Abstract
Bladder cancer (BC) is a prevalent type of tumor in the urinary system, and it has been discovered that long non‐coding RNA (lncRNA) plays a significant role in its occurrence and development. However, thus far, no reports have been published on the involvement of LINC00461 in BC. Here, we found that LINC00461 levels were upregulated in BC tissues and cell lines. Besides, knockdown of LINC00461 inhibited BC cell proliferation, migration, invasion through epithelial-mesenchymal transition (EMT), and slowed down tumor growth in vivo. Moreover, we found that LINC00461 regulated HNRNPUL1 expression through miR-518b sponge activity, and the miR-518 inhibitor could reverse the inhibitory effects of LINC00461 knockdown on BC cell proliferation, migration, and EMT. Our results suggest that LINC00461 may serve as a potential biomarker and therapeutic target for BC.
Supplementary Information
The online version contains supplementary material available at 10.1007/s12672-024-01294-5.
Keywords: LINC00461, EMT, MiR-518b, HNRNPUL1
Introduction
Bladder cancer (BC) is a common malignant tumor in the urinary and reproductive system, with about 500,000 new cases and 200,000 deaths worldwide every year [1]. In fact, its incidence ranks tenth among malignant tumors, making it a serious health concern. Most patients are initially diagnosed with non-muscle invasive bladder cancer (NMIBC). However, the treatment for NMIBC, which is mainly traditional transurethral bladder tumor resection (TUBRT), presents a significant challenge as there is a high risk of recurrence and progression after surgery. Shockingly, around 20% of NMIBC patients progress to muscle invasive bladder cancer (MIBC) within 5 years [2]. Although there are treatment methods such as radical cystectomy and chemotherapy, patients with metastasis have poor prognosis, and the 5-year survival rate is less than 50% [3]. Therefore, there is a critical need to find early diagnostic biomarkers or treatment targets that can improve patient prognosis.
LncRNA refers to a type of non-coding RNA that is typically longer than 200 nucleotides, and it is known to play a crucial role in various physiological processes, including transcription, translation, and chromatin modification [4, 5]. Additionally, lncRNA can act as a competing endogenous RNA (ceRNA) by binding to miRNA, which in turn regulates the protein levels of coding genes and thereby impacts the onset and progression of tumors [6]. Among these, LINC00461 is an essential RNA that is located on the fifth chromosome in humans [7]. For instance, in breast cancer tissues and cell lines, LINC00461 is significantly upregulated and promotes the tumor's progression by acting as a ceRNA of miR-30a-5p [8]. In liver cancer, LINC00461 has been found to regulate LRIG2 and promote the growth and invasion of liver cancer by “sponging” miR-149-5p [9]. However, its biological function in BC remains unclear.
Materials and methods
Patient tissue specimens and cell lines
Tumor and paracancerous tissues were collected from 32 patients diagnosed with BC at Shaoxing People’s Hospital between January 2020 and October 2022. All samples were histopathologically diagnosed by two experienced pathologists and stored in liquid nitrogen. All patients provided written informed consent before the operation, and no preoperative treatment was received. All experiments were approved by the Ethics Committee of our hospital. All the cell lines (5637, EJ, UMUC3, T24, SV-HUC-1) were obtained from the American Type Culture Collection (ATCC, Manassas, USA). 5637, EJ, UMUC3, and SV-HUC-1 cells were cultured in RPMI-1640 medium (Biochannel, Nanjing, China), while T24 cells were cultured in DMEM High Glucose (Biochannel, Nanjing, China) supplemented with 10% FBS (Biochannel, Nanjing, China) at 37 °C with 5% CO2.
Cell transfection
The siNC, siLINC00461, miR-518b inhibitor and miR-518b mimic were purchased from RiboBio (Guangzhou, China). The pcDNA3.1-NC and pcDNA3.1-LINC00461 (overLINC00461) were purchased from GenePharma (Shanghai, China). The sequences of siRNAs, inhibitor and mimic were shown in Supplementary Table 1. EJ and UMUC3 cells were cultured in 6-well plates (2 × 105 cells), before transfection cells density need reach 40–50%. The mixture including synthesized nucleotide and transfection reagent (RiboBio Guangzhou, China) were added to the plates, and the transfected cells were harvested 24 h or 48 h later to subsequent experiments.
qRT-PCR
We employed the RNA-quick purification kit (ESscience Biotech, Shanghai, China) to extract RNA from tissues and cells. Subsequently, cDNA was synthesized using the PrimeScript RT Reagent Kit (Takara, Beijing, China) and RT‐qPCR was conducted using TB Green Premix Ex Taq™ II (Takara, Beijing, China). The results were analyzed using the Light Cycler 480II PCR (Roche, Switzerland) and the expression levels of RNA were determined by the 2−ΔΔCq method. The primer sequences were shown in Supplementary Table 1.
Western blotting
The proteins were extracted by RIPA Lysis buffer (Beyotime, Shanghai, China) along with 1% Phosphatase inhibitor cocktail (MedChemExpress, USA). The proteins were then separated by 10% SDS-PAGE and transferred to PVDF membranes, before being sealed with 5% non-fat milk for 2 h. Subsequently, the membranes were incubated with primary antibodies overnight at 4 °C. The following day, secondary antibody (Proteintech Group, Chicago, USA) was used to incubate the membrane for 1 h at 37 °C, followed by 3 rinses with TBST. Finally, the protein bands were visualized using the ECL kit (Servicebio, Wuhan, China) and captured on Tanon 5200 (Tanon, Shanghai, China). The primary antibodies included HNRNPUL1 (Cat. No: 10578-1-AP) and β-actin (Cat. No: 81115-1-RR) were purchased from Proteintech Group (Wuhan, China). The primary antibodies included E-cadherin (Cat. No: 3195T), N-cadherin (Cat. No: 13116T) and Vimentin (Cat. No: 5741T) get from Cell Signaling Technology (USA).
FISH assay
The cells (EJ, UMUC3) were fixed with 4% paraformaldehyde and permeabilized with 1 ml PBS containing 0.5% TritonX-100. Subsequently, hybridization was performed with 100 μl hybridization solution containing the 2.5 μl Ribo lncRNA FISH Probe Mix (RiboBio, Guangzhou, China). After 10 min of staining with 1 × DAPI dye solution, the cells were rinsed with PBS three times, and the round coverslip was sealed with an anti-fluorescence quencher. Finally, the images were captured under a confocal laser scanning microscope (Leica, Germany). The FISH Probe sequence of LINC00461 was shown in Supplementary Table 1.
Cell proliferation assay
Seed the transfected EJ and UMUC3 cells (5 × 103 cells/well) into a 96 well plate, respectively. After 24, 48, 72 or 96 h, 10 μl CCK-8 reagent (Medchemexpress, USA) was added into each well and the optical density (OD) value was measured at 450 nm using a microplate reader (Molecular Devices, USA).
Cell invasion assay
To assess cell invasion, each trans-well chamber was coated with matrigel (BD, USA). Subsequently, 1 × 105 cells in serum-free medium were seeded in the upper trans-well chambers (Corning, USA) and 500 μl RP-MI 1640 medium containing 10% FBS was added to the lower chamber. After 24 h of culture at 5% CO2 and 37 °C, the cells were fixed with 4% paraformaldehyde for 15 min, then stained with 0.1% crystal violet for 15 min. Finally, the remaining cells in the upper chamber were removed with a cotton swab and visualized under an inverted light microscope (Nikon, Japan).
Wound healing assay
EJ and UMUC3 cells were seeded in 6‐well plates and allowed to reach a density of more than 90%. Wounds were then generated using a pipette tip, and the images of the wound fields or cells were photographed after 0 and 24 h using a microscope (Nikon, Japan).
Dua‐luciferase reporter assay
Artificially synthesized fragments of HNRNPUL1 and LINC00461 containing putative miR-518b binding sites or their mutant sequences were inserted into the pmirGLO vector (Promega, Beijing, USA) to construct LINC00461-WT, HNRNPUL1-WT, LINC00461-MUT and HNRNPUL1-MUT. The pmirGLO vectors WT and MUT were co-transfected with miR-518b mimic or miR-NC into cells. After 48 h of transfection, luciferase activity was evaluated using the Dual-Luciferase Reporter Assay System (Promega, USA).
Tumor xenografts in nude mice
T24 cell suspension (0.2 ml, 2 × 106 cells) was subcutaneously injected into the right armpit area of three-week-old female BALB/c nude mice (Slac Laboratory Animal Company, Shanghai, China). After one week, the mice were randomly assigned into two groups: siLINC00461 group and control group. The mice were administered with vivo-optimized antisense oligonucleotides (ASOs) (10 nmol, dissolved in 30 μL of sterile PBS) or negative control (RiboBio, Guangzhou, China) twice a week for three weeks by intratumoral injection, respectively. The mice were euthanized by cervical dislocation four weeks after tumor cell injection and their tumor volume was measured once a week using the formula: volume = (length × width2)/2.
Statistical analysis
All of the data presented as mean ± standard deviation (SD) were processed using the SPSS 20.0 software (SPSS, USA). All experiments were performed in triplicate. Paired or unpaired Student’s t-test were used to compare differences between two groups. p < 0.05 indicated a statistically significant difference.
Results
LINC00461 expression is up-regulated in BC tissues and cell lines
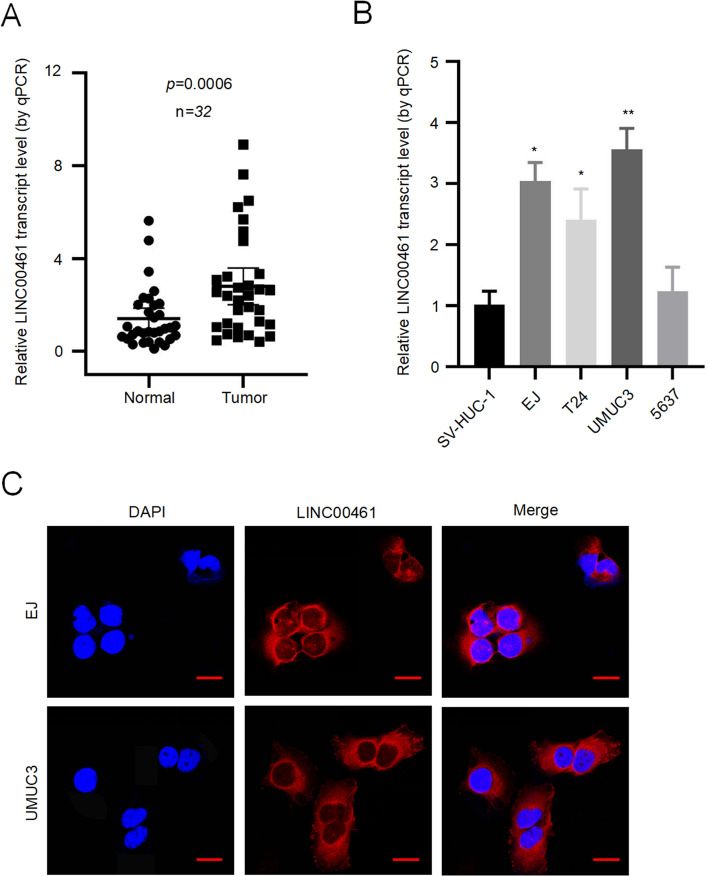
We observed that LINC00461 was highly expressed in BC tissues compared to normal tissues, as shown by qRT-PCR results (Fig. 1A). We also examined the expression of LINC00461 in BC cell lines (EJ, T24, UMUC3, and 5637) and normal urothelial cell line (SV-HUC-1). Our findings revealed that LINC00461 was significantly upregulated in cancer cells, especially in EJ and UMUC3 cells, compared to normal cells (Fig. 1B). Therefore, we selected these two cell lines for further experiments. FISH experiments were conducted, and the results showed that LINC00461 was primarily located in the cytoplasm (Fig. 1C).
Fig. 1.
LINC00461 was overexpressed in BC tissues and cell lines. A The expression of LINC00461 in BC tissues and normal tissues by qRT-PCR. Using paired Student’s t-test, n = 32, p = 0.0006. B The expression level of LINC00461 in BC cells (EJ, T24, UMUC3, 5637) and normal cell (SV-HUC-1), determined by qRT-PCR. C FISH assay to localize LINC00461 in BC cells (scale bar, 10 μm). *p < 0.05, **p < 0.01
Knockdown of LINC00461 inhibits BC cells proliferation, migration, invasion through EMT
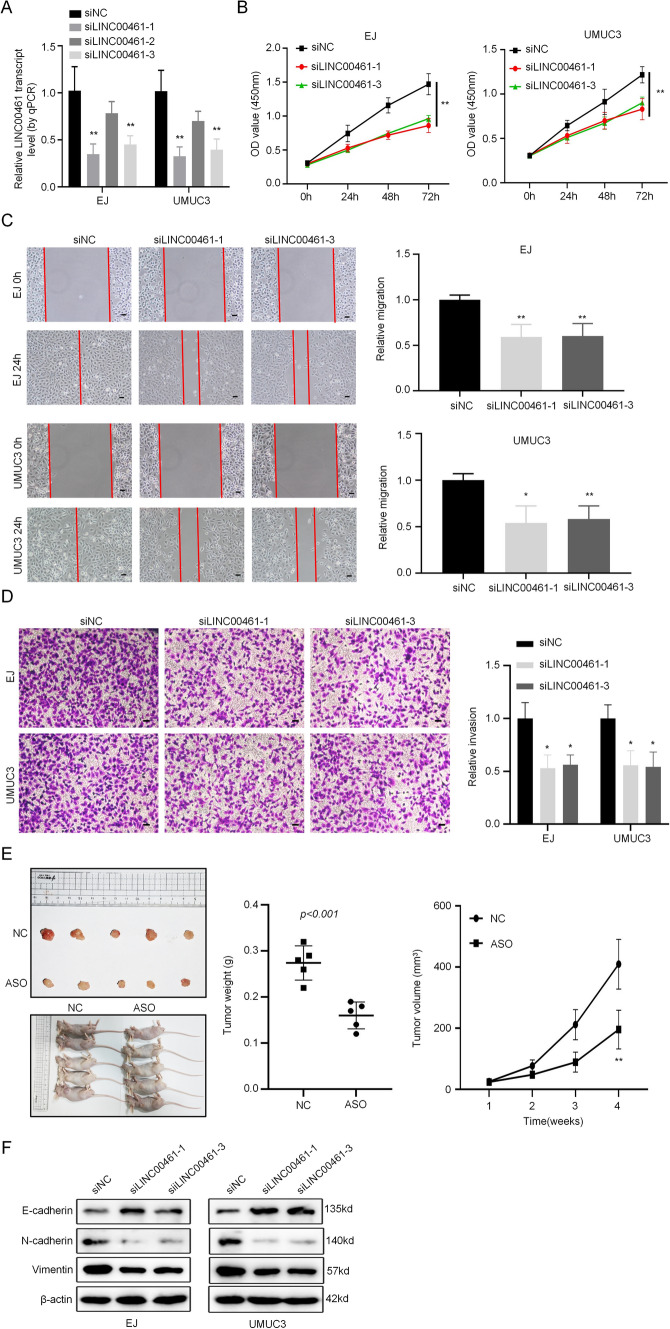
We investigated the potential biological roles of LINC00461 in BC cells by knocking down its expression using three different siRNAs in EJ and UMUC3 BC cells. The knockdown efficiency of LINC00461 was confirmed by qRT-PCR, and siLINC00461-1 and siLINC00461-3 showed greater knockdown efficiency compared to siLINC00461-2 (Fig. 2A), which were selected for subsequent experiments. CCK-8 assays revealed that the knockdown of LINC00461 attenuated the proliferative capacity of EJ and UMUC3 cells (Fig. 2B). Furthermore, we evaluated the effect of LINC00461 on BC cell migration and invasion using wound healing and cell invasion assays. The results showed that silencing LINC00461 significantly inhibited cell migration capacity in EJ and UMUC3 cells (Fig. 2C) and suppressed the invasive activity of both cell lines (Fig. 2D). Next, to investigate the effect of LINC00461 on the tumorigenicity of BC cells in vivo, we subcutaneously injected EJ/ASO or EJ/NC cells into BALB/c-nude mice. We found that knockdown of LINC00461 significantly reduced the tumor volume and growth rate compared to the control group (Fig. 2E). Finally, we analyzed the expression of EMT markers (E-cadherin, N-cadherin, and Vimentin) via Western blotting to determine whether LINC00461 regulates EMT in BC cells. Our results showed that LINC00461 knockdown inhibited the expression of N-cadherin and Vimentin but increased the expression of E-cadherin (Fig. 2F). These findings suggest that LINC00461 plays a regulatory role in cell proliferation, migration, and invasion in BC cells, and may serve as a potential therapeutic target for BC treatment. Consistently, overexpression of LINC00461 significantly promoted the cell proliferation and invasion ability of EJ and UMUC3 cells, and promoted the expression of N-cadherin and Vimentin but inhibited the expression of E-cadherin (Supplementary Fig. 1A–D).
Fig. 2.
Knockdown of LINC00461 inhibits BC cell proliferation, migration, invasion. A The efficiency of the LINC00461 knockdown was demonstrated by qRT-PCR. B The proliferation of EJ and UMUC3 following transfection with siLINC00461 was analyzed by CCK-8 assay. C, D Wound healing and cell invasion assays were utilized to assess the effect of knockdown of LINC00461 on BC cells migration and invasion (scale bar, 10 μm). E The image of subcutaneous tumor of nude mice injected with T24/siLINC00461 or T24/siNC cells, and the volume and weight of the tumors. F Protein levels of EMT markers (E-cadherin, N-cadherin, Vimentin) in cells transfected with siLINC00461 were confirmed by Western blotting. **p < 0.01
LINC00461 functions as a sponge for miR-518b to affect HNRNPUL1 expression in BC cells
A growing body of evidence suggests that lncRNA can act as a miRNA sponge, regulating downstream targets [10, 11]. To identify potential targets for LINC00461, we performed a bioinformatic analysis using the starbase V3.0 database, and found miR-518b was predicted to interact with LINC00461. To investigate the correlation between LINC00461 and miR-518b, we conducted a dual-luciferase reporter assay, which indicated that LINC00461 can directly bind to miR-518b (Fig. 3A, B). Additionally, qRT-PCR showed a decreased miR-518b expression level in tumor tissues and a negative correlation between LINC00461 expression and miR-518b expression in BC tissues (Fig. 3C, D). Then, we explored the effect of miR-518b on the biological function of bladder cancer. The results showed that overexpression of miR-518b significantly inhibited the proliferation and invasion of EJ and UMUC3 cells (Supplementary Fig. 2A, B). Moreover, our results showed that overexpression of miR-518b inhibited the expression of N-cadherin and Vimentin but increased the expression of E-cadherin (Supplementary Fig. 2C).
Fig. 3.
LINC00461 positively regulates HNRNPUL1 expression via sponging miR-518b. A Starbase predicted binding sites between LINC00461 and miR-518b. B Dua-luciferase reporter assay validated interaction between LINC00461 and miR-518b. C The expression of miR-518b in BC tissues and normal tissues by qRT-PCR (n = 32). D Pearson correlation analysis of the correlation between the relative expression levels of miR-518b and LINC00461. E Starbase predicted binding sites between HNRNPUL1-wild type or HNRNPUL1-mutant type and miR-518b. F Dua-luciferase reporter assay validated interaction between miR-518b and HNRNPUL1. G The inhibitory efficiency of the miR-518b inhibitor was demonstrated by means of qRT-PCR. H, I Detection of the expression of HNRNPUL1 in BC cells transfected with miR-518b inhibitor through qRT-PCR and Western blotting. J, K Detection of the expression of HNRNPUL1 in BC cells transfected with miR-518b mimic through qRT-PCR and Western blotting. L Protein levels of HNRNPUL1 in cells transfected with siLINC00461 were confirmed by Western blotting. *p < 0.05
We also predicted HNRNPUL1 as a potential target of miR-518b using the starbase V3.0 database, and a dual-luciferase reporter assay demonstrated that overexpression of miR-518b significantly reduced luciferase activity in a luciferase reporter harboring HNRNPUL1-WT but not in the reporter carrying HNRNPUL1-Mut (Fig. 3E, F). In addition, the expression levels of HNRNPUL1 were influenced by miR-518b, which were also influenced by LINC00461 (Fig. 3G–L). Furthermore, we found that protein expression level of HNRNPUL1 was significantly higher in 9/12 BC tumor tissues compared to adjacent normal tissues (Supplementary Fig. 3A).
Inhibiting miR-518b rescues the effects of LINC00461 knockdown on cell proliferation, invasion and EMT
To further validate our findings, we conducted rescue experiments. The CCK-8 assay revealed that knockdown of LINC00461 significantly inhibited cell growth. However, co-transfection of BC cells with the miR-518b inhibitor resulted in increased cell viability, indicating that the miR-518b inhibitor partially reversed the effects of LINC00461 silencing on cell proliferation (Fig. 4A). Similar results were obtained from our cell invasion assays, which showed that miR-518b inhibitors attenuated the impact of LINC00461 knockdown on cellular invasion (Fig. 4B). Furthermore, Western blotting analysis confirmed that LINC00461-silenced BC cells had significantly upregulated E-cadherin levels and significantly downregulated N-cadherin and vimentin levels. These effects were reversed in cells co-transfected with siLINC00461 and miR-518b inhibitors. Moreover, our Western blotting analysis revealed that downregulation of LINC00461 restored miR-518b inhibition-mediated repression of HNRNPUL1 protein levels (see Fig. 4C). These results suggest that the LINC00461/miR-518b/HNRNPUL1 axis may ultimately serve as a modulator of proliferation, invasion, and epithelial-mesenchymal transition (EMT) in BC cells.
Fig. 4.
Knockdown of LINC00461 inhibits BC cells EMT through miR-518b/HNRNPUL1 axis. A The miR-518b inhibitor partially reversed cell proliferation inhibition induced by silencing LINC00461. B The miR-518b inhibitor partially reversed cell invasion induced by silencing LINC00461 (scale bar, 10 μm). C The miR-518b inhibitor partially reversed the inhibition of EMT and HNRNPUL1 protein expression induced by silencing LINC00461. *p < 0.05, **p < 0.01
Discussion
Numerous studies have reported aberrant expression of lncRNA in BC, highlighting their role in the pathogenesis of this disease [12, 13]. The H19 gene, which is typically not expressed in adult tissues, is highly expressed in BC and was the first lncRNA identified as a BC oncogene [14]. Subsequently, other lncRNA, including LNMAT2, LNMAT1, and PTENP1, have been implicated in BC occurrence, development, diagnosis, and prognosis [15–17]. Additionally, increased expression of LINC00461 has been shown to promote cell proliferation, migration, and invasion in colorectal cancer cells by upregulating the expression of NFIB and CDK2 [18]. In this study, we found that LINC00461 was significantly overexpressed in BC tissues and cell lines and showed that it inhibits cell proliferation by regulating the expression of HNRNPUL1 via miR-518 targeting in BC. Our findings provide new insight into the novel tumor-inhibiting mechanism of LINC00461 in BC.
Heterogeneous nuclear ribonucleoprotein U-like 1 (HNRNPUL1) belongs to the HNRNP family, which is involved in various cellular processes, such as RNA metabolism, telomere elongation, DNA repair, and chromatin reorganization [19]. Among them, DNA damage are crucial in various malignant tumors, including BC. Recent studies have shown the involvement of non-coding RNA in DNA damage [20]. Interestingly, recent studies have shown that HNRNPs are closely involved in cancer, with HNRNPUL1 being highly expressed in various types of cancers [21]. Furthermore, the expression levels of HNRNPUL1 are also significantly associated with survival prognosis in esophageal cancer patients [22]. In light of these findings, our study focused on investigating the impact of the LINC00461/miR-518b/HNRNPUL1 axis on BC proliferation, invasion, and metastasis, as well as its connection with EMT, which broadens our understanding of non-coding RNA's role in BC development.
EMT is a process in which epithelial cells lose their polarity and adhesion ability, acquiring the migratory and invasive properties of mesenchymal cells. EMT plays a critical role in the development of normal tissues, wound healing, tumor metastasis, and organ fibrosis [23, 24]. In urogenital tumors, EMT induction has been shown to enhance the tumor's invasive and metastatic characteristics. Furthermore, EMT can lead to chemotherapy resistance due to the malignant behavior of tumor cells [25]. Recent research has found that non-coding RNAs (ncRNAs), such as lncRNA and microRNA, play a significant role in regulating EMT through various pathways [26]. These findings have opened up new possibilities for understanding the molecular mechanisms behind tumor metastasis, and have the potential to lead to the development of new therapeutic approaches for managing these challenging conditions.
In this study, we found that LINC00461 enhances BC cell proliferation, migration and invasion by sponging miR-518 to upregulate HNRNPUL1 expression and promote EMT in BC cells, suggesting that LINC00461 may serve as a potential biomarker and therapeutic target for BC.
Supplementary Information
Supplementary Material 1: Figure 1. Overexpression of LINC00461 promotes BC cells proliferation and invasion through EMT.The efficiency of overexpression of LINC00461 was demonstrated by qRT-PCR.The proliferation of EJ and UMUC3 following transfection with overLINC00461 was analyzed by CCK-8 assay.Trans-well assays were utilized to assess the effect of overexpression of LINC00461 on BC cells invasion.Protein levels of EMT markersin cells transfected with overLINC00461 were confirmed by Western blotting. *p < 0.05.
Supplementary Material 2: Figure 2. MiR-518b promotes BC cells proliferation and invasion through EMT.The proliferation of EJ and UMUC3 following transfection with miR-518b mimic was analyzed by CCK-8 assay.Trans-well assays were utilized to assess the effect of miR-518b mimic on BC cells invasion.Protein levels of EMT markersin cells transfected with miR-518b mimic were confirmed by Western blotting. *p < 0.05.
Supplementary Material 3: Figure 3. The expression level of HNRNPUL1 in BC tumor tissues.Western blotting assay was carried out to observe the protein expression level of HNRNPUL1 in bladder cancer tissues and their matched normal adjacent tissues collected from our hospital
Acknowledgements
Thanks for all participants involved in this study.
Author contributions
YZ was conceived the idea, participated in the design of the study, participated in the analysis and interpretation of data, and drafted the manuscript. KZ was conceived the idea, participated in the design of the study, and drafted the manuscript. JL conceived the idea, participated in the design of the study, and drafted the manuscript. CP was responsible for was responsible for Western blotting assay and participated in the analysis. JJ collected and classified the human BC tissue samples, and participated in the analysis of data. JC was in charge for cell experiment. YL was responsible for RNA extraction and PCR assay. GX collected the human BC tissue samples, and participated in the design of the study. SP was conceived the idea, participated in the design of the study, and drafted the manuscript.
Funding
This study was supported by Zhejiang Provincial Medicine, Health, and Science and Technology Project (No. 2022KY1297, No. 2023KY1253), and Zhejiang Province Traditional Chinese Medicine Science and Technology Plan Project (No. 2024ZL1135).
Data availability
All data related to the results of this study are available within the article.
Declarations
Ethics approval and consent to participate
The informed consent was signed by all participators. This study was proceeded after getting the approval of the Ethics Committee of Shaoxing People's Hospital. All methods in the study were carried out in accordance with the Helsinki guidelines and declaration or any other relevant guidelines. The study about animal experiment got authorization from the Experimental Animal Ethics Committee of Shaoxing People's Hospital and performed in accordance with the guidelines of the National Animal Care and Ethics Institution. It was carried out in compliance with the ARRIVE guidelines. Tumor size must not exceed 20 mm in any direction in an adult mouse according to the ethics committee. In our study, the maximal tumor size was not exceeded 20 mm.
Consent to participate
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yijie Zhou and Keyuan Zhao contributed equally to this work.
Contributor Information
Gang Xu, Email: shxdxy@163.com.
Shouhua Pan, Email: 13606550587@163.com.
References
- 1.Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. 10.3322/caac.21660 [DOI] [PubMed] [Google Scholar]
- 2.Abufaraj M, Dalbagni G, Daneshmand S, Horenblas S, Kamat AM, Kanzaki R, et al. The role of surgery in metastatic bladder cancer: a systematic review. Eur Urol. 2018;73(4):543–57. 10.1016/j.eururo.2017.09.030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Patel VG, Oh WK, Galsky MD. Treatment of muscle-invasive and advanced bladder cancer in 2020. CA Cancer J Clin. 2020;70(5):404–23. 10.3322/caac.21631 [DOI] [PubMed] [Google Scholar]
- 4.Yao RW, Wang Y, Chen LL. Cellular functions of long noncoding RNAs. Nat Cell Biol. 2019;21(5):542–51. 10.1038/s41556-019-0311-8 [DOI] [PubMed] [Google Scholar]
- 5.Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22(2):96–118. 10.1038/s41580-020-00315-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21(11):1253–61. 10.1038/nm.3981 [DOI] [PubMed] [Google Scholar]
- 7.Zhang Q, Zhong C, Shen J, Chen S, Jia Y, Duan S. Emerging role of LINC00461 in cancer. Biomed Pharmacother. 2022;152: 113239. 10.1016/j.biopha.2022.113239 [DOI] [PubMed] [Google Scholar]
- 8.Dong L, Qian J, Chen F, Fan Y, Long J. LINC00461 promotes cell migration and invasion in breast cancer through miR-30a-5p/integrin β3 axis. J Cell Biochem. 2019;120(4):4851–62. 10.1002/jcb.27435 [DOI] [PubMed] [Google Scholar]
- 9.Ji D, Wang Y, Li H, Sun B, Luo X. Long non-coding RNA LINC00461/miR-149-5p/LRIG2 axis regulates hepatocellular carcinoma progression. Biochem Biophys Res Commun. 2019;512(2):176–81. 10.1016/j.bbrc.2019.03.049 [DOI] [PubMed] [Google Scholar]
- 10.Nojima T, Proudfoot NJ. Mechanisms of lncRNA biogenesis as revealed by nascent transcriptomics. Nat Rev Mol Cell Biol. 2022;23(6):389–406. 10.1038/s41580-021-00447-6 [DOI] [PubMed] [Google Scholar]
- 11.Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172(3):393–407. 10.1016/j.cell.2018.01.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Chen C, Zheng H, Luo Y, Kong Y, An M, Li Y, et al. SUMOylation promotes extracellular vesicle-mediated transmission of lncRNA ELNAT1 and lymph node metastasis in bladder cancer. J Clin Invest. 2021;131(8): e146431. 10.1172/JCI146431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Li Y, Li G, Guo X, Yao H, Wang G, Li C. Non-coding RNA in bladder cancer. Cancer Lett. 2020;485:38–44. 10.1016/j.canlet.2020.04.023 [DOI] [PubMed] [Google Scholar]
- 14.Verhaegh GW, Verkleij L, Vermeulen SH, den Heijer M, Witjes JA, Kiemeney LA. Polymorphisms in the H19 gene and the risk of bladder cancer. Eur Urol. 2008;54(5):1118–26. 10.1016/j.eururo.2008.01.060 [DOI] [PubMed] [Google Scholar]
- 15.Chen C, Luo Y, He W, Zhao Y, Kong Y, Liu H, et al. Exosomal long noncoding RNA LNMAT2 promotes lymphatic metastasis in bladder cancer. J Clin Invest. 2020;130(1):404–21. 10.1172/JCI130892 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chen C, He W, Huang J, Wang B, Li H, Cai Q, et al. LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat Commun. 2018;9(1):3826. 10.1038/s41467-018-06152-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zheng R, Du M, Wang X, Xu W, Liang J, Wang W, et al. Exosome-transmitted long non-coding RNA PTENP1 suppresses bladder cancer progression. Mol Cancer. 2018;17(1):143. 10.1186/s12943-018-0880-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yu H, Ma J, Chen J, Yang Y, Liang J, Liang Y. LncRNA LINC00461 promotes colorectal cancer progression via miRNA-323b-3p/NFIB axis. Onco Targets Ther. 2019;12:11119–29. 10.2147/OTT.S228798 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gurunathan G, Yu Z, Coulombe Y, Masson JY, Richard S. Arginine methylation of hnRNPUL1 regulates interaction with NBS1 and recruitment to sites of DNA damage. Sci Rep. 2015;5:10475. 10.1038/srep10475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sidaway P. Bladder cancer: mutations in DNA-damage-repair pathways confer platinum sensitivity. Nat Rev Urol. 2017;14(4):197. 10.1038/nrurol.2017.22 [DOI] [PubMed] [Google Scholar]
- 21.Li H, Liu J, Shen S, Dai D, Cheng S, Dong X, et al. Pan-cancer analysis of alternative splicing regulator heterogeneous nuclear ribonucleoproteins (hnRNPs) family and their prognostic potential. J Cell Mol Med. 2020;24(19):11111–9. 10.1111/jcmm.15558 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Li J, Sang M, Zheng Y, Meng L, Gu L, Li Z, et al. HNRNPUL1 inhibits cisplatin sensitivity of esophageal squamous cell carcinoma through regulating the formation of circMAN1A2. Exp Cell Res. 2021;409(1): 112891. 10.1016/j.yexcr.2021.112891 [DOI] [PubMed] [Google Scholar]
- 23.Navas T, Kinders RJ, Lawrence SM, Ferry-Galow KV, Borgel S, Hollingshead MG, et al. Clinical evolution of epithelial–mesenchymal transition in human carcinomas. Cancer Res. 2020;80:304–18. 10.1158/0008-5472.CAN-18-3539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yang J, Antin P, Berx G, Blanpain C, Brabletz T, Bronner M, et al. Guidelines and definitions for research on epithelial–mesenchymal transition. Nat Rev Mol Cell Biol. 2020;21(6):341–52. 10.1038/s41580-020-0237-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.He P, Dai Q, Wu X. New insight in urological cancer therapy: from epithelial–mesenchymal transition (EMT) to application of nano-biomaterials. Environ Res. 2023;229: 115672. 10.1016/j.envres.2023.115672 [DOI] [PubMed] [Google Scholar]
- 26.Monteiro-Reis S, Lobo J, Henrique R, Jerónimo C. Epigenetic mechanisms influencing epithelial to mesenchymal transition in bladder cancer. Int J Mol Sci. 2019;20(2):297. 10.3390/ijms20020297 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material 1: Figure 1. Overexpression of LINC00461 promotes BC cells proliferation and invasion through EMT.The efficiency of overexpression of LINC00461 was demonstrated by qRT-PCR.The proliferation of EJ and UMUC3 following transfection with overLINC00461 was analyzed by CCK-8 assay.Trans-well assays were utilized to assess the effect of overexpression of LINC00461 on BC cells invasion.Protein levels of EMT markersin cells transfected with overLINC00461 were confirmed by Western blotting. *p < 0.05.
Supplementary Material 2: Figure 2. MiR-518b promotes BC cells proliferation and invasion through EMT.The proliferation of EJ and UMUC3 following transfection with miR-518b mimic was analyzed by CCK-8 assay.Trans-well assays were utilized to assess the effect of miR-518b mimic on BC cells invasion.Protein levels of EMT markersin cells transfected with miR-518b mimic were confirmed by Western blotting. *p < 0.05.
Supplementary Material 3: Figure 3. The expression level of HNRNPUL1 in BC tumor tissues.Western blotting assay was carried out to observe the protein expression level of HNRNPUL1 in bladder cancer tissues and their matched normal adjacent tissues collected from our hospital
Data Availability Statement
All data related to the results of this study are available within the article.