Abstract
The simplest approach to convey the results of scientific analysis, which can include complex comparisons, is typically through the use of visual items, including figures and plots. These statistical plots play a critical role in scientific studies, making data more accessible, engaging, and informative. A growing number of visual representations have been utilized recently to graphically display the results of oncologic imaging, including radiomic and radiogenomic studies. Here, we review the applications, distinct properties, benefits, and drawbacks of various statistical plots. Furthermore, we provide neuroradiologists with a comprehensive understanding of how to use these plots to effectively communicate analytical results based on imaging data.
Keywords: Statistical plot, waterfall plot, spider plot, heatmap, volcano plot, principal component, t-SNE
Introduction
Visual illustrations of quantitative information, including graphs and figures, are often practical and powerful methods of presenting scientific research results.1,2 These graphical illustrations are an essential part of medical publications. Up to half of a given biological article’s content can be related to figures, and most clinical trial articles report one to three figures.3,4
Plots and figures provide a visual representation of data, making it easier for readers to understand and interpret the results. They can also allow for a more efficient and concise way to present complex data, compared to describing it in text. Additionally, plots and figures can highlight patterns and trends in the data, making it easier to identify significant results and novel relationships. They also help to make the data more engaging and can help to increase readers' interest in a study.5,6
Recently, an increasing number of varied diagrams and plots have been used to visually display outcomes in radio-oncologic and radiogenomic studies. Thus, radiologists must be familiar with these kinds of data representations to fully understand the literature and be able to make accurate assessments of results. In this review, we describe the potential application of these various statistical plots in oncologic imaging research and provide guidance on their interpretations. We also highlight the specific features of each graph type and review potential advantages and drawbacks/pitfalls.
Graphical illustrations of tumor response
Waterfall plots
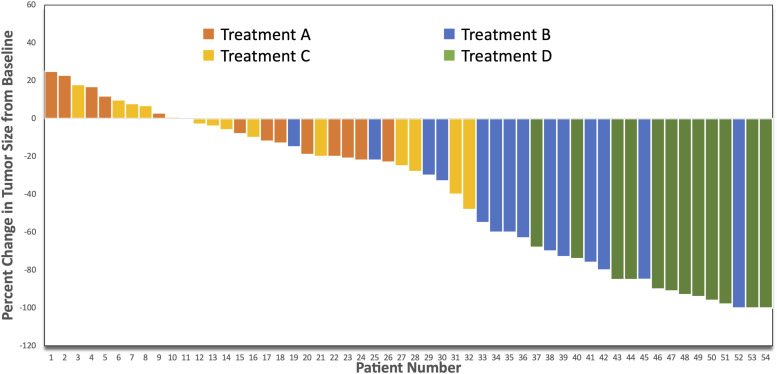
Waterfall plots have been used in radio-oncologic studies to show maximum tumor size change from baseline after therapy (Figure 1). Each bar represents a patient, and subjects are shown along the x-axis in increasing or decreasing order. Bars below the zero on the y-axis represent tumor shrinkage, while those above zero represent tumor expansion. In addition, bars could be labeled with different colors to add more information to the plot (e.g., clinical or molecular characteristics, treatment type, etc.). 7 Moreover, in radiomic studies that use a radiomic score to categorize patients into high-risk and low-risk groups, waterfall plots can be used to sort patients based on their scores in an ascending or descending order (Figure 2). 8 In this case, one could quickly identify the relative number of subjects in each group. Waterfall plots are employed in radiogenomic studies, too. For example, gene sets can be ordered in an increasing level of association with MRI features along the x-axis (Figure 3). 9
Figure 1.
Waterfall plot showing response to different kinds of treatments: percent change in tumor burden.
Figure 2.
Waterfall plot of the radiomics signature for each patient in the radiomics validation subset. Based on the radiomics signature cutoff of −4.0, patients were divided into a high-risk group (≥−4.0) and a low-risk group (<−4.0). The status of dead or censorship was marked with different colors. Reproduced from Sun et al., Radiology, Copyright 2021, Vol. 301, Pages 654–663, with permission from the Radiological Society of North America (RSNA®). 8
Figure 3.
Waterfall plots show statistics for MRI factor 1 (tumor size) used to select gene sets for display in heat map and table, for all gene sets in MsigDB c2.cgp, with selected gene sets in green. Waterfall plots show three types of association statistics on y-axis calculated for all gene sets, which are ordered in increasing level of association along x-axis: normalized enrichment statistic (NES), maximum enrichment statistic at (Max. ES at), and leading edge. Fill of maximum enrichment statistic waterfall plot is zero at middle, such that enrichment at top is shown downward and enrichment at bottom upwards. Reproduced from Bismeijer et al., Radiology, Copyright 2020, Vol. 296, Pages 277–287, with permission from the Radiological Society of North America (RSNA®) 9
Waterfall plots have some limitations. They can only represent the best on-study change in tumor burden compared to baseline for each patient; however, they cannot show tumor progression kinetics or dynamics. 10 Second, they may potentially overestimate the response rate based on visual assessment. For example, Kim et al. concluded that waterfall plots demonstrated visual response rates that were 6.1% higher than response rates based on investigator review and 12.0% higher than response rates based on central review. 11 Third, changes over time are not depicted on waterfall plots. 12 These limitations should be kept in mind, when using them in the context of clinical studies.
Spider plots
In oncologic imaging, spider plots have been utilized to illustrate a change in tumor volume over time compared to baseline tumor burden. They also attempt to normalize the initial tumor volumes and then track the percentage of change over time compared to baseline rather than absolute changes in volume. In contrast to waterfall plots, spider plots allow us to study the progression and duration of responses while indirectly identifying the length of the best response. 13 In addition, the coloring and shape of the end point, as well as the line style and line color, may all be used to provide extra information, giving researchers a variety of options (Figure 4). 14
Figure 4.
Spider plot displaying response to therapy over time in 7 subjects with low-grade glioma based on the Response Assessment in Neuro-Oncology (RANO) criteria by utilizing percent change in T2/FLAIR signal volume.
One issue with spider plots (which also affects waterfall plots) is that they show percentage change from baseline which is statistically inefficient and may be deceptive. 15 Unequal variances between baseline and subsequent measurements are hidden by presenting percentage change from baseline, resulting in a potentially optimistic, rather than fully accurate, assessment of data variability. In order to overcome this challenge, Mercier et al. suggested adding a simple spaghetti plot to depict the change of individual raw sum of lesion diameter (SLD) values across time. 12 Additionally, when there are a high number of patients, spider plots are more difficult to read since it becomes hard to follow individual lines, especially when they intersect. 14
Swimmer plots
Swimmer plots are a data visualization tool that employs horizontal bars, and various graphical symbols and colors to depict subjects’ responses to treatment (e.g., Response Assessment in Neuro-Oncology [RANO] measurements) over time (Figure 5). 16 Each patient is represented as a single horizontal bar, color-coded by the patient’s disease stage at the commencement of therapy at month zero, using time as the x-axis. The type of tumor response is shown by adding graphical symbols to the plot. Further information conveyed through a swimmer plot includes the duration of treatment, as well as the onset of response and its duration. Conventionally, patients are ordered according to how long their treatments have lasted. Thus, one is able to quickly identify which patients received therapy for a longer period of time.
Figure 5.
Swimmer plot of 30 patients with glioma at relapse treated with regorafenib sorted by overall survival after initiation of therapy. Time to progression ranged from 0.8 to 8.2 months. Patients with oligodendroglioma (#2, 4) were alive after 13.8 and 20.7 months, respectively. Most patients with glioblastoma (96%) and astrocytoma (75%) had died. Reproduced from Werner et al 18
However, when too many patients or variables are included, they become untidy and uninformative. 14 Furthermore, the data are presented in the form of categorical variables, such as a complete or partial response, and does not depict the extent of the response or how it varies numerically over time. 13 In addition, although it offers valuable information on ongoing therapy response, it offers little to no data on stable disease or responses to previous treatment(s). 17
Graphical illustrations of cancer genotypes and phenotypes
The molecular landscape of cancer is enormous and complex, making it a challenge to represent genetic, expressions, and proteomic data in visually simple ways. Heatmaps, volcano plots, and many other representations have been commonly employed to make this high-dimensional data easier to understand.
Heatmaps
Heatmaps have become a mainstay of oncologic research and are increasingly present in radiologic spaces with studies that look at radiomic and radiogenomic feature sets. Their popularity is likely related to their ability to condense high-dimensional data into easy-to-interpret matrices and their ease of generation with packages in commonly used statistical software. Their design typically includes patients or subjects in the columns of the matrix with rows representing individual features, which could be any number of radiomic or multi-omic data. Colors and their intensities and shades are used to represent levels of expression or the presence or absence of a mutation at each intersection (Figure 6). 19 Dot maps are similar in their use, and build on this by adding a dimension of effect size, which is represented by the size of the dot at the intersections of the grid. 20 Typically, features are clustered and organized so that patterns in the data can be easily recognized. Heatmap representations are familiar to readers across a variety of fields and can be used to represent a wide variety of matrix data. For example, in radiogenomic studies, it is not uncommon to see radiomic features mapped against molecular features, which allows readers a method of visualizing relationships between these large feature sets.
Figure 6.
Feature weight matrix. This heatmap demonstrates the relative influence of each feature on the overall clustering attempt. Darker cells depict higher values of influence as calculated by the scalar product between the feature vector and the principal component vector multiplied by the explained variance of the respective principal component. By performing this operation, the relative importance of certain types of features and imaging sequences can be understood. Feature names describe the attribute that they measure in the following way: ImagingSequence_FeatureType_Metric. Reprinted from Neoplasia, Vol. 36, Haldar et al., Unsupervised machine learning using K-means identifies radiomic subgroups of pediatric low-grade gliomas that correlate with key molecular markers, Pages 100869, Copyright 2023, with permission from Elsevier. 21
Despite their popularity, heatmaps lack robustness in their application as a visual tool as they allow users to alter many parameters related to their generation. For example, representations of similar datasets can vary in terms of color, arrangement of variables, ranges of color saturation, and dimensionality of the map. Furthermore, there is no standardized procedure for clustering features, so heatmaps generated from the same dataset but through different clustering techniques could appear vastly different. Additionally, if there are an extremely large number of features (e.g., thousands) it can be difficult to view more subtle trends. While this does not undermine their utility in representing patterns of relationships between subjects and high-dimensional features, it does limit one’s ability to compare maps between studies and requires the interpreter to pay attention to the methods used to create the representation.
Heatmaps projected onto anatomical atlases
A modification of the heatmap from its usual grid form involves overlaying it onto a standardized atlas or onto patient-specific images showing a tumor. These types of projections are familiar to radiologists (e.g., BOLD signal representations from fMRI). In recent oncological applications, radiogenomic works have been able to correlate imaging features with molecular markers, allowing for probability maps of certain molecular signatures onto tumor imaging. 22 Such representations use color to show the potential presence or absence of certain molecular markers within different regions of the tumor with color intensity/shade being related to the likelihood of expression (Figure 7). These representations are very intuitive and offer a spatial dimension to the data being presented. However, unlike traditional heatmaps which can show patterns in thousands of features all at once, these heatmap overlays are limited in showing one or a few features at a time.
Figure 7.
Radiogenomics maps resolve the regional intratumoral heterogeneity of EGFR amplification status in GBM. Shown are two different image-localized biopsies (Biopsy #1, Biopsy #2) from the same GBM tumor in a single patient. For each biopsy, T1+C images (left) demonstrate the enhancing tumor segment (dark green outline, T1W+Contrast) and the peripheral non-enhancing tumor segment (light green outline, T2W lesion). Radiogenomics color maps for each biopsy (right) also show regions of predicted EGFR amplification (amp, red) and EGFR wildtype (wt, blue) status overlaid on the T1+C images. For biopsy #1 (green square), the radiogenomics map correctly predicted low EGFR copy number variant (CNV) and wildtype status with high predictive certainty (p < 0.05). Conversely for biopsy #2 (green circle), the maps correctly predicted high EGFR CNV and amplification status, also with high predictive certainty (p < 0.05). Note that both biopsies originated from the non-enhancing tumor segment, suggesting the feasibility for quantifying EGFR drug target status for residual subpopulations that are typically left unresected followed gross total resection. Reproduced from Hu et al. 22
Circos (circular multitrack) plots
Circos plots are circular representations that can show relationships in complex data in a visually appealing and symmetric way. They were originally designed to represent genomic data and can capture information on copy number variations, gene interconnectedness, biochemical pathways, pathogenic pathways, and much more. 23 Their use in oncology has been primarily to organize the large amount of gene sequencing data that is collected on tumors. In radiogenomics, circos plots can be implemented as ways to relate genetic features with radiomic features (Figure 8). Notably, circos plots differ from similar-appearing circular dendrograms as they represent connectivity across objects represented in the periphery whereas in dendrograms, objects on the periphery are connected by a branching structure to the center of the plot.
Figure 8.
Radiogenomic associations in TCGA-TCIA GBM. Molecular omic features are represented on the top of the image, while imaging features are represented on the bottom. The arcs represent relations. (–) indicates a negative relation, (+) a positive relation, (m) mutation of the corresponding gene, (l) a low value of the corresponding feature, and (h) a high value. CER: Contrast-enhancing ratio, CEV: Contrast-enhancing volume, TCGA: The Cancer Genome Atlas, TCIA: The Cancer Imaging Archive, GBM: glioblastoma multiforme. Reproduced from Zanfardino et al. 24
While the information represented in circos plots is appealing and information-rich, criticism of these plots includes low resolution and the visual over-emphasis of connections between distant genes compared to those that are nearby one another on the plot. Further, the interpretation of specific circos plots varies as different colors, line weights and arrangements of variables can be selected as parameters for the plots.
Volcano plots
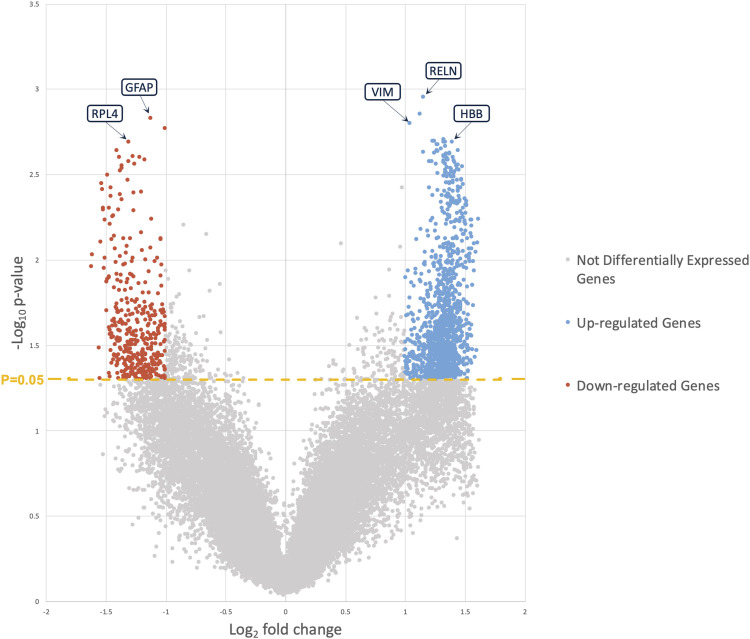
Volcano plots are modified scatter plots that allow for easy identification of changes in large datasets. These are most commonly employed with genomic, transcriptomic, proteomic, and metabolomic data that contain thousands of features. 25 Volcano plots highlight the highest magnitude changes in expressions of these data as well as the most significant changes. They accomplish this by plotting the magnitude of the change on the x-axis vs the negative log of the p-value of the change on the y-axis. The plots are centered at zero on the x-axis, with points to the right representing increased expression and those to the left representing decreased expression. Because lower p-values represent greater statistical significance, points located further along the y-axis are interpreted as changes that are most significant. This allows the reader to quickly focus on key changes and ignore the majority of points, which will remain clustered towards the center of the graph abutting the x-axis (Figure 9).
Figure 9.
Volcano plot visualizing fold-changes of RNA-seq data (log2 fold change, x-axis) vs statistical significance (-log10 of p-value, y-axis). The plot is colored such that those points having a fold-change<2 (log2 2= 1) or points having a “-log10 p-value<1.3” (log10 0.05=-1.3) are shown in gray.
These plots can be used to represent radiomic feature sets as well in similar ways. One possible use case would be to represent radiomic features extracted from follow-up scans of a tumor. Since features that change most significantly between scans would be highlighted, they can inform those seeking to identify features related to tumor progression and response that may not be otherwise apparent.
Although a volcano plot is a useful tool for visualizing the results of differential gene expression analysis, it has some limitations. One limitation is that the plot can be difficult to interpret when there are a large number of genes, as it can become cluttered and overwhelming. Additionally, volcano plots are impacted by outliers since they are built on p-values from a t test and fold-change values, both of which rely on classical location and scatter. 26 The plot can also inflate the false discovery rate, especially for features with the highest fold-change. 27
Graphical illustration of survival
Kaplan–Meier curves
Kaplan–Meier (KM) curves can be used to depict prediction of an event occurrence in a specific time period (times-to-event). 28 Although ubiquitous for illustrating predictions of patient or group survival across healthcare domains, the predicted event is not necessarily limited to survival outcomes and can be extended to any type of event, even non-medical fields. 29 Serial time, status at the serial time, and study group are the three main variables used to create a KM curve. Serial time is the time duration defined by the difference between the start point and endpoint dates. The start point of this duration is defined when a subject enters a study, such as the date of diagnosis of a condition, or complication. The endpoint can be the date that our event of interest happened for each subject or the date that the subject is censored from our study. Censoring means that the subject could not reach the endpoint (e.g., loss of follow-up) or pass the study’s end date without getting the endpoint. The endpoint can be the date of death, date of recurrence, or any other events based on the study investigator's decision. The serial time, also known as an “interval” for each subject, is plotted (horizontal lines) in the KM curve from the shortest to the longest duration without considering the study entry date. The censored subjects are depicted as tick marks in KM curves. 29 There are vertical lines in KM curves that connect the horizontal lines. These vertical lines show the changes in overall probability over the period of study. 30
KM curves are commonly used to estimate the overall survival and progression-free survival of patients with different types of cancer who received specific treatment. In oncologic imaging, KM curves can be used to estimate the effect of various imaging features on overall survival and progression-free survival. KM curves allow researchers and clinicians to compare the probability of any type of event, such as survival outcome, tumor response rate, tumor progression, tumor recurrence with specific demographic, or imaging prognostic factors.31,32
Figure 10 shows an example of KM survival analysis for predicting survival of diffuse glioma based on tumor relative cerebral blood volume (rCBV) values. 33 In this study, the median follow-up was 14.5 months (range, 0–76 months), and subjects were divided into two groups based on median rCBV (rCBV ≥5.195 marked with solid line vs rCBV <5.195 marked with dotted line). In this example, KM curve demonstrates that patients with an rCBV lower than the median (<5.195) had significantly better overall survival (Figure 10).
Figure 10.
Kaplan–Meier survival curves of 126 patients with diffuse gliomas. Survival curves are plotted according to the classification based on median rCBV values. Relative cerebral blood volume has a significant influence on overall survival, with a median survival of 11 months for tumors with perfusion values lower than the median rCBV. Used with permission of American Society of Neuroradiology, from AJNR, American journal of neuroradiology, Hilario et al., Vol. 35, Issue 6, Copyright 2014 33 ; permission conveyed through Copyright Clearance Center, Inc.
One limitation of the KM curve is that it does not take into account any covariates or other variables that may influence survival. To account for these variables, a Cox proportional hazards model can be used. 34 Cox proportional hazards model is used to estimate the relationship between the time to an event of interest and a set of predictor variables. It allows us to estimate the effect of predictor variables on the hazard rate. 35 Radiology studies use this model to develop predictive models for survival outcomes, based on a set of predictor variables such as imaging features, biomarkers, or genetic markers.
Another limitation is the fact that KM curve is a non-parametric estimator. In comparison to KM curve, Weibull distribution is a parametric model that assumes a specific functional form for the survival distribution and thus provides more descriptive information about its shape and scale, enabling inter-group comparisons, and yielding a smoother survival curve compared to the step function of the KM curve. 36
Graphical illustrations of treatment effect
Forest plots
Forest plots are mainly used to depict the results of meta-analysis studies (Figure 11). In this type of graph, each study is marked as a box (point estimate) in the middle of a horizontal line (95% confidence interval). The point estimate determines the effect size on the Y-axis. Effect size can be the risk ratio or odds ratio for binary outcomes or the mean difference for continuous outcomes. The box size (area) shows the study weight, representing the extent of information each study provides. The diamond (below and in parallel with the horizontal lines) indicates pooled results of all included studies. A vertical line perpendicular to the middle Y-axis shows “no effect” (1 for risk ratio or odds ratio in case of binary outcomes and 0 for the mean difference in case of continuous outcomes). If the horizontal line (95% confidence interval) from each study or the diamond crosses the “no effect” vertical line, the difference between the studies’ groups is not significantly different. There is another vertical line in the center of the diamond, which corresponds to the pooled value of point estimates. The amount of overlap between the studies (horizontal lines and point estimate) estimates heterogeneity among included studies; more overlap determines less heterogeneity between the studies.37–39 I-squared (I2) is a quantitative measure that reflects the heterogeneity among studies that ranges from 0% (lowest heterogeneity) to 100% (highest heterogeneity).
Figure 11.
Forest-plot of the area under the curve (AUC) of the receiver operator curve (ROC) of the different perfusion metrics in predicting IDH mutation status. IDH, isocitrate dehydrogenase, ktrans, volume transfer coefficient; rCBV, relative cerebral blood volume; Ve, fractional volume of the extravascular extracellular space; Vp, fractional blood plasma volume; 95%-CI, 95%-confidence interval. Reproduced from Van Santwijk et al. 45
In radiology, a forest plot can be used in meta-analysis to present and summarize the results of multiple studies, including the diagnostic performance (accuracy, sensitivity, specificity, positive and negative predictive values) of a specific imaging modality,40,41 the radiologic prognostic factors of disease,42,43 and radiologic evaluation of the effectiveness of various treatments (progression-free survival and overall survival). 44 Forest plot visually represents the result of different studies and allows the reader to compare the results of each study, evaluate the consistency of results across studies, and identify any outliers that may be influencing the overall results.
The main limitation of the forest plot is not showing the potential publication bias, which can be displayed by funnel plots.
Funnel plots
Funnel plots are a type of scatter plot to evaluate publication bias or heterogeneity between studies in a meta-analysis. Several factors can cause publication bias, such as a higher chance of publication for studies that report positive results or a lower chance of publication for a study supported by a drug company that did not show favorable results. Therefore, funnel plots can be used to evaluate the presence of small-study effects, which occur when small sample size studies tend to overestimate effect sizes compared to larger studies. This could help to ensure that the findings of the meta-analysis are as robust as possible and that the conclusions drawn are not impacted by the absence of certain studies. 46 Funnel plots consist of two axes; the x-axis represents the study’s result and is mostly depicted as the mean result log (such as odds ratio, risk ratio, or mean differences). The y-axis represents precision and is mostly shown as standard error or inverse standard error.46,47 In the funnel plot, the overall effect of all included studies in a meta-analysis is illustrated by a vertical line in the middle. There are also two sloped lines that meet each other at the vertical line on top and make a triangle with the x-axis (the reverse funnel). These two lines are indicators of pseudo-confidence intervals, including 95% of studies without any publication bias and heterogeneity. 47
A funnel plot can be used to evaluate the precision of the studies included in meta-analyses of diagnostic test accuracy (sensitivity and specificity) and treatment efficacy (progression-free survival or overall survival) studies. In addition, the funnel plot can evaluate the consistency of results across studies and compare the diagnostic performance of different imaging modalities. 48 For instance, in Figure 12, the funnel plot shows 28 studies included in a meta-analysis to assess the diagnostic accuracy of maximum relative cerebral blood volume to differentiate between glioma grades II and III. 49 In this meta-analysis, mean difference (MD) was considered an estimate of the effect size for each study. The accordant funnel plot visually shows symmetric distribution among included studies, demonstrating no significant publication bias. In addition to the visual inspection of a funnel plot, some statistical tests, such as Begg’s test or Egger’s test, can be applied to evaluate the publication bias. 50
Figure 12.
Funnel plot of 28 included studies (n = 727 patients) illustrated by open circles with the effect estimate mean difference (MD) of rCBVmax plotted on the horizontal axis, the standard error (SE) of the MD plotted on the vertical axis, and a triangular 95% confidence region. The study distribution is symmetric without apparent publication bias. Used with permission of American Society of Neuroradiology, from AJNR, American journal of neuroradiology, Delgado et al., Vol. 38, Issue 7, Copyright 2017 49 ; permission conveyed through Copyright Clearance Center, Inc.
One limitation of the funnel plot is that it is purely visual and subjective, which can cause misinterpretation, especially when the number of studies is small. 51
Violin plots
The violin plot combines the box plot and Kernel density or histogram plots to illustrate numeric data. The main specification of this type of graph is showing the distributional features of various sets of data. Violin plots consist of two axes; the x-axis shows different study groups, and the y-axis shows the quantitative variable which aims to be assessed. Similar to the box plot, the median and interquartile range (IQR) can be marked in violin plots. 52 A violin plot can be used in radiology studies to display the distribution of certain image features, such as intensity or texture, within a specific region of interest (ROI), and compare the distribution of image features or measurements across multiple patients or studies, which could help to identify patterns or trends that could inform future research or diagnostic decisions. 53 Another application in imaging studies could be the usage of violin plots for comparing radiologic indices obtained at different time points. For example, Figure 13 is a violin plot that compares various perfusion metrics (cerebral blood flow, bolus arrival time, and Buxton-modeled cerebral blood flow) across three different time points. 54
Figure 13.
Violin plots showing changes in perfusion metrics across time points for all patients’ brain regions. The horizontal line within the plot indicates the median. Darker data points indicate patients with CMS. A, CBF derived from single-PLD ASL. B, BAT derived from multi-TI ASL. C, CBF derived from multi-TI ASL. Horizontal significance bars show false discovery rate–adjusted P values from t test pair-wise comparisons (degree sign indicates P < .1; asterisk, P < .05). Pre-op indicates preoperative; Post-op, postoperative. Used with permission of American Society of Neuroradiology, from AJNR, American journal of neuroradiology, Toescu et al., Vol. 43, Issue 10, Copyright 2022 54 ; permission conveyed through Copyright Clearance Center, Inc.
Additionally, violin plots could be used to compare the distribution of radiologic tumor response to the treatment, such as progression-free survival or overall survival, across different patient subgroups or treatment regimens, to identify factors that may influence treatment response, such as patient characteristics, tumor radiologic features, or specific treatment components. 55
Although violin plots can depict data distribution, the frequency of data cannot be visualized. Moreover, they have a tendency to overestimate the impact of extreme outliers. 14
Graphical illustrations with dimensionality reduction and clustering
Principal component plot
Radiomic and radiogenomic studies that utilize artificial intelligence (AI) and machine learning methods often include high-dimensional datasets (the number of variables is close to or larger than the number of samples), which can include up to hundreds or thousands of extracted variables.56,57 To visualize or analyze high-dimensional data, dimensionality reduction methods can be used for transforming the data such that it can be shown in 2D or 3D plots. One technique is principal components analysis (PCA), which captures the essence of the data in a few principal components (PCs) based on the variance of the samples. The top PCs are those that account for the largest variance, and plotting the top PCs can allow identification of patterns in the dataset. 58
Principal component plots are scatter plots that show the top 2 or 3 PCs, with one on each axis (Figure 14). Original data points are projected to the space by transforming their values into PC scores with the fitted model. Data points which lie closer together are more similar, and those farther apart are more dissimilar. This can reveal natural clusters (spatial groupings of data points), which can be compared to known categories. For example, the top PCs based on extracted radiomic features can be plotted and data points can be labeled based on clinical or genomic factors, such as groups defined by high or low survival risk, treatment types or trial arm assignment, or presence/absence of a gene mutation (Figure 15).59–61 This allows the visual assessment of whether the PCs based on imaging features are related to these properties. Clusters (classes/labels) are usually denoted by different colors and/or symbols.
Figure 14.
An illustration of the perfusion time-series in tumorous subregions, that is, ET, NC, and ED (A); and the clustering of each tissue type using PC analysis (B), signifying the potential of the PCs in capturing tissue characteristics. PC1, PC2, and PC3 represent the first, second, and third principal components, respectively. ET Enhancing tumor, NC Necrotic core, ED paeritumoral edema. Reproduced from Akbari et al. 62
Figure 15.
Clustering projection and illustrative images. In the top chart, the final imaging-based clustering results are depicted here with each point representing a unique subject plotted against the first two principal components (PCs). Each color represents a cluster group (Cluster 1: Blue; Cluster 2: Orange; Cluster 3: Red). In this analysis, the first two PCs explain only 25 percent of the variance in the feature set which may explain the proximity of the clusters on this projection. Although the true clustering is done on 48 dimensions, separation of the subjects can be appreciated even on this two-dimensional projection of the data. Below the chart, representative images were selected from the T2 Axial MR images from 4 patients in each cluster. These patients were picked from the center-most regions of each cluster and can thus be presumed to be most representative of their groups. Although the full volume of tumor from all 4 modalities (T1 pre-contrast, T1 post-contrast, T2, and FLAIR) was utilized for this work, for illustrative purposes only the axial T2 slice demonstrating the largest diameter of tumor was selected for this figure. Reprinted from Neoplasia, Vol. 36, Haldar et al., Unsupervised machine learning using K-means identifies radiomic subgroups of pediatric low-grade gliomas that correlate with key molecular markers, Pages 100869, Copyright 2023, with permission from Elsevier. 21
One important limitation is that PCs are new variables that are linear combinations of the features, they do not correspond to any original features and so they may be less interpretable. Additionally, PCA has assumptions about the data which must be met in order for it to be utilized properly, and if more than two or three PCs are needed to explain the majority of the variance, it may not be informative to visualize these PCs.
Cluster plots (K-means)
Cluster plots are useful to assess whether and how plotted variables can split data samples into groups. These groups can be known a priori or determined via clustering analysis methods. 63 Cluster plots are scatter plots, where X and Y-axes correspond to different numerical variables and each point represents a data sample, with the addition that colors and/or symbols are used to depict the different groups (Figure 15). For high dimensional data, cluster analysis may be done on the principal components after performing dimension reduction via PCA.
Clustering methods are useful for determining meaningful groupings of unlabeled data samples and are prevalent in exploratory data analysis. One of the most widely used methods is K-means, which can be used to assign data points into spatially distinct groups.64,65 In K-means clustering, the number of resulting clusters (K) is fixed, and samples are iteratively grouped into clusters until there is a maximal separation between cluster centers, with minimal distance between points within a cluster and the cluster center (centroid). The general overlap between clusters and their constituent points indicates their separability, with more separated clusters forming more distinct groups, which is an indication of how well the variables can partition the data samples into the groups. Sometimes the cluster centroids are depicted with a unique color or symbol.
Note that although this plotting approach allows visualization of groupings based on two or three variables, it can also be an over-simplification as datasets may be better explained by more variables and a higher-dimensional space. Additionally, there may be no naturally occurring clusters in a dataset, in which case clustering methods would not be useful for interpretation.
Newer dimension reduction and displaying techniques: t-SNE and UMAP
Many genomic, molecular, and radiomic analyses involve high-dimensional data. Two of the newer methods for performing dimension reduction and plotting the results are t-SNE (t-distributed stochastic neighbor embedding) from 2008 and UMAP (Uniform Manifold Approximation and Projection) from 2018.66,67 Both techniques perform non-linear scaling of the data, whereas PCA explores relationships. t-SNE is a dimensionality reduction statistical method for visualizing high-dimensional data by giving each datapoint a location in 2D or 3D space. The technique finds clusters in data while reducing data dimensions, by trying to keep similar data close and dissimilar data apart in the graph. It has been used in a variety of medical and non-medical applications, but examples of use in neuro-oncology have been in analyzing tumor RNA-seq data, methylation profiles, and imaging-based texture analysis (Figure 16).68–70 t-SNE-based tumor classifications and data-driven identification of prognostic tumor subpopulations have influenced the newest classifications of brain tumors.71,72
Figure 16.
t-Distributed stochastic neighbor embedding (t-SNE) analysis of DNA methylation profiles of the investigated tumors alongside selected reference samples. Reference DNA methylation classes: high-grade astrocytoma with piloid features (ANA PA); diffuse high-grade glioma, H3.3 G34 mutant (DHG H3 G34); diffuse midline glioma H3 K27M mutant (DMG H3 K27); pediatric glioblastoma, IDH wildtype, subclass MYCN (GB pedMYCN); pediatric glioblastoma, IDH wildtype, subclass not otherwise specified sutbype A (GB pedNOS A); pediatric glioblastoma, IDH wildtype, subclass not otherwise specified sutbype B (GB pedNOS B); pediatric glioblastoma, IDH wildtype, subclass RTK1a (GB pedRTK1a); pediatric glioblastoma, IDH wildtype, subclass RTK1b (GB pedRTK1b); pediatric glioblastoma, IDH wildtype, subclass RTK1c (GB pedRTK1c); pediatric glioblastoma, IDH wildtype, subclass RTK2a (GB pedRTK2a); pediatric glioblastoma, IDH wildtype, subclass RTK2b (GB pedRTK2b); infant-type hemispheric glioma (IHG); hemispheric pilocytic astrocytoma (PA CORT); infratentorial pilocytic astrocytoma (PA INF); midline pilocytic astrocytoma (PA MID); pleomorphic xanthoastrocytoma (PXA). Reproduced from Guerrini-Rousseau et al. 73
UMAP is a new technique for dimensionality reduction and display, which on the surface looks very similar to t-SNE. However, it is a computationally much faster method with resultant better scalability to very large datasets compared to t-SNE, and it better preserves the global structure of the data. Areas of application are similar to that of t-SNE, and it has rapidly gained acceptance as a method for analysis and discovery in tumor biology (Figure 17).
Figure 17.
(a) UMAP of pediatric tumors and adult glioma subtypes from TCGA. Coloring in UMAP of pediatric tumors and adult glioma subtypes from TCGA by (b) number of point mutations and (c) number of gene fusions per tumor (d) number of genes with copies gained per tumor and (e) number of genes with copies deleted per tumor. Reproduced from Arora et al. 74
Both t-SNE and UMAP are extensively used in genomic and molecular studies. They have only been recently applied to radiological imaging and radiomic analyses, although it is expected that their usage will increase as larger imaging datasets are explored and analyzed. Potential challenges and pitfalls of these techniques include the difficulty of selecting optimal hyperparameters for the analysis, which can dramatically affect the results and visualization.
Conclusion
In the present paper, we have reviewed several types of statistical plots that are common in oncologic imaging research studies for visualizing data and analytical results. In addition to improving the clarity of one’s findings for publishing or presentation, the visual depiction of data also benefits our comprehension of underlying distributions and relationships in meaningful ways. When exploring raw data via numerical information alone, it would be challenging to discern and comprehend complex structures within the information. Data visualization can help to make these structures clear, allowing us to find patterns and discover new insights.
A growing amount of contemporary oncologic imaging, radiomic, and radiogenomic research depends on the conveyance of complicated data through a variety of graphs and plots. Although these plots are meant to present data in an easily digestible fashion, sometimes, they can be hard to interpret to the untrained eye. Therefore, radiologists must learn the newly developed graphical imagery and vocabulary to facilitate simple and comfortable communication. By understanding the advantages and limitations of statistical plots, radiologists will be able to choose the ones that will best highlight the findings of their research through improved data delivery.
Currently, there are several statistical software packages that enable researchers to prepare complex plots. However, new statistical plots with enhanced features are needed as more complex data are produced with the advancement of oncologic imaging studies, including those which utilize AI methods with large feature sets. Such plots and statistical software packages have the potential to be vastly employed if they are highly informative and user-friendly. In the words of Ivan Turgenev (1818–1883), “The drawing shows me at one glance what might be spread over ten pages in a book.”
Footnotes
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding: The author(s) received no financial support for the research, authorship, and/or publication of this article.
ORCID iD
Ali Nabavizadeh https://orcid.org/0000-0002-0380-4552
References
- 1.Schriger DL, Sinha R, Schroter S. and et al. From submission to publication: a retrospective review of the tables and figures in a cohort of randomized controlled trials submitted to the British Medical Journal. Ann Emerg Med 2006; 48(6): 750–756. DOI: 10.1016/j.annemergmed.2006.06.017. [DOI] [PubMed] [Google Scholar]
- 2.Tufte ER. The visual display of quantitative information. Graphics Press, 1983. [Google Scholar]
- 3.Polepalli Ramesh B, Sethi RJ, Yu H. Figure-Associated Text Summarization and Evaluation. PLoS One 2015; 10: e0115671. DOI: 10.1371/journal.pone.0115671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Pocock SJ, Travison TG, Wruck LM. Figures in clinical trial reports: current practice & scope for improvement. Trials 2007; 8: 36. DOI: 10.1186/1745-6215-8-36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Franzblau LE, Chung KC. Graphs, tables, and figures in scientific publications: the good, the bad, and how not to be the latter. J Hand Surg Am 2012; 37: 591–596. DOI: 10.1016/j.jhsa.2011.12.041. [DOI] [PubMed] [Google Scholar]
- 6.Durbin CG., Jr. Effective use of tables and figures in abstracts, presentations, and papers. Respir Care 2004; 49: 1233–1237. [PubMed] [Google Scholar]
- 7.Gillespie TW. Understanding waterfall plots. J Adv Pract Oncol 2012; 3: 106–111. [PMC free article] [PubMed] [Google Scholar]
- 8.Sun Q, Chen Y, Liang C, et al. biologic pathways underlying prognostic radiomics phenotypes from paired MRI and RNA sequencing in glioblastoma. Radiology 2021; 301: 654–663. DOI: 10.1148/radiol.2021203281. [DOI] [PubMed] [Google Scholar]
- 9.Bismeijer T, Van Der Velden BHM, Canisius S, et al. radiogenomic analysis of breast cancer by linking MRI phenotypes with tumor gene expression. Radiology 2020; 296: 277–287. DOI: 10.1148/radiol.2020191453. [DOI] [PubMed] [Google Scholar]
- 10.Lorusso PM, Anderson AB, Boerner SA, et al. Making the investigational oncology pipeline more efficient and effective: are we headed in the right direction? Clin Cancer Res 2010; 16: 5956–5962. DOI: 10.1158/1078-0432.ccr-10-1279. [DOI] [PubMed] [Google Scholar]
- 11.Kim MS, Prasad V. Assessment of accuracy of waterfall plot representations of response rates in cancer treatment published in medical journals. JAMA Netw Open 2019; 2: e193981. DOI: 10.1001/jamanetworkopen.2019.3981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Mercier F, Consalvo N, Frey N, et al. From waterfall plots to spaghetti plots in early oncology clinical development. Pharm Stat 2019; 18: 526–532. DOI: 10.1002/pst.1944. [DOI] [PubMed] [Google Scholar]
- 13.Castanon Alvarez E, Aspeslagh S, Soria JC. 3D waterfall plots: a better graphical representation of tumor response in oncology. Ann Oncol 2017; 28: 454–456. DOI: 10.1093/annonc/mdw656. [DOI] [PubMed] [Google Scholar]
- 14.Chia PL, Gedye C, Boutros PC, et al. Current and evolving methods to visualize biological data in cancer research. J Natl Cancer Inst 2016; 108: djw031. DOI: 10.1093/jnci/djw031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Vickers AJ. The use of percentage change from baseline as an outcome in a controlled trial is statistically inefficient: a simulation study. BMC Med Res Methodol 2001; 1: 87. DOI: 10.1186/1471-2288-1-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Phillips S. Swimmer plot: tell a graphical story of your time to response data using PROC SGPLOT. PharmaSUG, San Diego Available at http://www.pharmasugorg/proceedings/2014/DG/PharmaSUG-2014-DG07pdf 2014 [Google Scholar]
- 17.Lythgoe MP, Olivier T, Prasad V. The iceberg plot, improving the visualisation of therapy response in oncology in the era of sequence-directed therapy. Eur J Cancer 2021; 159: 56–59. DOI: 10.1016/j.ejca.2021.09.034. [DOI] [PubMed] [Google Scholar]
- 18.Werner JM, Wolf L, Tscherpel C, et al. Efficacy and tolerability of regorafenib in pretreated patients with progressive CNS grade 3 or 4 gliomas. J Neuro Oncol 2022; 159: 309–317. DOI: 10.1007/s11060-022-04066-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Schroeder MP, Gonzalez-Perez A, Lopez-Bigas N. Visualizing multidimensional cancer genomics data. Genome Med 2013; 5: 9. DOI: 10.1186/gm413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ewing AD, Houlahan KE, Hu Y, et al. Combining tumor genome simulation with crowdsourcing to benchmark somatic single-nucleotide-variant detection. Nat Methods 2015; 12: 623–630. DOI: 10.1038/nmeth.3407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Haldar D, Kazerooni AF, Arif S, et al. Unsupervised machine learning using K-means identifies radiomic subgroups of pediatric low-grade gliomas that correlate with key molecular markers. Neoplasia 2023; 36: 100869. DOI: 10.1016/j.neo.2022.100869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hu LS, Wang L, Hawkins-Daarud A, et al. Uncertainty quantification in the radiogenomics modeling of EGFR amplification in glioblastoma. Sci Rep 2021; 11: 3489. DOI: 10.1038/s41598-021-83141-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Krzywinski M, Schein J, Birol İ, et al. Circos: an information aesthetic for comparative genomics. Genome Res 2009; 19: 1639–1645. DOI: 10.1101/gr.092759.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zanfardino, Pane, Mirabelli, et al. TCGA-TCIA Impact on Radiogenomics Cancer Research: A Systematic Review. Int J Mol Sci 2019; 20: 6033. DOI: 10.3390/ijms20236033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Li W, Freudenberg J, Suh YJ, et al. Using volcano plots and regularized-chi statistics in genetic association studies. Comput Biol Chem 2014; 48: 77–83. DOI: 10.1016/j.compbiolchem.2013.02.003. [DOI] [PubMed] [Google Scholar]
- 26.Kumar N, Hoque MA, Sugimoto M. Robust volcano plot: identification of differential metabolites in the presence of outliers. BMC Bioinf 2018; 19: 128. DOI: 10.1186/s12859-018-2117-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ebrahimpoor M, Goeman JJ. Inflated false discovery rate due to volcano plots: problem and solutions. Brief Bioinform 2021: 22: bbab053. DOI: 10.1093/bib/bbab053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kaplan EL, Meier P. Nonparametric estimation from incomplete observations. J Am Stat Assoc 1958; 53: 457. DOI: 10.2307/2281868. [DOI] [Google Scholar]
- 29.Rich JT, Neely JG, Paniello RC, et al. A practical guide to understanding Kaplan-Meier curves. Otolaryngol Head Neck Surg 2010; 143: 331–336. DOI: 10.1016/j.otohns.2010.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Feinstein AR. Clinical epidemiology : the architecture of clinical research. Philadelphia, PA: W.B. Saunders Co., 1985. [Google Scholar]
- 31.Park SH, Han K, Park SY. Mistakes to avoid for accurate and transparent reporting of survival analysis in imaging research. Korean J Radiol 2021; 22: 1587–1593. DOI: 10.3348/kjr.2021.0579.20210819 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Stolberg HO, Norman G, Trop I. Survival analysis. AJR Am J Roentgenol 2005; 185: 19–22. DOI: 10.2214/ajr.185.1.01850019. [DOI] [PubMed] [Google Scholar]
- 33.Hilario A, Sepulveda JM, Perez-Nunez A, et al. A prognostic model based on preoperative MRI predicts overall survival in patients with diffuse gliomas. Am J Neuroradiol 2014; 35: 1096–1102. DOI: 10.3174/ajnr.a3837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Dudley WN, Wickham R, Coombs N. An introduction to survival statistics: kaplan-meier analysis. J Adv Pract Oncol 2016; 7: 91–100. DOI: 10.6004/jadpro.2016.7.1.8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Deo SV, Deo V, Sundaram V. Survival analysis-part 2: cox proportional hazards model. Indian J Thorac Cardiovasc Surg 2021; 37: 229–233. DOI: 10.1007/s12055-020-01108-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wan X, Peng L, Li Y. A review and comparison of methods for recreating individual patient data from published kaplan-meier survival curves for economic evaluations: a simulation Study. PLoS One 2015; 10: e0121353. DOI: 10.1371/journal.pone.0121353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Dettori JR, Norvell DC, Chapman JR. Seeing the forest by looking at the trees: how to interpret a meta-analysis forest Plot. Global Spine J 2021; 11: 614–616. DOI: 10.1177/21925682211003889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sedgwick P. How to read a forest plot in a meta-analysis. BMJ 2015; 351: h4028. DOI: 10.1136/bmj.h4028. [DOI] [PubMed] [Google Scholar]
- 39.Stephenson J. Explaining the forest plot in meta-analyses. J Wound Care 2017; 26: 611–612. DOI: 10.12968/jowc.2017.26.11.611. [DOI] [PubMed] [Google Scholar]
- 40.Ye Y, Yang Y, Gong J, et al. Comparing the diagnostic accuracy of PET and CMR for the measurement of left ventricular volumes and ejection fraction: a system review and meta-analysis. Nucl Med Commun 2022; 43: 1143–1154. DOI: 10.1097/mnm.0000000000001612. [DOI] [PubMed] [Google Scholar]
- 41.Lim SJ, Suh CH, Shim WH, et al. Diagnostic performance of T2* gradient echo, susceptibility-weighted imaging, and quantitative susceptibility mapping for patients with multiple system atrophy-parkinsonian type: a systematic review and meta-analysis. Eur Radiol 2022; 32: 308–318. DOI: 10.1007/s00330-021-08174-4. [DOI] [PubMed] [Google Scholar]
- 42.Kao YS, Lin KT. A meta-analysis of the diagnostic test accuracy of CT-based radiomics for the prediction of COVID-19 severity. Radiol Med 2022; 127: 754–762. DOI: 10.1007/s11547-022-01510-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hashemi-Madani N, Emami Z, Janani L, et al. Typical chest CT features can determine the severity of COVID-19: A systematic review and meta-analysis of the observational studies. Clin Imaging 2021; 74: 67–75. DOI: 10.1016/j.clinimag.2020.12.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhou W, Yue Y, Zhang X. Radiotherapy plus chemotherapy leads to prolonged survival in patients with anaplastic thyroid cancer compared with radiotherapy alone regardless of surgical resection and distant metastasis: a retrospective population study. Front Endocrinol 2021; 12: 748023. DOI: 10.3389/fendo.2021.748023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Van Santwijk L, Kouwenberg V, Meijer F, et al. A systematic review and meta-analysis on the differentiation of glioma grade and mutational status by use of perfusion-based magnetic resonance imaging. Insights into Imaging 2022; 13: 102. DOI: 10.1186/s13244-022-01230-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Hoffman JIE. Chapter 36 - Meta-analysis. In: Hoffman JIE. (ed). Basic biostatistics for medical and biomedical practitioners 2nd ed. Academic Press, 2019, pp. 621–629. [Google Scholar]
- 47.Sterne JA, Sutton AJ, Ioannidis JP, et al. Recommendations for examining and interpreting funnel plot asymmetry in meta-analyses of randomised controlled trials. BMJ 2011; 343: d4002. DOI: 10.1136/bmj.d4002. [DOI] [PubMed] [Google Scholar]
- 48.Wang X, Yang L, Wang Y. Meta-analysis of the diagnostic value of (18)F-FDG PET/CT in the recurrence of epithelial ovarian cancer. Front Oncol 2022; 12: 1003465. DOI: 10.3389/fonc.2022.1003465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Delgado AF, Delgado AF. Discrimination between Glioma Grades II and III Using dynamic susceptibility perfusion MRI: a meta-analysis. Am J Neuroradiol 2017; 38: 1348–1355. DOI: 10.3174/ajnr.a5218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Tatsioni A, Ioannidis JPA. Meta-analysis. In: Quah SR. (ed). International encyclopedia of public health. 2nd ed. Oxford: Academic Press, 2017, pp. 117–124. [Google Scholar]
- 51.Simmonds M. Quantifying the risk of error when interpreting funnel plots. Syst Rev 2015; 4: 24. DOI: 10.1186/s13643-015-0004-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Hintze JL, Nelson RD. Violin plots: a box plot-density trace synergism. Am Statistician 1998; 52: 181–184. DOI: 10.1080/00031305.1998.10480559. [DOI] [Google Scholar]
- 53.Liu Q, Huang Y, Chen H, et al. The development and validation of a radiomic nomogram for the preoperative prediction of lung adenocarcinoma. BMC Cancer 2020; 20: 533. DOI: 10.1186/s12885-020-07017-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Toescu SM, Hales PW, Cooper J, et al. Arterial spin-labeling perfusion metrics in pediatric posterior fossa tumor surgery. Am J Neuroradiol 2022; 43: 1508–1515. DOI: 10.3174/ajnr.a7637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Pantano F, Zalfa F, Iuliani M, et al. Large-scale profiling of extracellular vesicles identified miR-625-5p as a novel biomarker of immunotherapy response in advanced non-small-cell lung cancer patients. Cancers 2022; 14: 20220514. DOI: 10.3390/cancers14102435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Gillies RJ, Kinahan PE, Hricak H. Radiomics: images are more than pictures, they are data. Radiology 2016; 278: 563–577. DOI: 10.1148/radiol.2015151169.20151118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Lambin P, Leijenaar RTH, Deist TM, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol 2017; 14: 749–762. DOI: 10.1038/nrclinonc.2017.141. [DOI] [PubMed] [Google Scholar]
- 58.Ringnér M. What is principal component analysis? Nat Biotechnol 2008; 26: 303–304. DOI: 10.1038/nbt0308-303. [DOI] [PubMed] [Google Scholar]
- 59.Kickingereder P, Burth S, Wick A, et al. Radiomic profiling of glioblastoma: identifying an imaging predictor of patient survival with improved performance over established clinical and radiologic risk models. Radiology 2016; 280: 880–889. DOI: 10.1148/radiol.2016160845. [DOI] [PubMed] [Google Scholar]
- 60.Fathi Kazerooni A, Saxena S, Toorens E, et al. Clinical measures, radiomics, and genomics offer synergistic value in AI-based prediction of overall survival in patients with glioblastoma. Sci Rep 2022; 12: 8784. DOI: 10.1038/s41598-022-12699-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Madhogarhia R, Haldar D, Bagheri S, et al. Radiomics and radiogenomics in pediatric neuro-oncology: a review. Neurooncol Adv 2022; 4. DOI: 10.1093/noajnl/vdac083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Akbari H, Kazerooni AF, Ware JB, et al. Quantification of tumor microenvironment acidity in glioblastoma using principal component analysis of dynamic susceptibility contrast enhanced MR imaging. Sci Rep 2021; 11: 15011. DOI: 10.1038/s41598-021-94560-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Rizzo S, Botta F, Raimondi S, et al. Radiomics: the facts and the challenges of image analysis. Eur Radiol Exp 2018; 2: 36. DOI: 10.1186/s41747-018-0068-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Laudicella R, Agnello L, Comelli A. Unsupervised brain segmentation system using k-means and neural network. In: Mazzeo PL, Frontoni E, Sclaroff S, et al. (eds). Image Analysis and Processing ICIAP 2022 Workshops. Cham, 2022. Springer International Publishing, pp. 441–449. [Google Scholar]
- 65.Rathore S, Akbari H, Rozycki M, et al. Radiomic MRI signature reveals three distinct subtypes of glioblastoma with different clinical and molecular characteristics, offering prognostic value beyond IDH1. Sci Rep 2018; 8: 5087. DOI: 10.1038/s41598-018-22739-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Van der Maaten L, Hinton G. Visualizing data using t-SNE. J Mach Learn Res 2008; 9: 2579–2605. [Google Scholar]
- 67.McInnes L, Healy J, Melville J. Umap: uniform manifold approximation and projection for dimension reduction. arXiv preprint arXiv:180203426 2018. DOI: 10.48550/arXiv.1802.03426. [DOI]
- 68.Kobak D, Berens P. The art of using t-SNE for single-cell transcriptomics. Nat Commun 2019; 10: 5416. DOI: 10.1038/s41467-019-13056-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Kresbach C, Bronsema A, Guerreiro H, et al. Long-term survival of an adolescent glioblastoma patient under treatment with vinblastine and valproic acid illustrates importance of methylation profiling. Childs Nerv Syst 2022; 38: 479–483. DOI: 10.1007/s00381-021-05278-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Yu Y, Wu X, Chen J, et al. Characterizing brain tumor regions using texture analysis in magnetic resonance imaging. Front Neurosci 2021; 15: 634926–20210603. DOI: 10.3389/fnins.2021.634926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Abdelmoula WM, Balluff B, Englert S, et al. Data-driven identification of prognostic tumor subpopulations using spatially mapped t-SNE of mass spectrometry imaging data. Proc Natl Acad Sci U S A 2016; 113: 12244–12249. DOI: 10.1073/pnas.1510227113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Sturm D, Orr BA, Toprak UH, et al. New brain tumor entities emerge from molecular classification of CNS-PNETs. Cell 2016; 164: 1060–1072. DOI: 10.1016/j.cell.2016.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Guerrini-Rousseau L, Tauziède-Espariat A, Castel D, et al. Pediatric high-grade glioma MYCN is frequently associated with Li-Fraumeni syndrome. Acta Neuropathol Commun 2023; 11: 3–20. DOI: 10.1186/s40478-022-01490-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Arora S, Szulzewsky F, Jensen M, et al. An RNA seq-based reference landscape of human normal and neoplastic brain. Res Sq 2023. DOI: 10.21203/rs.3.rs-2448083/v1. [DOI] [PMC free article] [PubMed] [Google Scholar]