Abstract
Using the x-ray structure of the glycerol 3-phosphate transporter (GlpT), we devised a model for the distantly related oxalate transporter, OxlT. The model accommodates all earlier biochemical information on OxlT, including the idea that Lys-355 lies on the permeation pathway, and predicts that Lys-355 and a second positive center, Arg-272, comprise the binding site for divalent oxalate. Study of R272K, R272A, and R272Q derivatives verifies that Arg-272 is essential, and comparisons with GlpT show that both anion transporters bind substrates within equivalent domains. In 22 single-cysteine variants in TM7 and TM8, topology as marked by accessibility to Oregon green maleimide is predicted by the model, with similar concordance for 52 positions probed earlier. The model also reconciles cross-linking of a cysteine pair placed near the periplasmic ends of TM2 and TM7, and retrospective study of TM2 and TM11 confirms that positions supporting disulfide trapping lie at a helical interface. Our work describes a pathway to the modeling of OxlT and other transporters in the major facilitator superfamily and outlines simple experimental tests to evaluate such proposals.
Keywords: anion-binding, disulfide trapping, major facilitator superfamily, membrane protein, permeation pathway
OxlT, the oxalate/formate antiporter of Oxalobacter formigenes (1, 2), belongs to the major facilitator superfamily (MFS), a large and diverse collection encompassing 30–40% of known transporters and permeases (www.biology.ucsd.edu/~msaier/transport). The main biochemical mechanisms associated with transporters (uniport, antiport, and symport) may be found within the MFS, whose individual members display a broad catalog of substrate specificity, including simple sugars and amino acids, intermediary metabolites, and even neurotransmitters (3). All members of the MFS share an architectural theme in which a central loop connects two groups of (typically) six transmembrane α-helices. Moreover, the superfamily as a whole is characterized by a short motif (GXXXDK/R) at the cytoplasmic ends of TM2 and TM8 (3, 4), suggesting that these two six-helix clusters derived from a common ancestor; indeed, at times one finds a clear sequence homology between the N- and C-terminal domains (4, 5).
Insight into the structure of MFS transporters is presently limited. Helix organization, symmetry, and connectivity were established first for OxlT, in work based on a low-resolution (6.5 Å) structure obtained by electron crystallography (6, 7). Subsequently, higher resolution (3.2–3.5 Å) was achieved by x-ray crystallography of two other MFS members from Escherichia coli, the H+/lactose symporter (LacY) and the phosphate/glycerol 3-phosphate antiporter (GlpT) (8, 9). These latter achievements have prompted several recent attempts to use LacY or GlpT as structural templates for models of other systems (10–13). With this in mind and to provide a detailed perspective to guide further work, we selected the GlpT structure as a template for derivation of a homology model of OxlT, using the suite of modeling tools available via the Swiss-Model module of the Swiss Pro web site (14–16). In the present case, the derived homology model also served as the starting point for experiments that revealed important information. In particular, our work confirms the presence in OxlT of a bifunctional active site, comprised of two positively charged residues (Arg-272 and Lys-355), positioned 10 Å apart across the permeation pathway. This arrangement bears striking resemblance to that found in GlpT, where the anion-binding site incorporates two arginine residues separated by a comparable distance spanning the pathway (9, 17). Indeed, superposition of the GlpT and OxlT ligand-binding residues shows these active site components all point to the same region, together identifying a discoid roughly 10 Å in diameter and 3 Å in thickness as the main region involved in substrate (anion) binding.
Materials and Methods
Plasmids and Protein Expression. OxlT and its mutants, each with a polyhistidine (His-9) C-terminal extension (18), were housed in Strain XL3 (2) and grown in Luria–Bertani medium as described (18–20). Most OxlT variants were simple derivatives of a cysteineless C28G/C271A protein, but for disulfide trapping, this variant also carried a tandem duplication of the Factor Xa cleavage site (IEGR) in its central loop as well as a separate mutation (R248Q) eliminating an endogenous Factor Xa cleavage site (21). Cameleon or QuikChange mutagenesis (Stratagene) was confirmed by DNA sequencing.
Reconstitution of OxlT. Membrane protein was solubilized and reconstituted (1, 20) to give proteoliposomes loaded with 100 mM potassium oxalate and 50 mM potassium phosphate (pH 7). OxlT function was monitored by accumulation of 100 μM external [14C]oxalate in exchange for internal substrate, using proteoliposomes suspended in 100 mM potassium sulfate/50 mM potassium phosphate (pH 7) (1, 20); where noted, thiol-active agents were evaluated after a 10-min preincubation at 23°C with excess (100 μM) probe.
Site-Directed Fluorescence Labeling. Membranes suspended in 20 mM potassium phosphate (pH 8) were incubated with 40 μM Oregon green maleimide (OGM) for 10 min at 23°C before SDS/PAGE analysis of purified OxlT, as described (19, 22, 23). Fluorescence of SDS/PAGE gels was recorded by using a Fuji LAS 1000 system, whereas protein content was revealed by Coomassie brilliant blue staining. In all cases, the presence of an OGM-reactive cysteine was confirmed by parallel tests of denatured protein (data not shown; see ref. 22).
Disulfide Trapping. Cross-linking used membranes suspended in 20 mM Tris/HCl (pH 7.5) and exposed for 10 min at 23°C to 0.5 mM Cu(II)(1,10-phenanthroline)2, followed by a quench with 10 mM DTT and 10 mM EDTA (pH 7) (21). Aliquots incubated with or without Factor Xa protease (1 μg of protease per 10-μg sample) were processed for SDS/PAGE, with or without reductant (200 mM DTT), and after transfer to nitrocellulose, OxlT mobility was assessed by using antibody against the C-terminal polyhistidine extension.
Construction of the OxlT Homology Model. The “Project Mode” of the Swiss-Model sever was used to construct the OxlT homology model, with the structure of GlpT as template (15, 16). Initially, the N- and C-terminal halves of the OxlT sequence were separately overlapped at the MSF signature sequences in TM2 and TM8. Small gaps were then introduced to retain maximum homology according to clustalw (24), and this adjusted alignment was submitted to the Swiss-Model server to yield a first-generation model. A second-generation model was derived by insertion of a 3-aa gap into loop 6–7 to accommodate disulfide trapping of cysteines at positions 49 and 242 (see text). The final, third-generation model was obtained by using the “Build Loop” manipulation in deep viewer to rebuild loops 6–7, 7–8, 8–9, and 9–10, which had some residues with inappropriate ϕ/Ψ angles.
Chemicals. Toronto Research Chemicals provided carboxyethylmethanethiosulfonate (MTSCE) and methanethiosulfonateethylamine (MTSEA). Octyl-β-d-glucopyranside was from Roche–Calbiochem, and purified E. coli phospholipid was from Avanti Polar Lipids. [14C]Oxalate came from Perkin-Elmer Life Sciences, and Factor Xa protease was from New England Biolabs. OGM was purchased from Molecular Probes. Other reagents were of the highest purity available.
Results
Construction of the Homology Model. X-ray crystallography of two MFS members, LacY (8) and GlpT (9), provided an opportunity to construct a model for OxlT, a distantly related member of this broadly spread superfamily. Typically, such homology models are not considered useful if identity between template and target is <30% (25), and comparisons of the OxlT sequence with these potential templates indicated identities significantly lower than this nominal minimum: 18–20% for the comparison with GlpT (clustalw and t-coffee, respectively), and a somewhat lower value (16–18%) for that with LacY. Nevertheless, we pursued modeling of OxlT, using GlpT as the template, anticipating that the effort could be guided by parallel experiments. Furthermore, we speculated that these anion exchange transporters might share features not apparent from sequence identity, because they both use positively charged residues located in the hydrophobic sector as ligand-binding sites (9, 20).
Development of the OxlT homology model was an iterative process (see Materials and Methods) in which structural constraints provided by the Swiss-Model server were reframed after consideration of both preexisting and new experimental data. In the first stage, we estimated OxlT helix boundaries by using the GlpT template and two simple restrictions. First, we reasoned that the MFS signature sequence (GXXXDK/R) at the cytoplasmic boundaries of TM2 and TM8 would lie in comparable positions in the two molecules. Second, because OxlT and GlpT have very different central loops, we did not require connectivity in this region, but instead threaded the N- and C-terminal halves of the OxlT sequence independently. After introduction of small (one- to two-residue) gaps in the OxlT sequence at loops 8–9 and 9–10, to maximize agreement with the OxlT vs. GlpT clustalw alignment for TM9 and TM10, we submitted the alignment to the Swiss-Model server. This yielded a first-generation model in general agreement with existing experimental data, including OxlT topology as probed by site-directed fluorescence labeling (22) and the locations of several residues (Lys-355, Ser-359, Gln-63) expected to lie on the permeation pathway (19, 20, 23).
The preliminary model was also consistent with earlier observations that the TM2 and TM11 helical faces contain pathway-lining residues apposed in a way that supports disulfide trapping (21) (see below). In parallel work, we initiated trials of cross-linking of other targets, focusing on residues near the periplasmic boundaries of TM1 and TM7, and found that the G49C/N242C double-cysteine mutant displays cross-linking under the appropriate conditions (below). To accommodate this finding, we revised the first-generation model by inserting a three-residue gap into loop 6–7, effectively shortening the original OxlT central loop and driving the provisional TM7 sequence toward the periplasm by one helix turn; a similar effect was generated by rebuilding of loop 7–8. As a result, positions 49 and 242 were placed near the periplasmic border of their respective helices, and the periplasmic loop between TM7 and TM8 was correspondingly enlarged.
The modified alignment was resubmitted to the Swiss-Model sever for optimization, leading to a second-generation model that showed an excellent fit to all known experimental data, but which had, in the C-terminal half, a number of loop residues with ϕ/Ψ angles in the forbidden zone of a Ramachandran plot. A final, third-generation model was generated by using the “Build Loop” manipulation to rebuild loops 6–7, 7–8, 8–9, and 9–10, leading to an expansion of loop 7–8 by three residues formerly assigned to TM7 and incorporation into loop 8–9 of two residues from the cytoplasmic end of TM9.
It is instructive to compare the overall topology of the final model (Fig. 1A) with the model derived earlier from analysis of hydropathy (2), which had successfully guided much experimental work (19–23). Both models make comparable predictions as to the relative positions of residues critical to the study of OxlT. In particular, they each suggest that Gln-63, Ser-359 and Lys-355 are within the hydrophobic sector; as well, the homology model indicates these each face the permeation pathway, in accord with their accessibility to small hydrophilic probes (19, 20, 23). However, significant differences arise in considering the composition of both TM7 and TM8. For example, the nature of TM7 was altered to accommodate both cross-linking of positions 49 and 242 (below) and construction of loop 7–8. More striking, the idea that the MFS signature sequence is placed comparably in GlpT and OxlT results in wholesale reorganization of TM8, with the result that along with Lys-355 in TM11, the OxlT hydrophobic sector is now expected to have a second positive center, Arg-272, that might facilitate substrate (anion) binding (Fig. 1 B and C). For the same reason, whereas the earlier model had assigned Cys-271 to the cytoplasmic end of TM8, the current model places this residue within the body of that helix. Below, we describe tests conducted to determine whether these modifications are consistent with experimental findings.
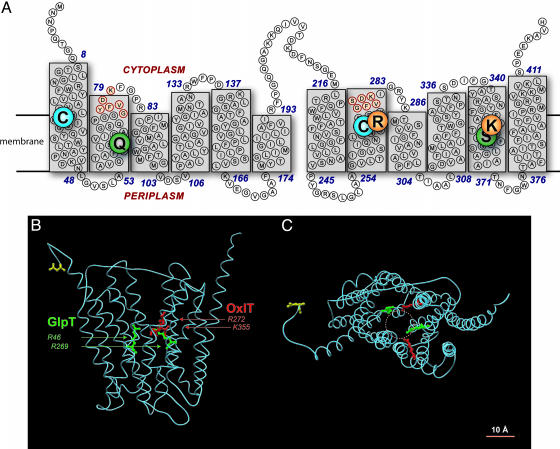
Fig. 1.
OxlT homology model. (A) OxlT topology. Boxes show transmembrane helices terminating in loops whose beginning and ending residue numbers are indicated; the MFS signature is noted in red. OxlT cysteines (Cys-28 and Cys-271) are shown by enlarged circles, as are the active-site residues, Arg-272 and Lys-355. Also emphasized are two residues (Gln-63 and Ser-359) known to lie on the permeation pathway. Solid lines indicate membrane thickness as reported for GlpT. (B) Ribbon tracing of the OxlT model, as viewed from the lipid phase, with the N-terminal asparagine (residue no. 2) indicated in yellow for reference. The extended side chains of the OxlT ligand-binding residues, Arg-272 and Lys-355, are shown in red. For comparison, ligand-binding residues in GlpT (Arg-46 and Arg-269) are shown in green. (C) OxlT as viewed from the cytoplasm and showing the active-site residues noted in B. In both OxlT and GlpT, these active-site residues are separated by 10 Å.
A Bifunctional Active Site. The homology model (Fig. 1) differs most strikingly from earlier models in its prediction that the OxlT hydrophobic core contains two positively charged residues, Arg-272 in TM8 and Lys-355 in TM11. The latter had been noted earlier (2, 20), and current evidence strongly supports the idea that Lys-355 facilitates binding of the anionic substrate, oxalate (20). Because oxalate is a dicarboxylate (-OOC-COO-) at physiological pH, Arg-272 is an attractive candidate as a second substrate-binding element; the homology model supports this speculation, because the positively charged guanidino group of Arg-272 faces the permeation pathway (Fig. 1C). To test this idea, we generated derivatives with either of three residues (Ala, Gln, Lys) at this position. After reconstitution of detergent extracts into oxalate-loaded proteolipomes, we readily monitored the oxalate self-exchange mediated by the parental protein, and although we found no detectable exchange for the R272A and R272Q variants, on extended assay we did observe oxalate transport by the R272K mutant (Fig. 2). Further kinetic study showed that relative to its parent, oxalate exchange by the R272K variant displays a 200-fold diminished catalytic function, reflecting both a lowered Michaelis constant and decreased maximal velocity for oxalate transport (mean values from four independent experiments showed Michaelis constants of 700 μM vs. 90 μM and maximal velocities of 50 vs. 1,430 nmol/min/mg, respectively) (Fig. 2, Insets).
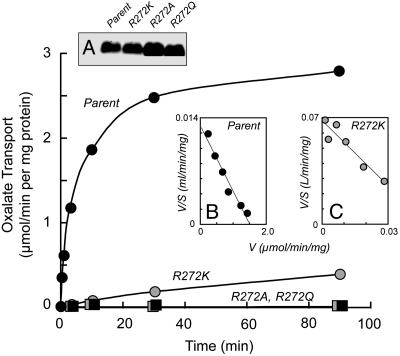
Fig. 2.
R272 is required for OxlT function. [14C]Oxalate transport by oxalate-loaded proteoliposomes for the indicated variants is shown. (Inset A) An immunoblot, developed by using antibody directed against the OxlT C-terminal polyhistidine tag, indicates reconstitution of comparable amounts of parental and R272K proteins and ≈2-fold higher recoveries for the R272A and R272Q derivatives. (Insets B and C) One of four experiments comparing the kinetics of [14C]oxalate exchange by parental and R272K proteins (see text).
The center of the GlpT substrate permeation pathway contains two basic residues (Arg-46, Arg-269) that serve as ligand-binding sites for inorganic phosphate or glycerol 3-phosphate (9, 17). The OxlT homology model suggests a parallel finding for OxlT, which requires both Arg-272 (Fig. 2) and Lys-355 (19, 20) for normal function. This parallel is greatly strengthened by a close examination of the GlpT and OxlT structures, which shows that although these active-site pairs originate from different locations in the two proteins (TM1 and TM7 in GlpT; TM8 and TM11 in OxlT), all four residues face the permeation pathway such that their side chains delimit a common domain, a discoid ≈10 Å in diameter and3Åin thickness, at the membrane center (Fig. 1 B and C). We conclude that GlpT and OxlT are organized in parallel fashion, and that in each case substrate (anion) binding occurs within a comparably structured domain.
Cysteine Accessibility in OxlT. Neither of the two cysteines in OxlT is required for normal function (19), but the placement of Arg-272 on the permeation pathway (Fig. 3) raises the possibility that Cys-271 might be an informative target for thiol-directed probes, in much the way that cysteine(s) placed along TM11 revealed its participation in substrate translocation (19, 20). We addressed the issue by generating the appropriate single-cysteine derivatives and probing their responses to three thiol-active agents [carboxyethylmethanethiosulfonate (MTSCE), methanethiosulfonateethylamine (MTSEA), and p-chloromercuribenzoic acid (pCMB)], each known to target the S359C variant on TM11 (19, 20), which served as the positive control in such work. We found that the lipid-soluble pCMB gave strong (≥90%) inhibition for all cysteine-containing variants, but did not affect the C27G/C271A cysteine-less derivative. By contrast, neither the parental wild-type protein nor its single-cysteine (C28G or C271A) or cysteineless derivatives responded to the water-soluble MTSEA and MTSCE; instead, these probes exerted the expected strong inhibition of the S259C mutant (data not shown). We conclude the resident OxlT cysteines are inaccessible to such hydrophilic probes.
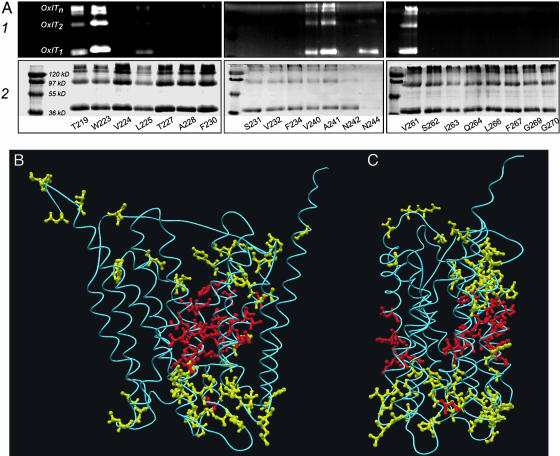
Fig. 3.
Accessibility to OGM. (A) Membranes from cells expressing the indicated single-cysteine derivatives were exposed to OGM, after which detergent-solubilized OxlT was purified and processed for SDS/PAGE. The fluorescence profile of the gel identifies proteins modified by OGM (Upper); Commassie brilliant blue staining of the same gel reveals total protein (Lower). Note that SDS/PAGE yields OxlT monomers, dimers, and oligomers (nmers), as indicated, because of an increased tendency for aggregation of mutants placed in the cysteine-less background (19). (B) Results of A mapped on the OxlT homology model as viewed from the lipid phase, together with 52 cases reported earlier and assigned to the cytoplasmic or periplasmic surfaces or to the hydrophobic core (22, 23). Coding with yellow and red indicates, respectively, residues that are or are not susceptible to in situ labeling by OGM. (C) Data from B as viewed from the lipid phase with the N terminus closest to the reader.
OGM Accessibility in OxlT. The topology initially suggested for OxlT was based, in part, on work in which cysteine substitution mutants were probed with OGM, an impermeant and fluorescent thiol-active agent (20, 22, 23). In that work, a collection of functional single-cysteine derivatives was evaluated for OGM accessibility, and reactive cysteines were assigned either a periplasmic or cytoplasmic location based on responses found by using intact cells (revealing positions facing the periplasm), or membrane preparations (in which both periplasmic and cytoplasmic surfaces are exposed). Cysteines that reacted only after protein denaturation were presumed to lie in the hydrophobic sector, where an absence of in situ reactivity was attributed to an elevated pK of the cysteine side chain or to inaccessibility due to the physical bulk of the probe. Data from this earlier work agree with predictions made from the OxlT homology model, suggesting that the method has merit as a tool to probe generic features of membrane topology. This concordance prompted us to test the homology model in regard to TM7 and TM8, because the model had altered the composition of these helices relative to earlier expectations. Accordingly, we generated and screened single-cysteine variants on TM7 and TM8 to identify those in which normal or near normal activity (≥20% parental specific activity) was retained; among these, we selected 22 derivatives, in some of which the target cysteine was presumed to lie near the OxlT cytoplasmic or periplasmic surfaces (likely accessible to OGM), whereas in others the target was expected to be in the hydrophobic interior (OGM inaccessible). After exposing membranes to OGM, accessibility was scored by examining SDS/PAGE fluorescence profiles of the purified proteins (Fig. 3A). The results agree with earlier findings. Thus, in all but one case, cysteines modified by this in situ test map to the cytoplasmic or periplasmic portions of OxlT, whereas those that do not react reflect positions within the hydrophobic core (Fig. 3 B and C). As noted in earlier work (22), one expects that in some instances the cysteinyl side chain will not be oriented suitably for OGM modification, even though the residue may lie outside the hydrophobic core. This appears to be the circumstance for position 242. Thus, although OGM does not modify the N242C derivative, that position can be modified in situ by the smaller impermeant probe, MTSET, which prevents the reaction of denatured protein with OGM (data not shown; see ref. 22).
These and earlier findings (20, 22, 23) are summarized by mapping of all 74 OGM-accessible (surface-exposed) or OGM-inaccessible (hydrophobic core) positions onto the OxlT homology model (Fig. 3 B and C). This maneuver shows a clear segregation of the two classes, with accessible positions lying at the cytoplasmic and periplasmic surfaces of the protein and inaccessible positions confined to the hydrophobic core. This finding suggests both that the homology model correctly predicts the patterns of accessibility for those residues tested (Fig. 3), and more generally that OGM accessibility is an informative tool for validation of similar models based on structural templates.
Proximity Revealed by Disulfide Trapping. As noted earlier, development of the homology model relied in part on a repositioning of TM7 to accommodate the results of induced disulfide trapping of a cysteine pair located near the ends of TM1 and TM7. In such work (Fig. 4), we used the Q63C/S359C cysteine pair (at the centers of TM2 and TM11) as a positive control (see ref. 21) and monitored disulfide formation by noting SDS/PAGE mobility after exposing oxidant-treated membranes to Factor Xa protease, using a specifically designed host protein containing the protease cleavage site in the OxlT central loop. In the absence of disulfide formation, Factor Xa cleavage yields a fragment (C-ter) that can be detected by antibody to the C-terminal polyhistidine affinity tag. In contrast, when a successful cross-link is present, protease cleavage reveals the C-ter fragment only if samples are also treated with DTT before electrophoresis. These expectations, along with behavior of the positive control, lead us to conclude that the G49C/242C pair, but not the G49C/L240C pair, shows successful disulfide trapping (Fig. 4A). To incorporate this information in building the homology model (Fig. 1), we moved TM7 residues toward the periplasm; because the position of Gly-49 was constrained by earlier work using both OGM accessibility (23) and disulfide trapping (21), that placed position no. 49 near the periplasmic end of TM2, where it can interact with position no. 370 at a comparable location near TM11. With this modification of TM7, linkage between cysteines at positions 49 and 242 was easily rationalized, because the two positions are at approximately the same depth relative to the membrane surface (near the periplasmic borders of TM2 and TM7), with side chain orientations that can accommodate a disulfide bridge (Fig. 4B).
Fig. 4.
Disulfide trapping at the OxlT periplasmic surface. (A) Membranes from cells expressing the indicated double-cysteine derivatives were exposed to copper phenanthroline to initiate disulfide trapping. After quenching the reaction, samples were incubated with or without Factor Xa protease and processed for SDS/PAGE (21), with reductant added before electrophoresis where indicated. Immunoblots were developed by using antibody directed against the OxlT C-terminal polyhistidine tag. (B) The OxlT model as viewed from the periplasmic surface (N terminus at the left). The residue pair that supports cross-linking (nos. 49 and 242) is shown in green; residue no. 240 is indicated in red. (C) Retrospective analysis of disulfide trapping in TM2 and TM11, using the ribbon diagram of Fig. 1B to display the target helices; TM2 residues highlighted in green cross-link with at least one of four residues in TM11 (shown in yellow), whereas TM2 residues indicated in red did not engage in cross-linking (21).
It was also of interest to carry out a retrospective analysis of cross-linking that had examined proximity between TM2 and TM11 (21). In that early study, TM2/TM11 double-cysteine variants were constructed such that multiple positions on TM2 were examined as potential partners for disulfide linkage with one of four possible positions in TM11. The observed patterns of disulfide trapping implied that TM2/TM11 helices abut one another; it was gratifying to find that the homology model required no further modification to incorporate these biochemical findings (Fig. 4C).
Discussion
Homology models provide insight that can corroborate (or challenge) earlier work and encourage the design of new experimental approaches for maximal impact. Such models are typically most useful if there is a relatively high degree of similarity between target and template (25), and the present case is exceptional in that, despite a low level of global identity between target (OxlT) and template (GlpT), the derived model readily accommodates findings from both earlier experiments and new studies. Such success presumably reflects that these transporters share not only a common membership in the MFS, but also a common biochemical mechanism (anion exchange), a comparably structured central binding domain (Fig. 3) and an equivalent set of conformational states. It is already clear that GlpT supports informative homology models for its immediate relatives in the organophosphate transporter family (ref. 10 and Q.Y. and P.C.M., unpublished data); the work described here extends this to the distantly related OxlT antiporter.
The MFS dominates the larger class of transporter proteins, and much historical work has been based on MFS members (see refs. 4 and 26), generating considerable interest in devising structural models of these targets. However, because crystallographic templates will likely be limited in number, it is essential to find independent ways to validate such exercises. For this reason, aside from developing a model for OxlT, we felt it important to explore three distinct kinds of experimental tests. Clearly, targeted mutagenesis was invaluable to analysis of the OxlT ligand-binding site (Fig. 2), and theoretical and experimental findings now strongly reinforce one another. Of equal impact was the broader use of cysteine-scanning mutagenesis coupled to labeling by OGM, which provided a reasonably clear experimental view of general topology (Fig. 3). In fact, the combined use of this method and that of disulfide trapping (Fig. 4) provided the main justification for the nature of TM7. We note that such experimental work led us to assign residues 217–244 to TM7, whereas in the recent model of Hirae and Subramaniam (12), this same helix involves residues 220–248, as in our initial attempts. Further support for the suggestion made here is found in earlier work (21) showing that position 248 is sufficiently exposed to serve as a cleavage site for Factor Xa; indeed, this residue is among the most accessible of those at the periplasmic surface (see Protein Data Base ID code 1ZC7). The two OxlT models diverge in other ways as well, largely as a result of their different underlying goals and assumptions. Thus, our work uses the entire GlpT structure as template to model a ligand-free state in which active-site residues are open to the cytoplasm. By contrast, Hirae and Subramaniam (12) address a conformation in which OxlT has bound ligand. As a result, this earlier model focuses on aspects of GlpT helix packing that, after an appropriate rigid-body rotation of N- and C-terminal domains (see below), best fit the OxlT 6.5-Å structure (6, 7). This finding led to the proposal that helix organization in the GlpT C-terminal domain (GlpTTM7–12) is the preferred scheme for both halves of OxlT (OxlTTM1–6 and OxlTTM7–12). That choice, derived from a structural alignment of GlpT and OxlT helices (12), is consistent with the known symmetry of MFS proteins (6–9), but necessarily leads to a model that differs from that shown here (Fig. 1) in both the general aspects of helix packing (especially in the OxlT N-terminal domain) and in the specifics of how residues in the OxlT N- and C-terminal domains articulate against one another. In particular, our use of the full GlpT backbone as template better accommodates the experimental finding of (interdomain) cross-linking between TM1 and TM7 and between TM2 and TM11 (Fig. 4), and also enables full extension of the Lys-355 side chain into the permeation pathway (Fig. 1).
A major impact of these models has been the inference that two positively charged residues in the OxlT hydrophobic sector, Arg-272 and Lys-355, serve to bind the divalent substrate, oxalate (-OOC-COO-). Several lines of evidence are now consistent with this idea. Thus, it is clear that Arg-272 is required for normal OxlT function (Fig. 2), as is Lys-355 (19, 20). Moreover, comparison of OxlT and GlpT (Fig. 1) reveals that the paired ligand-binding residues of both transporters extend into the permeation pathway in the same fashion. Notably, in the one case (OxlT), these basic side chains orient toward the periplasm, whereas in the other example (GlpT) they point toward the cytoplasm, yet each pair terminates within a small discoidal region (10 Å × 3 Å) that serves as the ligand-binding domain (Fig. 3). Importantly, such dimensions are consistent with those found for ligand association in oxalate binding proteins of known structure (27, 28).
The GlpT ligand-binding residues lie at symmetrical positions (on TM1 and TM7) in the N- and C-terminal halves of the protein, and it is proposed that binding of substrate (phosphate or glycerol 3-phosphate) triggers conformational changes allowing rigid-body movement of the two six-helix domains against one another (9, 29). For this reason, that the OxlT ligand-binding residues lie wholly in the OxlTTM7–12 domain (on TM8 and TM11) raises mechanistic questions. If, as one expects, rotation of the six-helix N- and C-terminal domains occurs in both systems, we anticipate that such events in OxlT will show a stronger dependence on interdomain helix–helix interactions than in GlpT, where substrate itself links the two six-helix clusters. It is also worth comment that the findings in OxlT remain compatible with the more general view that substrate-binding occurs at the interface of interacting domains (26), because in the MFS, a pseudo-twofold symmetry is reflected not only in the N- and C-terminal six-helix domains, but also within each six-helix cluster (6–9) as an internal 3 plus 3 symmetry. Accordingly, ligand-binding sites in OxlT resemble those in GlpT in that Arg-272 (TM8) and Lys-355 (TM11) are in symmetry-related positions at the interface of the paired three-helix units comprising the OxlTTM7–12 domain. In turn, this prompts a second mechanistic question, because one can imagine that articulation of the three-helix units (OxlTTM7–9 and OxlTTM10–12) against each other may also contribute to alternating accessibility of the binding site. Although such conformational movements are expected to be relatively modest owing to the short loops that connect these domains, some broad features of OxlT suggest that the required conformational changes may also be minimal. In particular, oxalate itself is relatively small, and the oxalate self-exchange reaction is unusually rapid (> 2,000 per s) (1, 30). In other systems, especially if interdomain loop size is increased, the functional impact of such internal mobility should not be neglected.
A separate question relates to whether a ligand-binding site defined by Arg-272 and/or Lys-355 also facilitates transport of the alternative physiological substrate, monovalent formate (HCOO-) (1), or whether there is an as yet unidentified site that guides movement of this secondary substrate. Unpublished data suggest that both formate and oxalate interact with the single central binding site (Q.Y., R. Sarker, and P.C.M., unpublished data), and resources such as those provided by the model described here should accelerate the experimental approach to this and related issues.
Acknowledgments
This work was supported by National Institutes of Health Grant GM24195 and National Science Foundation Grant MCB-0235305.
Author contributions: Q.Y., X.W., L.Y., E.M., Y.-M.K., and P.C.M. designed research; Q.Y., X.W., L.Y., M.M., and Y.-M.K. performed research; Q.Y., X.W., L.Y., M.M., Y.-M.K., and P.C.M. analyzed data; and P.C.M. wrote the paper.
Abbreviations: MFS, major facilitator superfamily; OGM, Oregon green maleimide.
Data deposition: The atomic coordinates and structure factors have been deposited in the Protein Data Bank, www.pdb.org (PDB ID code 1ZC7).
References
- 1.Anantharam, V., Allison, M. J. & Maloney, P. C. (1989) J. Biol. Chem. 264, 7244-7250. [PubMed] [Google Scholar]
- 2.Abe, K., Ruan, Z.-S. & Maloney, P. C. (1996) J. Biol. Chem. 271, 6789-6793. [DOI] [PubMed] [Google Scholar]
- 3.Pao, S. S., Paulsen, I. T. & Maier, M. H., Jr. (1998) Microbiol. Mol. Biol. Rev. 62, 1-34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Henderson, P. J. F. (1993) Curr. Opin. Cell Biol. 5, 708-721. [DOI] [PubMed] [Google Scholar]
- 5.Maiden, M. C., Davis, E. O., Baldwin, S. A., Moore, D. C. & Henderson, P. J. (1987) Nature 325, 641-643. [DOI] [PubMed] [Google Scholar]
- 6.Hirai, T., Heymann, J., Shi, D., Sarker, R. I., Maloney, P. C. & Subramaniam, S. (2002) Nat. Struct. Biol. 9, 597-600. [DOI] [PubMed] [Google Scholar]
- 7.Hirai, T., Heymann, J. A., Maloney, P. C. & Subramaniam, S. (2003) J. Bacteriol. 185, 1712-1718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Abramson, J., Smirnova, I., Kasho, V., Verner, G., Kaback, H. R. & Iwata, S. (2003) Science 301, 610-615. [DOI] [PubMed] [Google Scholar]
- 9.Huang, Y., Lemieux, M. J., Song, J., Auer, M. & Wang, D. N. (2003) Science 301, 616-620. [DOI] [PubMed] [Google Scholar]
- 10.Almqvist, J., Huang, Y., Hovmoller, S. & Wang, D. N. (2004) Biochemistry 43, 9289-9297. [DOI] [PubMed] [Google Scholar]
- 11.Vardy, E., Arkin, I. T., Gottschalk, K. E., Kaback, H. R. & Schuldiner, S. (2004) Protein Sci. 13, 1832-1840. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hirae, T. & Subramaniam, S. (2004) Biophys. J. 87, 3600-3607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lagerstedt, J. O., Voss, J. C., Wieslander, A. & Persson, B. L. (2004) FEBS Lett. 578, 262-268. [DOI] [PubMed] [Google Scholar]
- 14.Peitsch, M. C. (1995) Bio/Technology 13, 658-660. [Google Scholar]
- 15.Guex, N. & Peitsch, M. C. (1997) Electrophoresis 8, 2714-2723. [DOI] [PubMed] [Google Scholar]
- 16.Schwede, T., Kopp, J., Guex, N. & Peitsch, M. C. (2003) Nucleic Acids Res. 31, 381-3385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fann, M.-C., Davies, A. H., Varadhachary, A., Kuroda, T., Sevier, C., Tsuchiya, T. & Maloney, P. C. (1998) J. Membr. Biol. 164, 187-195. [DOI] [PubMed] [Google Scholar]
- 18.Fu, D. & Maloney, P. C. (1997) J. Biol. Chem. 272, 2129-2135. [DOI] [PubMed] [Google Scholar]
- 19.Fu, D. & Maloney, P. C. (1998) J. Biol. Chem. 273, 17962-17967. [DOI] [PubMed] [Google Scholar]
- 20.Fu, D., Sarker, R. I., Abe, K., Bolton, E. & Maloney, P. C. (2001) J. Biol. Chem. 276, 8753-8760. [DOI] [PubMed] [Google Scholar]
- 21.Kim, Y. M., Ye, L. & Maloney, P. C. (2001) J. Biol. Chem. 276, 36681-36686. [DOI] [PubMed] [Google Scholar]
- 22.Ye, L., Jia, Z., Jung, T. & Maloney, P. C. (2001) J. Bacteriol. 183, 2490-2496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ye, L. & Maloney, P. C. (2002) J. Biol. Chem. 277, 20372-20378. [DOI] [PubMed] [Google Scholar]
- 24.Thompson, J. D., Higgins, D. G. & Gibson, T. J. (1994) Nucleic Acids Res. 22, 4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Marti-Renom, M. A., Stuart, A. C., Fiser, A., Sanchez, R., Melo, F. & Sali, A. (2000) Annu. Rev. Biophys. Biomol. Struct. 29, 291-325. [DOI] [PubMed] [Google Scholar]
- 26.Maloney, P. C. (1994) Curr. Opin. Cell Biol. 6, 571-582. [DOI] [PubMed] [Google Scholar]
- 27.Kavanagh, K. L., Elling, R. A. & Wilson, D. K. (2004) Biochemistry 43, 879-889. [DOI] [PubMed] [Google Scholar]
- 28.Walsh, M. A., Otwinowski, Z., Perrakis, A., Anderson, P. M. & Joachimiak, A. (2000) Structure Fold. Des. 8, 505-514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lemieux, M. J., Huang, Y. & Wang, D. N. (2004) Curr. Opin. Struct. Biol. 14, 405-412. [DOI] [PubMed] [Google Scholar]
- 30.Ruan, Z. S., Anantharam, V., Crawford, I. T., Ambudkar, S. V., Rhee, S. Y., Allison, M. J. & Maloney, P. C. (1992) J. Biol. Chem. 267, 10537-10543. [PubMed] [Google Scholar]