Abstract
Purpose
The abundance and distribution of tumor-infiltrating lymphocytes (TILs) as well as that of other components of the tumor microenvironment is of particular importance for predicting response to immunotherapy in lung cancer (LC). We describe here a pilot study employing artificial intelligence (AI) in the assessment of TILs and other cell populations, intending to reduce the inter- or intra-observer variability that commonly characterizes this evaluation.
Design
We developed a machine learning-based classifier to detect tumor, immune, and stromal cells on hematoxylin and eosin-stained sections, using the open-source framework QuPath. We evaluated the quantity of the aforementioned three cell populations among 37 LC whole slide images regions of interest, comparing the assessments made by five pathologists, both before and after using graphical predictions made by AI, for a total of 1110 quantitative measurements.
Results
Our findings indicate noteworthy variations in score distribution among pathologists and between individual pathologists and AI. The AI-guided pathologist's evaluations resulted in reduction of significant discrepancies across pathologists: three comparisons showed a loss of significance (p > 0.05), whereas other four showed a reduction in significance (p > 0.01).
Conclusions
We show that employing a machine learning approach in cell population quantification reduces inter- and intra-observer variability, improving reproducibility and facilitating its use in further validation studies.
Keywords: Digital pathology, Machine learning, Pathology image, Computer-aided tool, Whole slide images, Lung cancer, NSCLC, Tumor-infiltrating lymphocytes, QuPath
Highlights
-
•
Research explores the impact of digitization on traditional pathology practices.
-
•
Data from H&E slides to validate ML tools for precise cell population analysis.
-
•
Pathologists' evaluations show higher accuracy and consistency with AI assistance.
-
•
Non-parametric tests confirm significant improvements with AI-aided assessments.
-
•
ML tools enhance pathology workflows, providing precise cell population quantification.
Introduction
The application of light microscopy for examining hematoxylin and eosin (H&E) tissue slides remains an essential practice in the field of cancer diagnosis. Such a technique requires the pathologist to accurately identify and distinguish the assorted histopathological components within complex images. For instance, LC histopathology is characterized by a complex interplay of tumor cells, immune cells (lymphocytes, plasma cells, macrophages, and granulocytes), fibroblasts, and endothelial cells.1 Many of said measures are valuable biomarkers in the clinical field: several studies show how the density of tumor-infiltrating lymphocytes (TILs) correlates with better clinical outcomes2,3 such as longer disease-free survival (DFS) or improved overall survival (OS) in multiple cancer types.4 Nonetheless, studies have found substantial inter-reader variability in the assessment of TILs density using H&E slides and this has limited the use of TILs density as a prognostic marker for non-small cell lung cancer (NSCLC).5
Moreover, tumor cell percentage within formalin-fixed paraffin-embedded samples is a critical factor in the biomarker's evaluations using Next Generation Sequencing (NGS) panels, such as the ability to detect important low-frequency somatic mutations in target cancer therapy.
Finally, it is known that evaluating the presence and quantity of different cell populations is a process that is time-consuming and subjective. Furthermore, it generates considerable inter- and intra-observer variation and is a qualitative or, at best, semi-quantitative approach.6, 7, 8
However, research in the field of diagnostic imaging is expanding with promising results. Although diagnosis based on examination of the classic histological slide using optical microscopy remains the gold-standard, in recent years, advances in the digitization of histological specimens have led to digital slides or whole slide images (WSIs) production.9, 10, 11 Recently the Food and Drugs Administration (FDA) approved the use of WSIs for primary diagnostic use.12,13 In a few years, most new pathology slides will be digitized, enabling the development and clinical implementation of various digital pathology-based diagnostic and prognostic biomarkers that are likely to provide decision support to traditional pathology interpretation in the clinical setting.1 New analytic approaches include the application of machine learning (ML) in image analysis in various fields such as pathology diagnostic14 and dermatology.15
The advancement of sophisticated ML algorithms has the potential to enhance the analysis of pathological images and assist pathologists in their diagnostic duties, particularly in identifying neoplasms, detecting tumor metastases, and carry out accurate quantifications of distinct cell populations, including tumor and inflammatory cells. Multiple articles have suggested ML methodologies to assist pathologists in quantifying tumor cell fractions on digital scans. However, these endeavors predominantly encounter a common constraint: establishing a dependable ground-truth definition.16,17 In this work, we evaluate whether ML-based tools may improve pathology workflows to enable in-depth, reproducible, and accurate characterization of the tumor microenvironment, all with a limited manual curation effort and an objective evaluation strategy.
Materials and methods
Whole slide dataset
This study used anonymized archived LC cases from the Regina Elena National Cancer Institute Biobank BBIRE. The dataset consists of scannable images from the surgical removal of LC (Table S1). The original diagnostic slides were derived from sections (4–5 μm thickness) cut from formalin-fixed paraffin-embedded whole-slide blocks H&E stained. Thirty-seven H&E slides were scanned at 0.25 μm /pixel (40×) acquisition resolution using an Aperio AT2 (Leica Biosytems Inc.). Default instrument settings were used for scanning parameters; default autofocus was used and the scanner automatically selected the focus points number, but in a few cases, manual focus was applied to optimize sharpness. A single focal plane was used, without Z-stacking. Images were stored in a proprietary format (.SVS) and uploaded to a local WSI management server (eSlideManager, Leica). Slides were scanned in anonymized format; they were retrospective archival material, and the results did not have any impact on the management of the patient. All patients included in the study provided written Broad Research Usage Informed Consent before surgery.
Machine learning
In this study, integrated tools from QuPath v.0.418 were used for image analysis. The scanned images of each slide were never analyzed in their entirety but rather an expert pathologist selected regions of interest (ROIs). The selection criteria for the ROIs were meticulously defined to ensure comprehensive representation of the tissue architecture and tumor characteristics. ROIs were chosen based on the presence of well-defined tumor regions, ensuring a diverse and representative dataset. Additionally, areas with sufficient cellularity and minimal artifacts were prioritized to facilitate accurate annotation and subsequent algorithm training. The target cell populations in this study include tumor cells, immune cells, and stromal cells such as fibroblast and endothelial cells. TILs were defined as mononuclear immune cells including lymphocytes and plasma cells.19 To ensure an accurate analysis of cell populations, we calculated the stain vector for each image to perform color deconvolution in order to perform a color normalization for each image. (Fig. 1A). This method allows us to separate the staining components20 and reduces staining variability and ensures more consistent and accurate tissue analysis (Supplementary Information).
Fig. 1.
Graphical Abstract. Depiction of all the steps devoted to the training and the evaluation of the cell classification model. (A) Pre-processing step; (B) training step; (C) test step.
To train and evaluate the cell classification model a total of 82 ROIs, which comprised 41 pairs, were utilized for training the classifier (Fig. 1B). The ROIs were manually drawn to construct the training set from 13 LC H&E digital slides. Each pair is characterized by a comparable number of cells and cell classes. One element of each pair was named Sample, whereas the other was assigned as Training. Sample regions provide learning input, whereas Training regions guide model parameter optimization for accurate classification. Next, cell detection within these areas was performed with the deep learning nuclei segmentation algorithm StarDist v.4.0.21 The same set of parameters were used for the segmentation of all the other ROIs drawn. Segmented nuclei were manually annotated within the Sample ROIs. The quantity of cells to be annotated in the Sample ROIs was not predetermined but rather deduced throughout the annotation process based on the software's ability to characterize cells in the Training. As a result of this dynamic annotation process, the annotation of 370 cells resulted adequate to learn enough features for cell type discrimination. Among these, 150 were labeled as tumor cells, 86 as immune cells, and 132 as stromal cells. Thereafter, a random forest classifier model was trained on the Training ROIs, defined in the previous step, using the integrated QuPath function Train object classifier. The whole training process is documented in the Supplementary Information. This methodology strongly reduces the time dedicated to manual annotation process, while also avoiding overfitting.
To test the classifier, 13 different WSIs of LC H&E distinct from those used in the training set were chosen. A total of 43 annotations were drawn and collectively named Test ROIs with an average area of 44,459 square micrometers and a total of 19,477 cells were identified (Fig. 1C). Each classification made by the classifier was critically reviewed by an expert pathologist, identifying false positives and negatives, as well as true positives and negatives for each cell type, to calculate accuracy, sensitivity, and F1 score (Table S2).
AI-pathologist comparative analysis
A ROI, a rectangle containing a representative tumor area, was selected on a further 37 digital slides for comparative analysis with pathologist. The ROI area (5.8 mm2) was the same for all 37 slide images, within the tumor without including significant areas of necrosis or background tissue. First, all 37 ROIs were classified using the trained model. The pathologists were asked to estimate the percentage of tumor, immune, and stromal cells present in each ROI in two rounds. In the first round (Round I), pathologists provided estimates without access to AI results. In the second round (Round II), they reconsidered their evaluations, incorporating graphical predictions generated by the AI. Additionally, each pathologist provided ratings on the efficacy of AI classification for each image.
Statistical analysis
Percentage assignments by pathologists and AI were analyzed by non-parametric statistical tests (Kruskal–Wallis and Wilcoxon rank-sum test α = 0.05). For the correlation study, Kendall's correlation coefficient was also performed. All the statistical surveys were conducted using R v. 4.2.2. Comprehensive details regarding the calculations can be found in Supplementary Information.
Results
The effect of AI integration on pathologist evaluations
We conducted a comparative analysis of tumor, immune, and stromal cell assessments in 37 ROIs related to lung cancer (LC), involving five pathologists. This assessment was carried out both before and after integrating AI-generated graphical predictions, resulting in a total of 1110 measurements. Overall, in Round II, there was a percentual increase in pathologists' assignments of tumor and immune cell classes (49.2% for tumor cells and 32.7% for immune cells in Round II, compared to 47.3% for tumor cells and 21.7% for immune cells in Round I), whereas there was a decrease in the percentual assignment of stromal cells (12.1% in Round II compared to 21.1% in Round I) (Fig. 2A).
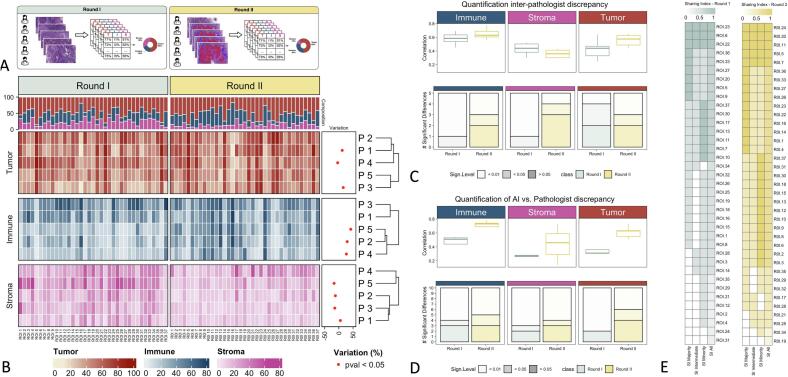
Fig. 2.
(A) Illustration of the pathologist's estimation rounds. Round I and Round II have been performed on the same ROIs with and without the visual support of AI-based cell classification. (B) Pathologist's estimations heatmap. The heatmap shows the percentage of the three cell types assigned by each of the pathologists (P1–P5) for all ROIs. For each ROI, the top annotation describes the average cell composition. Statistically significant percentual variation (across ROIs) in estimates provided by pathologists in Round I and Round II is described and shown in the right annotation. (C) Quantification of inter-pathologist discrepancy in Round I vs Round II. The boxplots represent Kendall correlation coefficients, calculated on the estimates given by all couples of pathologists (across ROIs). The bar plots represent the number of significant Wilcoxon tests performed on the estimates given by all couples of pathologists (across ROIs), at two different p-value thresholds (0.05 and 0.01). (D) Quantification of AI vs. Pathologist discrepancy. The boxplots represent Kendall correlation coefficients, calculated on the estimates given by the AI vs. each of the pathologists (across ROIs). The bar plots represent the number of significant Wilcoxon tests performed on the estimates given by the AI vs. each of the pathologists (across ROIs), at two different p-value thresholds (0.05 and 0.01). (E) Sharing Index (SI) heatmap in Round I and Round II for all three hierarchical positions as well as majority, minority, and intermediate hierarchical position individually. ROIs on the y-axis are sorted from the highest to the lowest global SI. A SI of 1 indicates complete agreement, whereas a SI of 0 indicates the maximum level of discrepancy across pathologists.
Our analysis revealed a significant variation in the pathologists' assignments between the two rounds (p-value < 0.05, Wilcoxon test). Specifically, a lower level of discrepancy was found across pathologists' assignments in the AI-aided assignments of Round II, because three comparisons showed a loss of significance (p > 0.05), whereas other four showed a reduction in significance (p > 0.01) (Fig. 2B).
Increased pathologist consensus regarding cell estimates
When comparing estimates between pairs of pathologists, there was an overall increase in concordance, as illustrated in Fig. 2C. Specifically, whereas the correlation between estimates for immune and stromal cells remained similar (box plots), the number of significant differences decreased (bar plots). In the case of tumor cells, the number of significant differences (Kruskal–Wallis tests p-values <0.05) remained unchanged, yet the correlation increased.
The assessment of pathologists' assignments compared to those generated by AI revealed a notable rise in the average correlation coefficient and a decrease in the number of significant differences across the three cellular populations (Fig. 2D). Taken together, these results indicate that the pathologist's percentage assignments were influenced remarkably by the AI-generated graphical prediction visualization in Round II.
AI performance in cell population hierarchy evaluation
Additionally, we aimed to evaluate AI performance based solely on the hierarchy of cell populations determined by pathologists, irrespective of their absolute counts. This approach led to the identification of three categories: majority, intermediate, and minority cell populations. Furthermore, we introduced the concept of a Sharing Index to quantify agreement among pathologists regarding hierarchical assignments through an index ranging from 0 to 1 (Supplementary Material).
In Round I, all five pathologists assigned the same hierarchy of cell populations in 3 out of 37 ROIs, which increased to 5 in Round II (Fig. 2E). The classifier also provided an identical hierarchy in this subset. The Sharing Index generally increased from Round I to Round II, with half of pathologists agreeing on either the majority or the minority class in 28 ROIs, as opposed to 16 ROIs in Round I. The AI assigned the same hierarchy as the majority of pathologists in 22 ROIs, the same hierarchy as the minority (i.e., at least one pathologist) in 6 ROIs, and a different hierarchy from all pathologists in 6 ROIs. These findings indicated an increase in cases of complete agreement and partial agreement among pathologists in Round II. Furthermore, in instances of absolute concordance, complete agreement was also observed between pathologists and AI.
Finally, the output of the ML classifier, assessed at the cell level by an expert pathologist, demonstrated high accuracy on the test set (Table S2). AI-assisted classifications received grades from pathologists that were consistently above passing grades, indicating that the classifier effectively categorized the segmented cells in most cases (Table S3).
Discussion
High levels of TILs have emerged as promising prognostic biomarkers in NSCLC patients.22 Additionally, mounting evidence suggests that TILs are strongly associated with favorable response rates to immunotherapy in NSCLC patients.23 The International Working Group on Immuno-Oncology has published a set of guidelines for the manual evaluation of cell populations, such as TILs in H&E slides.24,25 However, despite these standardization efforts, their translational adoption into clinical practice has been limited due to the subjective nature and degree of variability of the assessments.
Assessing cell populations in tissue samples by pathologists can often exhibit significant variability.26,27 Due to the difficulty of making such assessments, the adjacent microenvironment's cell populations are often overlooked.
In our study, we utilized the QuPath software to develop a classifier capable of numerically quantifying tumor, immune, and stromal cells in standard H&E histological sections of LC. Whereas the use of StarDist for nuclei detection and ML-based cell classification appear to be promising tools for cell quantification, potential limitations related to the reliability of the segmentation and classification algorithms must be considered. These limitations may stem from biological artifacts or overfitting of the ML models. To mitigate these limitations and maximize the classifier's capabilities, we took care to avoid artifact-rich areas during the segmentation process and followed a meticulous training procedure.
Our findings reveal that, despite employing the estimate stain vector functions, the classifier exhibited high sensitivity towards the presence of out-of-focus areas and variations in coloration. These variations may be attributed to dissimilar coloring protocols or technical issues, such as suboptimal section thickness and degraded dyes or color variation among different laboratories.28,29 To overcome this challenge, cutting-edge automatic algorithms have been developed to identify the optimal regions for such analyses, thereby standardizing the quality of whole-slide images utilized in research.30
Not only did the ML classifier showed a general high level of accuracy, but played a crucial role in enabling the pathologists to assign more values that are comparable during the second evaluation. As demonstrated there exists a discernible disparity in the assignments of each pathologist even with the AI utilization.
The AI-generated percentage for each cell population is primarily based on the area of segmented nuclei, whereas pathologists consider both the nuclei and the extracellular matrix when evaluating stromal cells. However, the percentages assigned to stroma by the pathologists were reduced (−42.7%) after viewing the AI's graphical predictions, suggesting a potential overestimation of stromal cells by the pathologists in Round I due to the challenge associated with detecting individual stromal cells in a complex context, at the expense of the other two cellular components (tumor cells and immune cells), with a percentage increase in Round II of +4% for tumor cells and+50.8% for immune cells. This disparity can be attributed to the challenge pathologists encounter in numerically and reproducibly quantifying different cell populations within a given area.31,32
In conclusion, our objective was not to create a completely automated ML classifier for cell populations but rather to develop a computer-assisted tool that can aid pathologists in accurately estimating the proportion of various cell populations, resulting in a marked reduction of inter- and intra-observer variability and a significant improvement in reproducibility. Particularly, this tool can be beneficial in scenarios such as determining the percentage of cancer cells for molecular testing or evaluating the degree of immune infiltrate as a prognostic indicator for immunotherapy response.
Funding
This work was supported by Ricerca Corrente grants (Ministero della Salute) for IRCCS Istituto Nazionale Tumori Regina Elena. MP was supported by the European Union (Next Generation EU), Italian NRRP project code IR0000031 - Strengthening BBMRI.it - CUP B53C22001820006. GC, MP, EG were supported at IFO-IRE via BBMRI-ERIC on the EUCAIM Cancer Imaging project, GA 101100633 on call DIGITAL-2022-CLOUD-AI-02.
Author contribution
EG and MP performed study concept and design; EG, MP, DG and MB performed development of methodology and writing, review, and revision of the paper; EG, DG, MB and BAMA, provided acquisition, analysis and interpretation of data, and statistical analysis; DG, MB, SB, BAMA, SDM, TD, EM, AA, PV, EP, MLV, PN, GC, MP provided technical and material support. All authors read and approved the final paper.
Declaration of competing interest
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Matteo Pallocca reports a relationship with Dexma srl that includes: consulting or advisory. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this article.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jpi.2024.100400.
Contributor Information
Enzo Gallo, Email: enzo.gallo@ifo.it.
Davide Guardiani, Email: davide.guardiani.roma@gmail.com.
Martina Betti, Email: martina.betti@ifo.it.
Brindusa Ana Maria Arteni, Email: anamaria.arteni@ifo.it.
Simona Di Martino, Email: simona.dimartino@ifo.it.
Sara Baldinelli, Email: sara.baldinelli@ifo.it.
Theodora Daralioti, Email: daraliotitheodora@yahoo.it.
Elisabetta Merenda, Email: elisabetta.merenda@uniroma1.it.
Andrea Ascione, Email: andrea.ascione@uniroma1.it.
Paolo Visca, Email: paolo.visca@ifo.it.
Edoardo Pescarmona, Email: edoardo.pescarmona@ifo.it.
Marialuisa Lavitrano, Email: marialuisa.lavitrano@unimib.it.
Paola Nisticò, Email: paola.nistico@ifo.it.
Gennaro Ciliberto, Email: gennaro.ciliberto@ifo.it.
Matteo Pallocca, Email: matteo.pallocca@cnr.it.
Appendix A. Supplementary data
Supplementary fig. 1.

The illustration shows one of the ROIs used for training and assessment. The sample ROI was used for the sample assignment. The selected nuclei exhibited a Point object within their respective areas. The training ROI (on the right) was employed for performance evaluations and the calculation of classification metrics. In this instance, the Live Prediction was active. The AI classified the nuclei in red as tumoral cells, the nuclei in blue as immune cells, and the nuclei in magenta as stromal cells.
The document outlines methods for color deconvolution, nuclei segmentation with StarDist, and training an object classifier for Tumor, Immune, or Stromal cell categorization. It includes a sharing index for pathologist agreement and statistical tests, ensuring improved accuracy and reproducibility in histological analyses.
Histological characteristics of WSIs in which ROIs were drawn and analyzed for the training set, test set, and comparison with pathologists' estimates.
Metrics for the AI-based classification in the Training Set (right) and Test Set (left). The metrics include precision, recall, and F1 score, which were calculated based on the model's predictions and the ground truth labels. The ground truth labels were determined by a senior pathologist, an expert in LC.
Average rates given by the five pathologists to the graphical prediction carried out by the AI on 37 ROIs. Pathologists assigned a rating between 1 and 5, with 1 indicating a completely incorrect classification, 2 indicating a classification with more errors than correct classifications, 3 indicating a classification that is sufficient or has fewer errors than correct classifications, 4 indicating a good classification but with some negligible errors, and 5 indicating a perfect classification.
Data availability
The datasets generated during and/or analyzed during the current study are available in the github repository DP_BBIRE_ML_CompositionClassifier [https://gitlab.com/bioinfo-ire-release/bbire].
References
- 1.Corredor G., Wang X., Zhou Y., et al. Spatial architecture and arrangement of tumor-infiltrating lymphocytes for predicting likelihood of recurrence in early-stage non-small cell lung cancer. Clin Cancer Res Off J Am Assoc Cancer Res. 2019;25(5) doi: 10.1158/1078-0432.CCR-18-2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Geng Y., Shao Y., He W., et al. Prognostic role of tumor-infiltrating lymphocytes in lung cancer: a meta-analysis. Cell Physiol Biochem Int J Exp Cell Physiol Biochem Pharmacol. 2015;37(4) doi: 10.1159/000438523. [DOI] [PubMed] [Google Scholar]
- 3.Bremnes R., Donnem T., Busund L. Importance of tumor infiltrating lymphocytes in non-small cell lung cancer? Ann Transl Med. 2016;4(7) doi: 10.21037/atm.2016.03.28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Bruni D., Angell H., Galon J. The immune contexture and Immunoscore in cancer prognosis and therapeutic efficacy. Nat Rev Cancer. 2020;20(11) doi: 10.1038/s41568-020-0285-7. [DOI] [PubMed] [Google Scholar]
- 5.Brambilla E., Le Teuff G., Marguet S., et al. Prognostic effect of tumor lymphocytic infiltration in Resectable non-small-cell lung cancer. J Clin Oncol Off J Am Soc Clin Oncol. 2016;34(11) doi: 10.1200/JCO.2015.63.0970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.van den Bent M. Interobserver variation of the histopathological diagnosis in clinical trials on glioma: a clinician’s perspective. Acta Neuropathol. 2010;120(3) doi: 10.1007/s00401-010-0725-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Elmore J., Longton G., Carney P., et al. Diagnostic concordance among pathologists interpreting breast biopsy specimens. JAMA. 2015;313(11) doi: 10.1001/jama.2015.1405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Linder N., Taylor J.C., Colling R., et al. Deep learning for detecting tumour-infiltrating lymphocytes in testicular germ cell tumours. J Clin Pathol. 2019;72(2):157–164. doi: 10.1136/jclinpath-2018-205328. [DOI] [PubMed] [Google Scholar]
- 9.Farris A.B., Cohen C., Rogers T.E., Smith G.H., From the Department of Pathology EU, Atlanta, Georgia Whole slide imaging for analytical anatomic pathology and telepathology: practical applications today, promises, and perils. Arch Pathol Lab Med. 2023;141(4):542–550. doi: 10.5858/arpa.2016-0265-SA. [DOI] [PubMed] [Google Scholar]
- 10.Evans A.J., Salama M.E., Henricks W.H., Pantanowitz L. Implementation of whole slide imaging for clinical purposes: issues to consider from the perspective of early adopters. Arch Pathol Lab Med. 2017;141(7) doi: 10.5858/arpa.2016-0074-OA. [DOI] [PubMed] [Google Scholar]
- 11.Fraggetta F., Garozzo S., Zannoni G., Pantanowitz L., Rossi E. Routine digital pathology workflow: the Catania experience. J Pathol Inform. 2017:8. doi: 10.4103/jpi.jpi_58_17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Evans A., Bauer T., Bui M., et al. US Food and Drug Administration approval of whole slide imaging for primary diagnosis: akey milestone is reached and new questions are raised. Arch Pathol Lab Med. 2018;142(11) doi: 10.5858/arpa.2017-0496-CP. [DOI] [PubMed] [Google Scholar]
- 13.Patel A., Balis U., Cheng J., et al. Contemporary whole slide imaging devices and their applications within the modern pathology department: aselected hardware review. J Pathol Inform. 2021:12. doi: 10.4103/jpi.jpi_66_21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Araújo T., Aresta G., Castro E., et al. Classification of breast cancer histology images using convolutional neural networks. PLoS One. 2017;12(6) doi: 10.1371/journal.pone.0177544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Esteva A., Kuprel B., Novoa R.A., et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115–118. doi: 10.1038/nature21056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Frei A., Oberson R., Baumann E., et al. Pathologist computer-aided diagnostic scoring of tumor cell fraction: a Swiss national study. Mod Pathol Off J US Can Acad Pathol, Inc. 2023;36(12) doi: 10.1016/j.modpat.2023.100335. [DOI] [PubMed] [Google Scholar]
- 17.Willis J., Anders R., Torigoe T., et al. Multi-institutional evaluation of pathologists’ assessment compared to immunoscore. Cancers. 2023;15(16) doi: 10.3390/cancers15164045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bankhead P., Loughrey M., Fernández J., et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017;7(1) doi: 10.1038/s41598-017-17204-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rakaee M., Adib E., Ricciuti B., et al. Association of machine learning-based assessment of tumor-infiltrating lymphocytes on standard histologic images with outcomes of immunotherapy in patients with NSCLC. JAMA Oncol. 2023;9(1) doi: 10.1001/jamaoncol.2022.4933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ruifrok A., Katz R., Johnston D. Comparison of quantification of histochemical staining by hue-saturation-intensity (HSI) transformation and color-deconvolution. Appl Immunohistochem Mol Morphol AIMM. 2003;11(1) doi: 10.1097/00129039-200303000-00014. [DOI] [PubMed] [Google Scholar]
- 21.Schmidt U., Weigert M., Broaddus C., Myers G. MICCAI 2018 - 21st International Conference, Granada, Spain, September 16–20, 2018, Proceedings, Part II. SpringerLink; 2018. Cell detection with star-convex polygons. [DOI] [Google Scholar]
- 22.Rakaee M., Kilvaer T., Dalen S., et al. Evaluation of tumor-infiltrating lymphocytes using routine H&E slides predicts patient survival in resected non-small cell lung cancer. Hum Pathol. 2018:79. doi: 10.1016/j.humpath.2018.05.017. [DOI] [PubMed] [Google Scholar]
- 23.Gataa I., Mezquita L., Rossoni C., et al. Tumour-infiltrating lymphocyte density is associated with favourable outcome in patients with advanced non-small cell lung cancer treated with immunotherapy. Eur J Cancer (Oxford, England: 1990) 2021:145. doi: 10.1016/j.ejca.2020.10.017. [DOI] [PubMed] [Google Scholar]
- 24.Dieci M., Radosevic-Robin N., Fineberg S., et al. Update on tumor-infiltrating lymphocytes (TILs) in breast cancer, including recommendations to assess TILs in residual disease after neoadjuvant therapy and in carcinoma in situ: a report of the International Immuno-Oncology Biomarker Working Group on Breast Cancer. Semin Cancer Biol. 2018;52(Pt 2) doi: 10.1016/j.semcancer.2017.10.003. [DOI] [PubMed] [Google Scholar]
- 25.Salgado R., Denkert C., Demaria S., et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann Oncol Off J Eur Soc Med Oncol. 2015;26(2) doi: 10.1093/annonc/mdu450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kos Z., Roblin E., Kim R., et al. Pitfalls in assessing stromal tumor infiltrating lymphocytes (sTILs) in breast cancer. NPJ Breast Cancer. 2020:6. doi: 10.1038/s41523-020-0156-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Wein L., Savas P., Luen S., Virassamy B., Salgado R., Loi S. Clinical validity and utility of tumor-infiltrating lymphocytes in routine clinical practice for breast cancer patients: current and future directions. Front Oncol. 2017:7. doi: 10.3389/fonc.2017.00156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Homeyer A., Geißler C., Schwen L., et al. Recommendations on compiling test datasets for evaluating artificial intelligence solutions in pathology. Mod Pathol Off J US Can Acad Pathol, Inc. 2022;35(12) doi: 10.1038/s41379-022-01147-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Altini N., Marvulli M., Zito F., et al. The role of unpaired image-to-image translation for stain color normalization in colorectal cancer histology classification. Comput Methods Prog Biomed. 2023 doi: 10.1016/j.cmpb.2023.107511. [DOI] [PubMed] [Google Scholar]
- 30.Janowczyk A., Zuo R., Gilmore H., Feldman M., Madabhushi A. HistoQC: an open-source quality control tool for digital pathology slides. JCO Clin Cancer Inform. 2019:3. doi: 10.1200/CCI.18.00157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Jahn S., Plass M., Moinfar F. Digital pathology: advantages, limitations and emerging perspectives. J Clin Med. 2020;9(11) doi: 10.3390/jcm9113697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Smits A., Kummer J., de Bruin P., et al. The estimation of tumor cell percentage for molecular testing by pathologists is not accurate. Mod Pathol Off J US Can Acad Pathol, Inc. 2014;27(2) doi: 10.1038/modpathol.2013.134. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The document outlines methods for color deconvolution, nuclei segmentation with StarDist, and training an object classifier for Tumor, Immune, or Stromal cell categorization. It includes a sharing index for pathologist agreement and statistical tests, ensuring improved accuracy and reproducibility in histological analyses.
Histological characteristics of WSIs in which ROIs were drawn and analyzed for the training set, test set, and comparison with pathologists' estimates.
Metrics for the AI-based classification in the Training Set (right) and Test Set (left). The metrics include precision, recall, and F1 score, which were calculated based on the model's predictions and the ground truth labels. The ground truth labels were determined by a senior pathologist, an expert in LC.
Average rates given by the five pathologists to the graphical prediction carried out by the AI on 37 ROIs. Pathologists assigned a rating between 1 and 5, with 1 indicating a completely incorrect classification, 2 indicating a classification with more errors than correct classifications, 3 indicating a classification that is sufficient or has fewer errors than correct classifications, 4 indicating a good classification but with some negligible errors, and 5 indicating a perfect classification.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available in the github repository DP_BBIRE_ML_CompositionClassifier [https://gitlab.com/bioinfo-ire-release/bbire].