Abstract
We have isolated several new clones of human ribosomal DNA. Each clone contains part of the external transcribed spacer, a complete 18 S-rRNA gene, the internal transcribed spacers, a complete 28 S-rRNA gene and a short downstream flanking region. We present a detailed map of the human ribosomal transcription unit with the locations of numerous useful restriction sites. In particular, a unique NheI site in the 5.8 S-rRNA gene enabled this gene to be mapped with respect to the 18 S-rRNA and 28 S-rRNA genes. The human 45 S-rRNA coding region is approx. 13,000 nucleotide residues long, of which the external transcribed spacer comprises approx. 3700 nucleotide residues and the first and second internal transcribed spacers comprise approx. 1070 and 1200 nucleotide residues respectively. A partial survey for sites of variation between clones has revealed a single point of variation among 18 S-rRNA gene sequences (a T/C variation at position 140), several sites of length variation in the regions of the transcribed spacers closely flanking the 18 S-rRNA genes, and some sites of length variation among 28 S-rRNA genes. Most of these sites of variation are associated with simple sequence tracts and are in regions that are known to undergo relatively rapid evolutionary divergence. In particular, the sites of variation among 28 S-rRNA genes occur in G + C-rich tracts whose lengths vary among vertebrates and that can be correlated with extensive hairpin structures previously observed by electron microscopy. Each of the clones so far surveyed in detail differs from the others in one or more respects.
Full text
PDF




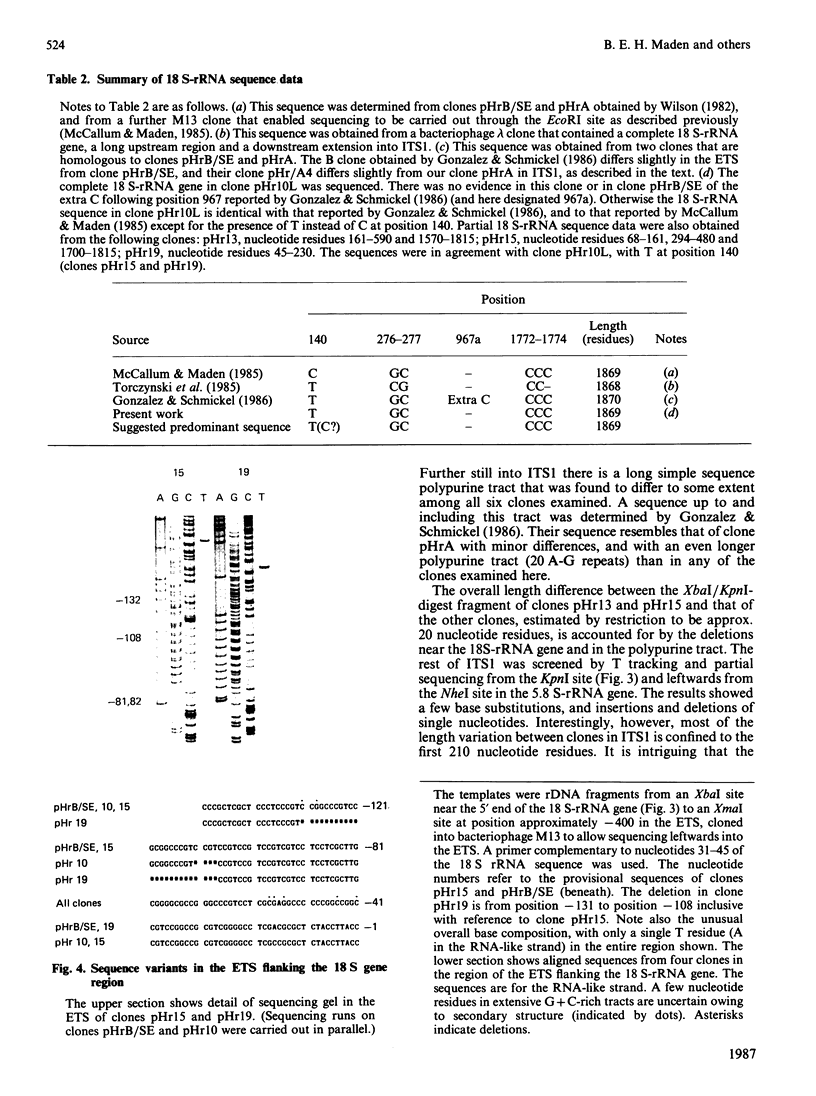



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan Y. L., Gutell R., Noller H. F., Wool I. G. The nucleotide sequence of a rat 18 S ribosomal ribonucleic acid gene and a proposal for the secondary structure of 18 S ribosomal ribonucleic acid. J Biol Chem. 1984 Jan 10;259(1):224–230. [PubMed] [Google Scholar]
- Chan Y. L., Olvera J., Wool I. G. The structure of rat 28S ribosomal ribonucleic acid inferred from the sequence of nucleotides in a gene. Nucleic Acids Res. 1983 Nov 25;11(22):7819–7831. doi: 10.1093/nar/11.22.7819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark C. G., Tague B. W., Ware V. C., Gerbi S. A. Xenopus laevis 28S ribosomal RNA: a secondary structure model and its evolutionary and functional implications. Nucleic Acids Res. 1984 Aug 10;12(15):6197–6220. doi: 10.1093/nar/12.15.6197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Financsek I., Mizumoto K., Mishima Y., Muramatsu M. Human ribosomal RNA gene: nucleotide sequence of the transcription initiation region and comparison of three mammalian genes. Proc Natl Acad Sci U S A. 1982 May;79(10):3092–3096. doi: 10.1073/pnas.79.10.3092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frischauf A. M., Lehrach H., Poustka A., Murray N. Lambda replacement vectors carrying polylinker sequences. J Mol Biol. 1983 Nov 15;170(4):827–842. doi: 10.1016/s0022-2836(83)80190-9. [DOI] [PubMed] [Google Scholar]
- Furlong J. C., Forbes J., Robertson M., Maden B. E. The external transcribed spacer and preceding region of Xenopus borealis rDNA: comparison with the corresponding region of Xenopus laevis rDNA. Nucleic Acids Res. 1983 Dec 10;11(23):8183–8196. doi: 10.1093/nar/11.23.8183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Furlong J. C., Maden B. E. Patterns of major divergence between the internal transcribed spacers of ribosomal DNA in Xenopus borealis and Xenopus laevis, and of minimal divergence within ribosomal coding regions. EMBO J. 1983;2(3):443–448. doi: 10.1002/j.1460-2075.1983.tb01442.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez I. L., Gorski J. L., Campen T. J., Dorney D. J., Erickson J. M., Sylvester J. E., Schmickel R. D. Variation among human 28S ribosomal RNA genes. Proc Natl Acad Sci U S A. 1985 Nov;82(22):7666–7670. doi: 10.1073/pnas.82.22.7666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez I. L., Schmickel R. D. The human 18S ribosomal RNA gene: evolution and stability. Am J Hum Genet. 1986 Apr;38(4):419–427. [PMC free article] [PubMed] [Google Scholar]
- Gorski J. L., Gonzalez I. L., Schmickel R. D. The secondary structure of human 28S rRNA: the structure and evolution of a mosaic rRNA gene. J Mol Evol. 1987;24(3):236–251. doi: 10.1007/BF02111237. [DOI] [PubMed] [Google Scholar]
- Hassouna N., Michot B., Bachellerie J. P. The complete nucleotide sequence of mouse 28S rRNA gene. Implications for the process of size increase of the large subunit rRNA in higher eukaryotes. Nucleic Acids Res. 1984 Apr 25;12(8):3563–3583. doi: 10.1093/nar/12.8.3563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan M. S., Maden B. E. Nucleotide sequence relationships between vertebrate 5.8 S ribosomal RNAs. Nucleic Acids Res. 1977 Jul;4(7):2495–2505. doi: 10.1093/nar/4.7.2495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loening U. E. Molecular weights of ribosomal RNA in relation to evolution. J Mol Biol. 1968 Dec;38(3):355–365. doi: 10.1016/0022-2836(68)90391-4. [DOI] [PubMed] [Google Scholar]
- Maden B. E., Forbes J. M., Stewart M. A., Eason R. 18S coding sequences in amplified ribosomal DNA from Xenopus laevis oocytes are highly homogeneous, unmethylated, and lack major open reading frames. EMBO J. 1982;1(5):597–601. doi: 10.1002/j.1460-2075.1982.tb01214.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maden B. E. Identification of the locations of the methyl groups in 18 S ribosomal RNA from Xenopus laevis and man. J Mol Biol. 1986 Jun 20;189(4):681–699. doi: 10.1016/0022-2836(86)90498-5. [DOI] [PubMed] [Google Scholar]
- Maden B. E. Methylation map of Xenopus laevis ribosomal RNA. Nature. 1980 Nov 20;288(5788):293–296. doi: 10.1038/288293a0. [DOI] [PubMed] [Google Scholar]
- McCallum F. S., Maden B. E. Human 18 S ribosomal RNA sequence inferred from DNA sequence. Variations in 18 S sequences and secondary modification patterns between vertebrates. Biochem J. 1985 Dec 15;232(3):725–733. doi: 10.1042/bj2320725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miesfeld R., Arnheim N. Identification of the in vivo and in vitro origin of transcription in human rDNA. Nucleic Acids Res. 1982 Jul 10;10(13):3933–3949. doi: 10.1093/nar/10.13.3933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raynal F., Michot B., Bachellerie J. P. Complete nucleotide sequence of mouse 18 S rRNA gene: comparison with other available homologs. FEBS Lett. 1984 Feb 27;167(2):263–268. doi: 10.1016/0014-5793(84)80139-8. [DOI] [PubMed] [Google Scholar]
- Schibler U., Wyler T., Hagenbüchle O. Changes in size and secondary structure of the ribosomal transcription unit during vertebrate evolution. J Mol Biol. 1975 May 25;94(3):503–517. doi: 10.1016/0022-2836(75)90217-x. [DOI] [PubMed] [Google Scholar]
- Smith G. P. Evolution of repeated DNA sequences by unequal crossover. Science. 1976 Feb 13;191(4227):528–535. doi: 10.1126/science.1251186. [DOI] [PubMed] [Google Scholar]
- Stewart M. A., Hall L. M., Maden B. E. Multiple heterogeneities in the transcribed spacers of ribosomal DNA from Xenopus laevis. Nucleic Acids Res. 1983 Feb 11;11(3):629–646. doi: 10.1093/nar/11.3.629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tautz D., Trick M., Dover G. A. Cryptic simplicity in DNA is a major source of genetic variation. Nature. 1986 Aug 14;322(6080):652–656. doi: 10.1038/322652a0. [DOI] [PubMed] [Google Scholar]
- Torczynski R. M., Fuke M., Bollon A. P. Cloning and sequencing of a human 18S ribosomal RNA gene. DNA. 1985 Aug;4(4):283–291. doi: 10.1089/dna.1985.4.283. [DOI] [PubMed] [Google Scholar]
- Torczynski R., Bollon A. P., Fuke M. The complete nucleotide sequence of the rat 18S ribosomal RNA gene and comparison with the respective yeast and frog genes. Nucleic Acids Res. 1983 Jul 25;11(14):4879–4890. doi: 10.1093/nar/11.14.4879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wellauer P. K., Dawid I. B., Kelley D. E., Perry R. P. Secondary structure maps of ribosomal RNA. II. Processing of mouse L-cell ribosomal RNA and variations in the processing pathway. J Mol Biol. 1974 Oct 25;89(2):397–407. doi: 10.1016/0022-2836(74)90527-0. [DOI] [PubMed] [Google Scholar]
- Wellauer P. K., Dawid I. B. Secondary structure maps of RNA: processing of HeLa ribosomal RNA. Proc Natl Acad Sci U S A. 1973 Oct;70(10):2827–2831. doi: 10.1073/pnas.70.10.2827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson G. N., Hollar B. A., Waterson J. R., Schmickel R. D. Molecular analysis of cloned human 18S ribosomal DNA segments. Proc Natl Acad Sci U S A. 1978 Nov;75(11):5367–5371. doi: 10.1073/pnas.75.11.5367. [DOI] [PMC free article] [PubMed] [Google Scholar]