Abstract
Objective
Clinical research networks facilitate collaborative research, but data sharing remains a common barrier.
Materials and Methods
The TriNetX platform provides real-time access to electronic health record (EHR)-derived, anonymized data from 173 healthcare organizations (HCOs) and tools for queries and analysis. In 2022, 4 pediatric HCOs worked with TriNetX leadership to found the Pediatric Collaboratory Network (PCN), facilitated via a multi-institutional data-use agreement (DUA). The DUA enables collaborative study design and execution, with institutional review board-approved transfer of complete datasets for further analyses on a per-protocol basis.
Results and Discussion
Of the 41.2 million children with TriNetX records, the PCN represents nearly 10%. The PCN assisted several early-career investigators to bring study concepts from conception to an international scientific meeting presentation and journal submission.
Conclusion
The PCN facilitates EHR vendor-agnostic multicenter pediatric research on the global TriNetX platform. Continued growth of the PCN will advance knowledge in pediatric health.
Keywords: pediatric, electronic health record, real-world data
Objective
Over the past 50 years, Pediatric Clinical Research Networks (PCRNs) have proliferated. Defined as “an organization of clinical sites and investigators that conducts or intends to conduct multiple collaborative research protocols,”1 PCRNs are vital to facilitate cross-institutional collaboration and enable sharing clinical data. Collaborative pediatric research permits rapid advancement in the treatment and management of conditions, particularly of rare diseases.
The TriNetX Pediatric Collaboratory Network (PCN) brings together healthcare organizations (HCOs) that contribute data to the TriNetX Global Health Research Network to share knowledge and enable pediatric-focused partnerships with limited institutional information technology (IT) support.
Background and significance
PCRNs have grown in scope, size, and volume with over 70 clinical research networks focusing on different clinical sub-specialties and geographical locations to study novel treatments for childhood disease. Among the first multi-institutional PCRNs in the United States was the National Cancer Institute’s (NCI) Clinical Trials Cooperative Group established in 19552 that provided a wealth of knowledge as “…one of the first comparative studies in the chemotherapy of malignant neoplastic disease,”3 while revealing complexities and challenges associated with multi-site research. These included logistical concerns, need for meetings to establish uniformity of procedures and interpretation of protocols, and the financial costs of planning and conducting larger-scale trials. In addition, the authors emphasized the critically important mechanics of a cooperative study conducted at different sites.3 Over the next few decades, other PCRNs were formed including the Children’s Oncology Group (1955),4 the Pediatric Rheumatology Collaborative Study Group (1973),5 the Rare Diseases Clinical Research Network (2003),6 the Pediatric Emergency Care Applied Research Network (2009),7 PEDSnet (2009),8 the Pediatric Trials Network (2010),9 and the Standardized Health data and Research Exchange (2018).10
Despite widespread adoption of PCRNs and their role in fostering collaboration and coherence, challenges remain. A survey of 70 PCRNs compared features of PCRNs and the perceived benefits of PCRN membership.11 Although 73% of those surveyed acknowledged “tangible benefits of such a coalition” (most notably collaboration on study proposals [60%], advocacy for networks [58%], connections with pediatric organizations [54%], and strategies for addressing institutional review board (IRB) issues [48%]), the telephone interviews revealed significant barriers to implementation and continuity. Of note, those interviewed expressed concerns related to (1) funding challenges (eg, funding core activities, individual studies, and cross-subsidization of PCRN expenses with local funds), (2) data management (eg, data flow, standardization, and informatics), (3) governance (standardization of protocol review and implementation, management of expectations, and review of external proposals), (4) inter-network relationships, and (5) ethical issues (eg, lack of local pediatric expertise).
The paucity of reusable and reliable platforms and infrastructure to facilitate data sharing via a collaborative network is a common barrier to multi-institutional collaboration. Such initiatives rely on grant funding that limits data sharing to focused disease-relevant domains. Furthermore, few platforms include both inpatient and outpatient data generated through real-world clinical care, and participating sites must often maintain significant IT technical support. Therefore, novel approaches are needed to address operational barriers.
In 2022, the PCN founding sites, Children’s National Hospital, West Virginia University Medicine Children’s, Children’s Health (TX), Johns Hopkins All Children’s Hospital (FL), and Johns Hopkins Children’s Center (MD) decided to address these challenges by leveraging their existing relationships with TriNetX (Figure 1). TriNetX was initially created to address problems and obstacles inherent in clinical trial protocol design, cohort identification, cohort analysis, study feasibility, and site selection processes and sought to imbue a more data-driven workflow into these activities.12,13 HCOs join TriNetX to address 3 primary use cases: (1) access to the TriNetX query and analytic tools to explore the de-identified patient data at their own HCO, (2) the ability to receive sponsored clinical trial opportunities from life sciences companies who query a network of participating HCOs, and (3) ease of inter-organizational collaboration and data querying due to the harmonization of HCO data to a common TriNetX data model. From its start in 2015, the TriNetX Global Health Research Network has expanded to comprise EHR vendor-agnostic data from approximately 214 million unique patients (of which over 41 265 348 are pediatric patients ≤21 years old) from across 173 HCOs worldwide.
Figure 1.
Map of current PCN member healthcare organizations (HCO). Tan: Children’s National Hospital; Yellow: West Virginial University Medicine Children’s; Red: Children’s Health; Blue: Johns Hopkins All Children’s Hospital/Johns Hopkins Children’s Center.
Methods
TriNetX platform
The TriNetX platform is a global, federated network where users can access continuously updated, anonymized electronic health records (EHR) data, including demographics, encounters, diagnoses, procedures, medications, laboratory values, vitals, and genomics. Healthcare organizations with limited IT and data resources benefit from the data harmonization and TriNetX’s EHR-agnostic infrastructure, which simplifies secure access to information shared from their data warehouses and EHRs. The TriNetX platform includes self-service tools that allow patient data to be aggregated and queried based on the inclusion/exclusion criteria for a given research project. Users can perform cohort comparison and analysis across clinical profiles and longitudinal measures. Publications drawing on TriNetX have demonstrated utility in answering research questions and gaining access to representative cohorts across diverse care settings.
PCN framework and infrastructure
The PCN, founded in 2022, permits federated pediatric queries across sites in TriNetX to facilitate collaboration between investigators and subject-matter experts with similar interests. As active contributors to the network, each PCN member applies a data-driven approach to their work, repurposing EHR data for clinical investigation. Although aggregated data from member sites is visible in the TriNetX Global Health Research Network, the “added value” of the PCN master data-use agreement (DUA) is to improve collaborations by making it simpler for investigators to meet experts at other PCN member sites. The PCN is visible in the global TriNetX portal as a selectable network, and results are returned immediately with patient counts and abundant clinical data for each participating site (Figure 2 and Table 1). A key feature is that no additional IRB reviews are needed to participate in or query the PCN. However, if a specific collaboration requires collection and sharing of additional data not already provided to TriNetX, IRB approval or a letter of exemption is necessary. A steering committee of institutionally designated representatives, including biostatistics experts, from the 4 core sites and 2 TriNetX representatives meets monthly to discuss data governance, coordination efforts, and new site engagement. To mitigate barriers to entry, the steering committee designed the membership process to be inclusive while ensuring that the structure and underlying mission of the PCN is maintained. Steering committee members can invite junior faculty to present research concepts and ongoing work that could benefit from PCN participation and PCN-based analyses. Investigators working under the master DUA can obtain aggregated counts at each of the PCN sites and ask the established point of contact at PCN sites for introductions to potential collaborators or learn how to share additional information unavailable through TriNetX. This might include, subject to IRB approval, obtaining de-identified data or validation of data at participating sites
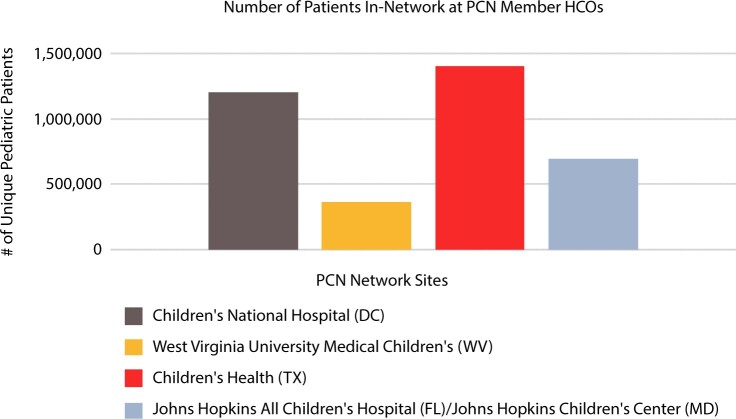
Figure 2.
Number of unique pediatric patients contributing data to PCN member healthcare organizations (HCOs).
Table 1.
An overview of the salient standards used in the TriNetX Pediatric Collaboratory Network, including associated sources and quantity of clinical observations (as of June 15, 2023).
| Data type | Source vocabulary | Number of facts |
|---|---|---|
| Diagnosis | ICD-10-CM | 83 000 766 |
| Procedures | ICD-10-PCS, CPT, HCPCS | 82 685 211 |
| Medications and vaccinations | RxNorm, OMOP RxNorm Extensions, CVX | 1 166 344 181 |
| Lab results and clinical findings | LOINC | 218 619 900 |
| Vital signs | LOINC | 116 212 872 |
| Oncology and chemotherapy lines | NAACCR, ICD-O topography and morphology, AJCC, SEER, etc | 94 377 |
| Total | 1 666 957 307 |
Abbreviations: AJCC, American Joint Committee on Cancer; CPT: Current Procedural Terminology; HCPCS, Healthcare Common Procedure Coding System; ICD-10-CM, International Classification of Diseases, Tenth Revision, Clinical Modification; ICD-10-PCS, International Classification of Diseases, Tenth Revision, Procedure Coding System; ICD-O, International Classification of Disease for Oncology; LOINC, Logical Observation Identifiers Names and Codes; NAACCR, North American Association of Central Cancer Registries; OMOP, Observational Medical Outcomes Partnership; SEER, Surveillance, Epidemiology, and End Results program of the National Cancer Institute.
Results
By relying on the Common Data Model provided by TriNetX, PCN members are able to focus efforts on data exploration and collaboration. One clear advantage is that early-career investigators can design and execute single-center and multicenter data queries and out-of-the-box analytics in support of their research aims with little to no investment (eg, research funding, personnel) (Figure 3). While members of the TriNetX Global Health Research Network can execute queries across the entire network, the community created by the PCN—which shares a common interest in pediatric research to learn from each other’s experiences and recommendations—may be of greatest utility. The PCN provides a consultative forum where junior investigators are able to optimize the design of their queries and analytic approaches as well as obtain guidance in preparing IRB protocols where warranted.
Figure 3.
Representation of the investigator’s path of inquiry to inform the development of a clinical trial and the utility of TriNetX during specific stages of the research development lifecycle.
PCN case study
In an example of a project brought to the PCN steering committee, an early-career investigator with limited prior experience in large healthcare database research sought to investigate the development of venous thromboembolism (VTE) events in pediatric patients with sickle cell disease (SCD)—a low-frequency complication in a rare pediatric disease. The specific aims of the project were to investigate the frequency of recurrent VTE and to identify prognostic factors associated with VTE recurrence among children <21 years of age with a diagnosis of SCD who had suffered a first (ie, index) VTE episode.
A query identifying children with SCD was built using a previously validated algorithm that was modified to exclude patients with International Classification of Diseases 10th revision (ICD-10) diagnosis codes of sickle cell trait or thalassemia without sickle cell disease. All available SCD ICD-10 codes were included in the study population (D57.0, D57.2, D57.4, D57.8). Index and recurrent VTE cases were likewise identified using ICD-10 codes (I82, I26, I67.6). Index VTE was defined as the first diagnosis of VTE in the patient’s EHR. The investigator presented her study design to the PCN consultative group to obtain feedback on query design and analytic considerations. To enhance the specificity of the search, recurrent VTE was defined as an acute VTE diagnosis that occurred 90 days after the index VTE and in a different anatomic location from the index VTE. The TriNetX “Explore Cohort” feature, a browser-based, real-time analytic that provides information on the key clinical characteristics of the cohort, such as demographics and laboratory values, was used to validate that identified patients were indeed children with SCD. Once validated, the investigator obtained the de-identified dataset from the TriNetX Research Network via a request to her local institutional Clinical Research Data Acquisition Core to perform the analysis in collaboration with institution-based biostatisticians.
The study’s results were presented as a selected oral abstract at the 63rd American Society of Hematology Annual Meeting,14 contributed to preliminary data for a National Institutes of Health Mentored Research Award (K23) application, and a manuscript describing the findings is in preparation.
In the first 30 months post-launch of the PCN, over 250 queries by 19 PCN users have been issued, steering committee presentations made by 4 junior investigators, and 3 NIH Patient-Oriented Research Career Development Award grant applications submitted utilizing preliminary data from PCN-consulted projects.
Discussion
The PCN is one of the few pediatric-focused, disease-agnostic collaborative clinical data networks. It leverages the robust data harmonization available through TriNetX’s Global Health Research Network to facilitate data sharing and empower investigator-initiated collaborative research. The rarity of many childhood diseases and the difficulty obtaining anonymized real-world data associated with these patients led the PCN founding members to seek simpler ways to establish inter-organizational collaboration. Additionally, TriNetX offers services to help PCN users build queries, assists in understanding the data PCN researchers are working with, and assigns dedicated account management teams to facilitate site-to-site introductions and subsequent inter-organizational discussions between researchers.
A key advantage of PCN participation is that it does not require additional data exportation from each organization but rather leverages the existing anonymized, harmonized data already shared by participating sites. The goal was to create a collaborative pediatric research community with minimal additional technical burden, with pre-established pan-network data use agreements in place, and to foster multi-institutional pediatric research collaboration. The PCN seeks to complement other well-established pediatric networks such as PEDSnet and ShaRE, to encourage collaboration in an easy-to-use common data environment, especially for users with limited data analysis experience and/or resources.
Conclusion
The PCN was established to be an accessible, robust ecosystem designed to address the unique challenges related to conducting multi-institutional, collaborative pediatric research. Providing real-time access to multi-site, anonymous EHR data for millions of patients, the PCN helps investigators answer imperative logistical research questions by supporting inclusion/exclusion criteria-based queries and providing powerful analytical tools, including the statistical power necessary for rare childhood conditions and low-frequency outcomes of interest. Additionally, the PCN has supported early-career investigators by providing access to consultation and facilitation for multicenter study planning from a seasoned community of pediatric researchers and biostatisticians. The network is open to HCOs that already contribute de-identified EHR-derived data to TriNetX’s broader platform and may join by entering a pan-PCN Data Use Agreement. The PCN seeks to grow its membership over the next several years to maximize impact in generating new knowledge across a broad array of pediatric health conditions and therapeutic areas.
Acknowledgments
The authors would like to thank John Esposito for his helpful insight, discussions, comments, and support.
Contributor Information
Jurran L Wilson, Center for Translational Research, Children’s National Hospital, Washington, DC 20010, United States.
Marisol Betensky, Department of Pediatrics, Division of Hematology, Johns Hopkins University School of Medicine, Baltimore, MD 21287, United States; Johns Hopkins All Children’s Institute for Clinical and Translational Research, St Peterburg, FL 33701, United States.
Sharda Udassi, Department of Pediatrics, WVU Medicine Children’s Hospital, Morgantown, WV 26505, United States.
Pavithra R Ellison, Department of Anesthesiology, WVU Medicine Children’s Hospital, Morgantown, WV 26505, United States.
Richard Lilienthal, TriNetX, LLC, Cambridge, MA 02140, United States.
Lindsay R Stahl, TriNetX, LLC, Cambridge, MA 02140, United States.
Matvey B Palchuk, TriNetX, LLC, Cambridge, MA 02140, United States.
Ayesha Zia, Department of Pediatrics, UT Southwestern Medical Center, Dallas, TX 75390, United States.
Deborah A Town, Research, Children’s HealthSM, Dallas, TX 75235, United States.
Wes Kimble, West Virginia Clinical and Translational Science Institute, Morgantown, WV 26506, United States.
Neil A Goldenberg, Johns Hopkins All Children’s Institute for Clinical and Translational Research, St Peterburg, FL 33701, United States; Departments of Pediatrics and Medicine, Johns Hopkins University School of Medicine, Baltimore, MD 21287, United States.
Hiroki Morizono, Center for Genetic Medicine Research, Children’s National Hospital, Washington, DC 20010, United States.
Author contributions
Jurran L. Wilson wrote the initial draft of the manuscript with Marisol Betensky and Neil A. Goldenberg providing case study information. Richard Lilienthal, Matvey B. Palchuk, and Lindsay R. Stahl contributed to operationalizing the Pediatric Collaboratory within the TriNetX platform. All authors provided critical feedback for establishing operating parameters of the Pediatric Collaboratory. All authors discussed the manuscript and contributed to the editing and final draft of the manuscript. Neil A. Goldenberg and Hiroki Morizono contributed equally to this paper as co-senior authors providing oversight and leadership responsibility for planning and execution.
Funding
This work was supported by National Institutes of Health National Center for Advancing Translational Science awards UL1TR001876 and 1UL1TR003098, National Institutes of Health National Human Genome Research Institute award 1U24HD106537, National Institutes of Health National Heart Lung and Blood Institute awards 1R01HL153963 and 1U01HL130048, National Institutes of Health National Institute of General Medical Services award 2U54GM104942-07, and American Heart Association award 20IPA35320263. The content of this publication is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Conflict of interest
A.Z. has received honoraria from Sanofi and Takeda for her role as an advisory board member for scientific advisory boards in the past 3 years. N.A.G. has received or has recently received consultancy fees from Anthos Therapeutics, Bayer, Boehringer-Ingelheim, Daiichi Sankyo, and the University of Colorado-affiliated Academic Research Organization CPC Clinical Research for roles in clinical trial planning or oversight committees (eg, advisory committee; steering committee; data and safety monitoring committee) in pharmaceutical industry-sponsored pediatric clinical trials of antithrombotics. All other authors have indicated they have no financial relationships or potential conflicts of interest relevant to this article to disclose.
Data availability
This paper describes the creation of a networked Collaboratory environment and no new data were generated or analyzed in support of this work. Access to the TriNetX environment is available from https://trinetx.com.
References
- 1. Institute of Medicine (US) Forum on Drug Discovery, Development and Translation . Transforming clinical research in the United States: challenges and opportunities: workshop summary. In: The National Academies Collection: Reports Funded by National Institutes of Health. Washington, DC: National Academies Press; 2010. [PubMed] [Google Scholar]
- 2. Schilsky RL, McIntyre OR, Holland JF, Frei E. 3rd. A concise history of the cancer and leukemia group B. Clin Cancer Res. 2006;12(11 Pt 2):3553s-3555s. 10.1158/1078-0432.CCR-06-9000 [DOI] [PubMed] [Google Scholar]
- 3. Frei E, Holland JF, Schneiderman MA, et al. A comparative study of two regimens of combination chemotherapy in acute leukemia. Blood. 1958;13(12):1126-1148. [PubMed] [Google Scholar]
- 4. O’Leary M, Krailo M, Anderson JR, Reaman GH.. Progress in childhood cancer: 50 years of research collaboration, a report from the Children’s Oncology Group. Semin Oncol. 2008;35(5):484-493. 10.1053/j.seminoncol.2008.07.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Brunner HI, Rider LG, Kingsbury DJ, PRCSG Advisory Council, et al. Pediatric Rheumatology Collaborative Study Group—over four decades of pivotal clinical drug research in pediatric rheumatology. Pediatr Rheumatol Online J. 2018;16(1):45. 10.1186/s12969-018-0261-x[published Online First: 20180711]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Krischer JP, Gopal-Srivastava R, Groft SC, Eckstein DJ, Rare Diseases Clinical Research Network. The Rare Diseases Clinical Research Network’s organization and approach to observational research and health outcomes research. J Gen Intern Med. 2014;29(Suppl 3):S739-S744. 10.1007/s11606-014-2894-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Bressan S, Klassen TP, Dalziel SR, et al. The Pediatric Emergency Research Network: a decade old and growing. Eur J Emerg Med. 2021;28(5):341-343. 10.1097/MEJ.0000000000000847 [DOI] [PubMed] [Google Scholar]
- 8. Forrest CB, Margolis PA, Bailey LC, et al. PEDSnet: a national pediatric learning health system. J Am Med Inform Assoc. 2014;21(4):602-606. 10.1136/amiajnl-2014-002743[published Online First: 20140512]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Connor EM, Smoyer WE, Davis JM, et al. Meeting the demand for pediatric clinical trials. Sci Transl Med. 2014;6(227):227fs11. 10.1126/scitranslmed.3008043 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Davis S, Ehwerhemuepha L, Feaster W, et al. Standardized Health data and Research Exchange (SHaRE): promoting a learning health system. JAMIA Open. 2022;5(1):ooab120. 10.1093/jamiaopen/ooab120 [published Online First: 20220117]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Slora EJ, Harris DL, Bocian AB, Wasserman RC.. Pediatric clinical research networks: current status, common challenges, and potential solutions. Pediatrics. 2010;126(4):740-745. 10.1542/peds.2009-3586[published Online First: 20100913]. [DOI] [PubMed] [Google Scholar]
- 12. Topaloglu U, Palchuk MB.. Using a federated network of real-world data to optimize clinical trials operations. JCO Clin Cancer Inform. 2018;2:1-10. 10.1200/CCI.17.00067 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Palchuk MB, London JW, Perez-Rey D, et al. A global federated real-world data and analytics platform for research. JAMIA Open. 2023;6(2):ooad035. 10.1093/jamiaopen/ooad035 [published Online First: 20230513]. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Betensky M, Amankwah E, Hankins JS, Goldenberg N.. High rate of recurrent venous thromboembolism is children with sickle cell disease and history of VTE: Analysis of the trinetx research network database. Blood. 2021;138(Supplement 1):778-778. 10.1182/blood-2021-154003 [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
This paper describes the creation of a networked Collaboratory environment and no new data were generated or analyzed in support of this work. Access to the TriNetX environment is available from https://trinetx.com.