Abstract
High heterogeneity in clinical benefit characterizes the use of immune checkpoint inhibitors (ICIs) in non-small cell lung cancer (NSCLC). We prospectively enrolled 113 advanced NSCLC patients treated with ICIs and performed liquid biopsy at the time of ICI start (T1), after 3 weeks (T2) and at the time of radiological evaluation (T3). Molecular variables were associated with outcome endpoints: cfDNA quantification, its dynamic change (∆T2–T1), variant allele frequency (VAF) of the gene with the highest frequency detected at baseline with NGS (maxVAF) and its dynamic change (∆T2–T1). At multivariate analysis, PD-L1 negativity, T1 cfDNA, cfDNA increase (∆T2–T1), and T2 VAF were significantly associated with shorter progression-free survival (PFS); PD-L1 negativity, squamous histology, T1 cfDNA, cfDNA (∆T2–T1) increase, and T2 maxVAF affected overall survival (OS). Among high PD-L1 expressing patients treated in first-line, elevated T2 maxVAF and cfDNA increase (∆T2–T1) correlated with worse PFS; higher T2 maxVAF and cfDNA increase (∆T2–T1) with worse OS. Derived integrated models were used to build nomograms and categorize different risk groups.
Subject terms: Cancer genomics, Translational research, Predictive markers, Non-small-cell lung cancer, Predictive markers
Introduction
The introduction of immunotherapy in the treatment of advanced non-small cell lung cancer (NSCLC) is one of the most important advances for patients’ outcomes and quality of life1,2.
Currently, most of the patients diagnosed with advanced NSCLC receive treatment with immune checkpoint inhibitors (ICIs) with an important exception represented by so-called oncogene-addicted diseases1.
In clinical practice, NSCLC patients need to be tested at least for the best-known druggable genes, including EGFR, ALK, ROS1, RET, MET, NTRK, BRAF, HER2 and KRAS actionable alterations3. The standard method of assessment is represented by tissue molecular characterization, although the use of liquid biopsy for molecular diagnosis is being implemented. In addition to supporting precision oncology, liquid biopsy has several research applications, among which its potential usefulness in the search for predictive biomarkers for immunotherapy.
Different treatment strategies including ICIs are currently available for first-line treatment: anti-PD-1/PD-L1 monotherapy, combination of chemotherapy and anti-PD-1 antibody, anti-PD-1 plus anti-CTLA4 plus a short course of chemotherapy1,4–10. Nowadays the medical choice among treatment options is based on PD-L1 expression, the only available predictive biomarkers in clinical practice, and toxicity profiling1. Great heterogeneity in response and duration of response has been observed. In particular, a subgroup of patients could even derive detrimental effects from immunotherapy single agent, while 15–20% of patients can have prolonged clinical benefit and are defined as long-survivors, even in the absence of more toxic combination strategies11–13.
To shed light on heterogeneity in clinical benefit, several biomarkers have been investigated and, overall, findings suggest that response to immunotherapy is likely not related to a single biomarker but rather to the dynamic interaction among factors related to host, tumour and microenvironment14,15. We thus speculated that circulating tumour DNA (ctDNA) could be considered as a surrogate of tumour burden and early treatment-related biological changes, potentially able to anticipate response and clinical benefit following ICIs13,16,17.
We performed a prospective study in advanced NSCLC patients treated with single-agent ICIs and integrated clinicopathological features and results of ctDNA monitoring in order to build a predictive model.
Methods
Patients’ enrolment and study design
Patients with advanced NSCLC and treated with ICIs at the Veneto Institute of Oncology IOV– IRCCS (Padua, Italy) were enrolled as part of the prospective study called MAGIC between January 2017 and August 201913,16. The main inclusion criteria were histological diagnosis of EGFR-ALK-ROS1 wild-type NSCLC, the feasibility of ICI treatment according to clinical practice, the possibility of an appropriate clinical and radiological follow-up and signed informed consent.
According to Italian regulatory indications, ICIs were administered after progression to chemotherapy independently of PD-L1 expression between January and July 2017. Since July 2017 patients were treated with first-line pembrolizumab in the presence of PD-L1 expression in at least 50% of tumour cells in tumour biopsies and with second-line ICIs (nivolumab, pembrolizumab and atezolizumab) in the presence of PD-L1 inferior to 50%.
PD-L1 expression levels and detection of EGFR/ALK/ROS1 actionable alterations were all assessed by the Pathology Unit according to clinical practice. PD-L1 was routinely assessed by using DAKO 22C3.
Liquid biopsies were collected at pre-planned time-points during ICIs: on the same day of the first administration of treatment (baseline or T1), after 2 weeks or 3 weeks according to treatment schedule (T2), at the time of first radiological evaluation (T3).
Radiological and clinical follow-ups were performed according to clinical practice.
The ethics committee of IOV evaluated and approved study design and informed consent (2016/82, 12/12/2016). Written informed consent was obtained from all patients before study entry. The study was conducted in accordance with the precepts of the Helsinki Declaration.
Plasma sample collection
Blood draws (~20 ml) were collected at each time point in two cell-free DNA BCT tubes (Streck Corporate, La Vista, NE, USA) and processed for plasma extraction within the next 24–72 h. As previously described13,16, two sequential centrifugations were performed to obtain plasma suitable for NGS analyses: first, blood draws were centrifuged at 2000 × g for 10 min at 4 °C to separate plasma, then plasma was centrifuged at 20,000 × g for 10 min at 4 °C to pellet cellular debris and the supernatant was stored at −80 °C, until cfDNA extraction. Aliquots of plasma-depleted blood were also stored at −80 °C, until they were used for CH analysis.
cfDNA extraction and quantity/quality assessment
cfDNA was extracted from 3–5 ml of plasma, using QIAamp Circulating Nucleic Acid Kit (Qiagen, Venlo, Netherlands) following the manufacturer’s instructions. Circulating nucleic acids were eluted in 30 µl of nuclease-free water. Quantity and amplificability of cfDNA were assessed by qPCR included in the Myriapod NGS-IL 56G Onco panel CE IVD kit (Diatech Pharmacogenetics, Jesi, Italy).
Genomic DNA extraction from plasma-depleted blood
Genomic DNA (gDNA) was extracted from 200 µl of plasma-depleted blood using a Qiagen DNA mini kit following manufacturers’ instructions. gDNA was eluted in 50 µl of buffer and quantified with Qubit dsDNA BR Assay Kit on a Qubit 4.0 Fluorometer (Thermo Fisher Scientific, Waltham, Massachusetts, USA).
DNA extraction from formalin-fixed paraffin-embedded (FFPE) tissue biopsies
DNA extraction from FFPE tissue biopsies was performed from five slides (10 µm thick) using QIAamp DNA Micro Kit (Qiagen), following the manufacturer’s instructions. Elution of DNA was done in 30 µl with nuclease-free water. Quantification and amplificability were done by qPCR included in the Myriapod NGS-IL 56 G Onco panel CE IVD kit.
Library preparation and sequencing
Targeted next generation sequencing (NGS) was performed in liquid biopsies collected at each time-point, gDNA from blood, and tumour DNA from FFPE tissue biopsies using Myriapod NGS-IL 56 G Onco panel CE IVD kit (Diatech) covering hot spot regions of 56 frequently mutated tumour suppressor genes and oncogenes in cancers (Supplementary Table 1). NGS libraries were prepared starting from 1.27 ng of cfDNA to a maximum of 25 ng. For gDNA from blood, the maximum input (25 ng) was always used. For FFPE tumour DNA, libraries were prepared starting from 3 ng to 25 ng. Eighteen Indexed libraries were pooled together, diluted at a final concentration of 12 pM, and sequenced on an Illumina MiSeqDx System using the 600 cycles MiSeq Reagent Kit v3 (Illumina) in pair-end mode (2 × 151 cycles). FastQ files obtained from sequencing were analysed using Myriapod NGS Data Analysis Software. Synonymous and intronic variants were excluded from analyses. The remaining mutations were consulted on Varsome Premium (Saphetor SA, Lausanne, Switzerland) to define their pathogenicity according to ACMG guidelines18. Only pathogenic or likely pathogenic mutations and variants of uncertain significance (VUS) were considered for longitudinal analyses.
Statistical analyses
Radiological response (RR), progression-free survival (PFS), and overall survival (OS) were the primary endpoints considered for the analysis.
Given the exploratory nature of the study, there was no formal hypothesis or power sample size calculation.
RR was graded as complete response (CR), partial response (PR), stable disease (SD), and progressive disease (PD) according to RECIST 1.1 criteria19. PFS was calculated as the time from the beginning of systemic treatment to the occurrence of radiological disease progression or death from any cause. OS was calculated as the time from the beginning of systemic treatment to death from any cause. Patients who did not develop a survival event during the study period were censored at the date of the last observation.
As molecular variables, we considered both cfDNA concentration in plasma (ng/ml) and variant allele frequency (VAF).
cfDNA genetic alterations not found in tumour tissue, were evaluated in plasma-depleted blood to exclude alterations related to clonal hematopoiesis (CH).
In the presence of multiple mutations, the one with the highest VAF at baseline was selected as the reference for NGS analysis and referred to in the manuscript as maxVAF.
When detected VAF of all tumour-associated gene alterations found was inferior to 0.5%, considered as the limit of detection (LOD) of the NGS assay, values were replaced with 0.025, corresponding to LOD/2. To define the LOD, we analysed samples carrying KRAS mutation according to NGS and confirmed by droplet digital PCR (ddPCR): the lowest VAF confirmed by ddPCR was 0.5%.
Continuous variables were described as median and interquartile range and their distributions according to different groups were compared using the Kruskal–Wallis test. Categorical variables were described as counts and percentages and compared between groups using the χ2 or Fisher exact test as appropriate. The median follow-up time was based on the reverse Kaplan–Meier estimator.
The correlation between cfDNA and maxVAF at each time point was calculated by using Spearman’s rank correlation coefficient.
Each biomarker was analysed for association with clinical outcome as a categorical variable according to high and low levels. Optimal cut-points were estimated by maximizing the discriminative ability of the Cox model for PFS and OS, and validated with bootstrapping.
The main effects of candidate clinical and molecular predictors were included in multiple Cox proportional hazards regression models for the survival outcome. No deviation from the proportional hazards assumption was found by the test statistic of Grambsch and Therneau.
A backward elimination with the Bayesian information criterion was applied to selecting all variables independently associated with the outcome. Results were displayed in terms of hazard ratios (HR) together with 95% CI. A nomogram of the final reduced model was generated for predicting the 6- and 12-month survival probabilities.
The ability of the models to differentiate between high-risk patients and low-risk patients, was calculated with Harrell’s concordance (C) index, adjusted through 1000 bootstrap resamples.
Lastly, patients were stratified into three risk groups according to optimal cut-points of their individual linear predictor scores, and the log-rank test was used to compare groups.
The nonparametric Kaplan–Meier method was used to estimate the survival probabilities and the median time was provided along with the corresponding 95% confidence interval (CI) estimated using the Brookmeyer-Crowley method.
All statistical tests used a two-sided 5% significance level. Statistical analyses were performed using RStudio (RStudio: Integrated Development for R. RStudio, Inc., Boston, MA).
Results
Patients, treatments and outcome
Among 149 advanced NSCLC patients complying with clinical inclusion criteria, 113 (76%) were considered for predictive analyses (Table 1 and Supplementary Table 2), based on the availability of complete clinicopathological data (including available PD-L1 evaluation) and liquid biopsy results. Patients with unknown PDL-1 data (24%) were excluded from our model, given its established predictive value. The main clinical features of the study population are summarized in Table 1.
Table 1.
Clinical features of the study population
| Included patientsa (N = 113, 76%) | Total (N = 149) | ||
|---|---|---|---|
| Age (years) | Median (Q1–Q3) | 67.0 (62.0, 73.0) | 67.0 (62.0, 74.0) |
| Sex | Male | 74 (65.5%) | 97 (65.1%) |
| Female | 39 (34.5%) | 52 (34.9%) | |
| Smoking | No | 13 (11.5%) | 19 (12.8%) |
| Former | 61 (54.0%) | 79 (53.0%) | |
| Yes | 39 (34.5%) | 51 (34.2%) | |
| ECOG-PS | 0 | 48 (42.5%) | 69 (46.3%) |
| 1–2 | 65 (57.5%) | 80 (53.7%) | |
| Histology | Non-squamous | 98 (86.7%) | 129 (86.6%) |
| Squamous | 15 (13.3%) | 20 (13.4%) | |
| PD-L1 | Negative | 38 (33.6%) | 38 (25.5%) |
| Positive | 75 (66.4%) | 75 (50.3%) | |
| Not available | – | 36 (24.2%) | |
| Extrathoracic sites | No | 44 (38.9%) | 62 (41.6%) |
| Yes | 69 (61.1%) | 87 (58.4%) | |
| N metastatic sites | 0–1 | 59 (52.2%) | 74 (49.7%) |
| 2–4 | 54 (47.8%) | 75 (50.3%) | |
| Treatment setting | 1 L | 50 (44.2%) | 50 (33.6%) |
| >1 L | 63 (55.8%) | 99 (66.4%) | |
| Follow-up (months) | Median (Q1–Q3) | 62.1 (59.4, 71.6) | 63.5 (60.5, 78.0) |
aIncluded patients refer to patients included in multivariate analyses, with all clinicopathological data available.
Fifty patients were treated with first-line pembrolizumab, while 63 were treated with ICIs following progression to first-line chemotherapy. Overall, the PR rate was 27.4% while 43 (38.1%) patients had PD as their best response. Disease control (PR + SD) was achieved in 34 patients out of 50 treated in the first line (Table 2). In the overall study population, with a median follow-up of 63.5 months, the median PFS was 6.3 months (95% CI: 5.3, 8.1), and the median OS was 12.0 months (95% CI: 8.1, 14.6) (Table 2).
Table 2.
Outcome of the study population, in terms of RR, PFS and OS
| 1 Treatment line (N = 50) | >1 Treatment line (N = 63) | Total (N = 113) | p value | ||
|---|---|---|---|---|---|
| RR | PR | 21 (42.0%) | 10 (15.9%) | 31 (27.4%) | 0.0160 |
| SD | 13 (26.0%) | 26 (41.3%) | 39 (34.5%) | ||
| PD | 16 (32.0%) | 27 (42.9%) | 43 (38.1%) | ||
| PFS (months) | Median (95% CI) | 9.0 (5.6, 15.4) | 5.5 (4.1, 6.8) | 6.3 (5.3, 8.1) | 0.0200 |
| Two years PFS | % (95% CI) | 22.0 (13.0, 37.1) | 12.7 (6.6, 37.1) | 16.8 (11.2, 25.3) | |
| OS (months) | Median (95% CI) | 13.2 (9.8, 32.1) | 10.8 (6.5, 14.5) | 12.0 (8.1, 14.6) | 0.0200 |
| Two years OS | % (95% CI) | 41.6 (29.9, 57.9) | 22.9 (14.5, 36.2) | 31.1 (23.6, 41.1) |
In the whole study population, 17 patients died within 12 weeks from the start of ICIs and two cases comply radiological criteria for hyper-progression11,13,20.
Among clinicopathological features evaluated, the absence of potentially druggable genetic alterations, PD-L1 levels, and receiving immunotherapy as first-line treatment were able to affect the probability of PFS with statistical significance at univariate analysis, as shown in Supplementary Table 3. Factors able to affect OS included age, levels of PD-L1, number of metastatic sites, and receiving immunotherapy as first-line treatment (Supplementary Table 4).
Results of longitudinal liquid biopsy
Initially, we sought to investigate inter-run variability and LOD of the NGS assay used to detect SNV. To this end, we analysed a positive control containing 11 mutations at known VAF, in multiple (n = 9) library preparations and sequencing runs. Results are displayed in Supplementary Table 5 and in general show marginal deviations in experimental VAF values compared to the expected VAF value, thus confirming the suitability of the method to track even small VAF changes. With regard to the LOD, we performed orthogonal validation of KRAS mutations detected by NGS in cfDNA samples by ddPCR and confirmed them down to a VAF of 0.5% (Supplementary Table 6). For this reason, we included in our model pathogenic mutations or VUS with maxVAF detected by NGS down to 0.5%.
Supplementary Table 2 summarises the results of longitudinal liquid biopsies. Median ctDNA quantification at baseline was 10.03 ng/ml (Q1–Q3: 6.68–15.18, Table 3a). Sixty seven out of 110 patients (60.9%) had detectable mutations at baseline. Of 43 patients (39.1%) with no detectable mutations at baseline, 9 were found positive at T2 and 4 at T3 (Supplementary Table 2). The concentration of cfDNA assessed at each time point was significantly correlated with the levels of VAF detected (Supplementary Table 7).
Table 3.
Liquid biopsy results and RR
| (a) Association of cfDNA quantification and maxVAF at T1 and T2 with RR | ||||||
|---|---|---|---|---|---|---|
| PR (N = 31) | SD (N = 39) | PD (N = 43) | Total (N = 113) | p value | ||
| maxVAF at T1 | N | 31 | 39 | 43 | 113 | <0.001 |
| Median (Q1, Q3) | 1.23 (0.02, 4.40) | 0.02 (0.02, 1.34) | 4.61 (0.02, 15.02) | 1.28 (0.02, 7.65) | ||
| maxVAF at T2 | N | 29 | 37 | 39 | 105 | <0.001 |
| Median (Q1, Q3) | 0.02 (0.02, 0.99) | 0.02 (0.02, 1.21) | 3.81 (1.06, 13.66) | 0.99 (0.02, 3.91) | ||
| maxVAF at T3 | N | 29 | 38 | 16 | 83 | 0.0010 |
| Median (Q1, Q3) | 0.02 (0.02, 0.02) | 0.02 (0.02, 1.43) | 4.84 (0.02, 10.79) | 0.02 (0.02, 1.66) | ||
| cfDNA at T1 | N | 31 | 39 | 43 | 113 | <0.001 |
| Median (Q1, Q3) | 8.81 (5.49, 12.34) | 8.28 (6.36, 10.90) | 13.80 (10.08, 24.34) | 9.91 (6.64, 15.21) | ||
| cfDNA at T2 | N | 29 | 37 | 39 | 105 | <0.001 |
| Median (Q1, Q3) | 8.50 (4.84, 13.01) | 8.22 (5.76, 12.86) | 14.97 (10.56, 51.69) | 10.81 (6.10, 17.88) | ||
| cfDNA at T3 | N | 29 | 38 | 16 | 83 | <0.001 |
| Median (Q1, Q3) | 6.10 (4.99, 8.15) | 10.06 (6.59, 12.51) | 16.66 (10.71, 32.31) | 8.25 (5.30, 13.08) | ||
| (b) Impact on PFS of molecular variables performed at the time of radiological assessment (T3) for patients achieving RR (univariate analysis) | |||||||
|---|---|---|---|---|---|---|---|
| PFS univariate analysis | |||||||
| n | Events | Median PFS (95% CI) | log-rank | HR (95% CI) | p value | ||
| cfDNA at T3 | <9.16 | 43 | 35 | 17.5 (10.0; 24.9) | 0.0023 | Ref | |
| ≥9.16 | 24 | 22 | 7.8 (5.8; 10.4) | 2.31 (1.33; 4.01) | 0.0029 | ||
| maxVAF at T3 | <0.025 | 48 | 39 | 11.7 (9.8; 17.9) | 0.0121 | Ref | |
| ≥0.025 | 19 | 18 | 6.8 (5.6; 11.3) | 2.04 (1.16; 3.60) | 0.0139 | ||
| (c) Impact on OS of molecular variables performed at the time of radiological assessment (T3) for patients achieving RR (univariate analysis) | |||||||
|---|---|---|---|---|---|---|---|
| OS univariate analysis | |||||||
| n | Events | Median (95% CI) | log-rank | HR (95% CI) | p value | ||
| cfDNA at T3 | <5.27 | 20 | 10 | 35.7 (14.5; NA) | 0.0049 | Ref | |
| ≥5.27 | 47 | 38 | 19.9 (12.9; 25.8) | 2.65 (1.31; 5.36) | 0.0066 | ||
| maxVAF at T3 | <0.025 | 48 | 32 | 28.5 (18.5; 37.7) | 0.0241 | Ref | |
| ≥0.025 | 19 | 16 | 16.3 (10.8; 21.2) | 2.19 (1.14; 4.19) | 0.0179 | ||
We considered basal liquid biopsy results in order to test the potential association with clinical features and tumour burden (Supplementary Table 8). Higher cfDNA concentration was detected among former smoker patients (p = 0.0250), whereas no association was observed with tumour burden-related factors. On the other side, patients with two or more metastatic sites have significantly higher VAF (p = 0.05, Supplementary Table 8).
Concordance between tissue genetic characterization and NGS in plasma
In a subset of patients, we investigated the concordance between genetic alterations detected in cfDNA and matched tumour samples. In our cohort of 113 patients, 11 (9.7%) patients were screened in tissue as per diagnostic routine by Sequenom MassARRAY® panel (Myriapod® Lung status kit, Diatech, Supplementary Table 2). In one case (#238), the maxVAF mutation found in liquid biopsy was not found in tissue, and this sample was then analysed in blood for potential CH origin (Supplementary Table 2). Moreover, 39 (34.5%) patients had FFPE tumour tissue available for NGS analysis by the 56 genes panel. Library preparation failed in 3 out of 39 cases because of poor DNA quality. In 36 patients library preparation and sequencing were successful. In one patient, no relevant mutations were found either in tissue biopsy or in liquid biopsy (T1). Twenty out of 36 patients (55.5%) had at least 1 mutation found both in tissue and in liquid biopsy (T1) (Supplementary Table 9). Globally 79 mutations were found in tissue and liquid biopsy in 36 patients. Fifteen mutations (of 10 patients) were present only in liquid biopsy and 34 mutations (of 18 patients) were present only in tissue. If we consider only mutations with the maxVAF at baseline, 18/23 (78.3%) were found also in tissue (Supplementary Table 9). Genetic alterations not found in tissue were evaluated as potential CH-related genes.
Analyses of CH-related genes
To assess the issue of potential clonal haematopoiesis-related genes, we performed NGS analysis in plasma-depleted blood to correct our model for the potential CH origin of the genetic alterations with the highest VAF at baseline. For CH analysis, we selected patients with no tissue information available (n = 43) and discordant cases between tissue and liquid biopsy (n = 5), while patients with concordance between tissue and liquid biopsy (n = 35) and patients with no relevant mutations found in liquid biopsy (n = 31) were excluded (Supplementary Table 2). Of the 48 patients subjected to CH analyses, in six cases the mutation with the highest VAF was also found in blood. In three cases (#230, #320 and #325) no other trackable mutation was available (Supplementary Table 2). In the other three cases (#132, #278, #313) the mutation with the second highest VAF was not detected in blood and it was chosen as trackable (Supplementary Table 2).
Liquid biopsy and outcome
Baseline (T1) and T2 cfDNA and maxVAF were significantly associated with RR, in particular patients experiencing PD as the best RR had higher cfDNA and higher maxVAF compared to those experiencing SD or PR (Table 3a).
For survival analyses, biomarkers were categorized according to median in-bag cut-point values from bootstrap validation to identify cut-offs defining high vs low levels for each variable that was used for subsequent analyses (Supplementary Tables 10 and 11).
At univariate survival analyses, higher cfDNA at baseline and T2 and maxVAF at baseline and T2, as well as the increase of cfDNA (ΔT2–T1) and of maxVAF (ΔT2–T1) from baseline were significantly associated with the risk of progression and shorter survival (Supplementary Tables 12 and 13).
We also considered the potential role of liquid biopsy performed at the time of RR assessment (T3) to further stratify the outcome of patients achieving RR (Table 3b, c). In particular, median PFS was 6.8 (95% CI: 5.6–11.3) months for patients with high maxVAF at T3, vs 11.7 (95% CI: 9.8–17.9) months in the presence of low maxVAF at T3 (p: 0.0139, Table 3b) and median OS was 16.3 (95% CI: 10.8–21.2) and 28.5 (95% CI: 18.5–37.7) months in the presence of high or low maxVAF at T3, respectively (p: 0.0241, Table 3c).
Integrated model of liquid biopsy results and clinical features
At multivariate analysis, including clinicopathological and molecular features, PD-L1 negativity, baseline cfDNA, increase of cfDNA (ΔT2–T1), and maxVAF at T2 were significantly associated with shorter PFS (Table 4); moreover, PD-L1 negativity, squamous histology, baseline cfDNA, increase of cfDNA (ΔT2–T1), and maxVAF at T2 were significantly associated with worse OS (Table 5).
Table 4.
Multivariate analysis including clinicopathological and molecular variables and their impact on PFS
| PFS | Multiple analysis | Shrunked | |||
|---|---|---|---|---|---|
| HR (95% CI) | p value | HR (95% CI) | p value | ||
| PD-L1 | Negative | Ref | Ref | ||
| Positive | 0.44 (0.28, 0.70) | 0.0001 | 0.48 (0.31, 0.77) | 0.0017 | |
| cfDNA at T1 | <8.11 | Ref | Ref | ||
| ≥8.11 | 1.65 (1.06, 2.59) | 0.0272 | 1.57 (1.01, 2.47) | 0.0480 | |
| cfDNA ∆T2–T1 | <−0.11 | Ref | Ref | ||
| ≥−0.11 | 1.98 (1.27, 3.08) | 0.0025 | 1.84 (1.18, 2.86) | 0.0070 | |
| VAF at T2 | <1.94 | Ref | Ref | ||
| ≥1.94 | 2.74 (1.75, 4.29) | 0.0000 | 2.46 (1.57, 3.85) | 0.0001 | |
| C-index | 0.718 | 0.707 |
Table 5.
Multivariate analysis including clinicopathological and molecular variables and their impact on OS
| OS | Multiple analysis | Shrunked | |||
|---|---|---|---|---|---|
| HR (95% CI) | p value | HR (95% CI) | p value | ||
| PD-L1 | Negative | Ref | Ref | ||
| Positive | 0.44 (0.28, 0.70) | 0.0001 | 0.50 (0.31, 0.79) | 0.0032 | |
| Histology | Non-squamous | Ref | Ref | ||
| Squamous | 3.05 (1.60, 5.83) | 0.0001 | 2.59 (1.35, 4.94) | 0.0040 | |
| cfDNA at T1 | <8.33 | Ref | Ref | ||
| ≥8.33 | 1.67 (1.05; 2.65) | 0.0308 | 1.50 (0.98; 2.47) | 0.0631 | |
| cfDNA ∆T2–T1 | <3.43 | Ref | Ref | ||
| ≥3.43 | 1.88 (1.19, 2.98) | 0.0067 | 1.71 (1.08, 2.71) | 0.0211 | |
| maxVAF at T2 | <1.94 | Ref | Ref | ||
| ≥1.94 | 2.74 (1.74, 4.33) | 0.0001 | 2.36 (1.50, 3.72) | 0.0002 | |
| C-index | 0.740 | 0.726 |
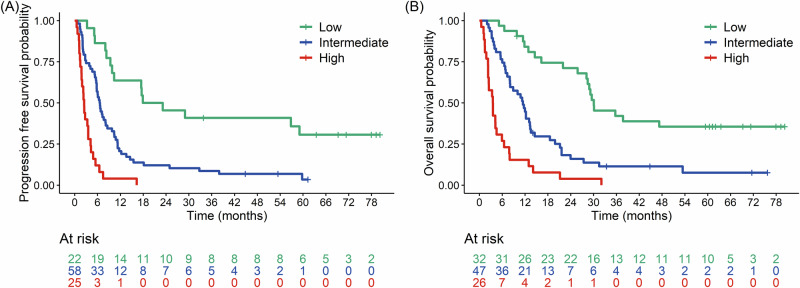
Supplementary Figs. 1 and 2 show related nomograms to assess risk factors and potentially anticipate outcomes following immunotherapy in clinical practice. Based on each patient’s total score in the nomogram, patients were stratified into three risk groups in order to early identify patients with a high probability of rapid progression and dismal outcome, patients with intermediate clinical benefit and patients likely to achieve durable clinical benefit (Fig. 1A, B and Supplementary Tables 14 and 15). In particular, the median PFS and OS for the low-risk group were 20.5 (95% CI: 10.3–NA) and 30.1 (95% CI: 28.2–NA) months, median PFS and OS for the intermediate group were 6.6 (95% CI: 5.8–8.6) and 11.4 (95% CI: 8.0–13.5) months; patients in the higher risk group had median PFS of 2.5 (95% CI: 1.8–4.2) and median OS of 3.5 (95% CI: 2.3–6.5) months.
Fig. 1. Risk stratification according to liquid biopsy results and clinicopathological features in the whole population.
A Kaplan–Meier curve for PFS of the study population stratified into three risk groups based on their individual score in the nomogram (Supplementary Figs. 1 and 2). Factors affecting PFS in multiple regression models were: cfDNA concentration in plasma at baseline, maxVAF at T2, increase (∆T2–T1) of cfDNA, and PD-L1 expression. B Kaplan–Meier curves for the OS of the study population stratified into three risk groups based on their individual score in the nomogram (Supplementary Figs. 1 and 2). Factors affecting OS in multiple regression models were: cfDNA at baseline, increased (∆T2–T1) of cfDNA, maxVAF at T2, histology and PD-L1 expression.
Integrated model and first-line treatment
Subsequently, we tried to validate our integrated prediction model in a more homogeneous clinical setting.
Fifty patients treated with first-line pembrolizumab were included in the analysis. Median PFS was 9.0 months (95% CI: 5.6–15.4), and median OS was 13.2 months (95% CI: 9.8–32.1). Twenty-one (42.0%) patients achieved a PR, while 16 patients experienced PD as the best response (Table 2).
In the multiple Cox model, high maxVAF at T2 (HR = 2.25, 95% CI: 1.15─4.41), and an increase (ΔT2–T1) of cfDNA (HR = 2.25, 95% CI: 1.17─4.33) remained significantly associated with a worse PFS (Supplementary Table 16). Similarly, high levels of maxVAF at T2 (HR = 2.09, 95% CI: 1.07─4.11) and an increase (ΔT2–T1) of cfDNA (HR = 2.18, 95% CI: 1.08─4.38) remained significantly associated with a worse OS (Supplementary Table 16).
Figure 2 depicts a three-group risk stratification model based on a nomogram created for first-line treatment.
Fig. 2. Risk stratification according to liquid biopsy results and clinicopathological features among patients treated with first-line ICIs and high PD-L1 expression.
A Kaplan–Meier curve for PFS (A) of patients treated with first-line pembrolizumab and stratified into three risk groups based on their individual scores in the nomogram. Factors affecting PFS in multiple regression models were: maxVAF at T2 and increase (∆T2–T1) of cfDNA. B Kaplan–Meier curves for the OS of patients treated with first-line pembrolizumab and stratified into three risk groups based on their individual scores in the nomogram. Factors affecting OS in multiple regression models were: maxVAF at T2 and increase (∆T2–T1) of cfDNA.
In particular, median OS was 3.3 (95% CI: 1.5–NA) months among patients considered at high risk, 6.9 (95% CI: 5.5–21.2) months among patients considered at intermediate risk and 29.5 (95% CI: 14.6–NA) among patients considered at low risk (Fig. 2 and Supplementary 17–18).
Among patients achieving RR according to RECIST V1.1, the results of liquid biopsy, performed at the time of radiological evaluation (T3), were able to anticipate long-term clinical benefit. In particular, high maxVAF at T3 was associated with a median PFS of 7.0 (95% CI: 12.2–32.1) months and a median OS of 12.8 (95% CI: 6.5–NA) months vs a median PFS of 17.5 (95% CI: 10.3–37.90) months and a median OS of 35.7 (95% CI: 14.6–NA) months for patients with low maxVAF at T3 (Supplementary Tables 19 and 20).
Discussion
Predicting the clinical benefit of immunotherapy in advanced NSCLC is recognized as one of the main challenges for clinical and translational research in oncology.
The potentialities of using dynamic liquid biopsy during treatment with ICIs have already been discussed elsewhere17,21–24. Some data have already been published showing how changes in ctDNA during treatment with ICIs might be associated with outcomes in advanced NSCLCs and other solid tumours13,16,25–28. Even though the idea of using ctDNA early monitoring as surrogate markers of tumour burden during treatment is increasingly accepted by the scientific community, available data are characterized by high heterogeneity in methods, on-treatment time-point evaluation, patient population and clinical end-points. A recent analysis pooled and harmonized data from 200 patients included in five different trials and showed that the predictive role of dynamic analysis of ctDNA can be confirmed29. Nevertheless, a practical model to be tested in an interventional clinical trial and afterwards applied in clinical practice is still awaited.
In our study, we prospectively included a single-centre population of advanced NSCLC patients, treated according to clinical practice (outside interventional clinical trial). All the patients performed liquid biopsy at the pre-established time-points and we included in our integrated analysis only patients with all clinicopathological data available, including PD-L1 expression in tissue at baseline and lack for driver alterations, commonly tested in clinical practice in tissue.
As far as molecular variables are concerned, we first tested multiple candidates (mean and median VAF of all alterated genes as reference, maxVAF as reference absolute and relative changes in ctDNA and VAF assessed by different methods), which were all consistent in association with clinical outcome endpoints (data not shown) and selected the one most likely to be applicable to clinical practice (cfDNA and maxVAF as static parameter and absolute change in ctDNA and maxVAF as dynamic parameter). Consistently, we decided to use a targeted NGS kit including a panel designed to target genes recurrently mutated in tumours in order to focus on quantitative variations in plasma and increase the potential applicability of our results in clinical practice. Also, the choice of time points for liquid biopsy has been made taking into account clinical feasibility (i.e. when a patient came to the hospital to perform routine blood assessment). T2 has been chosen as the earliest time-point not requiring additional visits with respect to the planned treatment administration time-point, with the final aim to identify as early as possible patients who do not benefit from ICI. T3 has been planned at the time of radiological evaluation, mostly after two cycles, considered elsewhere as the optimal time-point30.
The selected liquid biopsy variables were associated with standard commonly recognized end-points (PFS, OS, RR according to RECIST v1.1) and afterwards integrated with clinicopathological features able to affect the same endpoints.
Notable, even among patients achieving RR according to RECIST v1.1, liquid biopsy results were able to further stratify patients and identify those with differential duration of clinical benefit, thus underlining the limits of using RR as a surrogate endpoint and as an indicator for clinical communication and follow-up planning during treatment.
Moreover, in our integrated model, we clearly demonstrated that longitudinal cfDNA analysis adds further information when commonly used clinicopathological predictive factors are available. In parallel, we demonstrated the role of evaluating both cfDNA concentration and NGS results. In this context, in a recent paper focussed on early identification of patients potentially deriving “detrimental effects” from ICI treatment, we previously demonstrated that cfDNA quantification can be considered a surrogate of tumour burden or biological changes induced by treatment13. On the other hand, we show here that NGS results, evaluated by maxVAF values and variations, are able to further stratify the risk of death and recurrence.
The most important strengths of our study are the methodology of enrolment and the potential applicability of the model. Notably, the choice of an early time-point (T2, three weeks after the start of ICIs) permits figuring out the potential applicability of the results on addressing early changes during first-line systemic treatment, with the final aim to maximize treatment efficacy and avoid to administer more toxic combination strategies when not needed.
On the other hand, the most important limitation of our study is the lack of a validation set of analyses to confirm the results. However, we performed a cross-validation within the dataset to assess the robustness of the optimal cut-points and we estimated the adjusted HR for the cut-points using a bootstrap shrinkage procedure to reduce overfitting.
In addition, further predictive information could be provided by including in the model qualitative information about genetic characterization, in order to test the potential influence of specific genetic alterations on the response to immunotherapy, such as STK11, KEAP1 and KRAS mutations31–34. Anyway, we believe that starting from information commonly available in clinical practice might help the applicability of the model.
Potential future clinical applications of the model in advanced NSCLC patients include the possibility to customize the first-line treatment approach by intensifying treatment after the first two cycles only in patients at high risk for short PFS and OS. As a matter of fact, in our series, we also estimated our model in a more homogenous population, being patients expressing PD-L1 ≥ 50% in tumour cells and treated in the first line with pembrolizumab. A second potential application includes the possibility of early switch to second-line treatment among non-progressing patients at high risk of relapse according to T3 liquid biopsy results.
In conclusion, we demonstrated the role of early monitoring of cfDNA concentration and maxVAF quantification in plasma in predicting the outcome of patients treated with ICIs and our integrated model warrants further validation as a tool to customize treatment in non-oncogene addicted NSCLC.
Supplementary information
Acknowledgements
This work was funded by the IOV intramural research grant 2017–5 × 1000 (MAGIC-2 to S.I. and L.B.) and Ricerca Corrente funding from the Italian Ministry of Health.
Author contributions
A.B.B.: methodology, validation, investigation, writing—original draft and writing—review and editing and visualisation. P.D.B.: methodology, validation, formal analysis, data curation, writing—original draft and writing—review and editing. E.Z.: methodology and investigation. I.A.: resources, data curation and editing. A.P.: resources and data curation. J.C.: investigation. L.M.: investigation. V.G.: resources. S.I.: conceptualisation, methodology, writing—review and editing, supervision, project administration and funding acquisition. L.B.: conceptualisation, methodology, data curation, writing—original draft, writing—review and editing, visualisation, resources, supervision, project administration and funding acquisition.
Data availability
The molecular dataset is summarized in supplementary material and raw FASTQ sequencing files are available in the NCBI Sequence Read Archive database within the BioProject accession number PRJNA1086903 at the following link: https://www.ncbi.nlm.nih.gov/sra/PRJNA1086903.
Competing interests
The authors declare no competing interests.
Consent for publication
The property of data belongs to the sponsor IOV and the authors have the right to publish the data.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
The online version contains supplementary material available at 10.1038/s41698-024-00704-9.
References
- 1.Hendriks, L. E. et al. Non-oncogene-addicted metastatic non-small-cell lung cancer: ESMO clinical practice guideline for diagnosis, treatment and follow-up. Ann. Oncol.34, 358–376 (2023). [DOI] [PubMed] [Google Scholar]
- 2.Govindan, R. et al. Society for immunotherapy of cancer (SITC) clinical practice guideline on immunotherapy for the treatment of lung cancer and mesothelioma. J. Immunother. Cancer10, e003956 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Hendriks, L. E. et al. Oncogene-addicted metastatic non-small-cell lung cancer: ESMO clinical practice guideline for diagnosis, treatment and follow-up☆. Ann. Oncol.34, 339–357 (2023). [DOI] [PubMed] [Google Scholar]
- 4.Reck, M. et al. Five-year outcomes with pembrolizumab versus chemotherapy for metastatic non-small-cell lung cancer With PD-L1 tumor proportion score ≥ 50. J. Clin. Oncol.39, 2339–2349 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Gandhi, L. et al. Pembrolizumab plus chemotherapy in metastatic non-small-cell lung cancer. N. Engl. J. Med.378, 2078–2092 (2018). [DOI] [PubMed] [Google Scholar]
- 6.Mok, T. S. K. et al. Pembrolizumab versus chemotherapy for previously untreated, PD-L1-expressing, locally advanced or metastatic non-small-cell lung cancer (KEYNOTE-042): a randomised, open-label, controlled, phase 3 trial. Lancet Lond. Engl.393, 1819–1830 (2019). [DOI] [PubMed] [Google Scholar]
- 7.Paz-Ares, L. et al. First-line nivolumab plus ipilimumab combined with two cycles of chemotherapy in patients with non-small-cell lung cancer (CheckMate 9LA): an international, randomised, open-label, phase 3 trial. Lancet Oncol.22, 198–211 (2021). [DOI] [PubMed] [Google Scholar]
- 8.Herbst, R. S. et al. Atezolizumab for first-line treatment of PD-L1-selected patients with NSCLC. N. Engl. J. Med.383, 1328–1339 (2020). [DOI] [PubMed] [Google Scholar]
- 9.Sezer, A. et al. Cemiplimab monotherapy for first-line treatment of advanced non-small-cell lung cancer with PD-L1 of at least 50%: a multicentre, open-label, global, phase 3, randomised, controlled trial. Lancet Lond. Engl.397, 592–604 (2021). [DOI] [PubMed] [Google Scholar]
- 10.Paz-Ares, L. et al. Pembrolizumab plus chemotherapy for squamous non-small-cell lung cancer. N. Engl. J. Med.379, 2040–2051 (2018). [DOI] [PubMed] [Google Scholar]
- 11.Ferrara, R. et al. Hyperprogressive disease in patients with advanced non-small cell lung cancer treated With PD-1/PD-L1 inhibitors or with single-agent chemotherapy. JAMA Oncol.4, 1543–1552 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Peters, S. et al. OA03.05 analysis of early survival in patients with advanced non-squamous NSCLC treated with nivolumab vs docetaxel in CheckMate 057. J. Thorac. Oncol.12, S253 (2017). [Google Scholar]
- 13.Zulato, E. et al. Longitudinal liquid biopsy anticipates hyperprogression and early death in advanced non-small cell lung cancer patients treated with immune checkpoint inhibitors. Br. J. Cancer127, 2034–2042 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hegde, P. S., Karanikas, V. & Evers, S. The where, the when, and the how of immune monitoring for cancer immunotherapies in the era of checkpoint inhibition. Clin. Cancer Res.22, 1865–1874 (2016). [DOI] [PubMed] [Google Scholar]
- 15.Waldman, A. D., Fritz, J. M. & Lenardo, M. J. A guide to cancer immunotherapy: from T cell basic science to clinical practice. Nat. Rev. Immunol.20, 651–668 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zulato, E. et al. Early assessment of KRAS mutation in cfDNA correlates with risk of progression and death in advanced non-small-cell lung cancer. Br. J. Cancer123, 81–91 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bonanno, L. et al. Liquid biopsy and non-small cell lung cancer: Are we looking at the tip of the iceberg? Br. J. Cancer127, 383–393 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med.17, 405–424 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer45, 228–247 (2009). [DOI] [PubMed] [Google Scholar]
- 20.Champiat, S. et al. Hyperprogressive disease is a new pattern of progression in cancer patients treated by anti-PD-1/PD-L1. Clin. Cancer Res.23, 1920–1928 (2017). [DOI] [PubMed] [Google Scholar]
- 21.Merker, J. D. et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists Joint Review. J. Clin. Oncol.36, 1631–1641 (2018). [DOI] [PubMed] [Google Scholar]
- 22.Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol.14, 531–548 (2017). [DOI] [PubMed] [Google Scholar]
- 23.Wan, J. C. M. et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer17, 223–238 (2017). [DOI] [PubMed] [Google Scholar]
- 24.Sivapalan, L. et al. Liquid biopsy approaches to capture tumor evolution and clinical outcomes during cancer immunotherapy. J. Immunother. Cancer11, e005924 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cabel, L. et al. Circulating tumor DNA changes for early monitoring of anti-PD1 immunotherapy: a proof-of-concept study. Ann. Oncol.28, 1996–2001 (2017). [DOI] [PubMed] [Google Scholar]
- 26.Raja, R. et al. Early reduction in ctDNA predicts survival in patients with lung and bladder cancer treated with durvalumab. Clin. Cancer Res.24, 6212–6222 (2018). [DOI] [PubMed] [Google Scholar]
- 27.Hwang, M. et al. Peripheral blood immune cell dynamics reflect antitumor immune responses and predict clinical response to immunotherapy. J. Immunother. Cancer10, e004688 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Assaf, Z. J. F. et al. A longitudinal circulating tumor DNA-based model associated with survival in metastatic non-small-cell lung cancer. Nat. Med.29, 859–868 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Vega, D. M. et al. Changes in circulating tumor DNA reflect clinical benefit across multiple studies of patients with non-small-cell lung cancer treated with immune checkpoint inhibitors. JCO Precis. Oncol. 10.1200/PO.21.00372 (2022). [DOI] [PMC free article] [PubMed]
- 30.Anagnostou, V. et al. ctDNA response after pembrolizumab in non-small cell lung cancer: phase 2 adaptive trial results. Nat. Med.29, 2559–2569 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pavan, A. et al. Role of next generation sequencing-based liquid biopsy in advanced non-small cell lung cancer patients treated with immune checkpoint inhibitors: impact of STK11, KRAS and TP53 mutations and co-mutations on outcome. Transl. Lung Cancer Res.10, 202–220 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lim, T. K. H. et al. KRAS G12C in advanced NSCLC: prevalence, co-mutations, and testing. Lung Cancer184, 107293 (2023). [DOI] [PubMed] [Google Scholar]
- 33.Skoulidis, F. et al. STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov.8, 822–835 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ricciuti, B. et al. Diminished efficacy of programmed death-(Ligand)1 inhibition in STK11- and KEAP1-mutant lung adenocarcinoma is affected by KRAS mutation status. J. Thorac. Oncol.17, 399–410 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The molecular dataset is summarized in supplementary material and raw FASTQ sequencing files are available in the NCBI Sequence Read Archive database within the BioProject accession number PRJNA1086903 at the following link: https://www.ncbi.nlm.nih.gov/sra/PRJNA1086903.