Abstract
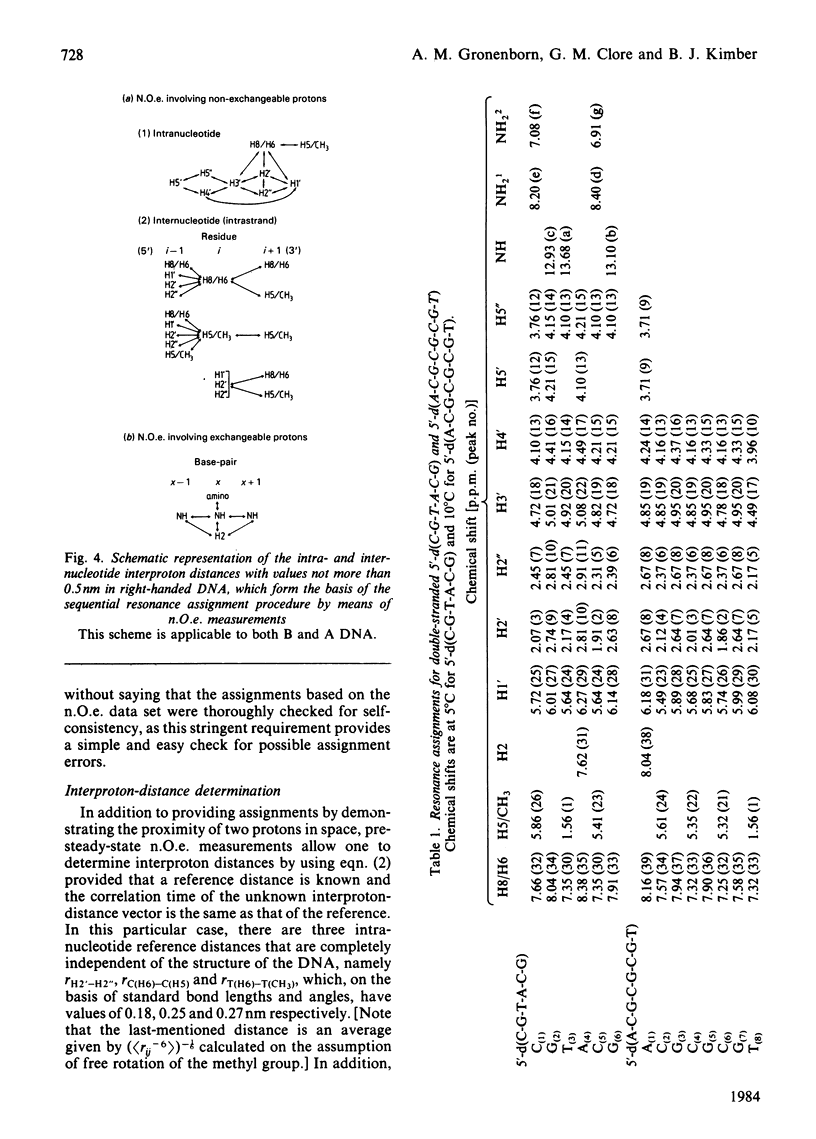
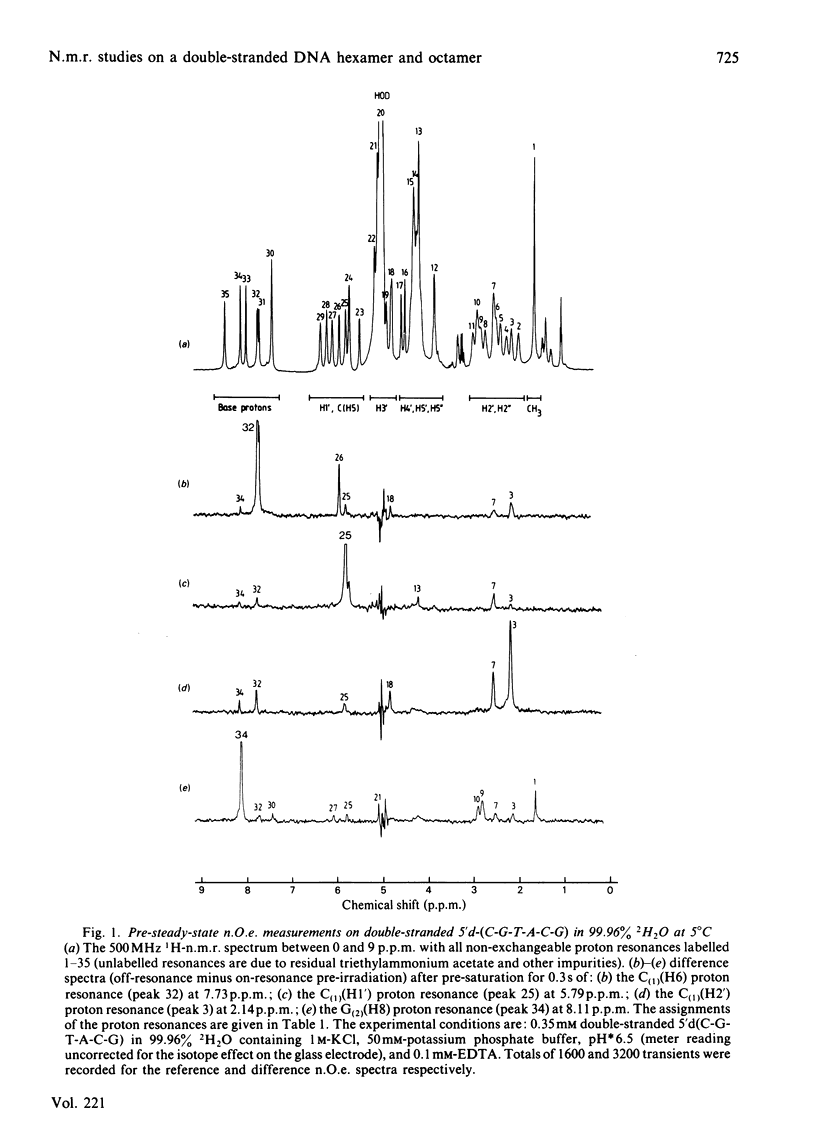
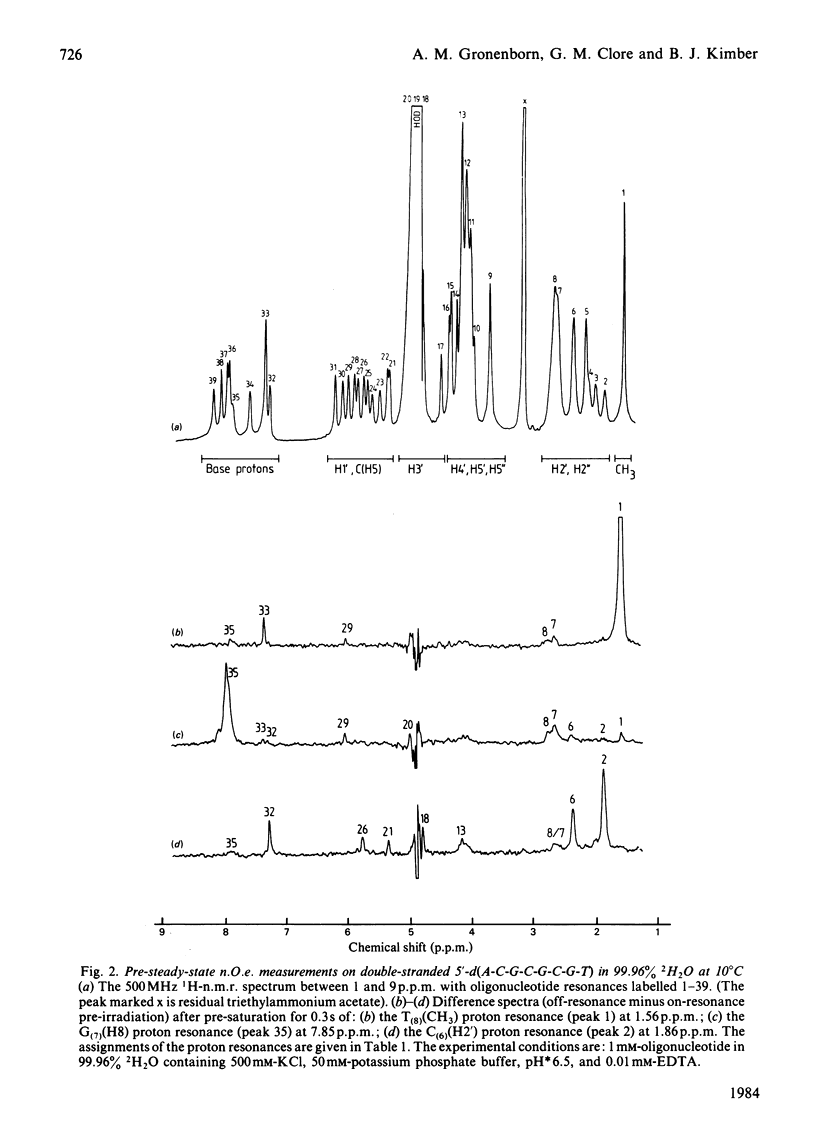
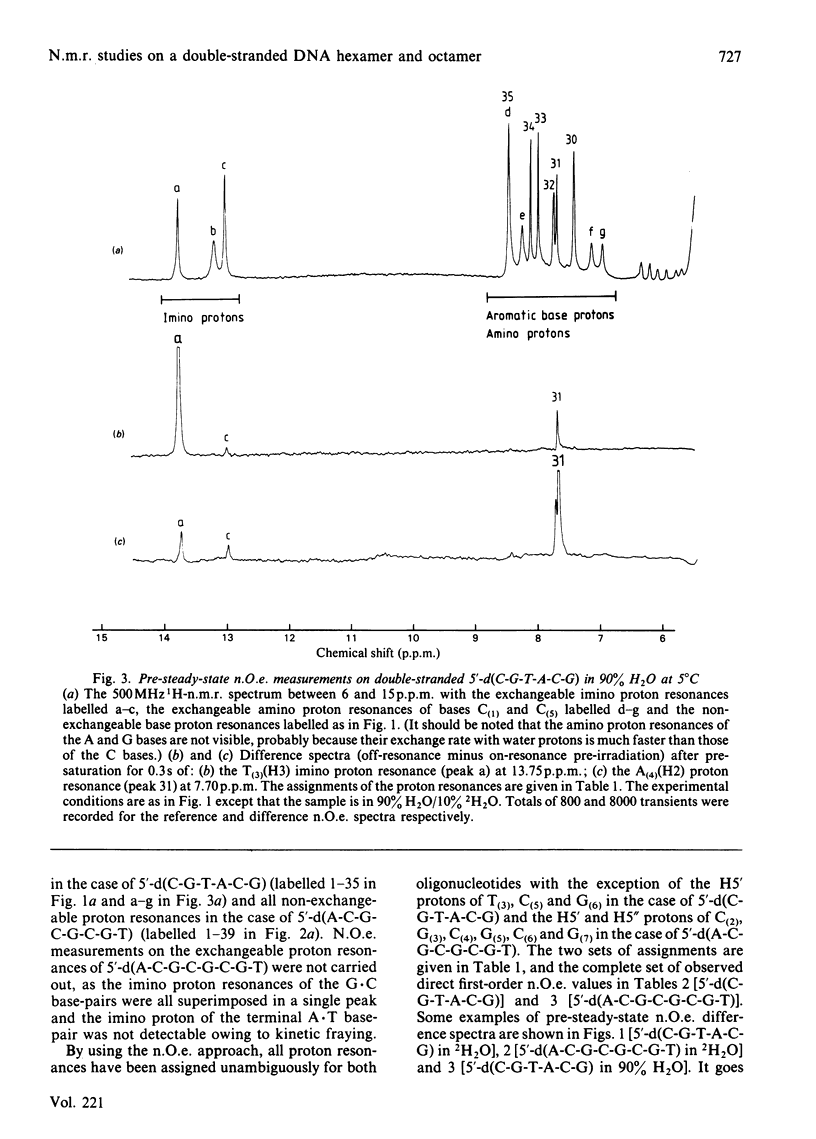
A 500 MHz 1H-n.m.r. study on two self-complementary alternating pyrimidine-purine oligodeoxyribonucleotides, 5'-d(C-G-T-A-C-G) and 5'-d(A-C-G-C-G-C-G-T), is presented. By using the proton-proton nuclear Overhauser effect virtually complete assignments are obtained and a large number of interproton distances [113 in the case of 5'-d(C-G-T-A-C-G) and 79 in the case of 5'-d(A-C-G-C-G-C-G-T)], both intra- and inter-nucleotide, are determined. The interproton-distance data are consistent with an overall right-handed B-DNA-type structure for both oligonucleotides, in agreement with their B-type c.d. spectra. However, whereas 5'-d(C-G-T-A-C-G) adopts a conventional B-type structure with a mononucleotide repeating unit, the interproton-distance data provide evidence that 5'-d(A-C-G-C-G-C-G-T) has a dinucleotide repeating unit consisting of alternation in glycosidic bond and sugar pucker conformations.
Full text
PDF










Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnott S., Chandrasekaran R., Birdsall D. L., Leslie A. G., Ratliff R. L. Left-handed DNA helices. Nature. 1980 Feb 21;283(5749):743–745. doi: 10.1038/283743a0. [DOI] [PubMed] [Google Scholar]
- Arnott S., Chandrasekaran R., Puigjaner L. C., Walker J. K., Hall I. H., Birdsall D. L., Ratliff R. L. Wrinkled DNA. Nucleic Acids Res. 1983 Mar 11;11(5):1457–1474. doi: 10.1093/nar/11.5.1457. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnott S., Hukins D. W. Optimised parameters for A-DNA and B-DNA. Biochem Biophys Res Commun. 1972 Jun 28;47(6):1504–1509. doi: 10.1016/0006-291X(72)90243-4. [DOI] [PubMed] [Google Scholar]
- Clore G. M., Gronenborn A. M. A nuclear-Overhauser-enhancement study of the solution structure of a double-stranded DNA undecamer comprising a portion of the specific target site for the cyclic-AMP-receptor protein in the gal operon. Sequential resonance assignment. Eur J Biochem. 1984 May 15;141(1):119–129. doi: 10.1111/j.1432-1033.1984.tb08166.x. [DOI] [PubMed] [Google Scholar]
- Clore G. M., Gronenborn A. M., Mitchinson C., Green N. M. 1H-NMR studies on nucleotide binding to the sarcoplasmic reticulum Ca2+ ATPase. Determination of the conformations of bound nucleotides by the measurement of proton-proton transferred nuclear Overhauser enhancements. Eur J Biochem. 1982 Nov;128(1):113–117. [PubMed] [Google Scholar]
- Clore G. M., Gronenborn A. M. Sequence-dependent structural variations in two right-handed alternating pyrimidine-purine DNA oligomers in solution determined by nuclear Overhauser enhancement measurements. EMBO J. 1983;2(12):2109–2115. doi: 10.1002/j.1460-2075.1983.tb01710.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R., Conner B. N., Wing R. M., Fratini A. V., Kopka M. L. The anatomy of A-, B-, and Z-DNA. Science. 1982 Apr 30;216(4545):475–485. doi: 10.1126/science.7071593. [DOI] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R. Structure of a B-DNA dodecamer. II. Influence of base sequence on helix structure. J Mol Biol. 1981 Jul 15;149(4):761–786. doi: 10.1016/0022-2836(81)90357-0. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Samson S., Dickerson R. E. Structure of a B-DNA dodecamer at 16 K. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4040–4044. doi: 10.1073/pnas.79.13.4040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drew H. R., Wing R. M., Takano T., Broka C., Tanaka S., Itakura K., Dickerson R. E. Structure of a B-DNA dodecamer: conformation and dynamics. Proc Natl Acad Sci U S A. 1981 Apr;78(4):2179–2183. doi: 10.1073/pnas.78.4.2179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gronenborn A. M., Clore G. M., Jeffery J. An unusual conformation of NAD+ bound to sorbitol dehydrogenase? A time-dependent transferred nuclear Overhauser effect study. J Mol Biol. 1984 Feb 5;172(4):559–572. doi: 10.1016/s0022-2836(84)80023-6. [DOI] [PubMed] [Google Scholar]
- Gronenborn A. M., Clore G. M., Jones M. B., Jiricny J. A nuclear Overhauser enhancement study on the imino proton resonances of a DNA pentadecamer comprising the specific target site of the cyclic AMP receptor protein in the ara BAD operon. FEBS Lett. 1984 Jan 9;165(2):216–222. doi: 10.1016/0014-5793(84)80172-6. [DOI] [PubMed] [Google Scholar]
- Gronenborn A. M., Clore G. M. Proton nuclear magnetic resonance studies on cyclic nucleotide binding to the Escherichia coli adenosine cyclic 3',5'-phosphate receptor protein. Biochemistry. 1982 Aug 17;21(17):4040–4048. doi: 10.1021/bi00260a020. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Reid B. R. Nuclear Overhauser assignment of the imino protons of the acceptor helix and the ribothymidine helix in the nuclear magnetic resonance spectrum of Escherichia coli isoleucine transfer ribonucleic acid: evidence for costacked helices in solution. Biochemistry. 1982 Oct 12;21(21):5129–5135. doi: 10.1021/bi00264a004. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Wemmer D. E., Chou S. H., Drobny G., Reid B. R. Assignment of the non-exchangeable proton resonances of d(C-G-C-G-A-A-T-T-C-G-C-G) using two-dimensional nuclear magnetic resonance methods. J Mol Biol. 1983 Dec 15;171(3):319–336. doi: 10.1016/0022-2836(83)90096-7. [DOI] [PubMed] [Google Scholar]
- Heerschap A., Haasnoot C. A., Hilbers C. W. Nuclear magnetic resonance studies on yeast tRNAPhe. III. Assignments of the iminoproton resonances of the tertiary structure by means of nuclear Overhauser effect experiments at 500 MHz. Nucleic Acids Res. 1983 Jul 11;11(13):4501–4520. doi: 10.1093/nar/11.13.4501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hogan M. E., Jardetzky O. Internal motions in deoxyribonucleic acid II. Biochemistry. 1980 Jul 22;19(15):3460–3468. doi: 10.1021/bi00556a009. [DOI] [PubMed] [Google Scholar]
- Klug A., Jack A., Viswamitra M. A., Kennard O., Shakked Z., Steitz T. A. A hypothesis on a specific sequence-dependent conformation of DNA and its relation to the binding of the lac-repressor protein. J Mol Biol. 1979 Jul 15;131(4):669–680. doi: 10.1016/0022-2836(79)90196-7. [DOI] [PubMed] [Google Scholar]
- Kuzmich S., Marky L. A., Jones R. A. Synthesis and physical characterization of the self-complementary, alternating pyrimidine/purine hexanucleotide d[CGTACG]. Nucleic Acids Res. 1982 Oct 25;10(20):6265–6271. doi: 10.1093/nar/10.20.6265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levitt M. Computer simulation of DNA double-helix dynamics. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 1):251–262. doi: 10.1101/sqb.1983.047.01.030. [DOI] [PubMed] [Google Scholar]
- Patel D. J. Antibiotic-DNA interactions: intermolecular nuclear Overhauser effects in the netropsin-d(C-G-C-G-A-A-T-T-C-G-C-G) complex in solution. Proc Natl Acad Sci U S A. 1982 Nov;79(21):6424–6428. doi: 10.1073/pnas.79.21.6424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel D. J., Kozlowski S. A., Bhatt R. Sequence dependence of base-pair stacking in right-handed DNA in solution: proton nuclear Overhauser effect NMR measurements. Proc Natl Acad Sci U S A. 1983 Jul;80(13):3908–3912. doi: 10.1073/pnas.80.13.3908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel D. J., Kozlowski S. A., Nordheim A., Rich A. Right-handed and left-handed DNA: studies of B- and Z-DNA by using proton nuclear Overhauser effect and P NMR. Proc Natl Acad Sci U S A. 1982 Mar;79(5):1413–1417. doi: 10.1073/pnas.79.5.1413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poulsen F. M., Hoch J. C., Dobson C. M. A structural study of the hydrophobic box region of lysozyme in solution using nuclear Overhauser effects. Biochemistry. 1980 Jun 10;19(12):2597–2607. doi: 10.1021/bi00553a011. [DOI] [PubMed] [Google Scholar]
- Redfield A. G., Gupta R. K. Pulsed NMR study of the structure of cytochrome c. Cold Spring Harb Symp Quant Biol. 1972;36:405–411. doi: 10.1101/sqb.1972.036.01.052. [DOI] [PubMed] [Google Scholar]
- Reid D. G., Salisbury S. A., Bellard S., Shakked Z., Williams D. H. Proton nuclear Overhauser effect study of the structure of a deoxyoligonucleotide duplex in aqueous solution. Biochemistry. 1983 Apr 12;22(8):2019–2025. doi: 10.1021/bi00277a044. [DOI] [PubMed] [Google Scholar]
- Roy S., Redfield A. G. Assignment of imino proton spectra of yeast phenylalanine transfer ribonucleic acid. Biochemistry. 1983 Mar 15;22(6):1386–1390. doi: 10.1021/bi00275a010. [DOI] [PubMed] [Google Scholar]
- Shakked Z., Rabinovich D., Kennard O., Cruse W. B., Salisbury S. A., Viswamitra M. A. Sequence-dependent conformation of an A-DNA double helix. The crystal structure of the octamer d(G-G-T-A-T-A-C-C). J Mol Biol. 1983 May 15;166(2):183–201. doi: 10.1016/s0022-2836(83)80005-9. [DOI] [PubMed] [Google Scholar]
- Tidor B., Irikura K. K., Brooks B. R., Karplus M. Dynamics of DNA oligomers. J Biomol Struct Dyn. 1983 Oct;1(1):231–252. doi: 10.1080/07391102.1983.10507437. [DOI] [PubMed] [Google Scholar]
- Wagner G., Wüthrich K. Sequential resonance assignments in protein 1H nuclear magnetic resonance spectra. Basic pancreatic trypsin inhibitor. J Mol Biol. 1982 Mar 5;155(3):347–366. doi: 10.1016/0022-2836(82)90009-2. [DOI] [PubMed] [Google Scholar]
- Weiss M. A., Patel D. J., Sauer R. T., Karplus M. Two-dimensional 1H NMR study of the lambda operator site OL1: a sequential assignment strategy and its application. Proc Natl Acad Sci U S A. 1984 Jan;81(1):130–134. doi: 10.1073/pnas.81.1.130. [DOI] [PMC free article] [PubMed] [Google Scholar]



