Abstract
Purpose.
Targeted therapy in translocation-associated sarcomas has been limited to oncogenic activation of tyrosine kinases or ligands, while gene fusions resulting in aberrant expression of transcription factors have been notoriously difficult to target. Moreover, secondary genetic alterations in sarcomas driven by translocations are uncommon, comprising mostly alterations in tumor suppressor genes (TP53, CDKN2A/B). Our study was triggered by an index patient showing a dramatic clinical response by targeting the secondary BRAF V600E mutation in a metastatic angiomatoid fibrous histiocytoma (AFH) harboring the typical EWSR1::CREB1 fusion.
Patient.
The patient, a 28-year-old female, was diagnosed with an AFH of the thigh and followed a highly aggressive clinical course, with rapid multifocal local recurrence within a year and widespread distant metastases (adrenal, bone, liver, lung). The tumor showed characteristic morphologic features, with histiocytoid cells intermixed with hemorrhagic cystic spaces and lymphoid aggregates. In addition to the pathognomonic EWSR1::CREB1 fusion, targeted DNA sequencing revealed in both primary and adrenal metastatic sites a hot spot BRAF V600E mutation and a CDKN2A/B deletion. Accordingly, the patient was treated with a BRAF-MEK inhibitor combination (encorafenib/binimetinib) showing an excellent but short-lived response.
Results.
Using a CRISPR-Cas9 approach, we introduced the BRAF c.1799 T>A point mutation in human embryonic stem (hES) cells harboring a conditional EWSR1 (exon7)::CREB1 (exon7) translocation and further differentiated to mesenchymal progenitors (hES-MP) before fusion expression. The cells maintained the fusion transcript expression and the AFH core gene signature, while responding to treatment with encorafenib and binimetinib.
Conclusion.
These results highlight that additional targeted DNA NGS in chemotherapy resistant translocation-associated sarcomas may reveal actionable oncogenic drivers occurring as secondary genetic events during disease progression.
Keywords: sarcoma, translocation, EWSR1, CREB, angiomatoid fibrous histiocytoma, BRAF V600E, MEK
INTRODUCTION
Recurrent gene fusions involving EWSR1/FUS with members of the cAMP response element binding protein (CREB) family (ATF1, CREB1 and CREM) drive the pathogenesis of multiple tumor types spanning a wide clinicopathologic spectrum and risk of malignancy 1. Among them, angiomatoid fibrous histiocytoma (AFH) is one of the tumors driven by EWSR1::CREB1 fusion, which typically arises in the superficial soft tissues of extremities in young adults and follows a benign clinical course in most cases 2. However, rare examples are associated with aggressive behavior and distant metastatic disease 3. In a recent study, we showed that CDKN2A/B homozygous deletions were the only recurrent secondary events in EWSR1::CREB1 fusion-positive AFH and present exclusively in metastatic AFH cases 1. One of these cases also showed a co-occurring TERT amplification. Targeted therapy in the context of translocation-associated sarcomas has been focused mainly on the oncogenic fusion and not on the secondary genetic alterations, in part because most secondary genetic events in fusion-positive sarcomas are not readily targetable. Moreover, effective therapies and clinical responses have only been encountered in targeting kinases or growth factors-associated fusions, but not transcription factors 4,5. In this study we report a remarkable clinical response in a metastatic AFH by targeting the BRAF V600E secondary mutation using a combination of BRAF and MEK inhibitors. As in vitro models of AFH harboring EWSR1::CREB1 fusion have been recently generated in human embryonic stem-derived mesenchymal progenitors (hES-MP) cells, we used this clinical case as a template for a double mutant (EWSR1::CREB1 fusion, BRAF V600E) preclinical model, as proof of principle for therapeutic targeting with the combination of BRAF and MEK inhibitors.
RESULTS
Index Patient Clinicopathologic Findings.
The patient presented with a painless right thigh mass, complicated by spontaneous rupture and subsequent regrowth. By MRI in 03/2020, it appeared as a 7 × 6.8 cm intramuscular hemorrhagic lesion, suspicious for hematoma. The mass was resected and microscopically was composed mainly of hemorrhagic spaces in keeping with a hematoma. The lesion recurred in 5 months and was re-resected in 08/2020 showing a 10.6 × 6.7 cm mass (Fig. 1). The recurrent lesion showed solid areas, composed of ovoid to epithelioid histiocytoid cells with pale cytoplasm and uniform nuclei with open chromatin. Although no nuclear pleomorphism or necrosis were discerned, the tumor showed brisk mitotic activity of 9 MF/10 HPFs (Fig. 2 A). MSK-Archer assay was performed on the first local recurrence, through which the correct AFH diagnosis was rendered. Findings showed an EWSR1::CREB1 transcript from the fusion of EWSR1 exons 1–7 with CREB1 exons 7–8 (Fig. 2 B) that is present in 90% of AFH cases 6,7. The MSK-IMPACT analysis on the same specimen confirmed the presence of the oncogenic fusion, a BRAF V600E mutation and a deletion of the CDKN2A/B locus. The BRAF mutation was further confirmed by IHC with a mutation-specific antibody (Fig. 2A, right panel). The patient developed two additional multifocal local recurrences and an adrenal metastasis by 08/2021. A biopsy of the adrenal mass confirmed metastatic disease. Further metastatic disease included liver, bone, soft tissue, and lung.
Figure 1. Radiologic imaging of local recurrence and adrenal metastasis pre- and post-therapy with encorafenib and binimetinib combination.
Upper panels show MRI of the right femur showing 2nd local recurrence (asterisk) with fluid-fluid level (left); large increase in size of the primary tumor mass in the thigh over one month period (center); and significant decrease in size after targeted therapy (right). Lower panels show CT scans for the same timepoints, with small adrenal gland metastasis (asterisk, left), with large increase in size (center) and subsequent decrease in size after targeted therapy (right).
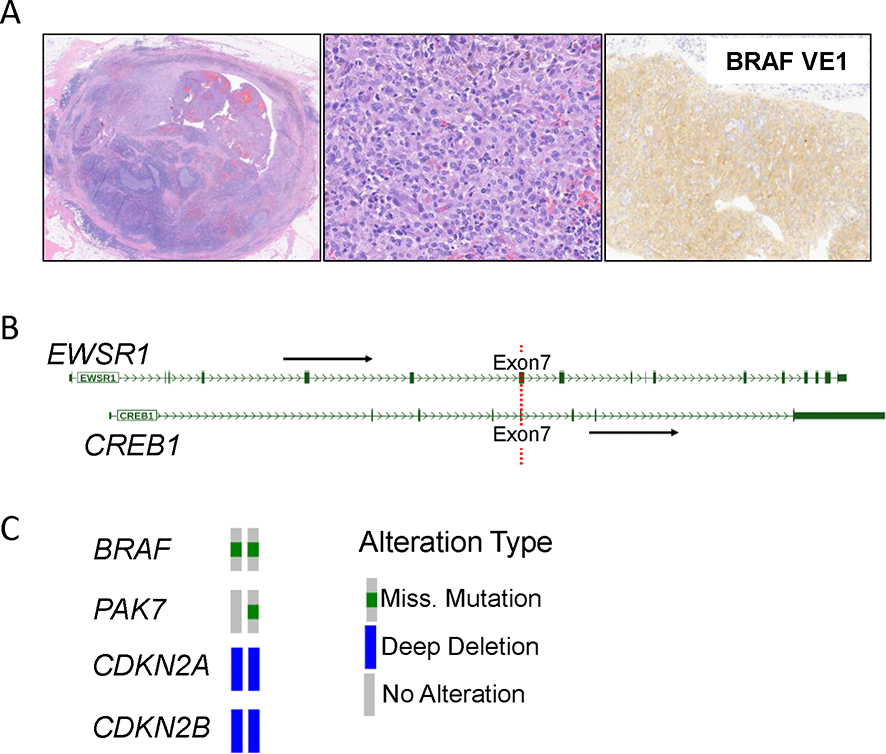
Figure 2. Pathologic features and molecular findings.
(A) Histologic images from the 2nd local recurrence showing at low power (left) a partly cystic-hemorrhagic and partly solid mass surrounded by a prominent rim of lymphoid aggregates. At higher magnification (center) the tumor is composed of sheets of plump histiocytoid cells with pale eosinophilic cytoplasm and reniform nuclei with open chromatin and mild atypia. Immunohistochemical stain with BRAF VE1 antibody shows diffuse, strong cytoplasmic positivity in the tumor cells (right). (B) Archer FusionPlex showed the presence of a fusion transcript composed of EWSR1 exon7 fused to CREB1 exon7. (C) IMPACT testing from the first local recurrence and the adrenal metastasis showed clonal BRAF V600E mutation and a CDKN2A/B deletion. The EWSR1::CREB1 fusion was also identified. The adrenal metastasis showed in addition a PAK7 mutation.
Treatment History and Follow-up
She was readmitted on 07/2021 with hemoglobin (Hgb) of 5 g/dL, evidence of bleeding both into the primary thigh tumor site and hemorrhagic adrenal metastasis, transfused and treated with 1 cycle of vincristine-doxorubicin-cyclophosphamide therapy for one cycle. After discharge, she was admitted again within two weeks with fever, anemia with Hgb 5 g/dL, and treated again with transfusion. No source of infection found but fever continued despite recovery of blood counts. IL6 was found elevated at 2814 pg/mL (upper limit of normal [ULN] 5), as well as ferritin 650 ng/ml (ULN 200), erythrocyte sedimentation rate > 100 mm/hr (ULN 20), and C-reactive protein elevated at 22.8 ng/mL (ULN 0.3). She was then started on tocilizumab for presumed paraneoplastic fever. Given further disease worsening despite chemotherapy, with knowledge of the BRAF V600E mutation, she started encorafenib and binimetinib on 08/2021 at FDA approved doses. Two months later, the thigh lesions demonstrated RECIST 1.1 response by imaging, fevers had dissipated, inflammatory markers improved (IL6 9.7 pg/mL, CRP 1.24 ng/mL), while other disease in adrenal gland was stable (Fig. 1). As recurrence/metastases were stable by imaging 2 and 4 months later, tocilizumab was discontinued, but shortly after on 03/2022 she presented again with fever, decrease in Hgb to 7.8 g/dL, increase in hemorrhagic adrenal metastasis and bloody ascites as evidence of disease worsening, with a progression-free survival of 7.0 months. Inflammatory markers were again markedly elevated with IL6 1038 and CRP 23.9. Radiation was used to treat the now 9 cm adrenal metastasis with stabilization in her hemoglobin. One month later, fevers continued, and she was again treated with tocilizumab with anaphylactoid reaction, managed successfully with a desensitization program. With further bleeding of the adrenal metastasis, she was taken to surgery for palliative adrenalectomy with improvement in her symptoms. MSK-IMPACT testing was repeated on the post-treatment adrenal gland metastasis, where a new PAK7 missense mutation of unknown significance was identified (Fig. 2C). She did well briefly, before presenting with chest wall pain, bone metastasis to rib and to T3 spine. She was treated with radiation to the bony metastatic sites. Encorafenib and binimetinib were continued, and she was considered for involvement on a phase 1 clinical trial of a novel BRAF inhibitor. The poor response to VAC chemotherapy vs prior response to binimetinib-encorafenib was the principal consideration for not pursuing traditional ifosfamide-based chemotherapy at that time. Despite being eligible, she opted to not enroll on the trial, due to complaints of fatigue and the desire to recover more post-radiation. Shortly thereafter, she presented with nausea, vomiting, abdominal pain and new liver metastatic disease and gastric outlet obstruction from external compression. Hyperbilirubinemia from liver metastatic disease and multisystem organ failure quickly ensued. She succumbed to metastatic disease on 08/2022, 30 months after diagnosis.
Generation and characterization of EWSR1::CREB1/ BRAF V600E double mutant model in hES cell lines.
We previously demonstrated that expression of the EWSR1::CREB1 fusion in hES-MP recapitulates the transcriptional signature of human AFH tumor. Therefore, we introduced the BRAF V600E mutation in hES-MP carrying the EWSR1::CREB1 translocation to study the effect to BRAF and MEK inhibitor in this in vitro model. The mutation was introduced by CRISPR-Cas9 method in human embryonic stem (hES) cells harboring the EWSR1 (exon7)::CREB1 (exon7) translocation using a 140 nt long single stranded DNA (ssDNA) with the nucleotide change c.1799 T>A. For screening purposes, a silent mutation disrupting the AluI restriction site was also introduced (Appendix Fig. 1A). Colonies from single cells were screened with allele specific primers followed by universal PCR and AluI digestion before sequencing to confirm the BRAF mutation (Appendix Fig. 1B). We were able to select one clone with homozygous and another with heterozygous mutation. In the heterozygous clone, a +1 bp insertion was identified in the second BRAF allele, corresponding to the double strand break (DSB) site. The BRAF V600E mutation is frequently heterozygous, even though the incidence of homozygous BRAF V600E mutations in tumors may be underestimated 8,9. By MSK-IMPACT analysis, the primary and metastatic tumor samples showed variant allele frequencies (VAF) of 33.9% and 33.5% for BRAF V600E, respectively, in keeping with a heterozygous mutation (Fig. 2C). Thus, we selected the heterozygous BRAF clone for further characterization. The expression of the BRAF V600E mutated protein was confirmed by western blot analysis (Appendix Fig. 1C) and the induction of the EWSR1::CREB1 fusion confirmed after expression of Cre recombinase by RT-PCR analysis (Appendix Fig. 1D).
Expression of the EWSR1::CREB1 fusion in BRAF V600E-mutant mesenchymal progenitor does not affect cell viability.
AFH, like many other sarcoma entities, is a tumor of mesenchymal origin. In prior studies we have characterized and validated the transcriptional profile of AFH tumors and selected the top upregulated genes as the ‘AFH core gene signature’ 6,10. The list included 4 genes: serum and glucocorticoid-regulated kinase 1 (SGK1), matrix-remodeling-associated protein 5 (MXRA5), solute carrier family 7 member 5 (SLC7A5) and dual specificity phosphatase 4 (DUSP4). This AFH signature was used as a reference in validating the reproducibility of our in vitro hES and hES-MP cell models after fusion induction, as compared to the human disease 10.
In hES-MPs the EWSR1::CREB1 fusion expression is retained for up to 30 days before cell growth arrest, with TP53 deletion only partially rescuing the phenotype 10. We differentiated BRAF-mutant hES cells to hES-MP 10,11 and expressed the EWSR1::CREB1 fusion by Cre transduction 12. The presence of the BRAF V600E mutation did not confer a growth advantage, with complete loss of the fusion transcript by 32 days post-induction (Fig. 3A). In the same experiment, the genes of the AFH signature were upregulated in a fusion-dependent manner, as demonstrated by the progressive loss of expression in the time course experiment (Fig. 3B). A similar result was obtained in cells transfected with a plasmid expressing the Cre recombinase and the mCherry marker with enrichment of fluorescent positive cells by FACS 10. (Appendix Fig. 2A and B). Overall results suggest that, in our in vitro model, the BRAF V600E oncoprotein does not rescue the survival of hES-MP expressing the EWSR1::CREB1 fusion protein.
Figure 3. EWSR1::CREB1 fusion expression and AFH core gene signature in BRAF V600E mutant mesenchymal cells.
(A) RT-PCR for EWSR1::CREB1 transcript and (B) qRT-PCR for AFH signature genes in a time course experiment after transduction with Cre recombinase in mesenchymal cells harboring the BRAF V600E mutation. Histograms represent the fold increase compared to -Cre condition at day 5 from 3 independent experiments. Statistical significance is calculated with a paired t-test comparing +Cre with the corresponding -Cre condition. *p<0.05, **p<0.01; when not indicated the difference is not statistically significant.
BRAF V600E mutation in hES-MP cells drives upregulation of genes involved in cellular transformation.
Whole transcriptome sequencing (RNAseq) was performed on BRAF V600E hES-MP cells 6 days after the expression of Cre recombinase and compared with cells expressing the fusion alone. Among the top upregulated genes, we identified PTPRN, a gene coding for a protein tyrosine phosphatase receptor involved in cell growth, differentiation, mitosis, and oncogenic transformation. Other genes included PLAT (plasminogen activator), VCAN (versican), and TGFB1 (transforming growth factor beta induced) with roles in cell migration, adhesion, and proliferation (Appendix Fig. 3A). Moreover, in the gene set enrichment analysis (GSEA), we identified enrichment in MAPK and TP53 pathways (Appendix Fig. 3B).
Cell viability after Inhibition of BRAF and MEK by encorafenib and binimetinib
To confirm that the hES-MP cells harboring the BRAF mutation respond to BRAF and MEK inhibitors, cells were exposed for 7 days to increasing concentration of encorafenib and binimetinib showing increased sensitivity, compared to control isogenic cell line (Fig. 4A). Cells were then subjected to a combination treatment using 250 nM encorafenib and 25nM binimetinib, with increased sensitivity compared to single treatment (Fig. 4B). The sensitivities of BRAF V600E mutant cells did not significantly changed after induction of the EWSR1::CREB1 fusion. (Fig. 4C). Finally, an increase in Annexin-V signal was observed in cells with BRAF mutation treated with binimetinib and with the combination, even if in cells expressing the fusion the increase is not statistically significant (Fig. 4D).
Figure 4. Viability and induction of apoptosis subsequent to encorafenib and binimetinib treatment in BRAF V600E mutant mesenchymal cells expressing EWSR1::CREB1 fusion protein.
(A) Cell viability of BRAF V600E mutant cells and parental cells after 7-day continuous treatment with increasing concentrations of encorafenib or binimetinib. (B) Cell viability after 7-day treatment with encorafenib (250 nM) and binimetinib (25 nM) in mesenchymal cells prior to induction of the EWSR1::CREB1 fusion. (C) Cell viability in a pool of Cre-transfected mesenchymal cells after 7-day treatment with encorafenib or binimetinib, at day 5 or day 12 post-transfection (e.v. empty vector). (D) Annexin V measurement of apoptosis in cells transfected with the Cre recombinase after 72-hour treatment with single dose of encorafenib, binimetinib or both. Each panel error bar represents the standard deviation from at least 3 independent experiments. For panel B and D unpaired t test * p< 0.05, ** p < 0.01, if not indicated the difference was not significant.
DISCUSSION
A long-standing challenge in sarcoma medical oncology is the variability of treatment response observed even in patients with similar histotypes, including translocation-associated sarcomas, suggesting that diverse co-occurring secondary alterations may impact on the clinical benefit to the various drug regimens applied. Sequencing the tumor genomes are paramount to unveil the underlying unique somatic alterations, particularly when the driver canonical fusion remains untargetable or otherwise chemo or radio-resistant, and thus other genomic alterations may be potentially targetable 13.
A recent study, investigating 1964 patients with soft tissue sarcoma by NGS, identified 11 patients (0.6%) harboring BRAF V600E mutations, among a large spectrum of histotypes. Among these, only one case of fusion-positive sarcoma was encountered (i.e. synovial sarcoma), while the remaining were soft tissue sarcomas with mostly complex genomic landscape 14. Moreover, like our case, the most co-occurring alteration in BRAF-mutant sarcomas was loss of CDKN2A in half of the cases, followed by TERT promoter mutations. A different study investigated the response of 4 sarcoma cell lines harboring BRAF V600E mutation, including one fusion-positive Ewing sarcoma (A673), to vemurafenib inhibition 15. Vemurafenib induced a strong cytostatic effect only in one liposarcoma cell line, but this result was achieved at a higher dose compared to BRAF V600E mutated melanoma cells. Although a limited growth inhibition was observed in the other three cell lines, reduced ERK phosphorylation was present in all, confirming the vemurafenib-induced MAPK pathway inhibition, despite continued cell proliferation. Of interest, although both BRAF V600E mutation and CDKN2A/B deletions were detected in the pre-therapy local recurrence, the post-therapy metastatic sample showed a new PAK7 missense mutation (p.H371R) of uncertain significance. PAK7 (a.k.a. PAK5) belongs to the p21-activated serine/threonine kinase (PAK) family and non-synonymous mutations in this gene have been documented in multiple cancers, with an incidence of up to 15% in melanomas 16.
The clinical presentation of this patient also highlights the challenge in making a correct diagnosis of AFH, which in this case both the radiographic and pathologic impressions was that of an organizing hematoma with rupture and locoregional spread. An additional unexpected adverse event during the patient clinical course was the development of a paraneoplastic cytokine release syndrome with elevated IL6. While fevers and elevations of cytokines are sometimes seen with bulky cancer metastases, they are not appreciated as a common event with AFH. The syndrome was reversed by immunosuppression using tocilizumab, an anti-IL-6 receptor antibody. Cytokine release syndrome (CRS), with elevated circulating levels of several cytokines including interleukin (IL)-6 and interferon γ, is a potentially life-threatening toxicity that has been observed following administration of natural and bispecific antibodies and adoptive T-cell therapies for cancer 17. The CRS symptoms seen in this patient appeared to be related to tumor recurrence/metastasis rather than treatment, which, if anything, in controlling the tumor also controlled the inflammatory syndrome 18–20. Although CREB1 and ATF1 members of the CREB family of transcription factors may participate interchangeably in gene fusions within the same tumor type, there is a striking propensity for the EWSR1::CREB1 fusion to occur in AFH and EWSR1::ATF1 in soft tissue clear cell sarcoma (CCS), while in gastrointestinal CCS (GI-CCS) the two fusion events occur with similar prevalence 1. AFH is a mesenchymal neoplasm of borderline malignant potential, typically occurring in the superficial soft tissues of children and adolescents 6 2.
In order to explore the significant clinical and pathologic promiscuity triggered by the EWSR1::CREB family of fusions, we have generated faithful preclinical models of AFH to model the EWSR1::CREB1 fusion in human embryonic stem-derived mesenchymal progenitor (hES-MP) cells 10. Several attempts to model fusion relevant sarcomas in non-transformed cell lines resulted in reduced cell proliferation and premature loss of cell viability 21,22. The induction of the EWSR1::CREB1 fusion in hES cells, while recapitulating the transcriptional profiles found in human AFH, with upregulation of SGK1, MXRA5, and CTSB, lead to premature cell growth arrest and loss of viability that was partially rescued by differentiation to hES-MP 10. Translocation-driven sarcomas have few or no secondary alterations and their role in transformation remains to be elucidated 1,23 22. We have previously demonstrated that TP53 deletion in hES-MP expressing the EWSR1::CREB1 oncogenic fusion, extends cell proliferation/viability. The BRAF V600E mutation despite its role in the constitutive activation of the MEK/ERK pathway and in cellular transformation in different tumor types 24 was not enough to induce cell proliferation in our AFH model. We have previously demonstrated that the induction of the EWSR1::CREB1 fusion in hES-MP cells induces overexpression of CDKN1A 10 the gene coding for p21, a regulator of cell cycle and target of TP53. The same upregulation was shown by RNAseq analysis on BRAF V600E cells. We hypothesize that in our in vitro model, the EWSR1::CREB1 protein activates various pathways inhibiting the cellular progression/proliferation and that a single mutation or deletion (e.g., BRAF V600E or TP53) is not enough to fully bypass this control.
Finally, treatment with either BRAF or MEK inhibition (encorafenib/binimetinib) was effective in reducing the viability of the cell line, with the combination treatment increasing the sensitivity. As many transcription factors, the EWSR1::CREB1 fusion protein is not directly druggable highlighting the importance of targeted sequencing on these genetically stable tumors to identify not recurrent secondary mutation that can exploit therapeutic vulnerabilities.
MATERIALS AND METHODS
Case Report.
The clinical and radiologic findings at diagnosis, local recurrence and metastasis and treatment modalities were retrieved from the patient clinical chart. The histologic slides together with immunohistochemical stains were re-reviewed by a sarcoma pathologist (CRA). The patient provided written consent for samples collection (IRB# 02–060).
Archer Assay.
Details of MSK-Fusion have been previously described 25. The solid fusion assay is a targeted RNA-based panel that utilizes the Archer Anchored Multiplex PCR (AMP) technology and next-generation sequencing to detect gene fusions in solid tumor and sarcoma samples.
MSK IMPACT.
Detailed descriptions of MSK-IMPACT workflow and data analysis, a hybridization capture-based targeted NGS assay using matched tumor and blood-derived normal DNA have been previously described 26. Oncoprint was generated from the cBio Cancer Genomics Portal 27.
Mammalian cell culture.
All experiments were approved by the Tri-SCI Embryonic Stem Cell Research Oversight Committee (ESCRO). Human embryonic stem cells (WA01, H1) are available from WiCell under a material transfer agreement. hES and hES-MP were cultured and maintained as previously described 11.
Preparation of ssDNA fragment and sgRNA plasmid.
3 μg of single strand DNA (ssDNA) fragment containing the mutations was synthesized by Genewiz and resuspended at 100 μg/ml. sgRNA sequences (5’-caccgtagctacagtgaaatctcga-3’, 5’-aaactcgagatttcactgtagctac-3’) were cloned into the dual Cas9/sgRNA expression vector pSpCas9(BB)-2A-GFP (PX458) (Addgene #48138) 28.
Generation of cell lines.
hES cells harboring the EWSR1 (exon 7)::CREB1 (exon 7) translocation were transfected by Amaxa nucleofector (Program A-024) with 1 μg of ssDNA and 3 μg of plasmid expressing sgRNA and Cas9 protein. GFP positive cells were sorted after 3 days and replated as single cells until colonies formation. The hES cells were differentiated to mesenchymal progenitors (hES-MP) as previously described 10.
PCR, RT-PCR and qRT-PCR analysis.
DNA and RNA were extracted as previously described 12. PCR and RT-PCR were performed using DreamTaq Green PCR master mix (Thermo Scientific). qRT-PCR was done as previously described 10. Primers are listed in Appendix Table 1, Appendix Table 2 and Appendix Table 3.
Cre expression.
hES cells were transfected and hES-MP transduced with Cre recombinase as previously described 10. Lipofectamine Stem Transfection Reagents (Invitrogen, STEM00015) were used for sensitivity and apoptosis assays.
Western blot.
Protein extraction and Western blot were performed as previously described 10. Antibodies are listed in Appendix Table 4.
Treatment with BRAF and MEK inhibitors.
Cells were treated for 7 days at increasing concentration of encorafenib (MedChem Express 502260171) and binimetinib (Selleck Chemicals S7007) and viability measured with CellTiter-Glo (Promega, G9242). In cells expressing the EWSR1::CREB1 fusion, the treatment started 5 days after nucleofection with Cre recombinase.
Apoptosis assay.
Cells were collected after 72 hrs drugs treatment and labeled using FITC Annexin V kit (Biolegend, 640905) and propidium iodide (PI), followed by flow cytometry. Data are presented as ratio of the percentage of AnnexinV+PI positive untreated versus treated cells.
RNA sequencing.
RNAseq was performed on BRAF V600E cells after fusion induction. All reads were aligned with STAR (ver 2.3) and BowTie2 against the human reference genome (hg19) 29. Differentially expressed genes (DEG) (log2 FC >2) from comparison with cells expressing the fusion alone were identified.
GSEA Analysis.
The gene lists were subjected to gene set enrichment analysis (GSEA) using Bioconductor clusterProfiler package to identify significant pathways 30. The curated gene sets and ontology gene sets 31 were used as the pathway database.
Statistical analysis.
Numerical data are shown as the mean ± s.d and differences between groups were determined using unpaired or paired t-test (in Graphpad). p-value <0.05 was considered statistically significant. If not specified, the analysis is not significant.
CONTEXT.
Key Objective.
Analyze the role of secondary alterations in fusion-driven sarcomas by generating faithful in vitro models for functional characterization.
Knowledge generated.
The clinical findings and the treatment history of a patient with a diagnosis of AFH sarcoma expressing the pathognomonic EWSR1::CREB1 fusion and the BRAF V600E mutation are described. A human cell line recapitulating the genetic and physiologic features of the tumor is generated and used to confirm the sensitivity to BRAF and MEK inhibitor treatment.
Relevance.
Our work demonstrates the importance of identifying targetable secondary mutations in difficult to treat-tumors to exploit therapeutic vulnerabilities and provides an invaluable tool for preclinical studies and drug discovery.
Funding Statement.
This work was supported by P50 CA217694 (CRA), P30 CA008748 (CRA, SH, RGM), Cycle for Survival (CRA), Kristen Ann Carr Foundation (CRA), St Baldrick’s Foundation (CRA). All other authors report no funding sources related to this study.
Appendix
Appendix Figure 1. Strategy for generating BRAF V600E mutation in hES cells and screening method.
(A) BRAF gene diagram (exon 15), highlighting the sgRNA (green box) and PAM (purple box) sequences. The Alu-I cutting site and codon for Valine amino acid are underlined. After HDR integration of ssDNA carrying the 2 mutant nucleotides (red), the Alu-I cutting site is lost, while the codon for the valine residue is substituted with the codon for glutamic acid. (B) Schematic representation of the PCR strategies for screening the BRAF V600E mutant. In the allele specific PCR, the primer pair is composed of a universal primer and a mutation-specific primer, which will amplify the mutant alleles. The positive clones were confirmed by an amplification across the targeted site and PCR product digestion with Alu-I. (C) Western blot with BRAF VE1 antibody on the hES BRAF V600E mutant clone used for experiments presented in Figs. 3 and 4. (D) EWSR1::CREB1 fusion RT-PCR showing comparable transcript levels in the BRAF V600E-mutant hES cells compared to the isogenic cell line.
Appendix Figure 2. Time course for EWSR1::CREB1 fusion and AFH core gene signature expression in Cre-transfected cells after sorting.
(A) RT-PCR and (B) qRT-PCR as described in Figure 3. BRAF V600E-mutant mesenchymal cells and isogenic cell lines transfected with a plasmid expressing the Cre-recombinase and an mCherry fluorescent marker. Three days after transfection, mCherry positive cells were sorted and replated for time course experiment and analysis. Histograms represent the fold increase compared to -Cre condition at day 6 of 1 experiment.
Appendix Figure 3. RNAseq analysis on hES-MP BRAF V600E cells expressing the EWSR1-CREB1 fusion.
(A) Histogram plots showing upregulation of PTPRN, PLAT, VCAN and TGFBI in BRAF V600E mutant cells expressing the EWSR1::CREB1 fusion compared to cells expressing fusion alone and (B) GSEA on deregulated genes from the same analysis.
Appendix Table 1.
List of primers for PCR analysis.
| PCR | Sequence |
|---|---|
| Allele-specific PCR | Forward 5’-CTCTTCATAATGCTTGCTCTG-3’ |
| Reverse 5’- CCATCGAGATTTCtCTGTgG-3’ * | |
| Universal PCR | Forward 5’-CCAATGAAGAGCCTTTACTGCT-3’ |
| Reverse 5’-GCATCTCAGGGCCAAAAAT-3’ |
Nucleotide in red indicate the point mutation introduced with ssDNA
Appendix Table 2.
List of primers for RT-PCR analysis.
| Gene/Gene Fusion | Forward Primer | Reverse Primer |
|---|---|---|
| EWSR1(ex7)::CREB1(ex7) gene fusion | 5’-CCTACCAGCTATTCCTCTACAC-3’ | 5’-CAGAGGCAGCTTGAACAAC-3’ |
| GAPDH ex6–8 gene | 5’-GAGGGGCCATCCACAGTCTTCT-3’ | 5’-GGAGCCAAAAGGGTCATCATCT-3’ |
Appendix Table 3.
List of primers for qRT-PCR analysis.
| Gene | Forward Primer | Reverse Primer |
|---|---|---|
| SGK1 ex4 | 5’-AGGATGGGTCTGAACGACTTT-3’ | 5’-GCCCTTTCCGATCACTTTCAAG-3’ |
| MXRA5 ex4 | 5’-TCAACGGCTTAACGTCTCTGA-3’ | 5’-CACGGATTTCCCTGCAAGTAA-3’ |
| SLC7A5 ex8 | 5’-CTTCTTCAACTGGCTCTGCG-3’ | 5’-CGATGCCACACTCCACG-3’ |
| DUSP4 ex2 | 5’-GCGGCTATGAGAGGTTTTCCTC-3’ | 5’-TGGTCGTGTAGTGGGGTCC-3’ |
| GAPDH ex6–8* | 5’-GAGGGGCCATCCACAGTCTTCT-3’ | 5’-GGAGCCAAAAGGGTCATCATCT-3’ |
Same sequences as in Appendix Table 2
Appendix Table 4.
List of antibodies for western blot analysis
| Antibody | Dilution |
|---|---|
| BRAF VEI (Abcam 228461) | 1:400 in 5% milk |
| Clathrin (BD Biosciences #610499) | 1:20,000 in TBS (0.1% Tween, 5% BSA) |
Footnotes
Conflict of Interest Statement
All authors report no conflict of interests related to this study.
Ethics Approval / Consent to Participate
This study was approved by the Memorial Sloan Kettering Cancer Institute Institutional Review Board.
References
- 1.Dermawan JK, Vanoli F, Herviou L, et al. : Comprehensive genomic profiling of EWSR1/FUS::CREB translocation-associated tumors uncovers prognostically significant recurrent genetic alterations and methylation-transcriptional correlates. Mod Pathol 35:1055–1065, 2022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Rekhi B AC, Chen G. : WHO Classification of Tumors Editorial Board. Soft Tissue and bone tumors. International Agency for Research on Cancer, 2020 [Google Scholar]
- 3.Cazzato G, Lupo C, Casatta N, et al. : Angiomatoid Fibrous Histiocytoma (AFH) of the Right Arm: An Exceptional Case with Pulmonary Metastasis and Confirmatory EWSR1::CREB1 Translocation. Diagnostics (Basel) 12, 2022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Kummar S, Shen L, Hong DS, et al. : Larotrectinib efficacy and safety in adult patients with tropomyosin receptor kinase fusion sarcomas. Cancer 129:3772–3782, 2023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Beaziz J, Battistella M, Delyon J, et al. : Long-Term Outcome of Neoadjuvant Tyrosine Kinase Inhibitors Followed by Complete Surgery in Locally Advanced Dermatofibrosarcoma Protuberans. Cancers (Basel) 13, 2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Antonescu CR, Dal Cin P, Nafa K, et al. : EWSR1-CREB1 is the predominant gene fusion in angiomatoid fibrous histiocytoma. Genes Chromosomes Cancer 46:1051–60, 2007 [DOI] [PubMed] [Google Scholar]
- 7.Rossi S, Szuhai K, Ijszenga M, et al. : EWSR1-CREB1 and EWSR1-ATF1 fusion genes in angiomatoid fibrous histiocytoma. Clin Cancer Res 13:7322–8, 2007 [DOI] [PubMed] [Google Scholar]
- 8.Sapkota B, Hill CE, Pollack BP: Vemurafenib enhances MHC induction in BRAF(V600E) homozygous melanoma cells. Oncoimmunology 2:e22890, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Rubinstein JC, Sznol M, Pavlick AC, et al. : Incidence of the V600K mutation among melanoma patients with BRAF mutations, and potential therapeutic response to the specific BRAF inhibitor PLX4032. J Transl Med 8:67, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Vanoli F, Meskauskaite B, Herviou L, et al. : Generation of human embryonic stem cell models to exploit the EWSR1-CREB fusion promiscuity as a common pathway of transformation in human tumors. Oncogene 40:5095–5104, 2021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Vanoli F, Herviou L, Tsuda Y, et al. : Generating in vitro models of NTRK-fusion mesenchymal neoplasia as tools for investigating kinase oncogenic activation and response to targeted therapy. Oncogenesis 12:8, 2023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Vanoli F, Tomishima M, Feng W, et al. : CRISPR-Cas9-guided oncogenic chromosomal translocations with conditional fusion protein expression in human mesenchymal cells. Proc Natl Acad Sci U S A 114:3696–3701, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Iyer G, Hanrahan AJ, Milowsky MI, et al. : Genome sequencing identifies a basis for everolimus sensitivity. Science 338:221, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kobayashi H, Zhang L, Okajima K, et al. : BRAF mutations and concurrent alterations in patients with soft tissue sarcoma. Genes Chromosomes Cancer 62:648–654, 2023 [DOI] [PubMed] [Google Scholar]
- 15.Gouravan S, Meza-Zepeda LA, Myklebost O, et al. : Preclinical Evaluation of Vemurafenib as Therapy for BRAF(V600E) Mutated Sarcomas. Int J Mol Sci 19, 2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.LaPak KM, Vroom DC, Garg AA, et al. : Melanoma-associated mutants within the serine-rich domain of PAK5 direct kinase activity to mitogenic pathways. Oncotarget 9:25386–25401, 2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee DW, Gardner R, Porter DL, et al. : Current concepts in the diagnosis and management of cytokine release syndrome. Blood 124:188–95, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Villiger PM, Cottier S, Jonczy M, et al. : A simple Baker’s cyst? Tocilizumab remits paraneoplastic signs and controls growth of IL-6-producing angiomatoid malignant fibrous histiocytoma. Rheumatology (Oxford) 53:1350–2, 2014 [DOI] [PubMed] [Google Scholar]
- 19.Potter SL, Quintanilla NM, Johnston DK, et al. : Therapeutic response of metastatic angiomatoid fibrous histiocytoma carrying EWSR1-CREB1 fusion to the interleukin-6 receptor antibody tocilizumab. Pediatr Blood Cancer 65:e27291, 2018 [DOI] [PubMed] [Google Scholar]
- 20.Markham SK, Mani A, Bauer J, et al. : Surface Texturing Design to Enhance Echogenicity of Biopsy Needles During Endoscopic Ultrasound Imaging. Ultrasound Med Biol 46:2453–2463, 2020 [DOI] [PubMed] [Google Scholar]
- 21.Brunet E, Simsek D, Tomishima M, et al. : Chromosomal translocations induced at specified loci in human stem cells. Proc Natl Acad Sci U S A 106:10620–5, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sole A, Grossetete S, Heintze M, et al. : Unraveling Ewing Sarcoma Tumorigenesis Originating from Patient-Derived Mesenchymal Stem Cells. Cancer Res 81:4994–5006, 2021 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tirode F, Surdez D, Ma X, et al. : Genomic landscape of Ewing sarcoma defines an aggressive subtype with co-association of STAG2 and TP53 mutations. Cancer Discov 4:1342–53, 2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Ascierto PA, Kirkwood JM, Grob JJ, et al. : The role of BRAF V600 mutation in melanoma. J Transl Med 10:85, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhu G, Benayed R, Ho C, et al. : Diagnosis of known sarcoma fusions and novel fusion partners by targeted RNA sequencing with identification of a recurrent ACTB-FOSB fusion in pseudomyogenic hemangioendothelioma. Mod Pathol 32:609–620, 2019 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cheng DT, Mitchell TN, Zehir A, et al. : Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): A Hybridization Capture-Based Next-Generation Sequencing Clinical Assay for Solid Tumor Molecular Oncology. J Mol Diagn 17:251–64, 2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Cerami E, Gao J, Dogrusoz U, et al. : The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–4, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Vanoli F, Jasin M: Generation of chromosomal translocations that lead to conditional fusion protein expression using CRISPR-Cas9 and homology-directed repair. Methods 121–122:138–145, 2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Antonescu CR, Le Loarer F, Mosquera JM, et al. : Novel YAP1-TFE3 fusion defines a distinct subset of epithelioid hemangioendothelioma. Genes Chromosomes Cancer 52:775–84, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Yu G, Wang LG, Han Y, He QY: clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16:284–7, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Subramanian A, Tamayo P, Mootha VK, et al. : Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–50, 2005 [DOI] [PMC free article] [PubMed] [Google Scholar]