Abstract
Systemic lupus erythematosus (SLE) is a multifactorial disease characterized by the convergence of genetic, immunological, and viral elements resulting in a complex interaction of both internal and external factors. The role of the Epstein-Barr virus (EBV) and human endogenous retroviruses (HERV-E) as triggers and maintenance elements in the pathogenesis of SLE has been widely recognized. Previous studies have independently evaluated the effects of EBV and HERV-E in this disease. In this work, for the first time, these viral factors are jointly investigated in SLE patients. This study aimed at assessing the differential expression of immune regulatory genes and the incidence of specific viral pathogens (EBV and HERV-E), alongside the detailed characterization of surface markers in T- and B-lymphocytes in patients with SLE and control participants. A comparative analysis between patients with SLE and control participants was performed, evaluating the expression of phenotypic markers and genes involved in the immune response (TNF-α, IL-2, IL-6, IL-10, IFNG, TLR3), as well as HERV-E gag and EBV viral genes (LMP1 and BZLF1).A significant association between SLE and EBV was found in this study. A notable increase in EBV LMP1 gene expression was observed in patients with SLE . Also, a significant overexpression of HERV-E was observed, in addition to a considerable increase in the distribution of the cell surface marker CD27 + on T- and B-lymphocytes, observed in individuals with SLE compared to the control group. This study provides evidence regarding the role that EBV virus plays in lymphocytes in the context of SLE, highlighting how both the virus and the host gene expression may influence disease pathogenesis by altering immune regulatory pathways mediated by TNF-α, IFN-γ, and IL-10, as well as parallel overexpression of HERV-E gag. The decrease in TLR3 could indicate a compromised antiviral response, which could facilitate viral reactivation and contribute to disease activity.
Keywords: Lupus, T lymphocytes, B lymphocytes, Epstein-Barr virus (EBV), Human endogenous retrovirus (HERV-E)
Subject terms: Genetics, Immunology, Nephrology, Rheumatology
Introduction
Systemic lupus erythematosus (SLE) is a complex autoimmune disease whose progress is influenced by a series of multifactorial elements. The production of autoantibodies against nuclear antigens and the presence of antiphospholipid antibodies are crucial, thus triggering inflammatory and thrombotic processes. Among the most severe and prognostically unfavorable tissue manifestations of SLE is lupus nephritis (LN), affecting 50–70% of patients1–3. In SLE, chronic inflammation plays a vital role in the expression of human endogenous retrovirus elements group E (HERV-E) that are integrated into the human genome, with the possibility of exacerbation under altered immune conditions4. Viral infections, especially with the Epstein-Barr virus (EBV), contribute to an abnormal activation of the immune system and destabilization of the immune balance in susceptible individuals5. These pathogens can serve as initial triggers and perpetuating factors in the autoimmune response, thus intensifying the production of autoantibodies and the dysregulation of T- and B-lymphocytes6–8. In this context, the immune system of patients with SLE is characterized by hypersensitivity, in which minor stimuli, including viral infections and inflammatory responses, may result in an exaggerated immunologic reaction. This interaction between inflammation, the expression of endogenous retroviruses, and genetic and environmental factors highlights the complex and unpredictable nature of SLE9,10. Despite the complex pathophysiological heterogeneity of SLE, the underlying mechanisms are not yet fully defined11,12.
Among the immune-regulatory mechanisms, IL-6 and IL-10 cytokines, tumor necrosis factor-α (TNF-α), and interferon gamma (IFN-γ) are disrupted in SLE13. IL-10 is a powerful regulator of B lymphocytes, and may be negatively regulated by IFN-γ, which is overexpressed in SLE14,15. The interaction between the virome and the immune system is complex and multifaceted. Viruses can modulate the immune response through various mechanisms, including the direct activation of pattern recognition receptors, manipulation of cellular signaling pathways, and alteration of the composition and function of immune cell populations. These interactions can have both beneficial and detrimental consequences for the host, depending on the type of virus, the viral load, and the individual's immune status16. As for viral infections, EBV and cytomegalovirus (CMV), and genomic viral elements, such as HERV-E are considered potential factors related to SLE, involving lytic cycles between shorter periods, with autoantigenic cross-reactivity in the case of EBV and CMV9,10, while HERV-E is associated with global hypomethylation states, thus allowing for a greater expression of HERV-E mRNA. EBV dysregulation has been associated with the development of autoimmune diseases4,17; after the early lytic infection, EBV establishes a latent and persistent infection in memory B cells. In this latency state, the virus remains immortalized within the host cells throughout the individual's life, alternating between phases of lytic activity and periods of latency, with occasional reactivation6. This study focuses on the description of the relative expression of human immune regulation genes (TNFA, IL-6, IL-10, IFNG, and TLR3), HERV-E gag, and EBV viral genes (LMP1 and BZLF1) related to the latency and reactivation activities, respectively. All these aspects are evaluated in peripheral blood mononuclear cells (PMBC), in addition to a phenotypic characterization of T- and B-lymphocyte subpopulations in patients with SLE and healthy individuals within a study population.
Method
Study population
Four ml of venous whole blood was collected in vacutainer tubes with EDTA anticoagulant from a total of 55 patients with SLE and 61 healthy control individuals. Patients diagnosed with SLE met the criteria established by the American College of Rheumatology18,19, they were classified based on the presence or absence of documented renal involvement through renal biopsy, following the guidelines of the consensus from the Group for Systemic Autoimmune Diseases (GEAS) of the Spanish Society of Nephrology (SEN)20, and were selected from a clinic in Barranquilla, Colombia. Participants recruited for the control group were individuals with no reported autoimmune diseases. Patients and control individuals showing the presence of infectious processes at the moment of sampling were excluded, as well as those who did not give their consent to participate in the study. The study was approved by the confirm that all methods used in this study were conducted in accordance with relevant guidelines and regulations, under the approval of the Scientific Committee of the Faculty of Basic Sciences at Universidad Simón Bolívar Ethics Committee in Clinical Research of the Costa S.A.S., in the city of Barranquilla, Colombia, on September 22, 2022, through Minute No. 390. All study participants accepted and signed the informed consent.
Total RNA extraction
Total RNA extraction was performed from 300 µl of peripheral blood from patients and control individuals and preserving it in EDTA at 4 °C, using TRIzol™ reagent (Invitrogen™) as indicated by21. The extracted RNA product was later eluted in 50 µl of nuclease-free water and treated with (2 U) DNAse I (Promega™) incubated at 37 °C for 30 min. DNAse was heated for 10 min at 85 °C to inactivate it.
RT-PCR
The relative gene expression of genes (TNFA; IL-2; IL-6; IL-10; IFNG; TLR3, HERV-E gag; LMP1, and BZLF1) together with a normalizing gene (GAPDH) was assessed using real-time reverse transcription-polymerase chain reaction (RT-PCR), and direct identification of the presence of EBV infection was measured with quantitative PCR (qPCR) by detection of the viral gene LMP1. Specific primers used in the study are indicated in Table 1.
Table 1.
Primer sequences used in the study for expression assays.
| Gene | Forward and reverse primers | Accession/location |
|---|---|---|
| GAPDH | 5′ TTGCCATCAATGACCCCTTCA3′ | NM_001357943.2 |
| 5′ CGCCCCACTTGATTTTGGA3′ | ||
| TNFA | 5′ CTTCTGCCTGCTGCACTTTG3′ | NM_000594.4 |
| 5′ CCTCAGCTTGAGGGTTTGCT3′ | ||
| IFNG | 5′ AGTTATATCTTGGCTTTTCA3′ | NM_000619 |
| 5′ ACCGAATAATTAGTCAGCTT3′ | ||
| IL_6 | 5′ GGTACATCCTCGACGGCATCT3′ | XM_054358146 |
| 5′ GTGCCTCTTTGCTGCTTTCAC3′ | ||
| IL_10 | 5′ ATGCCCCAAGCTGAGAACCAAGACCCA3′ | NM_001382624 |
| 5′ TCTCAAGGGGCTGGGTCAGCTATCCCA3′ | ||
| TLR3 | 5′ GCTGCAGTCAGCAACTTCAT3′ | NM_003265.3 |
| 5′ AGGAAAGGCTAGCAGTCATCC3′ | ||
| HERV-E gag | 5′ CACATGGTGGAG AGTCGTGTTT3′ | GenBank: M10976.1 |
| 5′ GCTTGCGGCTTTTCAGTATAGG3′ | ||
| LMP1 | 5′ CCCTTTGTATACTCCTACTGATGATCAC3′ | NC_007605.1 Gene ID: 3783750 |
| 5′ ACCCGAAGATGAACAGCACAAT3′ | ||
| BZLF1 | 5′ ACGCACACGGAAACCACAA3′ | NC_007605.1 Gene ID: 3783744 |
| 5′ CTTAAACTTGGCCCGGCATT3′ |
Based on the purified RNA products, RT-qPCR was performed in a total reaction volume of 20 µl, using 2 µl of RNA template and 10 µl of the iTaq™ Universal SYBR® Green Supermix (BIO-RAD, USA) (containing 0.2 mM dNTPs, 2 U of iTaq™ DNA Polymerase, 1.5 mM MgCl2, and SYBR® Green I), as well as 2 U of RT enzyme (BIO-RAD). In addition, 0.5 µM final concentration of each primer pair per reaction was added and used in separate reactions for each gene of interest in the study.
Amplification cycles were programmed on the thermal cycler CFX96 TOUCH™ (BIO-RAD) as follows: an initial retrotranscription reaction was carried out at 50 °C for 30 min, followed by denaturation at 95 °C for 1 min. followed by 50 cycles consisting of a denaturation at 95 °C for 20 s, annealing at 58 °C for 20 s, and an extension at 72 °C for 10 s. Fluorescence was measured at the end of each extension cycle at 72 °C. Finally, a melting curve, with a temperature range between 65 °C to 95 °C (0.5 °C increments) was performed. The Ct cutoff value was set at 35. The change in the relative ARNm expression was estimated with the 2−ΔΔCT method22. All reactions were carried out in triplicate.
Enzyme-linked immunosorbent assay (ELISA) for antibodies to EBV.
EBV serology was performed by ELISA, detecting IgG EBNA1, EA and VCA, antibodies (Vircell S.L. RefG1105; G1205; M1005), strictly according to the specifications provided by the manufacturer. The plates were read at a wavelength of 450 nm using a CLARIOstar Plus multimode luminescence microplate reader.
Flow cytometry analysis
EDTA-preserved peripheral blood samples from a subset of the study population (5 SLE patients and 5 controls) were used in the flow cytometry assay. A VersaFix solution (Versalyse + 0.2% formaldehyde) and a 1X PBS – 2% BSA solution was used to prepare the samples followed by centrifugation for 8 min at 300 × g. The resulting pellet was resuspended in 1X PBS, and the cells were transferred to cytometry tubes. Surface monoclonal antibodies were added to distinguish the total lymphocyte population (T and B lymphocytes) by labeling CD3 and CD19 antigens on T and B lymphocytes, respectively. The determination of CD4 + T, CD8 + T, and B lymphocyte subpopulations—active and inactive—is based on the expression of specific markers. For the T line, anti-CD4, anti-CD8, anti-HLA-DR, and anti-CD38 antibodies were used. For the B line, anti-CD19, anti-CD27, anti-CD25, and anti-CD20 antibodies were used, besides CD 81. Details of each fluorophore are described in Table 2. The cytometric reading was performed once the separation and labeling process was completed using Beckman Coulter flow cytometry equipment (Navios Flow Cytometer). The results were analyzed by Kaluza C 1.2.1 cytometric analysis software (https://www.mybeckman.co/flow-cytometry/software/kaluza/downloads) and represented as percentages, reflecting the proportion of cells in each subpopulation in relation to the total number of lymphocytes.
Table 2.
Presentation of the anti-CD implemented in the detection of cell populations, under the antibody-fluorochrome.
| Antibody | Fluorochrome | Detectable cell type and/or activation | Beckman coulter life sciences product no |
|---|---|---|---|
| Anti-CD45 | ORANGE KROME | All hematopoietic cells | A96416 |
| Anti-CD3 | ECD | T lymphocyte lineage | A07748 |
| Anti-CD81 | APC | Signal transduction events mediating | B19717 |
| Anti-CD4 | PC5,5 | CD4 + T lymphocyte lineage | C15632 |
| Anti-CD8 | FITC | CD8 + T lymphocyte lineage | 6,603,861 |
| Anti-HLA-DR | PACIFIC BLUE | CD8 + T lymphocytes, CD4 + T lymphocytes lineage and active B lymphocytes | A74781 |
| Anti-CD19 | APC 700 | B lymphocyte lineage | B49212 |
| Anti-CD20 | ALEXA FLOUR 750 | Mature B lymphocyte lineage | B49209 |
| Anti-CD25 | PE | Treg CD4 + T-lymphocytes lineage | A07774 |
| Anti-CD27 | PC7 | Member of the tumor necrosis factor receptor superfamily | B49205 |
Statistical analysis
Statistical analysis of all results was conducted using STATGRAPHICS Centurion XVI statistical software (https://www.statgraphics.com/download-statgraphics-centurion-xvi), with all data expressed as average values and standard deviation. The differences observed in the expression of genes between patients and control subjects were analyzed using the parametric statistical test (t-test) for data with normal distribution, and the Mann–Whitney test for data with non-normal distributions. In addition, Fisher's exact test for analyzing active and inactive cases in the context of EBV infection in patients and controls. We also applied Pearson's correlation analysis to identify the relationship between some variables. To this end, the results of the gene expression were transformed into logarithmic values, for a symmetrical distribution of the data, including normal distribution verification. A t-test was also performed to compare the percentage of the different T- and B-lymphocyte populations with their corresponding CD markers. Statistical significance was accepted when the p-value was < 0.05 with 95% confidence.
We confirm that all methods used in this study were conducted in accordance with relevant guidelines and regulations, under the approval of the Scientific Committee of the Faculty of Basic Sciences at Universidad Simón Bolívar.
Results
Demographic and clinical variables
We assessed demographic and clinical variables, including age, gender, the Systemic Lupus Erythematosus Disease Activity Index (SLEDAI), (Table 3). The age and female to male ratio was comparable between the SLE and control groups and the SLEDAI parameter for the patients was 12.3 + /- 8,76. Patients with SLE and LN primarily received medical treatments with hydroxychloroquine, prednisone, prednisolone, azathioprine, and mycophenolate. However, there was no significant representation of treatments to allow for group separation. Due to the mixed nature of the population in the Colombian Caribbean, which exhibits a triple admixture of European, Native American, and African ancestry, no ethnical differentiation was established in our results23.
Table 3.
Characteristics of the study population: patients (SLE) and control subjects. Among the SLE patients, specific classifications for lupus nephritis are included, distinguishing patients into NL class II, III, or IV.
| Study population | Sample (n) | Age Average, Range, Standard deviation) | Female/Male | SLEDAI-2 K (Average, Range, Standard deviation) | Years of disease (Average, Range, Standard deviation) | Lupus nephritis. NL |
|---|---|---|---|---|---|---|
| SLE | 55 |
37,61 (21 – 72), + /- 12.15 |
50/5 |
12,3 (3–33) + /- 8,76 |
4,82 (1 – 21), ± 4,72 |
NL II: 4 NL III:14 NL IV: 18 |
| Control subjects | 61 |
35,60 (18–61) + /- 11.08 |
57/4 | N/A | N/A | N/A |
Expression profiles of immune and viral Biomarkers
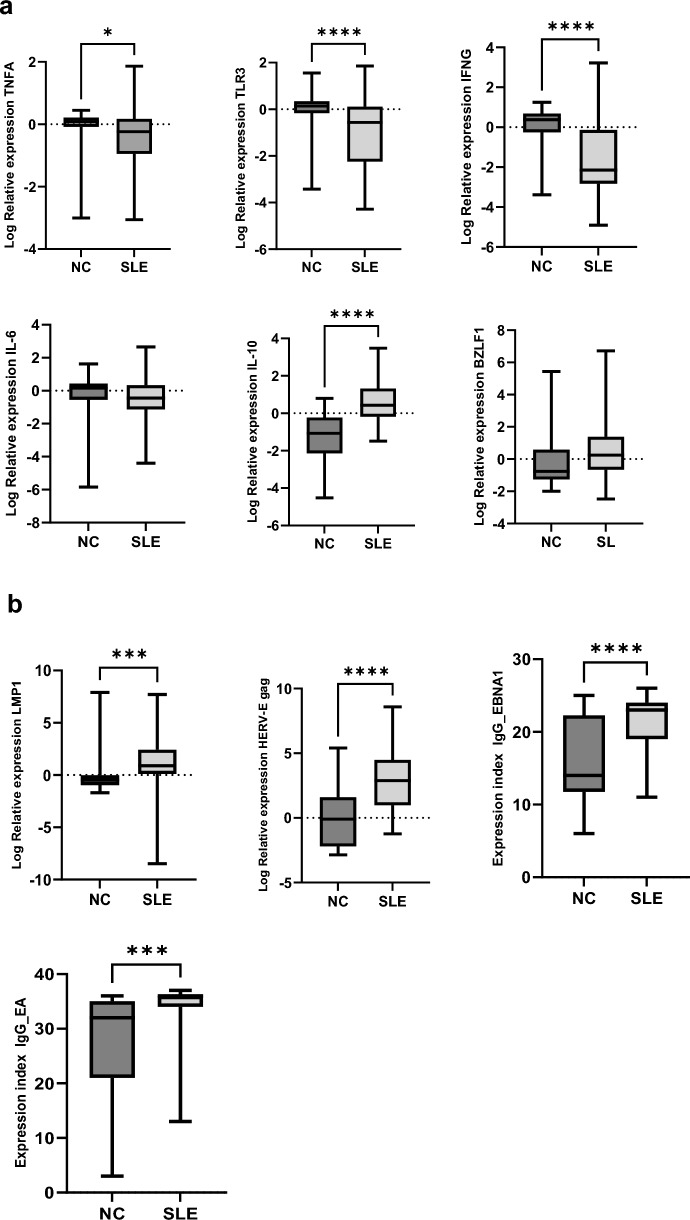
This study explored immunological gene expression signals and EBV infection signatures between patients with SLE and control individuals without autoimmune disorders. We observed a significant reduction in TNFA expression in patients with SLE compared to control individuals (p < 0.05, Fig. 1). Furthermore, we evaluated the relative expression of IL-6, IL-10, IFNG-γ, TLR3, HERV-E gag, LMP1, and BZLF1 from both groups (SLE n = 55; control individuals n = 61), showing significant differences in several of them (Fig. 1). In patients with SLE, we identified higher expression of IL-10, HERV-E gag, and LMP1, and lower expression of IFNG and TLR3. While no statistically significant differences were found in the expression of BZLF1 and IL-6 (p > 0.05). We found a significant inverse relationship between TNFA- and IL-10 by Pearson’s analysis in SLE patients (p < 0.05).
Fig.1.
Box-whisker plot of relative expression of genes (A) (TNFA; TLR3; IFNG; IL-06; IL-10) (HERV- E gag, LMP1, and BZLF1), (B) ratio of circulating plasma IgG EBNA1, and IgG EA in patients (SLE) and control subjects (NC). * p < 0.05, *** p < 0.001, **** p < 0.0001.
EBV seropositivity in SLE patients
Regarding the EBV serology, the analysis revealed distinct patterns suggestive of different infection stages. The simultaneous detection of IgG EBNA1, IgG EA, and IgG VCA indicates an active infection. Furthermore, the simultaneous presence of antibodies to both VCA and EBNA1 suggested a past infection (Table 4). It’s noteworthy that a high proportion of both SLE patients (98%) and control subjects (96%) exhibited seropositivity for EBV antibodies. However, we observed notable differences in viral activity between the two groups. Viral gene expression was detectable in 33% of SLE patients compared to 16% of controls. Moreover, a significant majority of SLE patients (93%) displayed evidence of ongoing EBV infection, contrasting with 64% in the control group (Table 4), statistically significant p < 0.05 in Fisher’s exact test. Finally, the prevalence of anti- EBNA1 IgG and anti-EA IgG was notably higher in SLE patients (Fig. 1) supporting an active infection state. Additionally, the expression of LMP1 was significantly elevated in the SLE group (p < 0.05) again suggesting viral maintenance.
Table 4.
ELISA test results for the detection of IgG EBNA1, IgG EA, and IgG VCA antibodies in patients with SLE and control subjects.
| IgG EBNA1 | IgG EA | IgG VCA | Classification | SLE n (%) | CONTROL n (%) | Fisher’s exact test |
|---|---|---|---|---|---|---|
| + /− | − | + | Previous infection | 5,2% | 28,5% | |
| + | + | + | Active infection | 94,5% | 64% | P < 0,05 |
| − | − | − | negative | 1,7% | 3,2% |
antibodies in patients with SLE and control subjects.
B -lymphocytes showed a higher expression of CD27
Flow cytometry was performed in selected subgroup (SLE n = 5 vs control individuals n = 5), obtaining total event readings in the CD45 + pan-leukocyte panel (SLE: x̄ = 111,654 ± 10,338; control subjects: x̄ = 87,275 ± 14,235). SLE and control groups showed a similar distribution of both CD45 + lymphocytes and cellular complexity and size. The proportions of CD3 + T and CD19 + B lymphocytes were also comparable in both groups. However, a significant difference in TNF-R expression was observed in CD20 + and CD27 + B lymphocytes (SLE: x̄ = 5.4% ± 2.83; NC: x̄ = 1.10% ± 1.53, p < 0.018, Fig. 2). No significant differences were found in HLA-DR, CD25, and CD81 markers (p > 0.05).
Fig.2.
Distribution of cell populations and their surface markers CD20 + (B lymphocyte); CD4 + (T helper lymphocyte); CD3 + (T lymphocyte); CD27 + (tumor necrosis factor receptor; TNF). Box-whisker plot of percentage distribution (%) of cell surface markers in patients (SLE) and control subjects (CN): A: CD4 + T lymphocytes expressing CD27 + . CD20 + B lymphocytes expressing CD27 + . *** p < 0.001, **** p < 0.0001.
Discussion
Understanding the pathophysiology of systemic lupus erythematosus (SLE) remains a challenge, given its multifaceted nature involving genetic, immunological, and environmental factors24–26. In our study, we sought to shed light on this complex condition by evaluating a cohort of individuals diagnosed with SLE, utilizing the Systemic Lupus Erythematosus Disease Activity Index (SLEDAI) score as a measure of disease activity. Our findings revealed that the SLE group exhibited predominantly active disease, as indicated by an average SLEDAI score of 12 points. Notably, 64% of these individuals presented with classification II, III, and IV nephropathies, with respective proportions of 7, 26, and 31% among the total study population.
Studies have reported that TNF-α may contribute to susceptibility to SLE through certain polymorphisms27, through elevated serum levels28, or by effects on T lymphocytes highly susceptible to TNF-α29. Furthermore, dysregulated production of TNF-α and IFN-γ, coupled with aberrant B-cell responses, has been implicated in the immunological dysfunction observed in SLE patients30. These findings underscore the intricate interplay between TNF-α and the immune dysregulation characteristic of SLE, shedding light on potential targets for therapeutic intervention. Previous studies show upregulation of these cytokines in Europeans28,31; however, in our study, where our population is different from those studies, we observed that the relative expression of both TNFA and INFG decreased significantly in patients with SLE compared to control individuals17, However, TNF-α also functions as an inflammatory mediator and inducer of apoptosis32, while deficient TNF-α production leads to the absence of both germinal centers and follicular dendritic cells33, which in murine models has been associated with lupus development34.
We examined immune changes in SLE with active or latent EBV infection. Previous studies indicate that EBV reduces IFN-γ response35–38. In our study we also show that IFN-γ is down regulated in the SLE group as well as having an active viral infection showed by the presence of anti-EBNA1, anti-EA, and anti-VCA IgGs (Table 4). The reduction of IFN-γ by EBV would probably dysregulate the already exacerbated immune response in individuals with SLE maintaining an overstimulated immune activity particular to SLE7; In the control group, both IFNG and TNFA are not reduced, maintaining low viral expression levels. Most of the population shows seropositivity, but active infection is significantly lower. Statistical relationships between IFNG and active EBV infection could not be established due to the near-total presence of infection in the patients.
We identified a significantly over expression of LMP1 in patients with SLE compared to control subjects. LMP1 is a viral protein implicated in B-cell transformation and viral maintenance39. Similar to our findings, others have demonstrated that EBV infection in individuals with lupus show a 10- to 100-fold higher expression of LMP1 compared to their control groups40–42, assessed through the viral load in peripheral blood, the frequency of infected B cells, and the amount of virus in serum. It is also important to highlight that this increase in gene expression seems not to be dependent on immunosuppressive therapy that may be ongoing to treat SLE40,41,43. These findings becomes relevant since LMP1 is a latent EBV protein with a high potential for altering cellular signal transduction pathways, including blocking intracellular DNA sensors, such as TLR9, and transcription factors, such as IRF3/744 . These are crucial pathways to promote the proliferation of target cells and, simultaneously, interfere with the regulated processes of apoptosis45. The influence of LMP1 is exerted through its expression in the plasma membrane, activating signaling pathways, such as NF- κB, protein kinases JNK, and p3845,46. In SLE patients, an increase in LMP1 could favor an increase in autoreactive B-cell survival, suggesting a mechanism for the higher activity of immune responses seen in SLE patients47.
Different HERV-E retrotransposons are more common in autoimmune disorders, indicating that genome methylation affects their expression, with DNA hypomethylation linked to SLE pathogenesis48. Our results show evidence of a significant relative overexpression of HERV-E gag in patients with SLE, also confirmed by other authors showing increased mRNA expression of HERV-E in CD4 + T cells of patients with lupus4. Since HERV-E proteins may be structurally similar to autoantigens and trigger autoimmunity through molecular mimicry, potentially serving as a new therapeutic target in lupus49,50.
There is evidence supporting the role of IL-10 in the promotion of growth and the transformation of auto-reactive B cells into plasma cells in lupus, which, in turn, influences the progress of the disease51,52. Our results show an increased expression of this cytokine in SLE patients, which is significantly negatively correlated with TNF according to Pearson’s analysis. Nonetheless, it is worth highlighting that studies on murine models indicate that IL-10 may also play a protective role in lupus as it has proinflammatory and anti-inflammatory effects53,54. This might also explain the lower expression of TNFA- seen in our studies as IL-10 inhibits TNF-α55,56 . It has been suggested that IL-10 could be a significant genetic candidate affecting susceptibility to systemic lupus erythematosus (SLE). IL-10 is a key cytokine in immune regulation, as it modulates T cell activity by reducing the production of pro-inflammatory cytokines, such as TNF-α57. Conversely, the influence of toll-like receptors (TLR) inside the cell, acting as nucleic acid sensors, are an aspect of interest in the pathogenesis of SLE; especially TLR3, which can sense double-stranded RNA (dsRNA) of viral origin, and influence cytokine production by NF-B signaling pathways58. This study shows a notable decrease in TLR3 expression in SLE patients. During active EBV infection, non-polyadenylated RNA forms loop structures simulating dsRNA, which activates TLR3 and promotes type 1 interferon production59, although our results cannot demonstrate such a relationship, given the near-complete activity of EBV in the patients, lupus patients show diminished TLR3 expression despite high EBV activity. This reduction could impair immune response and facilitate viral reactivation. Conversely, SLE patients with hepatitis C virus (HCV) have higher TLR3 expression compared to non-lupus controls also infected60 reflecting the heterogeneity of these patients’ response to infections.
The results of the distribution of the CD45 + pan-leukocyte panel among cell populations in general (neutrophils, monocytes, and lymphocytes) are an important parameter of association with the state of disease activity12. Conversely, an increase in the expression of CD27 in T and B lymphocytes suggests a crucial role in the immune activation process61,62. The differences in the distribution of CD27 + B cells are useful for the evaluation of the disease activity in patients with SLE63. On the other hand the average frequency of double-negative CD27- IgG- B cells is approximately 5% of peripheral blood B cells, and this proportion is relatively higher in older individuals (8.79–15.8%) and in patients with SLE64–66. Although no further statistical analyses were conducted due to the limitation of the flow cytometry data, the differences in the distribution of CD27 + B cells could be useful disease activity in SLE patients. It has been reported that in patients with SLE, CD27-high plasma cells increase in the periphery, correlating with SLEDAI and anti-dsDNA levels67. The different behaviors and possible effects of over and under-expressed genes in the context of lupus are represented in a unified manner based on the results obtained in Fig. 3.
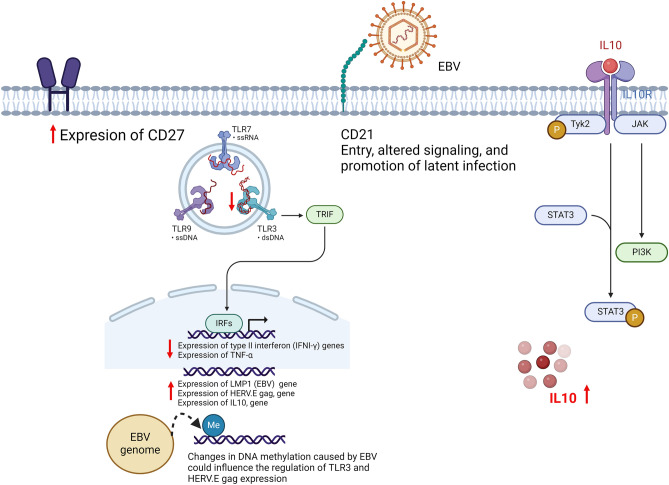
Fig.3.
Outline of relative gene expression (TNFA; TLR3; IFNG; IL-10; HERV- E gag, LMP1; CD27) showing imbalance in the context of patients with Systemic Lupus Erythematosus (SLE). The potential effect of EBV on DNA methylation is suggested by the dashed line. The arrow and red bars represent the critical points of affectation in patients with SLE. Created with BioRender.com.
The involvement of the EBV in the pathogenesis of SLE is further supported by the intricate interplay between the reactivation of HERVs and epigenetic dysregulation. The activation of HERV-E transcription, an endogenous retrovirus, by EBV viral proteins such as EBNA1 and LMP168, can trigger a cascade of events that contribute to the pathogenesis of SLE. The resulting overexpression of HERV-E can disrupt the regulation of DNA methylases, leading to global DNA hypomethylation, a hallmark of SLE69. DNA hypomethylation, in turn, can result in the dysregulation of genes critical for the immune response, including those encoding interferons and pro-inflammatory cytokines70. EBV infection also stimulates the production of inflammatory cytokines, such as IFN-γ, which can further modulate the expression and activity of methylases, amplifying epigenetic dysregulation and promoting the expression of genes associated with autoimmunity71.
Furthermore, the observed decrease in TLR3 expression in SLE patients, despite high EBV activity, adds another layer of complexity to this interaction. TLR3, a Toll-like receptor that recognizes double-stranded RNA, plays a crucial role in the antiviral immune response. The reduction in its expression could indicate an immune evasion mechanism employed by EBV, allowing its persistence and reactivation, which in turn perpetuates the chronic immune stimulation and epigenetic dysregulation observed in SLE. The interplay between EBV, HERVs, methylase regulation, inflammatory cytokine production, and decreased TLR3 expression paints a complex and multifaceted picture of SLE pathogenesis, highlighting the need for further research to fully understand these interactions and develop therapies targeting these mechanisms(Fig. 3).
The intricate relationship between EBV infection, HERV-E reactivation, and epigenetic dysregulation in SLE pathogenesis is further enriched by the unique contribution of this study. It stands as the first to comprehensively evaluate the combined interaction of EBV and HERV-E in the context of lupus, contrasting it with healthy controls. This approach provides a deeper understanding of the synergistic effects of these viral elements and their impact on the host immune response. By simultaneously examining viral gene expression, cytokine profiles, and lymphocyte phenotypes, this study offers a holistic view of the complex interplay between viral infection, epigenetic modifications, and immune dysregulation in SLE. These findings are essential for patient stratrification and the development of more target therapeutic interventions.
Acknowledgements
The authors wish to express their gratitude to the Colombian Association of Immunology, Simón Bolívar University, Clínica de la Costa S.A.S., and the Ministry of Science, Technology, and Innovation of Colombia (through the "Bicentennial Doctoral Excellence Scholarships," established in Article 45 of Law No. 1942 of 2018) for their invaluable support in the development and publication of this research. Their contribution has been crucial to the success of this work.
Author contributions
Y.B.L: Conceptualization, Investigation, Writing—original draft, Writing—review & editing. G.A.M: Conceptualization, Investigation, Supervision. L.P.L: Investigation, Writing—review & editing. L.G.E: Investigation, Supervision. E.Z.P: Conceptualization, Investigation, Writing—review & editing. N.S.L: Investigation, Writing. A.C.B: Conceptualization, Investigation, Supervision. A.J.A-H: Conceptualization, Supervision, Writing—review & editing. E.N.Q: Conceptualization, Investigation, Writing—review & editing. All authors reviewed the manuscript.
Data availability
The data supporting the findings of this study are available from the corresponding author, Y.B.L., upon request (yesit.bello@unisimon.edu.co).
Competing interests
The authors declare no competing interests.
Footnotes
The original online version of this Article was revised: The Acknowledgements section in the original version of this Article was omitted. Full information regarding the corrections made can be found in the correction for this Article.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
9/26/2024
A Correction to this paper has been published: 10.1038/s41598-024-73833-7
References
- 1.Karrar, S. & Cunninghame Graham, D. S. Abnormal B cell development in systemic lupus erythematosus. Arthritis Rheumatol.70, 496–507 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lugo, L. P., Olmos, Y. D. & Martínez, G. A. Biomarcadores en fluídos biológicos y su potencial uso como indicadores de nefritis lúpica en individuos con lupus eritematoso sistémico. Rev. Colomb. Nefrol.1, 39–47 (2014). [Google Scholar]
- 3.García Tello, A., Villegas Martínez, A. & González Fernández, A. F. Manifestaciones hematológicas en el lupus eritematoso sistémico. An.Med. Interna19, 53–57 (2002). [PubMed] [Google Scholar]
- 4.Wu, Z. et al. DNA methylation modulates HERV-E expression in CD4+ T cells from systemic lupus erythematosus patients. J. Dermatol. Sci.77, 110–116 (2015). [DOI] [PubMed] [Google Scholar]
- 5.Piroozmand, A., Kashani, H. H. & Zamani, B. Correlation between epstein-barr virus infection and disease activity of systemic lupus erythematosus: A cross-sectional study. Asian Pac. J. Cancer Prev.18, 523–527 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Thorley-Lawson, D. A. Epstein-Barr virus: Exploiting the immune system. Nat. Rev. Immunol.1, 75–82 (2001). [DOI] [PubMed] [Google Scholar]
- 7.Draborg, A. H. et al. Impaired cytokine responses to Epstein-Barr virus antigens in systemic lupus erythematosus patients. J. Immunol. Res.2016, e6473204 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Adamson, A. L. et al. Epstein-Barr virus immediate-early proteins BZLF1 and BRLF1 Activate the ATF2 transcription factor by increasing the levels of phosphorylated p38 and c-Jun N-terminal kinases. J. Virol.74, 1224–1233 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Guo, G. et al. The cytomegalovirus protein US31 induces inflammation through mono-macrophages in systemic lupus erythematosus by promoting NF-κB2 activation. Cell Death Dis.9, 1–15 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Laurynenka, V., Ding, L., Kaufman, K. M., James, J. A. & Harley, J. B. A high prevalence of anti-EBNA1 heteroantibodies in systemic lupus erythematosus (SLE) supports anti-EBNA1 as an origin for SLE autoantibodies. Front. Immunol.13, 830993 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wild, C. P. Complementing the genome with an “exposome”: The outstanding challenge of environmental exposure measurement in molecular epidemiology. Cancer Epidemiol. Biomark. Prev.14, 1847–1850 (2005). [DOI] [PubMed] [Google Scholar]
- 12.Gonzalez-Quintial, R. et al. Lupus acceleration by a MAVS-activating RNA virus requires endosomal TLR signaling and host genetic predisposition. PLoS ONE13, e0203118 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Richaud-Patin, Y., Alcocer-Varela, J. & Llorente, L. High levels of TH2 cytokine gene expression in systemic lupus erythematosus. Rev. Invest. Clin.47, 267–272 (1995). [PubMed] [Google Scholar]
- 14.Liu, T. F., Jones, B. M., Wong, R. W. S. & Srivastava, G. Impaired production of Il-12 in systemic lupus erythematosus. III: Deficient il-12p40 gene expression and cross-regulation of Il-12, Il-10 and ifn-γ gene expression. Cytokine11, 805–811 (1999). [DOI] [PubMed] [Google Scholar]
- 15.Hu, X. et al. IFN-γ suppresses IL-10 production and synergizes with TLR2 by regulating GSK3 and CREB/AP-1 proteins. Immunity24, 563–574 (2006). [DOI] [PubMed] [Google Scholar]
- 16.Mitchell, A. B., Oliver, B. G. G. & Glanville, A. R. translational aspects of the human respiratory virome. Am. J. Respir. Crit. Care Med.194, 1458–1464 (2016). [DOI] [PubMed] [Google Scholar]
- 17.Draborg, A. H., Duus, K. & Houen, G. Epstein-Barr virus in systemic autoimmune diseases. Clin. Dev. Immunol.2013, 535738 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Rigante, D. & Esposito, S. Infections and systemic lupus erythematosus: Binding or sparring partners?. Int. J. Mol. Sci.16, 17331–17343 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Petri, M. et al. Derivation and validation of the systemic lupus international collaborating clinics classification criteria for systemic lupus erythematosus. Arthritis. Rheum64, 2677–2686 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ruiz Irastorza, G. et al. Diagnosis and treatment of lupus nephritis consensus document from the systemic auto-immune disease group (GEAS) of the Spanish society of internal medicine (SEMI) and spanish society of nephrology (S.E.N.). Nefrologia32(1), 1–35 (2012). [DOI] [PubMed] [Google Scholar]
- 21.Rio, D. C., Ares, M., Hannon, G. J. & Nilsen, T. W. Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb. Protoc.2010, 5439 (2010). [DOI] [PubMed] [Google Scholar]
- 22.Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods25, 402–408 (2001). [DOI] [PubMed] [Google Scholar]
- 23.Moreno-Estrada, A. et al. Reconstructing the population genetic history of the Caribbean. PLoS Genet.9, e1003925 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Catalina, M. D., Owen, K. A., Labonte, A. C., Grammer, A. C. & Lipsky, P. E. The pathogenesis of systemic lupus erythematosus: Harnessing big data to understand the molecular basis of lupus. J. Autoimmun.110, 102359 (2020). [DOI] [PubMed] [Google Scholar]
- 25.Pan, L., Lu, M.-P., Wang, J.-H., Xu, M. & Yang, S.-R. Immunological pathogenesis and treatment of systemic lupus erythematosus. World J. Pediatr.16, 19–30 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Barbhaiya, M. & Costenbader, K. H. Environmental exposures and the development of systemic lupus erythematosus. Curr. Opin. Rheumatol.28, 497–505 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ramírez-Bello, J. et al. Tumor necrosis factor gene polymorphisms are associated with systemic lupus erythematosus susceptibility or lupus nephritis in Mexican patients. Immunol. Res.66, 348–354 (2018). [DOI] [PubMed] [Google Scholar]
- 28.Studnicka-Benke, A., Steiner, G., Petera, P. & Smolen, J. S. Tumour necrosis factor alpha and its soluble receptors parallel clinical disease and autoimmune activity in systemic lupus erythematosus. Br. J. Rheumatol.35, 1067–1074 (1996). [DOI] [PubMed] [Google Scholar]
- 29.Habib, H. M., Taher, T. E., Isenberg, D. A. & Mageed, R. A. Enhanced propensity of T lymphocytes in patients with systemic lupus erythematosus to apoptosis in the presence of tumour necrosis factor alpha. Scand. J. Rheumatol.38, 112–120 (2009). [DOI] [PubMed] [Google Scholar]
- 30.Mitamura, K. et al. Impaired tumour necrosis factor-alpha (TNF-alpha) production and abnormal B cell response to TNF-alpha in patients with systemic lupus erythematosus (SLE). Clin. Exp. Immunol.85, 386–391 (1991). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Davas, E. M. et al. Serum IL-6, TNFalpha, p55 srTNFalpha, p75srTNFalpha, srIL-2alpha levels and disease activity in systemic lupus erythematosus. Clin. Rheumatol.18, 17–22 (1999). [DOI] [PubMed] [Google Scholar]
- 32.Postal, M. & Appenzeller, S. The role of tumor necrosis factor-alpha (TNF-α) in the pathogenesis of systemic lupus erythematosus. Cytokine56, 537–543 (2011). [DOI] [PubMed] [Google Scholar]
- 33.Aringer, M. & Smolen, J. S. SLE - Complex cytokine effects in a complex autoimmune disease: Tumor necrosis factor in systemic lupus erythematosus. Arthritis. Res. Ther5, 172–177 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kontoyiannis, D. & Kollias, G. Accelerated autoimmunity and lupus nephritis in NZB mice with an engineered heterozygous deficiency in tumor necrosis factor. Eur. J. Immunol.30, 2038–2047 (2000). [DOI] [PubMed] [Google Scholar]
- 35.Liu, X., Sadaoka, T., Krogmann, T. & Cohen, J. I. Epstein-Barr virus (EBV) tegument protein BGLF2 suppresses type I interferon signaling to promote EBV reactivation. J. Virol.94, e00258-e320 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li, Y. et al. Epstein-Barr virus BZLF1-mediated downregulation of proinflammatory factors is essential for optimal lytic viral replication. J. Virol.90, 887–903 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Morrison, T. E., Mauser, A., Wong, A., Ting, J. P. & Kenney, S. C. Inhibition of IFN-gamma signaling by an Epstein-Barr virus immediate-early protein. Immunity15, 787–799 (2001). [DOI] [PubMed] [Google Scholar]
- 38.Hohenadl, C. et al. Transcriptional activation of endogenous retroviral sequences in human epidermal keratinocytes by UVB irradiation. J. Investig. Dermatol.113, 587–594 (1999). [DOI] [PubMed] [Google Scholar]
- 39.Ahsan, N., Kanda, T., Nagashima, K. & Takada, K. Epstein-Barr virus transforming protein LMP1 plays a critical role in virus production. J. Virol.79, 4415–4424 (2005). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Harley, J. B. & James, J. A. Everyone comes from somewhere: Systemic lupus erythematosus (SLE) and Epstein-Barr Virus, induction of host interferon (INF) and humoral anti-EBNA1 immunity. Arthritis. Rheum.62, 1571–1575 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kang, I. et al. Defective control of latent Epstein-Barr virus infection in systemic lupus erythematosus. J. Immunol.172, 1287–1294 (2004). [DOI] [PubMed] [Google Scholar]
- 42.Gross, A. J., Hochberg, D., Rand, W. M. & Thorley-Lawson, D. A. EBV and systemic lupus erythematosus: A new perspective. J. Immunol.174, 6599–6607 (2005). [DOI] [PubMed] [Google Scholar]
- 43.Moon, U. Y. et al. Patients with systemic lupus erythematosus have abnormally elevated Epstein-Barr virus load in blood. Arthritis Res. Ther.6, R295-302 (2004). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bentz, G. L., Shackelford, J. & Pagano, J. S. Epstein-Barr virus latent membrane protein 1 regulates the function of interferon regulatory factor 7 by inducing its sumoylation. J. Virol.86, 12251–12261 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Johansson, P., Jansson, A., Rüetschi, U. & Rymo, L. The p38 signaling pathway upregulates expression of the Epstein-Barr Virus LMP1 oncogene. J. Virol.84, 2787–2797 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yang, L. et al. EBV-LMP1 targeted DNAzyme enhances radiosensitivity by inhibiting tumor angiogenesis via the JNKs/HIF-1 pathway in nasopharyngeal carcinoma. Oncotarget6, 5804–5817 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Lam, N. & Sugden, B. LMP1, a viral relative of the TNF receptor family, signals principally from intracellular compartments. EMBO. J.22, 3027–3038 (2003). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Okada, M. et al. Role of DNA methylation in transcription of human endogenous retrovirus in the pathogenesis of systemic lupus erythematosus. J. Rheumatol.29, 1678–1682 (2002). [PubMed] [Google Scholar]
- 49.Talotta, R., Atzeni, F. & Laska, M. J. The contribution of HERV-E clone 4–1 and other HERV-E members to the pathogenesis of rheumatic autoimmune diseases. APMIS128, 367–377 (2020). [DOI] [PubMed] [Google Scholar]
- 50.Tugnet, N., Rylance, P., Roden, D., Trela, M. & Nelson, P. Human endogenous retroviruses (HERVs) and autoimmune rheumatic disease: Is there a link?. Open Rheumatol. J.7, 13–21 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sugita, K. et al. CD27, a member of the nerve growth factor receptor family, is preferentially expressed on CD45RA+ CD4 T cell clones and involved in distinct immunoregulatory functions. J. Immunol.149, 3208–3216 (1992). [PubMed] [Google Scholar]
- 52.Dörner, T. & Lipsky, P. E. Correlation of circulating CD27high plasma cells and disease activity in systemic lupus erythematosus. Lupus13, 283–289 (2004). [DOI] [PubMed] [Google Scholar]
- 53.Blenman, K. R. M. et al. IL-10 regulation of lupus in the NZM2410 murine model. Lab. Investig.86, 1136–1148 (2006). [DOI] [PubMed] [Google Scholar]
- 54.Ling, G.-S., Cook, H. T., Botto, M., Lau, Y.-L. & Huang, F.-P. An essential protective role of IL-10 in the immunological mechanism underlying resistance vs susceptibility to lupus induction by dendritic cells and dying cells. Rheumatol. (Oxford)50, 1773–1784 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Clarke, C. J. P., Hales, A., Hunt, A. & Foxwell, B. M. J. IL-10-mediated suppression of TNF-α production is independent of its ability to inhibit NFκB activity. Eur. J. Immunol.28, 1719–1726 (1998). [DOI] [PubMed] [Google Scholar]
- 56.Maiti, S., Dai, W., Alaniz, R. C., Hahn, J. & Jayaraman, A. Mathematical modeling of pro- and anti-inflammatory signaling in macrophages. Processes3, 1–18 (2015). [Google Scholar]
- 57.de Waal Malefyt, R. et al. Interleukin 10 (IL-10) and viral IL-10 strongly reduce antigen-specific human T cell proliferation by diminishing the antigen-presenting capacity of monocytes via downregulation of class II major histocompatibility complex expression. J. Exp. Med.174, 915–924 (1991). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Ranjith-Kumar, C. T. et al. Effects of single nucleotide polymorphisms on Toll-like receptor 3 activity and expression in cultured cells. J. Biol. Chem.282, 17696–17705 (2007). [DOI] [PubMed] [Google Scholar]
- 59.Iwakiri, D. et al. Epstein-Barr virus (EBV)-encoded small RNA is released from EBV-infected cells and activates signaling from Toll-like receptor 3. J. Exp. Med.206, 2091–2099 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Razin, M. et al. TLR3\TLR7 as differentially expressed markers among viral, nonviral, and autoimmune diseases in Egyptian patients. Viral Immunol.34, 607–621 (2021). [DOI] [PubMed] [Google Scholar]
- 61.Eliopoulos, A. G. & Young, L. S. LMP1 structure and signal transduction. Semin. Cancer Biol.11, 435–444 (2001). [DOI] [PubMed] [Google Scholar]
- 62.Caielli, S. et al. A CD4+ T cell population expanded in lupus blood provides B cell help through interleukin-10 and succinate. Nat. Med.25, 75–81 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Geginat, J. et al. IL-10 producing regulatory and helper T-cells in systemic lupus erythematosus. Semin. Immunol.44, 101330 (2019). [DOI] [PubMed] [Google Scholar]
- 64.Wei, C. et al. A new population of cells lacking expression of CD27 represents a notable component of the b cell memory compartment in systemic lupus erythematosus1. J. Immunol.178, 6624–6633 (2007). [DOI] [PubMed] [Google Scholar]
- 65.Wu, Y.-C.B., Kipling, D. & Dunn-Walters, D. K. The relationship between CD27 negative and positive B cell populations in human peripheral blood. Front. Immunol.10.3389/fimmu.2011.00081 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Li, Y., Li, Z. & Hu, F. Double-negative (DN) B cells: an under-recognized effector memory B cell subset in autoimmunity. Clin. Exp. Immunol.205, 119–127 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Jacobi, A. M. et al. HLA-DRhigh/CD27high plasmablasts indicate active disease in patients with systemic lupus erythematosus. Ann. Rheum. Dis.69, 305–308 (2010). [DOI] [PubMed] [Google Scholar]
- 68.Mameli, G. et al. Brains and peripheral blood mononuclear cells of multiple sclerosis (MS) patients hyperexpress MS-associated retrovirus/HERV-W endogenous retrovirus, but not Human herpesvirus 6. J. Gen. Virol.88, 264–274 (2007). [DOI] [PubMed] [Google Scholar]
- 69.Kitsou, K., Lagiou, P. & Magiorkinis, G. Human endogenous retroviruses in cancer: Oncogenesis mechanisms and clinical implications. J. Med. Virol.95, e28350 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Sun, B. et al. DNA methylation perspectives in the pathogenesis of autoimmune diseases. Clin. Immunol.164, 21–27 (2016). [DOI] [PubMed] [Google Scholar]
- 71.Relle, M., Foehr, B. & Schwarting, A. Epigenetic aspects of systemic lupus erythematosus. Rheumatol. Ther.2, 33–46 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
The data supporting the findings of this study are available from the corresponding author, Y.B.L., upon request (yesit.bello@unisimon.edu.co).