Abstract
Background and Aims
In coronavirus disease 2019 (COVID-19) patients, several serum biomarkers have been identified. Upon intensive care unit (ICU) admission, these laboratory markers become more crucial to distinguish between patients with severe cases of COVID-19. It might assist doctors in predicting the course of illnesses and treating patients appropriately. This work was to investigate the role of biomarkers in patients with COVID-19 classification admitted to the hospital and identified by reverse transcription polymerase chain reaction (RT-PCR).
Methods
Peripheral blood sample was taken from COVID-19 cases isolated on admission to determine C-reactive protein (CRP), D-dimer, Fibrinogen, neutrophil–lymphocyte ratio (NLR), leukocytes CRP ratio (LeCR), lymphocyte–CRP ratio (LCR), interleukin-6 (IL6), leukocytes interleukin 6 ratio (LeIL6), systemic inflammatory index (SII), platelet-to-lymphocyte ratio (PLR), and tissue plasminogen activator inhibitor one (tPAI-1). Follow-up for IL6, Ferritin, D-dimer, and tPAI-1 were determined on the 3rd and 7th days.
Results
Comparisons of severity revealed that hypertension, chronic obstructive pulmonary disease (COPD), and Ischemia were major risk factors in COVID-19 patients. There was a statistically significant difference between the test groups for fibrinogen (p < 0.000), IL6 (p < 0.009), LeCR (p < 0.006), and LCR (p < 0.011).
Conclusion
Based on laboratory test findings at the time of ICU admission, we can distinguish severe cases of COVID-19.
Keywords: COVID-19, intensive care unit, risk factors, RT- PCR, serum biomarkers
Introduction
Clinicians urgently need clinical laboratory predictors of disease progression toward the severe form to stratify risk, separate critical patients from mild/moderate coronavirus disease 2019 (COVID-19) patients, and differentiate critical patients from mild/moderate COVID-19 patients.1 A significant number of confirmed COVID-19 patients had laboratory abnormalities, including complete blood count (CBC) variants, cardiac and coagulation parameters, renal and liver function examinations, infection-related factors, as well as clinical symptoms and pulmonary computed tomography (CT) scans.2
Recently, a combination of laboratory tests has been used to detect and diagnose the hyperinflammatory state.2 Neutrophil-to-Lymphocyte Ratio (NLR), Lymphocyte to C Reactive Protein (LCR), Leukocyte to C Reactive Protein (LeCR), Platelet Lymphocyte Ratio (PLR), Systemic Inflammatory Index (SII), and Interleukins Leukocyte-6 (LeIL6) are examples of such pairs.3–5
Given that the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) undergoes continuous evolution, phylogenetic analysis was an additional focus of research to correlate the prognosis and death rate with the variants of the virus.6
Furthermore, the feasibility and reliability of capillary blood self-testing as an alternative to the more invasive venous blood testing serological assays for the presence of anti-SARS-CoV-2 for research and monitoring of the pandemic have been proposed.7
Surprisingly, some indicators of coagulation and fibrinolysis may provide a more reflective picture of hemostatic disorders in COVID-19 patients.8 D-dimer and tissue plasminogen activator inhibitor-1 (tPAI-1) are both such markers. As the role of reactive oxygen species (ROS) and cytokine storms in the genesis of COVID-19 becomes clearer, several arguments have emerged in favor of the use of N-acetyl cysteine (NAC) in COVID-19.9,10 It has been suggested that NAC may be explored in COVID-19 as a therapeutic modality to prevent excessive ROS production.11
Thus, this study was conducted to investigate the role of various biomarkers in the hope that they might aid in discriminating between severe and non-severe cases of COVID-19 individuals who were admitted to the hospital.
Methods
Study Design & Patients
A total of 101 patients with RT-PCR positive for COVID-19 enrolled in our study. All patients attended the National Liver Institute and Menoufia University Hospitals during the period study from August 2022 to February 2023.
Ethics Approval and Consent to Participate
All study protocols were carried out in compliance with the Helsinki Declaration and were authorized by Menoufia University’s National Liver Institutional Ethics Committee (Approval Number: 00211/2020). The patients were informed about the study’s benefits, potential risks, and all stages of the procedure. Before enrolling in the study, all patients signed a written informed agreement to participate in the study and for data dissemination. A parent or legal guardian of participants under 18 years of age also provided informed written consent.
Patients’ Criteria of Our Study
The study included patients with RT-PCR positive tests for COVID-19, while patients without any clinical features of COVID-19 or those who were not confirmed by RT-PCR tests were excluded.
The age of the included patients ranged from 9 to 75 years, and both genders were well represented, with 57 males (56.4%) and 44 females (43.6%). The body mass index (BMI) of the participants ranged from 16.02 to 42.22 kg/m², and 21 patients (20.8%) were classified as obese, having a BMI of 30 kg/m² or higher.
In terms of clinical comorbidities, hypertension was observed in 15 patients (14.9%), while chronic lung disease was present in 10 patients (9.9%). Additionally, 9 patients (8.9%) were diagnosed with diabetes, and ischemic heart disease was identified in 11 patients (10.9%).
All Subjects Included in the Current Study Were Subjected to the Following
On admission, nursing staff drew a blood sample for a differential CBC using an Automated Haematology Analyzer Sysmex XT 1800i (Sysmex Corporation, Kobe, Japan), a renal function test (RFT) and liver function test (LFT) using a Cobas 6000 analyzer (c501 module, Roche Diagnostics GmbH, D-68305 Mannheim, Germany), and D-dimer Enzyme Linked Immunosorbent Assay (ELISA) (kit cat#201-12-0091) was used for automated hemostasis testing of NLR, LeCR, LCR, IL6, SII, PLR, and tPAI-1.
Patients were tested for PCR-RT after being admitted to the hospital, and all results were positive. The tests for IL6, D-dimer, ferritin, and tPAI-1 were repeated on the 3rd and 7th days. The selection of the 3rd and 7th days for patient follow-up was informed by both clinical experience and findings from previous studies on COVID-19 progression. Clinically, the 3rd day marks an early stage where patients often show signs of stabilization or early deterioration, and monitoring key biomarkers such as D-dimer and IL-6 provides valuable insight into the patient’s inflammatory and coagulatory status. The 7th day is widely recognized as a critical point in disease progression when patients who will develop severe symptoms typically begin to exhibit more pronounced respiratory distress or other complications requiring intensive care.12,13
Cases were classified according to the need for hospitalization or intensive care admission (ICU) by the chest physician based on respiratory distress, tachypnea, and the need for high-flow oxygen supplementation. All cases followed the Menoufia University protocol in which they received therapeutic anticoagulants, an N-acetylcysteine infusion of 600 mg/8 hours, and corticosteroids, but antiviral treatment was determined by the chest physician and intensive care doctor.
Statistical Analysis Methods
The Statistical Package for Social Science (SPSS) program was used for statistical analysis (version 23, Inc., Chicago, IL, USA). Data was entered as either numerical or categorical, depending on the situation. Quantitative descriptive data is presented as mean ±SD and range, as well as qualitative data as frequency and percent (%). The Chi-square test was used to examine the connection between qualitative variables, and the Fisher's exact test was employed for 2×2 qualitative variables when more than 25% of the cells had an anticipated count less than 5. The Student’s t-test was used to compare the mean ± standard deviation (SD) of two sets of quantitatively normally distributed data; the Mann–Whitney test was used when the data was not normally distributed. The determination of cutoff values for biomarkers was performed using Receiver Operating Characteristic (ROC) curve analysis to assess their predictive ability for COVID-19 severity. The ROC curve was employed to calculate the area under the curve (AUC), which reflects the diagnostic accuracy of each biomarker. The optimal cutoff points were determined by maximizing the Youden Index, which balances sensitivity and specificity. In this analysis, assumptions included the continuous nature of biomarker levels and their association with disease severity. The clinical significance of these cutoffs lies in their ability to stratify patients based on the likelihood of severe or critical COVID-19 outcomes. These cutoffs, therefore, have practical applications in identifying patients at higher risk for severe complications, allowing for more targeted clinical interventions. The effect of intravenous N-acetylcysteine (IV NAC) on inflammation markers was assessed in COVID-19 patients over time. The data includes readings taken on admission (1st reading), the 3rd day (2nd reading), and the 7th day (3rd reading). P-value ≤0.05, considered statistically significant level.
Results
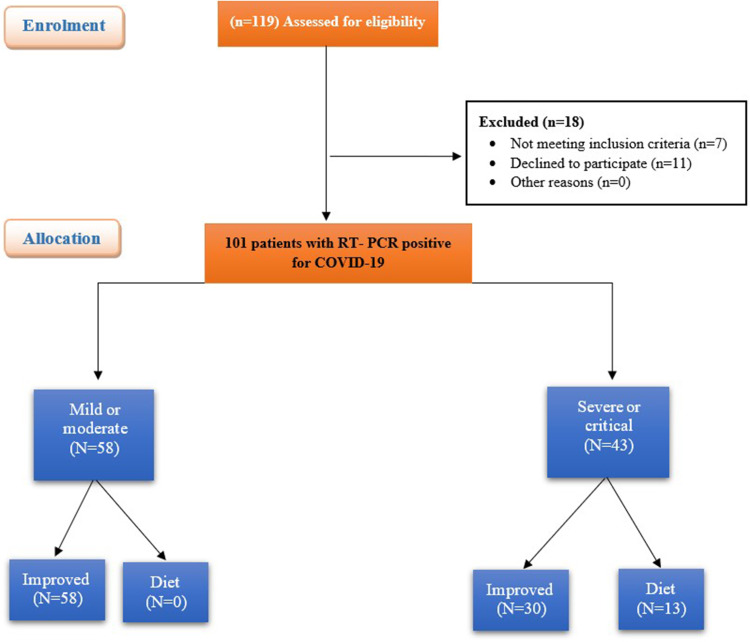
A flowchart of the study population is shown in Figure 1. Of the 119 patients with RT-PCR positive for COVID, eighteen patients were excluded from the study (11 patients declined consent and 7 patients did not meet the inclusion criteria). One hundred and one patients with RT-PCR positive for COVID participated in the study. The mean age of patients in the severe or critical group (51.83 ± 10.98 years) was higher than that of the mild or moderate group (39.34 ± 10.43 years). This difference was statistically significant (P = 0.0001). There was no significant difference in gender distribution between the groups, with males comprising 58.6% of the mild/moderate group and 53.5% of the severe/critical group (P = 0.607). BMI did not significantly differ between the two groups (P = 0.805) (Table 1).
Figure 1.
Flowchart of the studied groups.
Table 1.
Demographics and Clinical Features of COVID-19 Patients (n = 101)
| Clinical Variables | Mild or moderate (N=58) | Severe or critical (N=43) | P-value Mann–Whitney U-test |
|---|---|---|---|
| Age/year | 0.0001 | ||
| Mean ±SD. | 39.34± 10.43 | 51.83±10.98 | |
| Median (range) | 38.5(9–66) | 51.00(34–75) | |
| Gender, No. (%) | 0.607 (X2) | ||
| Male | 34 (58.6%) | 23(53.5%) | |
| Female | 24 (41.4%) | 20 (46.5%) | |
| BMI (kg/m2) | 0.805 | ||
| Mean ±SD. | 28.34± 5.46 | 27.62±4.07 | |
| Median (range) | 27.86(16.16–42.22) | 27.34(16.02–35.06) | |
| Hypertension | 0.009 (X2) | ||
| No | 54(93.1%) | 32 (74.4%) | |
| Yes | 4(6.9%) | 11 (25.6%) | |
| Chronic lung disease | 0.012 (X2) | ||
| No | 56 (96.6%) | 35 (81.4%) | |
| Yes | 2 (3.4%) | 8 (18.6%) | |
| Diabetes | 0.409 (X2) | ||
| No | 54 (93.1%) | 38 (88.4%) | |
| Yes | 4 (6.9%) | 5 (11.6%) | |
| Ischemic | 0.005 (X2) | ||
| No | 56 (96.6%) | 34 (79.1%) | |
| Yes | 2 (3.4%) | 9 (20.9%) | |
| Obese | 0.641 (X2) | ||
| No | 45 (77.6%) | 35 (81.4%) | |
| Yes | 13 (22.4%) | 8 (18.6%) | |
| Hospital stays/day | 0.01 | ||
| Mean ±SD. | 7.24± 0.94 | 8.72±3.20 | |
| Median (range) | 7(6–10) | 8.0(6–20) | |
| Stroke | 0.134 (X2) | ||
| No | 54 (93.1%) | 36 (83.7%) | |
| Yes | 4 (6.9%) | 7 (16.3%) | |
| Needs in hospital intubation | 0 (0.0%) | 9 (20.9%) | 0.0001 (X2) |
| Oxygen Saturation Admission | 0.0001 (t) | ||
| Mean ±SD. | 86.43± 4.28 | 82.44± 4.93 | |
| Median (range) | 86(77–95) | 84(75–95) | |
| Mortality, No. (%) | 0.0001 (X2) | ||
| Improved | 58 (100.0%) | 30(69.8%) | |
| Died | 0 (0.0%) | 13 (30.2%) |
Abbreviations: χ2, Chi-square; t, Independent test.
A significantly higher proportion of patients in the severe/critical group had hypertension (25.6%) compared to the mild/moderate group (6.9%), with a P-value of 0.009. The presence of chronic lung disease was more common in the severe/critical group (18.6%) compared to the mild/moderate group (3.4%), with a statistically significant difference (P = 0.012). Although the prevalence of diabetes was higher in the severe/critical group (11.6%) compared to the mild/moderate group (6.9%), this difference was not statistically significant (P = 0.409). Ischemic heart disease was significantly more prevalent in the severe/critical group (20.9%) compared to the mild/moderate group (3.4%), with a P-value of 0.005. The prevalence of obesity was similar between the two groups (22.4% in the mild/moderate group vs 18.6% in the severe/critical group), with no significant difference (P = 0.641) (Table 1).
The mean duration of hospital stay was significantly longer in the severe/critical group (8.72 ± 3.20 days) compared to the mild/moderate group (7.24 ± 0.94 days), with a P-value of 0.01. Stroke was more common in the severe/critical group (16.3%) compared to the mild/moderate group (6.9%), although this difference did not reach statistical significance (P = 0.134). None of the patients in the mild/moderate group required intubation, while 20.9% of the severe/critical group did, with a highly significant P-value of 0.0001 (Table 1).
The mean oxygen saturation on admission was significantly lower in the severe/critical group (82.44 ± 4.93%) compared to the mild/moderate group (86.43 ± 4.28%), with a P-value of 0.0001. All patients in the mild/moderate group survived (0% mortality), whereas 30.2% of patients in the severe/critical group died, with a highly significant P-value of 0.0001 (Table 1).
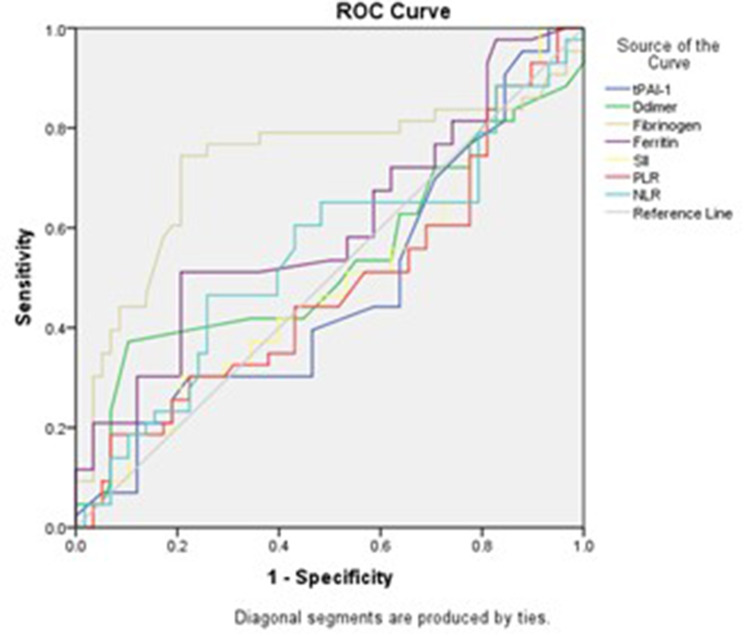
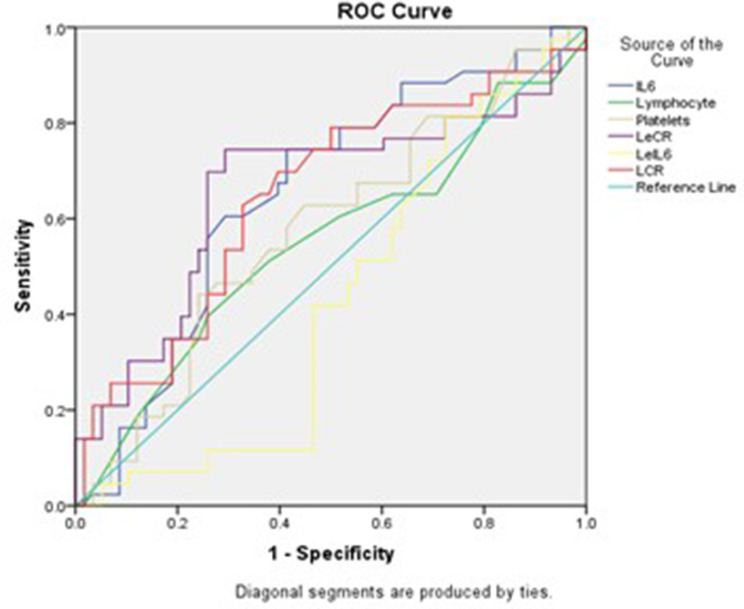
Considering the findings of ROC analysis, fibrinogen demonstrated the highest predictive value with a cut-off value of ≥862.5 mg/dL, an AUC of 0.728, a sensitivity of 74.4%, and a specificity of 79.3%. In contrast, tPAIS1, with a cut-off value of ≥0.95 ng/mL, an AUC of 0.476, showed poor predictive ability, with relatively low specificity (29%) despite a moderate sensitivity (69.8%). Ferritin exhibited moderate predictive capability with a cut-off value of ≥942 ng/mL, an AUC of 0.603, offering high specificity (79.3%) but lower sensitivity (51.2%). IL-6 emerged also as a moderately effective predictor of COVID-19 severity with a cut-off value of ≤11.25 pg/mL, an AUC of 0.653, a sensitivity of 72.1%, and a specificity of 58.6%. The leukocyte–CRP ratio (LeCR) also showed good predictive ability with a cut-off value of ≤0.34, an AUC of 0.660, a sensitivity of 74.4%, and a specificity of 70.7%. The lymphocyte–CRP ratio (LCR) presented moderate predictive power with a cut-off value of ≤0.62, an AUC of 0.649, with balanced sensitivity (65.1%) and specificity (63.8%). D-dimer showed limited discriminatory power with a cut-off value of ≥3.55 ng/mL, an AUC of 0.533, alongside modest sensitivity (53.5%) and specificity (44.8%). SII, PLR, and NLR all demonstrated weak discriminatory power, with AUC values close to 0.5, with sensitivities and specificities ranging between 43.1% and 65.1%. Other biomarkers, such as the leukocyte-IL-6 ratio (LeIL-6) and platelet count, showed weaker predictive abilities, with AUCs of 0.424 and 0.577, respectively (Table 2A and 2B, Figure 2 and 3).
Table 2.
Recommended Cut off Values for Biomarkers in Prediction of Severity of Covid-19 Patients
| Variable | AUC | Cutoff | Std. Error | Asymptotic Sig. b | Asymptotic 95% CI | Sensitivity | Specificity | |
| Lower Bound | Upper Bound | |||||||
| A | ||||||||
| tPAIS1 (ng/mL) | 0.476 | 0.95 | 0.059 | 0.675 | 0.360 | 0.591 | 69.8% | 29% |
| D-dimer1 (ng/mL) | 0.533 | 3.55 | 0.062 | 0.573 | 0.412 | 0.654 | 53.5% | 44.8% |
| Fibrinogen (mg/dL) | 0.728 | 862.5 | 0.057 | 0.000 | 0.618 | 0.839 | 74.4% | 79.3% |
| Ferritin (ng/mL) | 0.603 | 942 | 0.058 | 079 | 0.489 | 0.717 | 51.2% | 79.3% |
| SII (x10^9/L) | 0.489 | 2173.11 | 0.059 | 0.853 | 0.373 | 0.605 | 51.2% | 46.6% |
| PLR | 0.480 | 25.5778 | 0.060 | 0.731 | 0.363 | 0.597 | 51.2% | 43.1% |
| NLR | 0.545 | 8.1364 | 0.060 | 0.444 | 0.427 | 0.662 | 65.1% | 51.7% |
| B | ||||||||
| IL-6 (pg/mL) | 0.653 | 11.25 | 0.055 | 0.009 | 0.544 | 0.761 | 72.1% | 58.6% |
| Lymphocyte (10^9/L) | 0.548 | 9.5 | 0.059 | 0.414 | 0.431 | 0.664 | 51.2% | 62.1% |
| Platelets (10^9/L) | 0.577 | 268.5 | 0.058 | 0.190 | 0.463 | 0.690 | 62.8% | 55.2% |
| LeCR | 0.660 | 0.3418 | 059 | 0.006 | 0.545 | 775 | 74.4% | 70.7% |
| LeIL-6 | 0.424 | 0.9545 | 057 | 0.194 | 0.312 | 0.537 | 51.2% | 44.8% |
| LCR | 0.649 | 0.6235 | 0.056 | 0.011 | 0.538 | 0.759 | 65.1% | 63.8% |
Notes: PAIS1, PLR, NLR have at least one tie between the positive actual state group and the negative actual state group. Statistics may be biased. a. Under the nonparametric assumption. b. Null hypothesis: true area=0.5. Leukocytes CRP ratio (LeCR), lymphocyte–CRP ratio (LCR), interleukin (IL6), leukocytes interleukin 6 ratio (LeIL6) have at least one tie between the positive actual state group and the negative actual state group. Statistics may be biased. a. Under the nonparametric assumption. b. Null hypothesis: true area = 0.5.
Abbreviations: AUC, Area under curve; CI, Confidence Interval; Std, standard error.
Figure 2.
Receiver- operating characteristic curve (ROC) for biomarkers in prediction of severity of COVID-19 patients: tPAIS1: Tissue plasminogen activator inhibitor-1, measured in nanograms per milliliter (ng/mL); D-dimer: D-dimer level, measured in nanograms per milliliter (ng/mL); Fibrinogen: Fibrinogen level, measured in milligrams per deciliter (mg/dL); Ferritin: Ferritin level, measured in nanograms per milliliter (ng/mL); SII: Systemic immune-inflammation index, measured in billions per liter (10^9/L); PLR: Platelet–lymphocyte ratio; NLR: Neutrophil-lymphocyte ratio.
Figure 3.
Receiver- operating characteristic curve (ROC) for biomarkers in prediction of severity of COVID-19 patients: IL-6: Interleukin-6, measured in picograms per milliliter (pg/mL); Lymphocyte: Lymphocyte count, measured in billions per lite r (10^9/L); Platelets: Platelet count, measured in billions per liter (10^9/L); LeCR: Leukocyte–CRP ratio; LeIL-6: Leukocyte-Interleukin-6 ratio; LCR: Lymphocyte–CRP ratio.
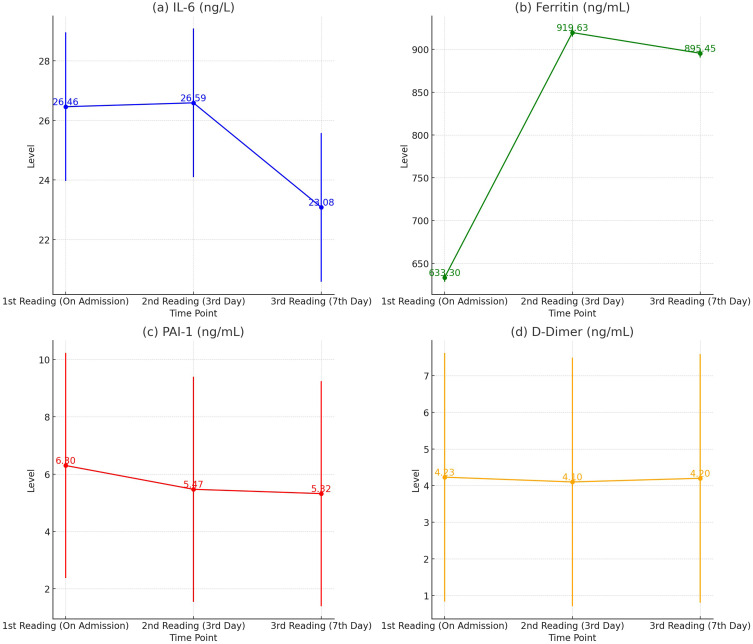
The assessment of the effect of IV NAC on inflammation markers showed that the IL-6 levels showed a slight but statistically significant decrease over the course of the study (P = 0.003). On admission, the mean IL-6 level was 26.46 ± 35.05 ng/l, which slightly increased to 26.59 ± 35.41 ng/l on the 3rd day, but then decreased to 23.08 ± 34.01 ng/l by the 7th day. The median values also showed a slight increase from 7.2 ng/l on admission to 10 ng/l on the 3rd day and then to 11 ng/l on the 7th day. Ferritin levels significantly increased from the time of admission through the 7th day (P = 0.0001). The mean ferritin level increased from 633.30 ± 382.32 ng/mL on admission to 919.63 ± 790.37 ng/mL on the 3rd day and slightly decreased to 895.45 ± 574.50 ng/mL by the 7th day. PAI-1 levels showed a statistically significant decrease over time (P = 0.003). On admission, the mean PAI-1 level was 6.30 ± 8.84 ng/mL, which decreased to 5.47 ± 7.43 ng/mL by the 3rd day and further to 5.32 ± 6.93 ng/mL by the 7th day. The D-dimer levels remained relatively stable but showed a slight and statistically significant decrease (P = 0.006). The mean D-dimer level was 4.23 ± 4.58 ng/mL on admission, slightly decreased to 4.10 ± 4.12 ng/mL on the 3rd day, and further to 4.20 ± 3.45 ng/mL by the 7th day (Table 3, Figure 4).
Table 3.
Effect of IV NAC on Inflammation Assessed by Serum Levels of IL6 (Ng/l) and Ferritin (Ng/Ml), t PAI.1 (Ng/Ml), D. Dimer and Clinical Outcome of COVID-19 Infection Patients
| Patient | 1st reading (On admission) | 2nd reading (3rd day) | 3rd reading (7th day) | P-value Friedman Test |
|---|---|---|---|---|
| IL-6 (pg/mL) | 26.46±35.05 | 26.59±35.41 | 23.08±34.01 | 0.003 |
| 101 | 101 | 101 | ||
| 7.2(0.50–155) | 10(1–130) | 11(0.7–140) | ||
| Ferritin (ng/mL) | 633.30±382.32 | 919.63±790.37 | 895.45±574.50 | 0.0001 |
| 101 | 101 | 101 | ||
| 540(219–3066) | 750(288–5455) | 800(129–3450) | ||
| PAI-1 (ng/mL) | 6.30±8.84 | 5.47±7.43 | 5.32±6.93 | 0.003 |
| 101 | 101 | 101 | ||
| 2(0.10–32) | 2(0.40–27) | 1.23(0.3–25) | ||
| D-Dimer (ng/mL) | 4.23±4.58 | 4.10±4.12 | 4.20±3.45 | 0.006 |
| 101 | 101 | 101 | ||
| 4(0.07–32) | 3(0.06–25) | 3.2(0.1–16) |
Figure 4.
Effect of IV NAC on inflammation assessed by serum levels of IL6 (ng/l) and ferritin (ng/mL), t PAI.1 (ng/mL), D. Dimer and clinical outcome of COVID-19 infection patients.
On admission, the mean IL-6 level was 26.46 ± 35.05 ng/l, which slightly increased to 26.59 ± 35.41 ng/l on the 3rd day, but then decreased to 23.08 ± 34.01 ng/l by the 7th day. Ferritin levels significantly increased from the time of admission through the 7th day (P = 0.0001). The mean ferritin level increased from 633.30 ± 382.32 ng/mL on admission to 919.63 ± 790.37 ng/mL on the 3rd day and slightly decreased to 895.45 ± 574.50 ng/mL by the 7th day. PAI-1 levels showed a statistically significant decrease over time (P = 0.003). On admission, the mean PAI-1 level was 6.30 ± 8.84 ng/mL, which decreased to 5.47 ± 7.43 ng/mL by the 3rd day and further to 5.32 ± 6.93 ng/mL by the 7th day. The D-dimer levels remained relatively stable but showed a statistically significant decrease (P = 0.006). The mean D-dimer level was 4.23 ± 4.58 ng/mL on admission, slightly decreased to 4.10 ± 4.12 ng/mL on the 3rd day, and further to 4.20 ± 3.45 ng/mL by the 7th day.
Discussion
Given the high fatality rates of COVID-19, as well as the high costs of admission and treatment, early identification of severe cases was required to avoid resource waste. As a result, any parameter that allows for the detection of severe instances is critical. As we evaluated risk variables on admission, we discovered that hypertension, chronic obstructive pulmonary disease (COPD), and ischemia were the most common. Our findings were strikingly similar to previous findings that COVID-19 infection was related to high rates of severity and mortality in COPD patients14 and hypertensive patients.15
There is a clear link between the clinical severity of COVID-19 disease and several biomarkers. As a result, a high fibrinogen level of ≥862.5 mg/dL may be employed as a prognostic marker for COVID-19. We believe fibrinogen is the most efficient biomarker for the following prognosis since it has the highest AUC = 0.728. This was significantly related to the clinical severity of the disease; Tang et al found that non-survivors had higher fibrinogen levels than survivors.16 In the context of coagulation biomarkers, the study of Ali et al17 observed significantly higher D-dimer levels in patients who succumbed to the disease compared to those who survived. In their study, elevated D-dimer, along with increased CRP and troponin levels, was strongly associated with disease severity and mortality. Our study suggested that while D-dimer remains an important marker with an AUC of 0.533, fibrinogen might offer superior predictive value in certain clinical settings.
For IL-6, LeCR and LCR values of ≤11.25 pg/mL, 0.34, and 0.62 with AUC s of 0.653, 0.660, and 0.649 suggested severity, which was consistent with Wu et al observation that combined ratios (LCR, LeCR) considerably decreased in severe compared to non-severe.18
Additionally, Liu19 declared that IL-6 could be a useful test for monitoring severe-type COVID-19.19 Further analysis revealed that the changes in the analysis of IL-6 follow-up revealed that the IL-6 mean in severe/non-severe cases was 17/33.3, p = 0.009 on admission, 19/32, p = 0.017 on day 3, and 17/27.5, p = 0.001 on day 7, and this was matched with results that observed IL-6 to be markedly decreased while the patient’s clinical picture is deteriorating.19
Our findings are consistent with the broader observation by Ali et al20 that unvaccinated patients experienced more severe symptoms and longer durations of illness during both the Delta and Omicron waves, highlighting the role of systemic inflammation and coagulation disturbances in severe COVID-19 cases. However, Ali et al20 specifically focused on CRP, WBCs, and PCR-Ct values as primary biomarkers. They reported lower CRP levels during the Omicron wave compared to the Delta wave, regardless of vaccination status. These differences may reflect the varying impact of different SARS-CoV-2 variants on inflammatory responses or the influence of vaccination on biomarker levels. Interestingly, Ali et al observed lower WBC counts in patients infected with the Delta variant compared to those with Omicron, a finding that was independent of vaccination status. In line with our study, Ali et al study highlighted the complexity of COVID-19’s clinical presentation and the role of various biomarkers in predicting disease severity.
We initiated NAC as a cornerstone in our treatment regimen in all hospitalized cases at a dose of 600 mg infusion every 8 hours, based on its therapeutic potential in reducing immunological activation and the resulting cytokine storm.21 This could explain why IL-6 levels remained steady during the follow-up period. NLR was found to be ≥8.1364 in our data for discriminating severe/non-severe cases, and this was documented by He et al, who documented NLR increased in severe compared to non-severe instances.22 In comparison, Yang et al discovered a cutoff of 3.3 for NLR to assist in estimating prognosis in critically ill patients.23 Patients with COVID-19.23 All of the enrolled individuals were routinely started on therapeutic LMWH. As a result, no significant variations in D-dimer and tPAI-1 findings were seen between severe and non-severe cases on admission. Following the same markers on the 3rd and 7th days, however, indicated a considerable drop in their levels, indicating an outstanding response to treatment.18
Additionally, Ji et al24 show patients who had direct vs indirect contact with Wuhan had significantly higher CRP and D-dimer levels but lower levels of LC, which does not indicate severity. When CRP levels were compared to other blood indicators in ratio, CRP changes were seen before others. ROC analysis by Tang et al16 showed CRP and disease progression have a very strong correlation. It’s interesting to note that the influenza A or B group exhibited higher absolute values of WCC and NLR than SARS-CoV-2. The severity of COVID-19 is anticipated to be predicted by increased IL-2R and IL-6 expression in serum found by Chen et al.25 Zhang et al26 reported in patients with COVID-19, D-dimer at admission levels greater than 2.0 g/mL could accurately predict in-hospital death and serve as an early indication for better care. Qin et al27 found IL-6 levels in COVID-19 that are severe and critical are significantly greater. Monitoring could aid in the early detection of serious illnesses. As for, D-dimer levels greater than 1 g/mL can aid doctors in early detection of individuals with poor prognosis, as concluded by Zhou et al.12 Although there was no statistical analysis observed despite the fact that there were clear differences in the lymphocytes (lymphocytopenia), platelet count (rose), and D-dimer (rose) in patients who suffered composite outcomes (ICU admission, invasive mechanical ventilation, and death).28
Our study’s evaluation of the effects of IV NAC on inflammation markers in COVID-19 patients offers a valuable point of comparison with the findings from the study by Ali et al29 on the side effects and serological biomarkers associated with the Pfizer/BioNTech COVID-19 vaccine. While the focus of Ali et al’s study was on the immune response and safety profile following vaccination, particularly the elevation of IgG antibodies and serum CRP levels, both studies provide insights into how different interventions—vaccination versus NAC treatment—modulate inflammatory responses in the context of COVID-19. Ali et al’s research demonstrated that the Pfizer/BioNTech vaccine elicited a strong immune response, evidenced by a significant increase in IgG antibodies and an elevated CRP level, particularly after the second dose. This controlled increase in CRP post-vaccination contrasts with the pathological rise in inflammatory markers like CRP and IL-6 that is commonly associated with severe COVID-19 cases and which NAC treatment in our study aimed to counteract. While the vaccine-induced CRP elevation observed by Ali et al29 reflects a normal and transient inflammatory response, our study’s findings show that in the setting of severe COVID-19, unchecked inflammation can lead to worsened outcomes unless actively managed by treatments like NAC. Moreover, the Ali et al29 study emphasized the importance of vaccination in preventing the severe inflammatory responses that necessitate treatments like NAC. By reducing the severity of the disease through effective immunization, the need for interventions aimed at curbing inflammation and oxidative stress, such as NAC, might be reduced.
Conclusions
In conclusion, our study provides clear evidence that the levels of specific biomarkers correlate with the severity of COVID-19 infection. These findings suggest that these biomarkers could be valuable tools for distinguishing severe cases of COVID-19 based on laboratory test results at the time of ICU admission. However, to enhance the utility of these biomarkers in daily clinical practice, further research is necessary.
Future studies should focus on validating these biomarkers in larger, more diverse populations to ensure their generalizability across different demographic and clinical settings. Additionally, longitudinal studies could explore how these biomarker levels change over the course of the disease and in response to various treatments, such as NAC therapy, to refine their prognostic value. Identifying a core set of laboratory biomarkers that can reliably predict prognosis and outcomes in hospitalized patients with severe COVID-19 would be a significant advancement in clinical practice, helping healthcare providers make more informed decisions and potentially improving patient outcomes.
Abbreviations
ICU, intensive care unit; COVID, Coronavirus disease; CRP, C-reactive protein; NLR, neutrophil–lymphocyte ratio; LeCR, leukocytes CRP ratio; LCR, lymphocyte–CRP ratio; IL6, interleukin-6; LeIL6, leukocytes interleukin 6 ratio; SII, systemic inflammatory index; PLR, platelet-to-lymphocyte ratio; tPAI-1, tissue plasminogen activator inhibitor one; COPD, Chronic obstructive pulmonary disease; CBC, complete blood count; CT, computerized tomography; ROS, Reactive oxygen species; RFT, renal function test; LFT, liver function test; CI, Confidence Interval; ROC, receiver operating characteristic; AUC, Area under curve.
Data Sharing Statement
All data and materials included in this work are available.
Ethics Approval and Consent to Participate
Our local Ethics Committee approved our study and a written consent for participation was obtained from all patients. All humans were used in accordance with the Helsinki Declaration of 1975.
Author Contributions
All authors made a significant contribution to the work reported, whether that is in the conception, study design, execution, acquisition of data, analysis and interpretation, or in all these areas; took part in drafting, revising or critically reviewing the article; gave final approval of the version to be published; have agreed on the journal to which the article has been submitted; and agree to be accountable for all aspects of the work.
Disclosure
The authors declare no competing interests in this work.
References
- 1.Lin S, Huang AC-C, Chiu T-H, et al. Clinical and laboratory predictors for disease progression in patients with COVID-19: a multi-center cohort study. Biomed J. 2023;46(1):100–109. doi: 10.1016/j.bj.2022.11.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Siavoshi F, Safavi-Naini SAA, Shirzadeh Barough S, et al. On-admission and dynamic trend of laboratory profiles as prognostic biomarkers in COVID-19 inpatients. Sci Rep. 2023;13(1):6993. doi: 10.1038/s41598-023-34166-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Abensur Vuillaume L, Lefebvre F, Benhamed A, et al. CREMS network (clinical research in emergency medicine and sepsis). Lymphocyte-to-C-reactive protein (LCR) ratio is not accurate to predict severity and mortality in patients with COVID-19 admitted to the ED. Int J Mol Sci. 2023;24(6):5996. doi: 10.3390/ijms24065996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kocaaslan R, Dilli D, Çitli R, et al. Diagnostic value of the systemic immune-inflammation index in newborns with urinary tract infection. Am J Perinatol. 2024;41(S 01):e719–e727. doi: 10.1055/s-0042-1757353 [DOI] [PubMed] [Google Scholar]
- 5.Erdogan A, Can FE, Gönüllü H, et al. Evaluation of the prognostic role of NLR, LMR, PLR, and LCR ratio in COVID-19 patients. J Med Virol. 2021;93(9):5555–5559. doi: 10.1002/jmv.27097 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Baggio S, Togni G, Eckerle I, Vuillemier N, Kaiser L, Gétaz L. Feasibility of home-based ELISA capillary blood self-testing for anti-SARS-CoV-2 antibodies. Pract Lab Med. 2022;31:e00290. doi: 10.1016/j.plabm.2022.e00290 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ali KM, Rashid PMA, Ali AM, et al. Clinical outcomes and phylogenetic analysis in reflection with three predominant clades of SARS-CoV-2 variants. Eur J Clin Invest. 2023;53(9):e14004. doi: 10.1111/eci.14004 [DOI] [PubMed] [Google Scholar]
- 8.Onishi T, Shimonishi N, Takeyama M, et al. The balance of comprehensive coagulation and fibrinolytic potential is disrupted in patients with moderate to severe COVID-19. Int J Hematol. 2022;115(6):826–837. doi: 10.1007/s12185-022-03308-w [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Alam MS, Hasan M, Maowa Z, et al. N-acetylcysteine reduces severity and mortality in COVID-19 patients: a systematic review and meta-analysis. J Adv Vet Anim Res. 2023;10(2):157–168. doi: 10.5455/javar.2023.j665 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Liu TH, Wu J-Y, Huang P-Y, et al. Clinical efficacy of N-acetylcysteine for COVID-19: a systematic review and meta-analysis of randomized controlled trials. Heliyon. 2024;10(3):e25179. doi: 10.1016/j.heliyon.2024.e25179 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Izquierdo-Alonso JL, Pérez-Rial S, Rivera CG, et al. N-acetylcysteine for prevention and treatment of COVID-19: current state of evidence and future directions. J Infect Public Health. 2022;15(12):1477–1483. doi: 10.1016/j.jiph.2022.11.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhou F, Yu T, Du R, et al. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395:1054–1062. doi: 10.1016/S0140-6736(20)30566-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.World Health Organization (WHO) guidelines on clinical management of COVID-19. Available from: https://www.who.int/teams/health-care-readiness/covid-19. Accessed October 18, 2024.
- 14.Alqahtani JS, Oyelade T, Aldhahir AM, et al. Prevalence, severity, and mortality associated with COPD and smoking in patients with COVID-19: a rapid systematic review and meta-analysis. PLoS One. 2020;15(5):e0233147–e0233147. doi: 10.1371/journal.pone.0233147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Tal-Singer R, Crapo JD. COPD at the time of COVID-19: a COPD foundation perspective. Chron Obstructive Pulm Dis. 2020;7(2):73–75. doi: 10.15326/2Fjcopdf.7.2.2020.0149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tang N, Li D, Wang X, Sun Z. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J Thromb Haemost. 2020;18(4):844–847. doi: 10.1111/jth.14768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Ali AM, Rostam HM, Fatah MH, et al. Serum troponin, D-dimer, and CRP level in severe coronavirus (COVID-19) patients. Immun inflamm DiS. 2022;10(3):e582. doi: 10.1002/iid3.582 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wu Z, McGoogan JM. Characteristics of and important lessons from the Coronavirus disease 2019 (COVID-19) outbreak in China: summary of a report of 72 314 cases from the Chinese center for disease control and prevention. JAMA. 2020;323(13):1239–1242. doi: 10.1001/jama.2020.2648 [DOI] [PubMed] [Google Scholar]
- 19.Liu T, Zhang J, Yang Y, et al. The Role of interleukin-6 in monitoring the severe case of coronavirus disease 2019. EMBO Mol Med. 2020;12(7):e12421. doi: 10.15252/emmm.202012421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ali AM, Tofiq AM, Rostam HM, et al. Disease severity and efficacy of homologous vaccination among patients infected with SARS-CoV-2 Delta or Omicron VOCs, compared to unvaccinated using main biomarkers. J Med Virol. 2022;94(12):5867–5876. doi: 10.1002/jmv.28098 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Mohanty RR, Padhy BM, Das S, et al. Therapeutic potential of N-acetyl cysteine (NAC) in preventing cytokine storm in COVID-19: a review of current evidence. Eur Rev Med Pharmacol Sci. 2021;25(6):2802–2807. doi: 10.26355/eurrev_202103_25442 [DOI] [PubMed] [Google Scholar]
- 22.He F, Deng Y, Li W. Coronavirus disease 2019: what we know? J Med Virol. 2020;92(7):719–725. doi: 10.1002/jmv.25766 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Yang A-P, Liu J-P, Tao W-Q, et al. The diagnostic and predictive role of NLR, d-NLR, and PLR in COVID-19 patients. Int Immunopharmacol. 2020;84:106504. doi: 10.1016/j.intimp.2020.106504 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ji W, Bishnu G, Cai Z, Shen X. Analysis clinical features of COVID-19 infection in secondary epidemic area and report potential biomarkers in evaluation. medRxiv. 2020. doi: 10.1101/2020.03.10.20033613 [DOI] [Google Scholar]
- 25.Chen N, Zhou M, Dong X, et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 2020;395(10223):507–513. doi: 10.1016/S0140-6736(20)30211-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhang L, Yan X, Fan Q, et al. D-dimer levels on admission to predict in-hospital mortality in patients with Covid-19. J Thromb Haemost. 2020;18(6):1324–1329. doi: 10.1111/jth.14859 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Qin C, Zhou L, Hu Z, et al. Dysregulation of immune response in patients with Coronavirus 2019 (COVID-19) in Wuhan, China. Clin Infect Dis. 2020;71(15):762–768. doi: 10.1093/cid/ciaa248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Guan W-J, Ni Z-Y, Hu Y, et al. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020;382(18):1708–1720. doi: 10.1056/NEJMoa2002032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Ali KM, Ali AM, Atta PM, et al. A study on the side effects caused by the Pfizer/BioNTech COVID-19 vaccine: focus on IgG antibodies and serological biomarkers. Cent Eur J Immunol. 2024;49(1):2–10. doi: 10.5114/ceji.2024.136382 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
All data and materials included in this work are available.