Abstract
Despite the increasing burden of dengue, the regional emergence of the virus in Kenya has not been examined. This study investigates the genetic structure and regional spread of dengue virus-2 in Kenya. Viral RNA from acutely ill patients in Kenya was enriched and sequenced. Six new dengue-2 genomes were combined with 349 publicly available genomes and phylogenies used to infer gene flow between Kenya and other countries. Analyses indicate two dengue-2 Cosmopolitan genotype lineages circulating in Kenya, linked to recent outbreaks in coastal Kenya and Burkina Faso. Lineages circulating in Western, Southern, and Eastern Africa exhibiting similar evolutionary features are also reported. Phylogeography suggests importation of dengue-2 into Kenya from East and Southeast Asia and bidirectional geneflow. Additional lineages circulating in Africa are also imported from East and Southeast Asia. These findings underscore how intermittent importations from East and Southeast Asia drive dengue-2 circulation in Kenya and Africa more broadly.
Subject terms: Phylogenetics, Molecular evolution, Dengue virus
There is limited data about local emergence and spread of dengue virus in East African countries. In this study, the authors investigate genetic and geographic epidemiology of dengue virus 2 in East Africa and report co-circulation of multiple distinct lineages introduced from regions in East and Southeast Asia.
Introduction
Dengue virus (DENV) is transmitted by mosquitoes and poses a significant public health risk, particularly in developing countries1,2. DENV is an arbovirus of the genus Flavivirus and family Flaviviridae3–5. The World Health Organization (WHO) estimates that two-fifths of the world’s population is at risk of infection by DENV6, with up to 390 million infections annually, of which only 25% are symptomatic, resulting in up to 25,000 deaths, and nearly 3 million DALYs attributed to DENV2,7. Symptomatic cases present with an acute illness characterized by flu-like symptoms, including fever and headache, called dengue fever (DF)8,9. This is often misdiagnosed as another febrile tropical disease such as malaria10. A small fraction of these cases progresses to severe dengue, characterized by abdominal pain, hemorrhage, and circulatory collapse, which can lead to death11,12.
Dengue virus is a positive-sense, single-stranded RNA virus of approximately 11 kb in length. The evolution and diversity of the virus are enhanced by the existence of four genetically similar but antigenically distinct serotypes, DENV-1, −2, −3, and −4, that are transmitted in both urban and sylvatic cycles13,14. DENV serotypes share approximately 70% of their genome and are further divided into genotypes; within-serotype genetic groupings that show ~6% divergence, and which are often geographically distinct, providing clues to virus emergence and spread15. The virus is thought to have diverged from a similar flavivirus approximately 1000 years ago, while the four serotypes are hypothesized to have independently diverged from sylvatic cycles approximately 125–325 years ago13. The subsequent geographic structure of the genotypes can be partially explained by the founder effect and genetic drift, as well as natural selection within specific geographic populations. The increasing spread and diversity of dengue virus are linked to climate change, the rise of large urban centers, and increasing globalization16,17.
Dengue virus infecting humans in Sub-Saharan Africa is primarily transmitted by Aedes aegypti mosquitoes, which also transmit chikungunya and Zika viruses4,18. The urban ecotype of the Ae. aegypti mosquito is well-adapted to human settlements and urban areas19,20. Adult Ae. aegypti female mosquitoes feed on human blood, which they use in egg production, and lay eggs preferentially in artificial containers20–22. As a result, domestic areas are particularly suitable for mosquito breeding and dengue transmission. Deployment of licensed dengue vaccines (Dengavaxia and Qdenga) is limited and there are safety concerns in populations of varying dengue immune status23–25.
The epidemiology of dengue in East Africa is not well understood26–28. This is partially a result of limitations in surveillance and competing public health priorities. Dengue virus is endemic to Kenya and outbreaks have been observed in the region since 1982, particularly along coastal cities and ports27,29–33. Cases occur throughout the year and peaks are observed following increased rainfall34,35. Urbanization is one of the major drivers of disease risk, and DENV outbreaks have been described in major coastal urban areas of Kenya such as Mombasa and Malindi as well as urban, inland areas such as Kisumu, though less frequently33,36. Despite the increasing burden of dengue in Kenya, the evolutionary characteristics of the virus, as well as the contribution of local and global level circulation patterns to overall disease spread remain largely unexamined. The aim of this study is to assess the underlying genetic structure of circulating DENV-2 and evaluate the extent of gene flow within Kenya, as well as the contribution of global and local circulation patterns to disease spread. We find that dengue virus serotype-2 circulating in Kenya is not monophyletic, consisting of at least two contemporary lineages. We further find that dengue-2 in other regions in Africa exhibits similar genetic diversity consisting of multiple lineages that circulate concurrently, some of which are associated with recent outbreaks. Finally, we show that this genetic diversity is maintained by intermittent introduction of dengue-2 from East and Southeast Asia and conclude that interrupting transmission from these regions might be an effective outbreak prevention strategy.
Results
Dengue virus 2 sequences isolated from Kenya
To investigate circulating dengue virus diversity in Kenya, we conducted prospective genomic sampling and used an amplicon-based approach to sequence PCR-positive samples. Successfully sequenced samples were combined with publicly available DENV-2 sequences for phylogenetic analyses. A total of 355 sequences were included in phylogenetic analyses. 43 of these samples were collected from 8 sites in Kenya, six of which are newly isolated sequences from our study sites of Msambweni (N = 1) and Ukunda (N = 5) (Fig. 1). Cycle threshold values of newly sequenced DENV-2 samples ranged from 18 to 32. The maximum possible coverage obtainable from our samples was ~95%, which was the proportion of the DENV-2 genome enriched by the amplicons. The average coverage from the samples included in the analysis was 90%, and the maximum coverage was 93% (Supplementary Table 1). An additional 4 DENV-1 positive and 1 DENV-3 positive samples were also obtained and are presented in separate work. The 37 DENV-2 sequences collected from Kenya but not as part of this study (from now on referred to as “non-study sites”), and 312 sequences collected from outside Kenya were retrieved from Genbank and included in our dataset. Samples from Thailand (N = 31), Brazil (N = 26), and Singapore (N = 21) contributed the largest number of global sequences to the analytic dataset after subsampling via a cluster-based approach. Global sequences were grouped into 16 geographically defined study regions based on preliminary analyses to reduce computational burden (Supplementary Table 3). The final sequence dataset consisted of 355 sequences from 62 countries globally and 8 distinct city locations within Kenya, collected between 1944 and 2022 (Supplementary Tables 3, 4, Supplementary Data 2).
Fig. 1. Locations of data collection sites.
Participants were recruited from outpatient clinics in dengue-endemic sites in Kenya: Chulaimbo, a rural inland site; Kisumu, an urban inland site; Msambweni, a rural coastal site; and Ukunda, an urban coastal site, as part of two arbovirus surveillance studies in Kenya. Data collection was conducted in all four sites from 2014 to 2018, and additional data collection was conducted in Kisumu and Ukunda from 2019 to 2022.
While the final dataset included sequences from all five known DENV-2 non-sylvatic genotypes, over 50% of included sequences (196 sequences) fell into Genotype II (Cosmopolitan), which also showed the broadest geographic distribution. Genotype V (Asian I) sequences were the most geographically restricted, with all 51 sequences from this genotype geocoded to Central Southeastern Asia or Eastern Asia. Genetically divergent sequences from the sylvatic genotype (Genotype VI) were not included in the final sequence dataset. All sequences from our study sites, as well as all non-study site Kenya sequences, fell into Genotype II (Cosmopolitan).
Maximum likelihood and Bayesian phylogenetic analyses
Maximum likelihood (ML) phylogenetic analysis resolved two distinct DENV-2 lineages circulating in Kenya (Fig. 2), as well as three additional lineages circulating in Western Africa, Southern Africa, and Eastern Africa. The two Kenya lineages (designated Lineages 1 and 2) were both in DENV-2 Genotype II (Cosmopolitan) (Fig. 2). The first of these two lineages (Lineage 1) consisted of non-study site sequences (collected from Wajir in central Kenya and Mombasa in coastal Kenya) that clustered closely with sequences collected from West and Central Africa (Fig. 2, Lineage 1). The second lineage (Lineage 2) consisted of 41 sequences collected from multiple coastal Kenya sites that clustered almost exclusively together with the exception of a single European sequence isolated from a traveler who recently visited Kenya (Fig. 2, Lineage 2)37. The coastal Kenyan clade shares ancestry with sequences from Southern Africa and Eastern Asia. Sequences within the coastal Kenyan clade were intermixed, with no apparent clustering pattern at the level of cities along this coastal region. This includes samples we collected within our study from the urban and rural coastal sites of Ukunda and Msambweni.
Fig. 2. Maximum-likelihood phylogenetic tree of DENV-2 constructed using six study sequences as well as 37 publicly available sequences from other sites in Kenya and 312 sequences from elsewhere, downloaded from Genbank.
a Maximum likelihood phylogenetic trees were constructed using six partial to complete DENV-2 genomes from the study (>80% coverage) as well as 349 partial to complete genomes (>60% coverage) genomes from publicly available sequences on Genbank, using RAxML-NG with the default convergence bootstrap cut-off of 0.03. Clades containing DENV-2 sequences isolated from Kenya (including study sequences) are highlighted and shown as insets b (Lineage 1) and c (Lineage 2). Sequences collected from study sites are starred. Tip colors indicate sequences collected from regions within Africa, as shown in the legend. As the focus of this analysis, Kenya is separated from other regions in Eastern Africa. DENV-2 genotypes (I–V) are shown as colored bars. Bootstrap support values are shown for nodes of interest.
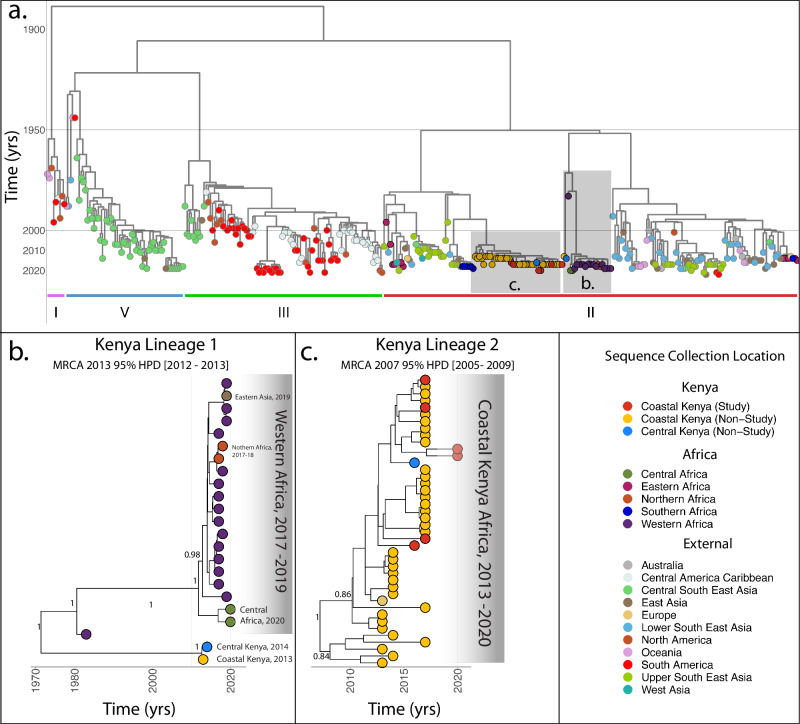
The final dataset yielded a relatively high temporal signal, with a correlation coefficient of 0.91 and R2 of 0.83 (Supplementary Fig. 1). Our Bayesian analyses yielded both an estimate of the evolutionary rate and a maximum clade credibility tree (MCC). The overall posterior mean evolutionary rate for DENV-2 inferred from the relaxed molecular clock across the entire dataset was 7.87 × 10−4 substitutions per site per year [95% HPD 7.30 × 10−4, 8.44 × 10−4]. The MCC tree under the relaxed molecular clock was topologically congruent with the ML tree and confirmed that DENV-2 isolates collected in Kenya are not monophyletic, consisting of at least two separate lineages (Fig. 3). Lineage 2’s date of origin corresponds to subsequent outbreaks along upper coastal and inland Kenya with an MRCA in 2007 [95% HPD 2005, 2009] (Fig. 3, panel c). Lineage 1’s long branch suggests that the lineage may have been circulating as early as the mid-1970s, although it was only sampled in Kenya more recently [MRCA 2013 95% HPD 2012, 2013] (Fig. 3 panel b). As noted in the ML results, the distribution of sequences within the most recent Cosmopolitan clade, Lineage 2, is generally not geographically structured, with sequences isolated from the coastal cities of Ukunda, Mombasa, Msambweni, Malindi, and Mtwapa intermixed (Fig. 3, Supplementary Fig. 3).
Fig. 3. Bayesian timed phylogenetic tree generated from phylogeographic analysis using six study sequences as well as publicly available sequences retrieved from Genbank.
a Timed phylogenetic trees were generated from six partial DENV-2 genomes from this study (>60% coverage) as well as 349 partial genomes (>60% coverage) from publicly available sequences on Genbank. Bayesian phylogeographic analyses were conducted on BEAST1.10.4 to determine tMRCAs as well as reconstruct ancestral sequence locations, with a general time reversible nucleotide substitution model with a Gamma distributed rate and a proportion of invariant sites (GTR + G4 + I) as determined by ModelTest-NG. Tips are colored by location and the legend shows the color corresponding to the specific region of collection of each sequence. Clades containing DENV-2 sequences isolated from Kenya (including study sequences) are highlighted and shown as insets b (Lineage 1), c (Lineage 2) with posterior node probabilities of key nodes shown. DENV-2 genotypes (I–V) are shown as colored bars.
Three additional lineages corresponding to sequences collected from other regions within Africa are also highlighted. The first of these lineages is closely related to Kenya-Lineage 2 and consists of sequences isolated from Western and Central Africa between 2017 and 2020, as well as an additional Western African sequence isolated in 1983. The date of the MRCA of this lineage is 1981 [95% HPD 1977–1983] (Fig. 6). The second of the three non-Kenya lineages consists of sequences isolated from Southern Africa, from 2018 to 2019, with an inferred MRCA in 2015 [95% HPD 2014–2016] (Fig. 6). Finally, the third of the non-Kenya lineages consists of sequences isolated from other locations within Eastern Africa from 2014 to 2015 with a MRCA date in 2012 [95% HPD 2011–2013] (Fig. 6). A final set of sequences were collected from regions in Eastern Africa between 1996 and 2018 (Supplementary Fig. 2).
Fig. 6. Bayesian timed phylogenetic tree showing DENV-2 lineages causing major recent outbreaks in West Africa (Burkina Faso, Senegal—2016−2019), Eastern Africa, and Southern Africa (Angola—2017−2019).
a Timed phylogenetic trees generated in BEAST1.10.4 from 355 DENV-2 genomes showing circulating genetic diversity of DENV-2 in Eastern (b), West (c), and Southern Africa (d).
Dengue virus 2 phylogeography
Phylogeographic analysis suggests that Kenya Lineage 2 was introduced into Kenya from East Asia (Fig. 5), while Kenya Lineage 1 was introduced into Kenya from Western Africa (Fig. 4). Both lineages are inferred to have been originally circulating in Oceania and Lower Southeast Asia with initial dispersals from these regions in the late 19th to mid-20th centuries (Figs. 4 and 5). Both circulating lineages are of the Cosmopolitan genotype. Geographic associations and their relative strength based on phylogeographic patterns are summarized on a regional scale in Supplementary Fig. 4. Geographic analyses based on phylogeographic patterns within Kenya suggest virus exportation from an initial introduction into Mombasa, along the coast to the cities of Malindi, Mtwapa, Msambweni, and Ukunda, as well as Lamu and Voi in central Kenya (Supplementary Fig. 5). Limited sequencing data collected from less densely sampled locations such as the central Kenya sites of Lamu, Wajir, and Voi, prevent more accurate phylogeographic inference specific to these sites. The additional non-Kenya lineages are inferred to share the same distal geographic origin of Lower Southeast Asia and Oceania, with the Western Africa lineage introduced directly into the region from Lower Southeast Asia, while the Eastern Africa and Southern Africa lineages were introduced from Eastern Asia (Fig. 7).
Fig. 5. Global geographic history of DENV-2 lineage 2 detected in Kenya.

Bayesian maximum clade credibility trees from phylogeographic analyses of 355 sequences were used to reconstruct the geographic history of DENV-2 lineage 1 circulating in Kenya detected in this study. Geolocation data from sequences was grouped into one of 16 study categories (Supplementary Table 3) to improve analytic tractability. a Each arrow in the map corresponds to a node on the MCC tree ancestral to the detected Kenyan lineage indicated in the plot, where a geographic transition between two locations occurred. Lines are annotated with inferred MRCA node date and 95% high posterior density (HPD) intervals. b Markov jump history between locations was recorded for inferred trees and is summarized in the bottom plot. The plot shows the inferred timing of Markov jumps representing transmission between two locations for the geographic history shown in the top plot. Vertical lines represent a geographic transition between two locations, while horizontal lines indicate times during which the ‘lineage’ remains in the same location, i.e. the reward times. Greater density of lines indicates greater certainty of inferred transmission events around a given date, while sparsity of lines indicates lower certainty.
Fig. 4. Global geographic history of DENV-2 lineage 1 detected in Kenya.

Bayesian maximum clade credibility trees from phylogeographic analyses of 355 sequences were used to reconstruct the geographic history of DENV-2 lineage 1 circulating in Kenya detected in this study. Geolocation data from sequences was grouped into one of 16 study categories (Supplementary Table 3) to improve analytic tractability. a Each arrow in the map corresponds to a node on the MCC tree ancestral to the detected Kenyan lineage indicated in the plot, where a geographic transition between two locations occurred. Lines are annotated with inferred MRCA node date and 95% high posterior density (HPD) intervals. b Markov jump history between locations was recorded for inferred trees and is summarized in the bottom plot. The plot shows the inferred timing of Markov jumps representing transmission between two locations for the geographic history shown in the top plot. Vertical lines represent a geographic transition between two locations, while horizontal lines indicate times during which the ‘lineage’ remains in the same location, i.e. the reward times. Greater density of lines indicates greater certainty of inferred transmission events around a given date, while sparsity of lines indicates lower certainty.
Fig. 7. Geographic history of DENV-2 lineages causing major recent outbreaks in West Africa (Burkina Faso, Senegal—2016−2019), Eastern Africa and Southern Africa (Angola—2017−2019).
Bayesian maximum clade credibility trees from phylogeographic analyses of 355 DENV-2 sequences were used to reconstruct the geographic history of the additional lineages causing recent outbreaks in a. Eastern, b Western and c Southern Africa. Each arrow in the map corresponds to a node on the MCC tree ancestral to the detected outbreak-causing lineage indicated in the plot, where a geographic transition between two locations occurred. Lines are annotated with inferred MRCA node date and 95% High Posterior Density (HPD) intervals. (i) indicates the transition from Western to Northern Africa occurring in 2016 95% HPD [2015−2017], and (ii) indicates the transition from Western to Central Africa occurring in 2012 [2010–2014]. Markov jump history between locations was recorded for inferred trees and is summarized in the plots on the right. The plot shows the inferred timing of Markov jumps representing transmission between two locations for the geographic history shown in left plot. Vertical lines represent a geographic transition between two locations, while horizontal lines indicate times during which the ‘lineage’ remains in the same location. Greater density of lines indicates greater certainty of inferred transmission events around a given date, while sparsity of lines indicates lower certainty.
We inferred rates of global gene flow with a discrete trait phylogeographic analysis and found a median transmission rate of 0.68 transmission events/year [Range 0.40, 3.00]. The maximum transmission rates were observed from East Asia to Upper Southeast Asia [transmission rate 3.00, 95% HPD 0.95, 5.52] and Lower Southeast Asia to Eastern Asia [transmission rate 2.32, 95% HPD 0.90, 4.16]. The maximum inferred transmission rate into a Kenya site was 0.73 transmission events/year [95% HPD 0, 3.00] from Eastern Asia, and the maximum transmission rate into a non-Kenya African region was 1.48 transmission events/year [95% HPD 0.39–3.10] from East Asia into East Africa (Supplementary Fig. 4).
Discussion
Over the last few decades, the importance of arboviruses as a public health priority has increased, as shown by several recent large outbreaks of arboviruses, including Zika, dengue, and chikungunya viruses38,39. In this study, we sequenced viral isolates from clinical study sites in rural and urban Kenya. Combined with previously published sequences from Kenya and elsewhere, we determined the underlying evolutionary characteristics driving DENV-2 circulation in the region as well as the regional and local dynamics of spread. Our analysis indicates the existence of multiple independent lineages of DENV-2 within East Africa, including two Genotype II (Cosmopolitan) lineages (Figs. 2 and 3, Lineages 1 and 2) circulating in Kenya, as well as three additional lineages of DENV-2 circulating in Western, Southern, and Eastern Africa. Of the two Kenyan Cosmopolitan genotype lineages, the more recent one (Lineage 2) was introduced in the early 2000s, while the earlier one (Lineage 1) may have been introduced as early as the 1970s (Fig. 3). Results suggest that this older Cosmopolitan lineage (Lineage 1), which was detected in Mombasa (Coastal Kenya) and Wajir (Central Kenya), has been maintained in the population at low levels, possibly circulating undetected in Central Kenya, and is not tied to any recent outbreaks within Kenya. The more recent lineage (Lineage 2) is associated with outbreaks and dengue circulation that have been observed in coastal Kenya32,33,40. Finally, we find that DENV-2 circulation dynamics in Kenya and Africa more broadly are strongly linked to circulation in East and Southeast Asia, with multiple independent introductions occurring from Eastern Asia and Southeastern Asia at different points over the last ten to thirty years (Figs. 6 and 7).
Our analysis detected two related cocirculating lineages of DENV-2 in Kenya. Lineages 1 and 2 (CII-lineage and CI-lineage from Langat et al, 2020) fall into the Cosmopolitan genotype, which has been shown to be uncharacteristically geographically dispersed, unlike other genotypes that tend to exhibit stronger geographic structure. The observed DENV-2 lineages circulating along coastal Kenya show little geographic segregation at the city level, suggesting constant bidirectional gene flow through the urban and rural locations along the coast such as Msambweni, Ukunda, Malindi, and Mombasa (Fig. 3, Supplementary Fig. 3). Comparisons between data collected in this study and existing data indicate that recent DENV-2 outbreaks in Malindi in 2017 and Mtwapa in 2014 are likely driven by relatively recent introductions and circulation of Lineage 2 (Malindi and Mtwapa: Fig. 5, Supplementary Fig. 5), while Lineage 1 is not associated with any known, recent outbreaks in Kenya, though it is associated with a 2017 Western Africa outbreak centered in Burkina Faso41. A multi-country dengue outbreak centered around Burkina Faso and Central Africa was reported in 2023, though no sequence data from this outbreak are publicly available yet to characterize the causative lineages, particularly whether or not this outbreak is related to Lineage 142. Limited reporting and surveillance of DENV-2 earlier on in its circulation, associated with Lineage 1, may have resulted in missed DENV-2 cases resulting in more sparse characterization of local circulation in phylogeographic analyses. Additional work is required to determine whether the continuing circulation of the older lineage at lower levels is due to lower outbreak potential, with more recent outbreak-causing lineages being more transmissible or otherwise more likely to cause outbreaks.
DENV-2 dynamics in the region are strongly tied to DENV-2 dynamics in East and Southeast Asia. Phylogeographic analyses with the current sequencing dataset suggest that Lineage 1, the oldest circulating lineage detected in this analysis, may have been introduced from Singapore (coded as Lower Southeast Asia) via West Africa in the 1970s, while Lineage 2 may have been introduced from China (coded as East Asia) in the last twenty to thirty years (Fig. 4). The more recently introduced lineages may be tied to increased travel, trade, and globalization. Phylogeographic analyses suggest that both lineages were introduced to Mombasa (Supplementary Fig. 3). Though other, unsampled locations may have also played a role in the introduction of both lineages, large urban centers such as Mombasa are often implicated in DENV importations at regional levels. Phylogeographic analyses also indicate that these large centers within Kenya, such as Mombasa may export the disease back into Asia (Supplementary Fig. 3). These results reinforce the role of large urban centers as potential seeding points of DENV-2 spread in the region. Despite understanding the role of these large urban centers, specific mechanisms of transmission between larger urban areas and their broader contribution to disease dynamics remain unexamined. Possible transmission routes include transmission via infected humans importing the disease to naïve areas and expanded vector ranges enabling local transmission in naïve areas following importation. Transovarial transmission (i.e. infected eggs) in vectors might further enable over-wintering and contribute to the establishment of endemic disease cycles. Transmission via infected humans has been observed, most notably with importations into colder areas less suitable for the dengue vector, such as European locations37,43.
While all four DENV serotypes have been observed in Kenya to date, DENV-2 has been shown to be the most frequent33. Previous analyses have detected DENV at several locations on the coast of Kenya, while very few studies have evaluated DENV in inland and western locations30,31,40. The burden of dengue in these inland or western locations ascertained from the handful of studies available may be severely underestimated32,33. Our data suggest that DENV circulating in inland locations may be maintained by introductions from the more connected coastal cities, but possibly at lower levels than the observed introductions into larger, coastal cities. The lower levels of circulation in inland and western regions may also be due to the vector distributions and ranges within Kenya. Coastal Kenya is generally more suitable for the Aedes vector due to higher rainfall and temperature during a larger portion of the year. Reduced DENV-2 circulation in inland Kenya may then be a result of shorter intervals of favorable temperature for vector breeding, or reduced availability of water holding containers, particularly those in non-human settings that would be filled by rainfall, resulting lower vector populations during most parts of the year20,21,44. In order to determine if our current results, which suggest lower circulation of DENV-2 into inland locations, are due to actual lower prevalence of DENV-2 in these locations rather than sampling, more work focusing on these locations is needed.
The additional non-Kenya lineages shown (Fig. 6) highlight dengue circulation associated with other African regions. The Western Africa lineage (Fig. 6) is associated with a 2017 outbreak in Western Africa reported in Letizia et al., which resulted in additional downstream spread of dengue into regions of North and Central Africa41,45 (Fig. 5). The Southern Africa lineage (Fig. 6) is associated with a 2017 outbreak in Angola reported in Neto et al. Finally, the additional Eastern Africa lineage is associated with dengue circulation and outbreaks in other Eastern African countries such as La Reunion and Tanzania46,47. The geographic and evolutionary features of these outbreaks are similar to those of the outbreaks reported in Kenya, with all three lineages tied to Lower Southeast Asia and East Asia, via introductions in the early to mid-2000s. The Western Africa lineage is a notable exception, with an inferred MRCA in the 1970s and possible introduction as early as the mid-1900s. Additional sporadic detection of DENV-2 as well as DENV-1 and DENV-3 in the region, including the continued circulation of dengue in La Reunion46, a recent outbreak in Tanzania47, and a recent outbreak in Chad and Sudan (https://iris.who.int/handle/10665/373147), suggests that dengue may be circulating in several other areas, but undetected48. This suggests a high risk of additional dengue outbreaks as well as risk of establishment of the virus in endemic, and potentially hyperendemic cycles in several countries in the region.
A major limitation of our study is sequence data availability. Despite the importance of DENV circulation, sequence data on DENV in Africa are sparse. Previous DENV sequence surveillance has focused on the envelope gene to evaluate molecular epidemiology and transmission pathways. In our study, we used as much sequence material as we were able to retrieve (>60% of the genomes), optimizing the amount of information at the potential cost of lower confidence at finer geographic scales, such as within specific sites. In addition, timed phylogenies benefit from a broad temporal as well as spatial scale of data collection. Due to the endemic and epidemic patterns of dengue virus circulation in the region, nucleotide data were only available for select years and locations, and data were missing for certain sites in several years. In our study, sequences collected from adjacent rural and urban locations along coastal Kenya clustered together, suggesting frequent gene flow between the locations, and so with a few sequences from either location, we can adequately identify both locations’ contributions to regional spread, reducing the impact of missing data at this level. To specifically disentangle patterns of gene flow between adjacent urban and rural locations, increased data collection at this local scale is required. While the impact of missing spatial data can be mitigated by data availability from other, nearby locations, missing temporal data can be more consequential: our study included no sequence data from Kenya from before 2013, though outbreaks had been reported and current analyses indicate that DENV was circulating in the population with introductions in the mid-2000s and the mid-1970s. As a result, we can make no conclusions about the circulation and spread of earlier lineages and how these may have contributed to observed, contemporary patterns of disease spread. Active surveillance that includes sequencing of circulating DENV, particularly in western and inland Kenya, would help characterize a more complete circulating diversity of the virus, including previously undetected extant lineages, which would be key to resolving earlier circulation patterns of the virus.
In this study, we use molecular data collected from patients to evaluate the evolutionary characteristics and spread of DENV-2 in Kenya. While our results generally match previously reported work, our analyses additionally identify the concurrent circulation of two distinct, contemporary DENV-2 lineages. This suggests that in addition to the previously reported findings indicating the importance of large urban centers in DENV-2 importations, large urban locations may act as foci for local and non-overlapping networks of spread, for example, in the coastal and inland/western regions of Kenya. Given the different ecological and demographic contexts, these separate networks of spread, regional to local, and local-coast to local-coast or local-west to local-west, may have different drivers of disease transmission. Reducing regional and local expansion of DENV-2 may then need to focus on interventions targeted at the specific network of spread and corresponding drivers of transmission within these networks. As DENV continues to expand its global range, understanding regional and local drivers of viral gene flow and disease spread will become vital to designing effective interventions and disease control.
Methods
Clinical surveillance
Data were collected as part of two arbovirus surveillance studies (The burden of Chikungunya and Dengue transmission, infection and disease in Kenya and Disentangling the human vector relationship to disrupt dengue and chikungunya virus outbreaks in Kenya, Technical University of Mombasa, NIH Project Number: R01AI102918), from outpatient clinics in dengue-endemic sites in Kenya: Chulaimbo, a rural inland site; Kisumu, an urban inland site; Msambweni, a rural coastal site; and Ukunda, an urban coastal site, as shown in Fig. 1. Child (1–17 years) and adult ( > 17 years) participants who presented with an acute febrile illness (reported during the previous 2 weeks and including temperature 38 °C at presentation) were recruited into the study between January 2014 and February 2022. Written informed consent was obtained at recruitment from the participant or relevant guardian if the participant was under the age of 18 years. A comprehensive clinical examination was conducted, including collection of demographic and clinical history data via survey forms, and blood draw for serological and molecular testing. Demographic data were stored in RedCap (Version 12.2.11) hosted at Stanford University, while human samples were frozen, shipped to Stanford University, and stored at −70 °C until use49. Ethical approval and oversight for data collection for this study were obtained from the Institutional Review Board of Stanford University (IRB eProtocol # 31488), as well as the Kenya Medical Research Institutes (KEMRI SSC 2611) and Technical University of Mombasa Ethical Review Committee (TUM/ERC EXT/004/2019).
Molecular testing
Nucleic acid material was extracted from blood using RNA extraction kits, including the Qiagen RNAEasy kit or the Mag-bind Viral RNA/DNA kit based on availability, and converted into complementary DNA (cDNA) using a High-Capacity cDNA Reverse Transcription Kit according to manufacturer instructions. Extracted samples were tested for viral presence using a SuperScript III One-Step qPCR assay with a published primer and probe set capable of detecting all four dengue virus serotypes50. The RT-qPCR reaction was run in a 25 µl final reaction volume consisting of 5 µl of sample, 12.5 µl of pre-mix qPCR buffer, 2 µl of the primer/probe mix (final concentration of 300 nM), 0.5 µl of polymerase mix and 5 µl of water per reaction. The reaction was run on a BIORAD CFX384 qPCR machine with the following cycles: 52 °C for 15 min, 94 °C for 1 min, and 45 cycles of 94 °C for 15 s, 55 °C for 40 s, 68 °C for 20 s and the dengue specific probe was detected on the FAM channel. The assay included a set of primers and probes to target RNAseP, which was used as an internal control. Dengue-positive samples were serotyped via an additional RT-qPCR reaction with a set of published serotype-specific primers and probes51. The qPCR reaction was run in a 25 µl final reaction volume consisting of 5 µl of sample, 12.5 µl of pre-mix qPCR buffer, 2 µl of the primer/probe mix, 0.5 µl of polymerase mix and 5 µl of water per reaction. The reaction was run on a BIORAD CFX384 qPCR with the following cycles: 50 °C for 30 min, 95 °C for 2 min, and 45 cycles of 95 °C for 15 s and 60 °C for 1 min. Dengue serotypes 1–4 were detected on the FAM, HEX, Texas Red, and Cy5 channels, respectively.
Dengue virus 2 viral amplification
While three dengue serotypes were detected in this study, DENV-2 was most prevalent and is the focus of this analysis. DENV-2 positive samples identified in the serotype-specific qPCR reaction were run through a viral amplicon enrichment protocol using custom primers. Custom DENV-2 enrichment primers were designed to capture the circulating diversity of DENV-2 using PrimalScheme, a tool that designs a set of primers allowing enrichment of viral genomes in overlapping amplicons spanning the genome in two reaction tubes52. A detailed description of the primer design process is included in the supplementary materials (Supplementary Methods). Briefly, primers were designed to enrich the genome in 400 base pair (bp) amplicons with an average overlap of 75 bp between amplicons (Supplementary Data 1). The primers were designed based on a set of 24 genomes retrieved from GenBank (https://www.ncbi.nlm.nih.gov/genbank/) corresponding to the regional DENV-2 circulating diversity previously identified33. The primers were divided into two pools such that overlapping amplicons were in separate reactions and diluted to a final reaction concentration of 15 nM. Samples were enriched in one of two pools using the NEB Inc. Q5 High Fidelity Master mix, in 25 µl final reaction volume consisting of 12.5 µl of master mix, 5 µl of water, 1.5 µl of primer mix, and 6 µl of sample template. The reaction conditions were 98 °C for 30 s, 37 cycles of 95 °C for 15 s, and 65 °C for 5 min as suggested by the PrimalScheme protocol. Each reaction tube was run on a gel afterward, and the presence of a 400 bp band was used as indicative of a successful reaction. The final set of samples included in the sequencing pipeline was a convenience sample of isolates passing all quality control steps up to this point.
Next-generation sequencing and bioinformatic analyses
Samples were prepared for sequencing based on the output of quality control steps. Purified, enriched cDNA was cleaned using AmpureX beads and quantified using a Qubit4 assay. Library preparation was conducted using the SparQ library preparation kit and TruSeq adapters according to manufacturer instructions with some modifications. TruSeq adapters were diluted based on input sample concentrations. After library preparation, sample concentration was re-quantified using a Qubit4 assay, and fragment size distribution and concentration were quantified using Agilent Bioanalyzer. Concentration was further quantified using the KAPA Hyper prep kit. Prepared samples were pooled in equimolar amounts and sequenced on Illumina’s MiSeq platform using a 2 × 250 bp V2 kit and paired-end dual-indexed sequencing at Stanford University. Water blanks and No Template Controls (NTCs) were included in all reactions from RNA extraction to sequencing and were analyzed through the same bioinformatics workflows to identify possible contamination. Analyses indicated minimal contamination.
Sequencing data was processed using custom scripts based on the iVar bioinformatics pipeline52. Raw reads were downloaded from Illumina’s Basespace platform and aligned to a reference downloaded from GenBank corresponding to a 2015 DENV-2 sample (Accession ID: MT076937.1) using BWA MEM53. Adapters and enrichment primer sequences were trimmed from aligned reads using the iVar trim command, and low-quality reads were trimmed using a PHRED score cutoff of 20. Reads whose length was <10 bases after trimming and quality filtering were discarded. Trimmed reads were sorted using samtools and consensus genomes were generated using samtools-mpileup and ivar-consensus commands54. The consensus base at a position was called for a minimum depth of 10 reads and at a frequency of at least 80%. samtools was used to retrieve coverage and depth statistics and Fastqc was used to calculate and visualize read quality data54,55. Consensus genomes for the six final sets of genomes sequenced as part of this study were exported for phylogenetic analysis and also uploaded to GenBank.
Phylogenetic and phylogeographic analyses
Consensus genomes generated from this study were combined with publicly available complete or partially complete (>60% of the complete genome) DENV-2 genomes downloaded from Genbank. Additional genomes were retrieved from Genbank from other regions known to be connected to Kenya from previous work as well as other similar sequences identified by subjecting sample sequences to NCBI’s BLASTN tool. A hierarchical clustering approach was used on the preliminary dataset of non-Kenya sequences to define clusters of high similarity based on distance-based similarity measures, and randomly sample a fraction of the sequences from within clusters in order to trim the sequence dataset. To do this, geographic location information was used to create location-specific datasets that were separately aligned and examined for similarity. Similar clusters in each location were down-sampled. This served to reduce the size of the dataset while maintaining information about the geographic spread contained in sequences. Location and date of collection corresponding to each genome were also retrieved from Genbank and stored in a separate dataset. In order to explore DENV-2 spread between global locations and locations within Kenya, location information was coded at the city level for locations within Kenya and at regional and sub-regional levels for locations outside Kenya (Supplementary Tables 3 and 4). Sequences in the final database were genotyped using Genome Detective’s Dengue Typing tool (https://www.genomedetective.com/app/typingtool/dengue/), and TempEst (version 1.5.3) was used to evaluate the suitability of our data for timed-phylogeny analyses with root-to-tip regressions (Supplementary Fig. 1)56,57. Samples with divergence much greater than expected, as indicated by the TempEst residuals, were excluded from the analysis. RDP4 was used for recombination detection and 13 sequences with evidence of recombination were excluded from the analysis (Supplementary Methods)58. The final dataset consisted of 355 genotyped samples, annotated with dates and locations (Supplementary Data 2).
Genomes were aligned using MAFFT (version 7.310), and the alignment was examined and manually curated using AliView, including the exclusion of 3’ and 5’ untranslated regions. Initial phylogenetic trees were estimated using a maximum likelihood approach with RAxML-NG on the Stanford Computational Genomics Cluster59. Modeltest-NG (version 0.1.7) was used to determine the best nucleotide substitution model on the maximum likelihood tree60,61. Bayesian tree estimation was conducted in BEAST 1.10.4 using a relaxed molecular clock and the General Time Reversible nucleotide substitution model with a Gamma distributed rate and invariant sites (GTR + G4 + I), as determined by ModelTest-NG (Supplementary Table 2). Three independent, 150 million-step Markov Chain Monte Carlo (MCMC) procedures were run, with sampling every 15,000 generations and 10% burn-in. Output convergence and effective sample sizes (>200 ESS) were evaluated in Tracer, and independent runs were down-sampled (to sampling once every 30,000 generations) and combined in LogCombiner. Final tree data was generated in TreeAnnotator. Maximum clade credibility (MCC) phylogenetic trees were visualized and modified using FigTree and the R ggtree package, and resulting maximum likelihood trees were rooted based on the tempest-derived outgroup. Evolutionary characteristics including evolution rates and times to most recent common ancestor (tMRCA) were calculated based on the MCC tree with a focus on circulating Kenyan lineages (i.e., groups of sequences within the genotypes descended from a single ancestral introduction into Kenya, and exhibiting continued general circulation in Kenya). Uncertainty was expressed using 95% high posterior density (HPD) intervals.
Estimation of the phylogeography of DENV-2 was performed using BEAST1.10 and jointly estimated along with the tree topology using Bayesian Stochastic Search Variable Selection (BSSVS) procedure to evaluate support, with location included as a discrete trait and an asymmetric substitution model to account for different rates of inward versus outward gene flow, and ancestral state reconstruction. The final analysis included 24 discrete locations, including 8 cities within Kenya and the remaining 16 global regions, and sequences collected from 1944 to 2022 (Supplementary Table 3, 4). SpreadD3 was used to assign Bayes Factors for the strength of evidence, and geographic spread rates were visualized using the ggplot and sf packages in R62–64. Maximum transmission rates (transmission events per year between any two locations) and 95% HPD intervals are reported between global sites and local sites separately. A record of the Markov jumps across the history of the phylogeny was also stored, and jumps corresponding to the geographic and evolutionary history of lineages circulating within Kenya and other African regions are shown. Auxiliary data analysis and final data visualization was conducted in R (Version 4.1.2), and all.xml files generated as part of these analyses are available online (https://github.com/SindisoNyathi/doctoralrepo_dengue_phylo/tree/main/beast_xmls).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Description of Additional Supplementary Files
Acknowledgements
We would like to thank all the study participants who contributed sample material to this study from the communities in Kisumu, Ukunda, Msambweni and Chulaimbo, Kenya. We would also like to thank M.K. Sahoo and Dr. R. Verma for their input on refining the laboratory methods. S.N. was funded by the Abbott Laboratories Graduate Student Fellowship and the Centre for Computational, Evolutionary and Human Genetics Fellowship at Stanford University. A.D.L. was funded for this study by an NIH grant (NIH Project Number: R01AI102918).
Author contributions
S.N. conducted the laboratory work, bioinformatics, data analysis, and writing. I.R., K.W., and P.T. contributed to and supervised laboratory work, and bioinformatic analysis and provided input on writing. F.M., B.N., D.V., A.D.L supervised recruitment, sample collection, and follow-up of study participants. J.M., P.A., P.S. Musunzaji., P.C., P.W.M., P.S. Maina., C.N., S.M., Z.J., and D.V. recruited study participants and collected samples and demographic data from study participants. E.M., S.B., J.A., and A.D.L. supervised the research project and contributed to the study design, data analysis, writing, and review of the manuscript.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Data availability
The sequences generated as part of this study have been deposited to Genbank with accession numbers: PP998553, PP998554, PP998555, PP998556, PP998557, PP998558 (https://www.ncbi.nlm.nih.gov/genbank/). All other sequences used in this analysis are available in the supplementary materials (Supplementary Data 2).
Code availability
All.xml files generated as part of this study specifying the parameters of the Beast runs for phylogenetic and phylogeographic analyses are available online and can be downloaded at (https://github.com/SindisoNyathi/doctoralrepo_dengue_phylo/tree/main/beast_xmls).
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors jointly supervised this work: Jason R. Andrews, A. Desiree LaBeaud.
Supplementary information
The online version contains supplementary material available at 10.1038/s41467-024-51018-0.
References
- 1.Benelli, G. & Mehlhorn, H. Declining malaria, rising of dengue and Zika virus: insights for mosquito vector control. Parasitol. Res.115, 1747–1754 (2016). 10.1007/s00436-016-4971-z [DOI] [PubMed] [Google Scholar]
- 2.Bhatt, S. et al. The global distribution and burden of dengue. Nature496, 504–507 (2013). 10.1038/nature12060 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Paixão, E. S., Teixeira, M. G. & Rodrigues, L. C. Zika, chikungunya and dengue: the causes and threats of new and reemerging arboviral diseases. BMJ Glob. Health3, e000530 (2018). 10.1136/bmjgh-2017-000530 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Patterson, J., Sammon, M. & Garg, M. Dengue, zika and chikungunya: emerging arboviruses in the new world. West. J. Emerg. Med.17, 671–679 (2016). 10.5811/westjem.2016.9.30904 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Sukhralia, S. et al. From dengue to Zika: the wide spread of mosquito-borne arboviruses. Eur. J. Clin. Microbiol. Infect. Dis.38, 3–14 (2019). 10.1007/s10096-018-3375-7 [DOI] [PubMed] [Google Scholar]
- 6.World Health Organisation. Dengue and Severe Denguehttps://www.who.int/news-room/fact-sheets/detail/dengue-and-severe-dengue (2022).
- 7.Guo, C. et al. Global epidemiology of dengue outbreaks in 1990–2015: a systematic review and meta-analysis. Front. Cell. Infect. Microbiol.7, 1–11 (2017). 10.3389/fcimb.2017.00317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Quirine, A. et al. Contributions from the silent majority dominate dengue virus transmission. PLOS Pathogens14, 82–86 (2018). [DOI] [PMC free article] [PubMed]
- 9.Salles, T. S. et al. History, epidemiology and diagnostics of dengue in the American and Brazilian contexts: a review. Parasit. Vectors11, 264 (2018). 10.1186/s13071-018-2830-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Amoako, N. et al. Detection of dengue virus among children with suspected malaria, Accra, Ghana. Emerg. Infect. Dis.24, 1561–1564 (2018). 10.3201/eid2408.180341 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Boyles, S. M. et al. Under-the-radar dengue virus infections in natural populations of Aedes aegypti mosquitoes. mSphere5, e00316-20 (2020). 10.1128/mSphere.00316-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pawitan, J. A. Dengue virus infection: predictors for severe dengue. Acta Med. Indones.43, 7 (2011). [PubMed] [Google Scholar]
- 13.Holmes, E. C. & Twiddy, S. S. The origin, emergence and evolutionary genetics of dengue virus. Infect. Genet. Evol.3, 19–28 (2003). 10.1016/S1567-1348(03)00004-2 [DOI] [PubMed] [Google Scholar]
- 14.Rico-Hesse, R. Microevolution and virulence of dengue viruses. Adv. Virus Res.59, 315–341 (2003). 10.1016/S0065-3527(03)59009-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Costa, R. L., Voloch, C. M. & Schrago, C. G. Comparative evolutionary epidemiology of dengue virus serotypes. Infect. Genet. Evol.12, 309–314 (2012). 10.1016/j.meegid.2011.12.011 [DOI] [PubMed] [Google Scholar]
- 16.Lee, S. A. et al. Effect of climate change, connectivity, and socioeconomic factors on the expansion of the dengue virus transmission zone in 21st century Brazil: an ecological modelling study. Lancet Planet. Health5, S14 (2021). 10.1016/S2542-5196(21)00098-X [DOI] [Google Scholar]
- 17.Messina, J. P. et al. Global spread of dengue virus types: mapping the 70 year history. Trends Microbiol.22, 138–146 (2014). 10.1016/j.tim.2013.12.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kraemer, M. U. G. et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. Albopictuse. eLife4, 1–18 (2015). 10.7554/eLife.08347 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hammond, S. N. et al. Characterization of Aedes aegypti (Diptera: Culcidae) production sites in urban Nicaragua. J. Med. Entomol.44, 851–60 (2007). 10.1093/jmedent/44.5.851 [DOI] [PubMed] [Google Scholar]
- 20.Ngugi, H. N. et al. Risk factors for Aedes aegypti household pupal persistence in longitudinal entomological household surveys in urban and rural Kenya. Parasit. Vectors13, 499 (2020). 10.1186/s13071-020-04378-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ndenga, B. A. et al. Characteristics of Aedes aegypti adult mosquitoes in rural and urban areas of western and coastal Kenya. PLoS ONE12, 1–14 (2017). 10.1371/journal.pone.0189971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Stewart Ibarra, A. M. et al. Dengue vector dynamics (Aedes aegypti) influenced by climate and social factors in Ecuador: implications for targeted control. PLoS ONE8, e78263, (2013). 10.1371/journal.pone.0078263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wilder-Smith, A. Dengue vaccine development: status and future. Bundesgesundheitsbl63, 40–44 (2020). [DOI] [PMC free article] [PubMed]
- 24.Scott, L. J. Tetravalent dengue vaccine: a review in the prevention of dengue disease. Drugs76, 1301–1312 (2016). 10.1007/s40265-016-0626-8 [DOI] [PubMed] [Google Scholar]
- 25.Thomas, S. J. & Rothman, A. L. Trials and tribulations on the path to developing a dengue vaccine. Vaccine33, D24–D31 (2015). 10.1016/j.vaccine.2015.05.095 [DOI] [PubMed] [Google Scholar]
- 26.Baba, M., Villinger, J. & Masiga, D. Repetitive dengue outbreaks in East Africa: a proposed phased mitigation approach may reduce its impact. Rev. Med. Virol.19, 57–64 (2009). [DOI] [PubMed] [Google Scholar]
- 27.Pollett, S. et al. The evolution of dengue-2 viruses in Malindi, Kenya and greater East Africa: epidemiological and immunological implications. Infect. Genet. Evol.90, 104617–104617 (2021). 10.1016/j.meegid.2020.104617 [DOI] [PubMed] [Google Scholar]
- 28.Simo, F. B. N. et al. Dengue virus infection in people residing in Africa: a systematic review and meta-analysis of prevalence studies. Sci. Rep.9, 13626 (2019). 10.1038/s41598-019-50135-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ellis, E. M. et al. A household serosurvey to estimate the magnitude of a dengue outbreak in Mombasa, Kenya, 2013. PLoS Negl. Trop. Dis.9, 1–10 (2015). 10.1371/journal.pntd.0003733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Johnson, B. K., Musoke, S., Ocheng, D., Gichogo, A. & Rees, P. H. Dengue-2 virus in Kenya. Lancet320, 208–209 (1982). [DOI] [PubMed]
- 31.Kamau, E. et al. Complete genome sequences of dengue virus type 2 strains from Kilifi, Kenya. Microbiol Resour Announc.8, 24–26 (2019). [DOI] [PMC free article] [PubMed]
- 32.Vu, D. M. et al. Unrecognized dengue virus infections in children, Western Kenya, 2014–2015. Emerg. Infect. Dis.23, 1915–1917 (2017). 10.3201/eid2311.170807 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shah, M. M. et al. High dengue burden and circulation of 4 virus serotypes among children with undifferentiated fever, Kenya, 2014–2017. Emerg. Infect. Dis.26, 2638–2650 (2020). 10.3201/eid2611.200960 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lutomiah, J. et al. Dengue outbreak in Mombasa City, Kenya, 2013–2014: entomologic investigations. PLoS Negl. Trop. Dis.10, e0004981 (2016). 10.1371/journal.pntd.0004981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Nosrat, C. et al. Impact of recent climate extremes on mosquito-borne disease transmission in Kenya. PLoS Negl. Trop. Dis.15, e0009182 (2021). 10.1371/journal.pntd.0009182 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Blaylock, J. M. et al. The seroprevalence and seroincidence of dengue virus infection in western Kenya. Travel Med. Infect. Dis.9, 246–248 (2011). 10.1016/j.tmaid.2011.06.005 [DOI] [PubMed] [Google Scholar]
- 37.Alfsnes, K. et al. Tracing and tracking the emergence, epidemiology and dispersal of dengue virus to Africa during the 20th century. One Health13, 100337 (2021). 10.1016/j.onehlt.2021.100337 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Russo, G., Subissi, L. & Rezza, G. Chikungunya fever in Africa: a systematic review. Pathog. Glob. Health114, 111–119 (2020). 10.1080/20477724.2020.1748965 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chang, C., Ortiz, K., Ansari, A. & Gershwin, M. E. The Zika outbreak of the 21st century. J. Autoimmun.68, 1–13 (2016). 10.1016/j.jaut.2016.02.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Langat, S. K. et al. Origin and evolution of dengue virus type 2 causing outbreaks in Kenya: evidence of circulation of two cosmopolitan genotype lineages. Virus Evolution6, 1–9 (2020). [DOI] [PMC free article] [PubMed]
- 41.Letizia, A. G. et al. Retrospective genomic characterization of a 2017 dengue virus outbreak, Burkina Faso. Emerg. Infect. Dis.28, 1198–1210 (2022). 10.3201/eid2806.212491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Manigart, O., Ouedraogo, I., Ouedraogo, H. S., Sow, A. & Lokossou, V. K. Dengue epidemic in Burkina Faso: how can the response improve? Lancet Lond. Engl.403, 434–435 (2024). 10.1016/S0140-6736(23)02803-9 [DOI] [PubMed] [Google Scholar]
- 43.Heath, C. J. et al. Evidence of transovarial transmission of chikungunya and dengue viruses in field caught mosquitoes in Kenya. PLoS Negl. Trop. Dis.14, e0008362 (2020). 10.1371/journal.pntd.0008362 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Huber, J. H., Childs, M. L., Caldwell, J. M. & Mordecai, E. A. Seasonal temperature variation influences climate suitability for dengue, chikungunya, and Zika transmission. PLoS Negl. Trop. Dis.12, 1–20 (2018). 10.1371/journal.pntd.0006451 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Neto, Z. et al. Molecular and genomic investigation of an urban outbreak of dengue virus serotype 2 in Angola, 2017–2019. PLoS Negl. Trop. Dis.16, e0010255 (2022). 10.1371/journal.pntd.0010255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Vincent, M. et al. From the threat to the large outbreak: dengue on Reunion Island, 2015 to 2018. Eurosurveillance24, 1900346, (2019). 10.2807/1560-7917.ES.2019.24.47.1900346 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kelly, M. E. et al. Molecular characterization and phylogenetic analysis of dengue fever viruses in three outbreaks in Tanzania between 2017 and 2019. PLoS Negl. Trop. Dis.17, e0011289 (2023). 10.1371/journal.pntd.0011289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mwanyika, G. O. et al. Dengue virus infection and associated risk factors in africa: a systematic review and meta-analysis. Viruses13, 536 (2021). 10.3390/v13040536 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Harris, P. A. et al. Research electronic data capture (REDCap)—a metadata-driven methodology and workflow process for providing translational research informatics support. J. Biomed. Inform.42, 377–381 (2009). 10.1016/j.jbi.2008.08.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Waggoner, J. J. et al. Single-reaction multiplex reverse transcription PCR for detection of zika, Chikungunya, and dengue viruses. Emerg. Infect. Dis.22, 1295–1297 (2016). 10.3201/eid2207.160326 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Waggoner, J. J. et al. Single-reaction, multiplex, real-time RT-PCR for the detection, quantitation, and serotyping of dengue viruses. PLoS Negl. Trop. Dis.7, 1–9 (2013). 10.1371/journal.pntd.0002116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Grubaugh, N. D. et al. An amplicon-based sequencing framework for accurately measuring intrahost virus diversity using PrimalSeq and iVar. Genome Biol.20, 1–19 (2019). [DOI] [PMC free article] [PubMed]
- 53.Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics26, 589–595 (2010). 10.1093/bioinformatics/btp698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics25, 2078–2079 (2009). 10.1093/bioinformatics/btp352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Andrews, S. et al. FastQChttps://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2012).
- 56.Fonseca, V. et al. A computational method for the identification of Dengue, Zika and Chikungunya virus species and genotypes. PLoS Negl. Trop. Dis.13, e0007231 (2019). 10.1371/journal.pntd.0007231 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol.2, vew007 (2016). 10.1093/ve/vew007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Martin, D. P., Lemey, P. & Posada, D. Analysing recombination in nucleotide sequences. Mol. Ecol. Resour.11, 943–955 (2011). 10.1111/j.1755-0998.2011.03026.x [DOI] [PubMed] [Google Scholar]
- 59.Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics35, 4453–4455 (2019). 10.1093/bioinformatics/btz305 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Darriba, D. et al. ModelTest-NG: a new and scalable tool for the selection of DNA and protein evolutionary models. Mol. Biol. Evol.37, 291–294 (2020). 10.1093/molbev/msz189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Posada, D. & Crandall, K. A. MODELTEST: testing the model of DNA substitution. Bioinformatics14, 817–818 (1998). 10.1093/bioinformatics/14.9.817 [DOI] [PubMed] [Google Scholar]
- 62.Nahata, K. D. et al. SPREAD 4: online visualisation of pathogen phylogeographic reconstructions. Virus Evol. 8, veac088 (2022). [DOI] [PMC free article] [PubMed]
- 63.Pebesma, E. Simple features for R: standardized support for spatial vector data. R. J.10, 439 (2018). 10.32614/RJ-2018-009 [DOI] [Google Scholar]
- 64.Yu, G. Using ggtree to visualize data on tree‐like structures. Curr. Protoc. Bioinform.69, e96 (2020). 10.1002/cpbi.96 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of Additional Supplementary Files
Data Availability Statement
The sequences generated as part of this study have been deposited to Genbank with accession numbers: PP998553, PP998554, PP998555, PP998556, PP998557, PP998558 (https://www.ncbi.nlm.nih.gov/genbank/). All other sequences used in this analysis are available in the supplementary materials (Supplementary Data 2).
All.xml files generated as part of this study specifying the parameters of the Beast runs for phylogenetic and phylogeographic analyses are available online and can be downloaded at (https://github.com/SindisoNyathi/doctoralrepo_dengue_phylo/tree/main/beast_xmls).