Abstract
Endoplasmic reticulum (ER) homeostasis in the hypothalamus has been implicated in the pathogenesis of diet-induced obesity (DIO) and type 2 diabetes; however, the underlying molecular mechanism remain vague and debatable. Here we report that SEL1L-HRD1 protein complex of the highly conserved ER-associated protein degradation (ERAD) machinery in POMC-expressing neurons ameliorates diet-induced obesity and its associated complications, partly by regulating the turnover of the long isoform of Leptin receptors (LepRb). Loss of SEL1L in POMC-expressing neurons attenuates leptin signaling and predisposes mice to HFD-associated pathologies including fatty liver, glucose intolerance, insulin and leptin resistance. Mechanistically, nascent LepRb, both wildtype and disease-associated Cys604Ser variant, are misfolding prone and bona fide substrates of SEL1L-HRD1 ERAD. In the absence of SEL1L-HRD1 ERAD, LepRb are largely retained in the ER, in an ER stress-independent manner. This study uncovers an important role of SEL1L-HRD1 ERAD in the pathogenesis of central leptin resistance and leptin signaling.
Subject terms: Endoplasmic reticulum, Obesity, Protein aggregation, Hormone receptors
The importance of protein degradation in the ER during leptin signaling has been unclear. Here, the authors show that SEL1L-HRD1 ER-associated degradation modulates leptin signaling in POMC neurons by degrading misfolded leptin receptors in the ER.
Introduction
Hypothalamic neurons play key roles in regulating metabolic processes and energy balance, adapting to changes in diet and body weight1. In conditions such as diet-induced obesity (DIO), the hypothalamus may undergo maladaptive changes that exacerbate weight gain and insulin resistance, contributing to the development and progression of type-2 diabetes2–8. Proteostasis within the endoplasmic reticulum (ER) is essential for maintaining cellular function and overall physiological homeostasis9–15. It has been proposed that the hypothalamic unfolded protein response (UPR), an ER quality-control mechanism activated by the buildup of misfolded proteins in the ER, may significantly contribute to the development of DIO and type-2 diabetes through the regulation of inflammation and leptin resistance16–22. However, other studies have reported that UPR might have a protective role in similar experimental settings23,24. Consequently, the significance and underlying mechanisms of ER quality control pathways in DIO remain a subject of debate.
In addition to the UPR, ER-associated degradation (ERAD) is a constitutively active and highly conserved mechanism that targets unfolded or misfolded proteins in the ER for degradation by the cytosolic proteasomes25–32. Among the many putative ERAD complexes, the SEL1L-HRD1 protein complex represents the most evolutionarily conserved branch26,28,32–34. In this complex, SEL1L is an obligatory cofactor for the E3 ligase HRD1 by mediating substrate recruitment and stabilizing HRD1 ERAD complex29–31,35–37. Recent studies using cell type-specific SEL1L or HRD1 knockout mouse models have highlighted the (patho-)physiological importance of the SEL1L-HRD1 ERAD pathway in a substrate-specific manner38–45. Particularly relevant to this study, we previously showed that SEL1L-HRD1 ERAD regulates water balance and food intake by controlling the maturation of prohormones proAVP and POMC, respectively42,43. POMC neuron-specific Sel1L deletion results in hyperphagia and age-associated obesity starting around 13 weeks of age when fed a low-fat chow diet42. Given the importance of POMC neurons in maintaining energy homeostasis under various nutritional status, one outstanding question is the relevance and significance of SEL1L-HRD1 ERAD in POMC neurons under pathophysiological conditions, such as DIO.
Here, we show that SEL1L-HRD1 ERAD in POMC-expressing neurons at the arcuate nucleus (ARC) of the hypothalamus controls DIO pathogenesis and leptin sensitivity via regulating the turnover of the long isoform of LepR (LepRb), responsible for leptin signaling46–49. POMC-specific Sel1L deficient (Sel1LPOMC) mice are hypersensitive to high fat diet (HFD) feeding. We further show that nascent LepRb protein is unstable and degraded by SEL1L-HRD1 ERAD, a process required for functional LepRb to reach the cell surface.
Results
HFD-induced transient upregulation of hypothalamic SEL1L-HRD1 ERAD expression
Given that the SEL1L-HRD1 protein complex is constitutively expressed in the ARC of the hypothalamus42, we investigated whether its expression in this region is regulated in response to overnutrition. To this end, we placed transgenic mice expressing the eGFP transgene under the control of POMC gene promoter42,50 on 60% HFD consisting of 60% calories derived from fat for 1 or 8 weeks. As expected, HFD feeding reduced the expression of Pomc, Npy and Agrp genes (Supplementary Fig. 1a) and increased the protein levels of POMC derivatives β-Endorphin and α-melanocyte-stimulating hormone (α-MSH) (Supplementary Fig. 1b–e). Moreover, HFD feeding enhanced neuronal activity in hypothalamic paraventricular nucleus (PVH) region as measured by nuclear c-FOS following both 1- and 8-week HFD (Supplementary Fig. 1d, e).
One-week HFD significantly induced Hrd1 mRNA level in the ARC, but not Sel1L mRNA level, while 8-week HFD feeding had no such effect (Fig. 1a). At the protein levels, both SEL1L and HRD1 proteins, were elevated in the ARC after 1-week HFD, and returned to the basal levels after 8-week HFD (Fig. 1b, c). Confocal microscopic analysis further showed the upregulation of SEL1L and HRD1 expression in the POMC-expressing neurons upon 1-week HFD, while returned to the basal level following 8-week HFD (Fig. 1d–g). In non-POMC neurons, HRD1 protein level was transiently upregulated, but not SEL1L (Fig. 1d–g). Hence, SEL1L-HRD1 expression in POMC-expressing neurons are responsive to acute, but not chronic, HFD feeding.
Fig. 1. Transient upregulation of SEL1L-HRD1 ERAD expression in the hypothalamus in response to high fat diet (HFD) feeding.
a Quantitative PCR (qPCR) analysis of Sel1L and Hrd1 mRNA levels in the ARC of the C57BL/6 J male mice fed on NCD, 1w- and 8w-HFD (n = 4 mice per group). b, c Representative images (b) and quantitation (c) of Western blot of SEL1L and HRD1 proteins in the ARC of the C57BL/6 J male mice fed on NCD, 1w- and 8w-HFD (n = 13 mice per group). d–g Representative image and quantitation of IF staining of SEL1L (d, e) and HRD1 (f, g) in the ARC of C57BL/6 J POMC-eGFP transgenic male mice fed on NCD, 1w- and 8w-HFD. Yellow arrows, GFP+ POMC neurons; white arrowheads, non-POMC neurons. e NCD, n = 372, 280 for POMC and non-POMC neurons; 1w-HFD, n = 335, 210 for POMC and non-POMC neurons; 8w-HFD: n = 350, 210 for POMC and non-POMC neurons. g NCD, n = 199, 376 for POMC and non-POMC neurons; 1w-HFD, n = 301, 284 for POMC and non-POMC neurons; 8w-HFD: n = 225, 211 for POMC and non-POMC neurons. NCD for normal chow diet; HFD for high fat diet; ARC, arcuate nucleus. arb. units, arbitrary units. Values, mean ± SEM. n.s., not significant; p values are as indicated unless ****p < 0.0001 by one-way ANOVA followed by Tukey’s multiple comparisons test (a, c, e, g). Source data are provided as a Source Data file.
Hypothalamic POMC-specific ERAD deficiency leads to early-set DIO and its pathologies
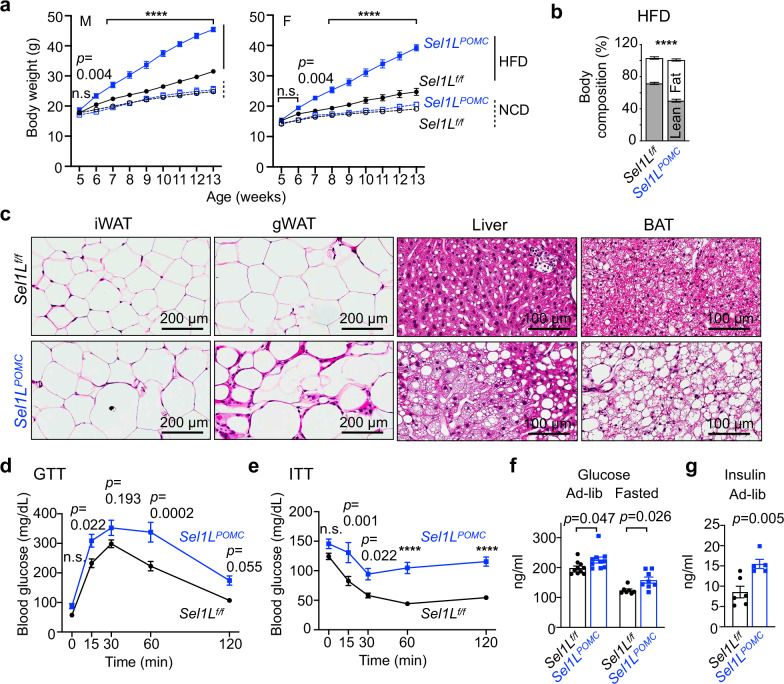
To delineate the importance of hypothalamic ERAD in DIO, we next examined the phenotypes of Sel1LPOMC mice generated by crossing Sel1Lf/f with the POMC-Cre mouse line42. The mice were subjected to an 8-week HFD or chow diet starting from 5 weeks of age. In line with our previous findings, Sel1LPOMC mice grew comparably to WT littermates on chow diet for the first 13 weeks of age42 (Fig. 2a). By contrast, soon after placed on HFD, Sel1LPOMC mice, both sexes, gained significantly more body weight than WT littermates (Fig. 2a). Body composition analysis showed that fat content was significantly increased in Sel1LPOMC mice, reaching over 50% of body mass after 8-week HFD with more lipid deposition in the livers, and both white and brown adipose tissues (WAT and BAT) (Fig. 2b, c and Supplementary Fig. 2a). HFD-fed Sel1LPOMC mice became highly glucose intolerant and insulin resistant (Fig. 2d, e), with elevated ad libitum and fasting blood glucose (Fig. 2f) as well as ad libitum insulin levels (Fig. 2g). In addition, Sel1LPOMC mice exhibited elevated levels of glucagon and corticosterone, while their rectal temperature 2 degrees Celsius lower compared to WT littermates (Supplementary Fig. 2b–d). Hence, we concluded that ERAD-deficient mice in POMC-expressing neurons are susceptible to DIO and its pathologies.
Fig. 2. Hypothalamic POMC-specific ERAD deficiency leads to DIO and its pathologies.
a Growth curve of Sel1Lf/f and Sel1LPOMC male (left) and female (right) mice fed NCD or HFD (NCD: n = 24 male and 16 female Sel1Lf/f mice, n = 9 male and 9 female Sel1LPOMC mice; HFD: n = 21 male and 10 female Sel1Lf/f mice, n = 18 male and 9 female Sel1LPOMC mice). b Body composition of Sel1Lf/f and Sel1LPOMC male mice after 8w-HFD (n = 4 Sel1Lf/f and n = 7 Sel1LPOMC). c H&E images of peripheral tissues from male mice fed HFD for 8 weeks (n = 3 mice per group). iWAT inguinal white adipose tissue, gWAT gonadal white adipose tissue, BAT brown adipose tissues. d, e GTT (d) and ITT (e) in 8w-HFD male mice fasted for 16 or 6 hrs prior to glucose (2 g/kg body weight) or insulin (1 unit/kg body weight) injections, respectively (n = 6 mice per group). f Serum glucose levels in 8w-HFD male mice either under ad-lib or 6h-fasting (ad-lib: n = 10 mice per group; fasting: n = 7,8 mice for Sel1Lf/f and Sel1LPOMC). g Ad-lib insulin levels in 8w-HFD male mice (n = 6, 5 mice for Sel1Lf/f and Sel1LPOMC). NCD for normal chow diet; HFD for high fat diet. Values, mean ± SEM. n.s. not significant; p values are as indicated unless ****p < 0.0001 by two-way ANOVA followed by Tukey’s multiple comparisons test (a) or Bonferroni’s multiple comparisons (b, d–f), or two-tailed Student’s t-test (g). Source data are provided as a Source Data file.
ERAD deficiency in POMC-expressing neurons triggers hyperphagia and leptin resistance
We next explored the possible mechanism underlying the susceptibility to DIO in Sel1LPOMC mice. Sel1LPOMC mice consumed ~ 40% more food daily, i.e., hyperphagia, at both 1- and 8-week HFD (Fig. 3a). To demonstrate the causal link between food intake and weight gain, we performed pair feeding (providing the same amount of the food consumed by WT littermates) after 8 weeks of ad libitum HFD feeding. Sel1LPOMC mice showed rapid weight gain under ad libitum; however, their weight gain significantly slowed down with pair feeding and returned to previous levels when placed back on ad libitum HFD (Fig. 3b). Moreover, weight gain of Sel1LPOMC mice was comparable to that of WT littermates with pair-feeding at the beginning of HFD feeding at 5 weeks of age (Fig. 3c). We then tested whether hyperphagia of Sel1LPOMC mice is caused by leptin resistance. Hyperleptinemia via leptin injection induces body weight loss in WT mice, but not Sel1LPOMC mice (Fig. 3d, e). Indeed, unlike WT mice, Sel1LPOMC mice continued to gain body weight following leptin injection (Fig. 3e). This difference in body weight gain was likely due to the differences in food intake in response to leptin injection (Fig. 3f), pointing to leptin resistance in Sel1LPOMC mice. Indeed, Sel1LPOMC mice developed increasingly severe hyperleptinemia with HFD feeding (Fig. 3g). Hence, we concluded that ERAD deficiency in POMC-expressing neurons triggers hyperphagia and leptin resistance.
Fig. 3. Hypothalamic ERAD deficiency triggers hyperphagia and leptin resistance.
a Daily food intake of Sel1Lf/f and Sel1LPOMC male mice at 1w- and 8w-HFD (1w-HFD: n = 11, 9 mice for Sel1Lf/f and Sel1LPOMC; 8w-HFD: n = 9, 11 mice for Sel1Lf/f and Sel1LPOMC). b Growth of male Sel1LPOMC mice under 8-week ad libitum, 8-week pair feeding and 1-week ad libitum on HFD (n = 3 mice, blue dots). Male Sel1Lf/f mice fed ad libitum with the same diets were included as controls (n = 3 mice, black dots). c Growth of Sel1Lf/f male mice ad libitum and Sel1LPOMC male mice either ad libitum or pair-feeding of HFD starting from 5 weeks of age (ad libitum: n = 11, 8 mice for Sel1Lf/f and Sel1LPOMC; pair feeding: n = 3 Sel1LPOMC). d, e, f Diagram of leptin sensitivity test (d). 12-week-old male mice on HFD were injected daily i.p. with vehicle PBS and then PBS or leptin (2 mg/kg body weight) for 3 days. e, f Body weight change (e), average daily food intake (f) following 3 daily vehicle or leptin injections of the male mice (PBS: n = 4 mice per group; leptin: n = 6, 4 mice for Sel1Lf/f and Sel1LPOMC). Body weight change was calculated by end point body weights minus starting point body weights. g Serum leptin levels in male mice fed on NCD, 1w- and 8w-HFD (NCD: n = 13, 11 mice for Sel1Lf/f and Sel1LPOMC; 1w-HFD: n = 7, 8 mice for Sel1Lf/f and Sel1LPOMC; 8w-HFD: n = 5, 8 mice for Sel1Lf/f and Sel1LPOMC). NCD for normal chow diet; HFD for high fat diet. Values, mean ± SEM. n.s. not significant; p values are as indicated unless ****p < 0.0001 by two-way ANOVA followed by Bonferroni’s multiple comparisons test (a, b, e, f) or Tukey’s multiple comparisons test (c, g). Source data are provided as a Source Data file.
The effect of hypothalamic SEL1L-HRD1 ERAD in DIO is mediated by central leptin resistance
To further establish the effect of leptin resistance in ERAD deficiency-associated DIO, we next performed parabiosis where two littermates were surgically stitched together to allow the sharing of the circulation (Fig. 4a). Following two weeks of recovery on chow diet, the parabionts were placed on HFD for 8 weeks (Fig. 4a). Two control parabionts, WT:WT (Group I) and Sel1LPOMC:Sel1LPOMC (Group II), gained weight as expected with the latter pair becoming obese (Fig. 4a, b). By contrast, in WT:Sel1LPOMC (Group III) parabionts, body weight gain for WT mice was attenuated compared to WT mice in WT:WT (Group I) parabionts, while body weight gain for Sel1LPOMC mice was comparable to that of Sel1LPOMC:Sel1LPOMC parabionts (Group II) (Fig. 4a, b). Body compositions (i.e., lean vs. fat) in parabionts were not affected by the partner (Fig. 4c). Moreover, serum leptin and insulin levels were highly elevated in Sel1LPOMC mice, but unaltered in WT mice regardless of the partners (Fig. 4d, e), likely due to short half-life of circulating hormones as previously reported51,52. Overall, these data suggested that Sel1LPOMC mice are defective in responding to circulating leptin.
Fig. 4. Hypothalamic SEL1L-HRD1 deficiency leads to DIO via leptin signaling.
a Schematic diagram for the experimental design (left) and pictures of Sel1Lf/f and Sel1LPOMC female parabionts following 8-week HFD (n = 3, 1, 5 pairs in groups I-III). b–e Body weights (b), body composition (c), serum leptin (d) and insulin (e) levels in female parabionts before and after parabiosis and 8-week HFD (n = 6, 2, 5 mice in groups I–III). NCD for normal chow diet; HFD for high fat diet. Values, mean ± SEM. n.s. not significant; p values are as indicated unless ****p < 0.0001 by two-way ANOVA followed by Bonferroni’s multiple comparisons test (b) or Tukey’s multiple comparisons test (c–e). Source data are provided as a Source Data file.
SEL1L-HRD1 ERAD deficiency impairs leptin-pSTAT3 signaling
We next asked how POMC-specific SEL1L-HRD1 ERAD regulates leptin sensitivity. As leptin induces phosphorylation of STAT3 (pSTAT3), we examined the levels of pSTAT3 in POMC neurons following leptin challenge. To visualize POMC neurons, we generated Sel1LPOMC mice with POMC-eGFP reporter (Sel1LPOMC;POMC-eGFP). HFD feeding progressively blunted leptin-induced pSTAT3 in POMC neurons of the ARC region of WT mice, but to a much greater extent, in Sel1LPOMC mice (Fig. 5a–d and Supplementary Fig. 3). In keeping with the notion that pSTAT3 a critical transcription factor for the Pomc gene53, hypothalamic Pomc mRNA expression was markedly decreased in Sel1LPOMC mice with HFD compared to WT littermates (Fig. 5e). Western blot analysis of pSTAT3 of the ARC region of Sel1LPOMC mice also showed a greater reduction of the percent of pSTAT3 following HFD feeding (Fig. 5f, g). Thus, our data suggested that SEL1L-HRD1 ERAD is required for leptin signaling in POMC-expressing neurons.
Fig. 5. Hypothalamic SEL1L-HRD1 ERAD deficiency impairs leptin-pSTAT3 signaling.
a–d Representative immunofluorescence (IF) staining of pSTAT3 (red) and eGFP (green) in Sel1Lf/f;Pomc-eGFP and Sel1LPOMC;Pomc-eGFP mice under NCD (a), 1w-HFD (b) and 8w-HFD (c). Mice were fasted overnight followed by i.p. injection with leptin (2 mg/kg body weight), with quantitation of percentage of pSTAT3+ POMC neurons shown in (d). Mice injected with PBS were included as negative controls (Supplemental Fig. 3). Yellow arrows, pSTAT3+ POMC neurons; white arrowheads, pSTAT3- POMC neurons. (NCD: n = 4 mice per genotype; 1w-, 8w-HFD and PBS: n = 3 mice per genotype). e Quantitative PCR (qPCR) analysis of Pomc mRNA levels in ARC of Sel1Lf/f and Sel1LPOMC male mice at 8w-HFD (n = 3 mice per group). f, g Representative Western blot analysis (f) and quantitation (g) of leptin-induced pSTAT3 in ARC of Sel1Lf/f and Sel1LPOMC mice under NCD or 8w-HFD (NCD: n = 2 mice per genotype for PBS, n = 4 mice per genotype for leptin; 8w-HFD: n = 4 mice per group). NCD for normal chow diet; HFD for high fat diet; ARC, arcuate nucleus. arb. units, arbitrary units. Values, mean ± SEM. n.s. not significant; p values are as indicated unless ****p < 0.0001 by two-way ANOVA followed by Bonferroni’s multiple comparisons test (d, e, g). Source data are provided as a Source Data file.
ERAD effect in POMC-expressing neurons is uncoupled from UPR or inflammation
Given the reported role of UPR in DIO pathogenesis16,18–21, we next asked whether ERAD deficiency activates UPR and if so, to what extent. There was no detectable activation of the PERK pathway as measured by phosphorylation of PERK and its downstream phosphorylation of eIF2α in the ARC of Sel1LPOMC mice (Fig. 6a and Supplementary Fig. 4a). Phosphorylation of IRE1α, on the other hand, was moderately elevated in the ARC of Sel1LPOMC mice, so was the splicing of Xbp1 mRNA (a downstream effector of IRE1α) as well as the ER chaperone BiP (Fig. 6b, c and Supplementary Fig. 4a–d). In vitro, treatment with an ER stress inducer thapsigargin (Tg) induced strong ER stress, but failed to affect leptin signaling in WT HEK293T cells transfected with mouse LepRb (mLepRb) (Fig. 6d and Supplementary Fig. 4e), indicating that UPR is not sufficient to induce leptin resistance. This finding contrasts with previous reports that have causally linked UPR to leptin resistance16,18–21.
Fig. 6. POMC-specific ERAD deficiency is associated with low levels of UPR and inflammation.
a–c Representative Western blot analysis and quantitation of the PERK pathway (a, n = 6 mice per group), and phostag-gel (P-T) analysis of IRE1α phosphorylation (b, n = 3 mice per group), BiP levels (b, n = 6 mice per group), and reverse transcriptase PCR (RT-PCR) analysis and quantitation of Xbp1 mRNA splicing (c, n = 5, 4 mice for Sel1Lf/f and Sel1LPOMC) in the ARC of Sel1Lf/f and Sel1LPOMC mice on 8w-HFD. Livers from male mice treated with tunicamycin (TM, 1 mg/kg, i.p.) for 24 hrs (Liver_TM), liver lysates treated with Lambda protein phosphatase (Liver_TM w/ λPP), and livers under basal condition (Liver_CON) were included as controls. d Representative Western blot and quantitation showing the lack of effect of ER stress on leptin signaling. WT HEK293T transfected with mLepRb were treated with thapsigargin (Tg) at the indicated concentration for 4 hrs and leptin for 30 min in serum free medium, followed by Western blot and RT-PCR analyses of UPR activation (n = 5, 2, 2 independent cell samples per group for Western blot, P-T gel, and RT-PCR). e Representative confocal images and quantitation of the number of GFP+ POMC neurons in Sel1Lf/f;POMC-eGFP and Sel1LPOMC;POMC-eGFP mice on 8w-HFD (n = 6, 9 mice for Sel1Lf/f;Pomc-eGFP and Sel1LPOMC;Pomc-eGFP). f, g Representative Western blot analysis (f) and quantitation (g) of inflammatory markers in the ARC of Sel1Lf/f and Sel1LPOMC male mice fed on 8w-HFD (n = 3 mice per group). arb. units, arbitrary units. HFD for high fat diet; ARC, arcuate nucleus. arb. units, arbitrary units. Values, mean ± SEM. n.s., not significant; p values are as indicated by two-way ANOVA followed by Tukey’s multiple comparisons test (d) or two-tailed Student’s t-test (a–c, e, g). Source data are provided as a Source Data file.
We found no significant POMC neuronal loss in the ARC of Sel1LPOMC;POMC-eGFP mice compared to WT littermates following 8-week HFD (Fig. 6e). Inflammatory markers were largely comparable in the ARC of Sel1LPOMC mice vs. WT littermates as measured by phosphorylation and protein levels of c-Jun N-terminal Kinase (JNK) as well as protein levels of I kappa B alpha (IκBα) (Fig. 6f–g). Astrogliosis in the ARC regions was comparable between WT and Sel1LPOMC mice, as demonstrate by Western blot and immunofluorescence staining of the astrocyte marker Glial Fibrillary acidic protein (GFAP) and/or microglia marker Ionized calcium-binding adaptor molecule 1 (IBA1) (Fig. 6f, g and Supplementary Fig. 4f). Taken together, we concluded that Sel1L deficiency in POMC-expressing neurons triggers leptin resistance independently of UPR, neuronal cell death, and inflammation.
LepRb protein is an endogenous substrate of SEL1L-HRD1 ERAD
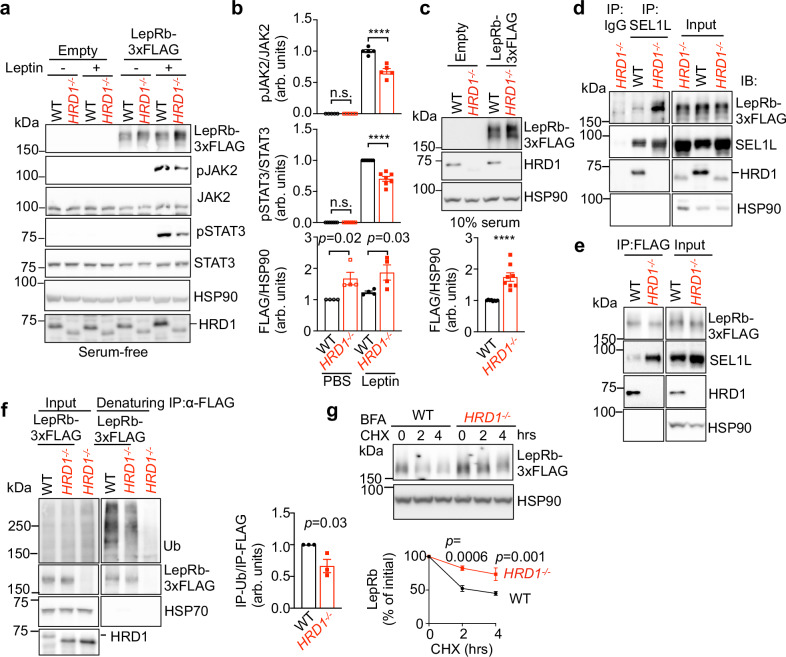
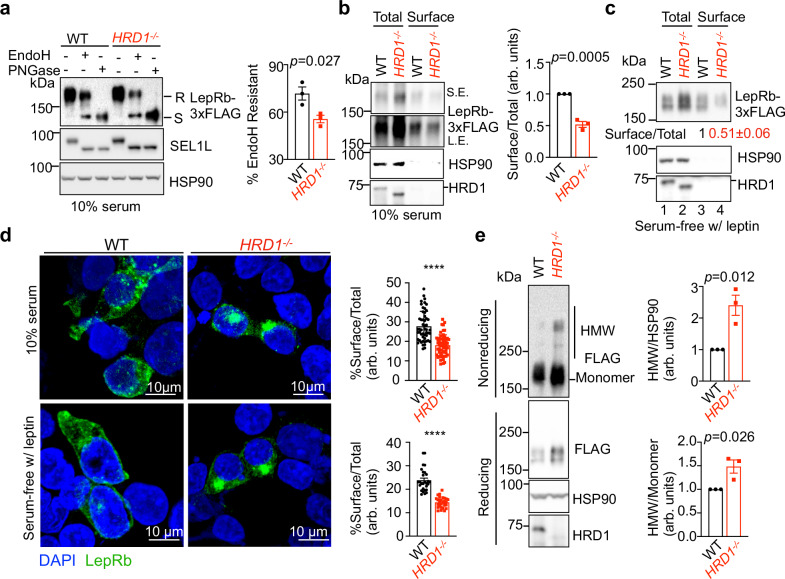
The aforementioned data suggested that SEL1L-HRD1 ERAD regulates leptin sensitivity upstream of STAT3. To further explore the underlying mechanism, we generated leptin-responsive LepRb-expressing HEK293T cells. In line with decreased leptin sensitivity in vivo, HRD1−/− HEK293T cells exhibited impaired phosphorylation of JAK2 and STAT3 compared to those in WT cells in response to leptin stimulation (Fig. 7a, b), despite having much higher LepRb protein levels (Fig. 7a–c). Moreover, SEL1L interaction with LepRb in LepRb-transfected cells was markedly enhanced in HRD1−/− cells compared to that in WT cells (Fig. 7d, e). LepRb was ubiquitinated in an HRD1-dependent manner (Fig. 7f) and was significantly stabilized in HRD1−/− cells compared to that in WT cells (Fig. 7g). As LepRb is a glycosylated protein, we next performed endoglycosidase H (EndoH) digestion assay to separate the fraction of LepRb that have exited the ER (endoH resistant) or retained in the ER (endoH sensitive). Intriguingly, EndoH resistant fraction of LepRb were significantly reduced by ~40% in HRD1−/− HEK293T cells compared to WT cells (Fig. 8a). This result was further confirmed using the surface biotinylation assay followed by immunoprecipitation with streptavidin-beads, which also showed reduced amount of surface LepRb in total protein levels by 50% in HRD1−/− cells compared to that of WT cells (Fig. 8b, c). Moreover, confocal microscopy following immunofluorescence staining showed an altered distribution of LepRb with increased intracellular, but decreased surface, expression in ERAD-deficient cells (Fig. 8d and Supplementary Fig. 5). In the absence of SEL1L-HRD1, LepRb protein was prone to form high molecular weight (HMW) aggregates via disulfide bonds (Fig. 8e). Taken together, our data showed that nascent LepRb protein is unstable and degraded by SEL1L-HRD1 ERAD, a process required for functional LepRb to reach the cell surface.
Fig. 7. SEL1L-HRD1 interacts with and targets LepRb for degradation.
a, b Representative Western blot (a) and quantitation (b) for leptin (100 nM, 30 min)-induced phosphorylation of JAK2-STAT3 in HEK293T transfected with mLepRb-3xFlag in serum-free medium. (n = 5,7,4 independent samples per group for pJAK2, pSTAT3 and mLepRb-3xFLAG). arb. units, arbitrary units. c Representative Western blot and quantitation of mLepRb-3xFLAG protein levels in transfected HEK293T in complete medium (DMEM w/ 10% serum) (n = 8 independent samples per group). arb. units, arbitrary units. d, e Immunoblot analysis following immunoprecipitation (IP) of SEL1L (d) or FLAG (e) from lysates of WT or HRD1-/- HEK293T transfected with mLepRb-3xFlag (n = 2, 3 independent samples for SEL1L-IP and FLAG-IP). f Immunoblot analysis of Ub following denaturing immunoprecipitation (IP) of Flag from lysates of HEK293T transfected with mLepRb-3xFlag (n = 3 independent samples per group). arb. units, arbitrary units. g Protein decay analysis and quantitation (below) of LepRb protein levels in LepRb-3xFlag-expressing HEK293T cells treated with Brefeldin-A and cycloheximide (CHX) for the indicated time periods (n = 4 independent samples per group). Values, mean ± SEM. n.s., not significant; p values are as indicated unless ****p < 0.0001 by two-tailed Student’s t-test (c, f) or two-way ANOVA followed by Bonferroni’s multiple comparisons test (b, g). Source data are provided as a Source Data file.
Fig. 8. SEL1L-HRD1 ERAD deficiency promotes the ER-retention and aggregation of LepRb.
a Representative Western blot and quantitation (right) of LepRb protein following treatment with EndoH or PNGase in HEK 293T transfected mLepRb-3xFlag (n = 3 independent samples per group). arb. units, arbitrary units. b, c Measurement of surface mLepRb levels in HEK293T transfected with mLepRb-3xFlag in DMEM with 10% serum (b), or serum-free DMEM treated with leptin for 30 min (c) followed by biotinylation for membrane surface proteins followed by immunoprecipitation by streptavidin-beads (b, n = 3 independent samples per group; c, n = 2 independent samples per group). arb. units, arbitrary units. d Representative confocal IF images of mLepRb-3xFlag in transfected HEK293T cells in DMEM with 10% serum or serum-free DMEM treated with 100 nM leptin for 30 min. Quantitation of percent of surface signals over total shown on the right (10% serum: n = 52 WT and 57 HRD1−/− cells; serum-free w/leptin: n = 28 cells per genotype). arb. units, arbitrary units. Split channels are shown in Supplementary Fig. 5. e Western blot analyses and quantitation (right) of mLepRb-3xFlag protein in transfected HEK293T cells on reducing and nonreducing SDS-PAGE (n = 3 independent samples per group). arb. units, arbitrary units. Values, mean ± SEM. n.s., not significant; p values are as indicated unless ****p < 0.0001 by two-tailed Student’s t-test (a, b, d, e). Source data are provided as a Source Data file.
SEL1L-HRD1 ERAD degrades and limits the HMW aggregation of human LepRb Cys604Ser (C604S) mutant
To demonstrate the clinical relevance of our findings, we examined whether a subset of LepRb disease variants are SEL1L-HRD1 ERAD substrates. Here, we focused on LepRb mutant C604S, a recessive point mutation identified in two brothers aged 1 and 5 years, both with severe obesity54,55, with unclear mechanism54,56–58. C604-C674 forms a disulfide bond in human LepRb corresponding to C602-C672 in mouse (Supplementary Fig. 6a, b)56–58. Indeed, leptin treatment triggered very subtle pSTAT3 in WT HEK293T cells expressing C602S mLepRb compared to those expressing WT mLepRb, which was further reduced in HRD1−/− cells (Supplementary Fig. 6c). Similar to WT mLepRb, C602S mLepRb was stabilized in the absence of HRD1 (Supplementary Fig. 6d). Notably, C602S mLepRb readily formed HMW aggregates in WT HEK293T cells, and to a much greater extent in HRD1−/− cells (Supplementary Fig. 6e). Hence, we concluded that C604S LepRb variant is also a substrate of SEL1L-HRD1 ERAD.
Discussion
This study reports an important role of hypothalamic ERAD in maintaining energy homeostasis under nutrient overload conditions. SEL1L-HRD1 ERAD defects in POMC neurons predispose mice to DIO and its pathologies, largely due to leptin resistance and hyperphagia. Moreover, this study identifies a regulatory mechanism for LepRb signaling. Our data show that native LepRb is an endogenous substrate of SEL1L-HRD1 ERAD. This degradation event is crucial for the production of functional LepRb proteins at the cell surface as ERAD deficiency results in LepRb being retained in the ER (Fig. 9). Pointing to the clinical relevance of our findings, human LepRb C604S variant is trapped in the ER and degraded by SEL1L-HRD1 ERAD, leading to reduced expression at the cell surface. Taken together, this study uncovers the importance of SEL1L-HRD1 ERAD in leptin signaling and leptin biology.
Fig. 9. The indispensable role of SEL1L-HRD1 ERAD in POMC-expressing neurons.
a In POMC-expressing neurons, SEL1L-HRD1 ERAD constitutively degrades misfolded LepRb, a process to ensure the production of functional LepRb at the cell surface. b The absence of SEL1L-HRD1 ERAD in POMC-expressing neurons causes the accumulation and aggregation of misfolded LepRb, which disrupts the proper folding of nascent LepRb. This impairment in LepRb biogenesis results in severe leptin resistance in mice when on a HFD. Created in BioRender. Mao, H. (2024) BioRender.com/a66y170.
In our Sel1LPOMC mouse model, the POMC promoter is active at embryonic day 10.5 (E10.5)59, resulting in Sel1L deletion in POMC-expressing progenitors. As these progenitors differentiate into various neuronal types, including LepRb-expressing POMC neurons as well as non-POMC neurons1,60–63, the impact of SEL1L-HRD1 ERAD in Sel1LPOMC mice may reflect effects on both POMC and non-POMC neurons. Notably, non-POMC neurons, such as nearly a quarter of the hypothalamic AgRP/NPY neurons60 or functionally distinct POMC neurons61,62, could also influence leptin signaling and food intake. Earlier studies have shown that mice with LepRb deficiency in POMC-expressing neurons (using the same POMC-cre line as ours) gain significantly higher body weight in both sexes at 4–6 weeks of age on a regular chow diet64,65 or HFD65. However, another study using the inducible POMC Cre-ERT2 line to delete LepRb in POMC neurons of adult mice found systemic insulin and leptin resistance but no significant body weight gain four weeks after tamoxifen administration66. The discrepancies may be attributed to the possible involvement of other non-POMC neurons in the studies using the original POMC-cre line64,65. Therefore, future research is needed to explore the role of SEL1L-HRD1 ERAD in other types of hypothalamic neurons.
Sel1L deficiency in POMC neurons is associated with minimal, if any, UPR activation, cell death or inflammation, which is in line with many recent studies of different cell and tissue types40,42–45,67. Hence, we conclude that the effect of SEL1L-HRD1 ERAD is independent of the IRE1α -XBP1 pathway of the UPR and cell death. Furthermore, we recently identified several hypomorphic variants of SEL1L and HRD1 in patients with ERAD-associated neurodevelopmental disorders with infancy onset (EDNI) syndrome68,69. Notably, neither patient fibroblasts nor knockin HEK293T cells carrying these variants exhibited any overt UPR. These findings point to cellular adaptation in response to ERAD deficiency and the accumulation of misfolded proteins. Adaptive mechanisms include upregulation of ER chaperones to increase folding efficiency, expansion of ER volume to dilute misfolded protein concentration, enhanced aggregation and sequestration of misfolded proteins to reduce proteotoxicity, and/or activation of ER-phagy to clear misfolded protein aggregates or damaged ER26,28,32,34,39,70.
Previous reports have suggested that UPR may play a causal role in leptin resistance due to impaired leptin signaling16,18,71. These studies used ER stress inducers like tunicamycin and thapsigargin which can be fraught with artefacts. Tunicamycin, for example, inhibits glycosylation of glycoproteins72, including LepRb, which may impair leptin signaling directly due to defective glycosylation and functionality of LepRb, rather than UPR activation. Additionally, the high doses of ER stress inducers used in these studies may not be physiologically relevant16,18,71. In our study, thapsigargin treatment induced ER stress responses in a dose-dependent manner but did not alter leptin signaling in WT HEK293T cells transfected with mLepRb, even at a very high level. Thus, these findings suggest that the UPR is unlikely to play a key role in leptin signaling. Further studies are needed to test this model.
This study demonstrates an important role of SEL1L-HRD1 ERAD in leptin signaling, at least in part via the regulation of the turnover of LepRb protein (Fig. 9). We previously showed that SEL1L-HRD1 ERAD is required for the posttranslational maturation of the prohormone POMC in mice on chow diet and that Sel1L deficiency in POMC neurons causes age-associated obesity in mice on chow diet due to the ER retention of POMC42. In DIO mouse models, we found defects in Sel1LPOMC mice occurring upstream of Pomc gene transcription as leptin-induced STAT3 phosphorylation is impaired in the absence of SEL1L-HRD1 ERAD. Further mechanistic studies identified that partial loss-of-function of LepRb results from impaired ER exit of nascent LepRb protein in SEL1L-HRD1 ERAD deficient cells. This study suggests that nascent LepRb proteins are prone to misfolding in the ER, likely due to the formation of multiple disulfide bonds and glycosylation. If not cleared efficiently, misfolded LepRb with highly reactive cysteine thiols may promote the formation of aggregates with aberrant inter- or intra-molecular disulfide-bonds42–44. Consequently, they rely on SEL1L-HRD1 ERAD to create an ER environment conducive for the proper folding and conformation of nascent LepRb (Fig. 9). In this study, due to the lack of an antibody for LepRb detection in vivo, we conducted extensive mechanistic investigations in vitro instead. We acknowledge the limitations of the in vitro system used, which may not fully represent in vivo neuronal systems. Nevertheless, the extensive use of in vivo and in vitro approaches in this study provides strong support for our overall conclusions.
Methods
Study approval
All animal experiments were performed in compliance with University of Michigan (Ann Arbor, MI) and University of Virginia Institutional Animal Care and Use Committee (#PRO00006888 and # 4459) guidelines.
Mice
POMC-specific Sel1L-deficient mice (Sel1LPOMC) and control littermates (Sel1Lf/f) on the C57BL/6 J background were generated by crossing Sel1Lfl/fl mice on the C57BL/6 J background36 with mice on the POMC-promoter–driven Cre line on the C57BL/6 J background (B6.FVB-Tg(Pomc-cre)1Lowl/J, JAX 010714)64. Sel1LPOMC mice were further crossed with Pomc-eGFP reporter mice on the C57BL/6 J background (C57BL/6J-Tg(POMC-EGFP)1Low/J, JAX 009593)73 to generate Sel1LPOMC;Pomc-eGFP and control Sel1Lf/f;Pomc-eGFP littermates. WT B6 mice were purchased from JAX and bred in our mouse facility. Mice were maintained on a normal chow diet (13% fat, 57% carbohydrate and 30% protein, PicoLab Rodent Diet 5L0D) and placed on a high-fat diet (HFD, calories provided by 60% fat, 20% carbohydrate and 20% protein, Research Diet D12492) from 5 weeks of age for 1 or 8 weeks. All mice were housed in a temperature-controlled room (20 °C–23 °C) with a 12-hr light/12-hr dark cycle and 40–60% humidity.
Food intake measurement and pair-feeding
To perform daily food intake measurement, mice were first acclimatized to single housing 24 h before the experiment. Daily food intake was measured 1 h before the onset of the dark cycle each day. For the pair-feeding at later stage of HFD feeding, Sel1LPOMC and WT littermates had free access to HFD for eight weeks and were then single housed and fed ~2.5 g HFD, which was determined by the average of daily food intake of WT littermates, at the start of the dark cycle. For the pair-feeding at early stage of HFD feeding, 5-week-old Sel1LPOMC mice were split into two groups: One group of Sel1LPOMC and WT littermates had continuous free access to food; the other group of Sel1LPOMC mice (pair-fed) was fed ~2.5 g at the start of dark hrs. Weekly bodyweight gains were monitored.
Leptin treatment in mice
Twelve-week-old mice were acclimatized to single housing with HFD and intraperitoneal (i.p.) injection (PBS) for three days, followed by three daily leptin injections (2 mg/kg body weight, R&D systems; catalog 498-OB-05M) 1 h before the onset of dark cycle as described42. Body weight and food intake were monitored daily during the treatment period. For pSTAT3 staining, mice were injected i.p. with 2 mg/kg leptin, followed by overnight fasting. Mice were anesthetized by isoflurane for fixation-perfusion 30 min after injection.
Parabiosis
Parabiosis surgery was performed as previously described74,75. The 7-week-old female mice from the same cage were anesthetized via isoflurane vaporizer and shaved thoroughly from 1 cm above the elbow to 1 cm below the knee on the side to be connected. On the shaved side, a longitudinal skin incision was performed followed by the gentle detachment of skin from the subcutaneous fascia to create free skin with 0.5 cm width. Mice were then joined by tightly attaching the olecranons and the knees. The skin was sutured ventrally from the elbow towards the knee using continuous dorsal stitches. After 2-week recovery, mice were fed on HFD for 8 weeks. Body weights were monitored and serum insulin and leptin levels were measured.
Tissue and blood collection
Briefly, blood was collected from anesthetized mice via cardiac puncture, transferred to 1.5 ml microcentrifuge tubes, kept at room temperature for 30 min prior to centrifugation at 2000 g for 15 min. Serum was aliquoted and stored at −80 °C until analysis. For brain microdissection, Adult Mouse Brain Slicer Matrix (BSMAA001-1, Zivic Instruments) was used to collect coronal brain slices containing ARC region with further microdissection to obtain ARC-enriched region. All tissues were snap-frozen in liquid nitrogen and stored at −80 °C before use.
Preparation of brain sections
Mice were anesthetized with isoflurane, perfused with PBS followed by 4% paraformaldehyde (PFA) (Electron Microscopy Sciences; catalog 19210) for fixation. Brains were then postfixed in 4% PFA for overnight at 4 °C, dehydrated in 15% sucrose overnight at 4 °C and then 30% sucrose overnight at 4 °C, and sectioned (30 μm) on a cryostat (Microm HM550 Cryostat, Thermo Fisher Scientific). The sections were stored in DEPC-containing anti-freezing media (50% 0.05 M sodium phosphate pH 7.3, 30% ethylene glycol, 20% glycerol) at −20 °C. Different brain regions were identified using the Paxinos and Franklin atlas. Counted as distance from bregma, the following coordinates were used: PVN (–0.82 mm to –0.94 mm) and ARC (–1.58 mm to –1.7 mm).
Western blot and antibodies
Frozen tissue or cells were homogenized by sonication in lysis buffer (150 mM NaCl, 50 mM Tris pH 7.5, 10 mM EDTA, 1% Triton X-100) with freshly added protease inhibitors (Sigma; catalog P8340), phosphatase inhibitors (Sigma; catalog P5726) and 10 mM N-ethylmaleimide (Thermo Scientific; catalog 23030). Lysates were incubated on ice for 30 min followed by centrifugation (13,000 g, 10 min at 4 °C). Supernatants were collected and analyzed for protein concentration using Bradford assay (Bio-Rad; catalog 5000006). For denaturing SDS-PAGE, samples were further supplemented with 1 mM DTT and denatured at 95 °C for 5 min in 5x SDS sample buffer (250 mM Tris-HCl pH 6.8, 10% sodium dodecyl sulfate, 0.05% Bromophenol blue, 50% glycerol, and 1.44 M β-mercaptoethanol). For non-reducing SDA-PAGE, samples were prepared in 5x non-denaturing sample buffer (250 mM Tris-HCl pH 6.8, 10% sodium dodecyl sulfate, 0.05% bromophenol blue, 50% glycerol). For phostag gel analysis based on phos-tag system as described76,77, SDS-PAGE gel was supplemented with 50 μM MnCl2 (Sigma) and 25 μM phostag reagent (NARD Institute; catalog AAL-107). Protein lysates from the livers of mice treated with tunicamycin (TM, 1 mg/kg, i.p.) for 24 h was used as a positive control for UPR. For phosphatase treatment, 100 µg tissue lysates were incubated with 1 µl lambda phosphatase (λPPase, New England BioLabs; catalog P0753S) in 1× PMP buffer (New England BioLabs; catalog B0761S) with 1 mM MnCl2 (New England BioLabs; catalog B1761S) at 30 °C for 30 min. Reaction was stopped by adding 5× SDS sample buffer and incubated at 90 °C for 5 min.
All samples were incubated in 65 °C for 10 min and run with 15-30 μg total lysate on SDS-PAGE gel for separation followed by electrophoretic transfer to PVDF membrane (0.45μm, Millipore; catalog IPFL00010). The blots were incubated in 2% BSA/Tri-buffered saline tween-20 (TBST) with primary antibodies overnight at 4 °C, washed with TBST followed by 1 hr incubation with goat anti-rabbit or mouse IgG HRP at room temperature. Blots were scanned by the ChemiDoc imaging system (Bio-Rad #1708265 and #12003153). Band density was quantitated using the Image Lab software (v6.0.1, Bio-Rad). Antibodies for Western blot were as follows: SEL1L (rabbit, 1:8000, Abclonal; catalog E112049), HRD1 (rabbit, 1:2000, ABclonal; catalog E15102), GRP78 BiP (rabbit, 1:5000, Abcam; catalog ab21685), HSP90 (rabbit, 1:5,000, Santa Cruz Biotechnology Inc.; catalog sc-7947), HSP70 (rabbit, 1:1000, Santa Cruz Biotechnology Inc.; catalog sc-1060), FLAG (mouse, 1:2000, Sigma-Aldrich; catalog F-1804), IRE1α (rabbit, 1:2,000, Cell Signaling Technology; catalog 3294), p-eIF2α (rabbit, 1:2000, Cell Signaling Technology; catalog 3597), eIF2α (rabbit, 1:2000, Cell Signaling Technology; catalog 9722), p-JNK (mouse, 1:2000, Cell Signaling Technology; catalog 9255), JNK (rabbit, 1:1000, Cell Signaling Technology; catalog 9252), PERK (Rabbit, 1:1000, Cell Signaling Technology; catalog 3192), pSTAT3 (Tyr705) (rabbit, 1:1000, Cell Signaling Technology; catalog 9131), STAT3 (rabbit, 1:1000, Cell Signaling Technology; catalog 9132), pJAK2 (Tyr1007/1008) (rabbit, 1:1000, Cell Signaling Technology; catalog 3771), JAK2 (rabbit, 1:1000, ABclonal; catalog A19629), Tubulin (mouse, 1:5000, Santa Cruz Biotechnology Inc.; catalog sc-5286), IκBα (rabbit, 1:1000, Cell Signaling Technology; catalog 9242) and IBA1 (rabbit, 1:1000, Proteintech; catalog 10904-1-AP). Secondary antibodies for Western blot were goat anti-rabbit IgG HRP and goat anti-mouse IgG HRP at 1:5,000 from Bio-Rad.
Denaturing IP for ubiquitination assay
HEK293T cells were transfected with LepRb-3xFLAG for 48 hrs before snap-frozen in liquid nitrogen. The sample lysates were prepared in NP-40 lysis buffer (50 mM Tris-HCl at pH7.5, 150 mM NaCl, 1% NP-40, 1 mM EDTA) with protease and phosphatase inhibitors, followed by centrifugation at 16,000 × g for 10 min. Protein concentrations of supernatants were measured using Bradford before the addition of 1% SDS and 5 mM DTT at 95 °C for 10 min. Lysates were diluted 1:10 with NP-40 lysis buffer with the final SDS concentration ≤0.1% and incubated with 20 μl anti-FLAG agarose overnight at 4 °C with gently rocking. The agarose beads were washed three times with NP-40 lysis buffer with protease and phosphatase inhibitors. The samples were eluted from beads in the SDS sample buffer at 95 °C for 5 min prior to be subjected for SDS-PAGE and immunoblotting.
Immunostaining and antibodies
For immunostaining of free-floating brain sections, samples were picked out of anti-freezing buffer followed by 3 washes with PBS. Free-floating sections were simultaneously incubated with primary antibodies in blocking buffer (0.3% donkey serum and 0.25% Triton X-100 in 0.1 M PBS) overnight at 4 °C. Following three washes with PBS, sections were incubated with secondary antibodies for 2 h at room temperature. Brain sections were then mounted on gelatin-coated slides (Southern Biotech; catalog SLD01-CS). Counterstaining and mounting were performed with mounting medium containing DAPI (Vector Laboratories; catalog H-1200) and Fisherfinest Premium Cover Glasses (Fisher Scientific; catalog 12-548-5 P). For immunostaining in cells, 24 h after transfection of LepRb-3xFLAG constructs, cells were placed on Poly-L-Lysine (Advanced Biomatrix; catalog 5048) coated Millicell EZ SLIDE 8-well glasses (Millipore; catalog PEZGS0816) for 24 h before treatment and fixation. Permeabilization was included and the overall process were the same as described above. Images were captured using the Nikon A1 confocal microscope at the University of Michigan Morphology and Image Analysis Core. To quantify immunoreactivity, identical acquisition settings were used for imaging each brain section from all groups within an experiment. The numbers of immunoreactivity-positive soma analysis and intensity of immunoreaction were quantified in 3D stack volumes after uniform background subtraction using the NIS Elements AR software (Nikon) and FIJI (ImageJ2 version 2.14.0, National Institute of Health, USA).
Antibodies for immunostaining were as follows: HRD1 (rabbit, 1:500, homemade), GRP78 BiP (rabbit, 1:500, Abcam; catalog ab21685), α-MSH (sheep, 1:2,000, Millipore; catalog AB5087), β-endorphin (rabbit, 1:2,000, Phoenix Pharmaceuticals; catalog H-022-33, provided by Carol Elias), and GFP (chicken IgY, 1:300, Abcam; catalog ab13970), p-Y705 STAT3 (rabbit, 1:200, Cell Signaling Technology; catalog 9145), GFAP (rabbit, 1:500, Agilent; catalog Z033429-2), FLAG (mouse, 1:500, Sigma-Aldrich; catalog F-1804), KDEL (rabbit, 1:500, Novus Biologicals; catalog NBP2-75549), eIF3η (goat, 1:500, Santa Cruz Biotechnology; catalog sc-16377).
Secondary antibodies for fluorescent immunostaining (all 1:500) were as follows: Anti-rabbit IgG Alexa Fluor 647; anti-goat IgG Alexa Fluor 488 & 647; anti-sheep IgG Cy5 were from Jackson ImmunoResearch. Donkey anti-mouse IgG Alexa flour 555 was from Invitrogen (catalog A32773) and goat anti-chicken IgY FITC was from Aves Labs (catalog F-1005).
Plasmids
mLepRb cDNA was provided by Dr. Martin Myers at University of Michigan Medical School. The LepRb coding region was amplified by PCR using a primer set containing HindIII and XbaI restriction site at 5’ and 3’ respectively.
F: 5′- CCG AAGCTT ATGATGTGTCAGAAATTCTATGTGGTT-3′
R: 5′- TGC TCTAGA CACAGTTAAGTCACACATCTTATT-3′
Both PCR products and the backbone vector p3xFLAG-CMV14 were digested using HindIII and XbaI restriction enzymes in the double digestion system from New England BioLabs. For construction of LepRb point mutants, quick change mutagenesis was performed using PFU DNA polymerase (600140, Agilent). The following primers were used for mutagenesis to construct LepRb-C602S:
F: 5′- CCTGCTGGTGTCAGACCTCAGTGCAGTCTATG-3′
R: 5′- CATAGACTGCACTGAGGTCTGACACCAGCAGG-3′
CRISPR/Cas9-based knockout (KO) in HEK293T cells
HEK293T cells were were originally obtained from ATCC (catalog CRL-3216) and cultured at 37 °C with 5% CO2 in DMEM with 10% fetal bovine serum (Fisher Scientific; catalog FB12999102). To generate HRD1-deficient HEK293T cells, sgRNA oligonucleotides designed for human HRD1 (5′-GGACAAAGGCCTGGATGTAC-3′) was inserted into lentiCRISPR v2 (plasmid 52961, Addgene). Cells transfected with empty plasmids without sgRNA were used as wild type control. Cells grown in 10 cm petri dishes were transfected with indicated plasmids using 5 μl 1 mg/ml polyethylenimine (PEI, Sigma) per 1 μg of plasmids for HEK293T cells. Cells were cultured 24 h after transfection in medium containing 2 µg/ml puromycin for 48 h and then in normal growth media.
Statistics and reproducibility
Results are expressed as the mean ± SEM unless otherwise stated. Statistical analyses were performed in GraphPad Prism version 8.0 (GraphPad Software Inc.). Comparisons between the groups were made by unpaired two-tailed Student’s t test for two groups, or one-way ANOVA or two-way ANOVA followed by multiple comparisons test for more than two groups. P value < 0.05 was considered as statistically significant. No data were excluded from the analyses. Sample size was determined by the formula of the power analysis, N = 8(CV)^2[1 + (1-PC)^2]/(PC)^2, to reach the error = 0.05, Power = 0.80, percentage change in means (PC) = 20%, co-efficient of variation (CV) = 10–15% (varies between the experiments). The exact number of mice and cultures were indicated in the figure legend. All experiments were repeated at least twice and/or performed with several independent biological samples, and representative data are shown. Mice in each group were randomly chosen based on the age, genotype and gender. The Investigators were not blinded to allocation during experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Source data
Acknowledgements
We thank Drs. Carol Elias, Richard Wojcikiewicz and Martin Myers for reagents; Drs. Peter Arvan, Carol Elias and Daniel Klionsky for critical comments and suggestions, and members of the Qi and Arvan laboratories for comments and technical assistance. This work was supported by NIH grants 1R01DK137794, 1R35GM130292, 1R01DK120047, and American Diabetes Association Career Development (1-12-CD-04) and Innovative Basic Science (1-19-IBS-23) Awards (L.Q.).
Author contributions
H.M. and G.H.K. designed the most of experiments and H.M., with the help of G.H.K. and L.P., performed most of the experiments and data analysis. G.H.K. performed parabiosis. H.M., with the help of G.H.K., wrote the methods and figure legends. L.Q. and H.M. wrote the manuscript. All authors have approved the manuscript.
Peer review
Peer review information
Nature Communications thanks Qingchun Tong and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Data availability
The materials and reagents used are either commercially available or available upon the request, with detailed information included in Methods. The predicted structure of mLepRb is available at AlphaFold ID AF-P48356-F1 [https://alphafold.ebi.ac.uk/entry/P48356]. All data supporting the findings and materials for the manuscript are available within the article and the Supplementary Information. Source data are provided with this paper.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
The online version contains supplementary material available at 10.1038/s41467-024-52743-2.
References
- 1.Toda, C., Santoro, A., Kim, J. D. & Diano, S. POMC neurons: from birth to death. Annu. Rev. Physiol.79, 209–236 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.McLean, F. H. et al. A high-fat diet induces rapid changes in the mouse hypothalamic proteome. Nutr. Metab.16, 26 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dalvi, P. S. et al. High fat induces acute and chronic inflammation in the hypothalamus: effect of high-fat diet, palmitate and TNF-alpha on appetite-regulating NPY neurons. Int. J. Obes.41, 149–158 (2017). [DOI] [PubMed] [Google Scholar]
- 4.Beutler, L. R. et al. Obesity causes selective and long-lasting desensitization of AgRP neurons to dietary fat. Elife9, e55909 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Horvath, T. L. et al. Synaptic input organization of the melanocortin system predicts diet-induced hypothalamic reactive gliosis and obesity. Proc. Natl Acad. Sci. USA107, 14875–14880 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Souza, G. F. et al. Defective regulation of POMC precedes hypothalamic inflammation in diet-induced obesity. Sci. Rep.6, 29290 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Poon, K. Behavioral feeding circuit: dietary fat-induced effects of inflammatory mediators in the hypothalamus. Front Endocrinol.11, 591559 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Velloso, L. A. & Schwartz, M. W. Altered hypothalamic function in diet-induced obesity. Int. J. Obes.35, 1455–1465 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhang, K. & Kaufman, R. J. From endoplasmic-reticulum stress to the inflammatory response. Nature454, 455–462 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chaudhari, N., Talwar, P., Parimisetty, A., Lefebvre d’Hellencourt, C. & Ravanan, P. A molecular web: endoplasmic reticulum stress, inflammation, and oxidative stress. Front Cell Neurosci.8, 213 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Zhang, Y. et al. Synergistic mechanism between the endoplasmic reticulum and mitochondria and their crosstalk with other organelles. Cell Death Discov.9, 51 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Wu, H., Carvalho, P. & Voeltz, G. K. Here, there, and everywhere: the importance of ER membrane contact sites. Science361, eaan5835 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kornmann, B. et al. An ER-mitochondria tethering complex revealed by a synthetic biology screen. Science325, 477–481 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Rowland, A. A. & Voeltz, G. K. Endoplasmic reticulum-mitochondria contacts: function of the junction. Nat. Rev. Mol. Cell Biol.13, 607–625, (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Marchi, S., Patergnani, S. & Pinton, P. The endoplasmic reticulum-mitochondria connection: one touch, multiple functions. Biochim. Biophys. Acta1837, 461–469 (2014). [DOI] [PubMed] [Google Scholar]
- 16.Ozcan, L. et al. Endoplasmic reticulum stress plays a central role in development of leptin resistance. Cell Metab.9, 35–51 (2009). [DOI] [PubMed] [Google Scholar]
- 17.Purkayastha, S. et al. Neural dysregulation of peripheral insulin action and blood pressure by brain endoplasmic reticulum stress. Proc. Natl Acad. Sci. USA108, 2939–2944 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ramirez, S. & Claret, M. Hypothalamic ER stress: a bridge between leptin resistance and obesity. FEBS Lett.589, 1678–1687 (2015). [DOI] [PubMed] [Google Scholar]
- 19.Schneeberger, M. et al. Mitofusin 2 in POMC neurons connects ER stress with leptin resistance and energy imbalance. Cell155, 172–187 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ye, Z., Liu, G., Guo, J. & Su, Z. Hypothalamic endoplasmic reticulum stress as a key mediator of obesity-induced leptin resistance. Obes. Rev.19, 770–785 (2018). [DOI] [PubMed] [Google Scholar]
- 21.Zhang, X. et al. Hypothalamic IKKbeta/NF-kappaB and ER stress link overnutrition to energy imbalance and obesity. Cell135, 61–73 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Xiao, Y. et al. Knockout of inositol-requiring enzyme 1alpha in pro-opiomelanocortin neurons decreases fat mass via increasing energy expenditure. Open Biol.6, 160131 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Williams, K. W. et al. Xbp1s in Pomc neurons connects ER stress with energy balance and glucose homeostasis. Cell Metab.20, 471–482 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yao, T. et al. Ire1alpha in Pomc neurons is required for thermogenesis and glycemia. Diabetes66, 663–673 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Friedlander, R., Jarosch, E., Urban, J., Volkwein, C. & Sommer, T. A regulatory link between ER-associated protein degradation and the unfolded-protein response. Nat. Cell Biol.2, 379–384 (2000). [DOI] [PubMed] [Google Scholar]
- 26.Qi, L., Tsai, B. & Arvan, P. New insights into the physiological role of endoplasmic reticulum-associated degradation. Trends Cell Biol.27, 430–440 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Travers, K. J. et al. Functional and genomic analyses reveal an essential coordination between the unfolded protein response and ER-associated degradation. Cell101, 249–258 (2000). [DOI] [PubMed] [Google Scholar]
- 28.Hwang, J. & Qi, L. Quality control in the endoplasmic reticulum: crosstalk between ERAD and UPR pathways. Trends Biochem. Sci.43, 593–605 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Carvalho, P., Goder, V. & Rapoport, T. A. Distinct ubiquitin-ligase complexes define convergent pathways for the degradation of ER proteins. Cell126, 361–373 (2006). [DOI] [PubMed] [Google Scholar]
- 30.Gardner, R. G. et al. Endoplasmic reticulum degradation requires lumen to cytosol signaling. Transmembrane control of Hrd1p by Hrd3p. J. Cell Biol.151, 69–82 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hampton, R. Y., Gardner, R. G. & Rine, J. Role of 26S proteasome and HRD genes in the degradation of 3-hydroxy-3-methylglutaryl-CoA reductase, an integral endoplasmic reticulum membrane protein. Mol. Biol. Cell7, 2029–2044 (1996). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bhattacharya, A. & Qi, L. ER-associated degradation in health and disease—from substrate to organism. J. Cell Sci.132, jcs232850 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Brodsky, J. L. Cleaning up: ER-associated degradation to the rescue. Cell151, 1163–1167 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Christianson, J. C., Jarosch, E. & Sommer, T. Mechanisms of substrate processing during ER-associated protein degradation. Nat. Rev. Mol. Cell Biol.24, 777–796 (2023). [DOI] [PubMed] [Google Scholar]
- 35.Vashistha, N., Neal, S. E., Singh, A., Carroll, S. M. & Hampton, R. Y. Direct and essential function for Hrd3 in ER-associated degradation. Proc. Natl Acad. Sci. USA113, 5934–5939 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sun, S. et al. Sel1L is indispensable for mammalian endoplasmic reticulum-associated degradation, endoplasmic reticulum homeostasis, and survival. Proc. Natl Acad. Sci. USA111, E582–591 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lin, L. L. et al. SEL1L-HRD1 interaction is required to form a functional HRD1 ERAD complex. Nat. Commun.15, 1440 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Sha, H. et al. The ER-associated degradation adaptor protein Sel1L regulates LPL secretion and lipid metabolism. Cell Metab.20, 458–470 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wu, S. A. et al. The mechanisms to dispose of misfolded proteins in the endoplasmic reticulum of adipocytes. Nat. Commun.14, 3132 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Bhattacharya, A. et al. Hepatic Sel1L-Hrd1 ER-associated degradation (ERAD) manages FGF21 levels and systemic metabolism via CREBH. EMBO J.37, e99277 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wei, J. et al. HRD1-ERAD controls production of the hepatokine FGF21 through CREBH polyubiquitination. EMBO J.37, e98942 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kim, G. H. et al. Hypothalamic ER-associated degradation regulates POMC maturation, feeding, and age-associated obesity. J. Clin. Invest.128, 1125–1140 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Shi, G. et al. ER-associated degradation is required for vasopressin prohormone processing and systemic water homeostasis. J. Clin. Invest.127, 3897–3912 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Yoshida, S. et al. Endoplasmic reticulum-associated degradation is required for nephrin maturation and kidney glomerular filtration function. J. Clin. Invest.131, e143988 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Shrestha, N. et al. Sel1L-Hrd1 ER-associated degradation maintains beta cell identity via TGF-beta signaling. J. Clin. Invest.130, 3499–3510 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Liu, H., Du, T., Li, C. & Yang, G. STAT3 phosphorylation in central leptin resistance. Nutr. Metab.18, 39 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Baumann, H. et al. The full-length leptin receptor has signaling capabilities of interleukin 6-type cytokine receptors. Proc. Natl Acad. Sci. USA93, 8374–8378 (1996). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Chen, H. et al. Evidence that the diabetes gene encodes the leptin receptor: identification of a mutation in the leptin receptor gene in db/db mice. Cell84, 491–495 (1996). [DOI] [PubMed] [Google Scholar]
- 49.Uotani, S., Bjorbaek, C., Tornoe, J. & Flier, J. S. Functional properties of leptin receptor isoforms: internalization and degradation of leptin and ligand-induced receptor downregulation. Diabetes48, 279–286 (1999). [DOI] [PubMed] [Google Scholar]
- 50.Bumaschny, V. F. et al. Obesity-programmed mice are rescued by early genetic intervention. J. Clin. Invest122, 4203–4212 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Harris, R. B. Contribution made by parabiosis to the understanding of energy balance regulation. Biochim. Biophys. Acta1832, 1449–1455 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Harris, R. B., Zhou, J., Weigle, D. S. & Kuijper, J. L. Recombinant leptin exchanges between parabiosed mice but does not reach equilibrium. Am. J. Physiol.272, R1800–1808, (1997). [DOI] [PubMed] [Google Scholar]
- 53.Munzberg, H., Huo, L., Nillni, E. A., Hollenberg, A. N. & Bjorbaek, C. Role of signal transducer and activator of transcription 3 in regulation of hypothalamic proopiomelanocortin gene expression by leptin. Endocrinology144, 2121–2131 (2003). [DOI] [PubMed] [Google Scholar]
- 54.Saeed, S. et al. Genetic variants in LEP, LEPR, and MC4R explain 30% of severe obesity in children from a consanguineous population. Obesity23, 1687–1695 (2015). [DOI] [PubMed] [Google Scholar]
- 55.Saeed, S. et al. High morbidity and mortality in children with untreated congenital deficiency of leptin or its receptor. Cell Rep. Med.4, 101187 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Peelman, F., Zabeau, L., Moharana, K., Savvides, S. N. & Tavernier, J. 20 years of leptin: insights into signaling assemblies of the leptin receptor. J. Endocrinol.223, T9–23, (2014). [DOI] [PubMed] [Google Scholar]
- 57.Tsirigotaki, A. et al. Mechanism of receptor assembly via the pleiotropic adipokine Leptin. Nat. Struct. Mol. Biol.30, 551–563 (2023). [DOI] [PubMed] [Google Scholar]
- 58.Moharana, K. et al. Structural and mechanistic paradigm of leptin receptor activation revealed by complexes with wild-type and antagonist leptins. Structure22, 866–877 (2014). [DOI] [PubMed] [Google Scholar]
- 59.McNay, D. E., Pelling, M., Claxton, S., Guillemot, F. & Ang, S. L. Mash1 is required for generic and subtype differentiation of hypothalamic neuroendocrine cells. Mol. Endocrinol.20, 1623–1632 (2006). [DOI] [PubMed] [Google Scholar]
- 60.Padilla, S. L., Carmody, J. S. & Zeltser, L. M. Pomc-expressing progenitors give rise to antagonistic neuronal populations in hypothalamic feeding circuits. Nat. Med.16, 403–405 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yu, H., Rubinstein, M. & Low, M. J. Developmental single-cell transcriptomics of hypothalamic POMC neurons reveal the genetic trajectories of multiple neuropeptidergic phenotypes. Elife11, e72883 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Biglari, N. et al. Functionally distinct POMC-expressing neuron subpopulations in hypothalamus revealed by intersectional targeting. Nat. Neurosci.24, 913–929 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Padilla, S. L., Reef, D. & Zeltser, L. M. Defining POMC neurons using transgenic reagents: impact of transient Pomc expression in diverse immature neuronal populations. Endocrinology153, 1219–1231 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Balthasar, N. et al. Leptin receptor signaling in POMC neurons is required for normal body weight homeostasis. Neuron42, 983–991 (2004). [DOI] [PubMed] [Google Scholar]
- 65.Bell, B. B. et al. Differential contribution of POMC and AgRP neurons to the regulation of regional autonomic nerve activity by leptin. Mol. Metab.8, 1–12 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Caron, A. et al. POMC neurons expressing leptin receptors coordinate metabolic responses to fasting via suppression of leptin levels. Elife7, e33710 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhou, Z. et al. Endoplasmic reticulum-associated degradation regulates mitochondrial dynamics in brown adipocytes. Science368, 54–60 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wang, H. H. et al. Hypomorphic variants of SEL1L-HRD1 ER-associated degradation are associated with neurodevelopmental disorders. J. Clin. Invest.134, e170054 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Weis, D. et al. Biallelic Cys141Tyr variant of SEL1L is associated with neurodevelopmental disorders, agammaglobulinemia, and premature death. J. Clin. Invest.134, e170882 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Shrestha, N. et al. Integration of ER protein quality control mechanisms defines beta cell function and ER architecture. J. Clin. Invest.133, e163584 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Hosoi, T. et al. Endoplasmic reticulum stress induces leptin resistance. Mol. Pharm.74, 1610–1619 (2008). [DOI] [PubMed] [Google Scholar]
- 72.Heifetz, A., Keenan, R. W. & Elbein, A. D. Mechanism of action of tunicamycin on the UDP-GlcNAc:dolichyl-phosphate Glc-NAc-1-phosphate transferase. Biochemistry18, 2186–2192 (1979). [DOI] [PubMed] [Google Scholar]
- 73.Cowley, M. A. et al. Leptin activates anorexigenic POMC neurons through a neural network in the arcuate nucleus. Nature411, 480–484 (2001). [DOI] [PubMed] [Google Scholar]
- 74.Kamran, P. et al. Parabiosis in mice: a detailed protocol. J. Vis. Exp., 10.3791/50556 (2013). [DOI] [PMC free article] [PubMed]
- 75.Ji, Y. et al. Toll-like receptors TLR2 and TLR4 block the replication of pancreatic beta cells in diet-induced obesity. Nat. Immunol.20, 677–686 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Qi, L., Yang, L. & Chen, H. Detecting and quantitating physiological endoplasmic reticulum stress. Methods Enzymol.490, 137–146 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Yang, L. et al. A Phos-tag-based approach reveals the extent of physiological endoplasmic reticulum stress. PLoS ONE5, e11621 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The materials and reagents used are either commercially available or available upon the request, with detailed information included in Methods. The predicted structure of mLepRb is available at AlphaFold ID AF-P48356-F1 [https://alphafold.ebi.ac.uk/entry/P48356]. All data supporting the findings and materials for the manuscript are available within the article and the Supplementary Information. Source data are provided with this paper.