Abstract
Background and Aims
We aimed to identify serum metabolites associated with mucosal and transmural inflammation in paediatric Crohn disease [pCD].
Methods
In all, 56 pCD patients were included through a pre-planned sub-study of the multicentre, prospective, ImageKids cohort, designed to develop the Paediatric Inflammatory Crohn magnetic resonance enterography [MRE] Index [PICMI]. Children were included throughout their disease course when undergoing ileocolonoscopy and MRE and were followed for 18 months, when MRE was repeated. Serum metabolites were identified using liquid chromatography/mass spectroscopy. Outcomes included: PICMI, the simple endoscopic score [SES], faecal calprotectin [FCP], and C-reactive protein [CRP], to assess transmural, mucosal, and systemic inflammation, respectively. Random forest models were built by outcome. Maximum relevance minimum redundancy [mRMR] feature selection with a j-fold cross-validation scheme identified the best subset of features and hyperparameter settings.
Results
Tryptophan and glutarylcarnitine were the top common mRMR metabolites linked to pCD inflammation. Random forest models established that amino acids and amines were among the most influential metabolites for predicting transmural and mucosal inflammation. Predictive models performed well, each with an area under the curve [AUC] > 70%. In addition, serum metabolites linked with pCD inflammation mainly related to perturbations in the citrate cycle [TCA cycle], aminoacyl-tRNA biosynthesis, tryptophan metabolism, butanoate metabolism, and tyrosine metabolism.
Conclusions
We extend on recent studies, observing differences in serum metabolites between healthy controls and Crohn disease patients, and suggest various associations of serum metabolites with transmural and mucosal inflammation. These metabolites could improve the understanding of pCD pathogenesis and assessment of disease severity.
Keywords: Serum metabolites, machine learning, paediatric Crohn disease
Graphical Abstract
Graphical Abstract.
1. Introduction
Inflammatory bowel diseases [IBD], including Crohn disease [CD] and ulcerative colitis [UC], are debilitating gastrointestinal ailments with no permanent cure. These conditions are becoming more frequent, especially in paediatric patients.1 paediatric IBD [pIBD] is characterised by a more extensive and complicated disease course and increased risk of cancer and surgery.2
The pathogenesis of IBD is complex, involving genetic, host immune, and environmental factors, including the gut microbes. Metabolites produced by intestinal microbiota and through host-interacting metabolic processes may represent relevant pathogenic processes for host immune responses.3 Serum and faecal metabolites are capable of distinguishing pCD from paediatric UC [pUC] and from healthy controls.4 Moreover, metabolites have been proposed to aid in the diagnosis of IBD.5
Despite the potential applicability of metabolomics in increasing the understanding of the underlying pathogenesis of IBD, retrieving meaningful information related to host immune responses remains highly challenging.6 Bennike et al.,5 reported the status of proteomics IBD biomarkers and suggested the use of mass spectrometry for identifying and characterising novel biomarkers. Nanni et al.,7 used linear discriminant analysis [LDA] to classify serum samples among healthy subjects and IBD patients. Nikolaus et al.,8 studied the association between metabolites in IBD patients and clinical and serological features, finding that serum levels of tryptophan were significantly lower in patients with IBD. Importantly, there is a need to employ tools other than just univariate statistics, linear models, and correlation techniques, and particularly, to consider the highly correlated structure of metabolomic data, to reduce the probability of obtaining false positives and false negatives.
Hence, we aimed to assess relationships between serum metabolites and inflammation in a prospective pCD cohort using machine learning techniques, based on multivariate methods that consider all metabolites simultaneously. We hypothesizsd that serum metabolites could provide a unique opportunity to characterise pCD inflammation by identifying a range of molecules and metabolic changes. Furthermore, this is the first study to analyse metabolites in association with not only mucosal inflammation, but also transmural inflammation, measured by a validated Paediatric Inflammatory Crohn MRE Index [PICMI] for pCD.9
2. Materials and Methods
2.1. Study population
This is a planned sub-study of the prospective multi-centre ImageKids study, which aimed to develop the PICMI score through a cohort of 240 children [aged 6 to 18 years] with CD. To ensure inclusion of the full breadth of disease stages, patients were recruited a priori by strata of disease duration: 20% 0–3 months from diagnosis, 20% 3–24 months from diagnosis, 20% 24–36 months from diagnosis, and 40% thereafter.9 Patients were recruited at 22 centres in Europe, North America, Asia, and Australia, when undergoing both ileocolonoscopy and MRE. Half of the recruited children were prospectively followed for 18 months, when MRE and clinical/laboratory assessments were repeated. Only patients with paired serum specimens collected at study initiation [baseline] and follow-up [18 months] were included [56 patients; Figure 1].
Figure 1.
Study flowchart.
2.2. Outcomes
C-reactive protein [CRP] is a widely used serum biomarker that reflects systemic inflammation.10 Faecal calprotectin [FCP] is a commonly used biomarker that indicates bowel inflammation.11 The Simple Endoscopic Score for Crohn Disease [SES-CD] reflects the grade of severity of endoscopic mucosal inflammation.12 The Paediatric Inflammatory Crohn’s MRE Index [PICMI] directly reflects transmural inflammation in paediatric CD [pCD].9
Determinants of study outcome [PICMI, SES-CD, FCP, and CRP] are detailed in Focht et al.9 The total PICMI score is a cumulative sum of the weighted sub-scores of all the segments involved in CD in small and large bowel. Discriminative validity and determination of the four cut-off values of disease categories by PICMI [≤ 10 remission, 11–55 mild inflammation, 56–120 moderate inflammation, and >120 severe inflammation], were previously validated.9
2.3. Serum metabolites processing
Serum samples were analysed at the Metabolomics Innovation Centre [TMIC; University of Alberta] using the TMIC Prime Assay. Consequently, samples underwent a targeted, quantitative, metabolomics approach following a combination of direct injection mass spectrometry with a reverse-phase LC-MS/MS custom assay. This custom assay, in combination with an ABSciex 4000 QTrap [Applied Biosystems/MDS Sciex] mass spectrometer, was used to identify and quantity 130 different endogenous metabolites including amino acids, acylcarnitines, biogenic amines and derivatives, uraemic toxins, glycerophospholipids, sphingolipids, and sugars.13,14
Isotope-labelled and other internal standards were used for metabolite quantification. The custom assay contained a 96-deep–well plate with a filter plate attached with sealing tape, and reagents and solvents used to prepare the plate assay. The first 14 wells were used for one blank, three zero samples, seven standards, and three quality control samples. For all metabolites except organic acid, samples were thawed on ice and were vortexed and centrifuged at 13 000x g; 10 µL of each sample was loaded onto the centre of the filter on the upper 96-well plate and dried in a stream of nitrogen. Subsequently, phenyl-isothiocyanate was added for derivatisation. After incubation, the filter spots were dried again using an evaporator. Extraction of the metabolites was then achieved by adding 300 µL of extraction solvent. The extracts were obtained by centrifugation into the lower 96-deep–well plate, followed by a dilution step with MS running solvent.
For organic acid analysis, 150 µL of ice-cold methanol and 10 µL of isotope-labelled internal standard mixture was added to 50 µL of serum sample for overnight protein precipitation, which was then centrifuged at 13 000x g for 20 min; 50 µL of supernatant was loaded into the centre of wells of a 96-deep–well plate, followed by the addition of 3-nitrophenylhydrazine [NPH] reagent. After incubation for 2 h, BHT stabiliser and water were added before LC-MS injection.
Mass spectrometric analysis was performed on an ABSciex 4000 Qtrap® tandem mass spectrometry instrument [Applied Biosystems/MDS Analytical Technologies, Foster City, CA] equipped with an Agilent 1260 series UHPLC system [Agilent Technologies, Palo Alto, CA]. The samples were delivered to the mass spectrometer by an LC method followed by a direct injection [DI] method. Data analysis was done using Analyst 1.6.2.
2.4. Statistical analysis
The dataset included 130 serum metabolite concentrations at baseline and 18 months’ follow-up for each patient.
The human metabolome is very dynamic and time dependent, as the rate of chemical reactions occurring in the cell is affected by multiple biological processes.15 Agueusop et al.16 demonstrated the short-term variability of the human serum metabolome under differing nutritional and metabolic health status. Therefore, and due to the long time between the two study visits with multiple changing parameters that may affect serum metabolites [eg, medical therapy and diet], the two different visits were grouped in one dataset. To validate this assumption, we performed a baseline analysis, demonstrating that the relationship between the disease state in the intestine and the metabolomic profile does not appear to be affected by advanced therapies [biologics] on immune pathways affecting the metabolites. Supplementary Figure S1 shows the results of the analysis performed with a subgroup of patients at baseline who were not subjected to any biologic treatment.
Resulting from the brevity of the dataset, the Moderate and Severe class labels from PICMI score are underrepresented, emanating from the problem of unbalanced classes. Therefore, we clustered ‘Remission + Mild’ and ‘Moderate + Severe’, defined by acute inflammation and moderate inflammation, to allow for two contrasting groups [Supplementary Figure S2]. Table 1 summarises the cut-offs and outcomes. Participants were excluded from a particular model if data on that outcome variable were missing.
Table 1.
Outcomes and cut-offs.
| Categorical outcomes | Cut-offs PICMI | Cut-offs FCP μg/g of faeces |
Cut-offs CRP mg/L |
Cut-offs SES-CD |
|---|---|---|---|---|
| Moderate/severe disease | >56 | >250 | >7 | ≥ 7 |
| Inactive/mild disease | ≤55 | ≤250 | ≤7 | <7 |
CRP, C-reactive protein; FCP, faecal calprotectin; PICMI, Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index; SES-CD, simple endoscopic score.
Statistical analyses were performed using R [v4.1.2] and Python [v. 3.7.4] using numpy [v. 1.16.4], pandas [v. 0.25.1], and scikit-learn [v. 0.21.2] packages.
We used the uniform manifold approximation and projection technique [UMAP] to analyse the metabolomics sample distribution between patients with moderate/severe and inactive/mild bowel inflammation.
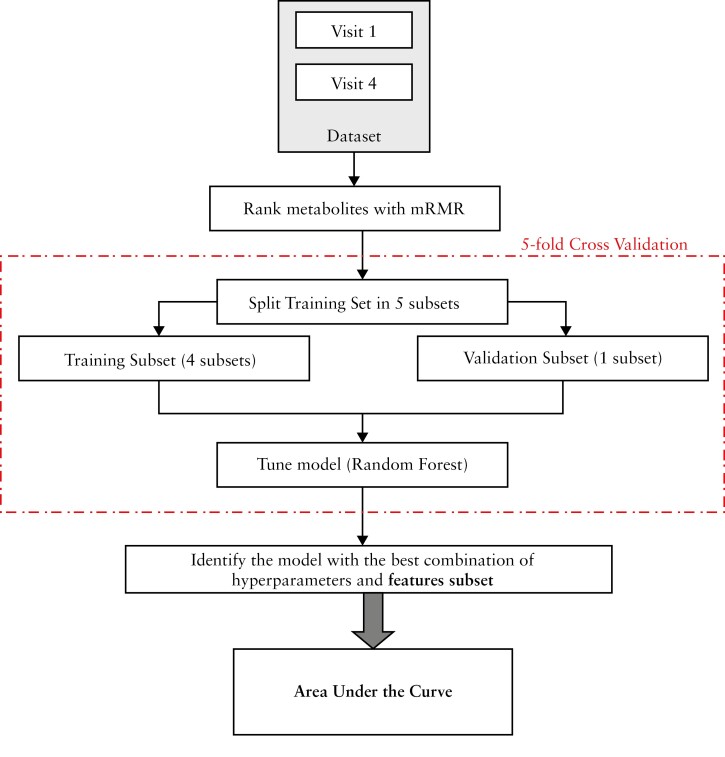
Using PICMI, SES-CD, FCP, and CRP outcomes, we employed a typical supervised machine learning pipeline [Figure 2] to build four random forest [RF] models to predict moderate/severe and inactive/mild inflammation from metabolomics composition. We performed a minimum redundancy maximum relevance [mRMR] feature selection approach to examine which metabolites are most associated with inflammation.17 We selected RF, as this model has been useful in evaluating the importance of features in previous omics studies.18–20 To learn which combination of metabolites can contribute to predicting inflammation, we divided the training set into five, equal-sized, disjoint subsets to perform 5-fold cross-validation. During this process, we left out one subset [validation subset], and combined the other four subsets to form the training subset. For more information regarding the learning process, please refer to Supplementary Figure S3.
Figure 2.
Set of action for model learning and evaluation. Maximum relevance minimum redundancy [mRMR] feature selection and a cross-validation scheme were used to optimise the model [and find the best subset of features and hyperparameter settings]. We then assess its performance, as measured by the area under its Receiver Operating Characteristic [ROC] curve.
The models were evaluated, yielding an area under the receiver operating curve [AUC-ROC]. We quantified the importance of specific metabolites with impurity-based importance [mean decrease Gini index] and ranked a list of metabolites with their relative importance to the model.
Finally, we used data from each visit [ie, study initiation at baseline and 18 months follow-up] to identify if clinical outcomes [ie, PICMI, FCP, and CRP] improved between timepoints. For FCP and CRP, we used the cut-off values presented in Table 1. For PICMI, any movement in the ordered categories defined by the cut-off values in the total PICMI score9 [≤10 remission, 11–55 mild inflammation, 56–120 moderate inflammation, and >120 severe inflammation] was used to define the ‘improving’ label [eg, moving from moderate to mild inflammation from baseline to follow-up].
The metabolomic composition from the ‘improving’ category was evaluated using non-metric multidimensional scaling [NMDS] according to the Bray–Curtis distance. In addition, we used the permutational multivariate analysis of variance [PERMANOVA] to test group differences.
2.5. Data availability
Raw metabolite data are available within the article’s Supplementary materials. Please contact corresponding authors for further data on the cohort.
3. Results
A total of 56 children met the eligibility criteria and were included in the study [Table 2]. Importantly, FCP and CRP measurements were available for only some patients, and SES-CD was only available at baseline [Table 2]. Consequently, for FCP, CRP, and SES-CD, the number of patients included in the analysis is lower than the number of patents included in PICMI analysis.
Table 2.
Characteristics of ImageKids cohort with serum samples available in baseline and follow-up [count [%], medians [minimum and maximum range], and means ± standard deviation, are reported as appropriate].
| Baseline visit data | Follow-up visit at 18 months data |
p-Valuea Baseline vs follow-up |
|
|---|---|---|---|
| Number of patients [n] | 56 | 56 | |
| Age [years] | 14.8 2.5 | 16.3 2.5 | 0.0005 |
| Male sex | 33 [59%] | ||
| BMI [%] | 18.5 3.4 | 20.1 3.6 | 0.01 |
| FCP μg/g of faeces | 869.5 [19.5, 4252.5] | 583.9 [19.5, 2152.0] | 0.1 |
| Remission/mild disease | 11 [20%] | 16 [29%] | |
| Moderate/severe disease | 39 [70%] | 18 [32%] | |
| Missing | 6 [10%] | 22 [39%] | |
| CRP mg/L | 22.8 [0.2, 188.0] | 7.5 [0.2, 115.0] | 0.005 |
| Remission/mild disease | 21 [38%] | 37 [67%] | |
| Moderate/severe disease | 27 [48%] | 13 [23%] | |
| Missing | 8 [14 %] | 6 [10%] | |
| PICMI | 65.9 [0.0, 282.0] | 41.8 [0.0, 210.0] | 0.02 |
| Remission/mild disease | 31 [55%] | 39 [70%] | |
| Moderate/severe disease | 25 [45%] | 17 [30%] | |
| SES-CD | 10.4 [0.0, 34.0] | - | |
| Remission/mild disease | 23 [42%] | - | |
| Moderate/severe disease | 23 [42%] | - | |
| Missing | 10 [18%] | ||
| wPCDAI | 31.5 [0.0, 80.0] | 6.8 [0.0, 25.0] | 0.0001 |
| Disease duration at enrolment | |||
| 0–3 months | 13 [23%] | ||
| 3–24 months | 14 [25%] | ||
| 24–36 months | 6 [10%] | ||
| >36 months | 23 [42%] | ||
| Country of origin | |||
| Israel | 19 [34%] | ||
| Canada | 16 [29 %] | ||
| USA | 6 [11%] | ||
| Germany | 3 [5%] | ||
| Spain | 8 [14 %] | ||
| Poland | 3 [5%] | ||
| Australia | 1 [2 %] | ||
| Disease locationb | |||
| L1 [ileal] | 11 [19%] | ||
| L2 [colonic] | 8 [14%] | ||
| L3 [ileocolonic] | 33 [59%] | ||
| L4 [upper disease] | |||
| L4a | 14 [25%] | ||
| L4b | 8 [14%] | ||
| P [perianal disease] | 2 [4%] | ||
| Disease behaviourb,c | |||
| B1 [inflammatory] | 43 [77%] | ||
| B2 [stricturing] | 9 [16%] | ||
| B3 [penetrating] | 3 [5%] | ||
| B2B3 [both penetrating and stricturing] | 1 [2%] | ||
| Medicationc | |||
| Steroids | 6 [10%] | 5 [8%] | |
| Thiopurine | 15 [27%] | 14 [25%] | |
| Methotrexate | 4 [7%] | 3 [5%] | |
| Biologics | 31 [56%] | 34 [62%] | |
| Enteral nutrition | 7 [12%] | 8 [14%] | |
BMI, body mass index; CRP, C-reactive protein; FCP, faecal calprotectin; PICMI, Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index; SES-CD, simple endoscopic score; wPCDAI, weighted Pediatric Crohn Disease Activity Index.
aBy Mann–Whitney U test.
bBy the Paris Classification.
cAt time of measurement.
3.1. Ranking serum metabolites in association with pCD inflammation
Global metabolomic profiling did not separate patients by disease duration strata [data not shown]. Then, metabolites from patients with paired samples were combined and UMAP analysis was used to cluster profiles by disease severity. Metabolomic samples did not cluster effectively by severity state, as measured by PICMI score, SES-CD, FCP, or CRP in embedded projections [Supplementary Figure S4].
We then used a typical, supervised, machine learning pipeline to maximise the separation capacity by disease severity [Figure 2]. Using the mRMR feature selection method, we identified the 30 most significant endogenous metabolites differentiating moderate/severe from inactive/mild classes. The classes were categorised using the activity indices [SES-CD and PICMI] and biomarkers [CRP and FCP], given that each reflects different aspects of CD activity and could be associated with different metabolomic profiles. The Venn diagram in Figure 3 displays the logical relationship of the highest scoring features in the dataset by PICMI score, SES-CD, FCP, and CRP. Importantly, tryptophan and glutarylcarnitine are metabolites common to PICMI, SES-CD, FCP, and CRP.
Figure 3.
Logical relationship of the 30 top mRMR metabolites. Each sphere contains the 30 top mRMR metabolites by outcome. The overlaps between the spheres represents the intersection of common top mRMR metabolites. The metabolites appearing in red have been previously reported to be associated with IBD at the Human Metabolome Database [HMD]. mRMR, maximum relevance minimum redundancy; IBD, inflammatory bowel disease. The Venny (Venny 2.1.0 [csic.es ]) online tool was used to generate the Venn Diagram.
In order to focus on the outcomes that are most relevant for detecting active inflammation, we investigated the relationship between the highest scoring metabolites by PICMI and SES-CD. Supplementary Figure S5 shows the intersection of the top 30 mRMR metabolites by PICMI and SES-CD outcomes. This result reveals the overlap between carnitine, acetate, and butyrate derivatives.
From the learning process, we found that the list of appropriate subsets of features to include for predicting inflammation contained 32 metabolites for PICMI, CRP, and FCP, and 65 for SES-CD. The complete list of feature subsets for each model is presented in Supplementary Table S2.
To rank the importance of the metabolites presented in Table S2, we quantified the predicted contribution by metabolite. We present a variable importance plot for each model. These plots rank the top 30 metabolites in descending predicting importance from 0 to 1, with 1 being the most important. Betaine [p < 0.05], tetradecadienylcarnitine [p < 0.05], trimethylamine N-oxide [p < 0.01], and sphingomyelin 16:1OH [p < 0.01] are the most influential predictors by PICMI, SES-CD, CRP, and FCP, respectively. Glutarylcarnitine appeared as an important predictor in all models [Figure 4]. The performances of the models were evaluated using an area under the receiver operating curve [AUC-ROC]. The model given by CRP class performed best with an AUC of 0.83 [Supplementary Figure S6].
Figure 4.
Importance of metabolites determined by the different prediction models. Top ranking metabolites are expressed with their relative importance for [A] Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index [PICMI], [B] simple endoscopic score [SES-CD], [C] C-reactive protein [CRP], and [D] faecal calprotectin [FCP]. The differences between the classes were tested using Mann–Whitney U test; *p < 0.05; **p < 0.01.
3.2. Serum metabolites and biological pathways
We then investigated the relationship between the top predicting serum metabolites, disease phenotypes, and biological pathways, by performing a metabolite set enrichment analysis [MSEA] by using the web-based platform MetaboAnalyst [https://www.metaboanalyst.ca]. The enriched pathway analysis showed a significant association [p < 0.05] with inflammation [Supplementary Figure S7.1, S7.2, and S7.4] and with coeliac disease [Supplementary Figure S7.3]. Interestingly, recent studies have reported an overlap between IBD and coeliac disease pathogenesis.21
We also employed the metabolic pathway analysis [MetPA] module of MetaboAnalyst. A list of the pathways identified and their relative impact is presented in Figure 5. The significant pathways [p < 0.05] in Figure 5 showed that the metabolites associated with pCD mainly resulted in perturbations in the citrate cycle [TCA cycle], aminoacyl-tRNA biosynthesis, tryptophan metabolism, butanoate metabolism, and tyrosine metabolism.
Figure 5.
Metabolic pathway analysis [MetPA] as generated by MetaboAnalyst based on the important metabolites. Matched pathways are displayed as circles. The colour and size of each circle are based on p-value and pathway impact value, respectively. A] MetPA by Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index [PICMI] metabolites. B] MetPA by simple endoscopic score [SES-CD] metabolites. C] MetPA by C-reactive protein [CRP] metabolites. D] MetPA by faecal calprotectin [FCP] metabolites. Pathway analysis as generated by MetaboAnalyst web-based platform (MetaboAnalyst).
3.3. Serum metabolites associated with clinical improvement
There were significant differences between disease severity from baseline to the follow-up visit [Table 2]. Therefore, we used the categories for PICMI are given in ref 9 and for FCP and CRP in Table 1 to identify patients with improved clinical outcomes from baseline [T0] to follow-up [T1] visit. The non-metric multidimensional scaling [NMDS] according to the Bray–Curtis distance showed two clusters separated by the metabolome profile of patients at baseline and follow-up visit [Figure 6].
Figure 6.
Embedded projections demonstrate a zone of improvement between baseline and follow-up visit. Metabolomic profiles given by [A] Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index [PICMI], [B] C-reactive protein [CRP], and [C] faecal calprotectin [FCP] classes.
We paired the data in the model and tested the differences in metabolome within each participant across visits. The metabolite signatures at baseline and follow-up visits were not significantly different [PERMANOVA p-values > 0.05]; however, the p-values are low enough to suggest an interesting result regarding the metabolome profiles from the two timepoints that visually clustered into two groups [Figure 6]. Figure 6 also shows that most of patients at follow-up visit had significantly reduced PICMI, CRP, and FCP [Table 2]. Therefore, to further investigate differences in the metabolomic profiles between the two groups, we performed partial least square discriminate analysis [PLS-DA] to calculate the variable importance in projection [VIP]. The serum metabolites that contributed the most to the classification of the groups were identified using the VIP score [Figure 7].
Figure 7.
Variable of Importance [VIP] scatter score plots depicting the most significant metabolites [VIP > 1.0] that explain the metabolite composition in the improvement group. Metabolomic composition given by [A] Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index [PICMI], [B] C-reactive protein [CRP], and [C] faecal calprotectin [FCP] classes. The relative abundance of metabolites is indicated by a coloured scale from blue to red representing the low and high, respectively. VIP score calculated using MetaboAnalyst web-based platform (MetaboAnalyst).
Metabolites with greater VIP scores contributed more to the separation between the two groups. Spermidine, spermine, putrescine, valerylcarnitine, and tryptophan were important metabolites in the discrimination of patient profiles which presented improvement from baseline to follow-up visit. To identify shared down-regulated pathways of the metabolites present in Figure 7 we employed the metabolic pathway analysis [MetPA] module of MetaboAnalyst. The significant pathways [p < 0.05] in Supplementary Figure S8 showed that the metabolites associated with pCD improvement mainly resulted in perturbations in beta-alanine metabolism, arginine metabolism, and butanoate metabolism.
4. Discussion
This is the first study to associate serum metabolites with both mucosal and transmural inflammation. This was done by including patients with pCD with different levels of activity using the PICMI score, reflecting transmural inflammation. Serum metabolite profiles have the potential to aid in understanding the course of CD and its relation to severity.22 They can distinguish between active IBD disease and remission, as shown in adults,23 and can discriminate paediatric CD from UC and healthy controls.4
We observed that tryptophan associated with both mucosal and transmural inflammation. In agreement with our findings, tryptophan was lower in the blood of IBD patients compared with controls.24 Moreover, Nikalaus and colleagues8 used standard statistics to demonstrate that increased tryptophan metabolism is associated with CD and UC activity. Given our results and prior evidence suggesting that tryptophan might play a role in IBD pathogenesis,8,25 further investigation into the role of tryptophan in pCD mucosal and transmural inflammation is warranted.
Other serum metabolites have consistently been observed with possible disease-associated metabolic changes in IBD patients. Table 3 presents a summary of studies that link our results [Figure 3] with previously reported metabolic changes and include the group of patients and the type of sample.
Table 3.
Differential abundance of amino acids, amines, tricarboxylic acid cycle intermediates, acylcarnitines, and sphingolipids in IBD patients.
| Metabolites | Associated class [Figure 4] | Patients | Sample | Reference |
|---|---|---|---|---|
| Amino acids | ||||
| Proline | CRP | IBD vs control | Serum | 26 |
| Histidine | PICMI, CRP, FCP | IBD vs control | Serum | 27 |
| Tyrosine | PICMI and SES | IBD vs control | Serum | 26 |
| Isoleucine | CRP | IBD vs control | Serum | 28 |
| Tryptophan | PICMI, SES, CRP, FCP | IBD vs control | Serum | 24 |
| Symmetric dimethylarginine | CRP | IBD vs control | Serum | 29 |
| Amines | ||||
| Trimethylamine-N-oxide [TMAO] | PICMI, SES, CRP | IBD vs control | Serum and Plasma | 30 |
| Choline | CRP and FCP | CD vs UC | Serum | 28 |
| Tricarboxylic acid cycle intermediates | ||||
| alpha-Ketoglutaric | SES and CRP | CD vs UC and control | Plasma | 23 |
| Succinic acid | PICMI | CD vs UC and control | Plasma | 23 |
| Citric acid | PICMI, CRP, FCP | Active IBD vs IBD in remission | Plasma | 27 |
| Acylcarnitines | ||||
| Valerylcarnitine | PICMI and CRP | CD vs UC | Plasma | 31 |
| Hexadecenoylcarnitine | PICMI and CRP | CD vs UC | Plasma | 31 |
| Sphingolipids | ||||
| Sphingomyelin | PICMI, FCP, CRP | pCD vs control | Plasma | 32 |
CD, Crohn Disease; CRP, C-reactive protein; FCP, faecal calprotectin; IBD, inflammatory bowel disease; pCD, pediatric Crohn Disease; PICMI, Pediatric Inflammatory Crohn Magnetic Resonance Enterography Index; SES-CD, simple endoscopic score; UC, ulcerative colitis.
Interestingly, Supplementary Figure 5S shows that carnitine, acetate, and butyrate derivatives associate with both transmural and mucosal inflammation. Consequently, previous studies from serum samples have reported increased levels of carnitine derivatives in CD patients.31 Other metabolites that are commonly disturbed in IBD patients are short-chain fatty acids [acetate and butyrate]. Increased levels of beta-hydroxybutyric acid and p-hydroxyphenylacetic acid are associated with intestinal inflammation relief,33,34 whereas decreased levels are associated with gut inflammation in IBD patients.35
For transmural inflammation, we found that some amino acids and lysophosphatidylcholines related metabolites were among the highest predictors [Figure 4A]. Betaine has been previously related with intestinal barrier functions36 and lysophosphatidylcholines are lipid intermediates involved in immune signalling which have been previously linked to the intestinal mucus of IBD patients.37 For mucosal inflammation, we found that acylcarnitines and tricarboxylic acid cycle intermediates [TCA] related metabolites were among the highest predictors [Figure 4B]. In fact, acylcarnitines might be responsible for direct immune cell activation and can exacerbate the inflammation of some tissues.38
The results from the analysis performed with CRP reinforced the importance of amino acids, acylcarnitines, and TCA metabolites in IBD inflammation. Trimethylamine-N-oxide [TMAO], tetradecanoylcarnitine, and citric acid were among the highest predictors in the CRP model [Figure 4C]. Finally, some sphingolipids ranked highly in the FCP model [Figure 4D]. Consequently, reduced levels of certain sphingolipids, such as sphingomyelin, have been reported in pCD patients.32 We found that serum metabolites possess a good discriminatory power to predict pCD mucosal and transmural activity [Supplementary Figure 6S] with areas under the curves up to 0.83. Therefore in the future, serum metabolite data should be considered, among other types of features [ie, clinical, imaging, etc.], for building machine learning tools to find the best combination of features to predict pIBD severity.
We found that metabolites were enriched in pathways related to inflammatory diseases and some bowel related disorders [Supplementary Figure S7]. The metabolic pathways identified in our study [Figure 5] suggest that pCD might induce changes in aminoacyl-tRNA biosynthesis, the citrate [TCA cycle], and tryptophan, tyrosine, and phenylalanine metabolism. The latter are aromatic amino acids that have been linked to IBD through several connections, including diet-microbiome interactions related to indole pathways.39
The longitudinal design of our study allowed us to explore changes of metabolites in association with concomitant changes in disease activity. We found that polyamide metabolites are associated with improvement in inflammation [Figure 7]. Spermidine, spermine, sarcosine, and putrescine are polyamides that are essential for cell homeostasis and have been previously related to regulation of inflammation40 and modulation of oxidative stress.41 Moreover, metabolites associated with pCD improvement mainly represented perturbations in beta-alanine metabolism and butanoate metabolism [Supplementary Figure 8S]. Importantly, beta-alanine has been previously related to attenuation of inflammation,42 and butanoate metabolism as a key pathway improving ulcerative colitis in paediatric patients.43
There are several limitations to our study, including the modest cohort size and the time lag between the two visits, as well as the uncontrolled use of other medical and dietary treatment and the effects these may have on immune response, and by extension on the blood metabolome. The fact that we considered each patient at each visit as an individual warrants caution in establishing causal associations but does improve power. Moreover, the models that were constructed and used in this study do not consider the combination [linear or non-linear] of relevant features such as sex, age, BMI, and medication use. It is important to be aware that the feature selection and feature importance processes are based on mathematical methods to obtain some level of accuracy, but do not necessarily establish casual relationships. Furthermore, the exploratory nature of this work allows for hypothesis generation, but larger studies are required to validate our findings. Therefore, our findings would have benefited from validation in a separate cohort, but prospective studies collecting such detailed data are uncommon, particularly in children.
The strengths of this study include the prospective follow-up of the cohort, central and double readings of the MRE using the PICMI, which is a standardised quantification tool in pCD, and the availability of a large array of serum metabolites. Moreover, the machine learning pipeline that was employed in this work enabled us to assess association between metabolites and disease activity by simultaneously including all metabolites as features. Also, the mRMR and cross-validation methods warranted overfitting while ensuring robustness of the models. Finally, to our knowledge this is the first time that metabolomic profiles are associated with transmural and mucosal inflammation in pCD patients, so that the aetiology basis is broadened.
Future work would ideally include a larger dataset with classes that are proportionally represented by the samples. Having more data might support our findings in a better way and deduce more relationships between the metabolites and the severity scale. A boon to us would be the ability to obtain data from several visits, for performing time-series analysis. This would provide insight into the specific set of metabolites that contribute to the change in the status of the patient from more severe to less severe. On the statistical front, we would like to concentrate on including other predictors such as clinical, microbiome, and medication use. This might help to build a classifier able to predict disease severity above chance. Finally, for future work we would recommend including microbiome with metagenomics data. This would allow indentification of hypothetical connections between gut microbial metabolites and human proteins with potential association with transmural and mucosal inflammation in pCD.
5. Conclusions
To conclude, we used a novel approach to identify serum metabolites related to pCD inflammation. We identified some amino acids, TCA, amines, acylcarnitines, and sphingolipids as important predictors for pCD inflammation. Betaine, tryptophan, and some lysophosphatidylcholines were among the highest predictors for transmural inflammation, whereas acylcarnitines metabolites were among the highest predictors for mucosal inflammation. Moreover, the metabolomic pathway analysis suggests that the aminoacyl-tRNA biosynthesis, citrate cycle [TCA cycle], butanoate, phenylalanine, tyrosine, and tryptophan are important metabolic processes for understanding the molecular mechanisms in inflammation and pCD.
The findings of this work aim to motivate further research to explore the metabolome profile differences between mucosal and transmural inflammation in pCD. Having a better characterisation of different metabolome profiles would help define pCD pathogenesis, further our understanding of pCD progression, and have more information for clinicians to select the most appropriate therapy for individuals.
Supplementary Data
Supplementary data are available at ECCO-JCC online.
Acknowledgements
We would like to thank the patients, families, coordinators, and investigators of the ImageKids study for allowing this work to take place, as well as the Metabolomic Innovation Centre, especially Mrs Rupasri Mandal.
Contributor Information
Ricardo G Suarez, Department of Paediatrics, University of Alberta, Edmonton, AB, Canada.
Namitha Guruprasad, Department of Computing Science, University of Alberta, Edmonton, AB, Canada.
Ganesh Tata, Department of Computing Science, University of Alberta, Edmonton, AB, Canada.
Zhengxiao Zhang, College of Ocean Food and Biological Engineering, Jimei University, Xiamen, Fujian, China.
Gili Focht, Juliet Keidan Institute of Paediatric Gastroenterology Hepatology and Nutrition, Hebrew University, Jerusalem, Israel.
Daniel McClement, Department of Physiology, University of Alberta, Edmonton, AB, Canada.
Víctor Manuel Navas-López, Paediatric Gastroenterology and Nutrition Unit, HRU, Málaga, Spain.
Sibylle Koletzko, Department of Paediatrics, Dr. von Hauner Children’s Hospital, LMU University Hospital, LMU Munich, Munich, Germany; Department of Paediatrics, Gastroenterology and Nutrition, School of Medicine, Collegium Medicum University of Warmia and Mazury, Olsztyn, Poland.
Anne M Griffiths, Division of Paediatric Gastroenterology, Hospital for Sick Children, Toronto, ON, Canada.
Oren Ledder, Juliet Keidan Institute of Paediatric Gastroenterology Hepatology and Nutrition, Hebrew University, Jerusalem, Israel.
Lissy de Ridder, Erasmus Medical Center, Sophia Children’s Hospital, Rotterdam, The Netherlands.
David Wishart, Department of Computing Science, University of Alberta, Edmonton, AB, Canada.
Ben Nichols, Department of Clinical Nutrition, University of Glasgow, Glasgow, UK.
Konstantinos Gerasimidis, Department of Clinical Nutrition, University of Glasgow, Glasgow, UK.
Dan Turner, Juliet Keidan Institute of Paediatric Gastroenterology Hepatology and Nutrition, Hebrew University, Jerusalem, Israel.
Eytan Wine, Department of Paediatrics, University of Alberta, Edmonton, AB, Canada; Department of Physiology, University of Alberta, Edmonton, AB, Canada.
Funding
The ImageKids study was supported by an educational grant from AbbVie, which was not involved in any part of the study including protocol preparation, data acquisition, analyses, or manuscript preparation. RGS was funded through a graudate student aware by the Women and Children Health Research Insititute at the University of Alberta.
Conflict of Interest
RS, NG, GT, ZZ, DM, DW, VMN, SK, OL, and BN have no conflict to disclose in relation to the content of this manuscript. GF reports consultation fees from Abbvie and Lilly. AMG reports consultant fees from Abbvie, Amgen, Bristol Myers Squibb, Lilly, Janssen, Merck, Pfizer; speaker fees from Abbvie, Janssen, Nestlé; investigator-initiated research grant from Abbvie. LdR received personal fees, consultation and congress support from Shire, Malinckrodt, Nestlé, Celltrion, Merck, Janssen biologics, Abbvie, and Pfizer. KG has received research grants and speaker fees from Nestle Health Sciences, Nutricia-Danone, Baxter, Mylan. DT reports consultation fee, research grant, royalties, or honorarium from Janssen, Pfizer, Hospital for Sick Children, Ferring, Abbvie, Takeda, Atlantic Health, Shire, Celgene, Lilly, Roche, ThermoFisher, BMS. EW reports personal fees from AbbVie, Janssen, Nestle Health Sciences, BioJamp, Pfizer, and Mead Johnson Nutrition.
Author Contributions
DT, GF, VMN, SK, AMG, OL, LdR, KG, and EW conceived the project and organised patient enrolment and sample collection. Metabolite data were processed by DW and analysed by RS, with support from NG, GT, DM, BN, and ZZ. RS and EW interpreted the results and wrote the manuscript, with critical revision and approval from all other authors.
References
- 1. Ng SC, Shi HY, Hamidi N, et al. Worldwide incidence and prevalence of inflammatory bowel disease in the 21st century: a systematic review of population-based studies. Lancet 2017;390:2769–78. [DOI] [PubMed] [Google Scholar]
- 2. Vernier–Massouille G, Balde M, Salleron J, et al. Natural history of paediatric Crohn’s disease: a population-based cohort study. Gastroenterology 2008;135:1106–13. [DOI] [PubMed] [Google Scholar]
- 3. Blacher E, Levy M, Tatirovsky E, Elinav E.. Microbiome-modulated metabolites at the interface of host immunity. J Immunol 2017;198:572–80. [DOI] [PubMed] [Google Scholar]
- 4. Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V.. Faecal and serum metabolomics in paediatric inflammatory bowel disease. J Crohns Colitis 2016;11:321–34. [DOI] [PubMed] [Google Scholar]
- 5. Bennike T, Birkelund S, Stensballe A, Andersen V.. Biomarkers in inflammatory bowel diseases: Current status and proteomics identification strategies. World J Gastroenterol 2014;20:3231–44. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Magnúsdóttir S, Thiele I.. Modeling metabolism of the human gut microbiome. Curr Opin Biotechnol 2018;51:90–6. [DOI] [PubMed] [Google Scholar]
- 7. Nanni P, Parisi D, Roda G, et al. Serum protein profiling in patients with inflammatory bowel diseases using selective solid-phase bulk extraction, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry and chemometric data analysis. Rapid Commun Mass Spectrom 2007;21:4142–8. [DOI] [PubMed] [Google Scholar]
- 8. Nikolaus S, Schulte B, Al-Massad N, et al. Increased tryptophan metabolism is associated with activity of inflammatory bowel diseases. Gastroenterology 2017;153:1504–16.e2. [DOI] [PubMed] [Google Scholar]
- 9. Focht G, Cytter-Kuint R, Greer MLC, et al. Development, validation, and evaluation of the paediatric inflammatory Crohn’s magnetic resonance enterography index from the ImageKids Study. Gastroenterology 2022;163:1306–20. [DOI] [PubMed] [Google Scholar]
- 10. Chen P, Zhou G, Lin J, et al. Serum biomarkers for inflammatory bowel disease. Front Med [Lausanne] 2020;7:1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Bjarnason I. The use of fecal calprotectin in inflammatory bowel disease. Gastroenterol Hepatol [N Y] 2017;13:53–6. [PMC free article] [PubMed] [Google Scholar]
- 12. Koutroumpakis E, Katsanos K.. Implementation of the simple endoscopic activity score in Crohn’s disease. Saudi J Gastroenterol 2016;22:183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Foroutan A, Guo AC, Vazquez-Fresno R, et al. Chemical composition of commercial cow’s milk. J Agric Food Chem 2019;67:4897–914. [DOI] [PubMed] [Google Scholar]
- 14. Foroutan A, Fitzsimmons C, Mandal R, et al. The bovine metabolome. Metabolites 2020;10:233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Færgestad EM, Langsrud Ø, Høy M, et al. Analysis of megavariate data in functional genomics. In: Comprehensive Chemometrics. Oxford: Elsevier; 2009: 221–78. [Google Scholar]
- 16. Agueusop I, Musholt PB, Klaus B, Hightower K, Kannt A.. Short-term variability of the human serum metabolome depending on nutritional and metabolic health status. Sci Rep 2020;10:16310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Ding C, Peng H.. Minimum redundancy feature selection from microarray gene expression data. J Bioinform Comput Biol 2005;03:185–205. [DOI] [PubMed] [Google Scholar]
- 18. Hageman J. Relevant metabolites’ selection strategies. In: Metabolomics Perspectives. Cambridge, MA: Elsevier; 2022: 381–98. [Google Scholar]
- 19. Acharjee A, Larkman J, Xu Y, Cardoso VR, Gkoutos GV.. A random forest based biomarker discovery and power analysis framework for diagnostics research. BMC Med Genomics 2020;13:178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Liang G. Altered gut bacterial and metabolic signatures and their interaction in inflammatory bowel disease. Synth Syst Biotechnol 2021;6:377–83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Shi Y, Feng S, Yan M, Wei S, Yang K, Feng Y.. Inflammatory bowel disease and celiac disease: a bidirectional Mendelian randomization study. Front Genet 2022;13:1–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Zhang A, Sun H, Wang X.. Serum metabolomics as a novel diagnostic approach for disease: a systematic review. Anal Bioanal Chem 2012;404:1239–45. [DOI] [PubMed] [Google Scholar]
- 23. Scoville EA, Allaman MM, Brown CT, et al. Alterations in lipid, amino acid, and energy metabolism distinguish Crohn’s disease from ulcerative colitis and control subjects by serum metabolomic profiling. Metabolomics 2018;14:17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Whiley L, Nye LC, Grant I, et al. Ultrahigh-performance liquid chromatography tandem mass spectrometry with electrospray ionization quantification of tryptophan metabolites and markers of gut health in serum and plasma—application to clinical and epidemiology cohorts. Anal Chem 2019;91:5207–16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Dudzińska E, Szymona K, Kloc R, et al. Increased expression of kynurenine aminotransferases mRNA in lymphocytes of patients with inflammatory bowel disease. Therap Adv Gastroenterol 2019;12:175628481988130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Schicho R, Shaykhutdinov R, Ngo J, et al. Quantitative metabolomic profiling of serum, plasma, and urine by 1 H NMR spectroscopy discriminates between patients with inflammatory bowel disease and healthy individuals. J Proteome Res 2012;11:3344–57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Dawiskiba T, Deja S, Mulak A, et al. Serum and urine metabolomic fingerprinting in diagnostics of inflammatory bowel diseases. World J Gastroenterol 2014;20:163–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Williams HRT, Willsmore JD, Cox IJ, et al. Serum metabolic profiling in inflammatory bowel disease. Dig Dis Sci 2012;57:2157–65. [DOI] [PubMed] [Google Scholar]
- 29. Owczarek D, Cibor D, Mach T.. Asymmetric dimethylarginine [ADMA], symmetric dimethylarginine [SDMA], arginine, and 8-iso-prostaglandin F2α [8-iso-PGF2α] level in patients with inflammatory bowel diseases. Inflamm Bowel Dis 2010;16:52–7. [DOI] [PubMed] [Google Scholar]
- 30. Wilson A, Teft WA, Morse BL, et al. Trimethylamine-N-oxide: a novel biomarker for the identification of inflammatory bowel disease. Dig Dis Sci 2015;60:3620–30. [DOI] [PubMed] [Google Scholar]
- 31. Lai Y, Xue J, Liu CW, et al. Serum metabolomics identifies altered bioenergetics, signaling cascades in parallel with exposome markers in Crohn’s disease. Molecules 2019;24:449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Daniluk U, Daniluk J, Kucharski R, et al. Untargeted metabolomics and inflammatory markers profiling in children with Crohn’s disease and ulcerative colitis—A Preliminary Study. Inflamm Bowel Dis 2019;25:1120–8. [DOI] [PubMed] [Google Scholar]
- 33. Wu X, Xu J, Li J, et al. Bacteroides vulgatus alleviates dextran sodium sulfate-induced colitis and depression-like behaviour by facilitating gut-brain axis balance. Front Microbiol 2023;14:1–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Chen G, Ran X, Li B, et al. Sodium butyrate inhibits inflammation and maintains epithelium barrier integrity in a TNBS-induced inflammatory bowel disease mice model. EBioMedicine 2018;30:317–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bauset C, Gisbert-Ferrándiz L, Cosín-Roger J.. Metabolomics as a promising resource identifying potential biomarkers for inflammatory bowel disease. J Clin Med 2021;10:622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Wu J, He C, Bu J, et al. Betaine attenuates LPS-induced downregulation of Occludin and Claudin-1 and restores intestinal barrier function. BMC Vet Res 2020;16:75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Ehehalt R, Wagenblast J, Erben G, et al. Phosphatidylcholine and lysophosphatidylcholine in intestinal mucus of ulcerative colitis patients. A quantitative approach by nanoelectrospray‐tandem mass spectrometry. Scand J Gastroenterol 2004;39:737–42. [DOI] [PubMed] [Google Scholar]
- 38. Dambrova M, Makrecka-Kuka M, Kuka J, et al. Acylcarnitines: nomenclature, biomarkers, therapeutic potential, drug targets, and clinical trials. Pharmacol Rev 2022;74:506–51. [DOI] [PubMed] [Google Scholar]
- 39. Ghiboub M, Boneh RS, Sovran B, et al. Sustained diet-induced remission in paediatric Crohn’s disease is associated with kynurenine and serotonin pathways. Inflamm Bowel Dis 2023;29:684–94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Bussière FI, Chaturvedi R, Cheng Y, et al. Spermine causes loss of innate immune response to helicobacter pylori by inhibition of inducible nitric-oxide synthase translation. J Biol Chem 2005;280:2409–12. [DOI] [PubMed] [Google Scholar]
- 41. Gobert AP, Wilson KT.. Polyamine- and NADPH-dependent generation of ROS during Helicobacter pylori infection: a blessing in disguise. Free Radic Biol Med 2017;105:16–27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Ostfeld I, Hoffman JR.. The effect of β-alanine supplementation on performance, cognitive function and resiliency in soldiers. Nutrients 2023;15:1039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Zhou Z, Cao J, Liu X, Li M.. Evidence for the butyrate metabolism as key pathway improving ulcerative colitis in both paediatric and adult patients. Bioengineered 2021;12:8309–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.