Abstract
Background
New-onset atrial fibrillation (NOAF) is the most common arrhythmia in critically ill patients admitted to intensive care and is associated with poor prognosis and disease burden. Identifying high-risk individuals early is crucial. This study aims to create and validate a NOAF prediction model for critically ill patients using machine learning (ML).
Methods
The data came from two non-overlapping datasets from the Medical Information Mart for Intensive Care (MIMIC), with MIMIC-IV used for training and subset of MIMIC-III used as external validation. LASSO regression was used for feature selection. Eight ML algorithms were employed to construct the prediction model. Model performance was evaluated based on identification, calibration, and clinical application. The SHapley Additive exPlanations (SHAP) method was used for visualizing model characteristics and individual case predictions.
Results
Among 16,528 MIMIC-IV patients, 1520 (9.2%) developed AF post-ICU admission. A model with 23 variables was built, with XGBoost performing best, achieving an AUC of 0.891 (0.873–0.888) in validation and 0.769 (0.756–0.782) in external validation. Key predictors included age, mechanical ventilation, urine output, sepsis, blood urea nitrogen, percutaneous arterial oxygen saturation, continuous renal replacement therapy and weight. A risk probability greater than 0.6 was defined as high risk. A friendly user interface had been developed for clinician use.
Conclusion
We developed a ML model to predict the risk of NOAF in critically ill patients without cardiac surgery and validated its potential as a clinically reliable tool. SHAP improves the interpretability of the model, enables clinicians to better understand the causes of NOAF, helps clinicians to prevent it in advance and improves patient outcomes.
Supplementary Information
The online version contains supplementary material available at 10.1186/s13054-024-05138-0.
Keywords: New-onset atrial fibrillation, Critically ill patients, Machine learning, Predictive models, MIMIC database
Introduction
New-onset atrial fibrillation (NOAF) is defined as the occurrence of atrial fibrillation (AF) in patients with no prior history of this condition. During AF, the loss of atrial function and the increase in ventricular rate can result in decreased cardiac output and hemodynamic disturbances [1]. NOAF is the most common arrhythmia encountered in critically ill patients admitted to the intensive care unit (ICU). The reported incidence of NOAF in this population varies widely, ranging from 1.7% to 43.9%, with significant heterogeneity among studies [2]. Research suggests that in patients with septic shock, the presence of NOAF serves as a marker of disease severity and represents an additional organ failure [3]. Furthermore, multiple studies have demonstrated a strong association between NOAF during critical illness and an increased risk of stroke, heart failure (HF), and both short-term and long-term mortality [4, 5]. While numerous studies have investigated NOAF following cardiac surgery [6, 7], research on critically ill patients who have not undergone cardiac surgery remains comparatively scarce. The identification and management of such patients and corresponding interventions continue to be challenging. Although several studies have shown a reduction in NOAF among high-risk patients, the quality of evidence supporting these findings is low [8, 9]. Notably, the incidence of NOAF among critically ill non-cardiac surgery patients is remarkably high [10]. In real-world clinical settings, the majority of critically ill patients developing NOAF are those with infections or other non-cardiac conditions, and these patients often do not receive timely, specialized intervention from cardiovascular specialists [11]. This gap in care underscores the pressing need for early identification of patients at high risk for AF within routine ICU settings and the exploration of potential targeted interventions.
Machine learning (ML) is gaining prominence in the field of medicine, demonstrating impressive results in predicting survival and prognosis among cancer patients [12]. In recent years, several ML models have been developed to identify individuals at risk of AF. However, these models are primarily limited to the general population or patients undergoing cardiac surgery [13, 14], with few models designed for routine identification of AF risk in the intensive care unit (ICU) setting. Furthermore, most studies rely on bedside electrocardiogram (ECG) for AF detection [15], which, despite its high accuracy, may not provide clinicians with sufficient information to prevent the onset of AF due to its short-term nature.
Despite the high accuracy achieved by ML models, the influence of individual variables on these models often remains unknown. This lack of transparency limits the application of ML in clinical practice [16]. SHapley Additive exPlanations (SHAP) combines optimal credit allocation with local explanations to visually represent the importance of each variable in the model [17], thereby providing a more interpretable output.
Therefore, this paper aims to build a model to identify NOAF risk groups in critically ill patients using ML methods, and to visually interpret the model using SHAP methods to assist clinicians in the clinical identification and intervention of high-risk groups.
Materials and methods
Data source
The data used to construct the model came from the Medical Information Mart for Intensive Care IV (MIMIC-IV, version: v2.2) [18, 19], which contains clinical information on 431,231 hospital admissions for 299,712 patients admitted to Beth Israel Deaconess Medical Center from 2008 to 2020. We also performed external validation using a subset of the MIMIC-III database [20], which included 26,836 admissions for 23,692 patients between 2001 and 2008, and there was no overlap with patients with MIMIC-IV. The corresponding author (Bing Xiao) passed the Collaborative Institutional Training Initiative (CITI) program exam and obtained a certificate (Record ID: 57,440,109). As the MIMIC database is de-identified, we do not need to obtain informed consent from patients. We made a verbal report to our hospital’s ethics committee and did not need to go through the normal approval process.
Participants
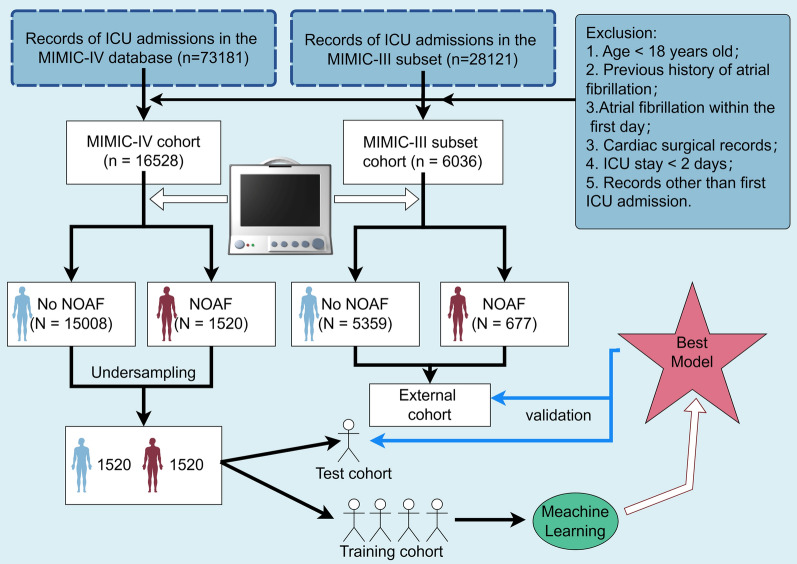
Patients who met the following criteria were included in the study: (1) Patients were older than 18 years; (2) Patients had been admitted to the ICU for more than two days; (3) Absence of atrial fibrillation event within first day; (4) Patients had not undergone cardiac surgery, including valve surgery and coronary artery bypass grafting; (5) Patients had no history of AF; and (6) For patients with multiple ICU admissions, only ICU admission records from the patient’s first admission were included. Figure 1 illustrated the patient screening process.
Fig. 1.
Patient screening flow from the MIMIC database. NOAF: new-onset atrial fibrillation
Data extraction and outcomes
Structured Query Language (SQL) in PostgreSQL was used to extract data from two databases on patients admitted to the ICU during the first 24 h. The variables extracted in this study were: (1) Demographic information: age, gender, race, and weight; (2) Comorbidities: myocardial infarction (MI), heart failure with reduced ejection fraction (HFrEF), heart failure with preserved ejection fraction (HFpEF), peripheral arterial disease, cerebrovascular disease, chronic lung disease, chronic kidney disease, chronic liver disease, hypertension, diabetes mellitus, sepsis; (3) Laboratory indicators: hemoglobin, white blood cells (WBC), platelets, blood urea nitrogen (BUN), creatinine, glucose, anion gap, potassium, sodium, calcium, creatine phosphate kinase (CK_CPK), creatine kinase isoenzymes (CK_MB), N-terminal pro-brain natriuretic peptide (NT-proBNP), urine output; (4) Vital signs: heart rate (HR), respiratory rate (RR), systolic blood pressure (SBP), diastolic blood pressure (DBP), temperature, percutaneous arterial oxygen saturation (SpO2); (5) Interventions: mechanical ventilation, continuous renal replacement therapy (CRRT), vasopressors, antibiotics. The maximum and minimum values of the first day were taken for multiple measurements, except for SpO2. To reduce the impact of missing data on model construction, the KNNImputer (KNN) method was used to impute data missing less than 20% and discard data missing more than 20% (Fig. S1).
The primary outcome was NOAF occurring after the first day of ICU admission, defined by heart rate status recorded at the nurse’s bedside [21].
Statistical analysis and model development
The Kolmogorov–Smirnov test was used for continuous variables. As continuous variables were all non-normal, the median (interquartile range) was used for description and the Mann–Whitney U test was used to compare differences between groups. Categorical variables were expressed as percentages (%) and Pearson chi-squared tests were used to compare differences between groups.
Due to class imbalance in the dependent variables, undersampling was used to resample the data to balance the data. The sample data were divided into a training set and an internal validation set by fivefold cross-validation sampling. In the case of many features, the lasso was used to select features, which is a method to introduce L1 regularization, select features and reduce dimensions by compressing coefficients, screening features with large contributions and eliminating redundant features.
In this study, eight ML algorithms, extreme gradient boosting (XGBoost), support vector machine (SVM), adaptive boosting (Adaboost), multilayer perceptron (MLP), neural network (NN), naive bayes (NB), logistic regression (LR) and gradient boosting machine (GBM), were used to construct the prediction model. The variables selected by lasso were included in the model. Ten-fold cross-validation was used to ensure the stability of the model. Grid tuning parameters were used to select the best tuning parameters for each algorithm. In the process of parameter adjustment, the highest area under the curve (AUC) of receiver operating characteristic (ROC) was selected as the optimal model. The models were built on the training set, and the internal validation set and external validation set were validated on the best model. The performance of the predictive model was assessed by AUC of the ROC curve, sensitivity, specificity, recall, F1 score, accuracy and recall. In addition, a decision curve analysis (DCA) and calibration curve were plotted to demonstrate the true clinical utility. To determine the optimal threshold probability for our model, we generated a clinical impact curve (CIC). This sophisticated visualization tool enabled us to rigorously assess and identify the most effective decision threshold for clinical application [22].
Using the SHAP method, a swarm diagram was drawn to show the contribution of each feature to the prediction results. SHAP evaluations of selected cases showed how much a feature affected a particular sample and helped us understand the model’s decision-making process. Finally, we used feature recursive elimination to further filter variables to lay out a simplified version of the model.
All statistical analyses were performed in R software (version: 4.3.3) and two-sided p-values less than 0.05 was considered significant.
Results
Baseline characteristics
After screening, a total of 16,528 MIMIC-IV patients were included in the study, and 1520 (9.2%) developed AF after ICU admission. A total of 6037 patients were drawn from the MIMIC-III subset with the same inclusion criteria, and 677 (11.2%) developed AF. Differences in baseline characteristics between MIMIC-IV and MIMIC-III subset patients were shown in Table S1.
Table 1 showed baseline information of all patients enrolled in MIMIC-IV database. Notably, older and white patients were more prone to developing NOAF during hospitalization. These patients experienced prolonged hospital and ICU stays, with substantially higher in-hospital mortality rates (28.42% vs. 11.63% in non-NOAF patients). NOAF patients also exhibited higher incidences of MI, HFrEF, HFpEF, peripheral arterial disease, chronic lung disease, chronic kidney disease, diabetes, hypertension, sepsis. On the first day of ICU admission, NOAF patients more frequently required interventions such as vasopressors, antibiotics, mechanical ventilation, and CRRT. Laboratory and vital signs assessments revealed that NOAF patients had lower levels of hemoglobin, platelets, SBP, DBP, MBP, temperature, SpO2 and urine output. Conversely, they demonstrated elevated WBC, BUN, creatinine, glucose, anion gap, potassium, RR compared to patients without NOAF.
Table 1.
Comparison of baseline characteristics in the non-NOAF and NOAF groups
| Variables | New-onset atrial fibrillation | P | ||
|---|---|---|---|---|
| Total (n = 16,528) | Non-NOAF (n = 15,008) | NOAF (n = 1520) | ||
| Admission age, M (Q1, Q3) | 64.19 (51.98, 75.98) | 62.94 (50.80, 74.73) | 74.19 (65.31, 82.86) | < 0.001 |
| Weight, M (Q1, Q3) | 78.00 (65.30, 93.70) | 78.00 (65.30, 93.45) | 78.80 (65.00, 95.30) | 0.192 |
| Gender, n (%) | 0.752 | |||
| Female | 7686 (46.50) | 6985 (46.54) | 701 (46.12) | |
| Male | 8842 (53.50) | 8023 (53.46) | 819 (53.88) | |
| Race, n (%) | < 0.001 | |||
| White | 10,490 (63.47) | 9429 (62.83) | 1061 (69.80) | |
| Black | 1634 (9.89) | 1540 (10.26) | 94 (6.18) | |
| Asian | 515 (3.12) | 466 (3.11) | 49 (3.22) | |
| Hispanic | 607 (3.67) | 575 (3.83) | 32 (2.11) | |
| Other | 3282 (19.86) | 2998 (19.98) | 284 (18.68) | |
| Los hospital, M (Q1, Q3) | 9.20 (5.82, 16.06) | 8.97 (5.70, 15.75) | 12.03 (7.68, 20.08) | < 0.001 |
| Los icu, M (Q1, Q3) | 3.85 (2.71, 6.79) | 3.71 (2.66, 6.21) | 6.58 (3.86, 11.63) | < 0.001 |
| Hospital expire flag, n(%) | < 0.001 | |||
| No | 14,351 (86.83) | 13,263 (88.37) | 1088 (71.58) | |
| Yes | 2177 (13.17) | 1745 (11.63) | 432 (28.42) | |
| MI, n (%) | < 0.001 | |||
| No | 14,206 (85.95) | 13,027 (86.80) | 1179 (77.57) | |
| Yes | 2322 (14.05) | 1981 (13.20) | 341 (22.43) | |
| Chronic lung disease, n (%) | < 0.001 | |||
| No | 12,520 (75.75) | 11,476 (76.47) | 1044 (68.68) | |
| Yes | 4008 (24.25) | 3532 (23.53) | 476 (31.32) | |
| Chronic renal disease, n (%) | < 0.001 | |||
| No | 13,716 (82.99) | 12,585 (83.86) | 1131 (74.41) | |
| Yes | 2812 (17.01) | 2423 (16.14) | 389 (25.59) | |
| Diabetes, n (%) | < 0.001 | |||
| No | 12,207 (73.86) | 11,152 (74.31) | 1055 (69.41) | |
| Yes | 4321 (26.14) | 3856 (25.69) | 465 (30.59) | |
| Chronic liver disease, n (%) | 0.463 | |||
| No | 14,107 (85.35) | 12,800 (85.29) | 1307 (85.99) | |
| Yes | 2421 (14.65) | 2208 (14.71) | 213 (14.01) | |
| Peripheral vascular disease, n (%) | < 0.001 | |||
| No | 14,888 (90.08) | 13,578 (90.47) | 1310 (86.18) | |
| Yes | 1640 (9.92) | 1430 (9.53) | 210 (13.82) | |
| Cerebrovascular disease, n (%) | 0.267 | |||
| No | 13,217 (79.97) | 11,985 (79.86) | 1232 (81.05) | |
| Yes | 3311 (20.03) | 3023 (20.14) | 288 (18.95) | |
| Hypertension, n (%) | < 0.001 | |||
| No | 6772 (40.97) | 6327 (42.16) | 445 (29.28) | |
| Yes | 9756 (59.03) | 8681 (57.84) | 1075 (70.72) | |
| HFrEF, n (%) | < 0.001 | |||
| No | 14,462 (87.50) | 13,265 (88.39) | 1197 (78.75) | |
| Yes | 2066 (12.50) | 1743 (11.61) | 323 (21.25) | |
| HFpEF, n (%) | < 0.001 | |||
| No | 15,012 (90.83) | 13,728 (91.47) | 1284 (84.47) | |
| Yes | 1516 (9.17) | 1280 (8.53) | 236 (15.53) | |
| Sepsis, n (%) | < 0.001 | |||
| No | 6759 (40.89) | 6416 (42.75) | 343 (22.57) | |
| Yes | 9769 (59.11) | 8592 (57.25) | 1177 (77.43) | |
| Hemoglobin min, M (Q1, Q3) | 10.40 (8.70, 12.10) | 10.50 (8.80, 12.10) | 9.90 (8.47, 11.53) | < 0.001 |
| Hemoglobin max, M (Q1, Q3) | 11.70 (10.10, 13.40) | 11.70 (10.10, 13.40) | 11.40 (9.88, 13.03) | < 0.001 |
| WBC min, M (Q1, Q3) | 9.50 (6.90, 12.90) | 9.40 (6.90, 12.80) | 10.20 (7.20, 13.90) | < 0.001 |
| WBC max, M (Q1, Q3) | 12.70 (9.20, 17.50) | 12.60 (9.10, 17.40) | 13.60 (9.90, 18.90) | < 0.001 |
| Platelets min, M (Q1, Q3) | 184.00 (129.00, 245.00) | 185.00 (130.00, 246.00) | 171.00 (117.00, 232.25) | < 0.001 |
| Platelets max, M (Q1, Q3) | 219.00 (161.00, 288.00) | 220.00 (162.00, 289.00) | 210.00 (153.00, 279.25) | 0.001 |
| BUN min, M (Q1, Q3) | 17.00 (11.00, 27.00) | 16.00 (11.00, 26.00) | 22.00 (15.00, 37.00) | < 0.001 |
| BUN max, M (Q1, Q3) | 20.00 (14.00, 33.00) | 19.00 (13.00, 32.00) | 27.00 (18.00, 45.00) | < 0.001 |
| Aniongap max, M (Q1, Q3) | 16.00 (14.00, 19.00) | 16.00 (14.00, 19.00) | 17.00 (14.00, 20.00) | < 0.001 |
| Aniongap min, M (Q1, Q3) | 13.00 (11.00, 15.00) | 13.00 (11.00, 15.00) | 14.00 (11.00, 16.00) | < .001 |
| Creatinine min, M (Q1, Q3) | 0.90 (0.70, 1.30) | 0.80 (0.60, 1.20) | 1.00 (0.70, 1.70) | < 0.001 |
| Creatinine max, M (Q1, Q3) | 1.00 (0.80, 1.60) | 1.00 (0.80, 1.50) | 1.25 (0.90, 2.20) | < 0.001 |
| Glucose min, M (Q1, Q3) | 113.00 (95.00, 136.00) | 113.00 (95.00, 136.00) | 118.00 (96.75, 143.00) | < 0.001 |
| Glucose max, M (Q1, Q3) | 148.00 (120.00, 194.00) | 147.00 (119.00, 192.00) | 159.00 (129.00, 209.00) | < 0.001 |
| Sodium min, M (Q1, Q3) | 137.00 (134.00, 140.00) | 137.00 (134.00, 140.00) | 137.00 (134.00, 140.00) | 0.008 |
| Sodium max, M (Q1, Q3) | 140.00 (137.00, 143.00) | 140.00 (137.00, 143.00) | 140.00 (137.00, 143.00) | 0.205 |
| Potassium min, M (Q1, Q3) | 3.80 (3.50, 4.20) | 3.80 (3.50, 4.20) | 3.90 (3.50, 4.30) | < 0.001 |
| Potassium max, M (Q1, Q3) | 4.40 (4.00, 4.90) | 4.30 (4.00, 4.80) | 4.50 (4.10, 5.20) | < 0.001 |
| Urine output, M (Q1, Q3) | 1585.00 (970.00, 2450.00) | 1625.00 (1000.00, 2495.00) | 1200.00 (692.00, 1900.00) | < 0.001 |
| HR min, M (Q1, Q3) | 69.00 (60.00, 81.00) | 70.00 (60.00, 81.00) | 68.00 (60.00, 80.00) | 0.114 |
| HR max, M (Q1, Q3) | 103.00 (90.00, 117.00) | 103.00 (90.00, 117.00) | 101.00 (89.00, 117.00) | 0.058 |
| SBP min, M (Q1, Q3) | 91.00 (82.00, 103.00) | 92.00 (82.00, 104.00) | 86.00 (77.00, 97.00) | < 0.001 |
| SBP max, M (Q1, Q3) | 150.00 (135.00, 166.00) | 150.00 (135.00, 166.00) | 148.00 (133.00, 165.00) | 0.006 |
| DBP min, M (Q1, Q3) | 46.00 (40.00, 54.00) | 47.00 (40.00, 55.00) | 43.00 (36.00, 49.00) | < 0.001 |
| DBP max, M (Q1, Q3) | 88.00 (77.00, 101.00) | 88.00 (77.00, 101.00) | 84.00 (72.88, 98.00) | < 0.001 |
| MBP min, M (Q1, Q3) | 60.00 (52.00, 68.00) | 60.00 (53.00, 68.00) | 56.00 (49.00, 63.00) | < 0.001 |
| MBP max, M (Q1, Q3) | 104.00 (93.00, 117.00) | 104.00 (93.00, 117.00) | 101.00 (90.00, 116.00) | < 0.001 |
| RR min, M (Q1, Q3) | 12.00 (10.00, 15.00) | 12.00 (10.00, 15.00) | 13.00 (10.00, 15.00) | 0.001 |
| RR max, M (Q1, Q3) | 27.00 (24.00, 32.00) | 27.00 (24.00, 32.00) | 28.00 (24.00, 33.00) | < 0.001 |
| Temperature min, M (Q1, Q3) | 36.50 (36.17, 36.72) | 36.50 (36.22, 36.72) | 36.39 (35.83, 36.67) | < 0.001 |
| Temperature max, M (Q1, Q3) | 37.33 (37.00, 37.89) | 37.33 (37.00, 37.89) | 37.39 (37.00, 37.89) | 0.81 |
| SpO2 min, M (Q1, Q3) | 93.00 (90.00, 95.00) | 93.00 (90.00, 95.00) | 92.00 (89.00, 94.00) | < 0.001 |
| Mechanical ventilation, n (%) | < 0.001 | |||
| No | 8075 (48.86) | 7599 (50.63) | 476 (31.32) | |
| Yes | 8453 (51.14) | 7409 (49.37) | 1044 (68.68) | |
| CRRT, n (%) | < 0.001 | |||
| No | 15,381 (93.06) | 14,119 (94.08) | 1262 (83.03) | |
| Yes | 1147 (6.94) | 889 (5.92) | 258 (16.97) | |
| Vasopressors, n (%) | < 0.001 | |||
| No | 11,567 (69.98) | 10,785 (71.86) | 782 (51.45) | |
| Yes | 4961 (30.02) | 4223 (28.14) | 738 (48.55) | |
| Antibiotic, n (%) | < 0.001 | |||
| No | 6944 (42.01) | 6425 (42.81) | 519 (34.14) | |
| Yes | 9584 (57.99) | 8583 (57.19) | 1001 (65.86) | |
Z: Mann–Whitney test, χ2: Chi-square test
M: Median, Q1: 1st Quartile, Q3: 3st Quartile
NOAF New-onset atrial fibrillation, Los Length of stay, MI Myocardial infarction, CRRT Continuous renal replacement therapy, HFrEF Heart failure with reduced ejection fraction, HFpEF Heart failure with preserved ejection fraction, WBC White blood cell, BUN Blood urea nitrogen, HR Heart rate, RR Respiratory rate, SBP Systolic blood pressure, DBP Diastolic blood pressure, MBP Mean blood pressure, SpO2 Percutaneous arterial oxygen saturation
Feature selection
Lasso regression was used to screen the relevant features of the training set, and the characteristics of the variable coefficients were shown in Fig. 2A. The iterative analysis was performed using a tenfold cross-validation method. The 23 variables closely associated with NOAF were admission_age, race, weight, urine output, WBC_max, BUN_min, potassium_min, HR_min, HR_max, SBP_min, DBP_max, MBP_min, RR_min, temperature_min, temperature_max, SpO2_min, chronic_liver_disease, HFrEF, HFpEF, sepsis, mechanical_ventilation, CRRT, vasopressors.
Fig. 2.
Lasso regression-based variable screening. A. Variation characteristics of variable coefficients; B. The process of selecting the optimal value of the parameter λ in the lasso regression model is carried out by the cross-validation method
Model performance comparisons
We constructed eight ML models to identify the risk of NOAF in critically ill patients in the ICU. Figure 3 displayed the discriminative performance of eight models in terms of ROC curves. All eight models showed considerable prediction performance for new-onset AF, with the XGBoost model exhibiting best performance. The XGBoost model achieved an AUC of 0.891 [95% confidence interval (CI): 0.878–0.903], setting the benchmark for NOAF prediction. Following closely, the GBM model showed comparable efficacy with an AUC of 0.877 (95% CI: 0.864–0.891), outperforming the remaining algorithms. The remaining models, while still demonstrating good predictive power, ranked as follows in descending order of performance: Adaboost (AUC = 0.859, 95% CI: 0.845–0.873), NN (AUC = 0.825, 95% CI: 0.809–0.841), MLP (AUC = 0.807, 95% CI: 0.789–0.824), NB (AUC = 0.792, 95% CI: 0.775–0.810), SVM (AUC = 0.788, 95% CI: 0.770–0.806) and LR (AUC = 0.786, 95% CI: 0.769–0.804).
Fig. 3.

ROC curves for the machine learning models. XGBoost: extreme gradient boosting; SVM: support vector machine; Adaboost: adaptive boosting; MLP: multilayer perceptron; NN: neural network; NB: naive bayes; LR: logistic regression; GBM: gradient boosting machine; ROC: receiver operating characteristic; AUC: area under the curve
Table 2 showed detailed performance metrics for the eight models. The XGBoost model exhibited superior overall performance (sensitivity: 0.826, specificity: 0.775). Notably, the XGBoost achieved the highest F1 score (0.805) and accuracy (0.801), while also boasting the highest recall rate (0.826) among all models evaluated. The calibration curves for all eight models were illustrated in Fig. 4A, providing crucial insights into their predictive reliability. Six of the eight models, excluding NB and Adaboost models, demonstrated favorable consistency between predicted probabilities and observed outcomes.
Table 2.
Performances of the machine learning models for predicting NOAF
| Model | Sensitivity | Specificity | F1 score | Accuracy | Recall |
|---|---|---|---|---|---|
| Adaboost | 0.804 | 0.743 | 0.780 | 0.773 | 0.804 |
| GBM | 0.815 | 0.763 | 0.794 | 0.789 | 0.815 |
| LR | 0.740 | 0.696 | 0.724 | 0.718 | 0.740 |
| MLP | 0.839 | 0.653 | 0.767 | 0.746 | 0.839 |
| NB | 0.691 | 0.736 | 0.707 | 0.713 | 0.691 |
| NN | 0.766 | 0.728 | 0.751 | 0.747 | 0.767 |
| SVM | 0.738 | 0.700 | 0.724 | 0.719 | 0.738 |
| XGBoost | 0.826 | 0.775 | 0.805 | 0.801 | 0.826 |
NOAF New-onset atrial fibrillation, XGBoost: Extreme gradient boosting; SVM Support vector machine, Adaboost: Adaptive boosting, MLP Multilayer perceptron; NN Neural network; NB Naïve bayes, LR Logistic regression, GBM Gradient boosting machine
Fig. 4.
Calibration capability and clinical benefit of the model. A. Calibration curve B. Clinical Impact Curve (CIC) C. Decision curve analysis (DCA), XGBoost: extreme gradient boosting; SVM: support vector machine; Adaboost: adaptive boosting; MLP: multilayer perceptron; NN: neural network; NB: naive bayes; LR: logistic regression; GBM: gradient boosting machine.
In terms of clinical applicability, except Adaboost, each model showed robust net benefit across a wide range of threshold probabilities, with the XGBoost model exhibited the highest net benefit and therefore selected as the optimal model for predicting NOAF (Fig. 4C). To further elucidate the model’s performance, we plotted the CIC for the XGBoost model (Fig. 4B), the x-axis showed different risk thresholds and their corresponding cost–benefit ratios, while the y-axis illustrated the number of positive patients identified by the model versus the actual true positives in a sample of 1000 individuals. This visualization revealed that as the threshold increased, the number of positive patients identified by the model converged towards the actual number of true positives. However, this convergence was accompanied by an escalation in the cost–benefit ratio associated with false positives. After careful consideration of these trade-offs, we established 0.6 as the optimal threshold for defining high-risk NOAF. This judicious selection stroked a balance between two critical factors: it mitigated the risk of excessive false-positive identifications that could result from an overly low threshold, while simultaneously avoiding the substantial losses associated with false-positive patients that might occur with an excessively high threshold.
External validation
Despite the inherent differences in baseline characteristics between the two datasets, our model demonstrated robust generalizability. The externally validated ROC curve yielded an AUC of 0.769 (95% CI: 0.755–0.782), as illustrated in Fig. S2.
Interpretability analysis
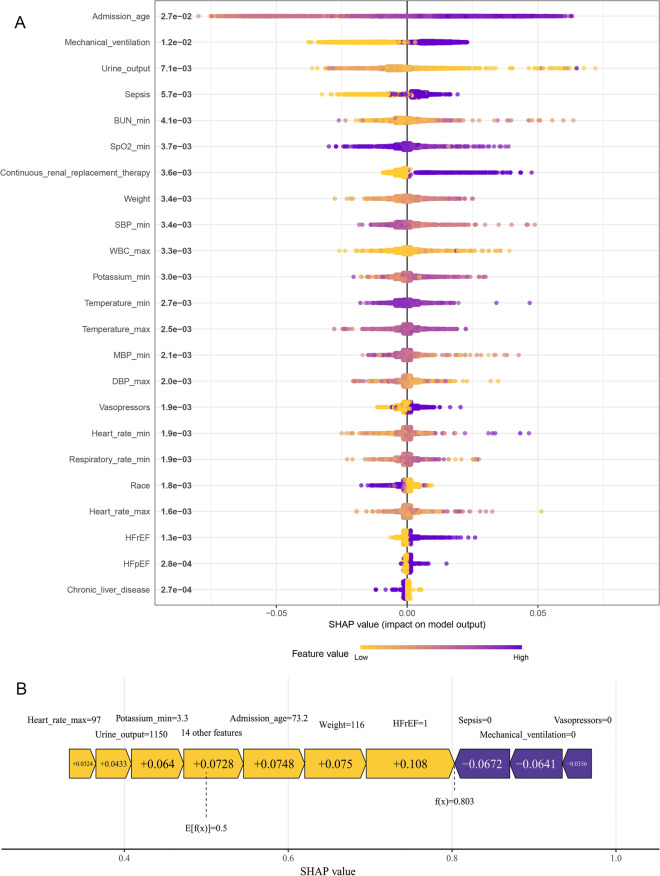
Figure 5A presented a comprehensive swarm plot illustrating the variables in the XGBoost model. The horizontal axis represented SHAP values, while the vertical axis displayed features sorted by their cumulative SHAP value impact. Each data pointed corresponds to a specific instance, with its position along the x-axis indicating the SHAP value for that particular instance and feature. Age, mechanical ventilation, urine output, sepsis, BUN, SpO2, CRRT, and weight emerged as the eight most important factors in predicting NOAF. Figure 5B offered a detailed case study, demonstrating the model’s prediction process for a specific patient. In this visualization, yellow indicators signified positive contributions to the prediction, while violet denoted negative influences. The f(x) value represented the actual SHAP value for each factor. Notably, for this particular patient, our XGBoost model predicted a higher risk of NOAF compared to the baseline. The key factors driving this prediction, as determined by their SHAP values, were HFrEF, sepsis, weight and age.
Fig. 5.
Visually interpret machine learning models using SHAP. A SHAP summary point. B SHAP force plot. SBP: systolic blood pressure; BUN: blood urea nitrogen; SpO2: percutaneous arterial oxygen saturation; WBC: white blood cell; MBP: mean blood pressure; DBP: diastolic blood pressure; HFrEF: heart failure with reduced ejection fraction; HFpEF: heart failure with preserved ejection fraction
Application of model
To enhance the clinical applicability of our model and facilitate rapid decision-making by clinicians, we used feature recursive elimination for a refined selection of variables (Fig. S3A). This optimization process allowed us to maximize model performance (ROC: 0.832) while streamlining the input to just 7 key variables (age, weight, mechanical ventilation, CRRT, vasopressors, HFrEF). To further improve accessibility and utility, we have deployed the optimized model on a dedicated website (https://7kdtqk-guanchengcheng.shinyapps.io/noaf3/). This user-friendly platform enabled clinicians to input a patient’s first-day metrics and promptly assess their risk of NOAF. Moreover, the tool provided a detailed breakdown of how each characteristic contributed to the overall risk assessment, offering valuable insights into the factors driving the prediction.
Discussion
We conducted a study to predict the risk of new-onset AF in critically ill patients. Eight ML algorithms were employed to construct predictive models by screening 23 clinical variables within the first 24 h of ICU admission. The results demonstrated that the XGBoost algorithm exhibited strong performance, with discrimination and calibration, and showed a substantial net benefit in clinical practice. The findings from the external validation cohort further confirmed the stability and accuracy of the model. To gain deeper insights into the model, we utilized the SHAP method for visualization. The colony plot analysis revealed that eight characteristics, namely age, mechanical ventilation, urine output, sepsis, BUN, SpO2, CRRT, and weight, had the most significant influence on the prediction of the XGBoost model. The SHAP force plot aimed to further elucidate the process of individualized AF risk prediction by the model, enabling us to comprehensively understand its underlying mechanism.
Current guidelines and studies rarely address the management of NOAF in the ICU setting separately. The available evidence is primarily derived from observational studies and expert consensus, lacking uniform treatment principles for NOAF [23]. Moreover, even after successful restoration of sinus rhythm through treatment, patients remain at a high risk of recurrence [24]. Consequently, early detection and intervention are crucial for patients at high risk of developing AF.
Attempts to build predictive models for AF risk have been made many years ago, and a retrospective study extracted data from ICU patients for the first eight hours and modelled them using SVM algorithms with an AUC of 0.73 [25]. In addition, some researchers used traditional logistic regression modelling to obtain a general AUC (0.836) [26], but both studies lacked external validation and interpretation of the model. Jarne Verhaeghe et al. constructed three CatBoost models for prediction, obtained AUC values of 0.81 [27], and interpreted the model using the SHAP method, but did not perform clinical transformation, which limits its application. Several other studies have used ECGs to construct models, but as mentioned above, short-term prediction did not give clinicians much time to process the situation, and clinical applications were limited.
The XGBoost algorithm is optimized based on gradient boosting decision trees, particularly in large datasets and complex feature spaces. In recent years, prediction models based on XGBoost have been widely used in the medical field, showing favorable performance in various areas such as septicemia, cardiovascular diseases, and kidney injury [28–30]. Compared to the traditional logistic regression algorithm, the XGBoost model can effectively capture non-linear relationships and build the final model by integrating several weak classifiers, resulting in better generalization ability. Moreover, the XGBoost model is robust to outliers and noisy data, further reducing noise in the dataset.
SHAP, as a method of interpreting ML model prediction, can help to understand the prediction process and the contribution of features of the model to some extent. Age had been considered the most critical factor in the development of AF, consistent with previous studies [31]. The structure of the atrium and the electrophysiological changes that occur with age made the conduction slow and the low voltage diffuse in some areas [32]. In addition, frailty and comorbidities associated with aging combine to reduce the body’s reserves for coping with stressful events and increase vulnerability to adverse outcomes such as oxidative stress, inflammation, interventions, and medications [33]. Renal insufficiency, manifested by decreased urine output, elevated BUN and the need for CRRT use, emerged as an important predictor of NOAF. AF has been observed to be prevalent among patients with chronic renal insufficiency [34], and in intensive care settings, critically ill patients with acute kidney injury (AKI), especially those receiving CRRT, have a higher incidence of NOAF [35], which may be attributed to intravascular volume depletion during RRT and electrolyte disturbances [36]. We observed that both HFrEF and HFpEF were risk factors for NOAF, and despite their distinct pathophysiological profiles, both forms of heart failure substantially elevated the risk of NOAF through complex mechanisms interacting with atrial structural remodeling, mitral regurgitation, and neurohumoral alterations [37]. Obese patients were often accompanied by epicardial fat deposition, systemic inflammation and elevated levels of oxidative stress, which promote the abnormality of atrial structure and electrophysiological function, and then induce AF [38]. Unfortunately, body mass index (BMI) data were not available due to the severe lack of height, but we still need to pay attention to the potential impact of overweight and obesity on AF. Interestingly, we observed that hypertension was excluded at variable screening, although it was recognized as a risk factor for AF, whereas lower SBP was associated with NOAF in the ICU. Consistent with our study, a retrospective study in elderly hemodialysis patients confirmed that lower pre-dialysis SBP was associated with a higher incidence of AF [39]. Another study, which identified single nucleotide polymorphism (SNP) loci associated with NOAF in intensive care patients, confirmed that genetic factors (SNPs) associated with ambulatory AF may play a small role in the development of new AF in critically ill patients in ICUs. Conversely, acute environmental factors and physiological stressors may be more important factors in the development of NOAF in critically ill patients [1, 40]. We suggest that hypotension implied an underlying circulatory disorder in the patient, by inducing a range of acute physiological stress responses and promoting the occurrence of NOAF in the ICU, a mechanism different from chronic atrial remodeling caused by hypertension. This study revealed a higher prevalence of sepsis and increased requirements for mechanical ventilation and vasopressor support among patients with NOAF. These findings were consistent with previous research and highlight the significant impact of acute illness factors, particularly sepsis, on patient outcomes. Notably, these acute factors appeared to exert a more profound influence on patient’s prognosis than the traditional cardiovascular comorbidities typically associated with AF [41, 42].
Our model had also achieved predictive performance in external validation, suggesting that our model has good generalizability. In addition, because the variables we had screened were commonly used clinical indicators, they were easy to measure and therefore easy to promote in hospitals at all levels. We had also designed a clinician-friendly interface that allows clinicians to easily predict AF risk and allocate resources based on predicted outcomes.
There were also limitations to our study. Primarily, the retrospective design introduced potential information bias, including data collection errors and missing data. While NOAF reported by bedside nurses demonstrated high accuracy, the possibility of omissions and false positives cannot be entirely excluded. Furthermore, due to the limitations inherent in the database, some certain potentially crucial indicators, including height, CK_MB and NT-proBNP, were missing, which might have inadvertently led to the omission of some key variables in our analysis, potentially impacting the comprehensiveness of our predictive model. Finally, although we used the MIMIC-III subset as an external validation, the data were from a single center and further large-scale prospective studies are needed to validate the accuracy of our model and its generalization to other populations. Nevertheless, we still believe that our model can help critical care physicians identify patients at high risk for AF in a timelier manner.
Conclusion
We developed a ML model to predict the risk of NOAF in critically ill patients without cardiac surgery and validated its potential as a clinically reliable tool. SHAP improves the interpretability of the model, enables clinicians to better understand the causes of NOAF, helped clinicians to prevent in advance and improves patient outcome.
Supplementary Information
Acknowledgements
We thank all the administrators who collected, organized and maintained the MIMIC database.
Author contributions
Bing Xiao contributed to the research design. Chengjian Guan, Angwei Gong and Yan Zhao contributed to data collection, data processing and graphing. Chen Yin, Lu Geng, Linli Liu, Xiuchun Yang contributed data proofreading and formal analysis. Chengjian Guan contributed to the writing of the manuscript. Xiao Bing and Jingchao Lu contributed to review and to edit. All authors have read and approved the final manuscript.
Funding
The project was supported by the S&T Program of Hebei No. 22377728D.
Availability of data and materials
The data for this study came from MIMIC database. Researchers need to request MIMIC data, so we can’t make it public. However, the corresponding author of this paper (Bing Xiao) may provide data to the researcher upon reasonable request after the researcher has applied through the MIMIC database.
Declarations
Ethics approval and consent to participate
The study was carried out in accordance with the Declaration of Helsinki. Due to the de-identification of the MIMIC repository, sensitive data is not involved, so we informed the Ethics Committee of this situation without a written report.
Informed consent
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Jingchao Lu, Email: ljchb2h@hebmu.edu.cn.
Bing Xiao, Email: xiaobing@hebmu.edu.cn.
References
- 1.Bosch NA, Cimini J, Walkey AJ. Atrial Fibrillation in the ICU. Chest. 2018;154(6):1424–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Wetterslev M, Haase N, Hassager C, et al. New-onset atrial fibrillation in adult critically ill patients: a scoping review. Intensive Care Med. 2019;45(7):928–38. [DOI] [PubMed] [Google Scholar]
- 3.Walkey AJ, Ambrus D, Benjamin EJ. The role of arrhythmias in defining cardiac dysfunction during sepsis. Am J Respir Crit Care Med. 2013;188(6):751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kim K, Yang PS, Jang E, et al. Long-term impact of newly diagnosed atrial fibrillation during critical care: a south korean nationwide cohort study. Chest. 2019;156(3):518–28. [DOI] [PubMed] [Google Scholar]
- 5.Bedford JP, Ferrando-Vivas P, Redfern O, et al. New-onset atrial fibrillation in intensive care: epidemiology and outcomes. Eur Heart J Acute Cardiovasc Care. 2022;11(8):620–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Burrage PS, Low YH, Campbell NG, et al. New-onset atrial fibrillation in adult patients after cardiac surgery. Curr Anesthesiol Rep. 2019;9(2):174–93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gaudino M, Di Franco A, Rong LQ, et al. Postoperative atrial fibrillation: from mechanisms to treatment. Eur Heart J. 2023;44(12):1020–39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Launey Y, Lasocki S, Asehnoune K, et al. Impact of low-dose hydrocortisone on the incidence of atrial fibrillation in patients with septic shock: a propensity score-inverse probability of treatment weighting cohort study. J Intensive Care Med. 2019;34(3):238–44. [DOI] [PubMed] [Google Scholar]
- 9.Wilson MG, Rashan A, Klapaukh R, et al. Clinician preference instrumental variable analysis of the effectiveness of magnesium supplementation for atrial fibrillation prophylaxis in critical care. Sci Rep. 2022;12(1):17433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wetterslev M, Hylander Møller M, Granholm A, et al. Atrial fibrillation (AFIB) in the ICU: incidence, risk factors, and outcomes: the international AFIB-ICU cohort study. Crit Care Med. 2023;51(9):1124–37. [DOI] [PubMed] [Google Scholar]
- 11.Walkey AJ, Benjamin EJ, Lubitz SA. New-onset atrial fibrillation during hospitalization. J Am Coll Cardiol. 2014;64(22):2432–3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Swanson K, Wu E, Zhang A, et al. From patterns to patients: advances in clinical machine learning for cancer diagnosis, prognosis, and treatment. Cell. 2023;186(8):1772–91. [DOI] [PubMed] [Google Scholar]
- 13.Ambale-Venkatesh B, Yang X, Wu CO, et al. Cardiovascular event prediction by machine learning: the multi-ethnic study of atherosclerosis. Circ Res. 2017;121(9):1092–101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lu Y, Chen Q, Zhang H, et al. Machine learning models of postoperative atrial fibrillation prediction after cardiac surgery. J Cardiothorac Vasc Anesth. 2023;37(3):360–6. [DOI] [PubMed] [Google Scholar]
- 15.Bashar SK, Hossain MB, Ding E, et al. Atrial fibrillation detection during sepsis: study on MIMIC III ICU data. IEEE J Biomed Health Inform. 2020;24(11):3124–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cabitza F, Rasoini R, Gensini GF. Unintended consequences of machine learning in medicine. JAMA. 2017;318(6):517–8. [DOI] [PubMed] [Google Scholar]
- 17.Lundberg SM, Erion G, Chen H, et al. From local explanations to global understanding with explainable AI for trees. Nat Mach Intell. 2020;2(1):56–67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Johnson AEW, Bulgarelli L, Shen L, et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci Data. 2023;10(1):1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Goldberger AL, Amaral LA, Glass L, et al. PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation. 2000;101(23):E215–20. [DOI] [PubMed] [Google Scholar]
- 20.Johnson AE, Pollard TJ, Shen L, et al. MIMIC-III, a freely accessible critical care database. Sci Data. 2016;3: 160035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ding EY, Albuquerque D, Winter M, et al. Novel method of atrial fibrillation case identification and burden estimation using the MIMIC-III electronic health data set. J Intensive Care Med. 2019;34(10):851–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kerr KF, Brown MD, Zhu K, et al. Assessing the clinical impact of risk prediction models with decision curves: guidance for correct interpretation and appropriate use. J Clin Oncol. 2016;34(21):2534–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Walkey AJ, Hogarth DK, Lip GYH. Optimizing atrial fibrillation management: from ICU and beyond. Chest. 2015;148(4):859–64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lancini D, Tan WL, Guppy-Coles K, et al. Critical illness associated new onset atrial fibrillation: subsequent atrial fibrillation diagnoses and other adverse outcomes. Europace. 2023;25(2):300–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.McMillan S, Rubinfeld I, Syed Z (2012) Predicting atrial fibrillation from intensive care unit numeric data. In: 2012 Computing in cardiology
- 26.Ortega-Martorell S, Pieroni M, Johnston BW, et al. Development of a risk prediction model for new episodes of atrial fibrillation in medical-surgical critically Ill patients using the AmsterdamUMCdb. Front Cardiovasc Med. 2022;9: 897709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Verhaeghe J, De Corte T, Sauer CM, et al. Generalizable calibrated machine learning models for real-time atrial fibrillation risk prediction in ICU patients. Int J Med Inform. 2023;175: 105086. [DOI] [PubMed] [Google Scholar]
- 28.Hou N, Li M, He L, et al. Predicting 30-days mortality for MIMIC-III patients with sepsis-3: a machine learning approach using XGboost. J Transl Med. 2020;18(1):462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Li J, Liu S, Hu Y, et al. Predicting mortality in intensive care unit patients with heart failure using an interpretable machine learning model: retrospective cohort study. J Med Internet Res. 2022;24(8): e38082. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhang Z, Ho KM, Hong Y. Machine learning for the prediction of volume responsiveness in patients with oliguric acute kidney injury in critical care. Crit Care. 2019;23(1):112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Schnabel RB, Yin X, Gona P, et al. 50 year trends in atrial fibrillation prevalence, incidence, risk factors, and mortality in the Framingham Heart Study: a cohort study. Lancet. 2015;386(9989):154–62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kistler PM, Sanders P, Fynn SP, et al. Electrophysiologic and electroanatomic changes in the human atrium associated with age. J Am Coll Cardiol. 2004;44(1):109–16. [DOI] [PubMed] [Google Scholar]
- 33.Brunker LB, Boncyk CS, Rengel KF, et al. Elderly patients and management in intensive care units (ICU): clinical challenges. Clin Interv Aging. 2023;18:93–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Ding WY, Gupta D, Wong CF, et al. Pathophysiology of atrial fibrillation and chronic kidney disease. Cardiovasc Res. 2021;117(4):1046–59. [DOI] [PubMed] [Google Scholar]
- 35.Hellman T, Uusalo P, Järvisalo MJ. New-onset atrial fibrillation in critically ill acute kidney injury patients on renal replacement therapy. Europace. 2022;24(2):211–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Buiten MS, de Bie MK, Rotmans JI, et al. The dialysis procedure as a trigger for atrial fibrillation: new insights in the development of atrial fibrillation in dialysis patients. Heart. 2014;100(9):685–90. [DOI] [PubMed] [Google Scholar]
- 37.Verhaert DVM, Brunner-La Rocca HP, van Veldhuisen DJ, et al. The bidirectional interaction between atrial fibrillation and heart failure: consequences for the management of both diseases. Europace. 2021;23(23 Suppl 2):ii40–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Shu H, Cheng J, Li N, et al. Obesity and atrial fibrillation: a narrative review from arrhythmogenic mechanisms to clinical significance. Cardiovasc Diabetol. 2023;22(1):192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chang TI, Liu S, Airy M, et al. Blood pressure and incident atrial fibrillation in older patients initiating hemodialysis. Clin J Am Soc Nephrol. 2019;14(7):1029–38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kerchberger VE, Huang Y, Koyama T, et al. Clinical and genetic contributors to new-onset atrial fibrillation in critically Ill adults. Crit Care Med. 2020;48(1):22–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Walkey AJ, Greiner MA, Heckbert SR, et al. Atrial fibrillation among Medicare beneficiaries hospitalized with sepsis: incidence and risk factors. Am Heart J. 2013;165(6):949-955.e3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bedford JP, Harford M, Petrinic T, et al. Risk factors for new-onset atrial fibrillation on the general adult ICU: a systematic review. J Crit Care. 2019;53:169–75. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The data for this study came from MIMIC database. Researchers need to request MIMIC data, so we can’t make it public. However, the corresponding author of this paper (Bing Xiao) may provide data to the researcher upon reasonable request after the researcher has applied through the MIMIC database.