Abstract
The gut-brain axis mediates bidirectional signaling between the intestine and the nervous system and is critical for organism-wide homeostasis. Here we report the identification of a peptidergic endocrine circuit in which bidirectional signaling between neurons and the intestine potentiates the activation of the antioxidant response in C. elegans in the intestine. We identify a FMRF-amide-like peptide, FLP-2, whose release from the intestine is necessary and sufficient to activate the intestinal oxidative stress response by promoting the release of the antioxidant FLP-1 neuropeptide from neurons. FLP-2 secretion from the intestine is positively regulated by endogenous hydrogen peroxide (H2O2) produced in the mitochondrial matrix by sod-3/superoxide dismutase, and is negatively regulated by prdx-2/peroxiredoxin, which depletes H2O2 in both the mitochondria and cytosol. H2O2 promotes FLP-2 secretion through the DAG and calciumdependent protein kinase C family member pkc-2 and by the SNAP25 family member aex-4 in the intestine. Together, our data demonstrate a role for intestinal H2O2 in promoting inter-tissue antioxidant signaling through regulated neuropeptide-like protein exocytosis in a gut-brain axis to activate the oxidative stress response.
Keywords: C. elegans, oxidative stress, neuropeptide, hydrogen peroxide, mitochondria, superoxide dismutase, peroxiredoxin, thioredoxin, protein kinase C, SNAP25, SOD-1, SOD-3, PRDX-2, TRX-3, PKC-2, AEX-4
Introduction
The gut-brain axis is critical for communication between the intestine and the nervous system to regulate behavior and maintain homeostasis, and altered gut-brain signaling is associated with neurodegeneration, obesity and tumor proliferation (Carabotti et al. 2015; Grenham et al. 2011; Mayer, Nance, and Chen 2022; Mehrian-Shai et al. 2019; Vitali et al. 2022). Over the last decade the importance of peptides that function as signals in gut-brain signaling has gained recognition. Numerous gut peptides are distributed throughout the gastrointestinal (GI) tract with regional specificity (Haber et al. 2017), and gut-secreted peptides can modulate neurocircuits that regulate feeding behavior and glucose metabolism (Batterham and Bloom 2003; Han et al. 2018; Song et al. 2019), inflammatory responses against pathogenic bacteria (Campos-Salinas et al. 2014; Yu et al. 2021), and satiety (Batterham et al. 2002; Chelikani, Haver, and Reidelberger 2005; Gibbs, Young, and Smith 1973; Lutz et al. 1995; Lutz, Del Prete, and Scharrer 1994; West, Fey, and Woods 1984). A gut-released peptide suppresses arousal through dopaminergic neurons during sleep in Drosophila (Titos et al. 2023). In C. elegans, gut-derived peptides regulate rhythmic behavior and behavioral responses to pathogenic bacteria (Lee and Mylonakis 2017; Singh and Aballay 2019; Wang et al. 2013). Conversely, peptides released from the nervous system regulate many aspects of intestinal function including gut mobility, inflammation and immune defense (Browning and Travagli 2014; Furness et al. 2014; Lai, Mills, and Chiu 2017). In C. elegans, the secretion of peptides from various neurons regulates the mitochondrial unfolded protein response (UPRmt), the heat shock response, and the antioxidant response in the intestine (Jia and Sieburth 2021; Maman et al. 2013; Prahlad, Cornelius, and Morimoto 2008; Shao, Niu, and Liu 2016). In spite of the many roles of peptides in the gut-brain axis, the mechanisms underlying the regulation of intestinal peptide secretion and signaling remain to be fully defined.
Hydrogen peroxide (H2O2) is emerging as an important signaling molecule that regulates intracellular signaling pathways by modifying specific reactive residues on target proteins. For example, H2O2-regulated phosphorylation of inhibitor of nuclear factor κB (NF-κB) kinase, leads to the activation of NF-κB during development, inflammation and immune responses. (Kamata et al. 2002; Oliveira-Marques et al. 2009; Takada et al. 2003). In addition, H2O2-induced tyrosine and cysteine modifications contribute to redox regulation of c-Jun N-terminal kinase 2 (JNK2), Src family kinase, extracellular signalregulated kinases 1 and 2 (ERK1/2), protein kinase C (PKC) and other protein kinases (Kemble and Sun 2009; Konishi et al. 1997; Lee et al. 2003; Nelson et al. 2018). H2O2 signaling has been implicated in regulating neurotransmission and transmitter secretion. H2O2 at low concentration increases neurotransmission at neuromuscular junctions without influencing lipid oxidation (A R Giniatullin and R A Giniatullin 2003; Giniatullin, Petrov, and Giniatullin 2019; Shakirzyanova et al. 2009), and enhanced endogenous H2O2 generation regulates dopamine release (Avshalumov et al. 2005; Avshalumov and Rice 2003; Bao, Avshalumov, and Rice 2005; Chen, Avshalumov, and Rice 2001, 2002). Acute H2O2 treatment increases exocytosis of ATP-containing vesicles in astrocytes (Li et al. 2019). Finally, mitochondrially-derived H2O2 regulates neuropeptide release from neurons in C. elegans (Jia and Sieburth 2021). Cellular H2O2 levels are tightly controlled through the regulation of its production from superoxide by superoxide dismutases (SODs) and cytoplasmic oxidases (Fridovich 1995, 1997; Messner and Imlay 2002; Zelko, Mariani, and Folz 2002), and through its degradation by catalases, peroxidases, and peroxiredoxins (Chance, Sies, and Boveris 1979; Marinho et al. 2014). In the intestine, endogenously produced H2O2 plays important roles as an antibacterial agent in the lumen, and in activating the ER unfolded protein response (UPRER) through protein sulfenylation (Botteaux et al. 2009; Corcionivoschi et al. 2012; Hourihan et al. 2016; Miller et al. 2020). Here we demonstrate a role of endogenous H2O2 signaling in the intestine in regulating the release of the intestinal FMRFamide-like peptide, FLP-2, to modulate a neurocircuit that activates the anti-oxidant response in the intestine in C. elegans. Intestinal FLP-2 signaling functions by potentiating the release of the antioxidant neuropeptide-like protein FLP-1 from AIY interneurons, which in turn activates the antioxidant response in the intestine. FLP-2 secretion from the intestine is rapidly and positively regulated by H2O2, whose levels are positively regulated by superoxide dismutases in the mitochondrial matrix and cytosol, and negatively regulated by the peroxiredoxin-thioredoxin system in the cytosol. Intestinal FLP-2 release is mediated by aex-4/SNAP25-dependent exocytosis of dense core vesicles and H2O2-induced FLP-2 secretion is dependent upon the production of intestinal diacylglycerol and on pkc2/PKCα/β kinase activity.
Results
Neuronal FLP-1 secretion is regulated by neuropeptide signaling from the intestine.
We previously showed that 10 minute treatment with the mitochondrial toxin juglone leads to a rapid, reversible and specific increase in FLP-1 secretion from AIY, as measured by a two-fold increase in coelomocyte fluorescence in animals expressing FLP-1::Venus fusion proteins in AIY (Fig. 1A and B, (Jia and Sieburth 2021)). Coelomocytes take up secreted neuropeptides by bulk endocytosis (Fares and Greenwald 2001) and the fluorescence intensity of Venus in their endocytic vacuoles is used as a measure of regulated neuropeptide secretion efficacy (Ailion et al. 2014; Ch’ng, Sieburth, and Kaplan 2008; Sieburth, Madison, and Kaplan 2006). To determine the role of the intestine in regulating FLP-1 secretion, we first examined aex-5 mutants. aex-5 encodes an intestinal subtilisin/kexin type 5 prohormone convertase that functions to proteolytically process peptide precursors into mature peptides in dense core vesicles (DCVs) (Edwards et al. 2019; Thacker and Rose 2000), and aex-5 mutants are defective in peptide signaling from the intestine (Mahoney et al. 2008). We found that aex-5 mutants expressing FLP-1::Venus in AIY exhibited no significant difference in coelomocyte fluorescence compared to wild type controls in the absence of juglone (Fig. 1B). However, coelomocyte fluorescence did not significantly increase in aex-5 mutants treated with juglone. Expression of aex-5 cDNA selectively in the intestine (under the ges-1 promoter) fully restored normal responses to juglone to aex-5 mutants, whereas aex-5 cDNA expression in the nervous system (under the rab-3 promoter) failed to rescue (Fig. 1B). Thus, peptide processing in intestinal DCVs is necessary for juglone-induced FLP-1 secretion from AIY.
Figure 1. Peptidergic gut-to-neuron FLP-2 signaling potentiates the oxidative stress response.
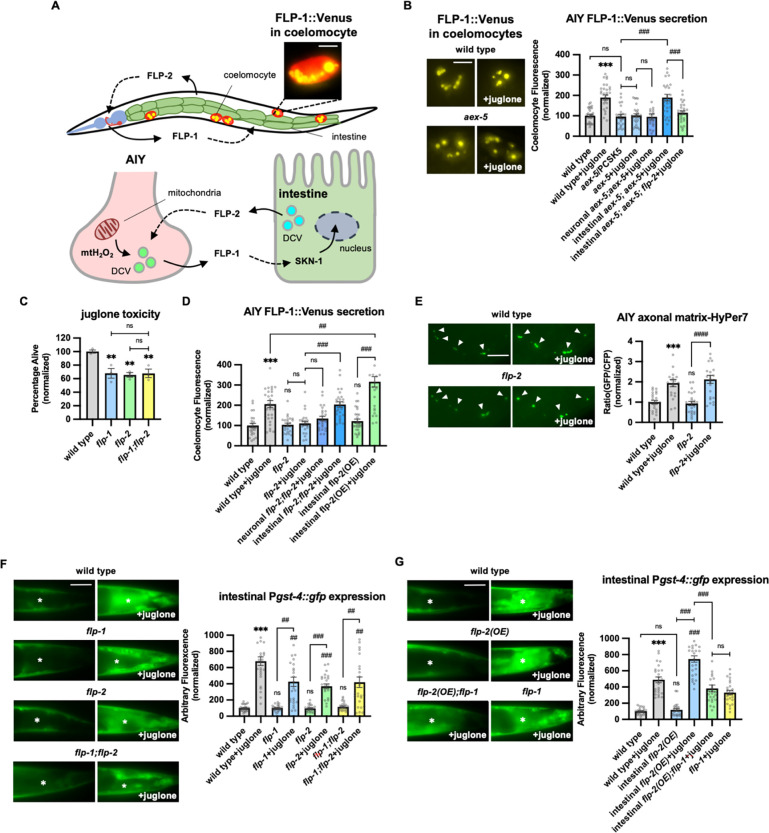
A (Top) Schematic showing the positions of AIY, intestine and coelomocytes of transgenic animals coexpressing FLP-1::Venus in the intestine and mCherry in coelomocytes. Representative image of the posterior coelomocyte that has taken up Venus into the endocytic compartment. Scale bar: 5μM. (Bottom) Schematic showing FLP-1 and FLP-2 peptides as inter-tissue signals in gut-intestine regulation of the antioxidant response.
B Representative images and quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-1::Venus fusion proteins in AIY following M9 or 300μM juglone treatment for 10min. Neuronal aex-5 denotes expression of aex-5 cDNA under the rab-3 promoter; intestinal aex-5 denotes expression of aex-5 cDNA under the ges-1 promoter. Unlined *** denotes statistical significance compared to “wild type”. n = 30, 30, 24, 30, 26, 30, 30 independent animals. Scale bar: 5μM.
C Average percentage of surviving young adult animals of the indicated genotypes after 16h recovery following 4h juglone treatment. Unlined ** denotes statistical significance compared to “wild type”. n = 213, 156, 189, 195 independent biological samples over three independent experiments.
D Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-1::Venus fusion proteins in AIY following M9 or 300μM juglone treatment for 10min. Neuronal flp-2 denotes expression of flp-2 gDNA under the rab-3 promoter; intestinal flp-2 denotes expression of flp-2 gDNA under the ges-1 promoter; intestinal flp-2(OE) denotes expression of flp-2 gDNA under the ges-1 promoter in wild type animals. Unlined *** and ns denote statistical significance compared to “wild type”. n = 20, 20, 25, 20, 20, 20, 25, 22 independent animals.
E Representative images and quantification of fluorescence of mitochondrial matrix-targeted HyPer7 in the axon of AIY following M9 or 300μM juglone treatment for 10min. Arrowheads denote puncta marked by mito::HyPer7 fusion proteins (Excitation: 500 and 400nm; emission: 520nm). Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation was used to measure H2O2 levels. Unlined *** and ns denote statistical significance compared to “wild type”. n = 24, 22, 25, 24 independent animals. Scale bar: 10μM. F Representative images and quantification of average fluorescence in the posterior intestine of transgenic animals expressing Pgst-4::gfp after 1h M9 or juglone exposure and 3h recovery. Asterisks mark the intestinal region used for quantification. Pgst-4::gfp expression in the body wall muscles, which appears as fluorescence on the edge animals in some images, was not quantified. Unlined *** and ns denote statistical significance compared to “wild type”; unlined ## and ### denote statistical significance compared to “wild type+juglone”. n = 25, 26, 25, 25, 25, 25, 25, 25 independent animals. Scale bar: 10μM.
G Representative images and quantification of average fluorescence in the posterior region of transgenic animals expressing Pgst-4::gfp after 1h M9 or juglone exposure and 3h recovery. Asterisks mark the intestinal region for quantification. Pgst-4::gfp expression in the body wall muscles, which appears as fluorescence on the edge animals in some images, was not quantified. Unlined *** denotes statistical significance compared to “wild type”; unlined ### denotes statistical significance compared to “wild type+juglone”. n = 23, 25, 25, 26, 24, 25 independent animals. Scale bar: 10μM.
B-G Data are mean values ± s.e.m normalized to wild type controls. ns. not significant, ** and ## P < 0.01, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test.
Next, we examined a number of mutants with impaired SNARE-mediated vesicle release in the intestine including aex-1/UNC13, aex-3/MADD, aex-4/SNAP25b, and aex-6/Rab27 (Fig. 2C (Iwasaki et al., 1997; Mahoney et al., 2006; Thacker & Rose, 2000; Thomas, 1990; Wang et al., 2013)), and they each exhibited no increases in FLP-1 secretion following juglone treatment above levels observed in untreated controls (Fig. S1A). NLP-40 is a neuropeptide-like protein whose release from the intestine is presumed to be controlled by aex-1, aex-3, aex-4 and aex-6 (Lin-Moore, Oyeyemi, and Hammarlund 2021; Mahoney et al. 2008; Shi et al. 2022; Wang et al. 2013). Null mutants in nlp-40 or its receptor, aex-2 (Wang et al. 2013), exhibited normal juglone-induced FLP-1 secretion (Fig. S1B). These results establish a gut-to-neuron signaling pathway that regulates FLP1 secretion from AIY that is likely to be controlled by peptidergic signaling distinct from NLP-40.
Figure 2. FLP-2 secretion from the intestine is stress regulated.
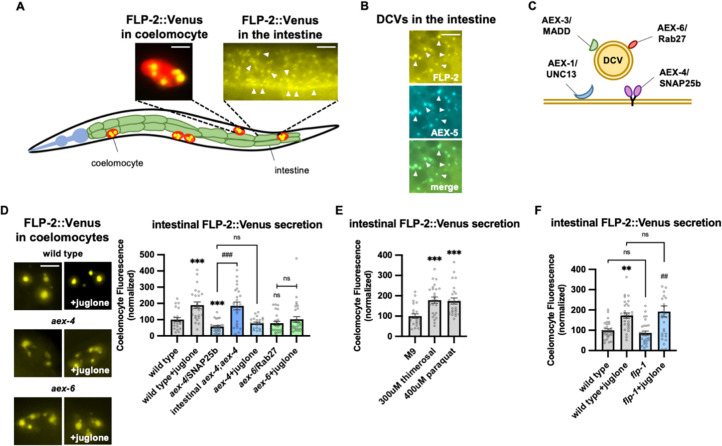
A Schematic showing the positions of intestine and coelomocytes of transgenic animals co-expressing FLP2::Venus in the intestine and mCherry in coelomocytes. Representative images of the posterior coelomocyte that have taken up Venus into the endocytic compartment (Scale bar: 5μM) and the posterior intestinal region showing the distribution of FLP-2::Venus in puncta in the intestine are shown (Scale bar: 15μM).
B Representative images of fluorescence distribution in the posterior intestinal region of transgenic animals co-expressing FLP-2::Venus and AEX-5::mTur2 fusion proteins. Arrowheads denote puncta marked by both fusion proteins. Scale bar: 5μM.
C Schematic showing the locations of AEX-1/UNC13, AEX-3/MADD, AEX-4/SNAP25 and AEX-6/Rab27 relative to a DCV.
D Representative images and quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone for 10min. Unlined *** and ns denote statistical significance compared to “wild type”. n = 29, 25, 24, 30, 23, 30, 25, 25, 25 independent animals. Scale bar: 5μM.
E Quantification of average coelomocyte fluorescence of transgenic animals expressing FLP-2::Venus fusion proteins in the intestine following treatment with M9 buffer or the indicated stressors for 10min. Unlined *** denotes statistical significane compared to “M9”. n = 23, 25, 25 independent animals.
F Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Unlined ** denotes statistical significance compared to “wild type”; unlined ## denotes statistical significance compared to “flp1”; a denotes statistical significance compared to “wild type+juglone”. n = 30, 30, 30, 30 independent animals.
D-F Data are mean values ± s.e.m normalized to wild type controls. ns. not significant, ** and ## P < 0.01, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test.
FLP-2 signaling from the intestine potentiates neuronal FLP-1 secretion and the oxidative stress response.
flp-1 protects animals from the toxic effects of juglone (Jia and Sieburth 2021). We reasoned that the intestinal signal that regulates FLP-1 secretion should also protect animals from juglone-induced toxicity. We identified the FMRF-amide neuropeptide-like protein, flp-2, in an RNA interference (RNAi) screen for neuropeptides that confer hypersensitivity to juglone toxicity upon knockdown (Jia and Sieburth 2021). flp-2 signaling has been implicated in regulating lifespan, reproductive development, locomotion during lethargus, and the mitochondrial unfolded protein response (UPRmt) (Chai et al. 2022; Chen et al. 2016; Kageyama et al. 2022; Shao et al. 2016). Putative flp2(ok3351) null mutants, which eliminate most of the flp-2 coding region, are superficially as healthy as wild type animals, but they exhibited significantly reduced survival in the presence of juglone compared to wild type controls (Fig. 1C). The reduced survival rate of flp-2 mutants was similar to that of flp-1 mutants, and flp-1; flp-2 double mutants exhibited survival rates that were not more severe than those of single mutants (Fig. 1C), suggesting that flp-1 and flp-2 may function in a common genetic pathway.
To determine whether flp-2 signaling regulates FLP-1 secretion from AIY, we examined FLP-1::Venus secretion. flp-2 mutants exhibited normal levels of FLP-1 secretion in the absence of stress, but FLP-1 secretion failed to significantly increase following juglone treatment of flp-2 mutants (Fig. 1D). flp-2 is expressed in a subset of neurons as well as the intestine (Chai et al. 2022), and flp-2 functions from the nervous system for its roles in development and the UPRmt (Chai et al. 2022; Chen et al. 2016; Kageyama et al. 2022; Shao et al. 2016). Expressing a flp-2 genomic DNA, fragment (containing both the flp-2a and flp-2b isoforms that arise by alternative splicing), specifically in the nervous system failed to rescue the FLP-1::Venus defects of flp-2 mutants, whereas expressing flp-2 selectively in the intestine fully restored juglone-induced FLP-1::Venus secretion to flp-2 mutants (Fig. 1D). These results indicate that flp2 signaling is dispensable for FLP-1 secretion from AIY under normal conditions, but that flp-2 originating from the intestine is necessary to increase FLP-1 secretion during oxidative stress.
To address how flp-2 signaling regulates FLP-1 secretion from AIY, we examined H2O2 levels in AIY using a mitochondrially targeted pH-stable H2O2 sensor HyPer7 (mito-HyPer7, Pak et al. 2020). Mito-HyPer7 adopted a punctate pattern of fluorescence in AIY axons, and the average fluorescence intensity of axonal mito-HyPer7 puncta increased about two-fold following 10 minute juglone treatment (Fig 1E), in agreement with our previous studies using HyPer (Jia and Sieburth 2021), confirming that juglone rapidly increases mitochondrial AIY H2O2 levels. flp-2 mutations had no significant effects on the localization or the average intensity of mito-HyPer7 puncta in AIY axons either in the absence of juglone, or in the presence of juglone (Fig 1E), suggesting that flp-2 signaling promotes FLP-1 secretion by a mechanism that does not increase H2O2 levels in AIY. Consistent with this, intestinal overexpression of flp-2 had no effect on FLP-1::Venus secretion in the absence of juglone, but significantly enhanced the ability of juglone to increase FLP-1 secretion (Fig. 1D). We conclude that both elevated mitochondrial H2O2 levels and intact flp-2 signaling from the intestine are necessary to increase FLP-1 secretion from AIY.
Previously we showed that FLP-1 signaling from AIY positively regulates the activation of the antioxidant transcription factor SKN-1/Nrf2 in the intestine. Specifically, flp-1 mutations impair the juglone-induced expression of the SKN-1 reporter transgene Pgst-4::gfp (Fig. 1F and (Jia and Sieburth 2021)). We found that mutations in flp-2 caused a similar reduction in juglone-induced Pgst-4::gfp expression as flp-1 mutants, and that flp-1; flp-2 double mutants exhibited similar impairments in juglone-induced Pgst-4::gfp expression as flp-1 or flp-2 single mutants (Fig. 1F). Conversely, overexpression of flp-2 selectively in the intestine elevated juglone-induced Pgst-4::gfp expression, without altering baseline Pgst-4::gfp expression, and the elevated Pgst-4::gfp expression in juglone-treated animals overexpressing flp-2 was entirely dependent upon flp-1 (Fig. 1G). It’s noteworthy that overexpressing flp-2 in the intestine did not enhance FLP-1::Venus release or Pgst-4::gfp expression in the absence of stress, indicating that the FLP-1 mediated anti-oxidant pathway by flp-2 is stress-activated. Together this data indicates that flp-2 signaling originating in the intestine positively regulates the stress-induced secretion of FLP-1 from AIY, as well as the subsequent activation of anti-oxidant response genes in the intestine. We propose that FLP-1 and FLP-2 define a bidirectional gut-neuron signaling axis, whereby during periods of oxidative stress, FLP-2 released from the intestine positively regulates FLP-1 secretion from AIY, and FLP-1, in turn, potentiates the antioxidant response in the intestine (Fig. 1A).
FLP-2 secretion from the intestine is H2O2-regulated.
To directly investigate the mechanisms underlying the regulation of FLP-2 secretion, we examined FLP-2::Venus fusion proteins expressed in the intestine under various conditions (Fig. 2A). FLP-2::Venus fusion proteins adopted a punctate pattern of fluorescence throughout the cytoplasm of intestinal cells and at the plasma membrane (Fig. 2A), and FLP-2::Venus puncta co-localized with the DCV cargo protein AEX-5/PCSK5 tagged to mTurquoise2 (AEX-5::mTur2, Fig. 2B). FLP-2::Venus fluorescence was also observed in the coelomocytes (marked by mCherry) (Fig. 2A, D), indicating that FLP-2 is released from the intestine. SNAP25 forms a component of the core SNARE complex, which drives vesicular membrane fusion and transmitter release (Chen and Scheller 2001; Goda 1997; Jahn and Scheller 2006). aex-4 encodes the C. elegans homolog of SNAP25, and mutations in aex-4 disrupt the secretion of neuropeptides from the intestine (Lin-Moore et al. 2021; Mahoney et al. 2008; Wang et al. 2013) (Fig. 2C). We found that aex-4 null mutations significantly reduced coelomocyte fluorescence in FLP-2::Venus expressing animals, and expression of aex-4 cDNA selectively in the intestine fully restored FLP-2 secretion to aex-4 mutants (Fig. 2D). Together these results suggest that intestinal FLP-2 can be packaged into DCVs that undergo release via SNARE-dependent exocytosis.
To test whether intestinal FLP-2 secretion is regulated by oxidative stress, we examined coelomocyte fluorescence in FLP-2::Venus-expressing animals that had been exposed to a number of different commonly used oxidative stressors. We found that 10 minute exposure to juglone, thimerosal, or paraquat, which promote mitochondria-targeted toxicity (Castello, Drechsel, and Patel 2007; Elferink 1999; Sharpe, Livingston, and Baskin 2012), each significantly increased Venus fluorescence intensity in the coelomocytes compared to untreated controls (Fig. 2D and E). We conducted four controls for specificity: First, juglone treatment did not significantly alter fluorescence intensity of mCherry expressed in coelomocytes (Fig. S2A). Second, impairing intestinal DCV secretion (by either aex-4/SNAP25 or aex-6/Rab27 mutations (Lin-Moore et al. 2021; Mahoney et al. 2006, 2008; Thomas 1990), blocked the juglone-induced increase in coelomocyte fluorescence in FLP-2::Venus-expressing animals (Fig. 2C). Third, nlp-40 and nlp-27 encode neuropeptide-like proteins that are released from the intestine but are not implicated in stress responses (Liu et al. 2023; Taylor et al. 2021; Wang et al. 2013). Juglone treatment had no detectable effects on coelomocyte fluorescence in animals expressing intestinal NLP-40::Venus or NLP-27::Venus fusion proteins (Fig. S2B and C), and NLP-40::mTur2 puncta did not overlap with FLP-2::Venus puncta in the intestine (Fig. S2D). Finally, flp-1 mutants exhibited wild type levels of FLP-2 secretion both in the absence and presence of juglone (Fig. 2E). The distribution of FLP-2::Venus puncta in the intestine was not detectably altered by juglone treatment. Together, these results indicate that acute oxidative stress selectively increases the exocytosis of FLP-2-containing DCVs from the intestine, upstream of flp-1 signaling.
SOD-1 and SOD-3 superoxide dismutases regulate FLP-2 release.
Juglone generates superoxide anion radicals (Ahmad and Suzuki 2019; Paulsen and Ljungman 2005) and juglone treatment of C. elegans increases ROS levels (de Castro, Hegi de Castro, and Johnson 2004) likely by promoting the global production of mitochondrial superoxide. Superoxide can then be rapidly converted into H2O2 by superoxide dismutase. To determine whether H2O2 impacts FLP-2 secretion, we first examined superoxide dismutase mutants. C. elegans encodes five superoxide dismutase genes (sod-1 through sod-5). sod-1 or sod-3 null mutations blocked juglone-induced FLP2 secretion without altering baseline FLP-2 secretion, whereas sod-2, sod-4, or sod-5 mutations had no effect on FLP-2 secretion in the presence of juglone (Fig. 3A and B, S3A). sod-1; sod-3 double mutants exhibited juglone-induced FLP-2 secretion defects that was similar to single mutants, without significantly altering FLP-2 secretion in the absence of stress (Fig. 3C). sod-1 encodes the ortholog of mammalian SOD1, which is a cytoplasmic SOD implicated in the development of amyotrophic lateral sclerosis (ALS) and cancer (Giglio et al. 1994; Papa, Manfredi, and Germain 2014; Wang et al. 2021; Zhang et al. 2007). SOD-1::fusion proteins adopted a diffuse pattern of fluorescence in intestinal cells, consistent with a cytoplasmic localization (Fig. 3D). Transgenes expressing the sod-1 cDNA selectively in the intestine fully rescued the juglone-induced FLP-2::Venus secretion defects of sod-1 mutants (Fig. 3A). sod-3 encodes a homolog of mammalian SOD2, which is a mitochondrial matrix SOD implicated in protection against oxidative stress-induced neuronal cell death (Fukui and Zhu 2010; Vincent et al. 2007). Intestinal SOD-3::GFP fusion proteins localized to round structures that were surrounded by the outer membrane mitochondrial marker TOMM-20::mCherry (Ahier et al. 2018), consistent with a mitochondrial matrix localization (Fig. 3E). Expression of sod-3 cDNA in the intestine fully restored juglone-induced FLP-2 release to sod-3 mutants (Fig. 3B). sod3 variants lacking the mitochondrial localization sequence (sod-3(DMLS)), were no longer localized to mitochondria (Fig. 3F) and failed to restore normal responsiveness to juglone to sod-3 mutants (Fig. 3B). Thus, the generation of H2O2 by either SOD-1 in the cytoplasm or by SOD-3 in the mitochondrial matrix is necessary for juglone to increase FLP-2 secretion.
Figure 3. SOD-1/SOD-3 mediates endogenous H2O2 regulates FLP-2 release from the intestine.
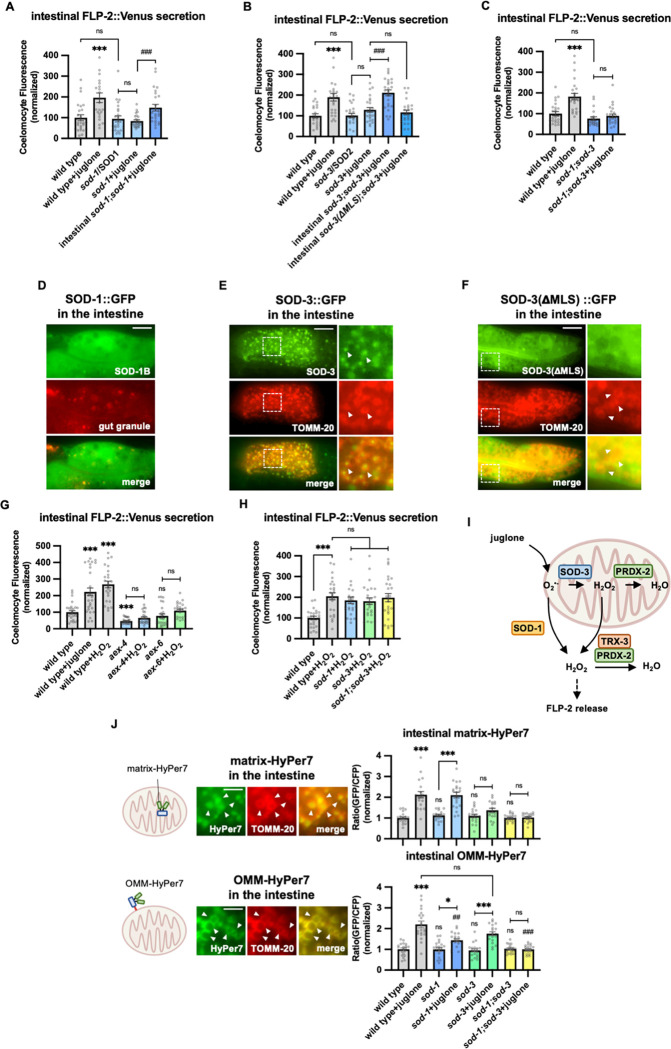
A Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Intestinal sod-1 denotes expression of sod-1b cDNA under the ges-1 promoter. Unlined *** denotes statistical significance compared to “wild type”. n = 25, 22, 24, 24, 25 independent animals.
B Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Intestinal sod-3 and sod3(ΔMLS) denote intestinal expression of sod-3 cDNA and sod-3(ΔMLS) variants, which lacks the mitochondrial localization sequence, under the ges-1 promoter. Unlined *** denotes statistical significance compared to “wild type”. n = 25, 25, 25, 25, 25, 25 independent animals.
C Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Unlined *** denotes statistical significance compared to “wild type”. n = 25, 25, 22, 25 independent animals.
D Representative images of fluorescence distribution in the posterior intestinal region of transgenic animals expressing SOD-1b::GFP fusion proteins in contrast against auto-fluorescence of gut granules. Scale bar: 10μM.
E Representative images of fluorescence distribution in the posterior intestinal region of transgenic animals co-expressing SOD-3::GFP and TOMM-20::mCherry (to target mitochondria) fusion proteins. Scale bar: 15μM.
F Representative images of fluorescence distribution in the posterior intestinal region of transgenic animals co-expressing SOD-3(ΔMLS)::GFP and TOMM-20::mCherry fusion proteins. Scale bar: 15μM.
G Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9, 300μM juglone or 1mM H2O2 treatment for 10min. Unlined *** and ns denote statistical significance compared to “wild type”. n = 29, 30, 25, 25, 25, 24, 25 independent animals.
H Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 1mM H2O2 treatment for 10min. n = independent animals. I Schematic showing that SOD-1 and SOD-3 mediate juglone-induced H2O2 production in promoting FLP2 release, and the PRDX-2/TRX-3 system detoxifies excessive H2O2.
J Schematic, representative images and quantification of fluorescence in the posterior region of the indicated transgenic animals co-expressing mitochondrial matrix targeted HyPer7 (matrix-HyPer7) or mitochondrial outer membrane targeted HyPer7 (OMM-HyPer7) with TOMM-20::mCherry following M9 or 300μM juglone treatment. Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation and 520nm for emission was used to measure H2O2 levels. Unlined *** and ns denote statistical significance compared to “wild type”. Unlined ## and ### denote statistical significance compared to “wild type+juglone”. (top) n = 20, 20, 18, 20, 19, 19, 20, 20 independent animals. (bottom) n = 20, 20, 19, 20, 20, 20, 20, 20 independent animals. Scale bar: 5μM.
A-C, G-H and J Data are mean values ± s.e.m normalized to wild type controls. ns. not significant, * P < 0.05, ## P < 0.01, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test.
Next, to determine if H2O2 can regulate FLP-2 secretion, we treated animals acutely with exogenous H2O2. We found that 10 minute H2O2 treatment increased FLP2::Venus secretion to a similar extent as 10 minute juglone treatment. aex-4/SNAP25, or aex-6/Rab27 mutants exhibited no increase in FLP-2 secretion in response to H2O2 treatment compared to untreated controls (Fig. 3G). In contrast, sod-1 or sod-3 mutants (or sod-1; sod-3 double mutants) exhibited an increase in FLP-2 secretion in response to H2O2 that was similar to that of wild type controls (Fig. 3H), suggesting that exogenous H2O2 can bypass the requirement of SODs but not SNAREs to promote FLP-2 secretion. Together these results suggest that H2O2 generated by SODs can positively regulate intestinal FLP-2 exocytosis from DCVs (Fig. 3I).
SOD-1 and SOD-3 regulate intestinal mitochondrial H2O2 levels.
To directly monitor H2O2 levels in the intestine, we generated transgenic animals in which HyPer7 was targeted to either the mitochondrial matrix (matrix-HyPer7) by generating fusion proteins with the cytochrome c MLS, or to the cytosolic face of the outer mitochondrial membrane (OMM-HyPer7) by generating fusion proteins with TOMM-20. When co-expressed in the intestine with the OMM marker TOMM-20::mCherry, matrixHyPer7 formed round structures throughout the cytoplasm that were surrounded by the OMM, and OMM-HyPer7 formed ring-like structures throughout the cytoplasm that colocalized with the OMM marker (Fig. 3J). Ten minute treatment with H2O2 significantly increased the fluorescence intensity by about two-fold of both matrix-HyPer7 and OMMHyPer7 without altering mitochondrial morphology or abundance (Fig. 3J, S3B and C), validating the utility of HyPer7 as a sensor for acute changes in H2O2 levels in and around intestinal mitochondria.
Juglone treatment for 10 minutes led to a similar two-fold increase in matrixHyPer7 fluorescence as H2O2 treatment (Fig. 3J). sod-3 mutations did not alter baseline H2O2 levels in the matrix, but they completely blocked juglone-induced increases H2O2 levels, whereas sod-1 mutations had no effect on either baseline or juglone-induced increases H2O2 levels (Fig. 3J). These results indicate that superoxide produced by juglone treatment is likely to be converted into H2O2 by SOD-3 in the mitochondrial matrix (Fig. 3I).
Next, we examined H2O2 levels on the outer surface of mitochondria using OMMHyPer7 and we found that juglone treatment led to a two-fold increase in OMM-HyPer7 fluorescence, similar to H2O2 treatment (Fig. 3J). sod-3 or sod-1 mutations did not alter baseline H2O2 levels on the OMM, but sod-1 single mutations attenuated, juglone-induced increases in OMM H2O2 levels, while sod-3 mutations had no effect (Fig. 3J). In sod-1; sod-3 double mutants, the juglone-induced increase in OMM H2O2 levels was completely blocked, whereas baseline H2O2 levels in the absence of stress were unchanged (Fig. 3J). These results suggest that sod-3 and sod-1 are exclusively required for H2O2 production by juglone and that both mitochondrial SOD-3 and cytosolic SOD-1 contribute to H2O2 levels in the cytosol. One model that could explain these results is that juglonegenerated superoxide is converted into H2O2 both by SOD-3 in the matrix, and by SOD1 in the cytosol, and that the H2O2 generated in the matrix can exit the mitochondria to contribute to cytosolic H2O2 levels needed to drive FLP-2 secretion (Fig. 3I).
The peroxiredoxin-thioredoxin system regulates endogenous H2O2 levels and FLP2 secretion.
To determine whether endogenous H2O2 regulates FLP-2 secretion, we examined mutations in the peroxiredoxin-thioredoxin system. Peroxiredoxins and thioredoxins detoxify excessive H2O2 by converting it into water and they play a critical role in maintaining cellular redox homeostasis (Netto and Antunes 2016) (Fig. 4A). C. elegans encodes two peroxiredoxin family members, prdx-2 and prdx-3, that are expressed at high levels in the intestine (Taylor et al. 2021). Null mutations in prdx-2 significantly increased FLP-2::Venus secretion compared to wild type animals in the absence of stress (Fig. 4B), whereas null mutations in prdx-3 had no effect on FLP-2 secretion (Fig. S4A). We observed a corresponding increase in both matrix-HyPer7 and OMM-HyPer7 fluorescence intensity in prdx-2 mutants (Fig. 4C and D), demonstrating that endogenous H2O2 is neutralized by peroxiredoxin and establishing a correlation between endogenous H2O2 levels and FLP-2 secretion. The increase in FLP-2 secretion in prdx-2 mutants was not further increased by juglone treatment (Fig. 4B). These results suggest that elevation in the levels of endogenously produced H2O2 in the intestine can positively regulate FLP2 secretion.
Figure 4. PRDX-2/PRDX and TRX-3/TRX regulate endogenous H2O2 and FLP-2 secretion.
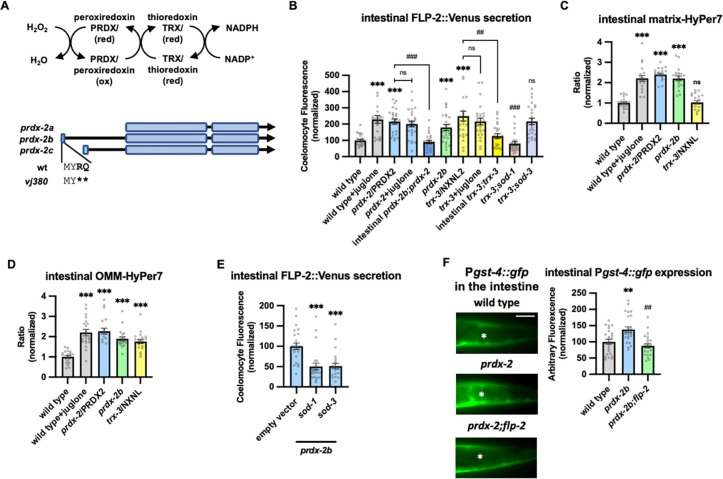
A (Top) Schematic showing the PRDX/TRX system in H2O2 detoxification. (Bottom) Schematic showing the three isoforms of prdx-2 transcripts and vj380 allele of prdx-2b knockout.
B Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Intestinal prdx-2b denotes expression of prdx-2b cDNA under the ges-1 promoter. Intestinal trx-3 denotes expression of trx-3 cDNA under the ges-1 promoter. Unlined *** denotes statistical significance compared to “wild type”; unlined ## and ### denote statistical significance compared to “trx-3”. n= 25, 23, 25, 25, 25, 25, 25, 25, 25, 25, 25 independent animals.
C and D Quantification of fluorescence in the posterior region of the indicated transgenic animals coexpressing matrix-HyPer7 (C) or OMM-HyPer7 (D) with TOMM-20::mCherry following M9 or 300μM juglone treatment. Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation and 520nm for emission was used to measure H2O2 levels. Unlined *** and ns denote statistical significance compared to “wild type. (C) n = 20, 20, 20, 20, 20 independent animals. (D) n = 20, 20, 20, 20, 20 independent animals. E Quantification of average coelomocyte FLP-2::Venus fluorescence of transgenic animals fed with RNAi bacteria targeting the indicated genes following M9 treatment for 10min. Unlined *** denotes statistical significance compared to “empty vector”. n = 25, 23, 24 independent animals.
F Representative images and quantification of average fluorescence in the posterior region of transgenic animals expressing Pgst-4::gfp after 1h M9 or juglone exposure and 3h recovery. Asterisks mark the intestinal region for quantification. Pgst-4::gfp expression in the body wall muscles, which appears as fluorescence on the edge animals in some images, was not quantified. Unlined ** denotes statistical significance compared to “wild type”, unlined ## denotes statistical analysis compared to “prdx-2b”. n = 25, 25, 25 independent animals. Scale bar: 10μM.
B-F Data are mean values ± s.e.m normalized to wild type controls. n.s. not significant, ** and ## P < 0.01, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test.
There are three isoforms of prdx-2 that arise by the use of alternative transcriptional start sites (Fig. 4A). Expressing the prdx-2b isoform selectively in the intestine fully rescued the elevated FLP-2::Venus secretion defects of prdx-2 mutants, whereas expressing prdx-2a or prdx-2c isoforms failed to rescue (Fig. 4B, S4B and C). To independently verify the role of prdx-2b function in FLP-2 release, we generated a prdx-2b-specific knockout mutant by introducing an in frame stop codon within the prdx2b-specific exon 1 using CRISPR/Cas9 (prdx-2b(vj380) Fig. 4A). prdx-2b(vj380) mutants exhibited increased H2O2 levels in the mitochondrial matrix and OMM (Fig. 4C and D), as well as increased FLP-2::Venus secretion compared to wild type controls that were indistinguishable from prdx-2 null mutants (Fig. 4B). prdx-2b mutations could no longer increase FLP-2 secretion when either sod-1 or sod-3 activity was impaired (Fig. 4E and Fig. S4D). Thus, the prdx-2b isoform normally inhibits FLP-2 secretion likely by promoting the consumption of H2O2 in the mitochondrial matrix and/or cytosol.
Once oxidized, peroxiredoxins are reduced by thioredoxins (TRXs) for reuse (Netto and Antunes 2016). TRX-3 is an intestine-specific thioredoxin promoting protection against specific pathogen infections (Jiménez-Hidalgo et al. 2014; Miranda-Vizuete, Damdimopoulos, and Spyrou 2000; Netto and Antunes 2016). Mutations in trx-3 elevated FLP-2::Venus release in the absence of juglone and expressing trx-3 transgenes in the intestine restored wild type FLP-2 release to trx-3 mutants (Fig. 4B). Juglone treatment failed to further enhance FLP-2::Venus release in trx-3 mutants (Fig. 4B). Mutations in cytoplasmic sod-1 but not in mitochondrial sod-3 reduced the elevated FLP-2::Venus release in trx-3 mutants to wild type levels (Fig. 4B). Mutations in trx-3 increased H2O2 levels in the OMM but had no effect on matrix H2O2 levels (Fig. 4C and D). Thus TRX-3 likely functions in the cytosol but not in the matrix to neutralize H2O2, and elevated H2O2 levels in the cytosol are sufficient to drive FLP-2 secretion without SOD-3-mediated H2O2 generation in the matrix.
Finally, to investigate the physiological significance of elevated endogenous H2O2 levels on the oxidative stress response we examined the effects of prdx-2b mutations on expression of gst-4. prdx-2b mutants had significantly increased Pgst-4::gfp expression in the intestine compared to wild type controls (Fig. 4F). The increased Pgst-4::gfp expression in prdx-2b mutants was completely dependent upon flp-2 signaling, since gst4 expression was reduced to wild type levels in prdx-2b; flp-2 double mutants (Fig. 4F). Together our data suggest that prdx-2b functions in the intestine to maintain redox homeostasis following SOD-1/SOD-3 mediated H2O2 production by regulating the secretion of FLP-2 (Fig. 3I).
PKC-2/PKCα/β mediates H2O2 induced FLP-2 secretion from the intestine.
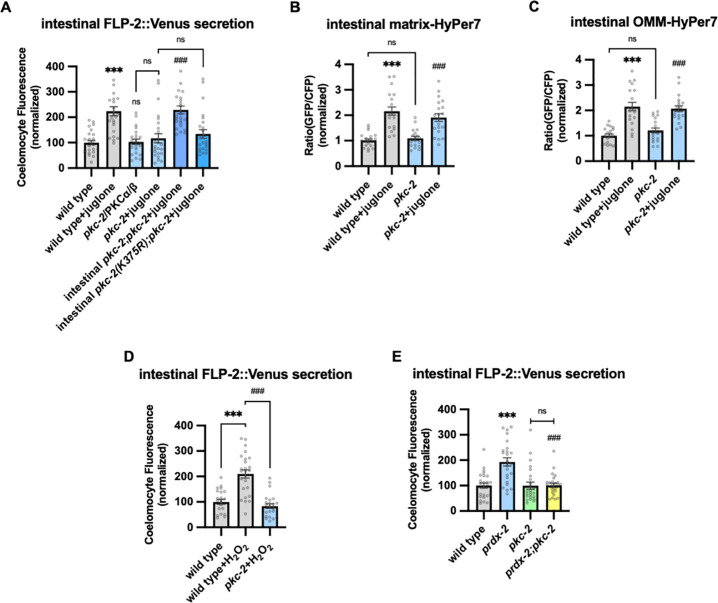
H2O2 functions as a cellular signaling molecule by oxidizing reactive cysteines to sulfenic acid, and this modification on target proteins can regulate intracellular signaling pathways (García Santamarina, Boronat i Llop, and Hidalgo Hernando 2014). One of the validated targets of H2O2 signaling is the protein kinase C (PKC) family of serine threonine kinases (Jia and Sieburth 2021; Konishi et al. 1997, 2001; Min, Kim, and Exton 1998). C. elegans encodes four PKC family members including pkc-1 and pkc-2, which are expressed at highest levels in the intestine (Islas-Trejo et al. 1997; Taylor et al. 2021). pkc-1 null mutants had no effect on baseline or juglone-induced FLP-2 secretion (Fig. S5A). pkc-2 null mutations did not alter baseline intestinal FLP-2 secretion, but they eliminated juglone-induced FLP-2 secretion (Fig. 5A). pkc-2 encodes a calcium and diacylglycerol (DAG) stimulated PKCα/β protein kinase C that regulates thermosensory behavior by promoting transmitter secretion (Edwards et al. 2012; Land and Rubin 2017). Expressing pkc-2 cDNA selectively in the intestine fully restored juglone-induced FLP-2 secretion to pkc-2 mutants (Fig. 5A), whereas expressing a catalytically inactive pkc2(K375R) variant (Van et al. 2021) failed to rescue (Fig. 5A). The intestinal site of action of pkc-2 is in line with prior studies showing that pkc-2 can function in the intestine to regulate thermosensory behavior (Land and Rubin 2017). pkc-2 mutants exhibited wild type H2O2 levels in the mitochondrial matrix and OMM of the intestine in both the presence and absence of juglone (Fig. 5B and C). Increasing H2O2 levels by either acute H2O2 treatment or by prdx-2 mutation failed to increase FLP-2 secretion in pkc-2 mutants (Fig. 5D and E). To determine whether pkc-2 can regulate the intestinal secretion of other peptides that are not associated with oxidative stress , we examined expulsion frequency, which is a measure of intestinal NLP-40 secretion (Mahoney et al. 2008; Wang et al. 2013). pkc-2 mutants showed wild type expulsion frequency (Fig. S5B), indicating that intestinal NLP-40 release is largely unaffected. Together, these results show that pkc-2 is not a general regulator of intestinal peptide secretion and instead functions downstream or in parallel to H2O2 to selectively promote FLP-2 secretion by a mechanism that involves phosphorylation of target proteins.
Figure 5. PKC-2/PKCα/β activation by H2O2 promotes FLP-2 secretion from the intestine.
A Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Intestinal pkc-2 denotes expression of pkc-2b cDNA under the ges-1 promoter. Intestinal pkc-2b(K375R) denotes expression of pkc2b(K375R) variants under the ges-1 promoter. Unlined *** and ns denote statistical significance compared to “wild type”; ### denote statistical significance compared to “pkc-2+juglone”. n = 24, 24, 25, 25, 25, 25 independent animals.
B and C Quantification of fluorescence in the posterior region of the indicated transgenic animals coexpressing matrix-HyPer7 (B) or OMM-HyPer7 (C) with TOMM-20::mCherry following M9 or 300μM juglone treatment. Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation and 520nm for emission was used to measure H2O2 levels. Unlined *** denotes statistical significance compared to “wild type”; unlined ### denotes statistical analysis compared to “pkc-2”. (B) n = 20, 20, 19, 20 independent animals, (C) n = 20, 20, 20, 20 independent animals.
D Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 1mM H2O2 treatment for 10min. n = 23, 25, 25 independent animals.
E Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 treatment for 10min. Unlined *** denotes statistical significance compared to “wild type”; unlined ### denotes statistical significance compared to “prdx-2”. n = 25, 25, 25, 25 independent animals.
A-E Data are mean values ± s.e.m normalized to wild type controls. ns. not significant, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test.
DAG positively regulates FLP-2 secretion.
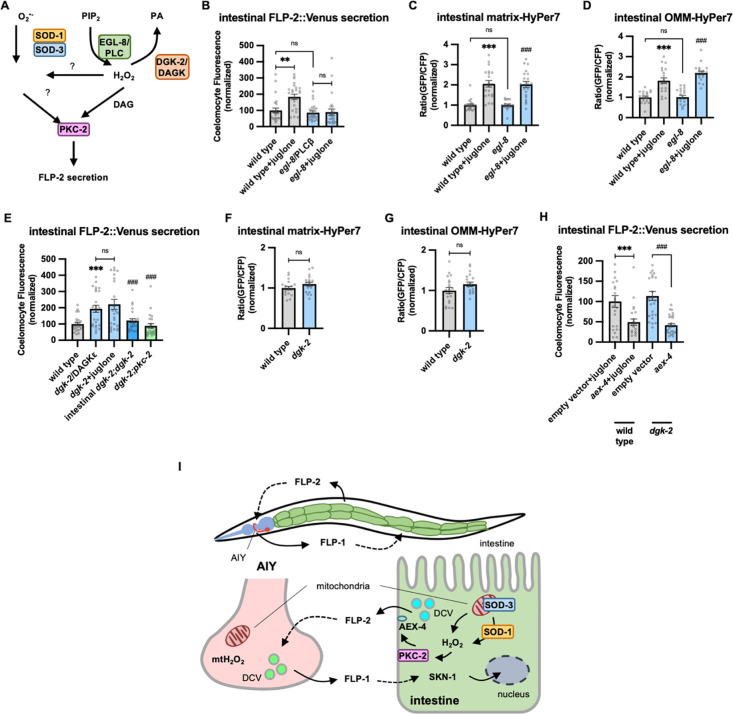
PKCα/β family members contain two N terminal C1 domains (C1A and C1B) whose binding to DAG promotes PKC recruitment to the plasma membrane (Burns and Bell 1991; Darby, Meng, and Fehrenbacher 2017; Johnson, Giorgione, and Newton 2000; Kim et al. 2016; Ono et al. 1989; Yanase et al. 2011). To address the role of DAG in promoting FLP2 secretion by PKC-2, we examined mutants that are predicted to have altered DAG levels. Phosphatidylinositol phospholipase C beta (PLCβ) converts phosphatidyl inositol phosphate (PIP2) to DAG and inositol triphosphate (IP3, Fig. 6A), and impairing PLC activity leads to reduced cellular DAG levels (Nebigil 1997). C. elegans encodes two PLC family members whose expression is enriched in the intestine, plc-2/ PLCβ and egl-8/ PLCβ (Taylor et al. 2021). plc-2 null mutants exhibited baseline and juglone-included FLP2 secretion that were similar to wild type controls (Fig. S6). egl-8 loss-of-function mutants exhibited wild type baseline FLP-2 secretion, but juglone-induced FLP-2 secretion was completely blocked (Fig. 6B). H2O2 levels in egl-8 mutants were similar to wild type controls, both in the presence and absence of juglone (Fig. 6C and D). Thus, egl-8/PLCb functions downstream of or in parallel to H2O2 production to promote FLP-2 secretion.
Figure 6. DAG promotes PKC-2 mediated FLP-2 secretion from the intestine.
A Schematic showing PLC and DGK mediates DAG metabolism and DAG functions in H2O2-mediated FLP2 signaling.
B Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or juglone treatment for 10min. n = 25, 25, 25, 25 independent animals.
C and D Quantification of fluorescence in the posterior region of the indicated transgenic animals coexpressing matrix-HyPer7 (C) or OMM-HyPer7 (D) with TOMM-20::mCherry following M9 or 300μM juglone treatment. Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation and 520nm for emission was used to measure H2O2 levels. Unlined *** denotes statistical significance compared to “wild type”; unlined ### denotes statistical significance compared to “egl-8”. (C) n = 22, 20, 20, 21 independent animals, (D) n = 20, 20, 20, 20 independent animals.
E Quantification of average coelomocyte fluorescence of the indicated mutants expressing FLP-2::Venus fusion proteins in the intestine following M9 or 300μM juglone treatment for 10min. Intestinal dgk-2 denotes expression of dgk-2a cDNA under the ges-1 promoter. Unlined *** denotes statistical significance compared to “wild type”; unlined ### denotes statistical significance compared to “dgk-2/DGKε”. n = 25, 25, 25, 25, 24 independent animals.
F and G Quantification of fluorescence in the posterior region of the indicated transgenic animals coexpressing matrix-HyPer7 (F) or OMM-HyPer7 (G) with TOMM-20::mCherry following M9 treatment. Ratio of images taken with 500nM (GFP) and 400nM (CFP) for excitation and 520nm for emission was used to measure H2O2 levels. (F) n = 20, 20 independent animals, (G) n = 20, 20 independent animals.
H Quantification of average coelomocyte fluorescence of the indicated transgenic animals fed with RNAi bacteria targeting the indicated genes in the intestine following M9 treatment for 10min. n = 25, 24, 25, 30 independent animals.
I (Top) Schematic showing the position of intestine and AIY neurons in FLP-1-FLP-2 mediated axis. (Bottom) Schematic showing endogenous H2O2 promotes PKC-2/AEX-4 mediated FLP-2 release from the intestine in FLP-1-FLP-2 regulated inter-tissue axis.
B-H Data are mean values ± s.e.m normalized to wild type controls. B-E and H ns. not significant, ** P < 0.01, *** and ### P < 0.001 by Brown-Forsythe and Welch ANOVA with Dunnett’s T3 multiple comparisons test. F and G ns. not significant by unpaired t test with Welch’s correction.
DAG kinase converts DAG into phosphatidic acid (PA), and is therefore a negative regulator or DAG levels (Fig. 6A) (van Blitterswijk and Houssa 2000; Topham 2006). In C. elegans, dgk-2/DGKε is the highest expressing DAG kinase in the intestine. Mutations in dgk-2 elevated FLP-2::Venus secretion (Fig. 6E) without altering H2O2 levels in the intestinal mitochondrial matrix or OMM (Fig. 6F and G). Expressing dgk-2 transgenes selectively in the intestine restored normal FLP-2::Venus secretion to dgk-2 mutants (Fig. 6E). Finally, the increase in FLP-2 secretion in dgk-2 mutants was not further increased by juglone treatment, but it was completely blocked by pkc-2 mutations or aex-4/SNAP25 mutations (Fig. 6E and H). These results show that FLP-2 secretion can be regulated bidirectionally by DAG, and they suggest that DAG and H2O2 function in a common genetic pathway upstream of pkc-2 to promote FLP-2 secretion (Fig. 6A).
Discussion
By screening for intercellular regulators of FLP-1 signaling from the nervous system in promoting the anti-oxidant response, we have uncovered a function for peptidergic signaling in mediating gut-to-neuron regulation of the anti-oxidant response in C. elegans. We identified the neuropeptide-like protein FLP-2 as an inter-tissue signal originating in the intestine to potentiate stress-induced FLP-1 release from AIY neurons and the subsequent activation of SKN-1 in the intestine. We found that H2O2 generated endogenously in the intestine or exogenously by acute oxidant exposure increases FLP2 secretion from intestinal DCVs. H2O2 promotes FLP-2 exocytosis through PKC-2, and AEX-4/SNAP25. The use of oxidant-regulated peptide secretion exemplifies a mechanism that can allow the gut and the nervous system to efficiently and rapidly communicate through endocrine signaling to promote organism-wide protection in the face of intestinal stress (Fig. 6I).
A new function for flp-2 signaling in the antioxidant response.
Previous studies have identified roles for flp-2 signaling in development and in stress responses. flp-2 promotes locomotion during molting (Chen et al. 2016) promotes entry into reproductive growth (Chai et al. 2022), regulates longevity (Kageyama et al. 2022), and activates the UPRmt cell-non autonomously during mitochondrial stress (Shao et al. 2016). The function we identified for flp-2 in the antioxidant response has some notable similarities with flp-2’s other functions. First, flp-2 mediates its effects at least in part by regulating signaling by other peptides. flp-2 signaling increases the secretion of the neuropeptide like protein PDF-1 during lethargus (Chen et al. 2016) and INS35/insulin-like peptide for its roles in reproductive growth choice and longevity (Kageyama et al. 2022), in addition to regulating AIY FLP-1 secretion (Fig. 5G).. Second, the secretion of FLP-2 is dynamic. FLP-2 secretion decreases during lethargus (Chen et al. 2016) and increases under conditions that do not favor reproductive growth (Kageyama et al. 2022), as well increasing in response to oxidants (Fig. 2C and D). However, in some instances, the regulation of FLP-2 secretion may occur at the level of flp-2 expression (Kageyama et al. 2022), rather than at the level of exocytosis (Fig. 2C). Finally, genetic analysis of flp2 has revealed that under normal conditions, flp-2 signaling may be relatively low, since flp-2 mutants show no defects in reproductive growth choice when animals are well fed (Chai et al. 2022), show only mild defects in locomotion during molting in non-sensitized genetic backgrounds (Chen et al. 2016), and do not have altered baseline FLP-1 secretion or antioxidant gene expression in the absence of exogenous oxidants (Fig. 1D and E). It is notable that increased ROS levels are associated with molting (Back et al. 2012; Knoefler et al. 2012), ageing (Back et al. 2012; Van Raamsdonk and Hekimi 2010), starvation (Tao et al. 2017), and mitochondrial dysfunction (Dingley et al. 2010), raising the possibility that flp-2 may be used in specific contexts associated with high ROS levels to affect global changes in physiology, behavior and development.
One major difference we found for flp-2 signaling in our study is that intestinal, but not neuronal flp-2 activates the oxidative stress response, whereas flp-2 originates from neurons for its reported roles in development and the UPRmt. The intestine is uniquely poised to relay information about diet to the rest of the animal, and secretion of a number of neuropeptide-like proteins from the intestine (e.g, INS-11, PDF-2 and INS-7) is proposed to regulate responses to different bacterial food sources (Lee and Mylonakis 2017; Murphy, Lee, and Kenyon 2007; O’Donnell et al. 2018). Since bacterial diet can impact ROS levels in the intestine (Pang and Curran 2014), secretion of FLP-2 from the intestine could function to relay information about bacterial diet to distal tissues to regulate redox homeostasis. In addition, the regulation of intestinal FLP-2 release by oxidants may meet a unique spatial, temporal or concentration requirement for activating the antioxidant response that cannot be met by its release from the nervous system.
AIY as a target for flp-2 signaling.
AIY interneurons receive sensory information from several neurons primarily as glutamatergic inputs to regulate behavior (Bargmann et al. 2007; Clark et al. 2006; Satoh et al. 2014). Our study reveals a previously undescribed mechanism by which AIY is activated through endocrine signaling originating from FLP-2 secretion from the intestine. FLP-2 could act directly on AIY, or it may function indirectly through upstream neurons that relay FLP-2 signals to AIY. frpr-18 encodes an orexin-like GPCR that can be activated by FLP-2-derived peptides in transfected mammalian cells (Larsen et al. 2013; Mertens et al. 2005), and frpr-18 functions downstream of flp-2 in the locomotion arousal circuit (Chen et al. 2016). frpr-18 is expressed broadly in the nervous system including in AIY (Chen et al. 2016), and loss-of-function frpr-18 mutations lead to hypersensitivity to certain oxidants (Ouaakki et al. 2023). FRPR-18 is coupled to the heterotrimeric G protein Gaq (Larsen et al. 2013; Mertens et al. 2005), raising the possibility that FLP-2 may promote FLP-1 secretion from AIY by directly activating FRPR-18 in AIY. However, flp-2 functions independently of frpr-18 in the reproductive growth circuit, and instead functions in a genetic pathway with the GPCR npr-30 (Chai et al. 2022). In addition, FLP-2-derived peptides (of which there are at least three) can bind to the GPCRs DMSR-1, or FRPR-8 in transfected cells (Beets et al. 2023). Identifying the relevant FLP-2 peptide(s), the FLP2 receptor and its site of action will help to define the circuit used by intestinal flp-2 to promote FLP-1 release from AIY.
FLP-1 release from AIY is positively regulated by H2O2 generated from mitochondria (Jia and Sieburth 2021). Here we showed that H2O2-induced FLP-1 release requires intestinal flp-2 signaling. However, flp-2 does not appear to promote FLP-1 secretion by increasing H2O2 levels in AIY (Fig 1E), and flp-2 signaling is not sufficient to promote FLP-1 secretion in the absence of H2O2 (Fig. 1D). These results point to a model whereby at least two conditions must be met in order for AIY to increase FLP-1 secretion: an increase in H2O2 levels in AIY itself, and an increase in flp-2 signaling from the intestine. Thus AIY integrates stress signals from both the nervous system and the intestine to activate the intestinal antioxidant response through FLP-1 secretion. The requirement of signals from multiple tissues for FLP-1 secretion may function to limit the activation of SKN-1, since unregulated SKN-1 activation can be detrimental to organismal health (Turner, Ramos, and Curran 2024). AIY shows a sporadic Ca2+ response regardless of the presence of explicit stimulation (Ashida, Hotta, and Oka 2019; Bargmann et al. 2007; Clark et al. 2006), and FLP-1 secretion from AIY is calcium dependent (Jia and Sieburth 2021)How mitochondrial H2O2 levels are established in AIY by intrinsic or extracellular inputs, and how AIY integrates H2O2 and flp-2 signaling to control FLP-1 secretion remain to be defined.
A role for endogenous H2O2 in regulated neuropeptide secretion.
Using HyPer7, we showed that acute juglone exposure results in a rapid elevation of endogenous H2O2 levels inside and outside intestinal mitochondria and a corresponding increase of FLP-2 release from the intestine that depends on the cytoplasmic superoxide dismutase sod-1, and mitochondrial sod-3. We favor a model whereby superoxide generated by juglone in the mitochondria is converted to H2O2 by SOD-3 in the matrix and by SOD-1 in the cytosol. In this case, both the superoxide generated by juglone and the H2O2 generated by SOD-3 would have to be able to exit the mitochondria and enter the cytosol. Superoxide and H2O2 can be transported across mitochondrial membranes through anion channels and aquaporin channels, respectively (Bienert and Chaumont 2014; Ferri et al. 2003; Han et al. 2003; Kontos et al. 1985). The observation that both SOD-1 and SOD-3 activity are necessary to drive FLP-2 release suggests that H2O2 levels much reach a certain threshold in the cytoplasm to promote FLP-2 release, and this threshold requires the generation of H2O2 by both SOD-1 and SOD-3.
We identified a role for the antioxidant peroxiredoxin-thioredoxin system, encoded by prdx-2 and trx-3, in maintaining low endogenous H2O2 levels in the intestine and in negatively regulating FLP-2 secretion. We showed that the prdx-2b isoform functions to inhibit FLP-2 secretion and to lower H2O2 levels in both the mitochondrial matrix and on the cytosolic side of mitochondria. These observations are consistent with a subcellular site of action for PRDX-2B in either the matrix only or in both the matrix and cytosol. In contrast, trx-3 mutations do not alter mitochondrial H2O2 levels, suggesting that TRX-3 functions exclusively in the cytosol. Thus, the PRDX-2B-TRX-3 combination may function in the cytosol, and PRDX-2B may function with a different TRX family member in the matrix. There are several thioredoxin-domain containing proteins in addition to trx-3 in the C. elegans genome (including trx-5/NXNL2) that could be candidates for this role. Alternatively, prdx-2 may function alone or with other redox proteins. PRDX-2 may function without thioredoxins in its roles in light sensing and stress response in worms (Li et al. 2016; Oláhová et al. 2008; Oláhová and Veal 2015). PRDX-2B contains a unique N terminal domain that is distinct from the catalytic domain and is not found on the other PRDX-2 isoforms. This domain may be important for targeting PRDX-2B to specific subcellular location(s) where it can regulate FLP-2 secretion.
Regulation of FLP-2 exocytosis by PKC-2/PKCα/β and AEX-4/SNAP25.
We demonstrated that pkc-2 mediates the effects of H2O2 on intestinal FLP-2 secretion, and H2O2- and DAG-mediated PKC-2 activation are likely to function in a common genetic pathway to promote FLP-2 secretion. Our observations that DAG is required for the effects of juglone (Fig. 6B), are consistent with a two-step activation model for PKC-2, in which H2O2 could first modify PKC-2 in the cytosol, facilitating subsequent PKC-2 recruitment to the membrane by DAG. Alternatively, DAG could first recruit PKC2 to membranes, where it is then modified by H2O2. We favor a model whereby H2O2 modification occurs in the cytosol, since H2O2 produced locally by mitochondria would have access to cytosolic pools of PKC-2 prior to its membrane translocation.
We defined a role for aex-4/SNAP25 in the fusion step of FLP-2 containing DCVs from the intestine under normal conditions as well as during oxidative stress. In neuroendocrine cells, phosphorylation of SNAP25 on Ser187 potentiates DCV recruitment into releasable pools (Nagy et al. 2002; Shu et al. 2008; Yang et al. 2007), and exocytosis stimulated by the DAG analog phorbol ester (Gao et al. 2016; Shu et al. 2008), without altering baseline SNAP25 function. Interestingly, the residue corresponding to Ser187 is conserved in AEX-4, raising the possibility that PKC-2 potentiates FLP-2 secretion by phosphorylating AEX-4. Since SNAP25 phosphorylation on Ser187 has been shown to increase its interaction with syntaxin and promote SNARE complex assembly in vitro (Gao et al. 2016; Yang et al. 2007), it is possible that elevated H2O2 levels could promote FLP-2 secretion by positively regulating SNARE-mediated DCV fusion at intestinal release sites on the basolateral membrane through AEX4/SNAP25 phosphorylation by PKC-2. Prior studies have shown that PKC-2 phosphorylates the SNARE-associated protein UNC-18 in neurons to regulate thermosensory behavior (Edwards et al. 2012; Land and Rubin 2017). Thus, PKC-2 may have multiple targets in vivo and target selection may be dictated by cell type and/or the redox status of the cell.
Similar molecular mechanisms regulating FLP-1 and FLP-2 release.
The molecular mechanisms we identified that regulate FLP-2 secretion from the intestine are similar in several respects to those regulating FLP-1 secretion from AIY. First, the secretion of both peptides is positively regulated by H2O2 originating from mitochondria. Second, in both cases, H2O2 promotes exocytosis of neuropeptidecontaining DCVs by a mechanism that depends upon the kinase activity of protein kinase C. Finally, the secretion of both peptides is controlled through the regulation of H2O2 levels by superoxide dismutases and by the peroxiredoxin-thioredoxin system. H2O2-regulated FLP-1 and FLP-2 secretion differ in the identity of the family members of some of the genes involved. prdx-3-trx-2 and sod-2 family members regulate H2O2 levels in AIY, whereas prdx-2-trx-3 and sod-1/sod-3 family members regulate H2O2 levels in the intestine. In addition, pkc-1 promotes H2O2 induced FLP-1 secretion from AIY whereas pkc-2 promotes H2O2 -induced FLP-2 secretion from the intestine. Nonetheless, it is noteworthy that two different cell types utilize largely similar pathways for the H2O2mediated regulation of neuropeptide release, raising the possibility that similar mechanisms may be utilized in other cell types and/or organisms to regulate DCV secretion in response to oxidative stress.
Materials and Methods
Strains and transgenic lines
C. elegans strains were maintained at 20°C in the dark on standard nematode growth medium (NGM) plates seeded with OP50 Escherichia coli as food source, unless otherwise indicated. All strains were synchronized by picking mid L4 stage animal either immediately before treatment (for coelomocyte imaging and intestine imaging) or 24h before treatment (for Pgst-4::gfp imaging). The wild type strain was Bristol N2. Mutants used in this study were out-crossed at least four times.
Transgenic lines were generated by microinjecting plasmid mixes into the gonads of young adult animals following standard techniques (Mello et al. 1991). Microinjection mixes were prepared by mixing expression constructs with the co-injection markers pJQ70 (Pofm-1::rfp, 25ng/μL), pMH163 (Podr-1::mCherry, 40ng/μL), pMH164 (Podr1::gfp, 40ng/μL) or pDS806 (Pmyo-3::mCherry, 20ng/μL) to a final concentration of 100ng/μL. For tissue-specific expression, a 1.5kb rab-3 promoter was used for panneuronal expression (Nonet et al. 1997), a 2.0kb ges-1 or a 3.5kb nlp-40 promoter was used for intestinal expression (Egan et al. 1995; Wang et al. 2013). At least three transgenic lines were examined for each transgene, and one representative line was used for quantification. Strains and transgenic lines used in this study are listed in the Supplementary Table.
Molecular Biology
All gene expression vectors were constructed with the backbone of pPD49.26. Promoter fragments including Prab-3, and Pges-1 were amplified from genomic DNA; genes of interest, including cDNA fragments (aex-5, snt-5, sod-1b, sod-3, isp-1, prdx-2a, prdx-2b, prdx-2c, trx-3, pkc-2b, dgk-2a, aex-4) and genomic fragments (flp-2, flp-40, nlp36, nlp-27) were amplified from cDNA library and genomic DNA respectively using standard molecular biology protocols. Expression plasmid of HyPer7 was designed based on reported mammalian expression plasmid for HyPer7 (Pak et al. 2020) and was synthesized by Thermo Fisher Scientific with codon optimization for gene expression in C. elegans. Plasmids and primers used in this study are listed in the Supplementary Table.
Toxicity Assay
A stock solution of 50mM juglone in DMSO was freshly made on the same day of liquid toxicity assay. 120μM working solution of juglone in M9 buffer was prepared using stock solution before treatment. Around 60–80 synchronized adult animals were transferred into a 1.5mL Eppendorf tube with fresh M9 buffer and washed three times, and a final wash was done with either the working solution of juglone with or M9 DMSO at the concentrations present in juglone-treated animals does not contribute to toxicity since DMSO treatment alone caused no significant change in survival compared to M9treated controls (Fig. S1C). Animals were incubated in the dark for 4h on rotating mixer before being transferred onto fresh NGM plates seeded with OP50 to recover in the dark at 20°C for 16 hours. Percentage of survival was assayed by counting the number of alive and dead animals. Toxicity assays were performed in triplicates.
RNAi Interference
Plates for feeding RNAi interference were prepare as described (Kamath 2003). Around 20–25 gravid adult animals with indicated genotype were transferred onto the RNAi plates that were seeded with HT115(DE3) bacteria transformed with L4440 vectors with targeted gene inserts or empty L4440 vectors. Eggs were collected for 4h to obtain synchronized populations. L4 stage animals were collected for further assays. RNAi clones were from Ahringer or Vidal RNAi library, or made from genomic DNA. Details were listed in the Supplementary Table.
Behavioral assays
The defecation motor program was assayed as previously described (Liu and Thomas 1994). Twenty to thirty L4 animals were transferred onto a fresh NGM plate seeded with OP50 E. coli and were stored in a 20°C incubator for 24 hours. After 24 hours, ten consecutive defecation cycles were observed from three independent animals and the mean and the standard error was calculated for each genotype. The pBoc and aBoc steps were recorded using custom Etho software (James Thomas Lab website:http://depts.washington.edu/jtlab/software/otherSoftware.html)
Microscopy and Fluorescence Imaging
Approximately 30–40 age matched animals were paralyzed with 30mg/mL 2,3butanedione monoxime (BDM) in M9 buffer and mounted on 2% agarose pads. Images were captured using the Nikon eclipse 90i microscope equipped with Nikon Plan Apo 20x, 40x, 60x, and 100x oil objective (N.A.=1.40), and a Photometrics Coolsnap ES2 camera or a Hamamatsu Orca Flash LT+CMOS camera. Metamorph 7.0 software (Universal Imaging/Molecular Devices) was used to capture serial image stacks and to obtain the maximum intensity projection image for analysis.
For transcriptional reporter imaging, young adult animals were transferred into a 1.5mL Eppendorf tube with M9 buffer, washed three times and incubated in 50μM working solution of juglone or M9 buffer control with equivalent DMSO for 1h in the dark on rotating mixer before recovering on fresh NGM plates with OP50 for 3h in the dark at 20°C. The posterior end of the intestine was imaged with the 60x objective and quantification for average fluorescence intensity of a 16-pixel diameter circle in the posterior intestine was calculated using Metamorph.
For coelomocyte imaging, L4 stage animals were transferred in fresh M9 buffer on a cover slide, washed six times with M9 before being exposed to 300μM juglone in M9 buffer (diluted from freshly made 50mM stock solution), 1mM H2O2 in M9 buffer, or M9 buffer. DMSO at the concentrations present in juglone-treated animals does not alter neuropeptide secretion since DMSO treatment alone caused no significant change in FLP-1::Venus or FLP-2::Venus coelomocyte fluorescence compared to M9-treated controls. (Fig. S1D and S2E). . Animals were then paralyzed in BDM and images of coelomocytes next to the posterior end of intestine were taken using the 100x oil objective. Average fluorescence intensity of Venus from the endocytic compartments in the posterior coelomocytes was measured in ImageJ.
For fusion protein fluorescence imaging, L4 stage animals were exposed to M9 buffer or indicated oxidants for 10min before being paralyzed in BDM and images taken of the posterior end of the intestine using 100x oil objective. For HyPer7 imaging, Z stacks were obtained using GFP (excitation/emission: 500nm/520nm) and CFP (excitation/emission: 400nm/520nm) filter sets sequentially, HyPer7 fluorescence signal was quantified as the ratio of GFP to CFP fluorescence intensity changes with respect to the baseline [(Ft − F0)/F0].
CRISPR/Cas9 Editing
prdx-2b(vj380) knock-out mutants were generated using a co-CRISPR protocol (Arribere et al. 2014). A sgRNA and a repair single stranded oligodeoxynucleotides (ssODN) targeting dpy-10 were co-injected with a sgRNA for genes of interest and a ssODN that induces homology-directed repair (HDR) to introduce Cas9 mediated mutagenesis. Fifteen young adult animals were injected to produce around thirty singled F1 animals carrying Dpy or Rol phenotype. F2 animals were genotyped for mutations based on PCR and enzyme digest. Homozygous mutants were outcrossed with wild type animals at least four times before being used for assays.
Statistics
Statistical analysis was performed on GraphPad Prism 9. Unpaired t-test with two tails was used for two groups and one-way ANOVA with multiple comparison corrections was used to three or more groups to determine the statistical significance. Statistical details and n are specified in the figure legends. All comparisons are conducted based on wild-type controls unless indicated by lines between genotypes. Bar graphs with plots were generated using GraphPad Prism 9.
Supplementary Material
Acknowledgements
C. elegans strains used in this study were provided by the Caenorhabditis Genetics Centre (CGC), which is funded by the NIH National Center for Research Resources (NCRR). We thank members of the Sieburth lab for critical reading and discussion of the manuscript. This work was supported by grants from National Institute of Health NINDS R01NS071085 and R01NS110730 to D.S.
Reference
- Giniatullin A R and Giniatullin R A. 2003. “Dual Action of Hydrogen Peroxide on Synaptic Transmission at the Frog Neuromuscular Junction.” The Journal of Physiology 552(1):283–93. doi: 10.1113/jphysiol.2003.050690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ahier Arnaud, Dai Chuan-Yang, Tweedie Andrea, Bezawork-Geleta Ayenachew, Kirmes Ina, and Zuryn Steven. 2018. “Affinity Purification of Cell-Specific Mitochondria from Whole Animals Resolves Patterns of Genetic Mosaicism.” Nature Cell Biology 20(3):352-. doi: 10.1038/s41556-017-0023-x. [DOI] [PubMed] [Google Scholar]
- Ahmad Taseer, and Suzuki Yuichiro J.. 2019. “Juglone in Oxidative Stress and Cell Signaling.” Antioxidants (Basel, Switzerland) 8(4):91. doi: 10.3390/antiox8040091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ailion Michael, Hannemann Mandy, Dalton Susan, Pappas Andrea, Watanabe Shigeki, Hegermann Jan, Liu Qiang, Han Hsiao-Fen, Gu Mingyu, Goulding Morgan Q., Sasidharan Nikhil, Schuske Kim, Hullett Patrick, Eimer Stefan, and Jorgensen Erik M.. 2014. “Two Rab2 Interactors Regulate Dense-Core Vesicle Maturation.” Neuron 82(1):167–80. doi: 10.1016/j.neuron.2014.02.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arribere Joshua A., Bell Ryan T., Fu Becky XH, Artiles Karen L., Hartman Phil S., and Fire Andrew Z.. 2014. “Efficient Marker-Free Recovery of Custom Genetic Modifications with CRISPR/Cas9 in Caenorhabditis Elegans.” [DOI] [PMC free article] [PubMed]
- Avshalumov Marat V., Chen Billy T., Koos Tibor, Tepper James M., and Rice Margaret E.. 2005. “Endogenous Hydrogen Peroxide Regulates the Excitability of Midbrain Dopamine Neurons via ATPSensitive Potassium Channels.” Journal of Neuroscience 25(17):4222–31. doi: 10.1523/JNEUROSCI.4701-04.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avshalumov Marat V., and Rice Margaret E.. 2003. “Activation of ATP-Sensitive K+ (KATP) Channels by H2O2 Underlies Glutamate-Dependent Inhibition of Striatal Dopamine Release.” Proceedings of the National Academy of Sciences - PNAS 100(20):11729–34. doi: 10.1073/pnas.1834314100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Back Patricia, De Vos Winnok H., Depuydt Geert G., Matthijssens Filip, Vanfleteren Jacques R., and Braeckman Bart P.. 2012. “Exploring Real-Time in Vivo Redox Biology of Developing and Aging Caenorhabditis Elegans.” Free Radical Biology and Medicine 52(5):850–59. doi: 10.1016/j.freeradbiomed.2011.11.037. [DOI] [PubMed] [Google Scholar]
- Bao Li, Avshalumov Marat V., and Rice Margaret E.. 2005. “Partial Mitochondrial Inhibition Causes Striatal Dopamine Release Suppression and Medium Spiny Neuron Depolarization via H2O2 Elevation, Not ATP Depletion.” The Journal of Neuroscience 25(43):10029–40. doi: 10.1523/JNEUROSCI.2652-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bargmann Cornelia I., Chalasani Sreekanth H., Chronis Nikos, Tsunozaki Makoto, Gray Jesse M., Ramot Daniel, and Goodman Miriam B.. 2007. “Dissecting a Circuit for Olfactory Behaviour in Caenorhabditis Elegans.” Nature 450(7166):63–70. doi: 10.1038/nature06292. [DOI] [PubMed] [Google Scholar]
- Batterham Rachel L., and Bloom Stephen R.. 2003. “The Gut Hormone Peptide YY Regulates Appetite.” Annals of the New York Academy of Sciences 994(1):162–68. doi: 10.1111/j.1749-6632.2003.tb03176.x. [DOI] [PubMed] [Google Scholar]
- Batterham Rachel L., Cowley Michael A., Small Caroline J., Herzog Herbert, Cohen Mark A., Dakin Catherine L., Wren Alison M., Brynes Audrey E., Low Malcolm J., Ghatei Mohammad A., Cone Roger D., and Bloom Stephen R.. 2002. “Gut Hormone PYY(3–36) Physiologically Inhibits Food Intake.” Nature (London) 418(6898):650-. [DOI] [PubMed] [Google Scholar]
- Beets Isabel, Zels Sven, Vandewyer Elke, Demeulemeester Jonas, Caers Jelle, Baytemur Esra, Courtney Amy, Golinelli Luca, Hasakioğulları İlayda, Schafer William R., Vértes Petra E., Mirabeau Olivier, and Schoofs Liliane. 2023. “System-Wide Mapping of Peptide-GPCR Interactions in C. Elegans.” Cell Reports (Cambridge) 42(9):113058–113058. doi: 10.1016/j.celrep.2023.113058. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bienert Gerd P., and Chaumont François. 2014. “Aquaporin-Facilitated Transmembrane Diffusion of Hydrogen Peroxide.” Biochimica et Biophysica Acta 1840(5):1596–1604. doi: 10.1016/j.bbagen.2013.09.017. [DOI] [PubMed] [Google Scholar]
- van Blitterswijk Wim J., and Houssa Brahim. 2000. “Properties and Functions of Diacylglycerol Kinases.” Cellular Signalling 12(9):595–605. doi: 10.1016/S0898-6568(00)00113-3. [DOI] [PubMed] [Google Scholar]
- Botteaux A., Hoste C., Dumont J. E., Van Sande J., and Allaoui A.. 2009. “Potential Role of Noxes in the Protection of Mucosae: H(2)O(2) as a Bacterial Repellent.” Microbes and Infection 11(5):537–44. doi: 10.1016/j.micinf.2009.02.009. [DOI] [PubMed] [Google Scholar]
- Browning Kirsteen N., and Travagli R. Alberto. 2014. “Central Nervous System Control of Gastrointestinal Motility and Secretion and Modulation of Gastrointestinal Functions.” Comprehensive Physiology 4(4):1339–68. doi: 10.1002/cphy.c130055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns D. J., and Bell R. M.. 1991. “Protein Kinase C Contains Two Phorbol Ester Binding Domains.” The Journal of Biological Chemistry 266(27):18330–38. doi: 10.1016/S0021-9258(18)55274-1. [DOI] [PubMed] [Google Scholar]
- Campos-Salinas Jenny, Cavazzuti Antonio, O’Valle Francisco, Forte-Lago Irene, Caro Marta, Beverley Stephen M., Delgado Mario, and Gonzalez-Rey Elena. 2014. “Therapeutic Efficacy of Stable Analogues of Vasoactive Intestinal Peptide against Pathogens.” The Journal of Biological Chemistry 289(21):14583–99. doi: 10.1074/jbc.M114.560573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carabotti Marilia, Scirocco Annunziata, Maselli Maria Antonietta, and Severi Carola. 2015. “The Gut-Brain Axis: Interactions between Enteric Microbiota, Central and Enteric Nervous Systems.” Annals of Gastroenterology : Quarterly Publication of the Hellenic Society of Gastroenterology 28(2):203–9. [PMC free article] [PubMed] [Google Scholar]
- Castello Pablo R., Drechsel Derek A., and Patel Manisha. 2007. “Mitochondria Are a Major Source of Paraquat-Induced Reactive Oxygen Species Production in the Brain.” The Journal of Biological Chemistry 282(19):14186–93. doi: 10.1074/jbc.M700827200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Castro Edouard, de Castro Sarah Hegi, and Johnson Thomas E.. 2004. “Isolation of Long-Lived Mutants in Caenorhabditis Elegans Using Selection for Resistance to Juglone.” Free Radical Biology & Medicine 37(2):139–45. doi: 10.1016/j.freeradbiomed.2004.04.021. [DOI] [PubMed] [Google Scholar]
- Chai Cynthia M., Torkashvand Mahdi, Seyedolmohadesin Maedeh, Park Heenam, Venkatachalam Vivek, and Sternberg Paul W.. 2022. “Interneuron Control of C. Elegans Developmental Decision-Making.” Current Biology 32(10):2316–2324.e4. doi: 10.1016/j.cub.2022.03.077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chance B., Sies H., and Boveris A.. 1979. “Hydroperoxide Metabolism in Mammalian Organs.” Physiological Reviews 59(3):527–605. doi: 10.1152/physrev.1979.59.3.527. [DOI] [PubMed] [Google Scholar]
- Chelikani Prasanth K., Haver Alvin C., and Reidelberger Roger D.. 2005. “Intravenous Infusion of Peptide YY(3–36) Potently Inhibits Food Intake in Rats.” Endocrinology (Philadelphia) 146(2):879–88. doi: 10.1210/en.2004-1138. [DOI] [PubMed] [Google Scholar]
- Chen Billy T., Avshalumov Marat V., and Rice Margaret E.. 2001. “H2O2 Is a Novel, Endogenous Modulator of Synaptic Dopamine Release.” Journal of Neurophysiology 85(6):2468–76. doi: 10.1152/jn.2001.85.6.2468. [DOI] [PubMed] [Google Scholar]
- Chen Billy T., Avshalumov Marat V., and Rice Margaret E.. 2002. “Modulation of Somatodendritic Dopamine Release by Endogenous H2O2: Susceptibility in Substantia Nigra But Resistance in VTA.” Journal of Neurophysiology 87(2):1155–58. doi: 10.1152/jn.00629.2001. [DOI] [PubMed] [Google Scholar]
- Chen Didi, Taylor Kelsey P., Hall Qi, and Kaplan Joshua M.. 2016. “The Neuropeptides FLP-2 and PDF-1 Act in Concert To Arouse Caenorhabditis Elegans Locomotion.” Genetics (Austin) 204(3):1151–59. doi: 10.1534/genetics.116.192898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Yu A., and Scheller Richard H.. 2001. “SNARE-Mediated Membrane Fusion.” Nature Reviews. Molecular Cell Biology 2(2):98–106. doi: 10.1038/35052017. [DOI] [PubMed] [Google Scholar]
- Ch’ng Queelim, Sieburth Derek, and Kaplan Joshua M.. 2008. “Profiling Synaptic Proteins Identifies Regulators of Insulin Secretion and Lifespan.” PLoS Genetics 4(11):e1000283. doi: 10.1371/journal.pgen.1000283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark Damon A., Biron David, Sengupta Piali, and Samuel Aravinthan D. T.. 2006. “The AFD Sensory Neurons Encode Multiple Functions Underlying Thermotactic Behavior in Caenorhabditis Elegans.” The Journal of Neuroscience 26(28):7444–51. doi: 10.1523/JNEUROSCI.1137-06.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corcionivoschi Nicolae, Alvarez Luis AJ, Sharp Thomas H., Strengert Monika, Alemka Abofu, Mantell Judith, Verkade Paul, Knaus Ulla G., and Bourke Billy. 2012. “Mucosal Reactive Oxygen Species Decrease Virulence by Disrupting Campylobacter Jejuni Phosphotyrosine Signaling.” Cell Host & Microbe 12(1):47–59. doi: 10.1016/j.chom.2012.05.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darby Lisa M., Meng Hongdi, and Fehrenbacher Jill C.. 2017. “Paclitaxel Inhibits the Activity and Membrane Localization of PKCα and PKCβI/II to Elicit a Decrease in Stimulated Calcitonin Gene-Related Peptide Release from Cultured Sensory Neurons.” Molecular and Cellular Neurosciences 82:105–17. doi: 10.1016/j.mcn.2017.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingley Stephen, Polyak Erzsebet, Lightfoot Richard, Ostrovsky Julian, Rao Meera, Greco Todd, Ischiropoulos Harry, and Falk Marni J.. 2010. “Mitochondrial Respiratory Chain Dysfunction Variably Increases Oxidant Stress in Caenorhabditis Elegans.” Mitochondrion 10(2):125–36. doi: 10.1016/j.mito.2009.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards Mark R., Johnson James R., Rankin Kimberley, Jenkins Rosalind E., Maguire Carl, Morgan Alan, Burgoyne Robert D., and Barclay Jeff W.. 2012. “PKC-2 Phosphorylation of UNC-18 Ser322 in AFD Neurons Regulates Temperature Dependency of Locomotion.” The Journal of Neuroscience 32(20):7042–51. doi: 10.1523/JNEUROSCI.4029-11.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards Samantha Louise, Mergan Lucas, Parmar Bhavesh, Cockx Bram, Wouter De Haes Liesbet Temmerman, and Schoofs Liliane. 2019. “Exploring Neuropeptide Signalling through Proteomics and Peptidomics.” Expert Review of Proteomics 16(2):131–37. doi: 10.1080/14789450.2019.1559733. [DOI] [PubMed] [Google Scholar]
- Egan Cay R., Chung May A., Allen Fran L., Heschl Mark FP, Van Buskirk Cheryl L., and McGhee James D.. 1995. “A Gut-to-Pharynx/Tail Switch in Embryonic Expression of the Caenorhabditis Elegans Ges-1 Gene Centers on Two GATA Sequences.” Developmental Biology 170(2):397–419. doi: 10.1006/dbio.1995.1225. [DOI] [PubMed] [Google Scholar]
- Elferink Jan GR. 1999. “Thimerosal A Versatile Sulfhydryl Reagent, Calcium Mobilizer, and Cell FunctionModulating Agent.” General Pharmacology 33(1):1–6. doi: 10.1016/S0306-3623(98)00258-4. [DOI] [PubMed] [Google Scholar]
- Fares Hanna, and Greenwald Iva. 2001. “Genetic Analysis of Endocytosis in Caenorhabditis Elegans: Coelomocyte Uptake Defective Mutants.” Genetics (Austin) 159(1):133–45. doi: 10.1093/genetics/159.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferri Domenico, Mazzone Amelia, Liquori Giuseppa Esterina, Cassano Grazia, Svelto Maria, and Calamita Giuseppe. 2003. “Ontogeny, Distribution, and Possible Functional Implications of an Unusual Aquaporin, AQP8, in Mouse Liver.” Hepatology (Baltimore, Md.) 38(4):947–57. doi: 10.1053/jhep.2003.50397. [DOI] [PubMed] [Google Scholar]
- Fridovich Irwin. 1995. “Superoxide Radical and Superoxide Dismutases.” Annual Review of Biochemistry 64(1):97–112. doi: 10.1146/annurev.bi.64.070195.000525. [DOI] [PubMed] [Google Scholar]
- Fridovich Irwin. 1997. “Superoxide Anion Radical (O·̄2), Superoxide Dismutases, and Related Matters.” The Journal of Biological Chemistry 272(30):18515–17. doi: 10.1074/jbc.272.30.18515. [DOI] [PubMed] [Google Scholar]
- Fukui Masayuki, and Zhu Bao Ting. 2010. “Mitochondrial Superoxide Dismutase SOD2, but Not Cytosolic SOD1, Plays a Critical Role in Protection against Glutamate-Induced Oxidative Stress and Cell Death in HT22 Neuronal Cells.” Free Radical Biology and Medicine 48(6):821–30. doi: 10.1016/j.freeradbiomed.2009.12.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furness John B., Callaghan Brid P., Rivera Leni R., and Cho Hyun-Jung. 2014. “The Enteric Nervous System and Gastrointestinal Innervation: Integrated Local and Central Control.” Advances in Experimental Medicine and Biology 817:39–71. doi: 10.1007/978-1-4939-0897-4_3. [DOI] [PubMed] [Google Scholar]
- Gao Jing, Hirata Makiko, Mizokami Akiko, Zhao Jin, Takahashi Ichiro, Takeuchi Hiroshi, and Hirata Masato. 2016. “Differential Role of SNAP-25 Phosphorylation by Protein Kinases A and C in the Regulation of SNARE Complex Formation and Exocytosis in PC12 Cells.” Cellular Signalling 28(5):425–37. doi: 10.1016/j.cellsig.2015.12.014. [DOI] [PubMed] [Google Scholar]
- Santamarina García, Sarela Susanna Boronat i Llop, and Hernando Elena Hidalgo. 2014. “Reversible Cysteine Oxidation in Hydrogen Peroxide Sensing and Signal Transduction.” doi: 10.1021/bi401700f. [DOI] [PubMed]
- Gibbs James, Young Robert C., and Smith Gerade P.. 1973. “Cholecystokinin Elicits Satiety in Rats with Open Gastric Fistulas.” Nature (London) 245(5424):323–25. doi: 10.1038/245323a0. [DOI] [PubMed] [Google Scholar]
- Giglio A. M., Hunter T., Bannister J. V., Bannister W. H., and Hunter G. J.. 1994. “The Copper/Zinc Superoxide Dismutase Gene of Caenorhabditis Elegans.” Biochemistry and Molecular Biology International 33(1):41–44. [PubMed] [Google Scholar]
- Giniatullin Arthur, Petrov Alexey, and Giniatullin Rashid. 2019. “Action of Hydrogen Peroxide on Synaptic Transmission at the Mouse Neuromuscular Junction.” Neuroscience 399:135–45. doi: 10.1016/j.neuroscience.2018.12.027. [DOI] [PubMed] [Google Scholar]
- Goda Yukiko. 1997. “SNAREs and Regulated Vesicle Exocytosis.” Proceedings of the National Academy of Sciences - PNAS 94(3):769–72. doi: 10.1073/pnas.94.3.769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grenham Sue, Clarke Gerard, Cryan John F., and Dinan Timothy G.. 2011. “Brain-Gut-Microbe Communication in Health and Disease.” Frontiers in Physiology 2:94–94. doi: 10.3389/fphys.2011.00094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haber Adam L., Biton Moshe, Rogel Noga, Herbst Rebecca H., Shekhar Karthik, Smillie Christopher, Burgin Grace, Delorey Toni M., Howitt Michael R., Katz Yarden, Tirosh Itay, Beyaz Semir, Dionne Danielle, Zhang Mei, Raychowdhury Raktima, Garrett Wendy S., Rozenblatt-Rosen Orit, Shi Hai Ning, Yilmaz Omer, Xavier Ramnik J., and Regev Aviv. 2017. “A Single-Cell Survey of the Small Intestinal Epithelium.” Nature (London) 551(7680):333–39. doi: 10.1038/nature24489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han Derick, Antunes Fernando, Canali Raffaella, Rettori Daniel, and Cadenas Enrique. 2003. “VoltageDependent Anion Channels Control the Release of the Superoxide Anion from Mitochondria to Cytosol.” The Journal of Biological Chemistry 278(8):5557–63. doi: 10.1074/jbc.M210269200. [DOI] [PubMed] [Google Scholar]
- Han Wenfei, Tellez Luis A., Perkins Matthew H., Perez Isaac O., Qu Taoran, Ferreira Jozelia, Ferreira Tatiana L., Quinn Daniele, Liu Zhong-Wu, Gao Xiao-Bing, Kaelberer Melanie M., Bohórquez Diego V., Shammah-Lagnado Sara J., de Lartigue Guillaume, and de Araujo Ivan E.. 2018. “A Neural Circuit for GutInduced Reward.” Cell 175(3):665–678.e23. doi: 10.1016/j.cell.2018.08.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hourihan John M., Moronetti Mazzeo Lorenza E., Fernández-Cárdenas L. Paulette, and Blackwell T. Keith. 2016. “Cysteine Sulfenylation Directs IRE-1 to Activate the SKN-1/Nrf2 Antioxidant Response.” Molecular Cell 63(4):553–66. doi: 10.1016/j.molcel.2016.07.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Islas-Trejo A., Land M., Tcherepanova I., Freedman J. H., and Rubin C. S.. 1997. “Structure and Expression of the Caenorhabditis Elegans Protein Kinase C2 Gene. Origins and Regulated Expression of a Family of Ca2+-Activated Protein Kinase C Isoforms.” The Journal of Biological Chemistry 272(10):6629–40. doi: 10.1074/jbc.272.10.6629. [DOI] [PubMed] [Google Scholar]
- Iwasaki Kouichi, Staunton Jane, Saifee Owais, Nonet Michael, and Thomas James H.. 1997. “Aex-3 Encodes a Novel Regulator of Presynaptic Activity in C. Elegans.” Neuron (Cambridge, Mass.) 18(4):613– 22. doi: 10.1016/S0896-6273(00)80302-5. [DOI] [PubMed] [Google Scholar]
- Jahn Reinhard, and Scheller Richard H.. 2006. “SNAREs - Engines for Membrane Fusion.” Nature Reviews. Molecular Cell Biology 7(9):631–43. doi: 10.1038/nrm2002. [DOI] [PubMed] [Google Scholar]
- Jia Qi, and Sieburth Derek. 2021. “Mitochondrial Hydrogen Peroxide Positively Regulates Neuropeptide Secretion during Diet-Induced Activation of the Oxidative Stress Response.” Nature Communications 12(1):2304–2304. doi: 10.1038/s41467-021-22561-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiménez-Hidalgo María, Kurz Cyril Léopold, Pedrajas José Rafael, Naranjo-Galindo Francisco José, González-Barrios María, Cabello Juan, Sáez Alberto G., Lozano Encarnación, Button Emma L., Veal Elizabeth A., Fierro-González Juan Carlos, Swoboda Peter, and Miranda-Vizuete Antonio. 2014. “Functional Characterization of Thioredoxin 3 (TRX-3), a Caenorhabditis Elegans Intestine-Specific Thioredoxin.” Free Radical Biology & Medicine 68(100):205–19. doi: 10.1016/j.freeradbiomed.2013.11.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson Joanne E., Giorgione Jennifer, and Newton Alexandra C.. 2000. “The C1 and C2 Domains of Protein Kinase C Are Independent Membrane Targeting Modules, with Specificity for Phosphatidylserine Conferred by the C1 Domain.” Biochemistry (Easton) 39(37):11360–69. doi: 10.1021/bi000902c. [DOI] [PubMed] [Google Scholar]
- Kageyama Natsumi, Nose Masayo, Ono Masahiro, Matsunaga Yohei, Iwasaki Takashi, and Kawano Tsuyoshi. 2022. “The FMRFamide-like Peptide FLP-2 Is Involved in the Modulation of Larval Development and Adult Lifespan by Regulating the Secretion of the Insulin-like Peptide INS-35 in Caenorhabditis Elegans.” Bioscience, Biotechnology, and Biochemistry 86(9):1231–39. doi: 10.1093/bbb/zbac108. [DOI] [PubMed] [Google Scholar]
- Kamata Hideaki, Manabe Tomoyuki, Oka Shin-ichi, Kamata Keiko, and Hirata Hajime. 2002. “Hydrogen Peroxide Activates IκB Kinases through Phosphorylation of Serine Residues in the Activation Loops.” FEBS Letters 519(1):231–37. doi: 10.1016/S0014-5793(02)02712-6. [DOI] [PubMed] [Google Scholar]
- Kamath R. 2003. “Genome-Wide RNAi Screening in Caenorhabditis Elegans.” Methods (San Diego, Calif.) 30(4):313–21. doi: 10.1016/s1046-2023(03)00050-1. [DOI] [PubMed] [Google Scholar]
- Kemble David J., and Sun Gongqin. 2009. “Direct and Specific Inactivation of Protein Tyrosine Kinases in the Src and FGFR Families by Reversible Cysteine Oxidation.” Proceedings of the National Academy of Sciences - PNAS 106(13):5070–75. doi: 10.1073/pnas.0806117106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Ki Mo, Heo Deok Rim, Kim Young-A., Lee Jun, Kim No Soo, and Bang Ok-Sun. 2016. “Coniferaldehyde Inhibits LPS-Induced Apoptosis through the PKC α/β II/Nrf-2/HO-1 Dependent Pathway in RAW264.7 Macrophage Cells.” Environmental Toxicology and Pharmacology 48:85–93. doi: 10.1016/j.etap.2016.10.016. [DOI] [PubMed] [Google Scholar]
- Knoefler Daniela, Thamsen Maike, Koniczek Martin, Niemuth Nicholas J., Diederich Ann-Kristin, and Jakob Ursula. 2012. “Quantitative In Vivo Redox Sensors Uncover Oxidative Stress as an Early Event in Life.” Molecular Cell 47(5):767–76. doi: 10.1016/j.molcel.2012.06.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konishi Hiroaki, Tanaka Motonari, Takemura Yukitoshi, Matsuzaki Hidenori, Ono Yoshitaka, Kikkawa Ushio, and Nishizuka Yasutomi. 1997. “Activation of Protein Kinase C by Tyrosine Phosphorylation in Response to H2O2.” Proceedings of the National Academy of Sciences - PNAS 94(21):11233–37. doi: 10.1073/pnas.94.21.11233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konishi Hiroaki, Yamauchi Emiko, Taniguchi Hisaaki, Yamamoto Toshiyoshi, Matsuzaki Hidenori, Takemura Yukitoshi, Ohmae Kyoko, Kikkawa Ushio, and Nishizuka Yasutomi. 2001. “Phosphorylation Sites of Protein Kinase C δ in H2O2-Treated Cells and Its Activation by Tyrosine Kinase in Vitro.” Proceedings of the National Academy of Sciences - PNAS 98(12):6587–92. doi: 10.1073/pnas.111158798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kontos Hermes A., Wei Enoch P., Ellis Earl F., Jenkins Larry W., Povlishock John T., Rowe G. Thomas, and Hess Michael L.. 1985. “Appearance of Superoxide Anion Radical in Cerebral Extracellular Space during Increased Prostaglandin Synthesis in Cats.” Circulation Research 57(1):142–51. doi: 10.1161/01.RES.57.1.142. [DOI] [PubMed] [Google Scholar]
- Lai N. Y., Mills K., and Chiu I. M.. 2017. “Sensory Neuron Regulation of Gastrointestinal Inflammation and Bacterial Host Defence.” Journal of Internal Medicine 282(1):5–23. doi: 10.1111/joim.12591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Land Marianne, and Rubin Charles S.. 2017. “A Calcium- and Diacylglycerol-Stimulated Protein Kinase C (PKC), Caenorhabditis Elegans PKC-2, Links Thermal Signals to Learned Behavior by Acting in Sensory Neurons and Intestinal Cells.” Molecular and Cellular Biology 37(19). doi: 10.1128/MCB.00192-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen Martha J., Lancheros Elizabeth Ruiz, Williams Tracey, Lowery David E., Geary Timothy G., and Kubiak Teresa M.. 2013. “Functional Expression and Characterization of the C. Elegans G-Protein-Coupled FLP-2 Receptor (T19F4.1) in Mammalian Cells and Yeast.” International Journal for Parasitology -- Drugs and Drug Resistance 3:1–7. doi: 10.1016/j.ijpddr.2012.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee Kiho, and Mylonakis Eleftherios. 2017. “An Intestine-Derived Neuropeptide Controls Avoidance Behavior in Caenorhabditis Elegans.” Cell Reports 20(10):2501–12. doi: 10.1016/j.celrep.2017.08.053. [DOI] [PubMed] [Google Scholar]
- Lee Yoon-Jin, Cho Hae-Nyun, Soh Jae-Won, Jhon Gil Ja, Cho Chul-Koo, Chung Hee-Yong, Bae Sangwoo, Lee Su-Jae, and Lee Yun-Sil. 2003. “Oxidative Stress-Induced Apoptosis Is Mediated by ERK1/2 Phosphorylation.” Experimental Cell Research 291(1):251–66. doi: 10.1016/S0014-4827(03)00391-4. [DOI] [PubMed] [Google Scholar]
- Li Guang, Gong Jianke, Lei Haoyun, Liu Jianfeng, and Xu X. Z. Shawn. 2016. “Promotion of Behavior and Neuronal Function by Reactive Oxygen Species in C. Elegans.” Nature Communications 7(1):13234. doi: 10.1038/ncomms13234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Zongwei, Gu Yong, Wen Ruoxue, Shen Fanxia, Tian Heng-Li, Yang Guo-Yuan, and Zhang Zhijun. 2019. “Lysosome Exocytosis Is Involved in Astrocyte ATP Release after Oxidative Stress Induced by H2O2.” Neuroscience Letters 705:251–58. doi: 10.1016/j.neulet.2019.03.046. [DOI] [PubMed] [Google Scholar]
- Lin-Moore Alexander T., Oyeyemi Motunrayo J., and Hammarlund Marc. 2021. “Rab-27 Acts in an Intestinal Pathway to Inhibit Axon Regeneration in C. Elegans.” PLoS Genetics 17(11):e1009877–e1009877. doi: 10.1371/journal.pgen.1009877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu D. W., and Thomas J. H.. 1994. “Regulation of a Periodic Motor Program in C. Elegans.” The Journal of Neuroscience 14(4):1953–62. doi: 10.1523/jneurosci.14-04-01953.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Junqiang, Zhang Pei, Zheng Zhongfan, Afridi Muhammad Irfan, Zhang Shan, Wan Zhiqing, Zhang Xiumei, Stingelin Lukas, Wang Yirong, and Tu Haijun. 2023. “GABAergic Signaling between Enteric Neurons and Intestinal Smooth Muscle Promotes Innate Immunity and Gut Defense in Caenorhabditis Elegans.” Immunity 56(7):1515–1532.e9. doi: 10.1016/j.immuni.2023.06.004. [DOI] [PubMed] [Google Scholar]
- Lutz T. A., Del Prete E., and Scharrer E.. 1994. “Reduction of Food Intake in Rats by Intraperitoneal Injection of Low Doses of Amylin.” Physiology & Behavior 55(5):891–95. doi: 10.1016/0031-9384(94)90076-0. [DOI] [PubMed] [Google Scholar]
- Lutz T. A., Geary N., Szabady M. M., Del Prete E., and Scharrer E.. 1995. “Amylin Decreases Meal Size in Rats.” Physiology & Behavior 58(6):1197–1202. doi: 10.1016/0031-9384(95)02067-5. [DOI] [PubMed] [Google Scholar]
- Mahoney Timothy R., Liu Qiang, Itoh Takashi, Luo Shuo, Hadwiger Gayla, Vincent Rose, Wang Zhao-Wen, Fukuda Mitsunori, and Nonet Michael L.. 2006. “Regulation of Synaptic Transmission by RAB-3 and RAB-27 in Caenorhabditis Elegans.” Molecular Biology of the Cell 17(6):2617–25. doi: 10.1091/mbc.E05-12-1170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahoney Timothy R., Luo Shuo, Round Elaine K., Brauner Martin, Gottschalk Alexander, Thomas James H., and Nonet Michael L.. 2008. “Intestinal Signaling to GABAergic Neurons Regulates a Rhythmic Behavior in Caenorhabditis Elegans.” Proceedings of the National Academy of Sciences - PNAS 105(42):16350–55. doi: 10.1073/pnas.0803617105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maman Moria, Marques Filipa Carvalhal, Volovik Yuli, Dubnikov Tatyana, Bejerano-Sagie Michal, and Cohen Ehud. 2013. “A Neuronal GPCR Is Critical for the Induction of the Heat Shock Response in the Nematode C. Elegans.” The Journal of Neuroscience 33(14):6102–11. doi: 10.1523/JNEUROSCI.4023-12.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marinho H. Susana, Real Carla, Cyrne Luísa, Soares Helena, and Antunes Fernando. 2014. “Hydrogen Peroxide Sensing, Signaling and Regulation of Transcription Factors.” Redox Biology 2(C):535–62. doi: 10.1016/j.redox.2014.02.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayer Emeran A., Nance Karina, and Chen Shelley. 2022. “The Gut-Brain Axis.” Annual Review of Medicine 73(1):439–53. doi: 10.1146/annurev-med-042320-014032. [DOI] [PubMed] [Google Scholar]
- Mehrian-Shai Ruty, Reichardt Juergen K. V., Harris Curtis C., and Toren Amos. 2019. “The Gut-Brain Axis, Paving the Way to Brain Cancer.” Trends in Cancer 5(4):200–207. doi: 10.1016/j.trecan.2019.02.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mello C. C., Kramer J. M., Stinchcomb D., and Ambros V.. 1991. “Efficient Gene Transfer in C.Elegans: Extrachromosomal Maintenance and Integration of Transforming Sequences.” The EMBO Journal 10(12):3959–70. doi: 10.1002/j.1460-2075.1991.tb04966.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mertens Inge, Meeusen Tom, Janssen Tom, Nachman Ronald, and Schoofs Liliane. 2005. “Molecular Characterization of Two G Protein-Coupled Receptor Splice Variants as FLP2 Receptors in Caenorhabditis Elegans.” Biochemical and Biophysical Research Communications 330(3):967–74. doi: 10.1016/j.bbrc.2005.03.071. [DOI] [PubMed] [Google Scholar]
- Messner Kevin R., and Imlay James A.. 2002. “Mechanism of Superoxide and Hydrogen Peroxide Formation by Fumarate Reductase, Succinate Dehydrogenase, and Aspartate Oxidase.” The Journal of Biological Chemistry 277(45):42563–71. doi: 10.1074/jbc.M204958200. [DOI] [PubMed] [Google Scholar]
- Miller Brittany M., Liou Megan J., Zhang Lillian F., Nguyen Henry, Litvak Yael, Schorr Eva-Magdalena, Jang Kyung Ku, Tiffany Connor R., Butler Brian P., and Bäumler Andreas J.. 2020. “Anaerobic Respiration of NOX1-Derived Hydrogen Peroxide Licenses Bacterial Growth at the Colonic Surface.” Cell Host & Microbe 28(6):789–797.e5. doi: 10.1016/j.chom.2020.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Min Do Sik, Kim Eung-Gook, and Exton John H.. 1998. “Involvement of Tyrosine Phosphorylation and Protein Kinase C in the Activation of Phospholipase D by H2O2 in Swiss 3T3 Fibroblasts.” The Journal of Biological Chemistry 273(45):29986–94. doi: 10.1074/jbc.273.45.29986. [DOI] [PubMed] [Google Scholar]
- Miranda-Vizuete Antonio, Damdimopoulos Anastasios E., and Spyrou Giannis. 2000. “The Mitochondrial Thioredoxin System.” Antioxidants & Redox Signaling 2(4):801–10. doi: 10.1089/ars.2000.2.4-801. [DOI] [PubMed] [Google Scholar]
- Murphy Coleen T., Lee Seung-Jae, and Kenyon Cynthia. 2007. “Tissue Entrainment by Feedback Regulation of Insulin Gene Expression in the Endoderm of Caenorhabditis Elegans.” Proceedings of the National Academy of Sciences - PNAS 104(48):19046–50. doi: 10.1073/pnas.0709613104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagy Gabor, Matti Ulf, Nehring Ralf B., Binz Thomas, Rettig Jens, Neher Erwin, and Sorensen Jakob B.. 2002. “Protein Kinase C-Dependent Phosphorylation of Synaptosome-Associated Protein of 25 kDa at Ser187 Potentiates Vesicle Recruitment.” The Journal of Neuroscience 22(21):9278–86. doi: 10.1523/JNEUROSCI.22-21-09278.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nebigil Canan G. 1997. “Suppression of Phospholipase C β, γ, and δ Families Alters Cell Growth and Phosphatidylinositol 4,5-Bisphosphate Levels.” Biochemistry (Easton) 36(50):15949–58. doi: 10.1021/bi971721m. [DOI] [PubMed] [Google Scholar]
- Nelson Kimberly J., Bolduc Jesalyn A., Wu Hanzhi, Collins John A., Burke Elizabeth A., Reisz Julie A., Klomsiri Chananat, Wood Scott T., Yammani Raghunatha R., Poole Leslie B., Furdui Cristina M., and Loeser Richard F.. 2018. “H2O2 Oxidation of Cysteine Residues in C-Jun N-Terminal Kinase 2 (JNK2) Contributes to Redox Regulation in Human Articular Chondrocytes.” The Journal of Biological Chemistry 293(42):16376–89. doi: 10.1074/jbc.RA118.004613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Netto Luis E. S., and Antunes Fernando. 2016. “The Roles of Peroxiredoxin and Thioredoxin in Hydrogen Peroxide Sensing and in Signal Transduction.” Molecules and Cells 39(1):65–71. doi: 10.14348/molcells.2016.2349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nonet Michael L., Staunton Jane E., Kilgard Michael P., Fergestad Tim, Hartwieg Erika, Horvitz H. Robert, Jorgensen Erik M., and Meyer Barbara J.. 1997. “Caenorhabditis Elegans Rab-3 Mutant Synapses Exhibit Impaired Function and Are Partially Depleted of Vesicles.” The Journal of Neuroscience 17(21):8061–73. doi: 10.1523/jneurosci.17-21-08061.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Donnell Michael P., Chao Pin-Hao, Kammenga Jan E., and Sengupta Piali. 2018. “Rictor/TORC2 Mediates Gut-to-Brain Signaling in the Regulation of Phenotypic Plasticity in C. Elegans.” PLoS Genetics 14(2):e1007213–e1007213. doi: 10.1371/journal.pgen.1007213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oláhová Monika, Taylor Sarah R., Khazaipoul Siavash, Wang Jinling, Morgan Brian A., Matsumoto Kunihiro, Keith Blackwell T., and Elizabeth A. Veal. 2008. “Redox-Sensitive Peroxiredoxin That Is Important for Longevity Has Tissue- and Stress-Specific Roles in Stress Resistance.” Proceedings of the National Academy of Sciences - PNAS 105(50):19839–44. doi: 10.1073/pnas.0805507105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oláhová Monika, and Veal Elizabeth A.. 2015. “A Peroxiredoxin, PRDX-2, Is Required for Insulin Secretion and Insulin/IIS-dependent Regulation of Stress Resistance and Longevity.” Aging Cell 14(4):558–68. doi: 10.1111/acel.12321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliveira-Marques Virgínia, Marinho H. Susana, Cyrne Luísa, and Antunes Fernando. 2009. “Role of Hydrogen Peroxide in NF-κB Activation: From Inducer to Modulator.” Antioxidants & Redox Signaling 11(9):2223–43. doi: 10.1089/ars.2009.2601. [DOI] [PubMed] [Google Scholar]
- Ono Yoshitaka, Fujii Tomoko, Igarashi Koichi, Kuno Takayoshi, Tanaka Chikako, Kikkawa Ushio, and Nishizuka Yasutomi. 1989. “Phorbol Ester Binding to Protein Kinase C Requires a Cysteine-Rich ZincFinger-Like Sequence.” Proceedings of the National Academy of Sciences - PNAS 86(13):4868–71. doi: 10.1073/pnas.86.13.4868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ouaakki Hafsa, Joshi Heetanshi, Rathor Laxmi, and Han Sung Min. 2023. “Frpr-18, a Neuropeptide Receptor, Regulates Organismal Lifespan and Stress Tolerance in C. Elegans.” microPublication Biology 2023. doi: 10.17912/micropub.biology.000840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pak Valeriy V., Ezeriņa Daria, Lyublinskaya Olga G., Pedre Brandán, Tyurin-Kuzmin Pyotr A., Mishina Natalie M., Thauvin Marion, Young David, Wahni Khadija, Gache Santiago Agustín Martínez, Demidovich Alexandra D., Ermakova Yulia G., Maslova Yulia D., Shokhina Arina G., Eroglu Emrah, Bilan Dmitry S., Bogeski Ivan, Michel Thomas, Vriz Sophie, Messens Joris, and Belousov Vsevolod V.. 2020. “Ultrasensitive Genetically Encoded Indicator for Hydrogen Peroxide Identifies Roles for the Oxidant in Cell Migration and Mitochondrial Function.” Cell Metabolism 31(3):642–653.e6. doi: 10.1016/j.cmet.2020.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pang Shanshan, and Curran Sean P.. 2014. “Adaptive Capacity to Bacterial Diet Modulates Aging in C. Elegans.” Cell Metabolism 19(2):221–31. doi: 10.1016/j.cmet.2013.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Papa Luena, Manfredi Giovanni, and Germain Doris. 2014. “SOD1, an Unexpected Novel Target for Cancer Therapy.” Genes & Cancer 5(1–2):15–21. doi: 10.18632/genesandcancer.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulsen Michelle T., and Ljungman Mats. 2005. “The Natural Toxin Juglone Causes Degradation of P53 and Induces Rapid H2AX Phosphorylation and Cell Death in Human Fibroblasts.” Toxicology and Applied Pharmacology 209(1):1–9. doi: 10.1016/j.taap.2005.03.005. [DOI] [PubMed] [Google Scholar]
- Prahlad Veena, Cornelius Tyler, and Morimoto Richard I.. 2008. “Regulation of the Cellular Heat Shock Response in Caenorhabditis Elegans by Thermosensory Neurons.” Science (American Association for the Advancement of Science) 320(5877):811–14. doi: 10.1126/science.1156093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satoh Yohsuke, Sato Hirofumi, Kunitomo Hirofumi, Fei Xianfeng, Hashimoto Koichi, and Iino Yuichi. 2014. “Regulation of Experience-Dependent Bidirectional Chemotaxis by a Neural Circuit Switch in Caenorhabditis Elegans.” The Journal of Neuroscience 34(47):15631–37. doi: 10.1523/JNEUROSCI.1757-14.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shakirzyanova A. V., Malomouzh A. I., Naumenko N. V., and Nikolsky E. E.. 2009. “The Effect of Hydrogen Peroxide on Spontaneous Quantal and Nonquantal Acetylcholine Release from Rat Motor Nerve Endings.” Doklady. Biological Sciences 424(1):18–20. doi: 10.1134/S0012496609010062. [DOI] [PubMed] [Google Scholar]
- Shao Li-Wa, Niu Rong, and Liu Ying. 2016. “Neuropeptide Signals Cell Non-Autonomous Mitochondrial Unfolded Protein Response.” Cell Research 26(11):1182–96. doi: 10.1038/cr.2016.118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharpe Martyn A., Livingston Andrew D., and Baskin David S.. 2012. “Thimerosal-Derived Ethylmercury Is a Mitochondrial Toxin in Human Astrocytes: Possible Role of Fenton Chemistry in the Oxidation and Breakage of mtDNA.” Journal of Toxicology 2012:373678–373678. doi: 10.1155/2012/373678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi Yanjun, Qin Lu, Wu Mengting, Zheng Junyu, Xie Tao, and Shao Zhiyong. 2022. “Gut Neuroendocrine Signaling Regulates Synaptic Assembly in C. Elegans.” EMBO Reports 23(8):e53267-n/a. doi: 10.15252/embr.202153267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shu Yilong, Liu Xin, Yang Yan, Takahashi Masami, and Gillis Kevin D.. 2008. “Phosphorylation of SNAP25 at Ser187 Mediates Enhancement of Exocytosis by a Phorbol Ester in INS-1 Cells.” The Journal of Neuroscience 28(1):21–30. doi: 10.1523/JNEUROSCI.2352-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sieburth Derek, Madison Jon M., and Kaplan Joshua M.. 2006. “PKC-1 Regulates Secretion of Neuropeptides.” Nature Neuroscience 10(1):49–57. doi: 10.1038/nn1810. [DOI] [PubMed] [Google Scholar]
- Singh Jogender, and Aballay Alejandro. 2019. “Intestinal Infection Regulates Behavior and Learning via Neuroendocrine Signaling.” eLife 8. doi: 10.7554/eLife.50033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song Youngmi, Koehler Jacqueline A., Baggio Laurie L., Powers Alvin C., Sandoval Darleen A., and Drucker Daniel J.. 2019. “Gut-Proglucagon-Derived Peptides Are Essential for Regulating Glucose Homeostasis in Mice.” Cell Metabolism 30(5):976–986.e3. doi: 10.1016/j.cmet.2019.08.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takada Yasunari, Mukhopadhyay Asok, Kundu Gopal C., Mahabeleshwar Ganapati H., Singh Sujay, and Aggarwal Bharat B.. 2003. “Hydrogen Peroxide Activates NF-Kappa B through Tyrosine Phosphorylation of I Kappa B Alpha and Serine Phosphorylation of P65: Evidence for the Involvement of I Kappa B Alpha Kinase and Syk Protein-Tyrosine Kinase.” The Journal of Biological Chemistry 278(26):24233–41. doi: 10.1074/jbc.M212389200. [DOI] [PubMed] [Google Scholar]
- Tao Jun, Wu Qin-Yi, Ma Yi-Cheng, Chen Yuan-Li, and Zou Cheng-Gang. 2017. “Antioxidant Response Is a Protective Mechanism against Nutrient Deprivation in C. Elegans.” Scientific Reports 7(1):43547–43547. doi: 10.1038/srep43547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor Seth R., Santpere Gabriel, Weinreb Alexis, Barrett Alec, Reilly Molly B., Xu Chuan, Varol Erdem, Oikonomou Panos, Glenwinkel Lori, Rebecca McWhirter Abigail Poff, Basavaraju Manasa, Rafi Ibnul, Yemini Eviatar, Cook Steven J., Abrams Alexander, Vidal Berta, Cros Cyril, Tavazoie Saeed, Sestan Nenad, Hammarlund Marc, Hobert Oliver, and Miller David M.. 2021. “Molecular Topography of an Entire Nervous System.” Cell 184(16):4329–4347.e23. doi: 10.1016/j.cell.2021.06.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thacker Colin, and Rose Ann M.. 2000. “A Look at the Caenorhabditis Elegans Kex2/Subtilisin-like Proprotein Convertase Family.” BioEssays 22(6):545–53. doi: 10.1002/(SICI)15211878(200006)22:6<545::AID-BIES7>3.0.CO;2-F. [DOI] [PubMed] [Google Scholar]
- Thomas J. H. 1990. “Genetic Analysis of Defecation in Caenorhabditis Elegans.” Genetics (Austin) 124(4):855–72. doi: 10.1093/genetics/124.4.855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Titos Iris, Juginović Alen, Vaccaro Alexandra, Nambara Keishi, Gorelik Pavel, Mazor Ofer, and Rogulja Dragana. 2023. “A Gut-Secreted Peptide Suppresses Arousability from Sleep.” Cell 186(10):2273–74. doi: 10.1016/j.cell.2023.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Topham Matthew K. 2006. “Signaling Roles of Diacylglycerol Kinases.” Journal of Cellular Biochemistry 97(3):474–84. doi: 10.1002/jcb.20704. [DOI] [PubMed] [Google Scholar]
- Turner Chris D., Ramos Carmen M., and Curran Sean P.. 2024. “Disrupting the SKN-1 Homeostat: Mechanistic Insights and Phenotypic Outcomes.” Frontiers in Aging 5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van An-Angela N., Kunkel Maya T., Baffi Timothy R., Lordén Gema, Antal Corina E., Banerjee Sourav, and Newton Alexandra C.. 2021. “Protein Kinase C Fusion Proteins Are Paradoxically Loss of Function in Cancer.” The Journal of Biological Chemistry 296:100445–100445. doi: 10.1016/j.jbc.2021.100445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raamsdonk Van, Michael Jeremy, and Hekimi Siegfried. 2010. “Reactive Oxygen Species and Aging in Caenorhabditis Elegans: Causal or Casual Relationship?” Antioxidants & Redox Signaling 13(12):1911–53. doi: 10.1089/ars.2010.3215. [DOI] [PubMed] [Google Scholar]
- Vincent Andrea M., Russell James W., Sullivan Kelli A., Backus Carey, Hayes John M., McLean Lisa L., and Feldman Eva L.. 2007. “SOD2 Protects Neurons from Injury in Cell Culture and Animal Models of Diabetic Neuropathy.” Experimental Neurology 208(2):216–27. doi: 10.1016/j.expneurol.2007.07.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vitali Roberta, Prioreschi Clara, Rebenaque Laura Lorenzo, Colantoni Eleonora, Giovannini Daniela, Frusciante Sarah, Diretto Gianfranco, Marco-Jiménez Francisco, Mancuso Mariateresa, Casciati Arianna, and Pazzaglia Simonetta. 2022. “Gut–Brain Axis: Insights from Hippocampal Neurogenesis and Brain Tumor Development in a Mouse Model of Experimental Colitis Induced by Dextran Sodium Sulfate.” International Journal of Molecular Sciences 23(19):11495-. doi: 10.3390/ijms231911495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Han, Girskis Kelly, Janssen Tom, Chan Jason P., Dasgupta Krishnakali, Knowles James A., Schoofs Liliane, and Sieburth Derek. 2013. “Neuropeptide Secreted from a Pacemaker Activates Neurons to Control a Rhythmic Behavior.” Current Biology 23(9):746–54. doi: 10.1016/j.cub.2013.03.049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Xiaowen, Zhang Hong, Sapio Russell, Yang Jun, Wong Justin, Zhang Xin, Guo Jessie Y., Pine Sharon, Van Remmen Holly, Li Hong, White Eileen, Liu Chen, Kiledjian Megerditch, Pestov Dimitri G., and Zheng X. F. Steven. 2021. “SOD1 Regulates Ribosome Biogenesis in KRAS Mutant Non-Small Cell Lung Cancer.” Nature Communications 12(1):2259–2259. doi: 10.1038/s41467-021-22480-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West D. B., Fey D., and Woods S. C.. 1984. “Cholecystokinin Persistently Suppresses Meal Size but Not Food Intake in Free-Feeding Rats.” American Journal of Physiology. Regulatory, Integrative and Comparative Physiology 246(5):776–R787. doi: 10.1152/ajpregu.1984.246.5.r776. [DOI] [PubMed] [Google Scholar]
- Yanase Yuhki, Hide Izumi, Mihara Shoji, Shirai Yasuhito, Saito Naoaki, Nakata Yoshihiro, Hide Michihiro, and Sakai Norio. 2011. “A Critical Role of Conventional Protein Kinase C in Morphological Changes of Rodent Mast Cells.” Immunology and Cell Biology 89(1):149–59. doi: 10.1038/icb.2010.67. [DOI] [PubMed] [Google Scholar]
- Yang Yan, Craig Tim J., Chen Xiaohui, Ciufo Leonora F., Takahashi Masami, Morgan Alan, and Gillis Kevin D.. 2007. “Phosphomimetic Mutation of Ser-187 of SNAP-25 Increases Both Syntaxin Binding and Highly Ca2+-Sensitive Exocytosis.” The Journal of General Physiology 129(3):233–44. doi: 10.1085/jgp.200609685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu Hong Bing, Yang Hyungjun, Allaire Joannie M., Ma Caixia, Graef Franziska A., Mortha Arthur, Liang Qiaochu, Bosman Else S., Reid Gregor S., Waschek James A., Osborne Lisa C., Sokol Harry, Vallance Bruce A., and Jacobson Kevan. 2021. “Vasoactive Intestinal Peptide Promotes Host Defense against Enteric Pathogens by Modulating the Recruitment of Group 3 Innate Lymphoid Cells.” Proceedings of the National Academy of Sciences - PNAS 118(41):1-. doi: 10.1073/pnas.2106634118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zelko Igor N., Mariani Thomas J., and Folz Rodney J.. 2002. “Superoxide Dismutase Multigene Family: A Comparison of the CuZn-SOD (SOD1), Mn-SOD (SOD2), and EC-SOD (SOD3) Gene Structures, Evolution, and Expression.” Free Radical Biology & Medicine 33(3):337–49. doi: 10.1016/S0891-5849(02)00905-X. [DOI] [PubMed] [Google Scholar]
- Zhang Fujian, Ström Anna-Lena, Fukada Kei, Lee Sangmook, Hayward Lawrence J., and Zhu Haining. 2007. “Interaction between Familial Amyotrophic Lateral Sclerosis (ALS)-Linked SOD1 Mutants and the Dynein Complex.” The Journal of Biological Chemistry 282(22):16691–99. doi: 10.1074/jbc.M609743200. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.