Abstract
To determine how neuronal circuits encode and drive behavior, it is often necessary to measure and manipulate different aspects of neurochemical signaling in awake animals. Optogenetics and calcium sensors have paved the way for these types of studies, allowing for the perturbation and readout of spiking activity within genetically defined cell types. However, these methods lack the ability to further disentangle the roles of individual neuromodulator and neuropeptides on circuits and behavior. We review recent advances in chemical biology tools that enable precise spatiotemporal monitoring and control over individual neuroeffectors and their receptors in vivo. We also highlight discoveries enabled by such tools, revealing how these molecules signal across different timescales to drive learning, orchestrate behavioral changes, and modulate circuit activity.
A central goal in molecular systems neuroscience is to determine what circuits, molecules, and proteins are responsible for driving behaviors of interest (1). The development of genetically encoded fluorescent indicators and their use in vivo in animal models allows visualization of spiking activity through the monitoring of intracellular calcium influx (2) or voltage-dependent membrane protein conformational changes (3) in response to an animal’s surrounding environment. Then, tools such as optogenetics (4) and chemogenetics (5–7) can be used to perturb these natural signaling dynamics to determine how they contribute to behavior. These established toolboxes of genetically encoded actuators and sensors, in combination with advanced viral targeting strategies (8), have transformed our ability to interrogate the functional roles of specific neuronal pathways and cell types in the mammalian brain.
However, numerous other types of signaling within neuronal circuits are involved in driving learned and innate behaviors—including neuromodulator and neuropeptide signaling. Neuromodulators and neuropeptides primarily act upon G protein-coupled receptors (GPCRs), which are a large superfamily of membrane proteins with a highly conserved structure consisting of seven transmembrane helices, three extracellular loops and three intracellular loops, and an intracellular C terminus, which interacts with G protein subunits (9). Ligand binding to the extracellular N terminus of the receptor induces a conformational change, leading to G protein activation and dissociation and the initiation of intracellular signaling cascades that modulate excitability, synaptic strength, and other network properties (10–13). Neuromodulators such as dopamine, serotonin, acetylcholine, and norepinephrine play critical roles in regulating behavioral brain states related to motivation (14), mood (15), and alertness (16). They also modulate synaptic plasticity to facilitate various types of learning (17). In addition, there are more than 100 known distinct neuropeptides (18), which are released by different neuronal cell types (19, 20). Neuropeptides are crucial for driving metabolic states, including hunger (neuropeptide Y) (21–23) and thirst (angiotensin II) (24, 25), and for driving innate behaviors such as maternal care (oxytocin) (26, 27).
Despite their importance in modulating behavior and neuronal circuit activity, until recently, relatively few technologies existed that enabled the study of neuromodulator and neuropeptide signaling within cell types in behaving animals (28–30). Several challenges specific to the diversity of their receptor targets, their release mechanisms, and their transmission properties in the brain (19, 31) hindered the development of such technologies. First, a given neuromodulator or neuropeptide can bind multiple receptors present across different brain regions. For example, dopamine can bind five different receptor subtypes (32), which recruit different G proteins and are associated with distinct behavioral phenotypes (33, 34). Second, neuromodulators and neuropeptides can be released by many types of neurons in different brain regions and are often co-released with several other neurochemicals (35–38). Third, unlike fast-acting neurotransmitters, both neuromodulators and neuropeptides can signal through volume transmission, traveling perisynaptically to bind their receptors (39). Moreover, whereas neuromodulators are typically released from small clear vesicles at axon terminals, neuropeptides are packaged into dense core vesicles that can be released from the soma, axons, or dendrites (19), making detection of their sparse release in vivo challenging. Therefore, recording or manipulating the spiking output of any given cell type alone is not sufficient to determine the signaling properties of specific neuromodulators or neuropeptides. Instead, it would be ideal to have tools that can monitor distinct classes of signaling molecules within a given brain region, and tools to manipulate the associated receptor signaling pathways in a temporally specific and cell type specific manner. In this review, we will focus on new tools leveraging chemical biology that enable us to record when and where neurochemicals are released, and to causally induce ligand and receptor signaling in awake and behaving animals. We will also highlight how these technologies allowed us to elucidate key physiological and pathological brain functions, as well as what future advancements are on the horizon for targeting and monitoring neurochemical signaling in vivo across the brain.
Sensing neuromodulators and neuropeptides
Fast-scan cyclic voltammetry has been considered the gold standard to measure neuromodulator release in awake and behaving animals (28, 30, 40). But despite having fast kinetics, it lacks the cellular and spatial specificity of genetically encoded activity probes, and it is difficult to isolate the release of individual neuromodulators because of their structural similarity and overlapping oxidation profiles. Biological sensors, including cell-based neurotransmitter fluorescence engineered reporters (CNiFERs) have also been developed to detect neuromodulator binding in vivo (41, 42); however, these require the infusion of genetically modified cells into the brain. With the advent of transgenic mouse strains for targeting specific neuromodulator- or neuropeptide-releasing cell populations, it became feasible to record the spiking activity of these cells in vivo by using optotagging and electrophysiology (43) or by using a calcium indicator and fiber photometry (44–46) or single-cell imaging (47, 48). Unfortunately, these approaches cannot provide a readout of the timescales and locations of neuromodulator or neuropeptide release.
Here, we describe a new generation of sensors that allows for the imaging of neuromodulator and neuropeptide dynamics in vivo. We also describe different classes of GPCR integrators, which have the capacity for stably tagging and manipulating neurons that have undergone receptor activation during a behaviorally-relevant time window.
Fluorescent neuromodulator and neuropeptide sensors
Three main classes of fluorescent neurochemical sensors adopted for use in mammalian brain tissue will be discussed here: (i) periplasmic binding protein (PBP) based sensors using the sensing fluorescent reporter (SnFR) design, (ii) GPCR-based sensors using the GPCR activation-based (GRAB) and Light designs, and (iii) functionalized single-wall carbon nanotubes (SWCNTs) (Fig. 1A).
Fig. 1. Fluorescent sensors for neuromodulator and neuropeptide release.
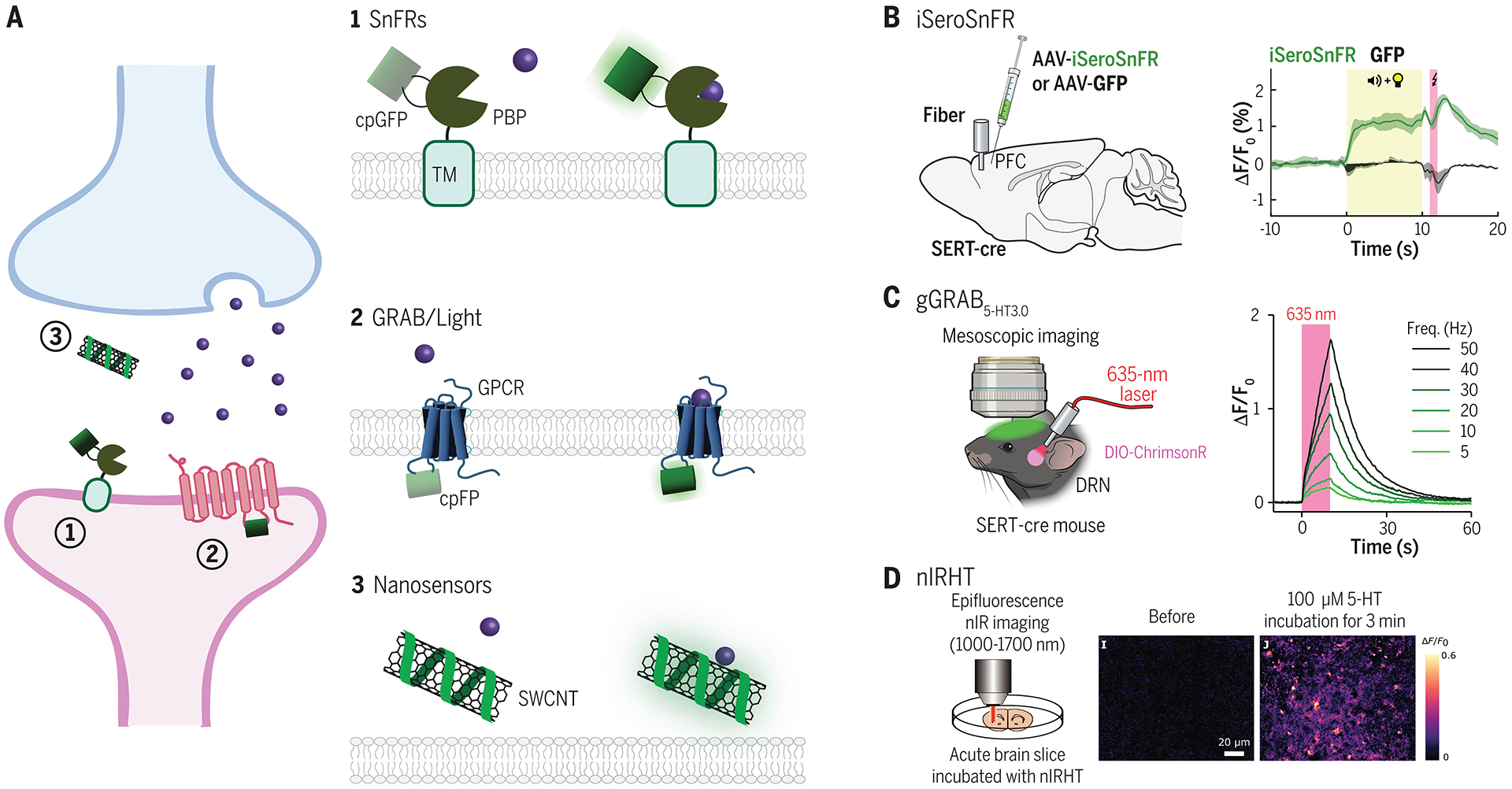
(A) Overview of main classes of sensor scaffolds used in brain tissue: 1. SnFRs are composed of a ligand-binding domain based on bacterial periplasmic binding proteins (PBP), which is fused to a circularly permutated GFP (cpGFP) and a transmembrane domain (TM). The cpGFP is located extracellularly. Upon ligand binding, a conformational change in the PBP binding pocket increases fluorescence emission of the cpGFP. 2. GRAB/Light sensors fuse a cpFP within the third intracellular loop of a GPCR. Ligand binding induces a conformational change leading to increased cpFP fluorescence. 3. Nanosensors are derived from polymer-functionalized semiconducting single-wall carbon nanotubes (SWCNTs), which are decorated with single stranded oligonucleotides that give the sensor affinity for specific ligands. Ligand binding alters the innate fluorescence of the SWCNT. (B) iSeroSnFR is a serotonin sensor based on an evolved PBP for acetylcholine. Because of its fast kinetics, iSeroSnFR can be used to record acute changes in serotonin dynamics during naturalistic behaviors such as aversive cue learning. Data shown from (57). (C) GRAB 5-HT3.0 (available as both a green and red version) is an improved serotonin sensor based on insertion of a cpFP into the third intracellular loop of the serotoning type 4 (5-HT4R) subtype. With a high affinity for serotonin, it can be used in vivo to monitor endogenous serotonin release. Either the red or green version can be combined with optogenetics to enable simultaneous read-out and control of serotonin signaling in vivo. Data shown from (75). (D) nIRHT is a SWCNT sensor for serotonin. It emits fluorescence in a nIR window at around 1200 nm. It has been validated in ex vivo mouse slice during bath serotonin application. Data shown from (86). See Table 1 for a list of genetically encoded neuromodulator and neuropeptide sensors designed for mammalian neuronal circuit studies.
PBP-based sensors.
One of the earliest engineered fluorescent sensors to image neurochemical release in vivo is iGluSnFR (49), which has since been optimized to enable the detection of glutamate release after a single action potential (50–52). This sensor utilizes a ligand-binding scaffold of a bacterial PBP, in this case Glt1, fused to a circularly permutated green fluorescent protein (cpGFP) located outside the extracellular membrane. Upon binding glutamate, a conformational change in the ligand-binding pocket increases the fluorescence of the cpGFP. PBPs, such as the one used in iGluSnFR, are attractive scaffolds for detecting neurochemicals for a number of reasons. First, because they come from bacterial organisms, they are unlikely to interact with any endogenous signaling pathways in model organisms or bind drugs that interact with these pathways (53, 54). Furthermore, they exhibit large conformational changes upon binding and thus have a large dynamic range (55). Using this backbone, SnFR tools have been expanded to image the dynamics of other neuromodulators and neurotransmitters, such as acetylcholine [iAchSnFR (53)] and γ-aminobutyric acid (GABA) [(iGABASnFR (56)].
A disadvantage with PBPs is that there are few that target neuromodulators present in model organisms, such as dopamine and serotonin. To overcome this limitation, machine learning techniques have been used to redesign the binding pocket of iAchSnFR to instead bind serotonin, giving rise to iSeroSnFR (57) (Fig. 1B). iSeroSnFR can detect endogenous serotonin release during behaviors such as sleep and fear learning. Studies have used it to reveal serotonin dynamics in the central amygdala during fear learning and recall (58), changes in serotonergic release in the orbitofrontal cortex after early life stress (59), and serotonin release in the nucleus accumbens (NAc) during social interactions (60). However, despite the success of SnFR sensors, re-engineering PBPs to bind a variety of different neuromodulators or neuropeptides remains a considerable challenge.
GPCR-based sensors.
Alternatively, GPCR modules can be used as ligand-binding scaffolds, as is the case with GRAB and Light sensors. GPCRs act as suitable sensor scaffolds, given the ability to tune their specificity and affinity for endogenous ligands through protein engineering. dLight1, a genetically encoded dopamine sensor, was developed by inserting a cpGFP into the third intracellular loop of the human type 1 dopamine receptor (61), effectively coupling a dopamine-induced conformational change to an increase in fluorescence emission. The cpGFP insertion site also blocks endogenous G protein binding, so that these sensors do not induce intracellular signaling cascades. dLight1 was validated in vivo by recording elevated fluorescence emission in response to reward, reward-predicting cues, and stimulation of dopamine releasing neurons (61). It has also been applied by other groups to demonstrate the role of dopamine in encoding saliency in the NAc core (62) and to elucidate dopamine dynamics in the dorsal and ventral striatum during self-paced exploratory behavior (63). By replacing the dopamine receptor component with other GPCRs, additional sensors for norepinephrine (64), serotonin (65, 66), and neuropeptides such as opioids (61, 67), gastrin-releasing peptide (68), and orexin (69) have also been engineered for in vivo use (Table 1).
Table 1. Genetically encoded neuromodulator and neuropeptide sensors.
A summary of recently developed fluorescent sensors for detecting neuromodulators, neuropeptides, or GPCR agonists in mammalian brain tissue. For a given sensor, only the most recently improved variant that has been validated is listed.
| Neurochemical | Sensor name | Initial in vivo validation in rodents | Ref. |
|---|---|---|---|
| Neuromodulators | |||
| Acetylcholine (Ach) | iAchSnFR | visual cortex, motor cortex, hippocampus | (53) |
| GRABAch3.0 | basolateral amygdala, hippocampus, somatosensory cortex, visual cortex | (77) | |
| Dopamine (DA) | dLight1.1 | nucleus accumbens | (61) |
| dLight1.2 | frontal/motor cortex | (61) | |
| dLight1.3b | nucleus accumbens | (29, 61) | |
| RdLight1 | nucleus accumbens | (82) | |
| gGRABDA3m | substantia nigra pars compacta | (73) | |
| gGRABDA3h | motor cortex | (73) | |
| rGRABDA2m | medial prefrontal cortex, nucleus accumbens, basolateral amygdala | (73) | |
| rGRABDA3m | medial prefrontal cortex, nucleus accumbens | (73) | |
| Histamine (HA) | GRABHA1m | preoptic area | (149) |
| GRABHA1h | medial prefrontal cortex | (149) | |
| Norepinephrine (NE) | GRABNE2m | locus coeruleus, medial prefrontal cortex, lateral hypothalamus, preoptic area, whole cortex | (79) |
| GRABNE2h | locus coeruleus, lateral hypothalamus | (79) | |
| nLightG | lateral hypothalamus, dorsal hippocampus | (64) | |
| nLightR | lateral hypothalamus | (64) | |
| Serotonin (5-HT) | iSeroSnFR | medial prefrontal cortex, dorsal striatum, orbitofrontal cortex, bed nucleus of the stria terminalis, basolateral amygdala | (57) |
| psychLight2 | dorsal raphe nucleus, bed nucleus of the stria terminalis, basolateral amygdala, orbitofrontal cortex, medial prefrontal cortex | (66) | |
| sDarken | prefrontal cortex | (65) | |
| gGRAB5-HT3.0 | whole cortex, basal forebrain, basal amygdala | (75) | |
| rGRAB5-HT1.0 | whole cortex, basal forebrain | (75) | |
| Neuropeptides | |||
| Corticotropin releasing factor (CRF) | GRABCRF1.0 | paraventricular nucleus, prefrontal cortex, motor cortex | (80) |
| Dynorphins | kLight1.3 | arcuate nucleus, hippocampus, nucleus accumbens | (67) |
| kLight1.3a | nucleus accumbens | (67) | |
| Enkephalins; β-endorphin | deltaLight | arcuate nucleus, nucleus accumbens | (67) |
| Gastrin-releasing peptide (GRP) | grpLight | auditory cortex | (68) |
| Orexins (Ox) | OxLight1 | nucleus accumbens, posterior insular cortex, basal forebrain, lateral hypothalamus, somatosensory cortex | (69) |
| Oxytocin (OT) | MTRIAOT | anterior olfactory nucleus, central amygdala | (150) |
| GRABOT1.0 | bed nucleus of the stria terminalis, ventral tegmental area, paraventricular nucleus, prefrontal cortex | (81) | |
| Somatostatin (SST) | GRABSST1.0 | basolateral amygdala | (80) |
GRAB sensors, which were developed at the same time as Light sensors, use a strategy similar to that of Light sensors. Both sensor classes are widely used to record neurochemical dynamics in vivo, differing mainly in the specific receptor subtypes used as scaffolds. GRABDA1 is an engineered human type 2 dopamine receptor also with a cpGFP inserted into its third intracellular loop. The original GRABDA1 variants were used in mice to image dopamine dynamics during aversive and rewarding stimuli, during male mating behaviors (70), and during sexual receptivity (71). Through modifications to the cpGFP and the use of different engineered dopamine receptors, the GRABDA sensors have since been improved to generate GRABDA2 (72) and gGRABDA3 variants (73). GRAB sensors have also been modified to detect the release of other neuromodulators, such as serotonin (Fig. 1C) (74, 75), acetylcholine (76, 77), and norepinephrine (78, 79). More recently, sensors have been designed for several neuropeptides as well, although only the somatostatin (SST) and corticotropin releasing factor (CRF) sensors (80) and the oxytocin sensor (81) have been tested so far in vivo (Table 1).
One limitation of these initial sensors was the lack of color-shifted variants, thus making it challenging to perform multiplexed imaging of different neurochemicals. Recently, a red-shifted RdLight1 (82) was introduced and combined with iGluSnFR to image both glutamate and dopamine dynamics in NAc. Red shifted nLightR (64), AChLightR (64), rGRABDA versions 1 to 3 (72, 73), and rGRAB5-HT1.0 (75) have also been engineered. However, a caveat to mApple-based fluorescent sensors such as RdLight1 and rGRABDA3 is that they exhibit photoswitching when activated by blue light, leading to artifactual shifts in the fluorescent signal that resemble neuromodulator signaling (73, 83). The rGRAB5-HT1.0 (75) as well as the rGRABDA1 and rGRABDA2 variants (73) have shown negligible photoactivation in cultured neurons (with the potential for a small negative-going photo artifact at higher blue light intensities). Thus, it is necessary to optimize the design of these sensors and carefully validate their performance in vivo before use for dual-color imaging and optogenetics.
SWCNT sensors.
Nanosensors represent another class of fluorescent sensors; they are not genetically encoded but have the advantage of emitting in the near-infrared (nIR) spectrum, potentially allowing for non-invasive imaging in the future (84). They consist of a backbone of polymer-functionalized semiconducting SWCNTs onto which single-stranded oligonucleotides are pinned, giving the sensor affinity for specific ligands. Beyenne et al. (85) used this design to create nIRCats, a nIR fluorescent sensor for catecholamines such as dopamine and norepinephrine. nIRCats demonstrated high affinity and sensitivity for dopamine after optogenetic stimulation of dopaminergic terminals (85). The same group designed a similar probe for serotonin, nIRHT, which proved effective ex vivo in mouse slice (Fig. 1D) (86). However, this tool has not yet been implemented in vivo in the brain. More recently, Xia et al. designed a nanosensor for Ach, which they used in vivo to measure Ach release in the peripheral nervous system (87). These nanosensors are fully synthetic and bioorthogonal, which make them well-suited for use in non-model organisms lacking genetic-targeting strategies. But a trade-off of theses sensors is that it is not possible to target them to specific cell types.
Applications of fluorescent sensors for biological discovery.
Both dLight and GRABDA, the first described GPCR sensors for dopamine, have now been broadly adopted in numerous biological settings. In addition, newer sensor variants have also been used to study the in vivo release of neuropeptides and other neuromodulators during specific behaviors. Recently, the delta-opioid receptor sensor, DeltaLight (67), was targeted to the arcuate hypothalamic nucleus (ARC) in mice to monitor opioid levels after feeding (Fig. 2A,B) (88). Although it was known that consuming palatable food increases endogenous opioid release in mice [as measured through serum and cerebrospinal fluid (89)], the precise dynamics and location of opioidergic signaling during food consumption had yet to be explored. Using deltaLight fiber photometry recordings, Sayar-Atasoy et al. showed that opioidergic signaling in the ARC increased during feeding in fasted mice, but not in ad libitum fed mice (Fig. 2C,D). As expected, consumption of palatable food still increased signaling in ad libitum fed mice. These data demonstrated that opioidergic signaling in the ARC is sensitive to feeding on the basis of either the nutritional status of the animal, or the palatability of the food.
Fig. 2. Application of sensors to study neurochemical signaling in vivo.
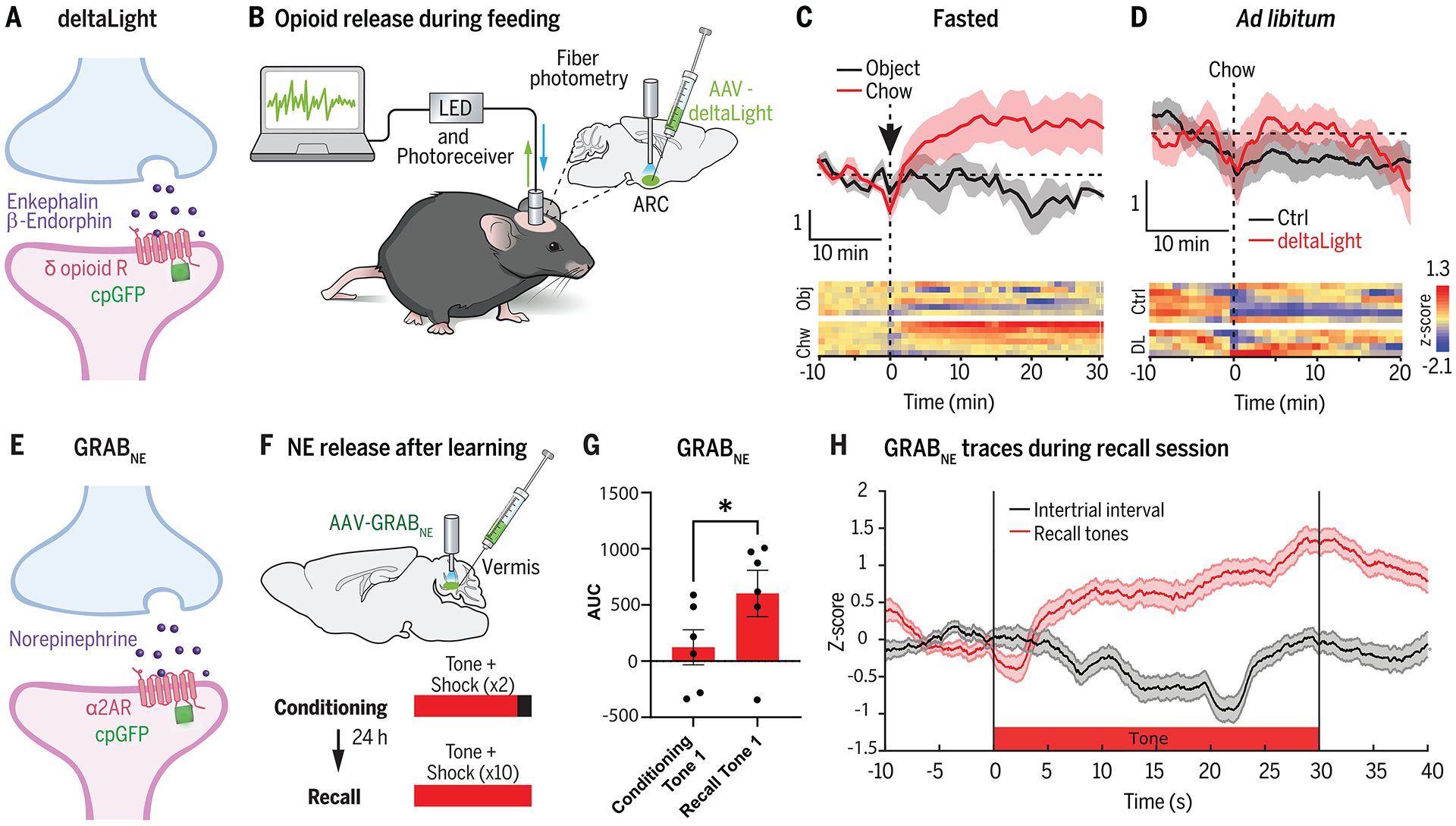
(A) DeltaLight (67) is an engineered delta-opioid receptor that acts as a sensor for endogenous enkephalin and β-endorphin neuropeptides. (B) DeltaLight was targeted to the arcuate hypothalamic nucleus (ARC) by using an adeno-associated virus (AAV) injection to monitor opioid levels after feeding using fiber photometry. (C,D) DeltaLight fluorescence increases were observed in fasted mice after food presentation compared with presentation of an inedible object. However, non-fasted mice fed ad libitum did not show an increase in deltaLight fluorescence to food presentation. Data shown from (88). (E) GRABNE (78) is an engineered adrenergic receptor which detects endogenously released norepinephrine (NE). (F) GRABNE was injected into the cerebellar vermis of mice to record fluorescence emission during fiber photometry. Mice underwent a cued fear conditioning learning paradigm, and NE responses were recorded during the tone presentations. (G) There was an increase in NE release to the tone after fear conditioning, as measured by the area under the curve (AUC) of the GRABNE photometry traces. Data shown from (90). (H) Time course of NE release during the tone during the recall session (compared with intertrial intervals with no tone). Data shown from (90).
Another recent study used the norepinephrine sensor GRABNE (78) in the cerebellar vermis of mice to study dynamics of norepinephrine release during fear learning (90). Previous immunohistochemistry studies had mapped tyrosine hydroxylase axon terminals in the cerebellum (91); however, it remained unclear what role cerebellar norepinephrine played in learning. Stanley et al. (90) recorded GRABNE activity in the cerebellum using by fiber photometry during a cued-tone fear conditioning paradigm (Fig. 2E,F). They found that although there was initially no norepinephrine response to the tone prior to learning, after fear conditioning, the tone elicited an increase in cerebellar norepinephrine release (Fig. 2G,H). These data show that fear learning can induce changes in norepinephrine signaling in the cerebellum, providing an important neuronal substrate to consider during fear learning.
Lastly, several sensors have been engineered to detect drug-induced changes in neurochemical signaling. psychLight2 (66), a fluorescent sensor based on the serotonin type 2A receptor (5-HT2AR), measures the binding dynamics of serotonergic psychedelics and can predict the behavioral head-twitch potency—a potential indication of hallucinations—of these drugs in mice. This sensor was used to screen for a non-hallucinogenic psychedelic analog with antidepressant-like effects. Additionally, the kLight sensor (67), which is based on the kappa-opioid receptor, was used to track morphine withdrawal–induced dynorphin release in the prefrontal cortex (92). Thus, these sensors are useful not only for monitoring endogenous neuromodulator and neuropeptide release but also for performing novel drug discovery and for tracking receptor-binding dynamics of different drugs in vivo (Table 1).
Integrators
Real-time fluorescent sensors are powerful tools for recording in vivo neurochemical dynamics during behavior. However, their activity readout is transient, and they do not provide the ability to subsequently manipulate or investigate the subset of neurons undergoing GPCR activation. Furthermore, because of imaging constraints, it is not currently feasible to perform whole-brain sensor imaging in the mammalian brain. Instead, integrators stably tag neurons undergoing GPCR activation during user-defined time windows, allowing for the expression of fluorescent tags or optogenetic channels for subsequent manipulation and investigation.
Transcriptional integrators.
Two transcriptional variants of GPCR-based integrators have been described thus far: SPARK (93, 94) and iTango2 (95). Both of these integrators are built off of the TANGO system (96), which has been extensively applied in cultured cells to identify drug-GPCR interactions. Thus far, only iTango2 has been applied in vivo (95). In both approaches, the C terminus of a GPCR is fused to a blue light–sensing protein, AsLOV2, which cages a tobacco etch virus (TEV) protease cleavage site that lies upstream of a non-native transcription factor. In the presence of blue light, AsLOV2 undergoes a conformational change to render the TEV cleavage site accessible for cleavage, but only if the TEV-protease is nearby. To bring the TEV protease within close proximity of its cleavage site in a ligand-dependent manner, the TEV protease is fused to a β-arrestin protein (which is recruited only to ligand-activated GPCRs). In the iTango2 design, a split version of the TEV protease is used to further reduce background protease activity—with the C-terminal half of TEV tethered to β-arrestin, and the N-terminal half of TEV inserted between the GPCR and the AsLOV2. Thus, in iTango2, the split TEV protease is only fully reconstituted upon ligand-GPCR activation (Fig. 3A). For both tools, in the presence of blue light and ligand-GPCR activation, the TEV protease cuts its cleavage site, releasing the transcription factor from the membrane so that it can enter the nucleus and drive reporter gene activation. The light inducibility of these tools renders them to be temporally specific, meaning that they can tag cells that are undergoing GPCR modulation within an experimenter-defined time window.
Fig. 3. Molecular integrators for neuromodulator and neuropeptide release.
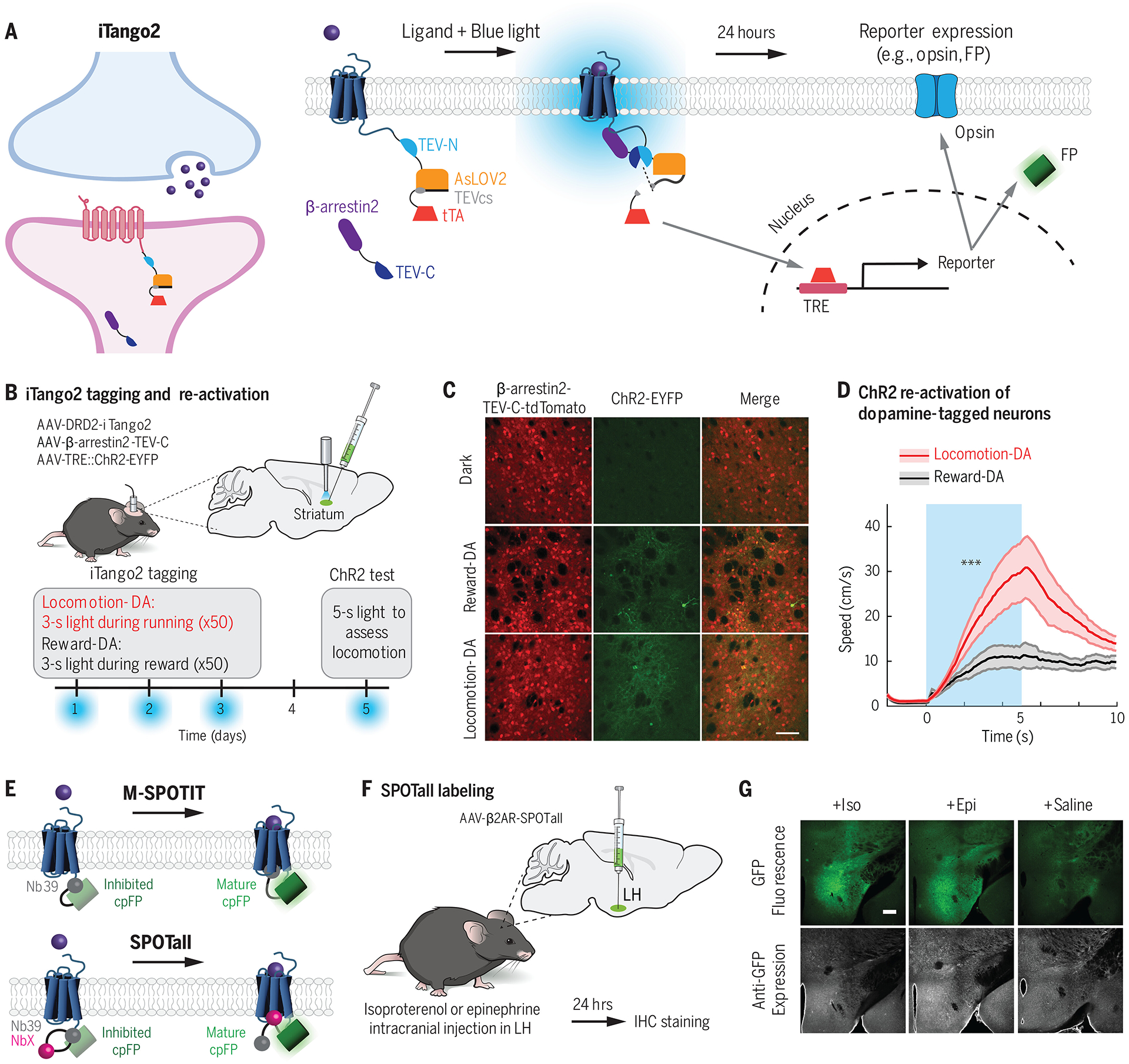
(A) iTango2 (95) is a transcriptional reporter system for detecting GPCR activation. In response to blue light and ligand-GPCR binding, a protease-mediated cleavage event occurs to release a membrane-bound transcription factor tTA. The tTA enters the nucleus where it drives expression of a TRE::reporter gene, such as an opsin or an FP. (B) DRD2-iTango2, β-arrestin2-TEV-C-TdTomato, and TRE::ChR2-EYFP viruses were injected bilaterally into the central striatum, and a fiberoptic cannula was implanted for blue light delivery. Water restricted mice were trained to run to receive a water reward. Blue light was delivered either during reward or locomotion across 3 days to tag neurons undergoing dopamine (DA) modulation. On a test day, blue light was delivered to activate ChR2-cells. ChR2, channelrhodopsin-2; EYFP, enhanced yellow fluorescent protein. (C,D) TRE::ChR2-EYFP expression was observed in mice tagged during either reward or locomotion. ChR2 stimulation of locomotion-DA neurons drove significantly more locomotion behavior compared with ChR2 stimulation of reward-DA neurons. Data shwon from (95). (E) M-SPOTIT and SPOTall (101) are integrators that rely on ligand-GPCR activation to drive the maturation of a fluorescent protein. cpGFP is fused to Nb39, a nanobody that binds the active μ-opioid receptor, and also happens to inhibit the maturation of cpGFP in this conformation. In M-SPOTIT, cpGFP-Nb39 is fused to the C terminus of the μ-opioid receptor. GPCR activation results in Nb39 binding, which no longer inhibits the cpGFP and thus allows it to mature and fluoresce. In SPOTall, different GPCR conformation-specific nanobodies (NbX) can be inserted between cpGFP and Nb39. Nb39 continues to block cpGFP maturation at rest, but upon ligand activation, NbX binds the GPCR and disrupts the Nb39-cpGFP interaction, resulting in enhanced GFP fluorescence. (F,G) SPOTall design for β2AR was tested in the lateral hypothalamus (LH) of mice. Elevated GFP fluorescence was observed in response to epinephrine (Epi) or isoproterenol (Iso) (which bind β2AR), but not in response to saline. Data shown from (101).
One of the first iTango2 applications in vivo was designed for the dopamine type 2 receptor (DRD2-iTango2; Fig. 3B). Lee et al. (95) tagged cells in the central striatum during either reward consumption or locomotion and expressed a channelrhodopsin for future reactivation. Stimulation of locomotion but not reward cells reliably increased the speed of movement, suggesting functionally-specific tagging (Fig. 3C,D). Since that study, an oxytocin receptor iTango2 system has also been described (OXTR-iTango2), which was used to tag neurons in the paraventricular nucleus of the hypothalamus that are activated during social interaction (97), as well as neurons in the ventral tegmental area to quantify elevated oxytocin signaling during pup calls leading to pup retrieval from mothers (98).
Unlike sensors which can only provide a readout of moment-to-moment neurochemical signaling, transcriptional integrators enable genetic access to cells targeted by neuromodulators, thus allowing tagging for post-mortem ex vivo studies such as RNA sequencing, proteomics, and electrophysiology. These tools provide the potential to isolate functional populations of cells and integrate behavioral studies with molecular and synaptic plasticity metrics. However, they are limited by slow integration times, requiring ~1 hour of light and neuromodulation to activate the construct (95). Furthermore, translation of tetracycline-responsive element (TRE)–dependent genes requires ~1 day after activation (95), meaning that the acute molecular signatures associated with specific behavioral states are unlikely to be present. Moreover, because they are blue light–dependent, they are restricted to one or a few brain regions.
Fluorescent protein integrators.
As an alternative to transcriptional reporters, recently Kroning et al. developed an indicator called M-SPOTIT (single-chain protein-based opioid transmission indicator tool), which measures activation of the μ-opioid receptor (μOR) (99, 100). M-SPOTIT fuses the C terminus of μOR to cpGFP and Nb39, a Gi-mimic nanobody that also interacts with cpGFP to inhibit its fluorophore maturation (Fig. 3E). When μOR is activated by a ligand, Nb39 binds to the GPCR instead, thus allowing cpGFP to mature and fluoresce. The group then extended this design to additional GPCRs, creating the SPOTall design, initially validated for β2-adrenergic receptor (β2AR) (101). Because Nb39 is specific to the μOR binding pocket, they added a conformation-specific nanobody for β2AR (Nb80) between the cpGFP and Nb39 (Fig. 3E). Nb80 binds only to the activated β2AR, thus releasing Nb39 from the cpGFP and allowing it to mature. This design can be potentially applied to any GPCR with the appropriate nanobody/receptor pair, and the authors have validated SPOTall designs for additional receptors in vitro. Both an improved M-SPOTIT2 and β2AR-SPOTall have been successfully tested in vivo (Fig. 3F,G).
These tools have a large dynamic range of fluorescence and require only ⁓30 s of receptor agonism to drive fluorophore activation. However, the incubation period required for cpGFP maturation is ~8 hours, resulting in a delay before signal readout. The SPOTall design also has the potential to initiate endogenous GPCR signaling cascades. Furthermore, because there is no temporal gating with these tools (compared with the blue light-gating used in iTango2), these probes are limited in terms of their dynamic range, given the potential for background labeling.
In general, although GPCR integrators are not as established tools as real-time GPCR sensors, the in vivo success of activity-based integrators of immediate early genes (102–104) or intracellular calcium (105–108) suggests that these tools are likely to become widely used in the future. Further improvements are still needed to increase the dynamic range of these tools and reduce background labeling in response to basal or tonically released neurochemical levels.
Manipulating neurochemical signaling during behavior
The aforementioned tools have given us insight into the behaviorally-relevant dynamics of neuromodulator release in vivo; however, these sensors do not allow for the causal investigation of specific neuromodulators during behavior—which would require the ability to manipulate either the ligand or the corresponding GPCR in a circuit-specific manner. Administration of synthesized agonists into the brain through a systemic cranial injection has been widely used to study the behavioral effects of neurochemical signaling in vivo (109). However, this approach lacks spatial and cell-type specificity, and implies the use of non-physiological local concentrations of the agonist. As an alternative, optogenetic stimulation of the subset of neurons that produce a given neuromodulator or neuropeptide can be used to drive neurochemical release (110), but this approach is confounded by the fact that most neurons co-release multiple neuromodulators and neuropeptides (111). Thus, there is a need for new synthetic drug- and light-gated probes and receptors that enable the timed released or modulation of a specific neurochemical or associated downstream receptor in behaving animals.
Engineered GPCR actuators
There are two main types of engineered GPCR actuators that have been widely adopted for use in neuronal circuit studies in vivo: (i) synthetic ligand-activated GPCRs, and (ii) light-activated chimeric GPCRs (Fig. 4A).
Fig. 4. Examining behavioral effects using drug- and light-activated chimeric GPCRs.
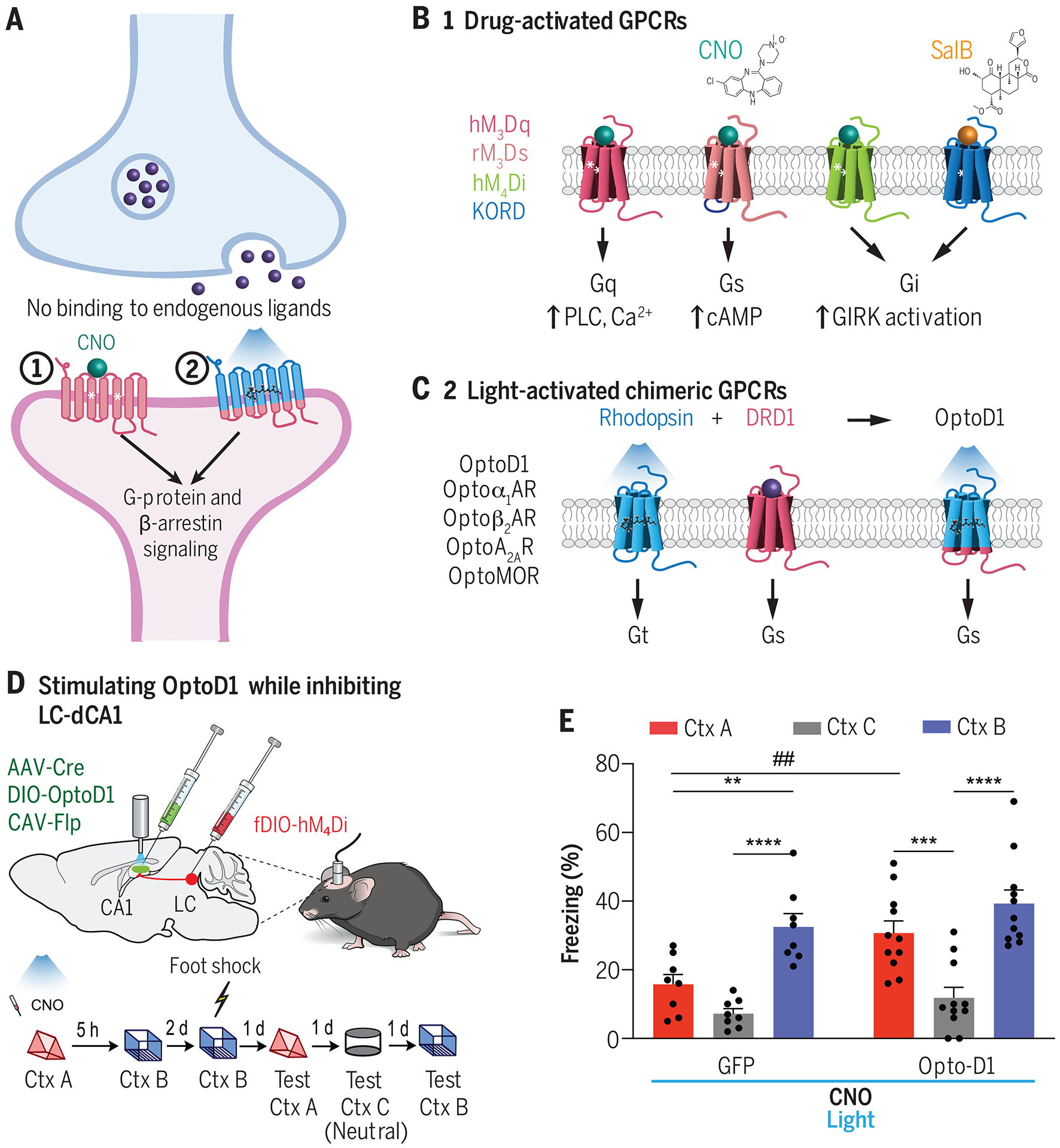
(A) Overview of two classes of engineered GPCR actuators for inducing neuromodulator or neuropeptide receptor signaling in mammalian brain tissue. (B) General designs of commonly used designer receptors exclusively activated by designer drugs (DREADDs) (6, 113). DREADDs activate a specific downstream signaling cascade dependent on the engineered receptor. (C) Design strategy for creating light activated GPCRs (OptoXRs) (120). OptoXRs are engineered chimeras between a rhodopsin (which is activated by blue light) and the intracellular loops of the desired endogenously present GPCR. Upon blue light irradiation, the OptoXR undergoes conformational activation, triggering the native downstream intracellular signaling cascades specific to the desired GPCR. Depicted is OptoD1, which mimics DRD1 signaling in response to blue light (44). (D, E) Chowdhury et al. (129) simultaneously used an inhibitory DREADD and OptoXR to demonstrate the role of a dopaminergic locus coeruleus-to-dorsal hippocampal (LC-dCA1) circuit in memory linking. Whereas hM4Di inhibition of LC-dCA1 neurons disrupts fear memory linking across two contexts, simultaneous OptoD1 stimulation can rescue the memory linking deficits. CNO, clozapine-N-oxide; Ctx A, context A; Ctx B, context B, Ctx C, context C (neutral). Data shown from (129).
Synthetic ligand-activated GPCRs.
Chemogenetic tools such as designer receptors exclusively activated by designer drugs (DREADDs) have been used as a noninvasive approach to modulate neuronal communication in vivo for well over a decade (5, 6, 112). These engineered GPCRs respond only to a synthetic ligand, clozapine-N-oxide (CNO). DREADDs were developed by performing directed evolution and rational engineering of various muscarinic acetylcholine receptor subtypes, which recruit different G-proteins to alter excitability in either direction (5, 113). Subsequently, a few additional DREADDs have been developed by using different GPCRs for in vivo use, including KORD (which is based on the k-opioid receptor that recruits Gi G proteins in response to salvinorin B) (114) and rM3Ds (which is based on a chimeric muscarinic/β-adrenergic receptor that recruits Gs G proteins in response to CNO; Fig. 4B) (115). An improved synthetic ligand, C21 (116), has also been reported to have fewer off-target effects and increased specificity for DREADDs.
Although DREADDs have been widely used in systems neuroscience as a general tool to either increase or decrease excitability within neuronal circuits (6, 117), thus far there is lack of availability for a broader variety of DREADDs across many different GPCRs—as there are with the fluorescent GPCR sensors—in part owing to the immense challenges in evolving GPCRs to gain specificity for an otherwise inert ligand.
Light-activated chimeric GPCRs.
As an alternative to the DREADDs approach, light-activated rhodopsin/GPCR chimeras (or OptoXRs) enable shorter time scales of GPCR activation, and are designed to mimic the signaling mechanisms of several different types of endogenous receptors (118). OptoXRs are engineered by creating a chimera of bovine rhodopsin with the intracellular loops and C-terminal tail of an endogenous GPCR to allow for blue light-dependent modulation of receptor signaling (Fig. 4C) (119). Some of the first OptoXRs have been engineered for in vivo studies on α1- and β2-adrenergic receptors (Optoα1AR and Optoβ2AR), which couple Gq and Gs G proteins, respectively (120). These OptoXRs maintained the biochemical signaling specificity of their endogenous counterparts (121) and could drive receptor-specific changes in neuronal firing and animal behavior. For example, Optoα1AR activation in the NAc drove elevated neuronal firing and conditioned place preference, whereas Optoβ2AR activation in the NAc drove a reduction in overall neuronal firing and no change in preference for the stimulation side.
Subsequently, numerous other OptoXRs and similar light-activated chimeric GPCRs have been developed for use in neuronal circuit studies during behavior (118), including for the dopamine type 1 receptor OptoD1 (44), adenosine 2A receptor OptoA2AR (122), and μOR OptoMOR (123). Recent work has also identified new Gi-coupled light-activated GPCRs based on parapinopsin (124), mosquito rhodopsin (125), and a ciliary opsin (126), which exhibit stable photoswitching properties that enable robust synaptic inhibition in vivo. Future studies could incorporate these Gi-coupled GPCRs as light-activated scaffolds to engineer improved OptoXRs.
Biological applications of GPCR actuators to reveal neurochemical signaling.
OptoXRs have been used in several contexts to elucidate how cell-type specific GPCR activation can modulate changes in intracellular signaling, firing, and overall animal behavior. The same OptoXR can recruit distinct downstream signaling pathways when activated in different brain areas, and as a result can also drive different behaviors. For example, Siuda et al. (127) showed that Optoβ2AR activation in CaMKIIα neurons in the basolateral amygdala (BLA), but not Optoβ2AR activation in all hSyn neurons in the central amygdala (CeA), drives sustained anxiogenic behavior in mice. However, Optoβ2AR activation in the BLA did not drive conditioned or real-time place aversion, suggesting that β2AR signaling in BLA-CaMKIIα neurons could be specific to anxiety and not general aversion. This finding is intriguing given that optogenetic stimulation of BLA-CaMKIIα spiking activity robustly drives fear learning (128), and could indicate that mechanisms independent of β2AR signaling in BLA neurons may mediate avoidance behaviors.
Another study by Li et al. (122) developed OptoA2AR to study the effects of adenosine 2A receptor signaling across different brain regions in mice. They found that OptoA2AR activation in the hippocampus recruits phosphorylated-CREB signaling, whereas OptoA2AR activation in the NAc instead recruits phosphorylated-MAPK signaling [CREB, cyclic adenosine monophosphate (cAMP) response element–binding protein; MAPK, mitogen-activated protein kinase]. Associated with these distinct biochemical changes, OptoA2AR stimulation in the hippocampus selectively impaired memory performance in a modified Y-maze test, whereas stimulation in the NAc did not impair memory performance in the same task, but instead enhanced locomotor activity.
Lastly, a recent study by Chowdhury et al. (129) used OptoD1 (44) to study how locus coeruleus-to-dorsal hippocampal (LC-dCA1) dopamine release modulates memory linking. First, they used a chemogenetic hM4Di DREADD in LC-dCA1 neurons, and found that silencing this projection disrupted fear memory linking across two different contexts experienced closely together in time. Because this circuit co-releases norepinephrine and dopamine, the authors asked whether activation of dopamine receptor signaling alone was sufficient to rescue memory linking deficits. They coupled hM4Di LC-dCA1 inhibition with optogenetic dopamine receptor activation by using OptoD1 and found that OptoD1 stimulation restored memory linking across the two contexts despite simultaneous LC-dCA1 inhibition (Fig. 4D,E).
These studies emphasize the spatiotemporal control of receptor signaling enabled by these new tools and highlight their ability to be used concurrently with other chemogenetic actuators to examine neuronal circuit mechanisms underlying behavior. They also represent examples in which broad optogenetic activation of a given neuronal cell type alone would have been insufficient to determine the cellular mechanism underlying an acute change in behavioral state or role in learning and memory.
Photocaging of GPCR ligands and neuropeptides
Although chemogenetics and OptoXRs enable the activation of an engineered GPCR in a given brain region or cell-type, one potential caveat is that these recombinant receptors might not fully mimic the natural signaling of the natively expressed receptors. One relevant question is whether these modified receptors are trafficked to the same subcellular locations as their endogenous counterparts when expressed virally. Furthermore, in some cases, it is not possible to restrict their expression only to the genetically defined cell-types that express the natural receptor. Instead of engineering activatable receptors, others have designed chemically modified receptor neuroeffectors (photoactivatable ligands) to allow for precise control over the release of a specific neuromodulator or neuropeptide (Fig. 5A).
Fig. 5. Photocaged ligands for spatiotemporal control of neurochemical signaling.
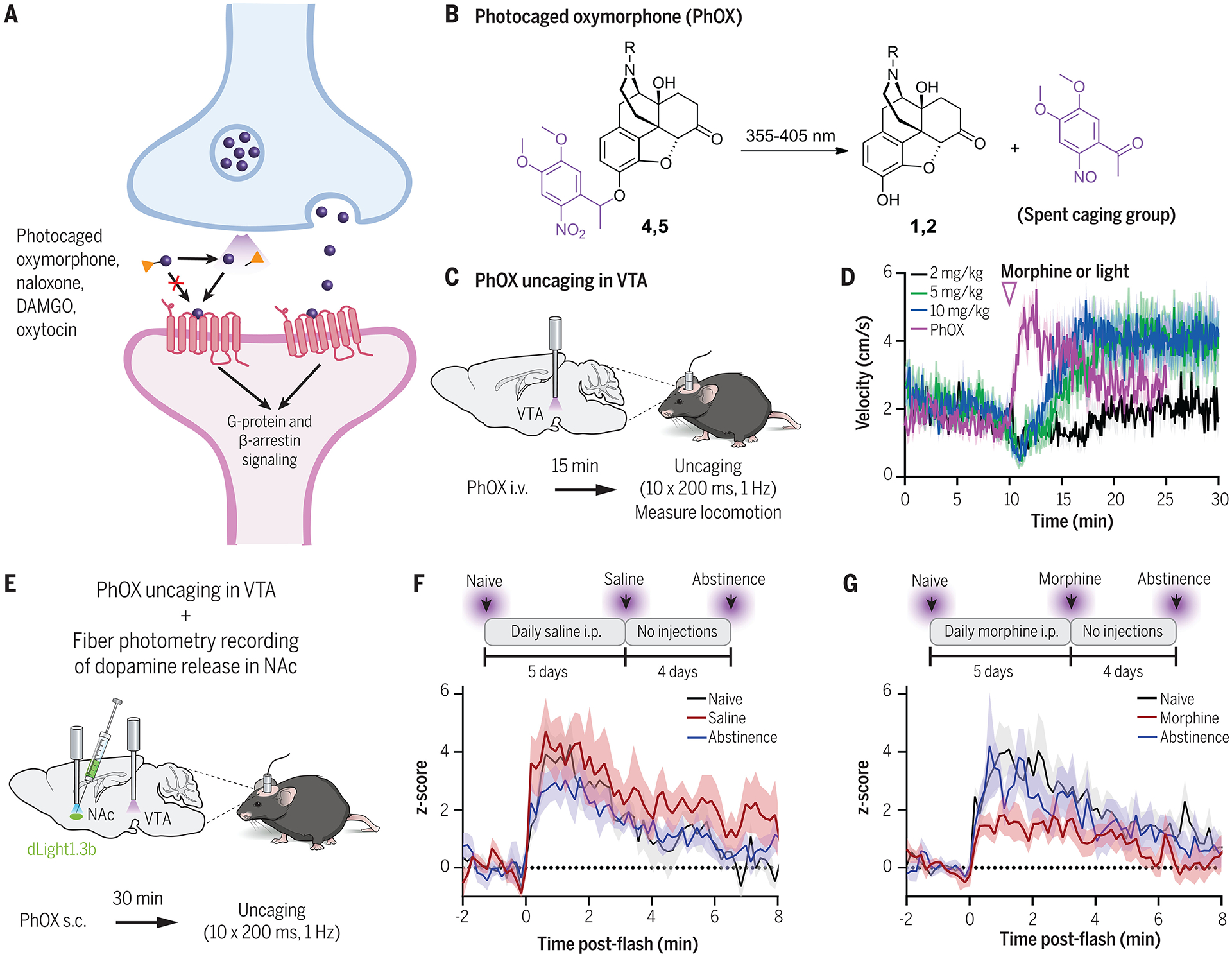
(A) Overview of photocaged ligands and peptides for acute neurochemical uncaging. In the absence of light, photoactivatable ligands are unable to bind to their receptors because of a photocleavable caging group. Light exposure cleaves off the caging group and allows the molecule to bind to endogenous receptors to trigger downstream signaling. (B) Chemical structure of a photocaged oxymorphone [PhOX (143)] before and after UV light activation. (C,D) PhOX uncaging in the VTA activates μ-opioid receptors and drives immediate behavioral changes in locomotion. Data shown from (143). (E-G) Simultaneous application of PhOX and a dopamine sensor, dLight1.3b (61), revealed how repeated morphine injections modulate opioid-induced VTA-to-NAc dopamine signaling in vivo. Dopamine release in the NAc was measured after acute PhOX activation in VTA at the naïve state (black), after 5 days of repeated saline (F) or morphine (G) injections (red), and after 4 days of abstinence from injections (blue). Mice undergoing repeated morphine injections showed a reduction in VTA opioid-NAc dopamine coupling. Data shown from (143).
Ligands can either be designed to be photoswitchable or photocaged. Photoswitchable ligands reversibly become “active” in response to one wavelength of light, and then become “inactive” in response to another wavelength of light (130). Photocaged ligands are inactive in the dark state, and then become irreversibly active upon photolysis and removal of a sterically hindering chemical group (131). Numerous different diffusible and tethered photoswitches have been developed for GPCRs and ion channels for use in neuron culture, invertebrates, larval zebrafish, and mouse (118, 132). Here, we will focus on photocaged ligands that have been adopted for use in mammalian circuits to study neuromodulator and neuropeptide signaling in vivo.
Photocaged endogenous ligands.
Several photocaged neuromodulators and neuropeptides have been described for application in vitro or ex vivo in mammalian brain tissue, including a photocaged dynorphin A(1–8) (133), [Leu5]-enkephalin (133, 134), dopamine (135, 136), orexin-B (137), and others (138). One of the first endogenous neuropeptides to be caged and applied in vivo is a caged oxytocin (139). Endogenously released oxytocin plays a vital role in maternal physiology and social behavior (140). However, it has been challenging to isolate its action within specific neuronal circuits, given the complex mechanisms by which it travels throughout the brain (141). Thus, to allow for spatiotemporal control of oxytocin signaling in vivo, Ahmed et al. (139) developed a photocaged oxytocin neuropeptide (cOT) by substituting in a modified unnatural amino acid at various key residues of the molecule. The final cOT1 design replaced a tyrosine residue with an ortho-nitrobenzyl-tyrosine, which undergoes photolysis within milliseconds upon treatment with ~405-nm ultraviolet (UV) light. Whereas the caged cOT1 is effectively inactive at the oxytocin receptor, the uncaged cOT1 has a similar affinity for the oxytocin receptor as the endogenous neuropeptide. To demonstrate its utility, the scientists injected cOT1 into the auditory cortex of pup-naïve female mice and delivered 0.5-s of uncaging LED light through an opto-fluidic implant. Mice receiving cOT1 uncaging exhibited accelerated pup retrieval compared with control mice receiving no LED light (139).
Photocaged drugs.
In addition to caging endogenous neuropeptides, others have also photocaged synthetic GPCR agonists and antagonists, such as the synthetic opiates DAMGO (142) and oxymorphone (143), and the opioid receptor antagonist naloxone (143, 144). Opioids are a class of clinically relevant analgesic drugs that bind opioid receptors in the brain. Although critical for pain management, they also have high potential for abuse and addiction, among other adverse effects. However, it can be challenging to study the mechanisms underlying their action because of the fact that these drugs diffuse throughout the brain on slow timescales. McClain et al. (143) developed a photocaged oxymorphone (PhOX; Fig. 5B) and naloxone (PhNX) to enable precise control over opioid release and binding across various brain regions in vivo. To make these drugs UV light-activatable, a dimethoxynitrophenethyl (DMNPE) caging group was used, which preserved the blood brain barrier penetrance of the caged drugs. The required duration of UV light in vivo is extremely brief (a single 200-ms flash) and allows for compatibility with other commonly used red and green fluorescent in vivo imaging probes.
To demonstrate the temporal specificity that could be achieved with PhOX, the authors compared locomotion changes induced by various concentrations of systemic oxymorphone injections, versus PhOX photoactivation in the ventral tegmental area (VTA). Mice displayed an increase in locomotion within <5 s of the first UV light flash with PhOX, whereas systemically administered morphine slowly induced locomotion changes over several minutes (Fig. 5C,D). PhOX-driven locomotion changes were abolished in mice treated with systemically administered naloxone, or treated with PhOX and PhNX together. This study also showed that PhOX and PhNX are compatible with real-time GPCR sensors for simultaneous measurements of neuronal circuit function in vivo. A critical question when studying the effects of opioids for clinical use is how repeated opioid use produces tolerance, thus requiring elevated dosage for pain treatment and enhanced potential for abuse. To ask whether prolonged MOR activation in the VTA alters mesolimbic dopamine signaling, the authors recorded dopamine release in the medial shell NAc by using dLight1.3b (61) while also acutely photoactivating PhOX in the VTA after multiple days of repeated morphine administration (Fig. 5E). They demonstrated that NAc dopamine signaling in response to a VTA-PhOX activation flash was diminished in mice that had undergone repeated morphine treatment compared with that observed in saline controls (Fig. 5F,G). These findings showed that opioid sensitivity within the mesolimbic dopamine circuitry is reduced by prolonged μOR activation.
The precise temporal resolution afforded by photocaged drugs/ligands provides sensitive readouts of neurochemical action in the brain, which may be important for future mechanistic studies and hypotheses. Furthermore, the compatibility of these photoactivatable ligands with complementary methodologies for recording neuronal signaling makes them particularly attractive for application in neuronal circuit studies. However, it should be noted that proper experimental design and data interpretation is critical because caged ligands still rely on dilution through diffusion and are only fully deactivated by cellular metabolism.
Conclusion
By bridging interdisciplinary approaches spanning chemistry, biology, and neuroscience, researchers have created a suite of technologies for exploring how neuromodulation and neuropeptide signaling in the brain modulates neuronal circuitry and animal behavior. These tools have started to reveal the exquisite roles that these molecules and their downstream receptors play in regulating both innate and learned behaviors. Looking to the future, multiplexed recording or manipulation of multiple molecules at once may reveal new principles of how neuromodulators and neuropeptides alter circuit plasticity in specific cell-types and at distinct subcellular locations. It will be critical to continue engineering sensors and actuators with excitation and activation spectra that avoid overlap with existing sensors for calcium, voltage, or neurotransmitters—for example, by using cp nIR fluorescent proteins (145) or new HaloTag-based sensors with nIR fluorescent HaloTag ligands (146). An additional limitation of existing sensors and actuators based on endogenous receptors is that they cannot distinguish between the promiscuous binding of multiple neuromodulators and neuropeptides to the same downstream receptor type. Machine learning- and artificial intelligence-based engineering (57, 147) and de novo protein design (148) may enable the creation of completely orthogonal tools that are tuned to only a single analyte and that can still maintain genetic encodability and cell type specificity. In this review, we focused on the application of these toolboxes in the central nervous system; however, it is important to note that neuromodulators and neuropeptides are released throughout the body as well, where they can have a profound effect on other systems, including the gut microbiome and the immune system. Understanding the interplay of brain-body neurochemical signaling will likely require novel tools that can function in the periphery as well as in the brain.
Acknowledgments:
We thank I. A. Ahmed, W. Wang, M. R. Banghart, M. R. Bruchas, and L. Tian for helpful discussions.
Funding:
This work was supported by the following sources:
Canadian Institutes of Health Research (Post-doctoral training award 202210MFE-491520-297096, J.M.)
National Science Foundation (Graduate Research Fellowship Program #000895154, M.A.)
Burroughs Wellcome Fund (Career Award at the Scientific Interface #1019469, C.K.K.)
Brain & Behavior Research Foundation (Young Investigator Grant #30238, C.K.K.)
Searle Scholars Program (SSP-2022-107, C.K.K.)
Arnold and Mabel Beckman Foundation (Beckman Young Investigator Award, C.K.K.)
National Institutes of Health (DP2MH136588 and R21DA059842, C.K.K., and T32MH082174 to M.A.)
Footnotes
Competing interests: Authors declare that they have no competing interests.
References.
- 1.Shen Y, Luchetti A, Fernandes G, Do Heo W, Silva AJ, The emergence of molecular systems neuroscience. Mol Brain 15, 7 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zhang Y et al. , Fast and sensitive GCaMP calcium indicators for imaging neural populations. Nature 615, 884–891 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Xu Y, Zou P, Cohen AE, Voltage imaging with genetically encoded indicators. Current Opinion in Chemical Biology 39, 1–10 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Deisseroth K, Optogenetics: 10 years of microbial opsins in neuroscience. Nature neuroscience 18, 1213 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Armbruster BN, Li X, Pausch MH, Herlitze S, Roth BL, Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc Natl Acad Sci U S A 104, 5163–5168 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Roth BL, DREADDs for neuroscientists. Neuron 89, 683–694 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Atasoy D, Sternson SM, Chemogenetic tools for causal cellular and neuronal biology. Physiological reviews 98, 391–418 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Fenno LE et al. , Comprehensive Dual- and Triple-Feature Intersectional Single-Vector Delivery of Diverse Functional Payloads to Cells of Behaving Mammals. Neuron 107, 836–853 e811 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rosenbaum DM, Rasmussen SG, Kobilka BK, The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Araneda R, Andrade R, 5-Hydroxytryptamine2 and 5-hydroxytryptamine1A receptors mediate opposing responses on membrane excitability in rat association cortex. Neuroscience 40, 399–412 (1991). [DOI] [PubMed] [Google Scholar]
- 11.Hernandez-Lopez S et al. , D2 dopamine receptors in striatal medium spiny neurons reduce L-type Ca2+ currents and excitability via a novel PLCβ1–IP3–calcineurin-signaling cascade. Journal of Neuroscience 20, 8987–8995 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Giessel AJ, Sabatini BL, M1 muscarinic receptors boost synaptic potentials and calcium influx in dendritic spines by inhibiting postsynaptic SK channels. Neuron 68, 936–947 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Maejima T, Masseck OA, Mark MD, Herlitze S, Modulation of firing and synaptic transmission of serotonergic neurons by intrinsic G protein-coupled receptors and ion channels. Frontiers in integrative neuroscience 7, 40 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Collins AL, Aitken TJ, Greenfield VY, Ostlund SB, Wassum KM, Nucleus accumbens acetylcholine receptors modulate dopamine and motivation. Neuropsychopharmacology 41, 2830–2838 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Carhart-Harris RL, Nutt D, Serotonin and brain function: a tale of two receptors. Journal of psychopharmacology 31, 1091–1120 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Eban-Rothschild A, Appelbaum L, de Lecea L, Neuronal mechanisms for sleep/wake regulation and modulatory drive. Neuropsychopharmacology 43, 937–952 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Likhtik E, Johansen JP, Neuromodulation in circuits of aversive emotional learning. Nature neuroscience 22, 1586–1597 (2019). [DOI] [PubMed] [Google Scholar]
- 18.Wang Y et al. , NeuroPep: a comprehensive resource of neuropeptides. Database 2015, bav038 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Van Den Pol AN, Neuropeptide transmission in brain circuits. Neuron 76, 98–115 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yao Z et al. , A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Clark JT, Kalra PS, Crowley WR, Kalra SP, Neuropeptide Y and human pancreatic polypeptide stimulate feeding behavior in rats. Endocrinology 115, 427–429 (1984). [DOI] [PubMed] [Google Scholar]
- 22.Semjonous NM et al. , Coordinated changes in energy intake and expenditure following hypothalamic administration of neuropeptides involved in energy balance. Int J Obes (Lond) 33, 775–785 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Horio N, Liberles SD, Hunger enhances food-odour attraction through a neuropeptide Y spotlight. Nature 592, 262-+ (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fitzsimons JT, Angiotensin, thirst, and sodium appetite. Physiol Rev 78, 583–686 (1998). [DOI] [PubMed] [Google Scholar]
- 25.Kinsman BJ et al. , Integration of Hypernatremia and Angiotensin II by the Organum Vasculosum of the Lamina Terminalis Regulates Thirst. J Neurosci 40, 2069–2079 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rilling JK, Young LJ, The biology of mammalian parenting and its effect on offspring social development. Science 345, 771–776 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Marlin BJ, Mitre M, D’Amour J A, Chao MV, Froemke RC, Oxytocin enables maternal behaviour by balancing cortical inhibition. Nature 520, 499–504 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Robinson DL, Venton BJ, Heien ML, Wightman RM, Detecting subsecond dopamine release with fast-scan cyclic voltammetry in vivo. Clinical chemistry 49, 1763–1773 (2003). [DOI] [PubMed] [Google Scholar]
- 29.Mohebi A et al. , Dissociable dopamine dynamics for learning and motivation. Nature 570, 65–70 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Phillips PE, Robinson DL, Stuber GD, Carelli RM, Wightman RM, Real-time measurements of phasic changes in extracellular dopamine concentration in freely moving rats by fast-scan cyclic voltammetry. Drugs of Abuse: Neurological Reviews and Protocols, 443–464 (2003). [DOI] [PubMed] [Google Scholar]
- 31.Nadim F, Bucher D, Neuromodulation of neurons and synapses. Current opinion in neurobiology 29, 48–56 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Xin J, Fan T, Guo P, Wang J, Identification of functional divergence sites in dopamine receptors of vertebrates. Computational biology and chemistry 83, 107140 (2019). [DOI] [PubMed] [Google Scholar]
- 33.Calabresi P, Picconi B, Tozzi A, Ghiglieri V, Di Filippo M, Direct and indirect pathways of basal ganglia: a critical reappraisal. Nature neuroscience 17, 1022–1030 (2014). [DOI] [PubMed] [Google Scholar]
- 34.Gerfen CR, Surmeier DJ, Modulation of striatal projection systems by dopamine. Annual review of neuroscience 34, 441–466 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sulzer D et al. , Dopamine neurons make glutamatergic synapses in vitro. Journal of Neuroscience 18, 4588–4602 (1998). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hnasko TS et al. , Vesicular glutamate transport promotes dopamine storage and glutamate corelease in vivo. Neuron 65, 643–656 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Smith CC, Greene RW, CNS dopamine transmission mediated by noradrenergic innervation. Journal of Neuroscience 32, 6072–6080 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kempadoo KA, Mosharov EV, Choi SJ, Sulzer D, Kandel ER, Dopamine release from the locus coeruleus to the dorsal hippocampus promotes spatial learning and memory. Proceedings of the National Academy of Sciences 113, 14835–14840 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Fuxe K et al. , The discovery of central monoamine neurons gave volume transmission to the wired brain. Progress in neurobiology 90, 82–100 (2010). [DOI] [PubMed] [Google Scholar]
- 40.Park J, Takmakov P, Wightman RM, In vivo comparison of norepinephrine and dopamine release in rat brain by simultaneous measurements with fast‐scan cyclic voltammetry. Journal of neurochemistry 119, 932–944 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Nguyen Q-T et al. , An in vivo biosensor for neurotransmitter release and in situ receptor activity. Nature neuroscience 13, 127–132 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Muller A, Joseph V, Slesinger PA, Kleinfeld D, Cell-based reporters reveal in vivo dynamics of dopamine and norepinephrine release in murine cortex. Nature methods 11, 1245–1252 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hughes RN et al. , Ventral tegmental dopamine neurons control the impulse vector during motivated behavior. Current Biology 30, 2681–2694. e2685 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Gunaydin LA et al. , Natural neural projection dynamics underlying social behavior. Cell 157, 1535–1551 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Chen Y, Lin Y-C, Kuo T-W, Knight ZA, Sensory detection of food rapidly modulates arcuate feeding circuits. Cell 160, 829–841 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Kim CK et al. , Simultaneous fast measurement of circuit dynamics at multiple sites across the mammalian brain. Nature methods 13, 325–328 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Howe MW, Dombeck DA, Rapid signalling in distinct dopaminergic axons during locomotion and reward. Nature 535, 505–510 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Engelhard B et al. , Specialized coding of sensory, motor and cognitive variables in VTA dopamine neurons. Nature 570, 509–513 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Marvin JS et al. , An optimized fluorescent probe for visualizing glutamate neurotransmission. Nat Methods 10, 162–170 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Helassa N et al. , Ultrafast glutamate sensors resolve high-frequency release at Schaffer collateral synapses. Proc Natl Acad Sci U S A 115, 5594–5599 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Marvin JS et al. , Stability, affinity, and chromatic variants of the glutamate sensor iGluSnFR. Nat Methods 15, 936–939 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Aggarwal A et al. , Glutamate indicators with improved activation kinetics and localization for imaging synaptic transmission. Nat Methods 20, 925–934 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Borden PM et al. , A fast genetically encoded fluorescent sensor for faithful in vivo acetylcholine detection in mice, fish, worms and flies. bioRxiv 2020.02.07.939504 [Preprint] (2020). 10.1101/2020.02.07.939504. [DOI] [Google Scholar]
- 54.McGirr A, LeDue J, Chan AW, Xie Y, Murphy TH, Cortical functional hyperconnectivity in a mouse model of depression and selective network effects of ketamine. Brain 140, 2210–2225 (2017). [DOI] [PubMed] [Google Scholar]
- 55.Dong C et al. , Fluorescence Imaging of Neural Activity, Neurochemical Dynamics, and Drug-Specific Receptor Conformation with Genetically Encoded Sensors. Annu Rev Neurosci 45, 273–294 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Marvin JS et al. , A genetically encoded fluorescent sensor for in vivo imaging of GABA. Nat Methods 16, 763–770 (2019). [DOI] [PubMed] [Google Scholar]
- 57.Unger EK et al. , Directed Evolution of a Selective and Sensitive Serotonin Sensor via Machine Learning. Cell 183, 1986–2002 e1926 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hon OJ et al. , Serotonin modulates an inhibitory input to the central amygdala from the ventral periaqueductal gray. Neuropsychopharmacology 47, 2194–2204 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Ramkumar R et al. , Long-term impact of early life stress on serotonin connectivity. Biological Psychiatry, (2024). [DOI] [PubMed] [Google Scholar]
- 60.Jang S et al. , Impact of the circadian nuclear receptor REV-ERBα in dorsal raphe 5-HT neurons on social interaction behavior, especially social preference. Experimental & Molecular Medicine 55, 1806–1819 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Patriarchi T et al. , Ultrafast neuronal imaging of dopamine dynamics with designed genetically encoded sensors. Science 360, (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Kutlu MG et al. , Dopamine release in the nucleus accumbens core signals perceived saliency. Curr Biol 31, 4748–4761 e4748 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Jørgensen SH et al. , Behavioral encoding across timescales by region-specific dopamine dynamics. Proceedings of the National Academy of Sciences 120, e2215230120 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Kagiampaki Z et al. , Sensitive multicolor indicators for monitoring norepinephrine in vivo. Nat Methods 20, 1426–1436 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Kubitschke M et al. , Next generation genetically encoded fluorescent sensors for serotonin. Nature Communications 13, 7525 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Dong C et al. , Psychedelic-inspired drug discovery using an engineered biosensor. Cell 184, 2779–2792 e2718 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Tian L et al. , Unlocking opioid neuropeptide dynamics with genetically-encoded biosensors. Research Square, doi: 10.21203/rs.21203.rs-2871083/v2871081 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Melzer S et al. , Bombesin-like peptide recruits disinhibitory cortical circuits and enhances fear memories. Cell 184, 5622–5634. e5625 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Duffet L et al. , A genetically encoded sensor for in vivo imaging of orexin neuropeptides. Nat Methods 19, 231–241 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Sun F et al. , A Genetically Encoded Fluorescent Sensor Enables Rapid and Specific Detection of Dopamine in Flies, Fish, and Mice. Cell 174, 481–496 e419 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Elias LJ et al. , Touch neurons underlying dopaminergic pleasurable touch and sexual receptivity. Cell 186, 577–590. e516 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Sun F et al. , Next-generation GRAB sensors for monitoring dopaminergic activity in vivo. Nat Methods 17, 1156–1166 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Zhuo Y et al. , Improved green and red GRAB sensors for monitoring dopaminergic activity in vivo. Nat Methods 21, 680–691 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Wan J et al. , A genetically encoded sensor for measuring serotonin dynamics. Nat Neurosci 24, 746–752 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Deng F et al. , Improved green and red GRAB sensors for monitoring spatiotemporal serotonin release in vivo. Nature Methods 21, 692–702 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Jing M et al. , A genetically encoded fluorescent acetylcholine indicator for in vitro and in vivo studies. Nat Biotechnol 36, 726–737 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Jing M et al. , An optimized acetylcholine sensor for monitoring in vivo cholinergic activity. Nat Methods 17, 1139–1146 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Feng J et al. , A Genetically Encoded Fluorescent Sensor for Rapid and Specific In Vivo Detection of Norepinephrine. Neuron 102, 745–761 e748 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Feng J et al. , Monitoring norepinephrine release in vivo using next-generation GRABNE sensors. Neuron, Mar 26:S0896–6273(0824)00155–00157 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Wang H et al. , A tool kit of highly selective and sensitive genetically encoded neuropeptide sensors. Science 382, eabq8173 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Qian T et al. , A genetically encoded sensor measures temporal oxytocin release from different neuronal compartments. Nature Biotechnology 41, 944–957 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Patriarchi T et al. , An expanded palette of dopamine sensors for multiplex imaging in vivo. Nat Methods 17, 1147–1155 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Taniguchi J et al. , Comment on ‘Accumbens cholinergic interneurons dynamically promote dopamine release and enable motivation’. bioRxiv, doi: 10.1101/2023.1112.1127.573485 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Hong G et al. , Through-skull fluorescence imaging of the brain in a new near-infrared window. Nat Photonics 8, 723–730 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Beyene AG et al. , Imaging striatal dopamine release using a nongenetically encoded near infrared fluorescent catecholamine nanosensor. Science Advances 5, (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Jeong S et al. , High-throughput evolution of near-infrared serotonin nanosensors. Sci Adv 5, eaay3771 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Xia J et al. , Imaging in vivo acetylcholine release in the peripheral nervous system with a fluorescent nanosensor. Proc Natl Acad Sci U S A 118, (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Sayar-Atasoy N et al. , Opioidergic signaling contributes to food-mediated suppression of AgRP neurons. Cell Rep 43, 113630 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Mizushige T et al. , Preference for dietary fat induced by release of beta-endorphin in rats. Life sciences 84, 760–765 (2009). [DOI] [PubMed] [Google Scholar]
- 90.Stanley AT, Post MR, Lacefield C, Sulzer D, Miniaci MC, Norepinephrine release in the cerebellum contributes to aversive learning. Nature Communications 14, 4852 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Carlson ES et al. , Catecholaminergic innervation of the lateral nucleus of the cerebellum modulates cognitive behaviors. Journal of Neuroscience 41, 3512–3530 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Abraham AD et al. , Release of endogenous dynorphin opioids in the prefrontal cortex disrupts cognition. Neuropsychopharmacology 46, 2330–2339 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Kim MW et al. , Time-gated detection of protein-protein interactions with transcriptional readout. Elife 6, (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Kim CK, Cho KF, Kim MW, Ting AY, Luciferase-LOV BRET enables versatile and specific transcriptional readout of cellular protein-protein interactions. eLife 8, e43826 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Lee D et al. , Temporally precise labeling and control of neuromodulatory circuits in the mammalian brain. Nat Methods 14, 495–503 (2017). [DOI] [PubMed] [Google Scholar]
- 96.Barnea G et al. , The genetic design of signaling cascades to record receptor activation. Proceedings of the National Academy of Sciences 105, 64–69 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Mignocchi N, Krüssel S, Jung K, Lee D, Kwon H-B, Development of a genetically-encoded oxytocin sensor. bioRxiv, 2020.07.14.202598 [Preprint] (2020). 10.1101/2020.07.14.202598. [DOI] [Google Scholar]
- 98.Valtcheva S et al. , Neural circuitry for maternal oxytocin release induced by infant cries. Nature 621, 788–795 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Kroning KE, Wang W, Designing a Single Protein-Chain Reporter for Opioid Detection at Cellular Resolution. Angew Chem Int Ed Engl 60, 13358–13365 (2021). [DOI] [PubMed] [Google Scholar]
- 100.Kroning KE, Li M, Petrescu DI, Wang W, A genetically encoded sensor with improved fluorescence intensity for opioid detection at cellular resolution. Chem Commun (Camb) 57, 10560–10563 (2021). [DOI] [PubMed] [Google Scholar]
- 101.Kroning K et al. , Single-chain fluorescent integrators for mapping G-protein-coupled receptor agonists. Proceedings of the National Academy of Sciences 121, e2307090121 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Liu X et al. , Optogenetic stimulation of a hippocampal engram activates fear memory recall. Nature 484, 381–385 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Guenthner CJ, Miyamichi K, Yang HH, Heller HC, Luo L, Permanent genetic access to transiently active neurons via TRAP: targeted recombination in active populations. Neuron 78, 773–784 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.DeNardo LA et al. , Temporal evolution of cortical ensembles promoting remote memory retrieval. Nature neuroscience 22, 460–469 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Wang W et al. , A light- and calcium-gated transcription factor for imaging and manipulating activated neurons. Nature biotechnology 35, 864–871 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Lee D, Hyun JH, Jung K, Hannan P, Kwon H-B, A calcium- and light-gated switch to induce gene expression in activated neurons. Nature Biotechnology, (2017). [DOI] [PubMed] [Google Scholar]
- 107.Kim CK et al. , A Molecular calcium integrator reveals a striatal cell type driving aversion. Cell 183, 2003–2019. e2016 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Sanchez MI, Nguyen Q-A, Wang W, Soltesz I, Ting AY, Transcriptional readout of neuronal activity via an engineered Ca2+-activated protease. Proceedings of the National Academy of Sciences 117, 33186–33196 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Gendreau PL, Petitto JM, Petrova A, Gariépy J-L, Lewis MH, D3 and D2 dopamine receptor agonists differentially modulate isolation-induced social-emotional reactivity in mice. Behavioural brain research 114, 107–117 (2000). [DOI] [PubMed] [Google Scholar]
- 110.Chen W et al. , The Roles of Optogenetics and Technology in Neurobiology: A Review. Front Aging Neurosci 14, 867863 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Hnasko TS, Edwards RH, Neurotransmitter corelease: mechanism and physiological role. Annual review of physiology 74, 225–243 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Conklin BR et al. , Engineering GPCR signaling pathways with RASSLs. Nature methods 5, 673–678 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Clark PJ, Brodnik ZD, Espana RA, Chemogenetic Signaling in Space and Time: Considerations for Designing Neuroscience Experiments Using DREADDs. Neuroscientist, 10738584221134587 (2022). [DOI] [PubMed] [Google Scholar]
- 114.Vardy E et al. , A New DREADD Facilitates the Multiplexed Chemogenetic Interrogation of Behavior. Neuron 86, 936–946 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Guettier JM et al. , A chemical-genetic approach to study G protein regulation of beta cell function in vivo. Proc Natl Acad Sci U S A 106, 19197–19202 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Bonaventura J et al. , High-potency ligands for DREADD imaging and activation in rodents and monkeys. Nat Commun 10, 4627 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Smith KS, Bucci DJ, Luikart BW, Mahler SV, DREADDS: Use and application in behavioral neuroscience. Behavioral neuroscience 135, 89 (2021). [DOI] [PubMed] [Google Scholar]
- 118.Marcus DJ, Bruchas MR, Optical Approaches for Investigating Neuromodulation and G Protein-Coupled Receptor Signaling. Pharmacol Rev 75, 1119–1139 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Kim J-M et al. , Light-driven activation of β2-adrenergic receptor signaling by a chimeric rhodopsin containing the β2-adrenergic receptor cytoplasmic loops. Biochemistry 44, 2284–2292 (2005). [DOI] [PubMed] [Google Scholar]
- 120.Airan RD, Thompson KR, Fenno LE, Bernstein H, Deisseroth K, Temporally precise in vivo control of intracellular signalling. Nature 458, 1025–1029 (2009). [DOI] [PubMed] [Google Scholar]
- 121.Siuda ER et al. , Optodynamic simulation of β-adrenergic receptor signalling. Nature communications 6, 8480 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Li P et al. , Optogenetic activation of intracellular adenosine A2A receptor signaling in the hippocampus is sufficient to trigger CREB phosphorylation and impair memory. Mol Psychiatry 20, 1481 (2015). [DOI] [PubMed] [Google Scholar]
- 123.Siuda ER et al. , Spatiotemporal control of opioid signaling and behavior. Neuron 86, 923–935 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Copits BA et al. , A photoswitchable GPCR-based opsin for presynaptic inhibition. Neuron 109, 1791–1809. e1711 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Mahn M et al. , Efficient optogenetic silencing of neurotransmitter release with a mosquito rhodopsin. Neuron 109, 1621–1635. e1628 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Wietek J et al. , A bistable inhibitory OptoGPCR for multiplexed optogenetic control of neural circuits. BioRxiv, (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Siuda ER, Al-Hasani R, McCall JG, Bhatti DL, Bruchas MR, Chemogenetic and Optogenetic Activation of Galphas Signaling in the Basolateral Amygdala Induces Acute and Social Anxiety-Like States. Neuropsychopharmacology 41, 2011–2023 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Johansen JP et al. , Optical activation of lateral amygdala pyramidal cells instructs associative fear learning. Proceedings of the National Academy of Sciences 107, 12692–12697 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Chowdhury A et al. , A locus coeruleus-dorsal CA1 dopaminergic circuit modulates memory linking. Neuron 110, 3374–3388 e3378 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Deal WJ, Erlanger BF, Nachmansohn D, Photoregulation of biological activity by photochromic reagents, III. Photoregulation of bioelectricity by acetylcholine receptor inhibitors. Proceedings of the National Academy of Sciences 64, 1230–1234 (1969). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Klán P et al. , Photoremovable protecting groups in chemistry and biology: reaction mechanisms and efficacy. Chemical reviews 113, 119–191 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Hull K, Morstein J, Trauner D, In vivo photopharmacology. Chemical reviews 118, 10710–10747 (2018). [DOI] [PubMed] [Google Scholar]
- 133.Banghart MR, Sabatini BL, Photoactivatable neuropeptides for spatiotemporally precise delivery of opioids in neural tissue. Neuron 73, 249–259 (2012). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Banghart MR, He XJ, Sabatini BL, A Caged Enkephalin Optimized for Simultaneously Probing Mu and Delta Opioid Receptors. ACS Chem Neurosci 9, 684–690 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Asad N et al. , Photoactivatable Dopamine and Sulpiride to Explore the Function of Dopaminergic Neurons and Circuits. ACS Chem Neurosci 11, 939–951 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Lee TH, Gee KR, Ellinwood EH, Seidler FJ, Combining ‘caged-dopamine’photolysis with fast-scan cyclic voltammetry to assess dopamine clearance and release autoinhibition in vitro. Journal of neuroscience methods 67, 221–231 (1996). [PubMed] [Google Scholar]
- 137.Duffet L et al. , A photocaged orexin-B for spatiotemporally precise control of orexin signaling. Cell Chem Biol 29, 1729–1738 e1728 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138.Layden AE et al. , A biomimetic C-terminal extension strategy for photocaging amidated neuropeptides. Journal of the American Chemical Society 145, 19611–19621 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 139.Ahmed IA et al. , Optopharmacological tools for precise spatiotemporal control of oxytocin signaling in the central nervous system and periphery. bioRxiv, doi: 10.1101/2022.1111.1110.516001 (2022). [DOI] [Google Scholar]
- 140.Marlin BJ, Froemke RC, O xytocin modulation of neural circuits for social behavior. Developmental Neurobiology 77, 169–189 (2017). [DOI] [PubMed] [Google Scholar]
- 141.Son S et al. , Whole-brain wiring diagram of oxytocin system in adult mice. Journal of Neuroscience 42, 5021–5033 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Ma X et al. , In vivo photopharmacology with a caged mu opioid receptor agonist drives rapid changes in behavior. Nat Methods 20, 682–685 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.McClain SP et al. , In vivo photopharmacology with light-activated opioid drugs. Neuron 111, 3926–3940 e3910 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Banghart MR, Williams JT, Shah RC, Lavis LD, Sabatini BL, Caged naloxone reveals opioid signaling deactivation kinetics. Molecular pharmacology 84, 687–695 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 145.Qian Y et al. , A genetically encoded near-infrared fluorescent calcium ion indicator. Nature methods 16, 171–174 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Deo C et al. , The HaloTag as a general scaffold for far-red tunable chemigenetic indicators. Nature chemical biology 17, 718–723 (2021). [DOI] [PubMed] [Google Scholar]
- 147.Wait SJ et al. , Machine learning-guided engineering of genetically encoded fluorescent calcium indicators. Nature Computational Science, 1–13 (2024). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Bennett NR et al. , Improving de novo protein binder design with deep learning. Nature Communications 14, 2625 (2023). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Dong H et al. , Genetically encoded sensors for measuring histamine release both in vitro and in vivo. Neuron 111, 1564–1576 e1566 (2023). [DOI] [PubMed] [Google Scholar]
- 150.Ino D, Tanaka Y, Hibino H, Nishiyama M, A fluorescent sensor for real-time measurement of extracellular oxytocin dynamics in the brain. Nature Methods 19, 1286–1294 (2022). [DOI] [PMC free article] [PubMed] [Google Scholar]


