Abstract
Objective
Arrhythmia detection and classification are challenging because of the imbalanced ratio of normal heartbeats to arrhythmia heartbeats and the complicated combinations of arrhythmia types. Arrhythmia classification on wearable electrocardiogram monitoring devices poses a further unique challenge: unlike clinically used electrocardiogram monitoring devices, the environments in which wearable devices are deployed are drastically different from the carefully controlled clinical environment, leading to significantly more noise, thus making arrhythmia classification more difficult.
Methods
We propose a novel hierarchical model based on CNN+BiLSTM with Attention to arrhythmia detection, consisting of a binary classification module between normal and arrhythmia heartbeats and a multi-label classification module for classifying arrhythmia events across combinations of beat and rhythm arrhythmia types. We evaluate our method on our proprietary dataset and compare it with various baselines, including CNN+BiGRU with Attention, ConViT, EfficientNet, and ResNet, as well as previous state-of-the-art frameworks.
Results
Our model outperforms existing baselines on the proprietary dataset, resulting in an average accuracy, F1-score, and AUC score of 95%, 0.838, 0.906 for binary classification, and 88%, 0.736, 0.875 for multi-label classification.
Conclusions
Our results validate the ability of our model to detect and classify real-world arrhythmia. Our framework could revolutionize arrhythmia diagnosis by reducing the burden on cardiologists, providing more personalized treatment, and achieving emergency intervention of patients by allowing real-time monitoring of arrhythmia occurrence.
Keywords: Machine learning, deep learning, electrocardiogram, wearable device, multi-label classification
Introduction
With the rapid development of wearable medical devices,1–3 electrocardiogram (ECG) monitoring is accessible to patients and doctors through wearable devices. A wearable ECG monitoring device is a crucial improvement for detection and classification since arrhythmia occurs rarely and sporadically. Due to limited time and a controlled environment, conventional ECG monitoring devices may never record arrhythmia heartbeats in a clinical setting. Thus, wearable ECG monitoring enables longer monitoring time and a more natural environment for the patients, increasing the probability of detecting arrhythmia events. However, wearable ECG monitoring devices pose several challenges. First, since the patient always wears the device and does different activities in various environments, the heartbeats recorded can be very noisy. This issue does not exist in the clinical ECG monitoring setting because the patient’s activities are limited, and the environment is carefully controlled to minimize the noise. Another challenge of wearable ECG monitoring devices is that significantly more data is produced due to prolonged monitoring duration. It becomes more labor-intensive for cardiologists to identify arrhythmia. As the adaptation of wearable devices increases, the workload will also increase, making arrhythmia detection and classification prohibitively expensive.
Additionally, arrhythmias have rhythm and beat types. Beat arrhythmias such as ventricular premature contraction (VPC) or atrial premature complexes (APCs) are not immediately life-threatening. Still, management or treatment is required if the frequency of occurrence of these beat arrhythmias increases significantly. In addition, observing the frequency of beat arrhythmias after heart-related (even if not necessarily heart-related) surgery, procedure, or treatment is essential in prognosis. In particular, when the frequency of beat arrhythmias such as VPC increases significantly, the probability of rhythm arrhythmias such as ventricular tachycardia (VT) also increases. Thus, detecting beat and rhythm arrhythmia together is clinically effective and especially suitable for ECG monitoring.
Moreover, detecting rhythm or beats alone does not capture beat arrhythmias occurring during rhythm arrhythmias. During atrial rhythm arrhythmias such as atrial fibrillation (AF) or atrial tachycardia (AT), ventricular beat arrhythmia such as VPC can occur. Conversely, atrial beat arrhythmia such as APC can occur during rhythm ventricular arrhythmias like VT. For clinicians, detecting all arrhythmia types gives a more comprehensive understanding of the patient’s heart condition and the characteristics of the beats and rhythms. Using this knowledge, clinicians can offer more personalized and effective treatment for the patient. Therefore, a multi-labeled approach to arrhythmia classification must enhance its practicality for clinical use.
To address the significant challenges in arrhythmia detection and classification using ECG data while providing more clinical usability, we explore deep-learning models that aim to adapt to the noise and automatically detect and classify multi-labeled arrhythmia heartbeats and rhythms. We propose a hierarchical model consisting of a binary classification module for detecting arrhythmia from normal heartbeats and a multi-label classification model for classifying arrhythmia heartbeats into various combinations of different arrhythmia anomalies. An illustration of our framework is shown in Figure 1. The binary classification module can notify doctors of potential arrhythmia patients, and the multi-label classification module can then provide information on the specific arrhythmia types, thus streamlining the decision process for the doctors and patients. We test our models with the hierarchical architecture on real-world ECG data collected by MEZOO’s wearable ECG monitoring device 4 and demonstrate high performances, proving the feasibility of automated arrhythmia detection and classification using our proposed method.
Figure 1.
(A)–(D) A flowchart of the proposed framework. (C) and (D) Proposed concepts of a hierarchical approach. (A) Multi-labeled wireless ECG arrhythmia raw data, (B) four-beat input data after preprocessing, (C) binary classification model for normal heartbeat and arrhythmia classification, (D) multi-class, multi-label arrhythmia classification model, (E) detailed structure of the proposed CNN+BiLSTM with attention model. ECG: electrocardiogram; CNN: convolutional neural network; BiLSTM: bidirectional long short-term memory.
Related works
Cardiac arrhythmia is a critical medical condition that requires timely diagnosis and intervention. ECG signals for arrhythmia detection have gained significant attention due to their non-invasive nature and high diagnostic accuracy. Several studies have explored the application of machine learning and deep learning techniques for classifying cardiac arrhythmia.
There have been significant advances in arrhythmia detection using deep learning over the past 7 years.5–12 Most of these were classification studies using Physionet’s MIT-BIH dataset.5,6 Although it is a well-organized dataset used as a benchmark by research groups, it has limitations. It has been a very long time since the experiment was conducted, and decades have passed since it was published. Recent state-of-the-art studies are reporting very high accuracy of over 99% in binary and multiclass arrhythmia classification using MIT-BIH dataset.7–9,13 For example, ResNet18 and EfficientNet-V2 have achieved high accuracy on the MIT-BIH dataset,10,11 but their performance metrics significantly decrease on the wearable ECG data. Additionally, MIT-BIH’s dataset only has ECG records from 47 people, and some specific arrhythmia types do not exist enough for the deep learning models to capture the complex relationships. These biases lead to difficulties in developing generalized algorithms for arrhythmia detection through deep learning.
One representative dataset other than the MIT-BIH is PhysioNet/Computing in Cardiology Challenge 2017 (CinC2017).14,5 It provides single-lead ECG data (AliveCor devices) from 8528 subjects with four types of heart rhythms (AF, normal, other rhythms, and noise). One study that achieved the highest results in the competition recorded 79% of the F1-score by using Resnet. 15 Further study introduced a 21-layer recurrent 1D CNN called RhythmNet and achieved 82% of score. 16 CinC2017 provides many ECG records but has the disadvantage that only AF is labeled among many types of arrhythmia.
One similar dataset to CinC2017 is the China Physiological Signal Challenge 2018 (CPSC2018). 17 It includes eight types of arrhythmia: AF, first-degree atrioventricular block (I-AVB), left bundle branch block (LBBB), Right bundle branch block (RBBB), premature atrial contraction (PAC), premature ventricular contraction (PVC), ST-segment depression (STD), and ST-segment elevated (STE). The training set contains 6877 (female 3178; male 3699) ECG recordings, and the test set contains 2954 ECG recordings. Twelve leads ECG recordings lasting from 6 to 60 seconds. The team that ranked first in CPSC2018 introduced the CNN+bidirectional RNN model and achieved an F1-score of 0.84. 18 The CPSC2018 dataset has the advantage of covering many types of arrhythmias and including many patients. However, it is a 12-lead ECG, which requires the patient to visit the clinic in person, attach electrodes, and examine it with the help of clinicians. Therefore, there are limitations in that it can only be acquired in a limited hospital environment, which is in a natural setting.
Recently, multi-class arrhythmia on wearable ECG data has been studied.19,20 In 2019, a 0.97 AUC score was achieved using a mobile ECG device called the Zio monitor with a 34-layer CNN model. 19 However, the work is limited to multi-class (not multi-label) classification, solely detecting rhythm arrhythmia types (without incorporating beat arrhythmia types) and producing predictions every 30 seconds. In comparison, our work focuses on the setting where multi-label arrhythmia classification on both rhythm arrhythmia and beat arrhythmia are required for the machine learning model.
Methods and materials
The nature of this study is to propose a novel machine-learning framework that achieves state-of-the-art performance on the proprietary dataset in a real-world setting. The dataset is used solely to benchmark rather than analyze the contents of the dataset.
Clinical data
MEZOO collected human subject data under the oversight and approval of the Institutional Review Board (IRB). The clinical data is received from MEZOO’s local clinic and Kosin University. MEZOO’s local clinic recruited 125 patients, 69 female and 56 male, with an average age of 51.20 and a standard deviation of 17.83. Data was obtained with the consent of individuals and provided by MEZOO after de-identification. Kosin University recruited 67 patients, 25 female and 42 male, with an average age of 59.09 and a standard deviation of 15.28. The study protocol was approved by the Ethics Committee of Kosin University Gospel Hospital (IRB No. 2022-08-029). The written informed consent was obtained from the participants prior to study initiation, and data was deidentified and anonymized. There are 192 patients, 94 female and 98 male, with an average age of 53.95 and a standard deviation of 17.36. The Rice University IRB reviewed the data used, and this activity was determined to be not human subjects research.
Data collection
In the first part of the data collection process, the data acquisition phase, the following steps were taken: (1) counseling, (2) completion of consent forms and attachment of wearable patch-type ECG, and (3) recording of ECG data. Physicians in private hospitals conducted counseling, while Holter lab technicians completed consent forms and equipment attachments. Completing the consent forms and attaching the equipment typically took < 10 minutes, and the recording of ECG data was averaged over 13 hours.
The experiment was conducted in 11 private hospitals (Roh Tae-Ho Paulo Internal Medicine, Baro Internal Medicine, etc.), and involved a total of 125 cases from 2 August 2021 to 18 October 2021. After the Hicardi+ equipment was attached at each hospital, participants returned home and recorded their ECG data during daily activities. The measurement was completed within 24 hours, after which the equipment was returned to the hospital.
Additionally, data were collected at Kosin University Hospital for outpatients who received a Holter prescription from the Department of Cardiology. This phase involved simultaneously attaching HiCardi+ and GE SEER Holter products to collect data over a 24-hour period, resulting in a total of 67 cases from 8 February 2023 to 30 June 2023.
In the second part, the data interpretation phase, the following steps were taken: (4) confirmation of cloud file creation, (5) initial ECG data reading, and (6) final ECG data interpretation. The Holter lab technicians confirmed cloud file creation and conducted the initial ECG data reading. Two cardiologists performed the final ECG data interpretation through the cloud-based monitoring server.
ECG monitoring device
Hicardi (MEZOO Co., Ltd., Wonju, Gangwon, Republic of Korea) is an 8 g, mm (without disposable electrode) wearable ECG monitoring patch device certified as a medical device by the Ministry of Food and Drug Safety of Korea (KFDA). This wearable device monitors and records ECG, respiration, skin surface temperature, and activity for up to 16 hours. The ECG signal is recorded with a 250 Hz sampling frequency and a 14-bit resolution. The data from the wearable patch is transferred through Bluetooth Low Energy (BLE) to a mobile gateway implemented as a smartphone application. The mobile gateway transmits the data to a cloud-based monitoring server. The cloud-based server, named “Livestudio,” enables medical staff to monitor patient data in real-time, view the transmitted ECG data files, click on the data to view and interpret the ECG data, and ultimately generate a final ECG report.
Pre-processing
The ECG data was received in .mat format. Each .mat file represents a patient. Each .mat file has seven keys, described in Table 1. Table 2 shows the detailed rhythm labels contained in the .mat files.
Table 1.
Description of the content of each .mat file.
| Keys | Description | Values |
|---|---|---|
| LeadOff | Records if the lead is in the correct position | An array of 0’s and 1’s, 1 meaning the lead is off, |
| to receive accurate ECG or not | 0 meaning the lead is not off | |
| Rpk_label | Labels the position of the R-peak | An array of 0’s and 1’s, 1 meaning the R-peak, |
| 0 meaning not the R-peak | ||
| Rthm_label | Lists all the arrhythmia types and their | A 2D-array of size 27, with 27 rhythm labels and |
| corresponding number in | their corresponding number | |
| dECG | The ECG data recorded | An array recorded at 250 Hz |
| data_lost | Records if the data is lost or not | An array of 0’s and 1’s, 1 meaning the data is lost, |
| one meaning the data is not lost | ||
| final_flag | The arrhythmia labels | A 2D-array of size (x, 27), where x is the length of dECG |
| fs | The recording frequency in Hz | 250 Hz |
2D: two-dimensional; ECG: electrocardiogram; dECG: Diploma in ECG Technology.
Table 2.
Description of the Rpk_label in each .mat file.
| Rpk_label | Label name | Label abbreviation |
|---|---|---|
| 0 | Normal | Normal |
| 1 | Pause | Pause |
| 2 | Ventricular Fib./Tach. | VFibTach. |
| 3 | Ventricular Premature Contraction | VPC |
| 4 | Ventricular Tachycardia | VTach |
| 5 | Ventricular Bigeminy | Bigem |
| 6 | Ventricular Trigeminy | Trigem |
| 7 | VPC Couplet | Couplet |
| 8 | Bradycardia | Brady |
| 9 | Atrial Tachycardia | ATach |
| 10 | Supraventricular Tachycardia | SVT |
| 11 | Paced Rhythm | Paced |
| 12 | Atrial Fibrillation/Flutter | AFib |
| 13 | Atrial Flutter | AFlut |
| 14 | Atrial Premature Contraction | APC |
| 15 | Run VPCs | VPCs |
| 16 | Sinus Tachycardia | STach |
| 20 | Block | Block |
| 21 | Ventricular Ectopic Beat | VEB |
| 22 | Supraventricular Ectopic Beat | SEB |
| 23 | Run APCs | APCs |
| 95 | Main-Rhythm | Main |
| 96 | Data-loss | loss |
| 97 | Artifact | Artifact |
| 98 | Lead-Off | LeadOff |
| 99 | In-progress | Inprogress |
| 100 | Unknown | Unknown |
VPC: ventricular premature contraction; APC: atrial premature complex.
In pre-processing, the ECG data, the Rpk_label, which indicates the location of the R-peak, is first used to locate each heartbeat. The midpoint points between R-peaks separate the heartbeats. Then, each of four consecutive heartbeats (with no overlapping) is taken as one input. Inputs that contains or are deleted. Inputs that contain are also removed since they correspond to the illegal flags of “Artifact,” “Lead-Off,” “In-progress,” and “Unknown.” Additionally, inputs with “NAN” values are removed. To standardize each input without losing temporal details within the four beats since heartbeat duration varies, we interpolate each input to 1024 samples. The reason for 1024 is that a beat’s average length is 228 samples. To maximize detail, minimize artificial interpolation, and facilitate input for model construction, 256 was chosen as the average heartbeat length, thus 1024 for four heartbeats. Then, z-score standardization is performed on each input individually. A label vector that represents arrhythmia types with 27 entries accompanies every input. Figure 2 shows an example of graphical pre-processing.
Figure 2.
Pre-processing procedure, including segmentation, resampling and standardization with example electrocardiogram (ECG) signal.
Hierarchical deep learning
We propose a novel hierarchical deep-learning approach to arrhythmia detection and classification. The first stage of our approach is detection, which involves a model performing binary classification between normal heartbeats and arrhythmia heartbeats. In the second stage, another model performs multi-label arrhythmia classification between different arrhythmia types and combinations, where potential arrhythmia types and combinations are generated for each input signal.
For binary classification, each input’s arrhythmia type label vector is simplified to a binary label representing arrhythmia anomaly. A total of of the data is normal heartbeats, and is arrhythmia heartbeats.
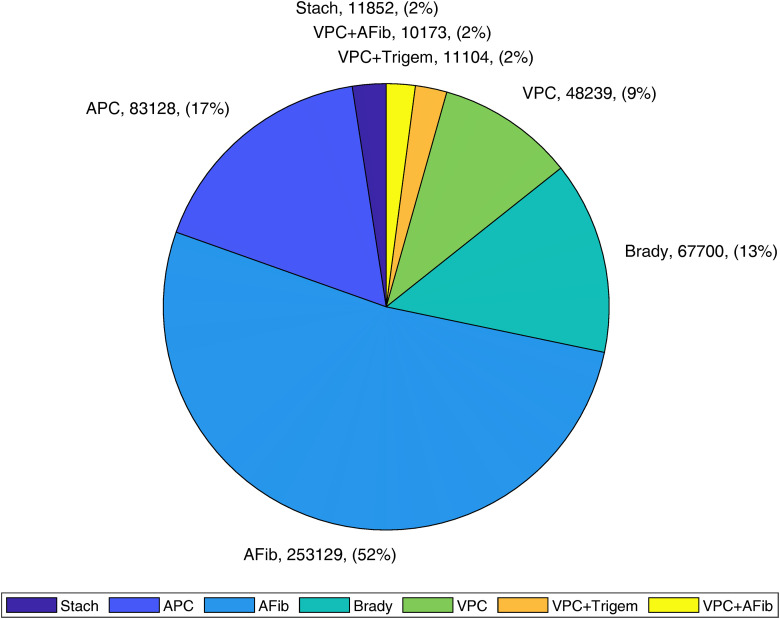
For multi-label classification, since there is limited data resulting in multi-label classification with only a few samples, multi-label classification is a significantly more challenging task than binary classification; we took the top seven multi-label classes as the target classes. The entire seven multi-label classes are sinus tachycardia, atrial premature contraction, atrial fibrillation/flutter, bradycardia, ventricular premature contraction, ventricular premature contraction and ventricular trigeminy, ventricular premature contraction and atrial fibrillation/flutter. The detailed distribution is shown in Figure 3.
Figure 3.
Distribution of top seven arrhythmia beat combinations.
Proposed model
CNN+BiLSTM with attention is a model with CNN layers in the first half, then bidirectional LSTM and attention layers in the second half. 21 The CNN component extracts local spatial features from the ECG signal. It comprises one or more convolutional layers, followed by pooling layers. These layers can capture local patterns and variations in the ECG signal’s amplitude and morphology. The BiLSTM layer is a recurrent neural network (RNN) type that can capture temporal dependencies and sequential patterns in the ECG signal. Bidirectional means it processes the input sequence in both forward and backward directions, allowing it to capture context from past and future time steps simultaneously. This is crucial for understanding the temporal dynamics of ECG signals, often characterized by complex rhythms and patterns. The attention mechanism is integrated into the model to dynamically weigh and focus on different parts of the input sequence. This helps the model give more attention to relevant segments of the ECG signal while ignoring noisy or less informative regions. Attention mechanisms are beneficial when dealing with long sequences like ECG data. The input ECG signal, a time series of voltage values, is fed into the model.
Our model is built based on the CNN+BiLSTM model proposed by Cheng et al. 22 The model includes three CNN blocks, three BiLSTM blocks, and three feedforward layers. Layer normalization and dropouts are added to all blocks to improve generalization. Kernel regularizer, bias regularizer, and activity regularizer are added to CNN blocks and feedforward layers. The activation function for CNN blocks and feedforward layers is Mish. 23 Additionally, the attention mechanism is added to each BiLSTM block. The learning rate is 0.0005, and the batch size is 64.
Statistical analysis
In this study, a statistical analysis and comparative study between patients and the control group is not applicable. Instead, the focus was on performance comparison by the various machine learning models and approaches on the proprietary dataset. Demographic data of the participants, including gender and age, are detailed in the “Methods and Materials” section. Figure 3 presents information regarding the types of arrhythmia data utilized.
Results
Evaluation metric
We use weighted accuracy, macro average F1-score, and macro average area under the curve (AUC) score to evaluate the performance of the five deep learning models experimented with, averaging over the six distinct singular arrhythmia types. These metrics give us an accurate insight into the model performance in this heavily imbalanced classification task.
Binary classification for arrhythmia detection
The preprocessed wearable ECG data used for binary classification includes all arrhythmia types and combinations from the data, where binary labels identifying the arrhythmia heartbeats are used in the classification. We used the CNN+BiLSTM with attention model to train the binary classification task between normal heartbeats and arrhythmia heartbeat by modifying the output layer to 1 unit. A five-fold cross-validation train-test split on the patient level was performed. For each split, of the patient’s data will be used for training, and the remaining for testing. The random seed for the five-fold split is set to . We used binary focal loss and class weights similar to the multi-label classification experiments to address the data imbalances. Additionally, due to the large size of the dataset, the number of normal heartbeats was randomly downsampled to 2 times the number of arrhythmia heartbeats, resulting in the distribution of the training dataset to be normal heartbeats and arrhythmia heartbeats for each cross-validation fold.
The results of the binary classification experiment are given in Table 3, where we demonstrate the weighted accuracy, macro F1-score, and macro AUC score of CNN+BiLSTM with attention in 15 runs across three random splits and five folds each. This method achieves an average weighted accuracy of , an average F1-score of , and an average AUC score of .
Table 3.
Weighted accuracy, macro F1-score, and macro AUC results of CNN+BiLSTM with attention on binary classification.
| Accuracy | F1-score | AUC | |
|---|---|---|---|
| seed0 | 0.951 0.02 | 0.859 0.06 | 0.914 0.04 |
| seed1 | 0.949 0.01 | 0.843 0.05 | 0.912 0.03 |
| seed2 | 0.949 0.02 | 0.813 0.06 | 0.893 0.04 |
| Overall | 0.95 0.02 | 0.838 0.06 | 0.906 0.04 |
AUC: area under the curve; CNN: convolutional neural network; BiLSTM: bidirectional long short-term memory.
Multi-label arrhythmia type classification
The preprocessed wearable ECG data used for multi-label classification includes the top seven most frequent combinations of arrhythmia types from the data. The input to the models is consecutive four heartbeats of shape (1024, 1) with no overlapping between inputs. Each input has a multi-hot encoded label of size 6 (the number of unique arrhythmia types). We do a five-fold cross-validation train-test split on the patient level for each deep learning method, including CNN+BiLSTM with attention, CNN+BiGRU with attention, ConViT, 24 EfficientNet, 25 and ResNet. 26 Therefore, for each split, 80% of the patient’s data will be used for training, and the remaining for testing. The random seed for the five-fold split is set to so that all methods will be trained and tested on the same five splits and repeated three times. The Adam optimizer is used to optimize all deep learning methods. Moreover, each method is fine-tuned on our dataset among the hyperparameters, including learning rate, batch size, kernel, or filter size, regularization, dropout rate, and activation functions listed in the “Method” section. To address the data imbalances across different arrhythmia types, we used binary focal loss for the multi-label problem, which has succeeded in many deep-learning problems. 27
The results of the multi-label classification experiment are shown in Table 4, where we demonstrate the weighted accuracy, macro F1-score, and macro AUC score of each model in three repetitive five-fold experiments. Overall, CNN+BiLSTM with attention has the highest average weighted accuracy, macro F1-score, and macro AUC score of all the models while having a lower standard deviation. CNN+BiLSTM with attention has high weighted accuracy and macro AUC score for the task, while the macro F1-score is not on par.
Table 4.
Experimental result of CNN+BiLSTM with attention, CNN+BiGRU with attention, ConViT, EfficientNet, and ResNet on multi-label classification between arrhythmia types and combinations.
| (a) Comparison of average weighted accuracy among the five deep learning methods across three cross-validation folds. The bolded number represents the best value for each seed. | |||||
|---|---|---|---|---|---|
| Accuracy | CNN+BiLSTM with attention | CNN+BiGRU with attention | ConViT | EfficientNetV2B0 | ResNet |
| Seed0 | 0.875 0.03 | 0.813 0.08 | 0.807 0.05 | 0.807 0.05 | 0.863 0.05 |
| Seed1 | 0.872 0.05 | 0.828 0.09 | 0.806 0.10 | 0.800 0.15 | 0.836 0.13 |
| Seed2 | 0.893 0.04 | 0.855 0.08 | 0.798 0.07 | 0.874 0.10 | 0.906 0.08 |
| Overall | 0.880 0.04 | 0.832 0.08 | 0.803 0.07 | 0.827 0.1 | 0.869 0.08 |
| (b) Comparison of macro average F1-score between the five different deep learning methods across three cross-validation folds. The bolded number represents the best value for each fold. | |||||
| F1-score | CNN+BiLSTM with attention | CNN+BiGRU with attention | ConViT | EfficientNetV2B0 | ResNet |
| Seed0 | 0.748 0.08 | 0.700 0.06 | 0.626 0.05 | 0.719 0.07 | 0.696 0.13 |
| Seed1 | 0.733 0.06 | 0.693 0.09 | 0.643 0.11 | 0.717 0.1 | 0.729 0.06 |
| Seed2 | 0.727 0.07 | 0.714 0.08 | 0.600 0.09 | 0.704 0.07 | 0.703 0.09 |
| Overall | 0.736 0.07 | 0.702 0.08 | 0.623 0.09 | 0.713 0.07 | 0.709 0.09 |
| (c) Comparison of macro average AUC score between the five different deep learning methods across three cross-validation folds. The bolded number represents the best value for each fold. | |||||
| AUC score | CNN+BiLSTM with attention | CNN+BiGRU with attention | ConViT | EfficientNetV2B0 | ResNet |
| Seed0 | 0.883 0.02 | 0.866 0.02 | 0.812 0.01 | 0.868 0.01 | 0.859 0.04 |
| Seed1 | 0.856 0.02 | 0.841 0.04 | 0.805 0.05 | 0.843 0.04 | 0.848 0.04 |
| Seed2 | 0.887 0.03 | 0.867 0.03 | 0.810 0.04 | 0.861 0.03 | 0.875 0.04 |
| Overall | 0.875 0.03 | 0.858 0.03 | 0.809 0.04 | 0.857 0.03 | 0.861 0.04 |
AUC: area under the curve; CNN: convolutional neural network; BiLSTM: bidirectional long short-term memory; BiGRU: bidirectional gated recurrent unit.
Additionally, we examined the sensitivity and specificity, as well as the confusion matrix of one fold out of the five-fold cross-validations we conducted, as shown in Table 5 and Figure 4. As we can see, the model generally performs well. The performance of Trigeminy is not very high, and the model will sometimes miss classifying multi-labeled classes by missing one class or including an additional class. One major reason is that even with the focal loss to address the imbalance issue, the data imbalances are too significant, and as we get more data from MEZOO, this issue can be alleviated.
Table 5.
Sensitivity and specificity result for one fold out of the five-fold cross-validation for each arrhythmia class of our framework on our real-world ECG dataset.
| Class | Sensitivity | Specificity |
|---|---|---|
| AFib | 0.97 | 0.95 |
| APC | 0.77 | 0.97 |
| Brady | 0.95 | 0.99 |
| VPC | 0.76 | 0.95 |
| STach | 0.99 | 1.00 |
| Trigem | 0.24 | 1.00 |
ECG: electrocardiogram; VPC: ven- tricular premature contraction; APC: atrial premature complex.
Figure 4.
Confusion matrix of one fold out of the five-fold cross-validation.
Benchmark our framework on the MIT-BIH database
As the MIT-BIH database is a commonly used benchmark for arrhythmia classification,28–31 we trained our model on the MIT-BIH database for multiclass classification of five classes: normal, VPC, ventricular escape, a fusion of ventricular and normal, and unclassified heartbeats. The preprocessing step involves the standard wavelet denoising the ECG records and standardizing using z-score normalization. After splitting the training and testing sets, we resample the training set to balance each of the five classes. A five-fold cross-validation is conducted, and the performance is measured in accuracy, sensitivity, specificity, and F1-score for us to compare with previous work.
The results for benchmarking our framework on the MIT-BIH database across five-fold cross-validation are 0.997 accuracy, 0.992 sensitivity, 0.998 specificity, and 0.993 F1-score. A per-class accuracy, sensitivity, specificity, and F1-score report are shown in Table 6 for one fold out of the five-fold cross-validation. The performance of our framework is similar to or better than previous state-of-the-art models,28–31 compared in Table 7.
Table 6.
Experimental result for one fold out of the five-fold cross-validation for each arrhythmia class of our framework on the MIT-BIH dataset.
| Class | Accuracy | Sensitivity | Specificity | F1-score |
|---|---|---|---|---|
| N | 0.994 | 0.994 | 0.994 | 0.993 |
| S | 0.999 | 0.999 | 0.999 | 0.998 |
| V | 0.999 | 0.998 | 0.999 | 0.999 |
| F | 0.996 | 0.984 | 0.997 | 0.983 |
| Q | 0.997 | 0.987 | 0.999 | 0.991 |
| Average | 0.997 | 0.992 | 0.998 | 0.993 |
Note: N means normal; S means VPC; V means ventricular escape; F means a fusion of ventricular and normal; and Q means unclassified heartbeat.
Table 7.
Performance comparison reported in related studies using the MIT-BIH ECG data.
| Methods | Accuracy | Sensitivity | Specificity | F1-score |
|---|---|---|---|---|
| Liang et al.* 28 | 0.84 | 0.99 | 0.85 | |
| Irfan et al. 29 | 0.994 | 0.984 | 0.996 | |
| Li et al. 30 | 0.996 | 0.938 | 0.993 | |
| Ribeiro et al. 31 | 0.985 | 0.998 | ||
| Ours | 0.997 | 0.992 | 0.998 | 0.993 |
Note: “ ” means the work’s paper does not give the metric. * Liang et al. used rhythmic labels instead of beat labels.
Benchmark previous work on multi-label arrhythmia type classification
To carefully compare the performance of our framework to the performance of previous state-of-the-art work on arrhythmia classification, we used the real-world multi-label arrhythmia dataset used in the previous section. We examined four state-of-the-art models in the literature and recorded their performance using accuracy, sensitivity, specificity, and F1-score across a five-fold cross-validation.
The results for benchmarking previous work on multi-label classification are shown in Table 8. As we can see, the performance of previous state-of-the-art models suffers as they encounter real-world wearable ECG signals. The noise, patient variability, and the multi-label nature decreased performance when compared to running on the MIT-BIH dataset. However, as we compare Table 4, which has our CNN+BiLSTM with Attention model, to the performance of previous state-of-the-art models, we see that our model has an advantage in arrhythmia classification.
Table 8.
Experimental result of previous related work compared to our framework on our multi-labeled real-world ECG data across five-fold cross-validation.
Discussions
Our contributions include addressing real-world ECG data from wireless portable ECG monitoring devices for arrhythmia deep learning classification. We have proven this by benchmarking our framework against previous state-of-the-art models on our real-world ECG and MIT-BIH clinical ECG datasets. Unlike the available public datasets collected in a clinical setting where the environment and patient movements are carefully controlled, real-world ECG data can have significantly more variability. The reason for such variability could be a diverse range of activities and environments affecting the ECG signal the monitored patient can perform in the real world. For example, suppose the patient is doing an intensive workout, such as running, which will make their heart beat faster. However, it can also cause the area or pressure of the contact surface for the monitor with the patient to change, resulting in different signals from the actual patient’s heartbeats. Moreover, the data becomes significantly more imbalanced due to longer monitoring duration. Therefore, real-world ECG data poses a challenge different from clinically collected ECG data.
We address these challenges by performing a prediction every four beats instead of every beat to address the data imbalances between normal heartbeats and arrhythmia heartbeats, as well as between different arrhythmia heartbeats. Additionally, the four beats in an input do not have equal lengths. Some beats can be faster, and others can be slower; capturing this temporal difference allows more details to remain in the data. Moreover, a longer input length provides more sophisticated convolution and time series analysis, enabling deeper models. Another reason is that our dataset contains Bigeminy and Trigeminy classes. Bigeminy has normal (sinus) beats and VPC beats appearing alternately, and Trigeminy has two normal beats and one abnormal one or two VPCs with one sinus beat. Therefore, observations of at least three beats are required to properly distinguish these classes clinically. Critically, this allows us to perform both rhythm and beat arrhythmia classification, which has significant clinical value.
Our hierarchical architecture, including a binary classification of normal and arrhythmia heartbeats and a subsequent multi-label arrhythmia classification between arrhythmia beats, allows a more efficient and streamlined workflow for arrhythmia patient care. The first binary classification part can alert doctors about potential arrhythmia patients, and then the detailed arrhythmia types can be predicted. This simplifies the decision-making process for doctors as they can better estimate a patient’s possible illness, compared to a more sophisticated model that predicts normal heartbeats alongside all types of arrhythmia heartbeats. Moreover, the sophisticated model will add computational cost and restrictions on the doctors and medical institutions as more heartbeats need to be fed into the model. In contrast, the predicted normal heartbeats in the hierarchical architecture will not go into the multi-label arrhythmia classification model with high probability.
MEZOO’s data demonstrates that arrhythmia types can concurrently occur in certain cases. Multi-label classification offers important clinical significance as patients with multiple arrhythmias may need management or treatment of all occurring arrhythmias rather than solely the main arrhythmia type. For example, different arrhythmia conditions are treated differently. Patients with VPC are treated drastically differently from patients with Atrial Fibrillation. With patients having both types of arrhythmia, but one is only detected, clinicians cannot offer accurate prognoses and formulate optimal treatment plans, resulting in serious medical consequences.
Clinical significance and impact
By leveraging advanced neural network architectures, this system enhances the precision and accuracy of arrhythmia detection, which is paramount for timely and effective patient management. The ability to continuously monitor and analyze ECG data in real time provides cardiologists with a powerful tool for early diagnosis and intervention, potentially reducing the incidence of adverse cardiac events. This approach not only supports the identification of common arrhythmias but also facilitates the recognition of more subtle and complex arrhythmic patterns that might otherwise go unnoticed in conventional monitoring settings. Notably, our results are derived from real-world data obtained over 24-hour periods during patients’ everyday activities rather than the brief ECG signals typically recorded during hospital visits. Clinically, this is significant because it captures a comprehensive view of the patient’s cardiac health, encompassing variations and anomalies that are more likely to occur in a naturalistic setting. Consequently, integrating this technology into clinical practice promises to elevate the standard of cardiac care, offering a robust solution for improving patient outcomes, personalizing treatment plans, and optimizing healthcare resources.
Limitations
Our study also has a few limitations. First, despite the overall robustness of our model, we observed some misclassifications, particularly in the VPC+Trigem and VPC+AFib classes. The confusion matrix shown in Figure 4 indicates that these classes are sometimes misclassified, but importantly, these misclassifications are typically between clinically related categories, such as different types of VPC-related conditions and AFib. However, it is known that when two or more arrhythmias coexist,32–34 clinicians might only label a subset of these arrhythmias or label additional arrhythmias that are not present. This kind of misspecified data might explain the observed overlap in our model’s classifications. More understanding of this interplay can assist clinicians in patient monitoring and treatment planning by leveraging our model’s multi-labeled predictions.
Second, our method is tested in the same environment as the training stage in a data center. The deployment of our method on wearable devices is an integral step in achieving commercial and clinical applications, including real-time ECG monitoring. We leave the actual deployment of the inference stage of our method on edge ECG monitor devices as future work.
Conclusion and future work
We propose a hierarchical machine learning model for arrhythmia detection and classification in the real-world wearable device setting. We compare our model with the benchmark arrhythmia prediction models, including CNN+BiGRU with attention, ConViT, EfficientNet, and ResNet, as well as previous state-of-the-art work, using proprietary data with wireless ECG data. Our model achieved excellent performance on both the MIT-BIH dataset and our proprietary data compared to the state-of-the-art arrhythmia classification models. Our results suggest the possibility of practical use of arrhythmia detection, including the user’s movements, daily noise, and device artifacts.
In the future, from a clinical and practical perspective, there is a need to develop algorithms for the classification of arrhythmia type with infrequent data, high-risk arrhythmia alarms, exploration of effective preprocessing methods, and real-time clinical decision support system (CDSS). Specifically, as we get more and more data from MEZOO, we will expand the multi-label classification beyond the top 7 most frequent arrhythmia types/combinations and conduct real-world deployment experiments of the hierarchical architecture. We will also improve the performance of our models through data augmentation techniques, novel architecture, and innovative preprocessing techniques. Edge device deployability is also a significant area of research; we envision a framework that can train, personalize, and infer on the ECG monitoring device locally without uploading data to ensure the privacy of every patient.
Acknowledgements
The authors would like to thank The Center for Research Computing at Rice University for providing hardware and technical assistance for experiments.
Footnotes
Contributorship: GZ and SL are both the first authors of this paper. JK, KP, HL, ZX, HS, and JS assisted in the experiment implementation. SPC and SII provided medical background and support for the paper. ICJ and VB are both corresponding authors of this article.
The author(s) declared no potential conflicts of interest for this article’s research, authorship, and/or publication.
Ethical approval: The Rice University IRB reviewed the data used, and this activity was determined to be Not Human Subjects Research.
Funding: The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was financially supported by the Ministry of Trade, Industry and Energy (MOTIE) and the Korea Institute for Advancement of Technology (KIAT) through the International Cooperative R&D program (Project Number: P0019781)
Guarantor: GZ.
ORCID iD: Guangyao Zheng https://orcid.org/0000-0002-8864-8578
References
- 1.Khan Y, Ostfeld AE, Lochner CM, et al. Monitoring of vital signs with flexible and wearable medical devices. Adv Mater 2016; 28: 4373–4395. [DOI] [PubMed] [Google Scholar]
- 2.Hung K, Zhang YT, Tai B. Wearable medical devices for tele-home healthcare. In: The 26th annual international conference of the IEEE engineering in medicine and biology society volume 2, pp.5384–5387. IEEE. [DOI] [PubMed]
- 3.Azariadi D, Tsoutsouras V, Xydis S, et al. ECG signal analysis and arrhythmia detection on iot wearable medical devices. In: 2016 5th International conference on modern circuits and systems technologies (MOCAST) pp.1–4. IEEE.
- 4.MEZOO Co. Ltd., 2023.
- 5.Goldberger AL, Amaral LAN, Glass L, et al. PhysioBank, PhysioToolkit, and PhysioNet. Circulation 2000; 101: e215–e220. [DOI] [PubMed] [Google Scholar]
- 6.Moody G, Mark R. The impact of the MIT-BIH arrhythmia database. IEEE Eng Med Biol Mag 2001; 20: 45–50. Conference Name: IEEE Engineering in Medicine and Biology Magazine. [DOI] [PubMed] [Google Scholar]
- 7.Mathunjwa BM, Lin YT, Lin CH, et al. ECG arrhythmia classification by using a recurrence plot and convolutional neural network. Biomed Signal Process Control 2021; 64: 102262. [Google Scholar]
- 8.Sahoo S, Dash M, Behera S, et al. Machine learning approach to detect cardiac arrhythmias in ECG signals: a survey. IRBM 2020; 41: 185–194. [Google Scholar]
- 9.Cheikhrouhou O, Mahmud R, Zouari R, et al. One-dimensional CNN approach for ECG arrhythmia analysis in fog-cloud environments. IEEE Access 2021; 9: 103513. Conference Name: IEEE Access. [Google Scholar]
- 10.Jing E, Zhang H, Li Z, et al. ECG heartbeat classification based on an improved ResNet-18 model. Comput Math Methods Med 2021; 2021: 6649970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.wFurqon M, Nugroho SMS, Rachmadi RF, et al. Arrhythmia classification using efficientnet-v2 with 2-D scalogram image representation. In: 2021 TRON Symposium (TRONSHOW) pp.1–9. IEEE.
- 12.Yildirim O, Baloglu UB, Tan RS, et al. A new approach for arrhythmia classification using deep coded features and LSTM networks. Comput Methods Programs Biomed 2019; 176: 121–133. [DOI] [PubMed] [Google Scholar]
- 13.Neri L, Oberdier MT, van Abeelen KC, et al. Electrocardiogram monitoring wearable devices and artificial-intelligence-enabled diagnostic capabilities: a review. Sensors 2023; 23: 4805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Clifford GD, Liu C, Moody B, et al. AF classification from a short single lead ECG recording: the PhysioNet/computing in cardiology challenge 2017. Comput Cardiol (2010) 2017; 44: 1–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Andreotti F, Carr O, Pimentel MAF, et al. Comparing feature-based classifiers and convolutional neural networks to detect arrhythmia from short segments of ECG. In: 2017 Computing in Cardiology (CinC). pp.1–4. DOI: 10.22489/CinC.2017.360-239. https://ieeexplore.ieee.org/document/8331748. ISSN: 2325-887X.
- 16.Xiong Z, Nash MP, Cheng E, et al. ECG signal classification for the detection of cardiac arrhythmias using a convolutional recurrent neural network. Physiol Meas 2018; 39: 094006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Liu F, Liu C, Zhao L, et al. An open access database for evaluating the algorithms of electrocardiogram rhythm and morphology abnormality detection. J Med Imaging Health Inform 2018; 8: 1368–1373. [Google Scholar]
- 18.Chen TM, Huang CH, Shih ESC, et al. Detection and classification of cardiac arrhythmias by a challenge-best deep learning neural network model. iScience 2020; 23: 100886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hannun AY, Rajpurkar P, Haghpanahi M, et al. Cardiologist-level arrhythmia detection and classification in ambulatory electrocardiograms using a deep neural network. Nat Med 2019; 25: 65–69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Rajpurkar P, Hannun AY, Haghpanahi M, et al. Cardiologist-level arrhythmia detection with convolutional neural networks. 2017. DOI: 10.48550/arXiv.1707.01836. http://arxiv.org/abs/1707.01836. ArXiv:1707.01836 [cs].
- 21.Liu G, Guo J. Bidirectional LSTM with attention mechanism and convolutional layer for text classification. Neurocomputing 2019; 337: 325–338. [Google Scholar]
- 22.Cheng J, Zou Q, Zhao Y. ECG signal classification based on deep CNN and BiLSTM. BMC Med Inform Decis Mak 2021; 21: 1–12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Misra D. Mish: a self regularized non-monotonic activation function. arXiv preprint arXiv:190808681 2019.
- 24.Gottesman Y. Interpretable ECG classification with 1D vision transformer. https://yonigottesman.github.io/ecg/vit/deep-learning/2023/01/20/ecg-vit.html, 2023.
- 25.Tan M, Le Q. Efficientnet: rethinking model scaling for convolutional neural networks. In: International conference on machine learning pp.6105–6114. PMLR.
- 26.He K, Zhang X, Ren S, et al. Deep residual learning for image recognition. In: Proceedings of the IEEE conference on computer vision and pattern recognition pp.770–778.
- 27.Lin TY, Goyal P, Girshick R, et al. Focal loss for dense object detection. In: 2017 IEEE international conference on computer vision (ICCV) pp.2999–3007. DOI: 10.1109/ICCV.2017.324.
- 28.Liang Y, Yin S, Tang Q, et al. Deep learning algorithm classifies heartbeat events based on electrocardiogram signals. Front Physiol 2020; 11: 569050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Irfan S, Anjum N, Althobaiti T, et al. Heartbeat classification and arrhythmia detection using a multi-model deep-learning technique. Sensors 2022; 22: 5606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Li X, Zhang F, Sun Z, et al. Automatic heartbeat classification using S-shaped reconstruction and a squeeze-and-excitation residual network. Comput Biol Med 2022; 140: 105108. [DOI] [PubMed] [Google Scholar]
- 31.Ribeiro HDM, Arnold A, Howard JP, et al. ECG-based real-time arrhythmia monitoring using quantized deep neural networks: a feasibility study. Comput Biol Med 2022; 143: 105249. [DOI] [PubMed] [Google Scholar]
- 32.Yanagisawa Y, Ibrahim W, Kumar N. A case of atrial fibrillation complicated by complete atrioventricular block. SAGE Open Med Case Rep 2023; 11: 2050313X231157486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Aro A, Eyob-Fesseha H, Haukka J, et al. How often atrial flutter and atrial fibrillation coexist? Results from a large nationwide ECG-based study. EP Europace 2023; 25: euad122.037. [Google Scholar]
- 34.Zhang D, Huang X. Treatment of atrial fibrillation with third-degree atrioventricular block by pacing his bundle and left bundle branch: Case report. Medicine 2020; 99: e21097. [DOI] [PMC free article] [PubMed] [Google Scholar]