Abstract
Up to now only one major type of core oligosaccharide has been found in the lipopolysaccharide of all Klebsiella pneumoniae strains analyzed. Applying a different screening approach, we identified a novel Klebsiella pneumoniae core (type 2). Both Klebsiella core types share the same inner core and the outer-core-proximal disaccharide, GlcN-(1,4)-GalA, but they differ in the GlcN substituents. In core type 2, the GlcpN residue is substituted at the O-4 position by the disaccharide β-Glcp(1-6)-α-Glcp(1, while in core type 1 the GlcpN residue is substituted at the O-6 position by either the disaccharide α-Hep(1-4)-α-Kdo(2 or a Kdo residue (Kdo is 3-deoxy-d-manno-octulosonic acid). This difference correlates with the presence of a three-gene region in the corresponding core biosynthetic clusters. Engineering of both core types by interchanging this specific region allowed studying the effect on virulence. The replacement of Klebsiella core type 1 in a highly type 2 virulent strain (52145) induces lower virulence than core type 2 in a murine infection model.
Lipopolysaccharide (LPS) is the major inmunodominant antigen in gram-negative bacteria. In the LPS of Enterobacteriaceae, three domains are recognized: the highly conserved and hydrophobic lipid A; the hydrophilic and highly variable O antigen; and the core, connecting lipid A and O antigen. The core domain is usually divided into inner and outer core. The inner core is highly conserved within the Enterobacteriaceae, but the outer core shows an increasingly recognized variability. The difference in variability among the three LPS domains has been attributed to the degree of exposure to the selective pressure of the environment where these cells live. The contribution to Enterobacteriaceae pathogenicity of lipid A and O antigen has been well studied, but so far little is known about the role of the core domain (30).
A possible role in pathogenesis or in adaptation to several body sites is suggested by the prevalence of some core types within clinical isolates. Thus, in Escherichia coli, there are five known core types (K-12, R1, R2, R3, and R4). All are found among commensal isolates (1), whereas the R1 and R3 types predominate among strains causing extraintestinal infections (2, 14) and verotoxigenic strains (8), respectively. Similarly, in Salmonella enterica two core types are known: the Typhimurium type predominates among isolates from human infections, whereas Arizonae IIIA predominates in other Salmonella enterica subspecies (30).
By contrast, no such core variability has been detected in other Enterobacteriaceae, such as Klebsiella pneumoniae (32). In this bacterial species, one major core structure has been found but with variation in the extent of nonstoichiometric substituents (Fig. 1A) (37, 40, 42); and so far, no differences in the core biosynthesis gene cluster (waa) (32) have been described among different K. pneumoniae isolates. However, differences in reactivity with a core-specific monoclonal antibody (38) suggest that there could be more than one major core type among K. pneumoniae isolates. K. pneumoniae is an important cause of nosocomial infections (10) at almost all body sites but with highest incidence in the urinary and respiratory tracts. Thus, it would not be surprising that different K. pneumoniae isolates could have different core LPS types related to preferred body site infection.
FIG. 1.
K. pneumoniae oligosaccharide (OS) core LPS structures. (A) Type 1 core (40, 42). Depending on the K. pneumoniae strain, residues J and K could be H or GalA, and residue P could be H or Hep. Broken lines denote the truncation level for the different core biosynthetic gene mutations (12, 18-20, 32). (B) Type 2 core found in K. pneumoniae 52145 (O1:K2). Oligosaccharides OS1 and OS2 from K. pneumoniae 52145 waaL mutant LPS obtained by complete O and N deacylation under alkaline conditions and mild acid hydrolysis, respectively. Letters denoting OS1 and OS2 residues are the same used in Fig. 4 and 5 (also Tables S2 and S3 in the supplemental material). Kdop, l,d-Hepp, Glcp, GlcNp, and GalAp.
In this work, we characterize genetically and chemically a second K. pneumoniae core LPS. Furthermore, we demonstrate that interchanging the two core types has measurable effects on K. pneumoniae virulence in an animal infection model.
MATERIALS AND METHODS
Bacterial strains, plasmids, and growth conditions.
Bacterial strains and plasmids used in this study are shown in Table 1. All strains were grown in LB broth and LB agar (24). LB media were supplemented with kanamycin (50 μg ml−1), ampicillin (100 μg ml−1), chloramphenicol (20 μg ml−1), and tetracycline (25 μg ml−1) when needed.
TABLE 1.
Bacterial strains and plasmids used
| Strain or plasmid | Relevant characteristics | Reference or source |
|---|---|---|
| K. pneumoniae | ||
| 52145 | Serovar O1:K2 (core type 2) | 25 |
| 52145ΔwaaF | Nonpolar waaF mutant | 19 |
| 52145ΔwaaC | Nonpolar waaC mutant | 19 |
| 52145ΔwabK | Nonpolar wabK mutant | This study |
| 52145ΔwaaL | Nonpolar waaL mutant | 19 |
| 52145ΔwabM | Nonpolar wabM mutant | This study |
| 52145ΔwaaQ (NC19) | Nonpolar waaQ mutant | 32 |
| 52145ΔwabG | Nonpolar wabG mutant | 19 |
| 52145ΔwabH | Nonpolar wabH mutant | This study |
| 52145Δorf10 (NC18) | Nonpolar orf10 mutant | 32 |
| 52145ΔwaaE (NC16) | Nonpolar orf10 mutant | 32 |
| 52145ΔwabK-M C3 | Nonpolar triple gene (wabK waaL wabM) deletion | This study |
| C3 | Serovar O8:K66 (core type 1) | 28 |
| C3ΔwabI-J | Nonpolar triple gene deletion (wabI waaL wabM) | This study |
| E. coli DH5α | F endA hsdR17(rK− mK−) supE44 thi-1 recAI gyr-A96 φ80lacZ | 15 |
| Plasmid | ||
| pKO3 | Cmr temperature sensitive replication sacB containing suicide plasmid | 22 |
| pKO3ΔwabK | Contains wabK deletion | This study |
| pKO3ΔwabM | Contains wabM deletion | This study |
| pKO3ΔwabH | Contains wabH deletion | This study |
| pKO3ΔwabK-M | Contains triple deletion: wabK-waaL52145-wabM | This study |
| pKO3ΔwabI-J | Contains triple deletion: wabI-waaLC3-wabJ | This study |
| pGEM-T easy | PCR generated DNA fragment cloning vector Ampr | Promega |
| pGEM-T-waaFC3 | 32 | |
| pGEM-T-waaCC3 | 32 | |
| pGEM-T-wabK | Amplification primers ORF4.A and ORF4.D | This study |
| pGEM-T-waaL52145 | ORF5.A and ORF5.D | This study |
| pGEM-T-wabM | ORF6.A and ORF6.D | This study |
| pGEM-T-wabI | ORFI.A and ORFI.R | This study |
| pGEM-T-waaLC3 | WaaLOR and WaalOF | This study |
| pGEM-T-wabJ | ORFJ.F and ORFJ.D | This study |
| pGEM-T-waaQC3 | 32 | |
| pGEM-T-wabGC3 | 32 | |
| pGEM-T-wabHC3 | ORF9.A and ORF9.D | This study |
| pGEM-T-orf10C3 | 32 | |
| pGEM-T-waaEC3 | 32 | |
| pGEM-T-wabK-waaL52145-wabM | ORF4.A and ORF6.D | This study |
| pGEM-T-wabI-waaLC3-wabJ | ORFI.A and ORFJ.D | This study |
| pGEM-T-orf4Sm-waaLSm-orf6Sm | KSM and MSM | This study |
General DNA methods.
Standard DNA manipulations were used (34). DNA restriction endonucleases, T4 DNA ligase, E. coli DNA polymerase (Klenow fragment), and alkaline phosphatase were used as recommended by the suppliers. To determine the LPS core biosynthetic gene cluster (waa) nucleotide sequence from K. pneumoniae 52145, specific DNA fragments were amplified by PCR from strain 52145 genomic DNA using primers designed from the known K. pneumoniae C3 sequence (32), followed by nucleotide sequence determination of each amplified fragment by the dideoxy termination method (35).
LPS isolation and electrophoresis.
To isolate LPS from the K. pneumoniae 52145 waaL mutant, the strain was grown in LB. LPS was extracted from dry cells (11.8 g) by the phenol-chloroform/light petroleum ether method (13), with a yield of 2.86% of the bacterial dry mass. For screening purposes, LPS was obtained after proteinase K digestion of whole cells (17). LPS samples were separated by sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis (PAGE) or SDS-Tricine-PAGE and visualized by silver staining as previously described (29, 39).
Preparation of oligosaccharides.
The LPS (50 mg) was hydrolyzed in 1% acetic acid (100°C, 60 min), and the precipitate was removed by centrifugation (8,000 × g, 30 min) and lyophilized to give lipid A (22 mg; 44% of LPS). The supernatant was evaporated to dryness, dissolved in water, and lyophilized (13 mg, 26% of LPS). Another fraction of LPS (200 mg) was first O delipidated by mild hydrazinolysis and then dephosphorylated with HF, reduced with NaBH4, and finally N delipidated by an alkaline hydrolysis with KOH as reported previously (7). The oligosaccharides were purified from salts by gel filtration chromatography on a Sephadex G10 column (Pharmacia), giving after lyophilization a residue of 25 mg.
Glycosyl and lipid analysis.
A sample (1 mg) of LPS was dried over P2O5 overnight and treated with 1 M HCl-CH3OH (1 ml) to obtain fatty acid methylesters and methylglycoside derivatives as reported previously (17).
Determination of absolute configuration of monosaccharides.
To determine the absolute configuration of galacturonic acid, glucosamine, and glucose, a sample of LPS (2 mg) was treated with 1 M HCl-MeOH (500 μl) for 20 h at 80°C. The sample was then hydrolyzed with a mixture of 2 M H2SO4-AcOH and derivatized as acetylated octyl glycosides as described previously (21).
Methylation analysis.
The alditol oligosaccharide mixture was N acetylated by dissolving a sample (2 mg) in dry methanol and treating it with 50 μl of acetic anhydride for 16 h. After evaporation of the solvents, the sample was methylated as reported previously (4). Linkage analysis was performed as follows: the methylated sample was reduced and carboxymethylated with lithium triethylborodeuteride (Super-Deuteride, Aldrich), subjected to mild acid hydrolysis to cleave ketosidic linkage, reduced again with NaBD4, and then totally hydrolyzed, reduced with NaBD4, and finally acetylated as described previously (11).
Gas chromatography-MS analysis.
An Agilent Technologies 5973N mass spectrometry (MS) instrument equipped with a 6850A gas chromatography and an RTX-5 capillary column (Restek, 30 m by 0.25 inside diameter; flow rate, 1 ml min−1; He as carrier gas) was used for analysis of alditol acetates and methyl glycoside acetates as previously described (17). Analysis of acetylated octyl glycosides was performed as follows: 150°C for 5 min, 150°C→240°C at 6°C min−1, 240°C for 5 min.
Nuclear magnetic resonance (NMR) spectroscopy.
Spectra were recorded on a Varian Inova 500 spectrometer equipped with a reverse probe at 25°C. 13C and 1H chemical shifts were measured in D2O using acetone, at δ 2.225 and 31.4 for proton and carbon, respectively, as an internal standard. Two-dimensional homo- and heteronuclear experiments (correlation spectroscopy [COSY], heteronuclear multiple bond correlation [HMBC], heteronuclear single quantum coherence [HSQC], HSQC-distortionless enhancement by polarization transfer spectroscopy [DEPT], HSQC-total correlation spectroscopy [TOCSY], rotating-frame nuclear Overhauser enhancement spectroscopy [ROESY], and TOCSY) were performed using standard pulse sequences available in the Varian software.
Mass spectrometry studies.
The matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrum was obtained on the Applied Biosystems 4700 proteomics analyzer with TOF/TOF optics. Calibration was done in positive mode on a peptide (Glu-fibrinopeptide) and used also in negative mode. The energy of laser was set to 4,900 in MS mode. The matrix was 2,5-dihydroxybenzoic acid at 25 mg m−1 in 20% acetonitrile in water. Spectra were acquired in negative mode.
K. pneumoniae mutant construction.
To obtain K. pneumoniae mutant strains with defects in core LPS, chromosomal in-frame nonpolar waa deletions were made (22). Sets of four primers (see Table S1 in the supplemental material), designed from the K. pneumoniae 52145 waa gene cluster sequence, were used in PCR DNA fragment amplifications to create the waa gene deletions. These constructs were subcloned in a suicide plasmid vector, pKO3 (22). The suicide plasmid derivatives obtained were used to introduce each mutation into wild-type K. pneumoniae strains by double recombination, as previously described (22).
Mutant complementation studies.
To complement the constructed mutants, each of the waa genes from K. pneumoniae strains C3 and 52145 were PCR amplified using pairs of specific primers (see Table S1 in the supplemental material). The amplified fragments were ligated to the vector pGEM-T. The recombinant plasmids were introduced into the mutants by electroporation, and the core LPS characteristics were studied.
Fifty-percent lethal dose (LD50).
Albino Swiss female mice (5 to 7 weeks old; Harlan Ibérica, S.L.) were injected intraperitoneally with 0.2 ml of the test samples (from 108 to 103 CFU ml−1). Mortality was recorded up to 7 days.
Dot blot hybridization.
The two DNA probes A and B used consisted of the digoxigenin-labeled PCR amplification products of strains C3 and 52145 produced with the DIG DNA labeling and detection kit (Boehringer, Mannheim, Germany). The 2,264-bp probe A, containing the strain C3 genes wabI, waaL, and wabJ, was obtained using primers WaaLOR and WaaLOF. The 2,267-bp probe B, containing the strain 52145 genes wabK, waaL, and wabM, was obtained by using primers ORF5.A and ORF5.D (see Table S1 in the supplemental material for primer sequence). Cell lysates were obtained by resuspending the cells of a 500-μl overnight culture in LB in 100 μl of 0.4 M NaOH and heating it for 30 min at 80°C. One microliter of cell lysate was blotted on a Hybond membrane and bound by UV cross-linking. Hybridization was done following standard protocols (34) with stringent washing at 65°C in 0.2× SSC (20× SSC is 3 M NaCl plus 0.3 M sodium citrate, pH 7.0). Probe that remained bound to homologous sequences was detected with the DIG DNA labeling and detection kit following the supplier's instructions.
RESULTS
Evidence for a second major core LPS type in K. pneumoniae.
Differences in reactivity with a core-specific monoclonal antibody among K. pneumoniae isolates (38) suggested the existence of different cores. A previous approach based on PCR amplification of genomic DNA fragments using K. pneumoniae C3-specific primers failed to detect any variability in the core LPS biosynthetic gene cluster (waa) from different K. pneumoniae strains (32). This failure could be due to the use of primers matching conserved regions shared by different putative K. pneumoniae waa gene clusters. Bacteriophage FC3-10 infects K. pneumoniae C3 and other strains using the core LPS as a receptor (5). Some naturally resistant strains became sensitive owing to mutations that interfere with O- and/or K-antigen biosynthesis, suggesting that these antigens might hinder the interaction between the phage and the core. By contrast, several K. pneumoniae strains, such as 52145, remained FC3-10 resistant after the introduction of these mutations. The strain C3 core LPS is of the K. pneumoniae core type 1. Thus, we reasoned that phage FC3-10-resistant K. pneumoniae strains that did not revert to phage sensitivity upon mutations precluding O- and/or K-antigen production might possess a different LPS core type.
A collection of K. pneumoniae strains from our laboratory and their O−, K−, and O− K− derivatives were screened for phage FC3-10 resistance. The LPS from K. pneumoniae FC3-10-resistant strains was extracted and compared to LPS from K. pneumoniae-sensitive strains that produce the classical core LPS by SDS-Tricine-PAGE. Some of these strains, such as 52145, showed LPS molecules with a slightly different pattern of gel migration. These differences in gel migration suggested differences in core LPS structure.
To test this hypothesis, strain 52145 was chosen for further study. Genomic DNA from strain 52145 was isolated and used in PCR amplification experiments using sets of primer pairs designed from the K. pneumoniae C3 waa gene cluster sequence. All but one of the primer pairs used allowed the amplification of DNA fragments of the same size as those of the C3 gene cluster. The primer pair WaaLOR-WaaLOF failed to generate DNA amplification using 52145 genomic DNA as template. This primer pair amplifies a DNA region containing the O-antigen ligase gene waaL and its two flanking genes when using strain C3 genomic DNA as a template, suggesting that there could be significant differences between the waa gene clusters of strains 52145 and C3.
K. pneumoniae 52145 waa gene cluster analysis.
The nucleotide sequence of the whole waa gene cluster from strain 52145 was determined in two steps. First the nucleotide sequence of the successfully amplified DNA fragments from the strain 52145 genome, using primer pairs S7-Nih5.A5, Nuk1-Kor, Nuk5-Nuk8, and Nuk7-Nuk10 (Fig. 2), was determined. Second, sequence-derived primers were used for further PCR amplification and sequence determination to close the sequenced gaps. Analysis of the data obtained revealed homologues of the reported strain C3 genes encoding l,d-heptose epimerase (hldD), heptosyltransferases I, II, and III (waaC, waaF, and waaQ), 3-deoxy-d-manno-oct+ulosonic acid (Kdo) transferase (waaA) (32), inner-core glucosyltransferase (waaE) (18, 20), galacturonyltransferase (wabG) (19), N-acetyl-glucosaminyltransferase (wabH) (12), and a gene of unknown function (orf10) similar to E. coli yibD (32) (Fig. 2). Although a putative waaL gene was also found, the 416-residue ORF552145 (WaaL) showed significant similarity to the putative O-antigen ligase (WaaL) of Serratia marcescens (6) among other Enterobacteriaceae but limited identity (19%) and similarity (38%) to the WaaL protein from strain C3.
FIG. 2.
Diagram of the K. pneumoniae 52145 waa gene cluster and comparison of this cluster to those of K. pneumoniae C3 (32) and S. marcescens N28b (6). White boxes denote PCR-amplified DNA fragments. Common inner core genes (black arrows), common outer-core genes (gray arrows). Common outer-core genes between K. pneumoniae 52145 and S. marcescens N28b (striped arrows). Cluster-specific genes (white arrows). K. pneumoniae 52145 waa gene cluster, GenBank accession no. AY823268.
By contrast, no gene homologues to K. pneumoniae C3 wabI and wabJ were detected in strain 52145, but two new genes were found (Fig. 2). The 390-residue ORF452145 (named WabK) showed significant levels of similarity (72%) and identity (55%) to a putative glucosyltransferase from the Serratia marcescens N28b waa gene cluster (ORF4Sm) (6). Similarly, the 330-residue ORF652145 (named WabM) showed again significant levels of similarity (64%) and identity (41%) to another putative glucosyltransferase from S. marcescens N28b (ORF6Sm) (6).
In both strains C3 and 52145, the gene pairs wabI-wabJ and wabK-wabM were found flanking the corresponding O-antigen ligase (waaLC3 and waaL52145) gene. The gene order and transcription direction were the same for these three genes in K. pneumoniae 52145 and S. marcescens N28b, while in the waaC3 gene cluster the wabJ gene was transcribed in the opposite direction (32) (Fig. 2).
Mutational and complementation studies.
To properly identify the additional genes from the waa52145 gene cluster, new nonpolar in-frame deletion mutants were constructed (see Materials and Methods). LPS from these mutants was analyzed by SDS-Tricine-PAGE (Fig. 3), revealing a gradation of electrophoresis migration patterns, in agreement with the postulated functions for the genes common to both K. pneumoniae waa clusters (Fig. 1A). In addition, each mutant was complemented by the corresponding K. pneumoniae C3 homologous genes (data not shown), except O-antigen ligase waaLC3 and waaL52145. The O− 52145ΔwaaL mutant was only partially complemented by waaLC3, while full restoration of O-antigen production was achieved by complementation with waaL52145. Since the O antigen is linked to the outer-core residue Kdo in K. pneumoniae strains containing type 1 core LPS (41), our results suggest differences in the O-antigen outer-core linkage region between strains C3 and 52145.
FIG. 3.
SDS-Tricine-PAGE analysis of LPS samples from K. pneumoniae 52145- and 52145-derived nonpolar core biosynthetic mutants.
As expected, mutants 52145ΔwabK and 52145ΔwabM were not complemented by any of the waaC3 genes, suggesting the absence of wabI (encoding the outer-core Kdo transferase) (12) and wabJ (encoding the outer-core heptosyltranferase) (12) in the waa52145 cluster. Thus, it seems likely that K. pneumoniae 52145 core LPS differs from the previously reported type 1 core in the distal outer-core residues Kdo and Hep.
Chemical characterization of the K. pneumoniae 52145 core LPS.
The LPS from the waaL52145 mutant was extracted by the PCP method. Sugar analysis of the oligosaccharide (OS) fractions revealed the presence of d-GalA, d-GlcN, d-Glc, l,d-Hep, and Kdo. Methylation analysis indicated the attachment points of each residue: GalA was present both as terminal and 4-linked, GlcN as 4-linked and 6-linked, l,d-Hep was present as 7-linked, 3,4-, and 3,7-linked, Glc as terminally and 6-linked, and Kdo as both terminally and 4,5-linked.
The core OS was obtained by degradation of LPS under both the alkaline (OS1) and acid conditions (OS2) (see Materials and Methods). The 1H-NMR spectra showed seven (A to G) (Fig. 4) and nine (H to R) (Fig. 5A) anomeric signals for OS1 and OS2, respectively. The complete assignment of all NMR signals from OS1 was achieved by two-dimensional homonuclear and heteronuclear experiments (COSY, TOCSY, HSQC-DEPT, HSQC-TOCSY, HMBC and ROESY), allowing us to show that in addition to the seven anomeric signals (A to G) occurring in the 1H spectrum (Fig. 1B and Fig. 4) (see Table S2 in the supplemental material), a further signal at 5.75 ppm (A4) occurred, which was correlated to the unsaturated C-4 of a threo-hex-4-enuronopyranosyl unit. This finding suggested that position 4 of galactopyranuronic acid is probably substituted by a sugar chain which was lost during the hot alkaline treatment by β-degradation during the alkaline hydrolysis of LPS (see Materials and Methods).
FIG. 4.
Anomeric region of the 1H-NMR spectra of OS1 obtained from K. pneumoniae 52145 waaL mutant LPS by complete O and N deacylation. Letters denote residues as in Fig. 1B OS1.
FIG. 5.
(A) Anomeric region of the 1H-NMR spectra and (B) ROESY spectra of OS2 obtained from K. pneumoniae 52145 waaL mutant LPS by mild acid hydrolysis. Letters denote residues as in Fig. 1B OS2.
The identification of the monosaccharide in OS1 and OS2 was obtained by proton and carbon chemical shift values (see Tables S2 and S3 in the supplemental materials), supported by the proton vicinal coupling constant values of the identified multiplet patterns. The anomeric configurations were assigned on the basis of observed 3JH,H values of proton anomeric signals and for Kdo residues in OS1 by the chemical shift values of the methylene protons at C-3. Residues present in both OS1 and -2 showed very similar chemical shift values (Fig. 4 and 5a) (see Tables S2 and S3 in supplemental materials).
The sequence of residues in OS1 (Fig. 1B) was deduced from the scalar interresidual correlations measured by a long-range 1H,13C-HMBC. A ROESY experiment confirmed this sequence by interresidual dipolar coupling and showed the spatial proximity of the C unit with the 4,5-linked Kdo residue, for which the HMBC did not show any diagnostic correlation. The sequence of residues in OS2 was deduced by interresidue nuclear Overhauser enhancement spectroscopy (NOEs) and confirmed by the long-range 13C-1H-HMBC, which also supported the attachment points of interglycosidic linkages suggested by the low-field chemical shift of the glycosylated carbons (see Table S3 in the supplemental material). In particular, the HMBC spectrum showed the following correlations: H-1 H/C-4 M, H-1 M/C4 I, H-1 I/C-3 L, H-1 L/C-3 N, H-1 P/C-4 N, H-1 R/C-7 O, and H-1 O/C-7 L, and the ROESY spectrum (Fig. 5B) showed the NOE contacts between H-1 of Q and the methylene protons of H, indicating the spatial proximity of these residues, for which the HMBC spectrum did not show any correlation. These experiments allowed to deduce the structure of OS1 and OS2 (Fig. 1B).
Mass spectroscopy studies.
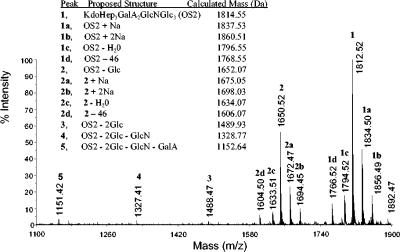
To determine the degree of heterogeneity in the core OS fraction, a negative-mode MALDI-TOF mass spectrum was obtained (Fig. 6). The spectrum showed a complex pattern of mass peaks, some containing one or two sodium ions. The major signal at m/z 1812.52 was in close agreement with a structure consisting of Glc3GlcN GalA2Hep3Kdo (calculated mass, 1814.55 Da). Satellite peaks at m/z 1794.52 and m/z 1766.52 correspond to the anhydro form (M-18) and a Kdo artifact (M-46), respectively (27). Thus, the core OS2 is the most abundant in K. pneumoniae C3 core LPS. Additional peaks (Fig. 6) could be attributed to molecular species lacking outer-core residues Glc, one or two, GlcN, and GalA. A low intense peak was found at m/z 1892.47, probably indicating the presence of a phosphate group in the core oligosaccharide.
FIG. 6.
Positive ion MALDI-TOF spectrum of acid-released core LPS oligosaccharides from K. pneumoniae 52145Δ waaL.
To confirm the sequence of residues of the deduced major core OS2, a MALDI-TOF/TOF experiment was performed. This technique has been reported to give higher fragmentation than other collision-induced dissociation techniques, giving both inter- and intraring fragments (23, 36). The MALDI-TOF/TOF mass spectrum of the (M+H)+ ion of OS2 at m/z 1815.37 (Fig. 7) contained a signal at m/z 486.17 corresponding to the sequence Glc-Glc-GlcN and signals at m/z 324.14 and 162.09 generated by its internal fragmentation. The signal at m/z 662.20 has been attributed to the B ion of the sequence Glc-Glc-GlcN-GalA, while the peaks at m/z 1222.32 and 1577.42 are diagnostic of the two branching points in OS2. The high intensity of the signals at m/z 162.09 and 486.17 can be due to the presence of HexN in these fragments. Moreover, it is not surprising that the peaks of the fragments corresponding to the branch points (1222.32 and 1577.42) are more intense than others (662.21 and 854.29).
FIG. 7.
Positive ion MALDI-TOF/TOF spectrum of m/z 1814.55 signal of acid-released core LPS oligosaccharide isolated from K. pneumoniae 52145ΔwaaL, in the positive ion mode. Insert shows the fragmentation pattern.
The tandem mass spectrum of the pseudomolecular ion (M+H)+ at m/z 1652.58 (Fig. 8) confirms the sequence and the branching of the deduced core OS2. The signal at m/z 1060.27 (Fig. 8), which corresponded to the ion at m/z 1222.32 in Fig. 7, clearly indicated that this OS has lost a terminal Glc. Moreover, the relative intensity of the signal at m/z 324.13, which is more intense than the same signal in Fig. 7, suggested that the cleavage is near the GlcN residue.
FIG. 8.
Positive ion MALDI-TOF/TOF spectrum of m/z 1652.07 signal of acid-released core LPS oligosaccharide isolated from K. pneumoniae 52145ΔwaaL, in the positive ion mode. Insert shows the fragmentation pattern.
Klebsiella pneumoniae type 2 outer-core glucosyltransferases.
The above results show that K. pneumoniae 52145 core LPS is a second core type in this species (type 2 core) and differs from the previously reported type 1 core by the presence of a terminal β-Glcp(1-6)-α-Glcp instead of Kdo(2) or α-Hep(1-4)-α-Kdo(2). Since both WabK and WabM show glycosyltransferase features, they are candidates implicated in the transfer of the outer-core Glc residues. If this hypothesis is true, the core oligosaccharides from wabK and wabM mutants should be shorter than that of wild type strain 52145 by either one or two Glc residues. As expected, major signals at m/z 1650.52 and 1488.47 (data not shown) were detected in the OS fractions from strains 52415ΔwabM and 52415ΔwabK, respectively. These two signals are in agreement with structures containing Glc2GlcNGalA2Hep3Kdo (calculated mass, 1,652.07 Da) and GlcGlcNGalA2Hep3Kdo (calculated mass, 1,489.93 Da). These data suggest that the WabM and WabK proteins are glucosyltransferases involved in the successive transfer of a Glc residue to the O-6 position of Glc and to the O-4 position of GlcN, respectively. Finally, it should be noted that the LPS from both 52415ΔwabM and 52415ΔwabK mutants are devoid of O1 antigen as judged by SDS-Tricine-PAGE analysis.
Engineering K. pneumoniae core LPS.
We decided to construct K. pneumoniae strains expressing heterologous core LPS to study the effects of core LPS type on its expression and virulence. Nonpolar triple deletion ΔwabI waaLC3 wabJ and ΔwabM waaL52145 wabK mutants were constructed from K. pneumoniae strains C3 and 52145, respectively (see Table S1 in the supplemental material for primers used in these constructions). As expected, LPS from mutants C3ΔwabI-J and 52145ΔwabM-K migrated faster than the corresponding wild-type LPS and were devoid of O antigen (Fig. 9). The three type 1 core-specific waa genes on pGEM-T-wabI-waaLC3-wabJ were transformed into 52145ΔwabM-K, resulting in full O1-antigen-containing LPS (Fig. 9). Similarly, C3ΔwabI-J cells transformed with pGEM-T-wabM-waaL52145-wabK showed full O8-antigen-containing LPS. Thus, the complemented triple mutants produce their parental O-antigen LPS independently of the complementing gene's origin (Fig. 9, data shown for 52145 and derivative strains). In addition, the engineered strains 52145ΔwabM-K(pGEM-T-wabI-waaLC3-wabJ) and C3ΔwabI-J(pGEM-T-wabM-waaL52145-wabK) produced the original wild-type K2 and K66 antigen, respectively. Thus, changing the core type among strains C3 and 52145 does not modify the nature of the parental O and K antigens.
FIG. 9.
SDS-PAGE (A) and SDS-Tricine-PAGE (B) analysis of LPS samples from K. pneumoniae type 2 core strains 52145 (O1:K2) and B5055 (O1:K2), 52145-derived triple core biosynthesis mutants, homologous and heterologous complemented mutants, and type 1 core strain C3 (O8:K66).
The effect on virulence caused by the interchange of core types was investigated using an experimental murine infection model. The 52145ΔwabM-K mutant showed a three-log increase in LD50 compared to wild type 52145 (Table 2). The 52145ΔwabM-K(pGEM-T-wabM-waaL52145-wabK) cells showed wild-type levels of virulence as measured by LD50. By contrast, introduction of the heterologous C3 genes in strain 52145ΔwabM-K by pGEM-T-wabI-waaLC3-wabJ transformation resulted in a two-log-higher LD50 value than that for wild-type 52145 (Table 2). In addition, introduction of the S. marcescens orf4Sm-waaLSm-orf6Sm genes into mutant 52145ΔwabK-M restored O-antigen production and wild-type LD50 values in the mouse assay (Table 2). This last result was expected because of the similarity shown between wabK and wabM with orf4Sm and orf6Sm, respectively. This similarity is in agreement with the presence of the outer-core disaccharide β-Glcp(1-6)-α-Glcp(1 in both K. pneumoniae 52145 and S. marcescens N28b. The results obtained in the LD50 experiments are not attributable to differences in O-antigen LPS amounts as judged by LPS gels (Fig. 9). Furthermore, enzyme immunoassay experiments using whole cells with specific monoclonal O1-antigen LPS antibody 2A4 (9) and polyclonal O8-antigen LPS serum showed no significant differences in the respective O-antigen LPS amounts between the wild type and complemented triple-mutant strains. Since wild-type strain 52145 and its derivative 52145ΔwabM-K(pGEM-T-wabI-waaLC3-wabJ) differ only in core type, it appears that the core nature is responsible for the differences in bacterial virulence, at least in the infection model used here.
TABLE 2.
LD50 for mice inoculated intraperitoneally with different K. pneumoniae strains
| Strain | LD50 CFU ml−1a |
|---|---|
| K. pneumoniae 52145 | 102.1 |
| 52145ΔwaaL | 105.1 |
| 52145ΔwabK | 105.5 |
| 52145ΔwabM | 105.2 |
| 52145ΔwabK-M | 105.5 |
| 52145ΔwabK-M + pGEM-T | 105.6 |
| 52145ΔwabK-M + pGEM-T-wabI-waaLC3-wabJ | 104.2 |
| 52145ΔwabK-M + pGEM-T-wabK-waaL52145-wabM | 102.2 |
| 52145wabK-M + pGEM-T-orf4Sm-waaLSm-orf6Sm | 102.3 |
LD50 were calculated as described previously (31). LD50 values are the means from three independent experiments. Maximum standard deviation, <100.5.
Distribution among Klebsiella strains of the type 1 and 2 core-specific genetic determinants.
The knowledge of both type 1 and 2 core waa gene clusters allowed the design of specific digoxigenin-labeled DNA probes (see Materials and Methods). The 2,264-bp type 1 and 2,994-bp type 2 core DNA probes encompassed the wabK waaL52145 wabM and wabI waaLC3 wabJ regions, respectively. The two probes were used in a dot blot assay to screen genomic DNAs from Klebsiella strains, including all the different O-antigen serotypes. One hundred Klebsiella strains were assayed (O1 [n = 34], O2 [n = 16], O2ac [n = 6], O3 [n = 18], O4 [n = 4], O5 [n = 13], O8 [n = 6], and O12 [n = 3] [16]) to establish their prevalence. None of the Klebsiella genomic DNAs reacted with both probes simultaneously, and only two strains failed to react with any probe. The type 1 core probe reacted with 79 Klebsiella strains (79%), while the type 2 core probe reacted with 19 strains (19%). It is important to point out that the percentage of Klebsiella strains with type 2 core-specific genes in strains with O1-antigen LPS increases to 29% (10 of 34 strains). The percentage of strains with type 2 core-specific genes was highest (83%; five of six strains) for serotype O1:K2 strains, besides their different geographical origin. Four of the five K. pneumoniae O1:K2 strains with type 2 core are highly virulent for mice.
DISCUSSION
We have identified in this study a second core type in K. pneumoniae 52145 that we have designated type 2 core. The elucidation of the chemical structure of type 2 core and the comparison of this structure with that of type 1 core revealed that both core types differ only in the outer-core region. The GlcpN residue is substituted at the O-6 position by either the disaccharide α-Hep(1-4)-α-Kdo(2 or a Kdo residue in the previously described K. pneumoniae type 1 core (Fig. 1A). By contrast, the GlcpN residue is substituted at the O-4 position by the disaccharide β-Glcp(1-6)-α-Glcp(1 in K. pneumoniae type 2 core (Fig. 1B). An identical disaccharide moiety has already been found in Serratia marcescens, indicating a close similarity of the biosynthetic pathways for S. marcescens and K. pneumoniae type 2 core LPS.
Comparison of the gene clusters encoding the core biosynthetic genes among strains producing type 1 (C3) and type 2 (52145) cores clearly showed that only two genes account for the differences between both core types. Strains containing type 1 core present the genes wabI (outer-core Kdo transferase) and wabJ (outer-core heptosyltransferase), while strains with type 2 core have the genes wabK and wabM. Evidence suggesting that the genes wabK and wabM encode glucosyltransferases involved in the transfer of the last two outer-core Glc residues of type 2 core is based on analysis of LPS from mutants 52415ΔwabK and -M by SDS-Tricine-PAGE and mass spectrometry. In agreement with the outer-core structural similarity between K. pneumoniae 52145 and S. marcescens N28b, wabK and wabM homologues were found in S. marcescens. These outer-core-specific genes are always found flanking the corresponding O-antigen ligase gene (waaL).
Previously it has been shown that in K. pneumoniae strains containing type 1 core, the O antigen is linked to the outer-core residue Kdo (41). K. pneumoniae 52145 mutants containing deletions in either wabK or wabM are devoid of O antigen, suggesting that in this strain, the O antigen is linked to the last Glcp outer-core residue. Interestingly, the similarity between the deduced putative O-antigen ligase proteins is quite low, and no full complementation was found between waaLC3 and waaL52145, suggesting some degree of specificity of the O-antigen ligase for each core type.
An analysis of the G+C percentage of each individual gene (Table 3) reveals an unusually low value for the contiguous genes wabK, waaL52145, and wabM, suggesting that they could have been acquired relatively recently by horizontal transfer, leading to the appearance of type 2 core in K. pneumoniae. In addition, this analysis suggests that this genetic information was not acquired from S. marcescens, because of the difference in G+C percentage between the two equivalent regions (Table 3).
TABLE 3.
Percent C+G in genes involved in the core LPS biosynthesis from K. pneumoniae strains 52145 and C3 and S. marcescens N28b
| Gene | % C+G in gene (designation) from:
|
||
|---|---|---|---|
| K. pneumoniae 52145 | K. pneumoniae C3 | S. marcescens N28b | |
| waa gene cluster | 55.80 | 57.80 | 59.23 |
| hldD | 55.20 | 55.60 | 56.65 |
| waaF | 61.19 | 61.11 | 63.90 |
| waaC | 62.82 | 64.12 | 62.32 |
| orf4 | 45.52 (wabK) | 52.70 (wabI) | 59.55 |
| waaL | 44.36 | 60.96 | 57.25 |
| orf6 | 46.42 (wabM) | 52.93 ca (wabJ) | 56.74 |
| orf7Sm | 57.44 c | ||
| waaQ | 60.72 | 60.93 | 62.14 |
| wabG | 61.97 | 62.96 | 61.88 |
| wabH | 61.37 | 61.39 | 59.58 |
| orf10Kp | 54.95 | 55.26 | |
| orf11Sm | 55.42 c | ||
| waaA | 60.98 | 59.10 | 61.35 |
| waaE | 59.72 | 59.74 | 61.5 |
| coaD | 55.91 | 56.27 | 60.29 |
c, open reading frame encoded by complementary-strand DNA.
The localization on a unique and small DNA region of the genes responsible for either type 1 or 2 core in K. pneumoniae allowed us to test for the first time the direct effect of core exchange in virulence. The results obtained (Table 2) strongly suggest that type 2 core enhances the killing ability of this species in the mouse model used, because replacing type 2 by type 1 core results in a two-log increase in LD50. The difference in virulence could be directly related to the core LPS structure or could be attributed to its effect on proper localization and/or folding of outer membrane components playing a role in virulence.
The type 2 core frequency among Klebsiella isolates was assayed by analysis of the presence of the type 1 and 2 core-specific genes. Preliminary results obtained with 100 Klebsiella strains from our laboratory suggest that they represent 19% of our Klebsiella collection, with a highest prevalence in serotype O1 strains (29%) and serotype O1:K2 strains (83%). Four of the five Klebsiella O1:K2 strains with type 2 core tested are highly virulent for mice. Furthermore, the presence of two Klebsiella strains that cannot be assigned to either type 1 or type 2 cores suggests the existence of additional Klebsiella core types. Since K. pneumoniae is an important cause of nosocomial infections ranging from urinary tract infections to septicemia and pneumonia (10) and can also produce primary liver infection in both healthy and unhealthy persons (3, 26, 33), future studies will be needed to determine if each core type is associated with groups of strains producing infections with specific clinical manifestations.
In conclusion, a second core LPS type from K. pneumoniae, probably arising by horizontal transfer, has been characterized both genetically and structurally. Genetic exchange of type 1 and 2 cores suggests that they are not equivalent in a mouse infection model.
Supplementary Material
Acknowledgments
This work was supported by Plan Nacional de I + D grants (Ministerio de Ciencia y Tecnología, Spain) and from Generalitat de Catalunya. L.I. and S.F. are recipients of predoctoral fellowships from Ministerio de Ciencia y Tecnología (Spain) and Universidad de Barcelona.
We also thank Maite Polo for her technical assistance.
Footnotes
Supplemental material for this article may be found at http://jb.asm.org/.
REFERENCES
- 1.Amor, K., D. E. Heinrichs, E. Frirdich, K. Ziebell, R. P. Johnson, and C. Whitfield. 2000. Distribution of core oligosaccharide types in lipopolysaccharides from Escherichia coli. Infect. Immun. 68:1116-1124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Appelmelk, B. J., Y. Q. An, T. A. Hekker, L. G. Thijs, D. M. MacLaren, and J. de Graaf. 1994. Frequencies of lipopolysaccharide core types in Escherichia coli strains from bacteraemic patients. Microbiology 140:1119-1124. [DOI] [PubMed] [Google Scholar]
- 3.Cahill, M., B. Chang, and A. Murray. 2000. Bilateral endogenous bacterial endophthalmitis associated with pyogenic hepatic abscess. Brit. J. Ophthalmol. 84:1436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ciucanu, I., and F. Kerek. 1984. A simple and rapid method for the permethylation of carbohydrates. Carbohydr. Res. 131:209-217. [Google Scholar]
- 5.Ciurana, B. 1990. Estudi de la membrana externa de Klebsiella pneumoniae en relació amb la patogenicitat. Ph.D. thesis. University of Barcelona, Barcelona, Spain.
- 6.Coderch, N., N. Piqué, B. Lindner, N. Abitiu, S. Merino, L. Izquierdo, N. Jimenez, J. M. Tomás, O. Holst, and M. Regué. 2004. Genetic and structural characterization of the core region of the lipopolysaccharide from Serratia marcescens N28b (serovar O4). J. Bacteriol. 186:978-988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Corsaro, M. M., R. Lanzetta, E. Parrilli, M. Parrilli, and M. L. Tutino. 2001. Structural investigation on the lipooligosaccharide fraction of psychrophilic Pseudoalteromonas haloplanktis TAC 125 bacterium. Eur. J. Biochem. 268:5092-5097. [DOI] [PubMed] [Google Scholar]
- 8.Currie, C. G., and I. R. Poxton. 1999. The lipopolysaccharide core type of Escherichia coli O157:H7 and other non-O157 verotoxin-producing E. coli. FEMS Immunol. Med. Microbiol. 24:57-62. [DOI] [PubMed] [Google Scholar]
- 9.Domenico, P., J. M. Tomas, S. Merino, X. Rubires, and B. A. Cunha. 1999. Surface antigen exposure by bismuth dimercaprol suppression of Klebsiella pneumoniae capsular polysaccharide. Infect. Immun. 67:664-669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Emori, T. G., and R. P. Gaynes. 1993. An overview of nosocomial infections, including the role of the microbiology laboratory. Clin. Microbiol. Rev. 6:428-442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Forsberg, L. S., U. Ramadas Bhat, and R. W. Carlson. 2000. Structural characterization of the O-antigenic polysaccharide of the lipopolysaccharide from Rhizobium etli strain CE3. J. Biol. Chem. 275:18851-18863. [DOI] [PubMed] [Google Scholar]
- 12.Frirdich, E., E. Vinogradov, and C. Whitfield. 2004. Biosynthesis of a novel 3-deoxy-D-manno-oct-ulosonic acid-containing outer core oligosaccharide in the lipopolysaccharide of Klebsiella pneumoniae. J. Biol. Chem. 279:27928-27940. [DOI] [PubMed] [Google Scholar]
- 13.Galanos, C., O. Lüderitz, and O. Westphal. 1969. A new method for the extraction of R lipopolysaccharides. Eur. J. Biochem. 9:245-249. [DOI] [PubMed] [Google Scholar]
- 14.Gibb, A. R., G. R. Barclay, I. R. Poxton, and F. Di Padova. 1992. Frequencies of lipopolysaccharide core types among clinical isolates of Escherichia coli defined with monoclonal antibodies. J. Infect. Dis. 166:1051-1057. [DOI] [PubMed] [Google Scholar]
- 15.Hanahan, D. 1983. Studies on transformation of Escherichia coli with plasmids. J. Mol. Biol. 166:557-580. [DOI] [PubMed] [Google Scholar]
- 16.Hansen, D. S., F. Mestre, S. Albertí, S. Hernández-Alles, D. Alvarez, A. Domenech-Sánchez, J. Gil, S. Merino, J. M. Tomás, and V. J. Benedí. 1999. Klebsiella pneumoniae lipopolysaccharide O typing: revision of prototype strains and O-group distribution among clinical isolates from different sources and countries. J. Clin. Microbiol. 37:56-62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hitchcock, P. J., and T. M. Brown. 1983. Morphological heterogeneity among Salmonella lipopolysaccharide chemotypes in silver-stained polyacrylamide gels. J. Bacteriol. 154:269-277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Izquierdo, L., N. Abitiu, N. Coderch, B. Hita, S. Merino, R. Gavín, J. M. Tomás, and M. Regué. 2002. The inner-core lipopolysaccharide biosynthetic waaE gene: function and genetic distribution among some Enterobacteriaceae. Microbiology 148:3485-3496. [DOI] [PubMed] [Google Scholar]
- 19.Izquierdo, L., N. Coderch, N. Piqué, E. Bedini, M. Corsaro, S. Merino, S. Fresno, J. M. Tomas, and M. Regué. 2003. The Klebsiella pneumoniae wabG gene: its role in the biosynthesis of the core lipopolysaccharide and virulence. J. Bacteriol. 185:7213-7221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Izquierdo, L., S. Merino, N. Coderch, M. Regué, and J. M. Tomás. 2002. The wavB gene of Vibrio cholerae and the waaE of Klebsiella pneumoniae codify for a beta-1,4-glucosyltransferase involved in the transfer of a glucose residue to the L-glycero-D-manno-heptose I in the lipopolysaccharide inner core. FEMS Microbiol. Lett. 216:211-216. [DOI] [PubMed] [Google Scholar]
- 21.Leontein, K., B. Lindberg, and J. Lönngren. 1978. Assignment of absolute configuration of sugars by g.l.c. of their acetylated glycosides from chiral alchols. Carbohydr. Res. 62:359-362. [Google Scholar]
- 22.Link, A. J., D. Phillips, and G. M. Church. 1997. Methods for generating precise deletions and insertions in the genome of wild-type Escherichia coli: application to open reading frame characterization. J. Bacteriol. 179:6228-6237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Mechref, Y., M. V. Novotny, and C. Krishnan. 2003. Structural characterization of oligosaccharides using MALDI-TOF/TOF tandem mass spectrometry. Anal. Chem. 75:4895-4903. [DOI] [PubMed] [Google Scholar]
- 24.Miller, J. H. 1972. Experiments in molecular genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 25.Nassif, X., J. M. Fournier, J. Arondel, and P. J. Sansonetti. 1989. Mucoid phenotype of Klebsiella pneumoniae is a plasmid-encoded virulence factor. Infect. Immun. 57:546-552. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ohmori, S., K. Shiraki, K. Ito, H. Inou, T. Ito, T. Sakai, K. Takase, and T. Nakano. 2002. Septic endophthalmitis and meningitis associated with Klebsiella pneumoniae liver abscess. Hepatol. Res. 22:307-312. [DOI] [PubMed] [Google Scholar]
- 27.Olsthoorn, M. M. A., J. Haverkamp, and J. E. Thomas-Oates. 1999. Mass spectrometric analysis of Klebsiella pneumoniae ssp. pneumoniae rough strain R20 (O1−:K20−) lipopolysaccharide preparations: identifications of novel core oligosaccharide components and three 3-deoxy-D-manno-oct-2-ulopyranosonic artifacts. J. Mass Spectrom. 34:622-636. [DOI] [PubMed] [Google Scholar]
- 28.Orskov, I., and F. Orskov. 1984. Serotyping of Klebsiella. Methods Microbiol. 14:143-164. [Google Scholar]
- 29.Pradel, E., and C. A. Schnaitman. 1991. Effect of rfaH (sfrB) and temperature on expression of rfa genes of Escherichia coli K-12. J. Bacteriol. 173:6428-6431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Raetz, C. R., and C. Whitfield. 2002. Lipopolysaccharide endotoxins. Annu. Rev. Biochem. 71:635-700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Reed, L. J., and C. H. Muench. 1938. A simple method of estimating fifty percent end points. Am. J. Hyg. 27:493-497. [Google Scholar]
- 32.Regué, M., N. Climent, N. Abitiu, N. Coderch, S. Merino, L. Izquierdo, M. Altarriba, and J. M. Tomás. 2001. Genetic characterization of the Klebsiella pneumoniae waa gene cluster, involved in core lipopolysaccharide biosynthesis. J. Bacteriol. 183:3564-3573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Saccente, M. 1999. Klebsiella pneumoniae in liver abscess, endophthalmitis, and meningitis in a man with newly recognized diabetes mellitus. Clin. Infect. Dis. 29:1570-1571. [DOI] [PubMed] [Google Scholar]
- 34.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: A laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 35.Sanger, F., S. Nicklen, and A. R. Coulson. 1977. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74:5463-5467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Spina, E., L. Sturiale, D. Romeo, G. Impallomeni, D. Garozzo, D. Waidelich, and M. Glueckmann. 2004. New fragmentation mechanisms in matrix-assisted laser/desorption ionization time-of-flight tandem mass spectrometry of carbohydrates. Rapid Commun. Mass Spectrom. 18:392-398. [DOI] [PubMed] [Google Scholar]
- 37.Süsskind, M., L. Brade, H. Brade, and O. Holst. 1998. Identification of a novel heptoglycan of α 1→2-linked D-glycero-D-manno-heptopyranose. J. Biol. Chem. 273:7006-7017. [DOI] [PubMed] [Google Scholar]
- 38.Trautmann, M., M. Ruhnke, T. Rukavina, T. K. Held, A. S. Cross, R. Marre, and C. Whitfield. 1997. O-antigen seroepidemiology of Klebsiella clinical isolates and implications for immunoprophylaxis of Klebsiella infections. Clin. Diagn. Lab. Immunol. 4:550-555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tsai, C. M., and C. E. Frasch. 1982. A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal. Biochem. 119:115-119. [DOI] [PubMed] [Google Scholar]
- 40.Vinogradov, E., M. Cedzynski, A. Ziolkowski, and A. Swierzko. 2001. The structure of the core region of the lipopolysaccharide from Klebsiella pneumoniae O3. 3-deoxy-alpha-D-manno-octulosonic acid (alpha-Kdo) residue in the outer part of the core, a common structural element of Klebsiella pneumoniae O1, O2, O3, O4, O5, O8, and O12 lipopolysaccharides. Eur. J. Biochem. 268:1722-1729. [DOI] [PubMed] [Google Scholar]
- 41.Vinogradov, E., E. Frirdich, L. L. MacLean, M. B. Perry, B. O. Petersen, J. O. Duus, and C. Whitfield. 2002. Structures of lipopolysaccharides from Klebsiella pneumoniae. Elucidation of the structure of the linkage region between core and polysaccharide O chain and identification of the residues at the nonreducing termini of the O chains. J. Biol. Chem. 277:25070-25081. [DOI] [PubMed] [Google Scholar]
- 42.Vinogradov, E., and M. B. Perry. 2001. Structural analysis of the core region of the lipopolysaccharides from eight serotypes of Klebsiella pneumoniae. Carbohydr. Res. 335:291-296. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.