 GH44, 1 GH45, and 1 GH51 were similar to cellulase protein sequences. Thus, at least 26% (24/91) of GHs identified in Fibrobacter sp. HC4 genome might be involved in cellulolysis. Twenty contained a signal peptide, suggesting that the proteins may be transferred to the bacterial surface or secreted into the environment (Table 3). This hypothesis was supported by the predicted localization; the majority (14) had a predicted extracellular localization extracellular, while the others were periplasmic or outer membrane. Also, some enzymes allowing the binding of catalytic enzymes to the substrate (CBMs) were identified: two CBM 30 domains and the CBM 11 domain which are associated with GH51. Seven fibro-slime domain-containing proteins, additional proteins that facilitate cell adhesion for cellulose degradation, were identified in the genome of Fibrobacter sp. HC4. We identified one gene coding for cellobiose phosphorylase that can hydrolase cellobiose into one glucose-1-phosphate and one glucose. The repertoire of CAZymes suggests that F. succinogenes strain HC4 is highly adapted to degrade cellulose.
TABLE 3.
Putative enzymes involved in cellulose and xylan degradation identified from Fibrobacter sp. HC4 genome
| Putative activity | CAZy family domain | Protein size (aa) | Signal peptides (aa) | Predicted localization | Accession | ||
|---|---|---|---|---|---|---|---|
| Specificity | Gene_ID | ||||||
| Cellulase | MMG03_00486 | Cellulase | GH5_10 | 672 | 1–25 | Extracellular | MCL4100917.1 |

|
MMG03_00749 | Endoglucanase | GH9 | 610 | 1–29 | Periplasmic | MCL4101172.1 |
| MMG03_00934 | Endoglucanase 3 | GH5 | 683 | 1–23 | Outer membrane | MCL4101356.1 | |
| MMG03_00968 | Cellulase | GH9 | 2,220 | 1–17 | Periplasmic | MCL4101390.1 | |
| MMG03_00969 | Cellulase | GH9 | 1,967 | 1–24 | Extracellular | MCL4101391.1 | |
| MMG03_01016 | Endoglucanase CelG | GH5_4 | 481 | 1–19 | Extracellular | MCL4101438.1 | |
| MMG03_01018 | Cellulase | GH5_4 | 710 | 1–23 | Extracellular | MCL4101440.1 | |
| MMG03_01129 | Endoglucanase Z | GH5_2 | 457 | 1–22 | Extracellular | MCL4101551.1 | |
| MMG03_01137 | Endoglucanase | GH9 | 636 | 1–26 | Periplasmic | MCL4101559.1 | |
| MMG03_01445 | Cellulase | CBM30-CBM30-CBM11-GH51 | 1,060 | 1–23 | Periplasmic | MCL4101862.1 | |
| MMG03_01558 | Cellulase | GH5_4 | 621 | 1–20 | Extracellular | MCL4101972.1 | |
| MMG03_01601 | Endo-1,4-beta-D-glucanase Y | GH8 | 547 | 1–21 | Extracellular | MCL4102015.1 | |
| MMG03_01637 | Endoglucanase | GH5_52 | 347 | Extracellular | MCL4102050.1 | ||
| MMG03_01847 | Cellulase | GH5_2 | 713 | 1–17 | Extracellular | MCL4102253.1 | |
| MMG03_01848 | Endo-1,4-beta-D-glucanase Y | GH8 | 486 | 1–18 | Extracellular | MCL4102254.1 | |
| MMG03_01949 | Endoglucanase | GH9 | 586 | Extracellular | MCL4102353.1 | ||
| MMG03_02027 | Cellulase | GH5_25 | 665 | 1–38 | Extracellular | MCL4102427.1 | |
| MMG03_02119 | Cellulase | GH5_2 | 775 | 1–20 | Extracellular | MCL4102518.1 | |
| MMG03_02213 | Cellulase | GH5_3 | 1,016 | 1–17 | Extracellular | MCL4102610.1 | |
| MMG03_02352 | Cellulase | GH45 | 524 | 1–23 | Outer membrane | MCL4102746.1 | |
| MMG03_02562 | Cellulase | GH5_2 | 361 | 1–19 | Extracellular | MCL4102950.1 | |
| MMG03_02679 | Cellulase | GH5_55-CBM4 | 541 | Extracellular | MCL4103066.1 | ||
| MMG03_02930 | Non-processive endocellulase | GH9 | 617 | Periplasmic | MCL4103314.1 | ||
| MMG03_02934 | Cellulase | GH44 | 604 | 1–27 | Periplasmic | MCL4103318.1 | |
| Phosphorylase | MMG03_01346 | Cellobiose phosphorylase | GH94 | 837 | Cytoplasmic | MCL4101765.1 | |
| Xylanase | MMG03_00112 | Endo-1,4-beta-xylanase | GH10 | 632 | 1–17 | Periplasmic | MCL4100547.1 |

|
MMG03_00659 | Reducing-end xylose-releasing exo-oligoxylanase Rex8A | GH8 | 370 | Extracellular | MCL4101087.1 | |
| MMG03_00765 | Endo-1,4-beta-xylanase | GH10-CBM6 | 642 | 1–20 | Extracellular | MCL4101188.1 | |
| MMG03_00766 | Beta-xylanase | GH10-CBM6 | 614 | 1–23 | Extracellular | MCL4101189.1 | |
| MMG03_00780 | Xylanase | GH30_8-CBM6 | 712 | 1–28 | Extracellular | MCL4101203.1 | |
| MMG03_00781 | Xylanase | GH43_16-CBM6-CBM6 | 755 | 1–22 | Extracellular | MCL4101204.1 | |
| MMG03_02381 | Reducing end xylose-releasing exo-oligoxylanase | GH8 | 427 | Extracellular | MCL4102774.1 | ||
| MMG03_02382 | Endo-1,4-beta-xylanase | GH10 | 551 | 1–18 | Periplasmic | MCL4102775.1 | |
| MMG03_02831 | Xylanase | GH43_29-CBM6-CBM6 | 692 | 1–20 | Extracellular | MCL4103217.1 | |
| β-xylosidase | MMG03_01122 | Xylanase or xylosidase | GH11-CBM4 | 576 | 1–23 | Extracellular | MCL4101544.1 |
| MMG03_00778 | Xylan 1,4-beta-xylosidase | GH43_10-CBM6 | 788 | 1–22 | Extracellular | MCL4101201.1 |
Although Fibrobacter sp. HC4 is specialized in cellulose degradation and does not grow on pure xylan, or xylose, enzymatic activities were detected with xylan using Congo Red. Thus, we looked at GHs potentially involved in xylan degradation. Forty-six GHs were identified as belonging to families which contain xylanase function in the genome of strain HC4: GH3, GH5, GH8, GH10, GH11, GH16, GH30, and GH43. Of these enzymes, two GH8, four GH10, one GH30, and two GH43 sequences were similar to those of xylanases and one GH43 to a xylan 1,4-beta-xylosidase by Blast (Table 3). The family GH11 contains only xylosidase and xylanase function according to CAZy database (http://www.cazy.org/). Of these putative xylanases, six were associated with CBM 6. At least 11 putative xylanases or xylosidases were predicted, supporting a xylanolytic activity for Fibrobacter sp. HC4. Nine proteins contained a signal peptide, and nine have a predicted extracellular localization, whereas only two would be periplasmic (Table 3).
Metabolic pathway for succinate and acetate production
All genes that allow the conversion of glucose into phosphoenoylpyruvate (PEP; glycolysis pathway) were identified. The genome of Fibrobacter sp. HC4 contained genes coding PEP carboxykinase and pyruvate kinase to transform PEP in pyruvate or in oxaloacetate, respectively (Fig. 4). The gene for pyruvate carboxylase, that can transform pyruvate in oxaloacetate, was also identified. The metabolism of pyruvate by Fibrobacter sp. HC4 could lead to acetate production as genes for pyruvate-flavodoxin oxidoreducatase, acetyl-coenzyme A synthetase, phosphate acetyltransferase, and acetate kinase were identified in the genome. Enzymes for succinate production from pyruvate, i.e., malate dehydratase, fumarate hydratase, and succinate dehydrogenase, were also identified from the genome of Fibrobacter sp. HC4. Genes of the nonoxidative branch of the pentose phosphate pathway were identified but not those of the oxidative branch. No xylose metabolizing enzymes (xylose isomerase, xylulokinase, and xylose permease) or transporters (D-xylose transport system substrate-binding protein, D-xylose transport system permease protein, and D-xylose transport system ATP-binding protein) were predicted either.
Fig 4.

Metabolism of glucose to succinate and acetate.
Functional activities of cellulase, cellobiosidase, and xylanase
To determine CMCase and xylanase activities, reduced carbohydrates released from CMC or beechwood xylan, respectively, were quantified. To compare these activities of Fibrobacter sp. HC4 with the type strain, the same assays were performed using F. succinogenes S85 (Fig. 5). For both strains, the fibrolytic activities were mostly extracellular. The extracellular CMCase activity of Fibrobacter sp. HC4 was higher than that of strain S85, while similar intracellular CMCase activities were measured for both strains. Consequently, Fibrobacter sp. HC4 had higher total CMCase activity than F. succinogenes S85 (P = 0.0043). Similarly, the extracellular xylanase activity of Fibrobacter sp. HC4 was higher than that of strain S85, while it was the opposite for the intracellular activity. Therefore, Fibrobacter sp. HC4 showed the highest capacity to degrade cellulose and xylan, consistent with the considerable number of putative cellulases and xylanases we found. In addition, the CMCase activity of Fibrobacter sp. HC4 was almost twice as important as that of xylanase, consistent with the finding that the strain possesses twice as many putative cellulases as xylanases.
Fig 5.

Enzyme-specific activities of Fibrobacter sp. HC4, the new strain, and F. succinogenes S85, the type strain, grown on cellulose (means and SD). (a) CMCase, (b) cellobiosidase, and (c) xylanase activities and schematic composition of substrates and target site for each enzyme class. P-values indicate the differences in extra or intracellular activities between strain HC4 and strain S85.
Similar total cellobiosidase activities were measured for Fibrobacter sp. HC4 and S85, although the intracellular activity of strain S85 was higher. GHs with cellobiosidase function have not been identified.
In vitro hay degradation enhanced in fecal microbial ecosystem in the presence of Fibrobacter sp. HC4 strain
To investigate the probiotic potential of Fibrobacter sp. HC4 in the case of large intestinal acidosis, we studied the effect of Fibrobacter sp. HC4 on the in vitro hay degradation, a substrate rich in cellulose, by an equine fecal microbial ecosystem. The feces used as inoculum contained 2.18 × 108 CFU/g of total cultivable bacteria, with only 2 × 105 MPN/g cellulolytic bacteria after the dietary challenge to the horse (Fig. 6a). The addition of Fibrobacter sp. HC4 at 7.7 × 107 MPN/mL increased gas production compared to the control without Fibrobacter sp. HC4 (P < 0.0001), from the first hours up to 36 h (Fig. 6b and c). The stationary phase started from 72 h in HC4 bottles, whereas gas production increased continuously until 150 h in the control bottles, without reaching the amount produced with the HC4-enriched ecosystem at the end of the test. Additionally, when Fibrobacter sp. HC4 was added to the fecal inoculum, more volatile fatty acids (VFAs) were produced compared to the control (P < 0.0001; Fig. 6d). Acetate, propionate, and butyrate concentrations were higher at 24 and 48 h in the bottles containing HC4 (Fig. 6d). In accordance, pH decreased more rapidly in the bottles containing Fibrobacter sp. HC4 than in control bottles (P < 0.0001; Fig. 6e). No lactate was detected. Dry matter disappearance was higher with the addition of Fibrobacter sp. HC4 than in control at 48 and 72 h (P = 0.0034; Fig. 6f). After 48 h, the addition of Fibrobacter sp. HC4 to the fecal inoculum did not alter the cultivable anaerobic bacterial populations (Fig. 6g).
Fig 6.

In vitro digestibility and gas production. (a) Composition for 1 g of added feces. (b) In vitro gas production kinetics. (c) Gas production rate every 12 h. (d) In vitro composition in VFAs. (e) pH evolution. (f) In vitro %dry matter disappearance (DMD) evolution. (g) Cultivable anaerobic bacterial populations at the end of in vitro fermentation. *, P < 0.1; **, P < 0.01; ***, P < 0.0001 HC4 vs control at each measurement point. Means and SD are presented.
DISCUSSION
To date, few cellulolytic bacteria inhabiting the large intestine of mammals have been isolated. Fibrobacter sp. HC4 is one of these rare strains to be deposited and cultivated. Our study provides new data on its metabolism and fibrolytic capacity. The isolation phase and all tests using Fibrobacter sp. HC4 were carried out under strictly anaerobic conditions. In fact, cellulolytic bacteria are considered to be extremely sensitive to O2 (19, 24–26). However, to our knowledge, their sensitivity to O2 has never been quantified. The sensitivity of bacteria to O2 is usually estimated by plating on Petri plates under aerobic conditions. In practice, it is measured in practice by the ability of microorganisms to grow on agar plates after a given period of exposure to air (27–29) or different percentages of O2 (30). Among strict anaerobic bacteria, several levels of sensitivity to O2 are distinguished, up to those called “Extremely Oxygen Sensitive” (EOS) (27, 30–32). Since Fibrobacter sp. HC4 does not grow on agar on Petri plates, we tested its O2 sensitivity in a liquid medium for the first time. Fibrobacter sp. HC4 did not grow in a non-degassed medium and did not resume growth when returned to an anaerobic environment after an hour in aerobic conditions. Therefore, we assumed that Fibrobacter sp. HC4 acted as an EOS bacterium. To cultivate it, we hypothesized that O2 in the liquid medium was negligible due to the presence of cysteine and of some sulfur compounds in the growth media. These compounds are antioxidant and could contribute to an early reduction of the aerobic medium, which is critical to support EOS bacteria (33). In addition, the Hungate tubes were saturated with CO2 to minimize the presence of O2. In support of these observations, we found that Fibrobacter sp. HC4 had genes that demonstrated its ability to survive under strict anaerobic conditions. We found in the genome of Fibrobacter sp. HC4 the enzymatic arsenal capable of reducing fumarate to succinate. This reduction was reported to be coupled with the oxidative decarboxylation of pyruvate to acetyl-CoA, allowing F. succinogenes to survive under anaerobic conditions (34). Malate dehydratase was present in the genome of the Fibrobacter sp. HC4. Thus, the NADH formed during the glycolysis pathway could be oxidized for the reduction of oxaloacetate in malate. Then, the fumarate hydratase identified in the genome of the Fibrobacter sp. HC4 could convert malate to fumarate, which is finally reduced to succinate the last acceptor of electrons generated by the oxidation of pyruvate (34, 35). In the metabolism of Fibrobacter sp. HC4, this coupling could be achieved by two oxidation-reduction reactions involving the cytochrome b556 subunit of the succinate dehydrogenase and the succinate dehydrogenase flavoprotein subunit that we identified (34, 36, 37).
Our results highlighted the high specialization of Fibrobacter sp. HC4 in cellulolysis. The genome of Fibrobacter sp. HC4 possesses 222 putative CAZymes of which many GHs can target cellulose. Fibrobacter sp. HC4 harbored 16 GH5, a multifunctional family, similar to other strains of F. succinogenes isolated from horses, and 13 protein sequences were close to those with cellulase functions. The majority of the putative cellulases found in the genome of HC4 belong to families GH5, GH8, and GH9 families similar to the endoglucanases previously identified in Fibrobacter genus in the literature (3, 23). In addition, the other enzymes we found in the genome of HC4 belonged to the same families (GH44, GH45, and GH51 families) as those previously reported for the Fibrobacter genus (3, 23). We confirmed the genomic potential of Fibrobacter sp. HC4 to degrade cellulose in further assays demonstrating the capacity of Fibrobacter sp. HC4 to excrete CMCase, to consume crystalline cellulose, as well as cellobiose and glucose, the two sugars released by cellulose degradation. In addition, we measured that hay degradation was improved in vitro when Fibrobacter sp. HC4 was added to a horse fecal inoculum. Fibrobacter sp. HC4 alone was able to degrade one-third of the DM in the hay, which had been washed to reduce the simple sugar content, after 150 h of culture. This confirmed the effective fibrolytic activity. Compared to the type strain F. succinogenes S85 originating from bovine rumen, Fibrobacter sp. HC4 showed higher CMCase activity. The major extracellular activities we measured were in accordance with predicted localizations of identified cellulases, with the majority predicted to be extracellular. These activities could also originate from outer membrane vesicles produced and released by F. succinogenes, which are enriched in CAZymes. The existence of these vesicles has been demonstrated in F. succinogenes S85 (38). The vesicles contain fibro-slime proteins, cellulases, and hemicellulases and are released from the bacterial cells to target plant cell walls, facilitating access to cellulose. Complex with fibro-slimes anchored to the peptidoglycan of the outer membrane would facilitate the adhesion of F. succinogenes S85 to cellulose (39), and proteins containing fibro-slime domains already identified in other genomes of Fibrobacter (23) have also been identified in the genome of Fibrobacter sp. HC4. The initial step of cellulose adhesion has a crucial role in the ability of F. succinogenes to degrade it (40). Cellulases are often associated with one or more CBMs that are noncatalytic proteins or internal peptides attached to the catalytic domain. In the genome of Fibrobacter sp. HC4, we identified two CBM 30 domains and the CBM 11 domain that were associated with GH51. CBM 11 and 30 were identified as binding to cellulose with F. succinogenes S85 (3). It was previously observed with other Fibrobacter strains that the gene encoding the endoglucanase GH51 carried the CBM 11 (23) and CBM 30 modules (3). Cellulose degradation by F. succinogenes could be mainly due to the action of endoglucanases. Similar total cellobiosidase activity was measured in equine and bovine strains, although no cellobiosidase (or exoglucanase) was identified in the genome of F. succinogenes S85 (3, 23) and not found in the genome of Fibrobacter sp. HC4. Although GH48 is present in cellulolytic Ruminococcus strains, no exoglucanases have been identified in Fibrobacter strains (23, 41, 42).
Like the other strains of F. succinogenes, strain HC4 possessed the enzymes to degrade xylan to access cellulose but not the enzyme to utilize the carbohydrates released from this degradation (3, 42). At least 10 putative xylanases were predicted in the genome of Fibrobacter sp. HC4 of which the majority is associated with CBM 6. It is one of the CBM families associated with the degradation of plant cell walls (43). The putative xylanases identified belonged to families GH8, GH10, GH30, and GH43. In the literature, hemicellulases identified from Fibrobacter belong to the GH2, GH3, GH10, GH11, GH18, GH43, and GH53 families (23). As xylan is the main source of hemicelluloses, only xylanases and xylosidases were investigated in this study. Xylose, the simple carbohydrate released by xylan degradation, can be used by other micro-organisms of the large intestinal microbiota. The degradation of other hemicelluloses (glucuronoxylan, arabinoxylan, glucomannan, and xyloglucan) by Fibrobacter sp. HC4 could be studied to verify the release of other sugars, i.e., mannose, galactose, rhamnose, and arabinose, which could also be utilized by other micro-organisms. In addition to facilitating access to cellulose for F. succinogenes, this hemicellulolytic capacity is of interest for the proper functioning of the intestinal microbiota.
In addition to releasing fermentable substrates used by other members of the equine gut microbiota via fiber degradation (8, 44), succinate and acetate produced by Fibrobacter sp. HC4 could also be used by other microorganisms. Succinate is an acid that acts as an intermediary in the metabolism of intestinal microbiota. In the ruminal (45) and human (46) microbiota, some bacterial populations used the succinate pathway to produce propionate. Acetate is also an intermediate used by butyrate-producing bacteria.
Beyond the equine species, promoting cellulose degradation in the large intestine and encouraging the production of VFA is a major health issue in many mammals. When Fibrobacter sp. HC4 was added to an equine fresh fecal microbial ecosystem, a higher total VFAs concentration and a higher disappearance of hay DM were observed compared to the ecosystem alone. The generation of VFAs resulting from the complex carbohydrates microbial degradation in the horse large intestine covers at least 50% of the host energy requirements (5). In horses, Fibrobacter sp. HC4 could be beneficial in case of large intestinal acidosis when cellulolytic bacteria concentrations are depressed.
In conclusion, using a multiple approach for investigating the capacity of strain Fibrobacter sp. HC4 to utilize cellulose, we confirmed its high specialization for adhering to cellulose fibers, secreting specific endoglucanases and producing high amounts of succinate and acetate. In addition, Fibrobacter sp. HC4 boosted the horse’s microbial ecosystem for more efficient hay utilization after a common challenge selected to reduce cellulolytic bacteria concentrations. Supplementation of such cellulolytic bacteria could be considered in the future for restoring large intestinal balance not only in horses but also in other mammals. This requires further in vivo studies to assess the effects of Fibrobacter sp. HC4 on the large intestinal microbiota and the host.
MATERIALS AND METHODS
The materials and methods used for the isolation, identification, and characterization of the new strain are shown in Fig. 7.
Fig 7.

Materials and methods for the isolation, identification, and characterization of the new strain.
Sample collection and enrichment of cellulose-degrading bacteria
The cecum digesta we used was collected from one cecum-cannulated horse as part of another experimental trial (47). All experimental procedures were evaluated and approved by the local ethical committee (Comité d’Ethique de l’Expérimentation Animale Grand Campus Dijon). In a sterile 25 mL flask saturated in CO2 to maintain anaerobic conditions, the sample was immediately stored at 38°C. Samples were transported back to the laboratory and inoculated within 1 h of collection. One gram of the collected sample was 10-fold diluted in an anaerobic mineral solution (48). Serial 10-fold dilutions down to 10−8 were carried out. All liquid and solid media were prepared, dispensed, and inoculated using strictly anaerobic techniques (24). Each dilution was inoculated in the specific medium for cellulolytic bacteria developed by Halliwell and Bryant (49) and modified by Julliand et al. (11) to be better adapted to the ecology of bacteria from the equine gut. All chemicals were purchased from Sigma-Aldrich unless otherwise specified, and equine cecal and bovine ruminal fluids were obtained from the filtrate of the digestive contents concerned. This medium contained, per liter: NaHCO3 59.51 mM, K2HPO4 2.58 mM, KH2PO4 3.30 mM, (NH4)2SO4 6.81 mM, NaCl 15.40 mM, MgSO4 0.75 mM, CaCl2 0.81 mM, resazurin 0.0044 mM, bovine ruminal fluid 200 mL/L, equine cecal fluid 200 mL/L, and cysteine hydrochloride 6.35 mM, filled to 1 L with distilled water. A piece of Whatman No. 1 filter paper cellulose (8 × 1 cm) was added to each tube before the addition of the medium (8 mL per tube). The medium tubes were then sterilized for 20 min at 120°C.
Cellulolytic bacteria isolation
After incubation at 38°C in the adapted medium and under anaerobic atmosphere for 7 days, two isolation methods were used from the highest dilution showing filter paper degradation: first, the roll tubes method described by reference (50) and then the enrichment method recently published by reference (15). Isolated strains were examined for purity, morphology and Gram staining by phase contrast microscopy in semi-synthetic medium (51) with 2% Sigmacell or piece of filter paper cellulose grown culture.
Genetic identification (sequencing of 16S rRNA genes) of cellulolytic isolates
Bacterial DNA was extracted using a DNA extraction kit (Instagene Matrix – 732–6030, Bio-rad) according to the manufacturer’s instructions. The concentration was measured using a BioPhotometer (Eppendorf). The 16S rRNA gene was amplified with universal primers 27f (5′-AGAGTTTGATCMTGGCTCAG-3′) and 1492r (5′-TACGGHTACCTTGTTACGACTT-3′) (52). PCR reactions were done in a total reaction volume of 50 µL containing: Taq polymerase buffer 10× MgCl2 (Core Kit 25, MP Biomedicals), 0.4 µL of dNTPs (Core Kit 25, MP Biomedicals), 1 µL of each primer, 5 µL of Taq polymerase (Core Kit 25, MP Biomedicals), and 5 µL of DNA. DNA amplification was achieved using a Thermal Cycler (CFX 96 Real Time System, Bio-Rad, Hercules, California, USA). The following program was used: denaturation for 4 min at 95°C; then 35 cycles consisting of 1 min at 94°C, 1 min at 55°C, and 2 min at 72°C; and a final step consisting of 15 min at 72°C and 30 min at 30°C (52). Amplification products were confirmed by gel electrophoreses with 2% agarose gel containing 0.002% gel red (GelRed, FluoProbes, INTERCHIM) in tris-borate-EDTA buffer (TBE) and stained with gel loading blue (GelPilot, QIAGEN). Subsequently, PCR products were sequenced by Genewiz using the Sanger method. Sequences were analyzed using BLAST from the NCBI website.
Enzymatic activity of isolated cellulolytic bacteria and F. succinogenes S85
These analyses were performed with our isolated strain and also with the reference strain from bovine rumen: F. succinogenes S85. The CMCase, xylanase, and fibrolytic activities were detected with Congo red. The 48-h cultures of isolated bacteria and of F. succinogenes S85 were deposited (10 µL) on agar plate with 0.5% of CMC (CMC sodium salt, medium viscosity, Sigma-Aldrich), with 0.3% of dried ground hay (neutral detergent fiber [NDF], 759.27, and acid detergent fiber [ADF], 480.39 g/kg DM) or with 0.3% of beechwood xylan (Megazyme International Ireland Ltd.) in an anaerobic glove box with a gas phase of 5% H2, 10% CO2, and 85% N2. After incubation at 38°C for 24 h, agar plates were strained with Congo red 0.2% to detect fibrolytic activity (53).
Bacterial enzymes were extracted from 48 h isolated strain cultures on filter paper medium centrifuged at 12,000 g for 20 min. The supernatants were frozen at −20°C. The cell pellets were suspended in 1/10 vol of phosphate buffer (pH 6.8), and membranes were weakened by two freeze-thaw cycles and then broken by sonication. The supernatants containing intra-cellular enzymes were harvested after centrifugation at 5,000 g for 2 min and were kept frozen at −20°C.
The CMCase and xylanase activities were assessed by incubating enzyme preparations for 20 min at 39°C in phosphate buffer (pH 6.8) with 1% of respectively CMC or xylan from beechwood (54). The reaction was stopped by adding 3,5 dinitrosalicylic acid (Sigma-Aldrich) and boiling for 5 min to measure the reducing sugars released by the Miller method (55) with glucose as the standard.
Cellobiosidase activity was measured by assaying the amount of para-nitrophenol released after incubation of enzyme extracts and para-nitrophenol substrates (Sigma-Aldrich) in solutions (5 mM) in phosphate buffer (pH 6.8) for 20 min at 39°C with shaking at 900 rpm (PHMP Microplate Thermostatic Shaker, Grant-bio, Royston, UK). The enzymatic reaction was stopped by increasing the pH by adding 0.4 M glycine/soda solution, pH 10.8. The amount of para-nitrophenol released was measured spectrophotometrically at 420 nm using a standard solution of para-nitrophenol (Sigma-Aldrich). Soluble proteins were determined using bovine serum albumin (Sigma-Aldrich) as the standard (56).
All cultures were performed in triplicate to determine the CMCase, cellobiosidase, and xylanase activities. Results are given in nanokatals (nkat) of glucose or xylose equivalent/mg of protein. One kat corresponds to 1 mol produced per second.
A mixed procedure on SAS (version 9.3, SAS Institute Inc., Cary, NC) was used to assess the differences in specific enzyme activities between strain HC4 and strain S85. The model included replicate as a random effect. The LS-means were compared between bacterial strain using the Tukey-Kramer method. Differences were considered significant at P < 0.05.
O2 sensitivity of isolated cellulolytic bacteria and F. succinogenes S85
To determine O2 sensitivity, 1 mL of 48 h cultures of isolated bacteria and 1 mL of 48 h cultures of F. succinogenes S85 were each inoculated in aerobic adapted medium described above (without adding CO2) and incubated at 38°C for 2 min, 1 h, 5 h, and 24 h with glucose or cellulose as substrate. After these incubation times, 1 mL of each condition was inoculated in the same medium in anaerobic conditions (deaerated under CO2) with cellulose as substrate. All cultures were performed in triplicate. Growth was verified by the filter paper degradation by bacteria and by microscopic observations after Gram staining.
Substrates consumed and degraded
The isolated strain was cultured in Lowe semi-synthetic medium (51) in which cellobiose was present or substituted by xylose, arabinose, glucose, fructose, mannose, rhamnose, maltose, saccharose, lactose, and raffinose (2 g/L) or by xylan from beechwood, Sigmacell (7 g/L). All cultures were performed in 10 mL of medium in Hungate tubes. Growth was monitored by optical density (OD) at 600 nm (CECIL 1011, Cecil Instrumentation Services Ltd, Cambridge, UK) every 24 h, and pH was measured after 120 h, at the end of growth (CyberScan pH 510, Eutech Instrument, Strasbourg, France). All cultures were performed in triplicate. If OD increased and pH decreased over time, growth on a substrate of the isolated strain was considered positive.
To observe whether the isolated strain degraded a complex fiber source, the strain was anaerobically cultivated on 1 g of hay previously washed and grounded to mimic mastication and foregut digestion in 100 mL of Lowe medium (51) in flasks with gas impermeable butyl stopper and aluminum crimp seals, and DM disappearance was measured after 150 h at 38°C. The hay was first washed at 60°C in the washing machine and dried at 80°C for 24 h before being ground. After those treatments, the hay contained (g/kg DM) the following: OM: 948, CP: 67.68, NDF: 759.27, ADF: 480.39, and water soluble carbohydrates (WSC): 71.21. All cultures were performed in triplicate. If DM disappearance increased over time, growth on the hay of isolated strain was considered positive.
Metabolic product analysis
Fermentation products present in culture with filter paper were quantified after 48 h. Acetate (C2), propionate (C3), butyrate (C4), and valerate (C5) concentrations were assayed by gas chromatography (Gas chromatograph—flame ionization detector Clarus 500, PerkinElmer, Courtaboeuf, France) (57). Succinate, lactate, and formate concentrations were measured spectrophotometrically at 340 nm (MRX revelation, Dynatech Laboratories, Guyancourt, France) in microplate using an enzymatic colorimetric method [Succinic acid Assay Kit, Formic Acid Assay kit, D-/L-Lactic Acid (D-/L-Lactate) (Rapid) Assay Kit, Megazyme International Ireland Ltd.]. All analyses were performed in duplicate on three different cultures. If metabolite concentration increased over time, the production of succinate, lactate, or formate by isolated strain was considered positive.
Characterization of the isolated strain’s growth on cellulose
The MPN (58) was determined using the modified Lowe medium after 15 days of incubation at 38°C (dilutions representing 10−6, 10−7, 10−8 and 10−9 of feces).
Phylogenetic trees of identified bacterial species
From full-length 16S rRNA gene sequence (accession number: OP018198) obtained from the whole genome (accession number: JALCZV000000000), a maximum likelihood analysis hierarchical clustering in MEGA 7.0 using 1,000 bootstrap replicates produced a dendrogram. Prior to the clustering analysis, the sequences were aligned in MEGA 7.0 with default parameters and cut off to an equal number of nucleotide positions (1,268 sites). A 16S rRNA gene sequence from B. fragilis type strain NCTC 9343 was included as an outgroup.
Investigation of carbon and nitrogen metabolism genes
Genes involved in carbon and nitrogen metabolism were investigated and identified, either by Genewiz, Germany, from their databases, or using sequences recorded in databases of strains close to the isolated strain (https://www.uniprot.org/ and https://www.ncbi.nlm.nih.gov/).
Investigation of CAZymes
The search for CAZymes from protein sequences of the whole genome was performed by Glycogenomics team of the Architecture and Function of Biological Macromolecules (AFMB) laboratory of Aix-Marseille University. Protein sequences were analyzed and compared to previously annotated proteins and saved in the internal CAZy database. The comparisons were performed by a script based on the modular structure of CAZymes and their classification. The annotation scripts compared each protein to the complete proteins in the CAZy database using BlastP (version 2.3.0+). Sequences with 100% coverage and more than 50% sequence identity to a sequence already in the CAZy database were automatically annotated with the same modularity. The search of GH was also done for other strains of Fibrobacter whose genome was available on public databases (NCBI and Joint Genome Institute [JGI]) by AFMB.
Protein sequences of GHs identified by AFMB were analyzed using BLAST from the NCBI website (https://blast.ncbi.nlm.nih.gov/Blast.cgi) or UniProt website (https://www.uniprot.org/blast/) to identify cellulase and xylanase putative functions of the new strain. We considered that two proteins could have the same function when their sequences were more than 60% identical. The site http://www.cazy.org/ was used to research the different functions of GH families (59).
To check whether the proteins identified as cellulase or xylanase contained signal peptides, we used SignalP 6.0 (https://services.healthtech.dtu.dk/services/SignalP-6.0/). SignalP 6.0 is a machine-learning model that detects signal peptide types and is applicable to metagenomic data (60). DeepLocPro 1.0 is a predictor of the subcellular localization of prokaryotic proteins (61), and it was used to predict the localization of the identified cellulases and xylanases.
In vitro digestibility and gas production
The effect of adding the strain HC4 to a complete fecal microbiobial ecosystem of horses was studied in vitro. The fecal inoculum was obtained from the feces of three horses submitted to an abrupt change of diet from a high-fiber to a high-starch diet, so there were few cellulolytic bacteria in the sampled fecal ecosystem (8) (Fig. 6a). Fifty milliliters of medium based on Lowe medium (51) and containing in final concentrations KCl 8.05 mM, NaCl 10.27 mM, MgSO4 4.15 mM, CaCl2 0.90 mM, NH4Cl 4.54 mM, KH2PO4 5.00 mM, Na2CO3 3.77 mM, NaHCO3 9.76 mM, resazurin 0.0044 mM, cysteine hydrochloride 3.17 mM, trace element solution (51) 10 mL, resazurine (0.1%) 1 mL, and 1 g of sterile dried ground hay as main carbon source were incubated with 1 g of fecal inoculum and 10 mL of selected cellulolytic bacteria culture. All chemicals were purchased from Sigma-Aldrich. The washed dried ground hay used was the same as that described above, and it was used as a substrate because it is rich in fiber. The hay and culture medium were sterilized by autoclaving to avoid the presence of uncontrolled initial microbiota.
The concentration of total cultivable anaerobic bacteria in the fecal inoculum was determined using four replicate roll tubes (50) prepared with dilutions representing 10−5, 10−6, and 10−7 mL of intestinal contents on a non-selective modified complete agar medium (11, 62) after 48 h of incubation at 38°C. Concentrations of lactate-utilizing bacteria were determined by the same method on a selective medium (63) after 48 h of incubation at 38°C. Concentrations of starch-utilizing bacteria were determined on a modified selective medium (51) containing 1% (wt/vol) soluble starch (ThermoFisher) as the main energy source after 48 h of incubation at 38°C. The MPN (58) of cellulolytic bacteria (dilutions representing 10−3, 10−4, 10−5, and 10−6 of feces) was determined using the modified broth medium containing filter paper (11, 49) after 15 days of incubation at 38°C. The same method was used to determine the MPN of the isolated cellulolytic bacteria inoculum with dilutions representing 10−7, 10−8, and 10−9 of bacterial culture.
Two controls were carried out in vitro: one positive with hay and fecal inoculum without addition of strain HC4 and one negative with fecal inoculum only. Each condition was replicated three times and incubated at 38°C. Fermentation was monitored following gas production by a pressure transducer in the headspace of the bottles at 3, 8, 12, 24, 30, 36, 48, 54, 60, 72, 80, 97, 123, and 150 h. After each measurement, the gas produced was extracted from each bottle using a syringe until the transducer was back to zero to avoid overpressure in the bottles. The pressure measures from same bottles were used to calculate the gas accumulation at each time and to know the gas production between two times. Every 24 h during the incubation, pH (Cyberscan pH 510 pH-meter, Eutech Instrument, Strasbourg, France), DM disappearance, and concentrations of VFAs (gas chromatography coupled with FID, Clarus 500, Perkin Elmer, Courtaboeuf, France) and lactate [Megazyme, D-/L-Lactic Acid (D-/L-Lactate) (Rapid) Assay Kit, Megazyme International Ireland Ltd.] were measured on three bottles for each condition.
A mixed procedure on SAS (version 9.3, SAS Institute Inc., Cary, NC) was used to evaluate the effect of HC4 strain addition and fermentation time on pressure, pH, DM disappearance, and concentrations of VFAs. The model included fermentation time, the addition of HC4 strain or not, and the interaction between time and presence of HC4 strain. The model also included replicates as a random effect and times as repeated measures. The LS-means were compared between bacterial strain using the Tukey-Kramer method. Differences were considered significant at P < 0.05.
ACKNOWLEDGMENTS
We thank Evelyne Forano, UMR MEDIS, INRAE, Univ. of Clermont Auvergne, for kindly providing us with the F. succinogenes S85 reference strain. We would also like to thank Aline Bonnotte, Plate-forme DImaCell, Institute AgroDijon, INRAE, Univ. Bourgogne FrancheComté, for SEM photographs. We acknowledge the CAZy bioinformatics facility at AFMB laboratory for the curation of CAZyme.
This work is part of the project Probio+: production et fonctionnalités innovantes 2019, supported by the Conseil Régional de Bourgogne Franche-Comté and the European Union through the PO FEDER-FSE Bourgogne 2014/2020 programs; European Regional Development Fund (BG0026784).
Contributor Information
Alicia Froidurot, Email: alicia.froidurot@lab-to-field.com.
Edward G. Dudley, The Pennsylvania State University, University Park, Pennsylvania, USA
DATA AVAILABILITY
The complete genome of Fibrobacter sp. HC4, sequenced and annotated by Genewiz, has been deposited in NCBI under accession number JALCZV000000000 and BioProject number PRJNA812370 (22).
REFERENCES
- 1. Flint HJ, Scott KP, Duncan SH, Louis P, Forano E. 2012. Microbial degradation of complex carbohydrates in the gut. Gut Microbes 3:289–306. doi: 10.4161/gmic.19897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Froidurot A, Julliand V. 2022. Cellulolytic bacteria in the large intestine of mammals. Gut Microbes 14:2031694. doi: 10.1080/19490976.2022.2031694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Suen G, Weimer PJ, Stevenson DM, Aylward FO, Boyum J, Deneke J, Drinkwater C, Ivanova NN, Mikhailova N, Chertkov O, Goodwin LA, Currie CR, Mead D, Brumm PJ. 2011. The complete genome sequence of Fibrobacter succinogenes S85 reveals a cellulolytic and metabolic specialist. PLoS One 6:e18814. doi: 10.1371/journal.pone.0018814 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Ze X, Le Mougen F, Duncan SH, Louis P, Flint HJ. 2013. Some are more equal than others. Gut Microbes 4:236–240. doi: 10.4161/gmic.23998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Julliand V, Grimm P. 2016. Horse species symposium: the microbiome of the horse hindgut: history and current knowledge. J Anim Sci 94:2262–2274. doi: 10.2527/jas.2015-0198 [DOI] [PubMed] [Google Scholar]
- 6. Goachet AG, Varloud M, Jacotot E, Julliand V.. 2004. Effet de l’administration orale de trois anthelminthiques sur la microflore colique du cheval.
- 7. Harlow BE, Lawrence LM, Flythe MD. 2013. Diarrhea-associated pathogens, lactobacilli and cellulolytic bacteria in equine feces: responses to antibiotic challenge. Vet Microbiol 166:225–232. doi: 10.1016/j.vetmic.2013.05.003 [DOI] [PubMed] [Google Scholar]
- 8. Julliand V, Grimm P. 2017. The impact of diet on the hindgut microbiome. J Equine Vet Sci 52:23–28. doi: 10.1016/j.jevs.2017.03.002 [DOI] [Google Scholar]
- 9. Collinet A, Grimm P, Julliand S, Julliand V. 2021. Multidimensional approach for investigating the effects of an antibiotic–probiotic combination on the equine hindgut ecosystem and microbial fibrolysis. Front Microbiol 12:1–14. doi: 10.3389/fmicb.2021.646294 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Lin C, Stahl DA. 1995. Taxon-specific probes for the cellulolytic genus Fibrobacter reveal abundant and novel equine-associated populations. Appl Environ Microbiol 61:1348–1351. doi: 10.1128/aem.61.4.1348-1351.1995 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Julliand V, de Vaux A, Millet L, Fonty G. 1999. Identification of Ruminococcus flavefaciens as the predominant cellulolytic bacterial species of the equine cecum. Appl Environ Microbiol 65:3738–3741. doi: 10.1128/AEM.65.8.3738-3741.1999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Daly K, Stewart CS, Flint HJ, Shirazi-Beechey SP. 2001. Bacterial diversity within the equine large intestine as revealed by molecular analysis of cloned 16S rRNA genes. FEMS Microbiol Ecol 38:141–151. doi: 10.1111/j.1574-6941.2001.tb00892.x [DOI] [Google Scholar]
- 13. Daly K, Shirazi-Beechey SP. 2003. Design and evaluation of group-specific oligonucleotide probes for quantitative analysis of intestinal ecosystems: their application to assessment of equine colonic microflora. FEMS Microbiol Ecol 44:243–252. doi: 10.1016/S0168-6496(03)00032-1 [DOI] [PubMed] [Google Scholar]
- 14. Hastie PM, Mitchell K, Murray J-AMD. 2008. Semi-quantitative analysis of Ruminococcus flavefaciens, Fibrobacter succinogenes and Streptococcus bovis in the equine large intestine using real-time polymerase chain reaction. Br J Nutr 100:561–568. doi: 10.1017/S0007114508968227 [DOI] [PubMed] [Google Scholar]
- 15. Neumann AP, McCormick CA, Suen G. 2017. Fibrobacter communities in the gastrointestinal tracts of diverse hindgut-fermenting herbivores are distinct from those of the rumen. Environ Microbiol 19:3768–3783. doi: 10.1111/1462-2920.13878 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Harlow BE, Lawrence LM, Flythe MD. 2015. Sample-handling factors affecting the enumeration of lactobacilli and cellulolytic bacteria in equine feces. J Equine Vet 35:744–748. doi: 10.1016/j.jevs.2015.07.011 [DOI] [Google Scholar]
- 17. Russell JB, Wilson DB. 1996. Why are ruminal cellulolytic bacteria unable to digest cellulose at low pH? J Dairy Sci 79:1503–1509. doi: 10.3168/jds.S0022-0302(96)76510-4 [DOI] [PubMed] [Google Scholar]
- 18. Sung HG, Kobayashi Y, Chang J, Ha A, Hwang IH, Ha JK. 2007. Low ruminal pH reduces dietary fiber digestion via reduced microbial attachment. Asian-Australas J Anim Sci 20:200–207. doi: 10.5713/ajas.2007.200 [DOI] [Google Scholar]
- 19. Hungate RE. 1946. The symbiotic utilization of cellulose. J Elisha Mitchell Sci Soc Chapel Hill N C 62:9–24. [PubMed] [Google Scholar]
- 20. Weimer PJ. 2022. Degradation of cellulose and hemicellulose by ruminal microorganisms. Microorganisms 10:2345. doi: 10.3390/microorganisms10122345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Wunderlich G, Bull M, Ross T, Rose M, Chapman B. 2023. Understanding the microbial fibre degrading communities & processes in the equine gut. Anim Microbiome 5:3. doi: 10.1186/s42523-022-00224-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Froidurot A, Jacotot E, Julliand V. 2022. Whole-genome sequencing and annotation of Fibrobacter succinogenes HC4, isolated from the horse cecum. Microbiol Resour Announc 11:e0044022. doi: 10.1128/mra.00440-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Neumann AP, Suen G. 2018. The phylogenomic diversity of Herbivore-associated Fibrobacter spp. is correlated to lignocellulose-degrading potential. mSphere 3:1–18. doi: 10.1128/mSphere.00593-18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Hungate RE. 1950. The anaerobic mesophilic cellulolytic bacteria. Bacteriol Rev 14:1–49. doi: 10.1128/br.14.1.1-49.1950 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Hungate RE. 1966. The rumen bacteria, p 8–90. In The rumen and its microbes. Elsevier. [Google Scholar]
- 26. Hungate RE. 1966. Ruminant functions related to rumen microbial activity, p 148–205. In The rumen and its microbes. Elsevier. [Google Scholar]
- 27. Lee A, Gordon J, Dubos R. 1968. Enumeration of the oxygen sensitive bacteria usually present in the intestine of healthy mice. Nature 220:1137–1139. doi: 10.1038/2201137a0 [DOI] [PubMed] [Google Scholar]
- 28. Flint HJ, Duncan SH, Scott KP, Louis P. 2007. Interactions and competition within the microbial community of the human colon: links between diet and health: minireview. Environ Microbiol 9:1101–1111. doi: 10.1111/j.1462-2920.2007.01281.x [DOI] [PubMed] [Google Scholar]
- 29. Duncan SH, Hold GL, Harmsen HJM, Stewart CS, Flint HJ. 2002. Growth requirements and fermentation products of Fusobacterium prausnitzii, and a proposal to reclassify it as Faecalibacterium prausnitzii gen. nov., comb. nov. Int J Syst Evol Microbiol 52:2141–2146. doi: 10.1099/00207713-52-6-2141 [DOI] [PubMed] [Google Scholar]
- 30. Loesche WJ. 1969. Oxygen sensitivity of various anaerobic bacteria. Appl Microbiol 18:723–727. doi: 10.1128/am.18.5.723-727.1969 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Ndongo S, Khelaifia S, Lagier J-C, Raoult D. 2020. From anaerobes to aerointolerant prokaryotes. Hum Microb J 15:100068. doi: 10.1016/j.humic.2019.100068 [DOI] [Google Scholar]
- 32. Rolfe RD, Hentges DJ, Barrett JT, Campbell BJ. 1977. Oxygen tolerance of human intestinal anaerobes1’ 2 Rial D. Am J Clin Nutr 30:1762–1769. doi: 10.1093/ajcn/30.11.1762 [DOI] [PubMed] [Google Scholar]
- 33. Andrade JC, Almeida D, Domingos M, Seabra CL, Machado D, Freitas AC, Gomes AM. 2020. Commensal obligate anaerobic bacteria and health: production, storage, and delivery strategies. Front Bioeng Biotechnol 8:550. doi: 10.3389/fbioe.2020.00550 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Miller TL. 1978. The pathway of formation of acetate and succinate from pyruvate by bacteroides succinogenes. Arch Microbiol 117:145–152. doi: 10.1007/BF00402302 [DOI] [PubMed] [Google Scholar]
- 35. Weimer PJ, Hatfield RD, Buxton DR. 1993. Inhibition of ruminal cellulose fermentation by extracts of the perennial legume cicer milkvetch (Astragalus cicer). Appl Environ Microbiol 59:405–409. doi: 10.1128/aem.59.2.405-409.1993 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Meinhardt SW, Glass TL. 1994. Characterization of the NADH dehydrogenase and fumarate reductase of Fibrobacter succinogenes subsp. succinogenes S85. Arch Microbiol 162:329–334. doi: 10.1007/BF00263780 [DOI] [PubMed] [Google Scholar]
- 37. Meinhardt SW, Glass TL. 1994. NADH-linked fumarate reductase and NADH dehydrogenase activities in Fibrobacter succinogenes. Curr Microbiol 28:247–251. doi: 10.1007/BF01575969 [DOI] [Google Scholar]
- 38. Arntzen MØ, Várnai A, Mackie RI, Eijsink VGH, Pope PB. 2017. Outer membrane vesicles from Fibrobacter succinogenes S85 contain an array of carbohydrate-active enzymes with versatile polysaccharide-degrading capacity. Environ Microbiol 19:2701–2714. doi: 10.1111/1462-2920.13770 [DOI] [PubMed] [Google Scholar]
- 39. Burnet MC, Dohnalkova AC, Neumann AP, Lipton MS, Smith RD, Suen G, Callister SJ. 2015. Evaluating models of cellulose degradation by Fibrobacter succinogenes S85. PLoS One 10:e0143809. doi: 10.1371/journal.pone.0143809 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Bayer EA, Lamed R, White BA, Flint HJ. 2008. From cellulosomes to cellulosomics. Chem Rec 8:364–377. doi: 10.1002/tcr.20160 [DOI] [PubMed] [Google Scholar]
- 41. Wilson DB. 2009. Evidence for a novel mechanism of microbial cellulose degradation. Cellulose 16:723–727. doi: 10.1007/s10570-009-9326-9 [DOI] [Google Scholar]
- 42. Abdul Rahman N, Parks DH, Vanwonterghem I, Morrison M, Tyson GW, Hugenholtz P. 2016. A phylogenomic analysis of the bacterial phylum fibrobacteres. Front Microbiol 6. doi: 10.3389/fmicb.2015.01469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Dai X, Tian Y, Li J, Luo Y, Liu D, Zheng H, Wang J, Dong Z, Hu S, Huang L. 2015. Metatranscriptomic analyses of plant cell wall polysaccharide degradation by microorganisms in the cow rumen. Appl Environ Microbiol 81:1375–1386. doi: 10.1128/AEM.03682-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Chaucheyras-Durand F, Sacy A, Karges K, Apper E. 2022. Gastro-intestinal microbiota in equines and its role in health and disease: the black box opens. Microorganisms 10:2517. doi: 10.3390/microorganisms10122517 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Wang L, Zhang G, Li Y, Zhang Y. 2020. Effects of high forage/concentrate diet on volatile fatty acid production and the microorganisms involved in VFA production in cow rumen. Animals (Basel) 10:223. doi: 10.3390/ani10020223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Flint HJ, Scott KP, Louis P, Duncan SH. 2012. The role of the gut microbiota in nutrition and health. Nat Rev Gastroenterol Hepatol 9:577–589. doi: 10.1038/nrgastro.2012.156 [DOI] [PubMed] [Google Scholar]
- 47. Grimm P, Combes S, Pascal G, Cauquil L, Julliand V. 2020. Dietary composition and yeast/microalgae combination supplementation modulate the microbial ecosystem in the caecum, colon and faeces of horses. Br J Nutr 123:372–382. doi: 10.1017/S0007114519002824 [DOI] [PubMed] [Google Scholar]
- 48. Bryant MP, Burkey LA. 1953. Cultural methods and some characteristics of some of the more numerous groups of bacteria in the bovine rumen. J Dairy Sci 36:205–217. doi: 10.3168/jds.S0022-0302(53)91482-9 [DOI] [Google Scholar]
- 49. Halliwell G, Bryant MP. 1963. The cellulolytic activity of pure strains of bacteria from the rumen of cattle. J Gen Microbiol 32:441–448. doi: 10.1099/00221287-32-3-441 [DOI] [PubMed] [Google Scholar]
- 50. Hungate RE. 1969. Chapter IV A roll tube method for cultivation of strict anaerobes, p 117–132. In Methods in microbiology [Google Scholar]
- 51. Lowe SE, Theodorou MK, Trinci APJ, Hespell RB. 1985. Growth of anaerobic rumen fungi on defined and semi-defined media lacking rumen fluid. Microbiology 131:2225–2229. doi: 10.1099/00221287-131-9-2225 [DOI] [Google Scholar]
- 52. Rousseaux S, Hartmann A, Soulas G. 2001. Isolation and characterisation of new Gram-negative and Gram-positive atrazine degrading bacteria from different French soils. FEMS Microbiol Ecol 36:211–222. doi: 10.1111/j.1574-6941.2001.tb00842.x [DOI] [PubMed] [Google Scholar]
- 53. Forano E, Broussolle V, Gaudet G, Bryant JA. 1994. Molecular cloning, expression, and characterization of a new endoglucanase gene from Fibrobacter succinogenes S85. Curr Microbiol 28:7–14. doi: 10.1007/BF01575979 [DOI] [Google Scholar]
- 54. Béra-Maillet C, Ribot Y, Forano E. 2004. Fiber-degrading systems of different strains of the genus Fibrobacter. Appl Environ Microbiol 70:2172–2179. doi: 10.1128/AEM.70.4.2172-2179.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Arnaut de Toledo V de A, Colla Ruvolo-Takasusuki MC, de Oliveira AJB, Dechechi E, Sanches Lopes SM. 2012. Spectrophotometry as a tool for dosage sugars in nectar of crops pollinated by honeybees. Macro To Nano Spectroscopy. doi: 10.5772/2503 [DOI] [Google Scholar]
- 56. Bradford MM. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. doi: 10.1006/abio.1976.9999 [DOI] [PubMed] [Google Scholar]
- 57. Jouany JP. 1982. Volatile fatty acid and alcohol determination in digestive contents, silage juices, bacterial cultures and anaerobic fermentor contents [microbial fermentation, biological liquids, analytic method using gas-liquid chromatography]. Sci Aliments [Google Scholar]
- 58. Clarke KR, Owens NJP. 1983. A simple and versatile micro-computer program for the determination of “most probable number.” J Microbiol Methods 1:133–137. doi: 10.1016/0167-7012(83)90031-3 [DOI] [Google Scholar]
- 59. Drula E, Garron ML, Dogan S, Lombard V, Henrissat B, Terrapon N. 2022. The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577. doi: 10.1093/nar/gkab1045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Teufel F, Almagro Armenteros JJ, Johansen AR, Gíslason MH, Pihl SI, Tsirigos KD, Winther O, Brunak S, von Heijne G, Nielsen H. 2022. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol 40:1023–1025. doi: 10.1038/s41587-021-01156-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Moreno J, Nielsen H, Winther O, Teufel F. 2024. Predicting the subcellular location of prokaryotic proteins with DeepLocPro. Bioinformatics. doi: 10.1101/2024.01.04.574157 [DOI]
- 62. Leedle JA, Hespell RB. 1980. Differential carbohydrate media and anaerobic replica plating techniques in delineating carbohydrate-utilizing subgroups in rumen bacterial populations. Appl Environ Microbiol 39:709–719. doi: 10.1128/aem.39.4.709-719.1980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Mackie RI, Heath S. 1979. Enumeration and isolation of lactate-utilizing bacteria from the rumen of sheep. Appl Environ Microbiol 38:416–421. doi: 10.1128/aem.38.3.416-421.1979 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The complete genome of Fibrobacter sp. HC4, sequenced and annotated by Genewiz, has been deposited in NCBI under accession number JALCZV000000000 and BioProject number PRJNA812370 (22).