Abstract
Background
Schizophrenia (SCZ) is a severe psychiatric disorder associated with alterations in early brain development. Details of underlying pathomechanisms remain unclear, despite genome and transcriptome studies providing evidence for aberrant cellular phenotypes and pathway deregulation in developing neuronal cells. However, mechanistic insight at the protein level is limited.
Methods
Here, we investigate SCZ-specific protein expression signatures of neuronal progenitor cells (NPC) derived from patient iPSC in comparison to healthy controls using high-throughput Western Blotting (DigiWest) in a targeted proteomics approach.
Results
SCZ neural progenitors displayed altered expression and phosphorylation patterns related to Wnt and MAPK signaling, protein synthesis, cell cycle regulation and DNA damage response. Consistent with impaired cell cycle control, SCZ NPCs also showed accumulation in the G2/M cell phase and reduced differentiation capacity. Furthermore, we correlated these findings with elevated p53 expression and phosphorylation levels in SCZ patient-derived cells, indicating a potential implication of p53 in hampering cell cycle progression and efficient neurodevelopment in SCZ.
Conclusions
Through targeted proteomics we demonstrate that SCZ NPC display coherent mechanistic alterations in regulation of DNA damage response, cell cycle control and p53 expression. These findings highlight the suitability of iPSC-based approaches for modeling psychiatric disorders and contribute to a better understanding of the disease mechanisms underlying SCZ, particularly during early development.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12888-024-06127-x.
Keywords: Proteomics, DigiWest, IPSC, Neural progenitors, Schizophrenia, Cellular signaling, Cell cycle, p53
Background
Schizophrenia spectrum disorders (SCZ) are a group of severe neurodevelopmental disorders with a highly heterogeneous clinical presentation of a broad range of symptoms including hallucinations, delusions, emotional blunting and cognitive deficits [1]. Current antipsychotic treatment options for SCZ are purely symptomatic and show only limited efficacy in alleviating negative and cognitive symptoms [2, 3]. Hence, there is an urgent need to develop causal treatments, which has so far been hampered by the still limited knowledge of the molecular mechanisms that are involved in the disease. As SCZ is regarded as a neurodevelopmental disorder, alterations occurring during early brain development are of particular interest. Genome-wide association and exome sequencing studies have linked > 100 risk loci to SCZ but information on the protein level remains limited [4, 5].
Induced pluripotent stem cells (iPSC) have emerged as a valuable source for phenotypic studies of disease-relevant cell types with a patient-specific genetic background. Transcriptomic analysis and shotgun proteomics of iPSC-derived neural progenitor cells (NPC) and neurons have previously helped to gain insight into potential disease-related processes. Most studies investigating the proteome in SCZ employ mass spectrometry approaches [6–8]. Deregulated expression of genes and proteins involved in protein synthesis, cell adhesion, regulation of the cytoskeleton, oxidative stress and neuronal differentiation have been identified in SCZ [8–11]. Additionally, deregulations in key signaling pathways have been described, among which the most prominent changes have been reported for proteins of the Wnt and Akt/ glycogen synthase kinase-3 beta (GSK3 beta) signaling pathways in both iPSC and patient studies [10, 12–16]. Postmortem studies have also linked the mitogen-activated protein kinase (MAPK)/ extracellular signal-regulated kinase (ERK) pathway to SCZ [17–19]. Despite the important insights that have been gained by these studies, many important questions regarding the connection between aberrant signaling and cellular (patho)-phenotypes remain unanswered. Moreover, with most studies focusing on the transcriptome level, proteomic studies remain underrepresented in the field. However, they are of great importance given that gene expression changes do not always translate to the protein level. Especially the study of phosphorylated proteins remains difficult using high throughput proteomic approaches but is of great importance for entangling mechanisms surrounding pathway activity. Applying such techniques to relevant neuronal cell types would greatly expand the knowledge of the biological processes at play during the early phases of disease development.
Here, we employ high-throughput Western Blotting (DigiWest) to study protein expression and phosphorylation of 133 proteins in SCZ patient-derived cells on the iPSC and NPC level in a targeted proteomics approach. The DigiWest is a multiplexed Western Blot derivative which transfers the Western Blot onto a bead-based system, thereby greatly increasing throughput while retaining the sensitivity of traditional Western Blotting. In this fashion, expression data from up to 200 proteins and phosphoproteins can be obtained in a targeted manner, allowing extensive analysis of cellular signaling pathways [20]. Applying this method, we aimed to compare differences in protein expression patterns between control (CTR) and SCZ in iPSC and NPC, respectively, to identify potential disease-relevant alterations in cellular signaling.
We report several aberrant SCZ-specific protein signatures, exclusive to NPC, with regards to signaling pathway activity. Most notably, we found proteomic alterations in cell cycle control, DNA-damage response regulation along with impaired differentiation capacity of SCZ NPC. Furthermore, we were able to correlate these alterations to p53 expression and phosphorylation levels in SCZ-derived cells, which provides further mechanistic insights into the early developmental stages of SCZ.
Methods
iPSC line information and maintenance
For detailed information on the three control and four patient-derived iPSC lines used see Additional file 1—Table S1. The CTR1 iPSC line was purchased from Thermo Fisher Scientific (Waltham, MA, USA—#A18945). The CTR2 iPSC line was a gift from the Tumorbiology group at NMI Reutlingen. For clinical patient data, see [21]. iPSC were maintained on plates coated with hESC-qualified Matrigel (Corning, Corning, NY, USA) in mTeSR Plus (STEMCELL Technologies, Vancouver, Canada). For single cell seeding, iPSC were enzymatically passaged using Accutase (Sigma-Aldrich, St. Louis, MO, USA).
NPC generation and maintenance
NPC were generated from iPSC following the embyroid body (EB) protocol of the STEMdiff™ SMADi Neural Induction Kit (STEMCELL Technologies) according to the manufacturer's instructions. The obtained NPC were cultured in STEMdiff™ Neural Progenitor medium (STEMCELL Technologies) on 6-well plates coated with 20% poly-l-ornithine (Sigma-Aldrich) for 2h at room temperature (RT) and 10 µg/ml Laminin (Sigma-Aldrich) at 37 °C overnight. NPC were enzymatically passaged with Accutase. For all experiments, only passage 2 NPC from three independent rounds of differentiation were used to ensure comparability of results.
DigiWest protein profiling
iPSC were obtained in triplicate and NPC samples from three independent differentiations. DigiWest was performed as published [20] using 12 µg of cellular protein. In brief, the NuPAGE system (Life Technologies) was used for gel electrophoresis and blotting onto PVDF membranes. Proteins were biotinylated on the membrane using NHS-PEG12-Biotin (50 µM) in PBST for 1 h. Sample lanes were cut into 96 strips (0.5 mm each) and placed in one well of a 96-well plate before adding 10 µl elution buffer (8 M urea, 1% Triton-X100 in 100 mM Tris–HCl pH 9.5). Each strip/protein fraction was incubated with 1 distinct Neutravidin-coated MagPlex bead population (Luminex, Austin, TX, USA). Coupling was performed overnight, and non-bound binding sites were blocked with 500 µM deactivated NHS-PEG12-Biotin for 1 h. By pooling all 96 protein-loaded bead populations, the original sample lane was reconstituted.
5 µl aliquots of bead mix were added to 96-well plates containing 50 µl assay buffer (Blocking Reagent for ELISA (Roche) supplemented with 0.2% milk powder, 0.05% Tween-20 and 0.02% sodium azide). Upon discarding of the assay buffer, 30 µl of primary antibody (diluted in assay buffer) was added per well. After overnight incubation at 15 °C, the bead-mixes were washed twice with PBST and species-specific PE-labelled (Phycoerythrin) secondary antibodies (Dianova, Hamburg, Germany) were added for 1 h at 23 °C. Beads were washed twice with PBST before readout on a Luminex FlexMAP 3D instrument.
One hundred thirty-seven primary antibodies (Additional file 2) were selected from a collection of > 1 500 available antibodies, all of which are performance-evaluated and routinely used in DigiWest. Selection was largely based on covering signaling pathways/cellular functions for which associations with SCZ have been described in previous genetic/transcriptomic/proteomic studies as mentioned elsewhere in this manuscript Refs [8–19, 22, 23]. Likewise, to accurately monitor pathway activity during neuronal differentiation, signaling proteins from neurodevelopmental pathways (Wnt [24], Hippo [25], Hedgehog [26], Smad [27] signaling) were additionally selected along with other common signaling and marker proteins. Pathway allocation of analytes was mapped based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [28, 29].
For peak integration, an Excel-based analysis tool was used. A total of 148 peaks were identified, with 133 (89.9%) generating reliable signals. Signal intensity was separately normalized to total protein amount on the beads within each differentiation. The software package MeV 4.9.0 was used for heatmap generation and statistical analysis [30]. For heatmaps, fluorescent signals were either median centered across samples for a given analyte (iPSC versus NPC) or centered around the average signal of the three CTR lines within each differentiation (CTR versus SCZ) before Log2 transformation. Raw and normalized DigiWest data can be found in Additional file 3 and 4, respectively.
Immunocytochemistry
For immunocytochemical staining, iPSC or passage-matched NPC were seeded on 96-well µClear™ plates (Greiner, Kremsmünster, Austria). Once 70–80% confluence was reached, cells were fixed with 4% PFA in PBS for 15 min at RT. Cells were washed 3 × with PBS and incubated with PBS + 1 × BMB blocking reagent (Roche) + 0.1% Triton (Carl Roth, Karlsruhe, Germany) for 30 min at RT. Primary antibodies were diluted in blocking/permeabilization solution and incubated overnight at 4 °C. Afterwards, cells were washed 3 × with PBS. Secondary antibodies were incubated in blocking/permeabilization solution for 2h at RT on an orbital shaker and washed with PBS. For nuclear staining, Hoechst 33,258 (Sigma-Aldrich) was diluted in PBS and incubated for 30 min at RT. A complete list of ICC antibodies used in this study can be found in Additional file 1 – Table S2.
Image acquisition and analysis of immunocytochemical staining
Z-Stack images were acquired from four sites per well with the ImageXpress Micro Confocal High-Content Imaging System (Molecular Devices, San Jose, CA, USA). To ensure unbiased image acquisition and analysis, all acquisition parameters for individual antibodies were kept constant within replicates. For image analysis, the software MetaXpress (Molecular Devices) was used. 2D projections were generated and intensity/size thresholding (for nuclear and spot-like staining patterns) was applied to generate a mask covering the fluorescent signal for each marker. Depending on the marker expression pattern (nuclear vs. cytosolic), different parameters were used for analysis. For nuclear stains and stainings in which cells/nuclei were either positive or negative for a marker, the mean stained area was analyzed per image. For stainings, which were present in all cells but varied in intensity between different cells, the mean intensity per image was analyzed. In any case, values from one well were averaged and normalized to Hoechst signal to account for differences in cell density. Data were centered around the mean signal across either all iPSC (Figs. 1C, 2C) or all CTR clones (Figs. 3E, 4H, 6B). Data was obtained from three biological replicates.
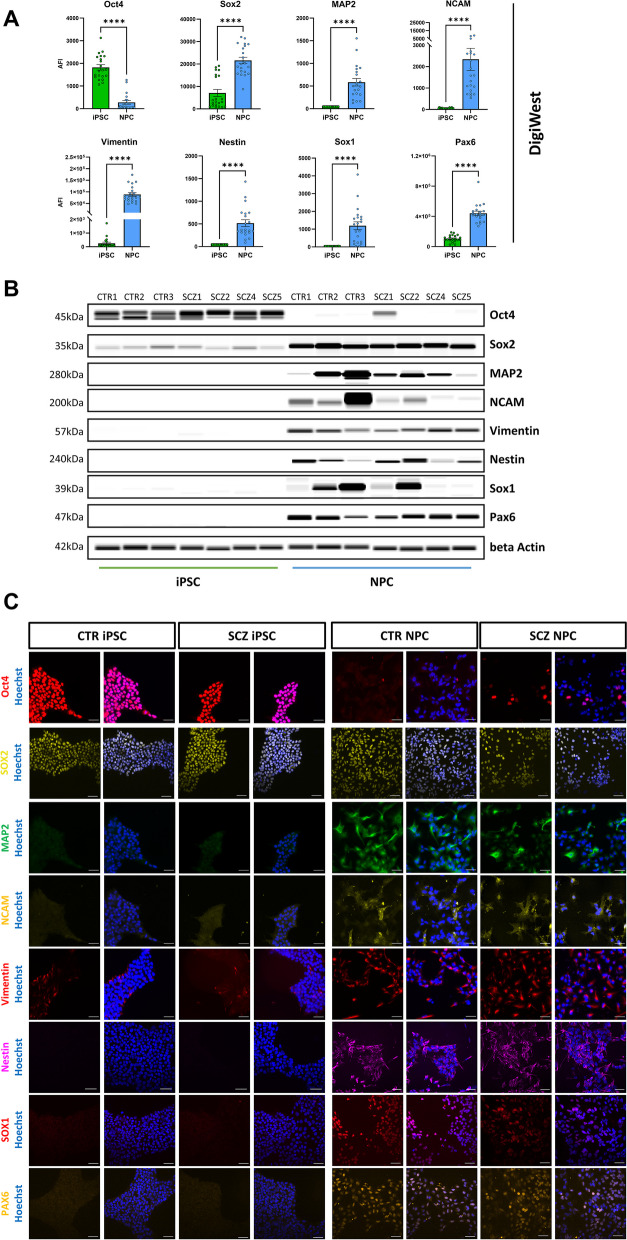
Fig. 1.
Marker expression in iPSC and NPC. A: DigiWest data (AFI = accumulated fluorescent intensity) of cell-type specific marker proteins in iPSC (n = 21) and NPC (n = 21); Mann–Whitney test. B: Western Blot mimic (gray-scale image) of markers displayed in A (exemplarily shown for one differentiation only). C: Example ICC images of marker expression in iPSC (left) and NPC (right) obtained by high-content microscopy. Scale bars: 50 µm. CTR and SCZ cells will be addressed separately at a later stage. *p < 0.05, ****p < 0.0001. Error bars: S.E.M
Fig. 2.
Expression signatures and pathway upregulation during differentiation. A: Heatmap and Hierarchical Cluster analysis of analytes significantly different between iPSC (n = 21) and NPC (n = 21) samples (Wilcoxon test, p < 0.001). B: DigiWest data (AFI = accumulated fluorescent intensity) for a subset of proteins with differential expression in iPSC (n = 21) and NPC (n = 21). Proteins are grouped according to their pathway allocation; Mann–Whitney test. C: Example ICC images of beta-catenin, LEF1 and p21 expression in iPSC (top) and NPC (bottom). Scale bars: 50 µm. D: Quantified ICC signals of proteins exemplarily shown in C (iPSC n = 21, NPC n = 21) obtained by high-content microscopy. Data are shown relative to mean iPSC signal; Mann–Whitney test. E–F: Volcano plot of separate iPSC vs NPC comparison for CTR (E—iPSC n = 9, NPC n = 9) and SCZ (F—iPSC n = 12, NPC n = 12) samples (Wilcoxon-Test, p < 0.01). Significantly upregulated proteins are shown in red, downregulated proteins in blue (analytes with FCs < I1I are excluded). Analytes with a significant interaction effect between cell type and disease allocation (p < 0.05, 2-Way-ANOVA) are highlighted (also see Additional File 5 – Figure S6 and Additional File 1—Table S3). *p < 0.05, ***p < 0.001, ****p < 0.0001. Error bars: S.E.M
Fig. 3.
SCZ-specific alterations in iPSC. A: Heatmap with Hierarchical Cluster analysis (HCL) of analytes significantly different (Wilcoxon Test, p < 0.05) between CTR (n = 9) and SCZ (n = 12) iPSC. Log2-transformed data is shown relative to mean signal across CTR lines of the respective differentiation. B: Volcano plot of comparison shown in A. C: DigiWest data (relative to control mean) of p53 and p53 – pS15 expression comparing CTR (n = 9) and SCZ (n = 12) iPSC; Mann–Whitney test. D: Example images of total p53 ICC staining in iPSC obtained by high-content microscopy. Scale bars: 50 µm. E: Quantified ICC signal of total p53 expression (relative to CTR mean) in CTR (n = 9) and SCZ (n = 12) iPSC (unpaired t-test). *p < 0.05, ***p < 0.001. Error bars: S.E.M
Fig. 4.
SCZ-specific alterations in NPC. A: Heatmap with Hierarchical Cluster analysis (HCL) of analytes significantly different (Wilcoxon Test, p < 0.05) between CTR (n = 9) and SCZ (n = 12) NPC. Log2-transformed data is shown relative to mean signal across CTR lines of the respective differentiation. B: Volcano plot of comparison shown in A. C: DigiWest data (relative to CTR mean) of Oct4, MAP2, NCAM and Sox1 expression in CTR (n = 9) and SCZ (n = 12) iPSC and NPC, respectively; Mann–Whitney test. D: Venn diagram showing the number of analytes differentially regulated between SCZ and CTR in the respective cell type. E: DigiWest data (relative to CTR mean) of p53 – pS15 and p53 (total) expression; CTR n = 9, SCZ n = 12, Mann–Whitney test. F: Quantified Western Blot signals of p53 – pS15 and p53 (total) expression in NPC (relative to CTR mean). Intensities were normalized to beta-Actin signal; CTR n = 6, SCZ n = 8, Mann–Whitney test. G: Western Blot images corresponding to quantification shown in F. H: Quantified ICC signal of total p53 expression (relative to CTR mean) in CTR (n = 9) and SCZ (n = 12) NPC (unpaired t-test). I: example images of total p53 ICC staining as obtained by high-content microscopy. Scale bars: 50 µm. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001. Error bars: S.E.M
Fig. 6.
Phenotypic cell cycle alterations in SCZ NPC correlate with p53 levels. A: Example images of G2/M regulatory proteins Aurora A and Cyclin B1 ICC staining in CTR and SCZ iPSC and NPC obtained by high-content microscopy. Scale bars: 50 µm. B: Quantified ICC signal of Aurora A and Cyclin B1 expression (relative to CTR mean) in CTR (n = 9) and SCZ (n = 12) in iPSC and NPC, respectively (Mann–Whitney test). C: Cell phase distribution of CTR and SCZ NPC (n = 5–8) in percent; 2-Way ANOVA with Tukey’s multiple comparisons test. D: Flow-cytometry cell cycle analysis of CTR and SCZ NPC. Nocodazole was used as a positive control. **p < 0.01 or as indicated. Error bars: S.E.M. E–G: Correlations between p53 – pS15 and E: p53 (total), F: differentiation markers and G: G2/M cell phase regulators. Spearman´s r; **p < 0.01, ***p < 0.001, ****p < 0.0001. The dashed red line indicates a simple linear regression applied to each XY correlation
Flow cytometry cell cycle analysis
Passage-matched NPC were seeded on 12-well plates at an appropriate density to ensure exponential growth until fixation (8.5 × 10^4/cm2). The following day, NPC were treated with DMSO or Nocodazole (Sigma-Aldrich, 125nM) for 24h. Afterwards, the cells were detached, counted and 6 × 10^5 cells per condition fixed with 4% PFA for 15 min at RT. Cells were washed 3 × with PBS and treated with 100 µg/ml RNAse A (New England Biolabs, Ipswich, MA, USA) and 0.1% Triton X-100 in TrypLE express (Thermo Fisher) at 37 °C for 30 min for RNA removal and cell permeabilization. For fluorescence-activated cell sorting (FACS), 3 × 10^5 cells were transferred per well of a 96-well plate and stained with 2 µg/ml propidium iodide (PI) solution (BioLegend, San Diego, CA, USA) in TrypLE express for 30 min at 37 °C. Analysis of PI staining was performed using the FACS Fortessa™ Cell Analyzer (BD Biosciences, Franklin Lakes, NJ, USA). A minimum of 15,000 events was recorded per well, from two wells per condition. Analysis of FACS data was performed using FlowJo 10 (BD Biosciences). The gating strategy is outlined in Additional file 5 – Figure S11A. Cell cycle phases were identified using the build-in univariate cell cycle analysis tool (Dean-Jett-Fox model, [31]).
Cell proliferation assay
Passage-matched NPC were seeded at a density of 2 × 10^4 cells/well in a 96-well plate in STEMdiff™ Neural Progenitor medium. For 72h, cells were cultivated in the IncuCyte® Live Cell Analysis System (Sartorius, Göttingen, Germany). Whole-well brightfield images were acquired in 4 h intervals with a 10 × objective in triplicates for each NPC line. Proliferation was analyzed using the IncuCyte® Basic Analyzer. Confluence was normalized to t = 0. Data was obtained from three biological replicates.
Western blot
Size-separation via SDS-PAGE was performed as described above (DigiWest) using 10 μg of protein and blotted onto a Nitrocellulose membrane (VWR). Blocking was performed for 1 h using 5% milk powder (Roth) in TBST. Primary antibodies (identical to those used in DigiWest) were diluted in 1% BSA (Roth) in TBST and incubated overnight at 4 °C. Blots were washed 5 × for 5 min in TBST before adding fluorescently labelled secondary antibodies for 1 h (donkey anti-Rabbit IgG (H + L) coupled to IRDye 800CW (LI-COR, Lincoln, NE, USA) and donkey anti-Mouse IgG (H + L) coupled to IRDye 680CW (LI-COR), each at a dilution of 1:10,000 in 5% milk powder in TBST) and detected on a LI-COR instrument. Before analysis, blots were washed 5 × for 5 min in TBST and bands were quantified using Image Studio. Uncropped Western Blot images can be found in Additional file 5 – Figure S15.
Neurite outgrowth assay
Passage-matched NPC were seeded into 48-well plates at 1 × 104 cells per well in Neural Progenitor medium. After 24 h, medium was changed to N2-medium (t = 0 h), consisting of DMEM/F12 + 15 mM HEPES, 1 × N2-supplement, 1% non-essential amino acids, 1% GlutaMAX, 1.5% glucose solution (all Thermo Fisher), 10 µM SB431542 (Bio-Techne, Minneapolis, MN, USA), 1 µM XAV939 (Sigma-Aldrich), 100 nM LDN193189 (STEMCELL Technologies), 10 ng/ml BDNF (Thermo Fisher). From then on, cells were cultured in the IncuCyte® Live Cell Analysis System (Sartorius) and imaged every 4 h in brightfield mode at 10 × magnification. Two wells were imaged per line, and 4 sites per well. After 24 h, a complete medium change with N2 medium was performed and cells were cultured until t = 40 h without further medium changes. Using the IncuCyte® NeuroTrack analysis module (Sartorius), neurite length (in mm) per number of cell body clusters was analyzed for each time point. Analysis parameters were defined as follows: cell body clusters with a minimum area of 200 µm, containing cells with a minimum width of 7 µm, were counted. Neurite width was defined as 1 µm. These parameters were analyzed for each of the four sites imaged and then averaged per well. Data was obtained from four biological replicates. Values at t = 0 h were set to 1.
Statistics
Statistical analysis was performed using GraphPad Prism 10 (Graphpad Software). Data was tested for normality using the Shapiro–Wilk Test. Only if both groups were normally distributed, groups were compared via unpaired, two-tailed t-test. If normality was not met, the two-tailed Mann–Whitney-U Test was used. For DigiWest data-based comparisons (used in heatmaps), the Wilcoxon-signed rank test was used. For neurite outgrowth and flow cytometry analysis, data was analyzed with a 2-way ANOVA and Tukey’s multiple comparisons test. N numbers and statistical details for each experiment can be found in the respective figure legend. In all cases, a p-value < 0.05 was considered significant unless stated otherwise.
Results
Characterization of developmental marker expression and signaling pathway activation during NPC differentiation via DigiWest
To characterize the differentiation process from iPSC into NPC, we examined the expression of 133 proteins from three healthy control lines and four SCZ patient lines. This set of proteins was composed of routinely used cell type markers for iPSC and NPC, as well as proteins involved in signaling pathways associated with neurodevelopment and previously linked to SCZ. First, we aimed to investigate the differences between the iPSC and NPC stage independent of disease allocation. Expression signatures of all measured proteins in NPC were consistent between three individual differentiations (Additional file 5 – Figure S1) where NPC of the respective differentiation do not cluster together, demonstrating high reproducibility. When comparing all iPSC (n = 21) and NPC (n = 21) samples, expectedly observed strong changes in the expression of cell-type specific markers. In NPC, DigiWest data shows downregulation of the pluripotency marker Oct4. Sox2 was expressed in both cell types but to a greater extent in NPC. Clear upregulation of neurodevelopmental/NPC-associated markers microtubule-associated protein 2 (MAP2), Vimentin, neural cell adhesion molecule (NCAM), paired-box protein Pax-6 (Pax6), SRY-Box Transcription Factor 1 (Sox1) and Nestin was observed (Fig. 1A, Additional file 5—Figure S2). Although variability in expression levels between individual patient-derived lines does persist, the two cell types are clearly distinguishable when displayed in a Western Blot-like format, as shown in Fig. 1B (Additional file 5 – Figure S3). Expression of these cell type-specific markers was confirmed by immunocytochemical staining (ICC) in both iPSC and NPC (Fig. 1C) with good comparability of DigiWest and ICC as exemplarily shown for four markers (Additional file 5 – Figure S4).
The high-throughput nature of DigiWest allowed us to assess expression of over 130 proteins and phosphoproteins and thus look beyond the expression of traditional cell type markers. We observed further pronounced and consistent expression differences between iPSC and NPC for various other proteins, leaving 61% (81/133) of analytes differentially expressed at a significance level of p < 0.001 (Fig. 2A, Additional file 5 – Figure S2), with the majority becoming strongly upregulated in NPC. These upregulations included key members of signaling pathways involved in neurodevelopment (Fig. 2B), such as Wnt signaling (active and total beta-catenin, Wnt3/7, low-density lipoprotein receptor-related protein 6 (LRP6 – pS1490), GSK3 beta, Dishelleved (Dvl) 2/3, lymphoid enhancer-binding factor 1 (LEF1), transcription factor (TCF) 1/7), Hippo signaling (large tumor suppressor kinase 1 (LATS1), Mob1, macrophage stimulating protein 1 (MST1), TEA domain transcription factor 1 (TEAD), kidney and brain expressed protein (KIBRA)) and Hedgehog signaling (glioma-associated oncogene (GLI)2/3, suppressor of fused protein (SUFU)). Notably, proteins involved in cell cycle regulation (cyclin-dependent kinase (CDK) 1/2/4/6, Cyclin D1/2, p21, Rb) were also strongly elevated in NPC (Fig. 2B). For select proteins we again validated the DigiWest results with ICC. Expression levels of beta-catenin and LEF1, both critical for Wnt signaling, and of the cell cycle modulator p21 were significantly elevated in NPC compared to iPSC (Fig. 2C + D).
When investigating the differentiation process from iPSC to NPC comparing patient and healthy control-derived cells, we mainly observed strong upregulations (CTR: 89.4%, SCZ: 92.4%) and only a handful of downregulations (CTR: 10.6%, SCZ: 7.6%) for both (Additional file 5 – Figure S5). As expected, most changes were shared between CTR and SCZ, given that we already observed highly consistent expression across most samples (see Fig. 2A). Regardless, some select analytes behaved differently in SCZ vs CTR-derived cells during development, as calculated by two-factor-ANOVA (interaction effect p < 0.05). These (e.g. LEF1, Smurf1, p21 or ERK1/2 – pT202/Y204) are highlighted in Fig. 2E/F and are shown in more detail in Additional file 5 – Figure S6/ Additional file 1 – Table S3. Overall, these data show that the DigiWest can clearly identify cell-type specific proteomic signatures in differentiated cells thus allowing extensive analysis of cellular signaling during the differentiation process.
SCZ iPSC show only few disease-associated alterations at the protein level
Next, we aimed to investigate disease-specific proteomic signatures at the iPSC and NPC stage, respectively. In iPSC, we found 11 (8%) analytes to be differentially expressed between CTR (n = 9) and SCZ (n = 12), with 5 becoming up- and 6 downregulated in SCZ (Fig. 3A + B). By far, the strongest effects in magnitude were observed for p53 phosphorylated at Ser15 (~ tenfold upregulation in SCZ), along with total levels of p53 (~ twofold upregulation in SCZ) (Fig. 3C). We confirmed upregulation of p53 in SCZ iPSC by ICC (Fig. 3D-E). Other de-regulated proteins showed significant, but less pronounced differences (Additional file 1 – Table S4). These data suggest that, except altered p53 expression and phosphorylation, there are relatively few prominent changes between CTR and SCZ on the iPSC level.
Reduced differentiation capacity of SCZ iPSC into neural progenitor cells
At the NPC stage, we again compared all SCZ (n = 9) and control (n = 12) samples and identified 30 analytes (23%) significantly different between the two groups (Fig. 4A + B), three times as many as in iPSC. Of these 30, 23 proteins were upregulated, and 7 were downregulated in SCZ. Crucially, several of the eight cell type-specific markers discussed previously (see Fig. 1) were among the differentially expressed proteins. On average, SCZ NPCs displayed higher levels of Oct4 and decreased levels of MAP2, NCAM and Sox1 compared to CTR NPC, while the other markers were not affected (Fig. 4C, Additional file 5 – Figure S7). Overall, this indicates decreased differentiation efficiency of SCZ NPC. Of note, considerably inter-donor variability was observed within both CTR as well as SCZ groups regarding the expression of these proteins with clones thus contributing to the observed SCZ phenotypes to varying extent (Additional file 5 – Figure S8A-B). To evaluate if decreased differentiation efficiency of SCZ iPSC impacts early neurodevelopmental processes, we tracked neurite outgrowth of NPC for 40 h. Starting 24 h after induction of neuronal differentiation, SCZ NPC showed a reduction of mean neurite length of increasing magnitude over time compared to CTR cultures (Additional file 5 –Figure S9).
Aberrant protein expression and dysregulated signaling pathway activity in SCZ NPC
Of all de-regulated analytes, most analytes (26/30) with differential regulation were exclusive to the NPC stage (Fig. 4D) and will be discussed in detail in Fig. 5, whereas only 4 SCZ-specific effects were conserved from the iPSC stage. Besides minor changes in CDK4 and ataxia-telangiectasia mutated kinase (ATM) expression (Additional file 5 – Figure S10A), these included highly increased levels of phosphorylated p53 (Ser15) and total p53 (Fig. 4E). Noteworthy, the average magnitude of change for these two analytes was twice as high in NPC (20-fold/sixfold) as in iPSC (tenfold/threefold). Similar results were obtained with traditional Western Blotting (Fig. 4F-G). We again confirmed p53 upregulation in NPC via ICC (F ig. 4H-I), also to a greater extent than in iPSC (threefold/1.8-fold). Overall, SCZ-specific alterations of protein expression mostly appear exclusively at the NPC stage, and in the case of p53 dysregulation are more pronounced compared to iPSC.
Fig. 5.
Pathway allocations of SCZ-specific proteins. A-E: DigiWest data (relative to CTR mean) of select proteins differentially expressed between CTR (n = 9) and SCZ (n = 12) in NPC only (see Fig. 4D), shown in direct comparison with iPSC (CTR n = 9, SCZ n = 12). Analytes are grouped based on pathway/cellular function. A: Cell cycle regulation, specifically G2/M phase transition, B: DNA damage response, C: Protein synthesis/translation, D: Wnt signaling, E: MAPK/Erk signaling. A complete list of all differentially expressed SCZ-specific analytes can be found in Additional file 1—Table S5. Either the Mann–Whitney test or unpaired t-test was used depending on data distribution. *p < 0.05, **p < 0.01, ***p < 0.001 or as indicated. Error bars: S.E.M
Furthermore, several of the dysregulated analytes exclusive to NPC were phospho-variants, suggesting deregulated cellular signaling. Accordingly, most of them could be attributed to distinct signaling pathways while others serve as regulators of cellular processes (Fig. 5A-E, Additional file 1 – Table S5). Among them were cell cycle-regulating proteins such as CDK1 – pY15, Cyclin B1 and Aurora Kinase A, as well as Histone H3 – pS10 (a marker for cells undergoing mitosis) all roughly showing a twofold increase in SCZ (Fig. 5A). Total CDK1, Aurora Kinase B and Cyclin E1 (Additional file 5 – Figure S10B) were also upregulated. Notably, they all play a critical role in regulating transition into G2/M phase. Interestingly, the relative inter-donor expression levels of Cyclin B1 and Aurora A, as well as p53 and p53 – pS15 (Additional file 5 – Figure S8C), showed strikingly similar patterns to the relative expression levels of the pluripotency marker Oct4.
Additionally, we observed several alterations in the DNA-Damage-response (DDR) pathway (Fig. 5B). Histone H2A.X (S139), a primary indicator for DNA damage, as well as phosphorylation of crucial DDR proteins Chk1 (S296), and ataxia telangiectasia and Rad3-related kinase (ATR—S428 and ATR – S1989) were elevated in SCZ cells. On the other hand, phospho-ATM (S1981) showed an opposing trend for downregulation (Additional file 5 – Figure S10B).
Upregulation was also observed for several proteins and phospho-variants involved in translation such as eukaryotic elongation factor 2 (eEF2 – pT56), eukaryotic Initiation Factor 2 (eIF2) alpha, the ribosomal protein S6, as well as eukaryotic translation initiation factor 4E (eIF4E) – pS209 (Fig. 5C) along with total eEF2 and eukaryotic elongation factor 2 kinase (eEF2K) – pS366 and (Additional file 5 – Figure S10B). Also, phosphorylation of eukaryotic translation initiation factor 4E-binding protein 1 (4E-BP1, T37/S46, p = 0.0585) and the inhibitory site of eIF2 alpha (S51, p = 0.0601) showed an up- and downregulatory trend, respectively.
Moreover, we found dysregulations in Wnt signaling (Fig. 5D + Additional file 5 – Figure S10B), such as decreased active beta-catenin (non-pS33/41/45), twofold elevated LEF1, reduced protein kinase A (PKA) C (T197), total PKA C and elevated Casein kinase 1 alpha.
Finally, SCZ NPC also displayed increased phosphorylation of key MAPK proteins (Fig. 5E), namely c-Raf (S289/296/301), Erk2 (T202/Y204), Erk1 (T202/Y204) and Ribosomal S6 kinase (RSK) 1 (T573). Notably, the change of Rsk1 phosphorylation was opposite to iPSC, in which a significant reduction was observed.
Overall, protein expression patterns in NPC derived from SCZ patients suggest cell-type specific dysregulations in translation and protein synthesis, increased DNA Damage and DDR, elevated MAPK/Erk signaling, altered Wnt signaling, and higher expression of proteins governing cell cycle control, specifically those involved in regulating G2/M transition.
SCZ NPC accumulate in G2/M phase of the cell cycle
As upregulation of cell cycle stage-specific mediators have so far not been phenotypically linked with SCZ using iPSC-derived cell types, we aimed to investigate this aspect in more detail. ICC staining for G2/M-associated proteins Aurora A and Cyclin B1 confirmed DigiWest data and revealed an increased expression in SCZ NPC, but not iPSC (Fig. 6A + B). Furthermore, we conducted FACS analysis of propidium iodide staining in NPC to evaluate cell cycle phase distribution. Here, we observed a significantly increased percentage of cells in the G2/M phase for SCZ NPC compared to controls, while S-phase and G0/G1 phase were unaltered (Fig. 6C-D, Additional file 5 – Figure S11B). In addition, CTR and SCZ NPC showed no difference in proliferation rates, thus excluding the possibility that the rise in expression is simply due generally increased cell cycle activity of SCZ NPC (Additional file 5 – Figure S12). We also confirmed elevated Cyclin B1 expression via Western Blotting, with SCZ NPC showing an up-regulatory trend (Additional file 5 – Figure S13).
Expression of differentiation markers and G2/M-phase proteins correlate with p53 levels
To investigate a potential link between p53 expression/phosphorylation and the observed cellular phenotypes, we performed correlation analyses in NPC across all donors among p53 – pS15, p53, differentiation markers as well as G2/M-specific cell cycle regulators (Additional file 5 – Figure S14). As p53 phosphorylation showed the most striking alteration in SCZ NPCs, its correlation analyses are shown in detail. We found a strongly significant positive correlation (Spearman´s r = 0.9671) with total p53 levels (Fig. 6E), indicating that p53 phosphorylation is dependent on p53 abundance. Furthermore, we observed a positive correlation with Oct4 (r = 0.8007) and negative correlations with MAP2 (r = -0.6272) and NCAM (r = -0.8289) and Sox1 (r = -0.6763), respectively (Fig. 6F), thus establishing a link between p53 and the observed reduced differentiation efficiency. p53 phosphorylation also strongly correlated with G2/M cell cycle proteins Aurora A (r = 0.8959) and Cyclin B1 (r = 0.8614), as well as moderately with CDK1 – pY15 (r = 0.5869, Fig. 6G).
Discussion
Here, we report the first protein profiling analysis focusing on cellular signal transduction to study early neurodevelopmental aberrations in SCZ using iPSC-derived cell types. In a targeted proteomics approach based on the DigiWest technology, we found deregulation of protein expression mainly in NPC derived from patients with SCZ. We demonstrate regulatory alterations in cell cycle control and DNA-damage response along with impaired differentiation capacity. Additionally, we correlatively linked these aberrations to p53 expression and phosphorylation levels in diseased cells. Furthermore, we confirmed previous observations of altered protein synthesis, WNT signaling and MAPK/Erk signaling in SCZ.
The efficient differentiation of iPSC into neuronal cell types is commonly assessed via immunostaining of few selected cell type-specific markers [32]. We demonstrate that DigiWest can recapitulate characteristic pluripotency and neural progenitor marker expression equally well when compared to ICC. We show activation of several key developmental pathways such as Wnt [33], Hedgehog [34], and Hippo [25] signaling, along with changes in cell cycle regulators [35, 36]. Moreover, cells derived from SCZ patients do differ in the expression of select Wnt, MAPK and cell cycle proteins during differentiation. While other studies have also reported decreased neuronal differentiation capacity of SCZ patient-derived iPSC [37, 38], a link to changes in developmental pathway activity underlying these phenotypes has not yet been proposed. Thus overall, DigiWest is a powerful, novel tool for iPSC-based studies as it can highlight disease-associated differences which are not necessarily visible by analysis of differentiation markers alone.
NPCs have frequently been used to recapitulate SCZ pathology in vitro at early neurodevelopmental stages, as it was previously demonstrated that gene expression signatures of iPSC-derived neurons are largely conserved in NPC [9] and that protein signatures of NPC show similarities to patient postmortem brains [11]. In line with this, we mostly observed prevalent changes exclusively in NPC, indicating that most alterations and dysfunctions in pathway regulation occur in differentiated cells, but are not yet present in iPSC. For instance, we provide evidence for regulatory changes of protein synthesis, MAPK signaling and Wnt signaling in SCZ NPC. In support of our findings, several reports using iPSC models of SCZ indicate changes in translation and protein synthesis [6, 8, 11] as well as altered WNT signaling in SCZ NPC [39], neurons [16] and patients [14].
Notably, we did find highly elevated expression levels of p53 and its phosphorylated variant p53-pS15 in SCZ already at the iPSC stage. p53 is a tumor suppressor protein with central roles in regulating cellular responses to genomic alterations and DNA damage along with controlling of the cell cycle and apoptosis. Importantly, p53 is only phosphorylated at Ser15 in response to DNA damage [37, 38]. Furthermore, this phosphorylation event stabilizes p53 levels by reducing interaction of p53 with its negative regulator mouse double minute 2 homolog (MDM2) [40]. p53 is mainly associated with tumorigenesis but also plays a role in brain development, neural stem cell regulation [41], and in several nervous system diseases including SCZ, autism, Parkinson´s disease, Alzheimer´s disease or epilepsy [42]. TP53 has been identified as a candidate risk gene in SCZ in gene association [43] as well as case–control/family studies [44] have specifically associated genetic variation in TP53 with SCZ. Further support for an involvement of p53 is given by extensive evidence showing a negative correlation between SCZ and cancer incidence [22, 45, 46].
It has been hypothesized that p53 upregulation in stem cells due to genotoxic stress impairs pluripotency and might lead to unwanted, unspecific differentiation [39]. This aspect could at least in part explain our finding of reduced differentiation efficiency of SCZ iPSC upon neural induction and the resulting lower state of maturity of SCZ NPC, as indicated by higher expression of OCT4 and lower expression of MAP2, NCAM and SOX1. Crucially, for MAP2 and NCAM, transcriptomic profiling of a different set of SCZ patient-derived NPC revealed similar effect [9]. The lower maturation state of SCZ NPC might impact the efficiency of early neurodevelopmental processes, as reflected in a reduction of neurite outgrowth which we and others [9, 47] have observed in SCZ. In accordance with our results, decreased MAP2 expression along with alterations in pathways governing neuronal differentiation have also been reported in SCZ iPSC-derived organoids [23, 48]. Moreover, impaired neuronal differentiation efficiency has been described for 22q11.2 deletion iPSC [38], a CNV associated with SCZ, and patient-derived glutamatergic neurons [37]. It can also be hypothesized that impaired neuronal maturation could influence other key neurodevelopmental processes such as neuronal synapse formation, which is impaired in SCZ [21]. This however needs further investigation. Importantly, the strong up-regulatory trend of p53 was the only notable SCZ-specific alteration present in both iPSC and NPC, highlighting its importance in the context of differentiation.
In addition, our results indicate that SCZ NPC show increased levels of damaged DNA and associated DNA-damage response. While the overarching picture of how DNA damage is implicated in neurogenesis and SCZ is still incomplete, it has been associated with the disease [49, 50]. For instance, genetic alterations in DNA repair enzymes were linked to SCZ [51] or can induce behavioral changes in mice [52]. A recent transcriptomic study of postmortem SCZ brain tissue also uncovered increased DNA damage repair [53]. Moreover, mass spectrometry analysis of SCZ-derived neural stem cells found alterations in pathways related to mitochondrial function, metabolic activity and DNA repair [54]. Lastly, reactive oxygen species and oxidative stress were shown to be elevated in SCZ NPC [9, 55] and animal models [56]. Thus, our data provide further evidence for a significant implication of DNA damage and repair mechanisms in SCZ pathology. Interestingly, we demonstrate that SCZ NPC display higher levels of G2/M-associated proteins, where cells are checked for DNA damage before cell cycle progression [57]. While we did not observe changes in basal proliferation rate of NPC, our FACS analysis showed a greater proportion of SCZ cells in G2/M phase—it is tempting to speculate that SCZ NPC become stuck in the G2/M phase, potentially leading to delayed cell cycle exit/progression during differentiation. Crucially, p53 can cause G2/M arrest in response to DNA damage [58]. Based on our data, we cannot infer whether cells arrest (and later attempt to recover) or exit the cell cycle temporarily; accordingly, cell cycle arrest has been described as a highly dynamic molecular state [59]. An association of DNA damage and G2/M phase in SCZ has thus far not been reported, although studies have shown alterations in cell cycle control in SCZ models [54, 60, 61] and the relevance of cell phase regulation and progression in neurodevelopment [62]. Most prior studies on SCZ proteomics/transcriptomics/genomics use shotgun approaches, merely reporting SCZ-specific changes on pathway-association level. With DigiWest being a targeted approach, we were able to focus on more detailed mechanisms of pathway activation – also by additionally measuring phosphorylation.
We identified such coherent proteomic changes while confirming previous findings from other omics approaches despite a small cohort size and notable inter-patient variability. Moreover, the four patients had diverse genetic backgrounds and disease manifestations, thus covering various clinical phenotypes, which is helpful in studying multifaceted diseases such as SCZ. Notably, for some of the measured analytes, we detected strong inter-donor variability, likely due to differences in the genetic background of each individual, which was previously shown to be the largest source of variability in a proteomic study on iPSC-derived astrocytes [63]. In addition, there are several other potential sources of inter-donor variability, including donor sex [64] and age, as exonic mutations can accumulate over time [65]. In our dataset, patient line SCZ1 displayed the strongest phenotype for key observations (highest p53 expression, lowest differentiation capacity, highest G2/M proportion), while the SCZ2 patient line showed a more CTR-like phenotype in certain aspects (also see Additional file 5 – Figure S8A). This needs to be kept in mind when interpreting the data as it might reflect the phenotype of certain subgroups of patients only. These limitations could be accounted for in future studies employing a larger number of patients to disentangle the relationship between individual sources of variation and SCZ phenotypes.
Although the aspects discussed above (p53, cell cycle, differentiation efficiency) all have individually been shown to be implicated in SCZ, our correlation data for the first time provide a possible connection between them. It can be speculated that patient-derived progenitor cells acquire DNA damage during early development and deal with it inadequately, causing cell cycle arrest or delay. This could hamper differentiation efficiency, resulting in less mature NPC and eventually neurons, which might contribute to the developmental defects associated with SCZ. In future studies it would be of interest to investigate whether our observed proteomic alterations and association with p53 will be confirmed in larger cohorts to further deepen the understanding of neurodevelopmental failure in SCZ from a mechanistic standpoint.
Conclusion
Using a high-throughput targeted proteomics approach, we uncover aberrant protein expression signatures in SCZ patient-derived NPC. Moreover, diseased cells displayed regulatory changes in cell cycle control along with impaired differentiation capacity. Using our proteomic data, we demonstrate a potential link of these phenotypes to aberrant p53 expression and phosphorylation. We ultimately hypothesize a potential interplay of these disease-specific alterations affecting the differentiation process during early neurodevelopment, which could be mechanistically implicated in the manifestation of developmental alterations occurring in SCZ. Further studies could explore the significance of our findings in larger cohorts using patient-derived neuronal cell types in iPSC-based model systems. Our study not only demonstrates the importance and significance of stem cell-based model systems and protein-based analytics in psychiatric disorders such as SCZ but also highlights our patho-mechanistic findings as accelerators in the search for potential drug targets.
Supplementary Information
Additional file 4: DigiWest Normalized Data
Additional file 5: Supplementary Figures
Acknowledgements
Not applicable.
Abbreviations
- 4E-BP1
Eukaryotic translation initiation factor 4E-binding protein 1
- ATM
Ataxia-telangiectasia mutated kinase
- ATR
Ataxia telangiectasia and Rad3-related kinase
- CDK
Cyclin-dependent kinase
- CTR
Control
- DDR
DNA damage response
- eEF2
Eukaryotic elongation factor 2
- eIF2
Eukaryotic initiation factor 2
- eIF4E
Eukaryotic translation initiation factor 4E
- ERK
Extracellular signal-regulated kinases
- FACS
Fluorescence-activated cell sorting
- GLI
Glioma-associated oncogene
- GSK3 beta
Glycogen synthase kinase-3 beta
- ICC
Immunocytochemical staining
- iPSC
Induced pluripotent stem cells
- KIBRA
Kidney and brain expressed protein
- LATS1
Large tumor suppressor kinase 1
- LEF1
Lymphoid enhancer-binding factor 1
- LRP6
Low-density lipoprotein receptor-related protein 6
- MAP2
Microtubule-associated protein 2
- MAPK
Mitogen-activated protein kinase
- MDM2
Mouse double minute 2 homolog
- MST1
Macrophage stimulating protein 1
- NCAM
Neural cell adhesion molecule
- NPC
Neural progenitor cells
- PAX6
Paired-box protein Pax-6
- PKA
Protein kinase A
- RSK1
Ribosomal S6 kinase 1
- RT
Room temperature
- SCZ
Schizophrenia
- SOX1
SRY-Box Transcription Factor 1
- SOX2
SRY-Box Transcription Factor 2
- SUFU
Suppressor of fused protein
- TCF1/7
Transcription factor 1/7
- TEAD1
TEA domain transcription factor 1
Authors’ contributions
Conceptualization: AS, JH, HV and MFT; methodology: AS, JH; formal analysis: AS and JH; investigation: AS and JH.; data curation: AS and JH; writing—original draft preparation: AS, JH and MFT; writing—review and editing: HV, RW, AJF, KS-L; material support: RW, AJF; clinical evaluation: RW, AJF; supervision: MFT, HV, AJF and KS-L; project administration: MFT and HV.
Funding
Open Access funding enabled and organized by Projekt DEAL. This work received financial support from the State of Baden-Wuerttemberg (Prädiktive Diagnostik von immunassoziierten Erkrankungen für die personalisierte Medizin, grant no. AZ 35–4223.10/8) and from the State Ministry of Baden-Wuerttemberg for Economic Affairs, Labour and Tourism.
Availability of data and materials
All DigiWest-related raw and normalized data generated or analyzed during this study are included within the article (and its supplementary information files). All other datasets (Western Blot, ICC and FACS) used and/or analyzed during the study are available from the corresponding author on reasonable request.
All other datasets (Western Blot, ICC and FACS) used and/or analyzed during the study are available from the corresponding author on reasonable request.
Declarations
Ethics approval and consent to participate
Ethics approval for the use of all patient-derived iPSC lines was granted from the ethics commission at the University Hospital and Medical Faculty of Tübingen University (311/2013BO1).
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Aaron Stahl, Email: a.stahl@uni-tuebingen.de.
Markus F. Templin, Email: markus.templin@nmi.de
References
- 1.Tandon R, Gaebel W, Barch DM, Bustillo J, Gur RE, Heckers S, et al. Definition and description of schizophrenia in the DSM-5. Schizophr Res. 2013;150(1):3–10. [DOI] [PubMed] [Google Scholar]
- 2.Carbon M, Correll CU. Thinking and acting beyond the positive: the role of the cognitive and negative symptoms in schizophrenia. CNS Spectr. 2014;19(Suppl 1):38–52 (quiz 35-7, 53). [DOI] [PubMed] [Google Scholar]
- 3.Stepnicki P, Kondej M, Kaczor AA. Current Concepts and Treatments of Schizophrenia. Molecules. 2018;23(8):2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Singh T, Poterba T, Curtis D, Akil H, Al Eissa M, Barchas JD, et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature. 2022;604(7906):509–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604(7906):502–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.English JA, Fan Y, Focking M, Lopez LM, Hryniewiecka M, Wynne K, et al. Reduced protein synthesis in schizophrenia patient-derived olfactory cells. Transl Psychiatry. 2015;5(10):e663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rodrigues JE, Martinho A, Santa C, Madeira N, Coroa M, Santos V, et al. Systematic Review and Meta-Analysis of Mass Spectrometry Proteomics Applied to Human Peripheral Fluids to Assess Potential Biomarkers of Schizophrenia. Int J Mol Sci. 2022;23(9):4917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Topol A, English JA, Flaherty E, Rajarajan P, Hartley BJ, Gupta S, et al. Increased abundance of translation machinery in stem cell-derived neural progenitor cells from four schizophrenia patients. Transl Psychiatry. 2015;5(10):e662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Brennand K, Savas JN, Kim Y, Tran N, Simone A, Hashimoto-Torii K, et al. Phenotypic differences in hiPSC NPCs derived from patients with schizophrenia. Mol Psychiatry. 2015;20(3):361–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Maschietto M, Tahira AC, Puga R, Lima L, Mariani D, Paulsen Bda S, et al. Co-expression network of neural-differentiation genes shows specific pattern in schizophrenia. BMC Med Genomics. 2015;8:23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Nascimento JM, Saia-Cereda VM, Zuccoli GS, Reis-de-Oliveira G, Carregari VC, Smith BJ, et al. Proteomic signatures of schizophrenia-sourced iPSC-derived neural cells and brain organoids are similar to patients’ postmortem brains. Cell Biosci. 2022;12(1):189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Emamian ES, Hall D, Birnbaum MJ, Karayiorgou M, Gogos JA. Convergent evidence for impaired AKT1-GSK3beta signaling in schizophrenia. Nat Genet. 2004;36(2):131–7. [DOI] [PubMed] [Google Scholar]
- 13.Freyberg Z, Ferrando SJ, Javitch JA. Roles of the Akt/GSK-3 and Wnt signaling pathways in schizophrenia and antipsychotic drug action. Am J Psychiatry. 2010;167(4):388–96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hoseth EZ, Krull F, Dieset I, Morch RH, Hope S, Gardsjord ES, et al. Exploring the Wnt signaling pathway in schizophrenia and bipolar disorder. Transl Psychiatry. 2018;8(1):55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Peng Y, Xu Y, Cui D. Wnt signaling pathway in schizophrenia. CNS Neurol Disord Drug Targets. 2014;13(5):755–64. [DOI] [PubMed] [Google Scholar]
- 16.Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, et al. Modelling schizophrenia using human induced pluripotent stem cells. Nature. 2011;473(7346):221–5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Funk AJ, McCullumsmith RE, Haroutunian V, Meador-Woodruff JH. Abnormal activity of the MAPK- and cAMP-associated signaling pathways in frontal cortical areas in postmortem brain in schizophrenia. Neuropsychopharmacology. 2012;37(4):896–905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kyosseva SV, Elbein AD, Griffin WS, Mrak RE, Lyon M, Karson CN. Mitogen-activated protein kinases in schizophrenia. Biol Psychiatry. 1999;46(5):689–96. [DOI] [PubMed] [Google Scholar]
- 19.Yuan P, Zhou R, Wang Y, Li X, Li J, Chen G, et al. Altered levels of extracellular signal-regulated kinase signaling proteins in postmortem frontal cortex of individuals with mood disorders and schizophrenia. J Affect Disord. 2010;124(1–2):164–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Treindl F, Ruprecht B, Beiter Y, Schultz S, Dottinger A, Staebler A, et al. A bead-based western for high-throughput cellular signal transduction analyses. Nat Commun. 2016;7:12852. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Breitmeyer R, Vogel S, Heider J, Hartmann SM, Wust R, Keller AL, et al. Regulation of synaptic connectivity in schizophrenia spectrum by mutual neuron-microglia interaction. Commun Biol. 2023;6(1):472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chou FH, Tsai KY, Su CY, Lee CC. The incidence and relative risk factors for developing cancer among patients with schizophrenia: a nine-year follow-up study. Schizophr Res. 2011;129(2–3):97–103. [DOI] [PubMed] [Google Scholar]
- 23.Notaras M, Lodhi A, Fang H, Greening D, Colak D. The proteomic architecture of schizophrenia iPSC-derived cerebral organoids reveals alterations in GWAS and neuronal development factors. Transl Psychiatry. 2021;11(1):541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mulligan KA, Cheyette BN. Neurodevelopmental Perspectives on Wnt Signaling in Psychiatry. Mol Neuropsychiatry. 2017;2(4):219–46. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sahu MR, Mondal AC. Neuronal Hippo signaling: From development to diseases. Dev Neurobiol. 2021;81(2):92–109. [DOI] [PubMed] [Google Scholar]
- 26.Gulino A, Di Marcotullio L, Ferretti E, De Smaele E, Screpanti I. Hedgehog signaling pathway in neural development and disease. Psychoneuroendocrinology. 2007;32(Suppl 1):S52–6. [DOI] [PubMed] [Google Scholar]
- 27.Hiew LF, Poon CH, You HZ, Lim LW. TGF-beta/Smad Signalling in Neurogenesis: Implications for Neuropsychiatric Diseases. Cells. 2021;10(6):1382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kanehisa M, Sato Y. KEGG Mapper for inferring cellular functions from protein sequences. Protein Sci. 2020;29(1):28–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, et al. TM4: a free, open-source system for microarray data management and analysis. Biotechniques. 2003;34(2):374–8. [DOI] [PubMed] [Google Scholar]
- 31.Fox MH. A model for the computer analysis of synchronous DNA distributions obtained by flow cytometry. Cytometry. 1980;1(1):71–7. [DOI] [PubMed] [Google Scholar]
- 32.Rust R, Weber RZ, Generali M, Kehl D, Bodenmann C, Uhr D, et al. Xeno-free induced pluripotent stem cell-derived neural progenitor cells for in vivo applications. J Transl Med. 2022;20(1):421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mulligan KA, Cheyette BNR. Wnt Signaling in Vertebrate Neural Development and Function. J Neuroimmune Pharmacol. 2012;7(4):774–87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Yang C, Qi Y, Sun Z. The Role of Sonic Hedgehog Pathway in the Development of the Central Nervous System and Aging-Related Neurodegenerative Diseases. Front Mol Biosci. 2021;8:711710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hardwick LJA, Ali FR, Azzarelli R, Philpott A. Cell cycle regulation of proliferation versus differentiation in the central nervous system. Cell Tissue Res. 2015;359(1):187–200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zaveri L, Dhawan J. Cycling to Meet Fate: Connecting Pluripotency to the Cell Cycle. Front Cell Dev Biol. 2018;6:57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Robicsek O, Karry R, Petit I, Salman-Kesner N, Muller FJ, Klein E, et al. Abnormal neuronal differentiation and mitochondrial dysfunction in hair follicle-derived induced pluripotent stem cells of schizophrenia patients. Mol Psychiatry. 2013;18(10):1067–76. [DOI] [PubMed] [Google Scholar]
- 38.Toyoshima M, Akamatsu W, Okada Y, Ohnishi T, Balan S, Hisano Y, et al. Analysis of induced pluripotent stem cells carrying 22q11.2 deletion. Transl Psychiatry. 2016;6(11):e934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Fu X, Wu S, Li B, Xu Y, Liu J. Functions of p53 in pluripotent stem cells. Protein Cell. 2019;11(1):71–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Shieh SY, Ikeda M, Taya Y, Prives C. DNA damage-induced phosphorylation of p53 alleviates inhibition by MDM2. Cell. 1997;91(3):325–34. [DOI] [PubMed] [Google Scholar]
- 41.Xiong Y, Zhang Y, Xiong S, Williams-Villalobo AE. A Glance of p53 Functions in Brain Development, Neural Stem Cells, and Brain Cancer. Biology. 2020;9(9):285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li H, Zhang Z, Li H, Pan X, Wang Y. New Insights into the Roles of p53 in Central Nervous System Diseases. Int J Neuropsychopharmacol. 2023;26(7):465–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zhuo C, Wang D, Zhou C, Chen C, Li J, Tian H, et al. Double-Edged Sword of Tumour Suppressor Genes in Schizophrenia. Front Mol Neurosci. 2019;12:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Ni X, Trakalo J, Valente J, Azevedo MH, Pato MT, Pato CN, et al. Human p53 tumor suppressor gene (TP53) and schizophrenia: case-control and family studies. Neurosci Lett. 2005;388(3):173–8. [DOI] [PubMed] [Google Scholar]
- 45.Ji J, Sundquist K, Ning Y, Kendler KS, Sundquist J, Chen X. Incidence of cancer in patients with schizophrenia and their first-degree relatives: a population-based study in Sweden. Schizophr Bull. 2013;39(3):527–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Mortensen PB. The incidence of cancer in schizophrenic patients. J Epidemiol Community Health. 1989;43(1):43–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Grunwald LM, Stock R, Haag K, Buckenmaier S, Eberle MC, Wildgruber D, et al. Comparative characterization of human induced pluripotent stem cells (hiPSC) derived from patients with schizophrenia and autism. Transl Psychiatry. 2019;9(1):179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Notaras M, Lodhi A, Fang H, Greening D, Colak D. The proteomic architecture of schizophrenia iPSC-derived cerebral organoids reveals alterations in GWAS and neuronal development factors. Transl Psychiatry. 2021;11(1):541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Markkanen E, Meyer U, Dianov GL. DNA Damage and Repair in Schizophrenia and Autism: Implications for Cancer Comorbidity and Beyond. Int J Mol Sci. 2016;17(6):856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Catts VS, Catts SV, Jablensky A, Chandler D, Weickert CS, Lavin MF. Evidence of aberrant DNA damage response signalling but normal rates of DNA repair in dividing lymphoblasts from patients with schizophrenia. World J Biol Psychiatry. 2012;13(2):114–25. [DOI] [PubMed] [Google Scholar]
- 51.Sujitha SP, Kumar DT, Doss CGP, Aavula K, Ramesh R, Lakshmanan S, et al. DNA Repair Gene (XRCC1) Polymorphism (Arg399Gln) Associated with Schizophrenia in South Indian Population: A Genotypic and Molecular Dynamics Study. PLoS ONE. 2016;11(1):e0147348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bjorge MD, Hildrestrand GA, Scheffler K, Suganthan R, Rolseth V, Kusnierczyk A, et al. Synergistic Actions of Ogg1 and Mutyh DNA Glycosylases Modulate Anxiety-like Behavior in Mice. Cell Rep. 2015;13(12):2671–8. [DOI] [PubMed] [Google Scholar]
- 53.Shishido R, Kunii Y, Hino M, Izumi R, Nagaoka A, Hayashi H, et al. Evidence for increased DNA damage repair in the postmortem brain of the high stress-response group of schizophrenia. Front Psychiatry. 2023;14:1183696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Zuccoli GS, Nascimento JM, Moraes-Vieira PM, Rehen SK, Martins-de-Souza D. Mitochondrial, cell cycle control and neuritogenesis alterations in an iPSC-based neurodevelopmental model for schizophrenia. Eur Arch Psychiatry Clin Neurosci. 2023;273(8):1649–64. [DOI] [PubMed] [Google Scholar]
- 55.da Silveira Paulsen B, de Moraes Maciel R, Galina A, Souza da Silveira M, dos Santos Souza C, Drummond H, et al. Altered oxygen metabolism associated to neurogenesis of induced pluripotent stem cells derived from a schizophrenic patient. Cell Transplant. 2012;21(7):1547–59. [DOI] [PubMed] [Google Scholar]
- 56.Cabungcal JH, Counotte DS, Lewis E, Tejeda HA, Piantadosi P, Pollock C, et al. Juvenile antioxidant treatment prevents adult deficits in a developmental model of schizophrenia. Neuron. 2014;83(5):1073–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Barnum KJ, O’Connell MJ. Cell Cycle Regulation by Checkpoints. In: Noguchi E, Gadaleta MC, editors. Cell Cycle Control: Mechanisms and Protocols. New York: Springer; 2014. p. 29–40. [Google Scholar]
- 58.Chen J. The Cell-Cycle Arrest and Apoptotic Functions of p53 in Tumor Initiation and Progression. Cold Spring Harb Perspect Med. 2016;6(3):a026104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Stallaert W, Taylor SR, Kedziora KM, Taylor CD, Sobon HK, Young CL, et al. The molecular architecture of cell cycle arrest. Mol Syst Biol. 2022;18(9):e11087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Fan Y, Abrahamsen G, McGrath JJ, Mackay-Sim A. Altered cell cycle dynamics in schizophrenia. Biol Psychiatry. 2012;71(2):129–35. [DOI] [PubMed] [Google Scholar]
- 61.Okazaki S, Boku S, Otsuka I, Mouri K, Aoyama S, Shiroiwa K, et al. The cell cycle-related genes as biomarkers for schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry. 2016;70:85–91. [DOI] [PubMed] [Google Scholar]
- 62.Cremisi F, Philpott A, Ohnuma S. Cell cycle and cell fate interactions in neural development. Curr Opin Neurobiol. 2003;13(1):26–33. [DOI] [PubMed] [Google Scholar]
- 63.Beekhuis-Hoekstra SD, Watanabe K, Werme J, de Leeuw CA, Paliukhovich I, Li KW, et al. Systematic assessment of variability in the proteome of iPSC derivatives. Stem Cell Res. 2021;56:102512. [DOI] [PubMed] [Google Scholar]
- 64.Waldhorn I, Turetsky T, Steiner D, Gil Y, Benyamini H, Gropp M, et al. Modeling sex differences in humans using isogenic induced pluripotent stem cells. Stem Cell Reports. 2022;17(12):2732–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lo Sardo V, Ferguson W, Erikson GA, Topol EJ, Baldwin KK, Torkamani A. Influence of donor age on induced pluripotent stem cells. Nat Biotechnol. 2017;35(1):69–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 4: DigiWest Normalized Data
Additional file 5: Supplementary Figures
Data Availability Statement
All DigiWest-related raw and normalized data generated or analyzed during this study are included within the article (and its supplementary information files). All other datasets (Western Blot, ICC and FACS) used and/or analyzed during the study are available from the corresponding author on reasonable request.
All other datasets (Western Blot, ICC and FACS) used and/or analyzed during the study are available from the corresponding author on reasonable request.