Abstract
Brain tumors, particularly glioblastoma (GBM), are devastating and challenging to treat, with a low 5-year survival rate of only 6.6%. Mouse models are established to understand tumorigenesis and develop new therapeutic strategies. Large-scale genomic studies have facilitated the identification of genetic alterations driving human brain tumor development and progression. Genetically engineered mouse models (GEMMs) with clinically relevant genetic alterations are widely used to investigate tumor origin. Additionally, syngeneic implantation models, utilizing cell lines derived from GEMMs or other sources, are popular for their consistent and relatively short latency period, addressing various brain cancer research questions. In recent years, the success of immunotherapy in specific cancer types has led to a surge in cancer immunology-related research which specifically necessitates the utilization of immunocompetent mouse models. In this review, we provide a comprehensive summary of GEMMs and syngeneic mouse models for adult brain tumors, emphasizing key features such as model origin, genetic alteration background, oncogenic mechanisms, and immune-related characteristics. Our review serves as a valuable resource for the brain tumor research community, aiding in the selection of appropriate models to study cancer immunology.
Keywords: adult brain tumor, genetically engineered mouse models, immunotherapy response, syngeneic implantation model
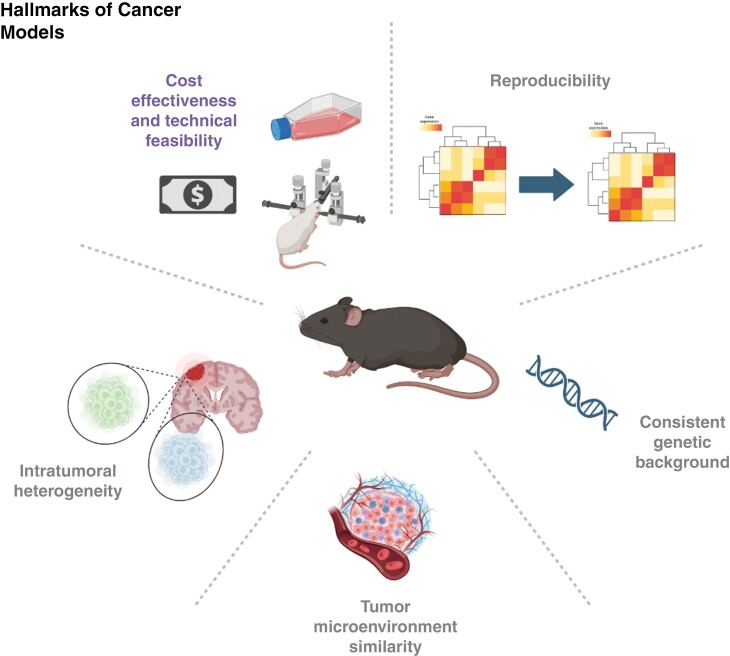
Brain tumor models underpin the investigation of tumor biology and the development of antitumor strategies. Consequently, ideal models should satisfy several requirements (see Figure 1): (I) genetic and; (II) tumor microenvironment (TME) features observed in patients; (III) intratumoral heterogeneity; (IV) reproducibility; and (V) cost-effectiveness and technical feasibility.1 However, available models remain imperfect due to insufficient recapitulation of the genetic and immune landscape of human tumors, at a reasonable cost and technical feasibility. Despite the advancement of three-dimensional (3D) ex vivo culture techniques, which better reflect tumoral architecture compared to traditional 2D cultures,2 in vivo models are indispensable in studying tumor–host immune interactions and developing immunotherapies. Tumor immunogenicity varies among patients with the same molecular cancer subtype and across different cancers, resulting in differential immune responses among the model diaspora.3 Tumor intrinsic and extrinsic factors influence immunotherapy efficacy. Increased tumor mutational burden (TMB) is hypothesized to promote tumor-specific neoantigens, thereby enhancing immune recognition and tumor destruction.4 Conversely, a hallmark of cancer is immune evasion often driven by oncogenic signaling.5 For instance, intrinsic factors like MYC/beta-catenin signaling promotes immune evasion through impaired dendritic cells (DCs) recruitment and T-cell function.6,7 Alternatively, loss of tumor suppressor function can impact host immune responses against cancer. For example, TP53 inactivation results in T-cells exclusion, and PTEN inactivation suppresses autophagy within the tumor.5 Additionally, extrinsic factors including tissue and tumor-associated microbiomes can influence oncogenesis by altering immune responses. Notably, the human gut microbiome is implicated in clinical responses to immune checkpoint blockade (ICB).8,9 The microbiome’s salutary effects may stem from its ability to activate innate and adaptive immunity to overcome tumor-mediated immunosuppression. Bacteroides cellulosilyticus was highly abundant in brain tumor patients who responded to ICB.9 Emerging evidence suggests that immune recognition of microbiota-derived molecules in the brain promotes T-regulatory cell migration to the CNS. The microbiome affects microglial maturation and function10and microbiome-derived short-chain fatty acids help regulate CNS homeostasis.11 Given that these factors affect antitumor immunity, it’s imperative for researchers to judiciously choose the appropriate mouse model that aligns with their research objectives.
Figure 1.
Factors that determine the rigor of a brain tumor model. Balancing technical and fiscal considerations, models ideally reflect the pathological features observed in patients such as the tumor microenvironment milieu, intratumoral heterogeneity, and the mutational landscape, in addition to being highly reproducible across research studies. Figure is generated by biorender.
In this comprehensive review, we provide an in-depth summary of genetically engineered mouse models (GEMMs) and syngeneic models specifically for adult brain tumors, with a specific emphasis on glioblastoma (GBM). For GEMMs, we extensively cover the methods of generation (Figure 2), the genetic background, and the potential impact of these oncogenes on the TME (also presented in Table 1). Syngeneic implantation models are given special attention by addressing the cell line origins and their driving genetic alterations. Furthermore, we summarize the immune-related characteristics and immunotherapy responses in syngeneic mouse models (see Table 2) to provide a thorough overview of each model’s immune aspects.
Figure 2.
Normal cells can transform into cancer cells through spontaneous carcinogenesis or exposure to carcinogenic chemicals and other agents. GEMMs can be created by introducing genetic material into normal cells using methods such as transposon-based systems, viral vectors, or electroporation. The temporal and spatial control of these genetic modifications allows for more accurate modeling of tumorigenesis
Table 1.
GEMM for Adult Brain Tumor
| Method | Oncogenic Factors | Tissue Specific | Medium Survival | Tumor Type | Genetic Background | Ref. | |
|---|---|---|---|---|---|---|---|
| SV40 T-Ag | Transgenic | SV40 T-Ag | GFAPR-drive | N/A | Astrocytomas, anaplastic astrocytomas |
C57BL/6 X DBA | 15 |
| PDGFR | Lentiviral deliver, CreERT2 | hPDGF-B, Cdkn2a kd | In situ injection | 60–80 days | Proneural GBM | C57BL/6J and 129S1, backcross to C57BL/6J | 28 |
| retroviral deliver | PDGF, ptenf/f, p53f/f | In situ injection | 27 days | C57BL/6J | 119 | ||
| Lentiviral deliver | PDGFA, STOPfl/fl-PDGFRα, TP53fl/fl | Spatiotemporal | 70 days | N/A | 27 | ||
| TP53 | Transgenic, CKO | p53∆E2–10/∆E2–10 or p53R172H, IDHWT,p53∆E5–6 | GFAP-cre | 256 days | GBM | C57BL/6 and 129S1/Svj | 120 |
| Transgenic, CKO | TP53–/–, NF1fl/fl | hGFAP-cre | 15 weeks | Grades 2–4 astrocytomas | C57BL/6, 129S4/SvJae,backcross to C57BL/6 | 31,32 | |
| TP53fl/+, NF1fl/fl | 22 weeks | GBM | |||||
| TP53fl/+, NF1+/fl | 30 weeks | GBM | |||||
| Transgenic, CKO | TP53–/–, NF1fl/fl; TP53–/+, NF1fl/+ | hGFAP-cre | 18 weeks | Grades 3–4 astrocytoma | Mixed genetic background of C57/BL6, Sv129, and B6/CBA | 35 | |
| Transgenic, inducible CKO | Nf1fl/+;p53fl/fl;Ptenfl/+and Nf1fl/fl;p53fl/fl | Nestin-cre-ERT2 | 46 weeks | High-grade astrocytoma | 129Svj/C57Bl6/B6CBA background | 34 | |
| CKO | TP53fl/fl, PTEN fl/+ | hGFAP-cre | 30 weeks | Anaplastic astrocytoma, GBM | FvB/C57Bl6 | 33 | |
| CKO | TP53fl/fl, PTEN fl/fl | Olig1-Cre | 90 days | GBM | C57BL/6J;129Sv | 121 | |
| CKO | TP53fl/fl, PTEN fl/fl | Olig2-tva-Cre | 115 days | GBM | |||
| MADR | NF1ko, p53ko, ptenko | In situ electroporation | N/A | Oligodendrogliomas | C57BL/6J | 36 | |
| Transgenic | NF1+/–, p53+/– | Globle | 7 months | Anaplastic astrocytomas, grade IV astrocytomas | Mixed C57BL/6, 129S4/SvJae, backcrossed to wild-type C57BL/6 (B6) | 31 | |
| SB | NRAS-GV12/shTP53/shATRX | In situ injection | 86 days | GBM | C57BL/6J | 37 | |
| NRAS-GV12/shTP53/Pdgfb | 74 days | GBM | C57BL/6J | ||||
| IDH1 IDH1 |
CKI | IDH1 R132H | Nestin-CreERT2 | 6–7 months | GBM precursors | C57BL/6 | 51 |
| SB | IDH1R132H, TP53sh, ATRXsh, NRAS G12V | In situ injection | 163 days | N/A | C57BL/6 | 54 | |
| IDH1WT, TP53sh, ATRXsh, NRAS G12V | 70 days | N/A | |||||
| CKI | IDH1 R132H (PM/Flex) | GFAP-creERT2 | N/A | N/A | B6N | 53 | |
| SB | NRAS-GV12/shTP53/shATRX/IDH1-R-132H | In situ injection | 213 days | N/A | C57BL/6J | 37 | |
| EGFR | RCAS/tv-a | EGFR, INK4a–ARF deletion | GFAP or nestin promoter | N/A | High-grade glioma | FVB/N, 129, and C57BL6 | 19 |
| Transgenic | V12Ha-ras, hEGFRvIII | hGFAP promoter | 2-4 weeks | Oligodendrogliomas | ICR | 21 | |
| Transgenic | v-erbB(transforming allele of EGFR) | S100β promoter | N/A | Low-grade oligodendroglioma | C57B6/JX DBA/2, back-crossed into FVB/N | 24 | |
| v-erbB, p53-/+ or ink4a/arf–/+, orink4a/arf–/– | GBM, high-grade anaplastic oligodendroglioma | p53–/– is FVB/N, ink4a/arf–/– is outbred | |||||
| Transgenic, adenoviruses delivered EGFRvIII | hGFAP:V12Ha-Ras, CMV:EGFRvIII | hGAP promoter | N/A | High-grade glioma | hGFAP:V12Ha-Ras is ICR | 23 | |
| Adenoviruses delivered Cre | Col1α1-EGFRvIII,EGFRwt, Ink4A/Arf–/–, pten–/– | IC injections | 6–8 weeks | GBM | N/A | 122 | |
| PiggyBac | CAG-STOPfl/fl-EGFRvIII, ptenfl/+ | Nestin-cre | 13 weeks | Spinal and brain glioma | Mixed FVB;C57BL/6J; C57BL/6J albino background | 22 | |
| PTEN | Transgenic, adenoviruses delivered cre | hGFAP:V12Ha-Ras; Ptenfl/fl | IC injection, hGFAP promoter | 2.5 weeks | GBM | hGFAP:V12Ha-Ras is ICR | 23 |
| RCAS virus | Kras, Ntv-a PTENf/f | RCAS-Cre, RCAS-Kras | N/A | GBM | Ptenf/f mice were a mixture of 129/Sv, CJ7, and C57BL6/J; tv-a transgenic mice were a mixture of FVB/N, C57BL6, BALB/C, and 129 | 38 | |
| CDKN2A | RCAS virus | INK4a-ARF–/–, KRAS + AKT | NESTIN or GFAP promoter | N/A | GBM | mixture of FVB/N, C57BL6, BALB/C, and 129 | 42 |
| H-ras | Transgenic | GFAP-HRasV12;p53KI/KI | GFAP-promoter | 10 weeks | astrocytoma | mixed 129SvJ/C57Bl6/CD1 | 56 |
| Transgenic | V12Ha-ras | hGFAP promoter | 4 weeks | Astrocytoma | ICR | 55 | |
| Lentivirus IC injection | H-RasV12, AKT,p53-/+ | GFAP-cre | 100–120 days | GBM | GFAP-cre and TP53–/– were maintained by backgrossing to C57BL/6 | 57 |
Abbreviations: CKI: conditional knock in; CKO: conditional knock out; MADR: mosaic analysis with dual recombinase; RCAS: replication-competent avian sarcoma-leukosis virus (ASLV) long-terminal repeat (LTR) with splice acceptor; SB: sleeping beauty.
Table 2.
Responses to Immunotherapy in Syngeneic Brain Tumor Mouse Models
| Tumor Model | Genetic Background | PD1 | CTLA4 | Others |
|---|---|---|---|---|
| Gl261 | C57BL/6J, carcinogen induced59 | Anti-PD1 (yes)90,123–125 | Anti-CTLA-4 (no)91,123 | anti-CD40 (no)73 |
| Anti-PD1 (yes, minor)86 | Anti-CTLA-4 (yes, minor)126 | Anti-CD40+ COX-2 inhibitor73(synergistic, minor) | ||
| Anti-PD1 (no)127–129 | anti-CTLA-4 (yes)130 | anti-OX40 (yes)125 | ||
| Anti-PD1 + anti-CTLA-4 (yes)84,126,131 | Anti-CTLA-4 + anti-PDL1(synergistic)130 | GVAX (yes, minor)125 | ||
| Anti-PD1+ anti TIGIT (yes, big)86 | Intratumoral IL-12 + anti-CTLA-4 (YES, big)91 | IDO inhibitor (no)130 | ||
| Anti-PD1+ anti TIGIT (yes, mild)127 | Anti-PD1 + anti-CTLA4 + CD40131 stimulation (no synergistic) | STING antonistic93 | ||
| Anti-PD1 + TMZ (synergistic)124 | Anti-PD1 + anti-CTLA4 + anti-IL6 (synergistic)131 | Intratumoral IL-12 (yes, minor)91 | ||
| Anti-PD1 + focal RT (synergistic)88 | Anti-PD1 + anti-CTLA4 + anti-IL6 + CD40 stimulation (overall survival)131 | mImp3 peptide vaccine + PolyIC (yes)132 | ||
| Anti-PD1 + GMCI (synergistic)101 | NKG2D CAR-T (yes, mild)133 | |||
| Anti-PD1 + Zika (synergistic)92 | Anti-IL6 + CD40 stimulation (synergistic)131 | |||
| Anti-PD1 + GVAX (synergistic)125 | Anti-IL6 (minor)131 | |||
| Anti-PD1 + anti-OX40 + GVAX (synergistic, dramatic)125 | CD40 stimulation (minor)131 | |||
| Anti-PD1 + anti-TIM3 + Focal RT87 (overall survival) | Anti-TIM3 (no)87 | |||
| Anti-PD1 + anti-CTLA4 + CD40 stimulation (no synergistic)131 | Anti-TIM3 + Focal RT (synergistic)87 | |||
| anti-PD1 + anti-CTLA4 + anti-IL6 (synergistic)131 | Anti-PD1 + anti-TIM3 + focal RT (overall survival)87 | |||
| Anti-PD1 + anti-CTLA4 + anti-IL6+ CD40 stimulation (overall survival)131 |
Anti-TIGIT (no)86,127 | |||
| Anti-PD1 + anti-TIM3 + Focal RT (overall survival)87 | Anti-PDL1 (yes)132 | |||
| Anti-PDL1-LNP/CDK5 inhibitor + RT (synergistic)83 | ||||
| Anti-CTLA-4 + anti-PDL1 (synergistic)130 |
||||
| CT2A | C57BL/6J. carcinogen induced58 | Anti-PD1 (yes, minor)100–102 | Anti-PD1 + anti-CTLA-4 + oncolytic virus delivery of IL12 (synergistic)97 | STING antonistic94 |
| Anti-PD1(NO)129 | Anti-PD1 + anti-CTLA-4 (no)97 | Anti-PDL1 (yes, minor)132 | ||
| Anti-PD1 + GMCI (synergistic)101 | Anti-CTLA-4 (no)102 | Neoantigen vaccine + PolyIC + anti-PDL1 (yes, synergistic)132 | ||
| Anti-PD1 + Zika (synergistic)92 | Anti-PDL1 + D2C7 (synergistic)102 | |||
| Anti-PD1 + anti-CTLA-4+ Oncolytic virus delivery of IL12 (synergistic)97 |
||||
| CT2A | C57BL/6J, carcinogen induced | Anti-PD1 + anti-CTLA-4(NO)97 | Anti-PDL1-LNP/CDK5 inhibitor + RT (synergistic)83 | |
| Anti-PD1 + D2C7(synergistic, moderate)102 | ||||
| 005GSC | FVB/N,C57BL/6, backcross to C57BL/657 | Anti-PD1 (no)98 | Anti-CTLA-4 (yes, minor)97 | Oncolytic virus delivery of IL12 (yes, minor)97 |
| Anti-PD1 (yes, minor)97 | Anti-PD1 + anti-CTLA-4 (yes)97 | CCR2 antagonist (NO)98 | ||
| Anti-PD1 + oncolytic virus delivery of IL12 (synergistic)97 | Anti-PD1 + anti-CTLA-4 + Oncolytic virus delivery of IL12 (overall long-term survival)97 | Anti-PDL1 (yes, minor)97 | ||
| Anti-PD1 + CCR2 antagonist (yes, modest)98 | Anti-CTLA-4 + oncolytic virus delivery of IL12 (synergistic)97 | Anti-PDL1 + oncolytic virus delivery of IL12 (synergistic)97 | ||
| Anti-PD1 + anti-CTLA-4(YES)97 | ||||
| SMA-560 | VM/Dk, spontaneous model | Anti-PD1 (yes, minor)102 | Anti-CTLA-4 (no)91,102 | Intratumoral IL-12 (yes, minor)91 |
| Anti-PD1 + D2C7 (immunotoxin) (synergistic, moderate)102 |
Anti-CTLA-4 (yes)104 | NKG2D CAR-T (yes)133 | ||
| Anti-CTLA-4 + D2C7 (synergistic, moderate)102 | Anti-PDL1 + D2C7(synergistic)102 | |||
| Intratumoral IL-12 + anti- CTLA-4 (YES, big)91 | ||||
| SB28 | C57BL/6, SB GEMM(NRasV12, TP53 shRNA, and mPDGF)73 | Anti-PD1 + irradiation (no synergistic)95 | Anti-PD1 + anti-CTLA-4 (NO)84 | FLT3L (yes, minor)95 |
| Anti-PD1 + anti-CTLA-4 (NO)84 | anti-CD40 (no)73 | |||
| Anti-CD40 + COX-2 inhibitor (synergistic, minor)73 | ||||
| Anti-PDL1 (yes, minor)95 | ||||
| KR158B | C57BL/6, 129S4/SvJae, backcross to C57BL/6, NF1 and TP53 KO transgenic mice31 | Anti-PD1 (no)90,98 | CCR2 antagonist (yes, modest)98 | |
| Anti-PD1 + CCR2 antagonist (synergistic)98 | ||||
| GEM tumor cell orthotopic injection | Pten–/–, Ink4a-Arf–/–, PDGFb | Anti-PD1 + anti-CTLA4 + CD40 stimulation + anti-IL6 (synergistic)131 | Anti-PD1 + anti-CTLA4 + CD40 stimulation + anti-IL6 (synergistic)131 | Anti-IL6 (minor)131 |
| Anti-PD1 + anti-CTLA4 + anti-IL6 (same as anti-IL6)131 | Anti-PD1 + anti-CTLA4 + anti-IL6(same as anti-IL6)131 | CD40 stimulation (no)131 | ||
| Anti-PD1 + anti-CTLA4 (no)131 | Anti-PD1 + anti-CTLA4 (no)131 | |||
| QPP4 | FvBXC57BL/6, QPP GEMM (Qki,TP53,PTEN triple ko)109 | Anti-PD1 (no)110 | Anti-CTLA4 (on)110 | |
| QPP5 | Anti-PD1 (NO)110 | Anti-CTLA4 (YES)110 | ||
| QPP7 | Anti-PD1 (yes)110 | Anti-CTLA4 (yes)110 | ||
| QPP8 | Anti-PD1 (no)110 | Anti-CTLA4 (no)110 |
Genetically Engineered Mouse Models
SV40 T-Antigen
The first brain tumor Genetically Engineered Mouse Model (GEMM) was established using egg microinjection of SV40-T-antigen DNA, a dominant-acting oncoprotein derived from the polyomavirus SV40.12 SV40 T-antigen has pleiotropic functions including regulation of viral genomic replication and host cell cycle. Importantly, SV40 T-antigen can induce malignant transformation by disrupting pRb and p53 tumor suppressors.13,14 The model’s clinical relevance is related to the minority of human brain tumor specimens that harbor DNA sequences and protein of SV40 T-antigen,13 suggesting that polyomavirus SV40 infection could stimulate brain tumor formation. First-generation SV40-T tumor models lacked tissue specificity, inducing cancer in various cell types. However, the second-generation models improved tissue specificity by combining CNS-specific regulatory elements with SV40 T-antigen, leading to CNS-specific malignancies.12,15 Pathologically, SV40-T infection generated diffuse cell proliferation in the periventricular subependymal zone with diffuse parenchymal invasion. Transformed cells exhibited secondary structuring similar to the histopathological features of human astrocytomas.15
Epidermal Growth Factor Receptor
Epidermal growth factor receptor (EGFR) is an oncogene commonly expressed in adult GBM and is associated with poorer prognosis. EGFR functions as a receptor tyrosine kinase (RTK) that binds epidermal growth factor as its ligand and downstream signaling activates cell proliferation, tumor invasion and inhibition of apoptosis.16 Genetic alterations of EGFR, including gene amplification, exon deletion, and point mutation, have been found in 30–50% of human GBM, promoting aberrant EGFR activation. Specifically, EGFRvIII, consisting of an intragenic deletion in exons 2 to 7, affects its extracellular domain, causing constitutive EGFR activation in a ligand-independent manner. Clinically, EGFRvIII is commonly expressed in GBM17 and emerges as a late molecular event after EGFR amplification.18
Given EGFR’s oncogenic role in brain tumors, models expressing mutant EGFR are desirable to recapitulate human gliomagenesis. A GEMM based on RCAS (replication-competent ASLV long terminal repeat with a splice acceptor) vector system for EGFR-driven glioma has been established.19 RCAS viruses only infect tumor virus A (TVA) receptor-expressing cells which confers tissue-specificity. Murine transgenic expression of TVA in specific cell types renders these cells susceptible to infection by RCAS viruses with genetic cargos.20 Paradoxically, multiple studies employing constitutively active GFAP-EGFRvIII resulted in failed or inefficient gliomagenesis, with significantly longer tumor latency of up to 60 weeks.21,22 When EGFR-driven models integrated coexpression of other predisposing mutations, this improved gliomagenesis efficiency. Mutant-activated EGFR expression with tumor suppressor ablation (p16Ink4a, p19Arf, and PTEN) or V(12)Ha-ras potentiated higher-grade glioma formation, but not for wildtype EGFR.21,23 These molecular alterations are nonspecific to cranial malignancies but also cooperate with spinal cord malignancies. For instance, PTEN loss is a codriver of EGFRvIII in spinal gliomagenesis rather than brain gliomagenesis.22 Related to EGFR, V-erbB is the oncogenic protein in the avian erythroblastosis virus. It acts as an auto-activating oncogene following significant deletion in its extracellular domain, and it shares similar molecular characteristics with EGFR. In tumor models and CNS stem cells, V-erbB expression, driven by the S100β promoter, results in low-grade oligodendroglioma development. Coexpression of V-erbB with INK4a/ARF or TP53 mutations accelerates gliomagenesis.24 Unfortunately, inconsistent tumor formation hinders model development, despite expression of tumor-associated mutations. For example, coexpression of mutant TP53 with constitutively active mutant EGFR failed to induce glioma unless CDK4 is expressed.19
PDGFR
PDGFR including its 2 isoforms, PDGFRα and PDGFRβ, is a cell surface receptor tyrosine kinase (RTK). Binding of its ligand, PDGF, to PDGFR results in receptor dimerization, transphosphorylation, and activation of intracellular signaling pathways, PI3K/AKT and RAS/MAPK.25
PDGFRα is the second most commonly amplified RTK gene in GBM, with 48% of PDGFRα-positive GBMs showing loss-of-function mutations in TP53.17,26 A GEMM driven by PDGFRα activation and TP53 knockout was generated using a lentiviral vector with a doxycycline-inducible PDGFRα and a constitutively active Cre. This vector was intracranially injected into transgenic mice expressing conditionally deleted alleles for PDGFRαfl/fl and TP53fl/fl, efficiently inducing gliomagenesis. However, activating PDGFRα with wildtype TP53 lacked tumor formation.27 In another model leveraging conditionally deleted Pten and TP53 alleles, retroviral-mediated intracranial PDGF delivery generated proneural gliomas with a median survival of 27 days, underscoring PDGFR’s cooperative role with other mutations. A lentiviral model introducing PDGFRβ and Cdkn2a short-hairpin RNA created GBM in mice, with a median survival of 77 days.28
TP53
TP53 is a transcription factor orchestrating multiple cellular functions as a tumor suppressor, including apoptosis, maintenance of genomic stability, inhibition of angiogenesis, and regulation of cell metabolism and the TME. Clinically, alterations of TP53 are commonly seen in 25–30% of primary GBM and 60–70% of grade IV astrocytomas. TP53 mutations in GBM most frequently occur in the DNA-binding domain, namely within 6 hotspot mutation sites (codons 175, 245, 248, 249, 273, and 282).29 These missense mutations result in the loss of DNA binding specificity and the gain of oncogenic functions. The p53-ARF-MDM2 pathway is dysregulated in 84% of GBM patients.30 In mice, germline loss of TP53 alone favors lymphoma and sarcoma development instead of gliomas. Similarly, loss of NF1 alone is insufficient to induce gliomagenesis, but heterozygous loss of wildtype alleles in TP53 and NF1 led to gliomagenesis in mice, generating the cell line, KR158.31 The sequence of TP53 loss relative to NF1 loss is significant for gliomagenesis in mice; losing TP53 before or concurrent to NF1 is necessary to form malignant astrocytomas.32 Loss of other tumor suppressors, like PTEN, with TP53 loss, significantly hastens gliomagenesis, resulting in higher-grade tumors and reduced survival.33
A neural stem/progenitor-based astrocytoma mouse model was generated by crossbreeding transgenic nestin-creERT2 mice with strains expressing inducibly floxed tumor suppressor alleles (Nf1fl/+; p53fl/fl; Ptenfl/+ or Nf1fl/fl; and p53fl/fl), resulting in high-grade astrocytomas.34 Additional heterozygous loss of PTEN with TP53 and NF1 mutations accelerates grade III astrocytomas formation, whereas loss of PTEN heterozygosity and Akt activation coincided with grade IV progression.35 It is well-established that neural stem cells (NSCs) contribute to the cancer stem cell (CSC) niche and tumor initiation in malignant brain tumors. Stemness was not sustained outside of the SVZ until QKi was additionally depleted with PTEN and TP53 because QKi acts as a significant regulator of NSC stemness.
Besides transgenic mice, other models utilized somatic tumorigenesis via NF1, TP53, and PTEN knock-out. Dual recombinase-mediated cassette exchange (MADR) is an in situ transgenesis method that permits stable labeling of mutant cells expressing single copy insertion of transgenic elements in precisely defined chromosomal loci. Recipient mice expressing loci with loxP and flp elements for insertion were electroporated in the early postnatal lateral ventricle with 3 plasmids: (1) smBFP2-P2A-SpCas9 for labeling and gene editing, (2) guide RNA targeting NF1, TP53, PTEN, and (3) Flp-Cre expression cassette.36 MADR-mediated tumor suppressor loss increased GBM’s penetrance to 92%.36 MADR confers several advantages by including single locus somatic modification, lineage tracing, dosage response of transgenes, personalized modeling, and adaptability to other strains.36 Tumors generated from the sleeping beauty (SB) transposon system, with NRAS-GV12/shTP53/shATRX (NPA) or NRAS-GV12/shTP53/PDGFb (NPD), have been reported to contain oncostream structures. These contain heterogeneous cellular populations including tumor cells, ACTA2+ mesenchymal cells, IBA1+ and CD68+ tumor-associated microglia/macrophages (TAMs), Nestin+ cells, and GFAP+ glial-derived cells, and positively correlate with tumor aggressiveness in human.37
PTEN
PTEN, a tyrosine phosphatase and tumor suppressor gene, governs numerous biological processes in cancer, including the maintenance of genomic stability, cell survival, migration, proliferation, and metabolism, and is frequently altered in brain cancer. Homozygous PTEN deletion under the control of the GFAP promoter is lethal in 6-week-old mice but not for heterozygous PTEN deletion. In the CNS, murine brain tissue expressing homozygously deleted PTEN showed significantly increased astrocytic proliferation and brain size but did not stimulate gliomagenesis. Like germline PTEN loss, somatic PTEN deletion through intracranial administration of CMV-cre adenoviral constructs into PTENf/f or PTENfl/+ mice did not induce gliomagenesis. Similar to other models, loss of PTEN with coexpressed V12Ha-Ras induced gliomas of varying grade, with heterozygously deleted PTEN generating lower grade astrocytomas, but homozygously deleted PTEN producing higher grade astrocytomas.23 In an RCAS-based system, conditional PTEN knockout with Kras expression in neural progenitors led to GBM formation. However, astrocyte-specific PTEN knockout did not result in tumor formation.38 As described previously, CNS-specific PTEN heterozygosity in an NF1/TP53-deficient astrocytoma model accelerated morbidity, and shortened survival with full penetrance of high-grade astrocytomas. Haploinsufficiency of PTEN accelerated the formation of grade III astrocytomas, while loss of PTEN heterozygosity progressed into grade IV astrocytomas.35 The dependency of PTEN loss on the coexpression of other genetic alterations must be considered when generating brain tumor models studying PTEN mutations.
CDKN2A/B-CDK4/6-RB
CDKN2A locus encodes 2 tumor suppressor proteins, p16INK4a and ARF.39 ARF regulates TP53 stability through interactions with MDM2 and ARF-BP1/Mule ubiquitin ligases.40 p16INK4a inhibits CDK4 and CDK6, the D-type cyclin-dependent kinases that initiate the phosphorylation of the tumor suppressor protein, RB. When hypophosphorylated, Rb inhibits genes related to S phase progression, thus promoting G1-phase arrest.41
About 77% GBM patient specimens harbor RB pathway aberrations, most commonly in deletion of the CDKN2A/CDKN2B locus on chromosome 9p21 (55% and 53%), followed by amplification of the CDK4 locus (14%).26 Leveraging the RCAS/tv-a system in mice, p16INK4a -ARF loss cooperated with Akt and Kras to increase tumor formation in glial progenitor cells. Akt and Kras alone are insufficient to induce gliomagenesis in astrocytes, but p16INKr4a-ARF loss promotes gliomagenesis suggesting that it sensitizes astrocytes to transformation.42 Transgenic mice expressing v-erbB with p16INK4a-ARF knockout increased the oligodendroglioma grade and penetrance.24
IDH1
The IDH1 (isocitrate dehydrogenase 1) mutation is characteristically found in >80% of grade IV astrocytomas and low-grade gliomas and is associated with increased overall survival. IDH1 mutational status is included in the pathological classification of glioma specimens given its prognostic value. IDH1 mutations frequently arise before the acquisition of TP53 mutations, loss of 1p/19q, and copy number alterations in PTEN and EGFR.43,44 IDH1 encodes isocitrate dehydrogenase 1, which catalyzes the oxidative carboxylation of isocitrate to α-ketoglutarate, resulting in the production of nicotinamide adenine dinucleotide phosphate (NADPH). This point mutation R132H is evolutionarily conserved and is localized to the substrate binding site, where it forms hydrophilic interactions with the alpha-carboxylate of isocitrate.45–47 Despite the loss of its canonical function, it obtained a new ability to catalyze the NADPH-dependent reduction of alpha-ketoglutarate to R(–)-2-hydroxyglutarate (2HG).48 However, the mechanism of how IDH1 R132H contributes to carcinogenesis remains unclear. It is thought that R-2-HG antagonizes α-KG by competitively inhibiting various α-KG-dependent dioxygenases, including the JmjC domain-containing histone demethylases (KDMs) and the TET (10–11 translocation) family of DNA hydroxylases that catalyze the sequential oxidation of 5-methlycytosine (5mC) to 5-hydroxymethylcytosine (5hmC), 5-formyl cytosine (5fC), and 5-carboxyl cytosine (5caC), promoting DNA demethylation.49 Another report showed 2HG binds to CDC42 and abolishes its association with MLK3, disrupting JNK-mediated apoptosis.50 Multiple models have been generated to elucidate the mechanisms related to mutant IDH1.
A conditional IDH1R132H GEMM was generated by crossing IDH1fl(R132H)/+ knock-in mice with tamoxifen-inducible nestin-CreERT2-expressing mice, which expresses Cre in the SVZ and the subgranular zone (SGZ) of the hippocampal dentate gyrus. Tamoxifen induction in 5-week-old mice produced phenotypes resembling gliomagenesis.51 Interestingly, offspring born from the parental crossing of IDH1fl(R132H)/+ mice with noninducing nestin-Cre mice died perinatally and exhibited brain hemorrhages.52 A similar model driving the conditional IDH1fl(R132H)/+ knock-in utilized GFAP-Cre–ERT2 mice to study this mutation.53 Analogous to other models, IDH1 mutations co-occur with other molecular lesions, including 1p/19q codeletion, inactivating TP53 mutations, and loss-of-function mutations in alpha thalassemia/mental retardation syndrome X-linked gene (ATRX) in glioma patients. To better recapitulate patient-relevant mutations, a nonviral gene delivery system, SB transposon, established a GEMM expressing IDH1R132H or wildtype IDH1 with TP53 and ATRX knock-down. IDH1R132H increases median overall survival without treatment and paradoxically elicits tumor radio-resistance.54 Clinically, this finding parallels the improved prognosis observed in patients with mutant IDH1-expressing brain tumors. IDH1 mutation status affects TME heterogeneity, with IDH1wt exhibiting greater heterogeneity.37 These models offer promising potential for understanding the favorable prognostic outcomes associated with IDH1 mutations, and mechanisms of coexpressed mutations like TP53, ATRX, etc.
Hras-V12
Though V12Ha-Ras is rarer in human GBMs, mouse models of transgenic GFAP-Harvey Ras (HRas)V12 exhibit MAPK pathway activation at comparable molecular levels to human GBMs,55,56 with higher doses of V12Ha-Ras promoting tumorigenesis.55
To generate a V12Ha-Ras-based model, mice with a GFAP-Cre background received a lentiviral construct harboring Cre-mediated expression of H-RasV12 or Akt cassettes to the hippocampus, SVZ, and cortex. In the hippocampus and SVZ, tumor formation failed or was significantly inefficient with individual or concurrent vector administration, respectively. Upon induction of TP53 mutations, tumor formation was significantly improved with shorter tumor latency. This model resulted in the formation of a cell line, 005GSC, generated from these tumors.57 Given this model’s dependency on TP53 alteration, a switchable TP53 functional model was developed that replaces endogenous Tp53 with 1 encoding the TP53ERTAM [estrogen receptor (ER)] fusion protein. p53ERTAM is functional only in the presence of the synthetic steroid ligand 4-hydroxytamoxifen (4-OHT).56 This model provides novel insight into the temporal sequence of tumor suppressor inactivation and oncogenic activation in glioma. Additionally, the model’s switchable functionality yields insight into how TP53 disrupts gliomagenesis following gene rescue.
Syngeneic Implantation Model
Syngeneic implantation models have been ubiquitously used in tumor research due to their shorter latency and consistently high tumor incidence.
Carcinogen-Induced and Spontaneous GBM Cell Lines
Carcinogen-induced cell lines most commonly used in neuro-oncology include GL261 and CT-2A. Both were generated by intracranially injecting methylcholanthrene into C57BL/6J mice and were maintained by serial transplantations of tumor pieces into the syngeneic mouse strain.58,59 The GL261 transplantation tumor model is the most extensively used GBM mouse model. However, it has been observed that GL261 exhibits a bulky growth pattern, forming a nodular mass without considerable parenchymal infiltration. These tumors rarely display necrosis and invasive growth, which are notable characteristics of human GBM. Further, it is molecularly driven by an activating Kras mutation, which is rarely observed in human GBM.17,60 Monolayer-cultured GL261 exhibits a range of intratumoral pathology, oscillating between mesenchymal-like (MES-like) and astrocyte-like (AC-like). In contrast, GL261-GSCs cultured in neurospheres (NS) and implanted GL261-GSCs shift towards oligodendrocyte-progenitor-like (OPC-like) and neural-progenitor-like (NPC-like) states, which more closely mimic patient samples.61 Despite the model’s divergent features from the clinical data, these models possess high utility as NSs.62 CT-2A NS were generated by serum-free media culturing.63 Unlike monolayer cultures of the parental line, CT-2A neurospheres preserve cancer stemness characteristics,63 providing insights into the glioma stem cell (GSC) niche. CT-2A recapitulates several features of human GBM, including high mitotic index and cell density, nuclear polymorphism, hemorrhage, pseudopalisading necrosis, and microvascular proliferation.64 This cell line demonstrates high intratumoral heterogeneity, displaying a differential expression of Sox9, Sox10, and GFAP across different cells within the tumor mass.64 However, conflicting results found that cells derived from sphere cultures were less tumorigenic, had similar temozolomide resistance and showed no significant differences in vascularization or immune cell infiltration compared to nonsphere cultures. Despite displaying elevated stemness markers and reduced sensitivity to radiation (RT), these findings challenge the notion that neurospheres are superior to 2D cultures.65 Since carcinogens stochastically induce mutations, excessive oncogenic alterations increase the model’s TMB, which is hypothesized to increase tumor immunogenicity and differ from patient samples which are not highly immunogenic. Aside from carcinogen-induced models, SMA-560 is one of the few models that spontaneously arose in VM/Dk mice.66 It was established as a cell line through serial transplantation of spontaneous tumor tissue in syngeneic mice.67 SMA-560 is highly tumorigenic and possesses astrocytic features but with low levels of S-100 and GFAP. However, the oncogenic driver mutation in this line is not well characterized. Two additional lines, SMA-497 and SMA-540, were also generated from VM/Dk mice, though not widely disseminated. Among these 3 lines, SMA-560 exhibits the most differentiated astrocytic features, whereas SMA-497 is the least differentiated and the least chemosensitive.68
Targeted Induced Genetic Alteration
The KR158B cell line is a commonly used mouse GBM cell line derived from a GEMM model expressing germline global NF1–/+ and TP53s/+ modifications. Tumor development in vivo results in the loss of NF1 and TP53 wildtype alleles, leading to homozygous NF1 and TP53 deletion.31 KR158B tumor tissue demonstrated tumoral and nontumoral cellular heterogeneity. Tumor cells exhibited both fast-cycling and slow-cycling characteristics, with slow-cycling cells (SCCs) displaying features reminiscent of CSCs, similar to human SCCs. Furthermore, tumors arising from gliomasphere cultures of KR158B exhibited more pronounced GBM characteristics, including extensive infiltration and well-defined pseudopalisading necrosis, compared to those from monolayer cultures.69 Similar to KR158, mut3 is derived from a GEMM that expresses heterozygous TP53–/+ and conditional cre-mediated NF1 knockout in the CNS (NF1f/+) with mut3 tumors losing both alleles.34 While KR158B and mut3 models lose TP53 and NF1, another model, mGB2, is derived from PTEN and TP53 double knock-out GEMM (Tlx-CreERT2/p53-floxed/pten-floxed). The original line, mGB0, is generated by tamoxifen-induced knockout of PTEN and TP53. Fully invasive mGB0 tumors are isolated after 10–24 months and 2 serial transplantations yielded the mGB2 line.70 mGB2 recapitulates human GBM characteristics including necrosis, invasion, intratumoral hemorrhages, and platelet aggregation.70,71 Unlike models that are reliant on an inducible approach, another GEMM model, SB28, is generated via the SB transposon integration system.72,73 Transfection reagent and plasmids containing SB transposon-flanked NRasV12, TP53 shRNA, and mPDGF were injected into lateral ventricles of mouse neonatal pups. With a note, NrasV12 mutation in human glioma is rarely found and tumor histology features are not well characterized.74 Other cell lines and culture conditions have innovated model approaches to better reflect human glioma biology. QPP cell lines are generated from a GEMM harboring triple knockout of Qki, TP53, and PTEN and are cultured specifically in NeuroCult serum-free medium. The QPP mouse model and associated syngeneic cell line mirror human GBM pathology and TME characteristics.75 To account for GSC biology, a cell line, 005GSC, was generated from the GFAP-cre TP53–/+ mouse that received hippocampal lentiviral injections containing H-rasV12 and AKT, resulting in CNS-specific expression. NSC culture conditions were utilized to generate this model since 005GSC tumor cells only survive 2 weeks in a monolayer. The advantages of NSC culture conditions include preservation of CSC features and expression of immature neural progenitor marker, nestin. 005GSC is histologically akin to human GBM, exhibiting tumor heterogeneity, invasiveness, vascularity, and an immunosuppressive TME.76 In comparison, bRiTs-G3s are generated in vitro first by retroviral transduction of constitutively active HRasV12 in neural stem/progenitor cells isolated from the SVZ in adult mice with homozygously deleted Ink4a/Arf loci. Transduced cells were cultured as neurospheres and implantation in mice resulted in highly invasive, hypervascular GBM-like tumors.77 Lastly, NSCL61 is another NSC-modified GBM cell line that overexpresses H-rasL61 in TP53-deficient NSC, which is highly enriched in GSC.78 The diverse methodology of model generation and culture conditions permits a detailed investigation of glioma heterogeneity. It is noteworthy that new in situ transgenesis methods increased the efficiency of GEMM generation. Besides in situ viral delivery of oncogenic factors, in situ electroporation is another option to rapidly create somatically transgenic mice. In utero electroporation (IUE) is a widely used method which when combined with genetic cargos, CRISPR, transposons, and MADR tools, offers multiple advantages including spatial and temporal specificity, rapid generation, flexible gene manipulation, wide applicability, and preservation of an immunocompetent context.36 IUE-mediated PiggyBac overexpression of RasV12 with HA-NFIA stably generated murine oligodendroglioma.79 Another model utilizing IUE involved electroporation of Cre-containing CRISPR-Cas9 vector into the frontal SVZ with highly efficient single-guide RNAs that target Trp53, Pten, and EGFR, with over 90% of electroporated mice developing brain tumors.80
Immune Therapy Response
GL261
Even though GL261 is a highly immunogenic cell line harboring 4932 nonsynomymous exome mutations, ICB in this syngeneic model shows equivocal efficacy dependent on dosing strategy, tumor inoculation route, or mice strains.81 Though considered a highly immune-infiltrated tumor, TAMs increased throughout the tumor development.61 Significant synergistic antitumor responses have been reported when combining PD1 blockade with other immune checkpoint inhibitors, including CTLA-4, OX40, and TIGIT. Concurrent ICB against PD1, OX40 and GVAX demonstrated increased overall survival. Unfortunately, combined ICB with inhibitors against IDO, CTLA-4, TIM3, or TIGIT showed inconsistent or no responses in this model.82–86 When combining ICB (PD1 or TIM3 blockade) with RT, these modalities produce synergistic therapeutic responses and survival benefits. Additional combinations like focal RT, anti-PD1 with anti-TIM3 or IDO inhibitors significantly increased survival.82,87,88 While RT may potentiate immunotherapy responses, it can also induce the expression of immune checkpoint molecules such as PD-L1 on TAMs. Given that CDK5 has a critical role in interferon-gamma (IFNγ)-stimulated PD-L1 production, inhibition of CDK5 using PD-L1-targeting lipid nanoparticles (LNPs) that encapsulate a CDK5 inhibitor could abrogate this RT response mechanism. Combining RT with PD-L1-targeting LNPs encapsulated with a CDK5 inhibitor synergistically enhanced survival in GL261 and CT2A83. Other conventional treatment modalities such as RT and chemotherapy synergize with cancer vaccines and ICB in GL261. Vaccines harboring either neoantigen peptide or whole irradiated tumor resulted in significant responses. Cancer vaccine efficacy was enhanced via intratumoral TMZ administration using convection-enhanced delivery in GL261, but not KR158B.89 In GL261, standard TMZ dosing induced T-cell exhaustion and attenuated ICB responses.90 This effect was not observed when using metronomic doses of TMZ and when combined with PD1 blockade, lower levels of T-cell exhaustion were observed relative to the standard dose.90 Standard TMZ dosing may have negatively impacted immunotherapy efficacy due to iatrogenic myelosuppression. However, intratumoral IL-12 administration significantly increased survival in GL261.91 Other immunotherapeutic combinations have been evaluated in GL261 such as Zika oncolytic virus and gene-mediated cytotoxic immunotherapy, which demonstrated synergistic antitumor efficacy.92 Potent immunostimulatory effects have also been observed with the stimulator of interferon genes (STING) DNA sensing pathway, and STING agonists have significantly increased survival in GL261 and CT2A, but this is dependent on NK cell populations.93,94 In sum, the ubiquitous use of GL261 in neuro-oncology has led to its application in evaluating multiple immunotherapy modalities, sustaining promising avenues of bench-to-bedside research.
SB28
SB28 is a poorly immunogenic, ICB-resistant line, with low MHC-I expression and modest CD8+ T-cell infiltration.84 Relative to GL261, SB28 has significantly fewer somatic mutations at approximately 108 in total and differs in its sensitivity to ICBs.84 When administering ICB, antitumor immune responses are not observed against SB28. PDL1 blockade alone has minor survival benefits, with no additional benefits derived from RT. Other models demonstrated that agonistic CD40 monoclonal antibody (mAb) activates myeloid cells and promotes antitumor immunity, but no response to anti-CD40 is observed in SB28. When CD40 blockade is combined with COX2 inhibition in SB28, a minor survival benefit is produced which has been linked to a reduction in immunosuppressive myeloid cells.73 Aside from the direct effects of immunotherapy, biological sex immensely impacts the immune response against cancer. This is self-evident in SB28 because blocking granulocytic myeloid-derived suppressor cells has shown sex-dependent survival benefit.85 Additionally, SB28’s immunogenicity is dependent on the tumor’s anatomic site. Unlike intracranial tumors, significant ICB responses occur against subcutaneously administered SB28. Mechanistically, subcutaneous SB28 has greater infiltration of DCs, T cells and NK cells. In an intracranial model, increased DC abundance mediated by FLT3 administration with RT or ICB failed to significantly increase survival in SB28. These findings epitomize the challenges of mounting effective antitumor immune responses in the brain, likely hindered by compensatory immune regulatory cells like T regulatory cells (T-regs).95
005GSC
Murine 005GSC is poorly immunogenic, lacking cell surface expression of MHC-I, CD40, CD80, and CD86. In contrast to SB28, 005GSC expresses increased levels of NK-cell receptor ligands, retinoic acid early inducible-1 (Rae-1) and CD155.76 ICB efficacy has been evaluated in 005GSC, but responses are checkpoint molecule dependent. Unlike PD1 blockade, CTLA-4 blockade displayed significant efficacy against orthotopically implanted 005GSC. αCTLA-4-specific responses were mechanistically reliant on CD4+ T-cell-secreted IFNγ that sensitized glioma cells to microglia phagocytosis.96 This model has also evaluated the impact of IL-12-expressing oncolytic viruses, resulting in macrophage polarization and intratumoral T-reg reduction.76,97 Modest survival benefits were achieved with combinatorial oncolytic virus and ICB. However, significant improvements were noted in triple combination with PD1 and CTLA-4 blockade, which was verified in CT2A.97 Lastly, targeted approaches against immunosuppressive cells were evaluated in 005GSC. Despite inhibiting immunosuppressive myeloid cells, CCR2 antagonism failed to significantly extend survival in 005GSC, but combinatorial approaches with ICB significantly improved survival.98 Aside from myeloid cells, other suppressive cells like T-regs limit immunotherapy responses. Significant therapeutic effects were observed in 005GSC by targeting T-regs, which were amplified with ICB.93 Unlike SB28, 005GSC as a model yielded mixed immunotherapy responses and is less immunogenic relative to GL261.
CT2A
CT2A is a less immunogenic line than GL261. Immunophenotyping of the TME revealed that CT2A has the lowest level of activated and resting microglia compared to GL261, 005GSC, and Mut3. However, CT2A has the highest level of TAMs and exhausted CD8 T cells.99 Given the high abundance of immunosuppressive cells, this model is highly relevant to immunotherapy evaluation since immunosuppression is a significant barrier to antitumor immune responses. Multiple studies have evaluated ICB using this model. For example, minor therapeutic responses were observed after the PD1 blockade, but no responses were observed after the CTLA-4 blockade.100–102 This observation is consistent with other models in that ICB responses are often checkpoint molecule-dependent. However, mechanisms underlying these differential responses across models are unknown but may underlie poor clinical responses. Other targeted therapies aimed at antagonizing oncogenic stimuli have been evaluated in CT2A. In CT2A and SMA-560 subcutaneous models, a novel mAb, D2C7, that targets wildtype EGFR and mutant EGFRvIII revealed a significant survival benefit dependent on CD4 and CD8 T-cells. The combination of D2C7 with ICB moderately improved survival, with significant survival benefit seen against PD1. However, combining D2C7 with mAbs against T-cell exhaustion did not augment the survival benefit.102 As previously discussed, oncolytic viruses have been evaluated with ICB in CT2A. PD1 blockade with oncolytic viral therapy showed significantly improved benefits compared to either monotherapy.92,97,101 These findings support the observations that combinatorial approaches with different immune modalities improve survival outcomes. Similarly, irradiation may enhance immunotherapy responses through its immune-stimulatory effects.82,83 Unlike models of 005GSC, the combination of anti-PD1 with anti-GITR, a T-reg targeting antibody, does not augment antitumor responses in CT2A. However, combining these antibodies with irradiation synergized to produce significant antitumor responses.93 Shifting to STING antagonism, significant therapeutic responses with STING antagonist monotherapy were noted in CT2A, similar to SB28 models.93
SMA-560
SMA-560 harbors 2171 nonsynomymous exome mutations, less than 50% of the total number observed in GL261. Additionally, this cell line expresses 2 neoantigen peptides: Odc1Q129L and E2f8K272R.103 Inconsistent ICB responses have been observed with modest responses after PD1 blockade and varied responses to CTLA-4 blockade.91,102,104 Additionally, ICB was combined with IL-12 administration. In SMA-560, intratumoral administration of IL-12 with CTLA-4 blockade dramatically increased survival and tumor eradication, but monotherapy failed to generate robust responses. This therapeutic effect is mediated by CD4+ cells, significantly decreasing FoxP3+ T-regs and increasing effector T cells.91
KR158B
Like other models, KR158B is a poorly immunogenic line and fails to generate therapeutic responses to ICB monotherapy.90,98,105 PD1 blockade administered with hematopoietic stem cell (HSC) transfer and lymphodepletive irradiation overcomes therapeutic resistance to ICB monotherapy.105 Aside from ICB, adoptive immunotherapies have been evaluated in KR158B, and monotherapy employing adoptively transferred tumor-reactive T-cells did not elicit survival benefits. However, treatment resistance to adoptively transferred T-cells was overcome by combining myeloablation with total body irradiation, HSC transfer, and DC vaccine.106,107 An important consideration is differential responses observed across models using the same treatment. Unlike in GL261, tumor vaccine combined with CED-TMZ treatment did not augment significant therapeutic effects against KR158B.89
Newly Developed Syngeneic model
With a note, freshly isolated tumor spheres from GEMMs were reportedly used in immunotherapy development. RCAS-based GBM neurospheres expressing PDGF-B, Cre, Ink4a-arf–/–, and PTENfl/fl were orthotopically implanted and ICB responses were achieved by targeting TAMs through IL-6 inhibition and CD40 stimulation. To study immune evasion mechanisms, GSCs were generated from novel NSCs engineered to express NF1–/–, PTEN–/– and EGFRvIII.108 Engineered NSCs were transplanted into immunocompetent mice and immune-escaped tumors were generated following serial transplantation.108 In this study, TAMs mediated tumor immune escape, with macrophage depletion via CSF1R blockade increasing overall survival. Interestingly, GSCs undergo transcriptional reprogramming consistent with myeloid “mimicry” hypothesized to help establish an immunosuppressive TME.108 Other models also efficiently replicate the human brain TME. As discussed earlier, QPP cell lines are derived from the QPP GEMM (Qki, TP53, and PTEN triple ko) mouse and maintained in neurosphere culture.109 The TME immunophenotypes of implanted QPP7 and QPP GEMM tumors reflected the immunosuppressive milieu of human gliomas.110 QPP4 and QPP8 resisted ICB, but QPP5 demonstrated responses to CTLA4 blockade. The only line to demonstrate sensitivity to PD1 and CTLA4 blockade is QPP7. Underlying this effect, QPP7 demonstrates higher immunogenicity relative to other lines likely due to the line’s higher neoantigen burden.110
Conclusion and Discussion
Mouse models serve a vital role in understanding tumorigenesis and identifying more efficacious therapeutic strategies. While human-derived tumor lines and immunocompromised mouse models have generated patient-derived xenografts (PDXs), they are hindered in their ability to adequately address critical questions in cancer immunology like the interplay among tumor, treatment and host immunity. As new immunotherapy strategies arise, there is increased demand for mouse models with available molecular, genetic, and immune status information along with sharing the methodology of model generation, the effects of introduced perturbations, and immunogenicity status. Despite the high model diversity of brain tumors, the field consistently relies on a minority of specific mouse models like GL261, which displays divergent immunogenicity and tumor neoantigen burden compared to patients.
Features of models exhibit strengths and weaknesses depending on the research objectives, but they generally facilitate scientific translation from bench to bedside. PDXs capture human intratumoral and intertumoral heterogeneity. However, these models typically require immunodeficient mice, and human–mouse tissue interactions can confound results. Humanized mouse models also recapitulate human tumor heterogeneity and include human immune components, ideally suitable for studying interactions between the immune system and tumors, though human-mouse tissue interactions may still confound results. GEMMs are advantageous for expressing specific mutations, with inducible functionalities. They are immune competent and avoid human–mouse interactions, but they are expensive to create, and inconsistent tumor formation limits their reproducibility. Although syngeneic implantation models surpass other models in time and cost-effectiveness, consistency, repeatability, and in vitro stability of glioma cell lines remain a challenge. Surgical interventions also disrupt the blood–brain barrier, with potential interactions with analgesic medications, tumor engraftment, and immunotherapy responses. Another critical gap remains in recapitulating brain tumor outcomes between older and younger adult patients. Models are frequently derived from adolescent-aged mice, in part, because aged mice are prohibitively expensive. Clinically, youthfulness bias in models harbors significance given that older age is associated with poorer brain tumor outcomes. Tumor-bearing brains are divergent in an age-dependent manner that impacts therapeutic efficacy.111 Future model efforts should address therapies in youthful and aged contexts to better translate preclinical immunotherapies to the appropriate patient population.
Mouse model selection should be based on the specific scientific question since no model contains every ideal aspect. Despite immune deficiencies in PDX models, it is still a favorable tool that recapitulates human tumor heterogeneity and genetics. Specifically, preclinical therapeutics like synNotch-CAR T-cells have been evaluated in PDX GBM models, demonstrating efficient tumor killing amid neoantigen heterogeneity.112 Supporting this premise that models promote translation, synNotch-CAR T-cells are being evaluated in a phase I clinical trial (NCT06186401) for patients diagnosed with EGFRvIII+ GBM.
In patients diagnosed with GBM, MGMT promoter methylation is associated with favorable prognoses following TMZ administration.113 Presently, human GBM cell lines and PDXs harboring MGMT promoter methylation have demonstrated increased TMZ sensitivity, but there is a paucity of GEMMs expressing MGMT methylation. Limited evidence suggests that MGMT expression may impact the immune landscape of primary GBM,114 but future efforts should be dedicated to generating models that express MGMT methylation.
Humanized mouse models exhibit features of human hematopoietic and immune cells. These models are advantageous in their unique human tumor-immune cell context and humanized GBM models reflect pathologic and immune-TME features observed in patients.115 PD1 blockade in these models reduced tumor size and immunosuppressive cells in the TME.116,117 These models have evaluated ICB with oncolytic viruses, in addition to STAT3 inhibition, which increased TILs within the TME.118
Our review serves as a valuable resource for cancer immunologists who seek to select appropriate mouse models for their specific research questions. We have provided a detailed summary of GEMMs and syngeneic mouse models for studying adult brain tumors. This review encompasses model characteristics, including the generation method, genetic background, oncogenic mechanisms, and immune-related features. Within the research community, there is a pressing need for the development of new mouse models that include clinically relevant genetic backgrounds and comprehensive profiling of immune-related characteristics. These advancements would significantly contribute to the progress and translation of cancer immunology studies.
Contributor Information
John Figg, University of Florida Brain Tumor Immunotherapy Program, Preston A. Wells, Jr. Center for Brain Tumor Therapy, Lillian S. Wells Department of Neurosurgery, McKnight Brain Institute, University of Florida, Gainesville, Florida, USA.
Dongjiang Chen, Division of Neuro-Oncology, Department of Neurological Surgery and Neurology, USC Keck Brain Tumor Center, University of Southern California Keck School of Medicine, Los Angeles, California, USA.
Laura Falceto Font, University of Florida Brain Tumor Immunotherapy Program, Preston A. Wells, Jr. Center for Brain Tumor Therapy, Lillian S. Wells Department of Neurosurgery, McKnight Brain Institute, University of Florida, Gainesville, Florida, USA.
Catherine Flores, University of Florida Brain Tumor Immunotherapy Program, Preston A. Wells, Jr. Center for Brain Tumor Therapy, Lillian S. Wells Department of Neurosurgery, McKnight Brain Institute, University of Florida, Gainesville, Florida, USA.
Dan Jin, University of Florida Brain Tumor Immunotherapy Program, Preston A. Wells, Jr. Center for Brain Tumor Therapy, Lillian S. Wells Department of Neurosurgery, McKnight Brain Institute, University of Florida, Gainesville, Florida, USA.
Funding
This work was supported by the National Institute of Neurological Disorders and Stroke (R01NS112315 and R01NS111033 to C.F.).
Conflict of interest statement
C.F. is a founder of iOncologi. Other authors declare no conflicts of interest.
Authorship statement
J.F., D.J., D.C., and L.F. contributed to drafting the manuscript. C.F. and D.J. contributed to revising the manuscript.
References
- 1. Akter F, Simon B, de Boer NL, et al. Pre-clinical tumor models of primary brain tumors: challenges and opportunities. Biochim Biophys Acta Rev Cancer. 2021;1875(1):188458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Barbosa MAG, Xavier CPR, Pereira RF, Petrikaite V, Vasconcelos MH.. 3D cell culture models as recapitulators of the tumor microenvironment for the screening of anti-cancer drugs. Cancers (Basel). 2021;14(1):190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Connolly KA, Fitzgerald B, Damo M, Joshi NS.. Novel mouse models for cancer immunology. Annu Rev Cancer Biol. 2022;6(1):269–291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Salmon H, Remark R, Gnjatic S, Merad M.. Host tissue determinants of tumour immunity. Nat Rev Cancer. 2019;19(4):215–227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Spranger S, Gajewski TF.. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat Rev Cancer. 2018;18(3):139–147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Spranger S, Bao R, Gajewski TF.. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature. 2015;523(7559):231–235. [DOI] [PubMed] [Google Scholar]
- 7. Ruiz de Galarreta M, Bresnahan E, Molina-Sanchez P, et al. Beta-catenin activation promotes immune escape and resistance to anti-pd-1 therapy in hepatocellular carcinoma. Cancer Discov. 2019;9(8):1124–1141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Villemin C, Six A, Neville BA, et al. The heightened importance of the microbiome in cancer immunotherapy. Trends Immunol. 2023;44(1):44–59. [DOI] [PubMed] [Google Scholar]
- 9. Dees KJ, Koo H, Humphreys JF, et al. Human gut microbial communities dictate efficacy of anti-PD-1 therapy in a humanized microbiome mouse model of glioma. Neurooncol Adv. 2021;3(1):vdab023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Erny D, Hrabe de Angelis AL, Jaitin D, et al. Host microbiota constantly control maturation and function of microglia in the CNS. Nat Neurosci. 2015;18(7):965–977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lin B, Ye Z, Ye Z, et al. Gut microbiota in brain tumors: An emerging crucial player. CNS Neurosci Ther. 2023;29(Suppl 1):84–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Brinster RL, Chen HY, Messing A, et al. Transgenic mice harboring SV40 T-antigen genes develop characteristic brain tumors. Cell. 1984;37(2):367–379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Rotondo JC, Mazzoni E, Bononi I, Tognon M, Martini F.. Association between simian virus 40 and human tumors. Front Oncol. 2019;9:670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Zhen H, Zhang X, Zhang Z, et al. Simian virus 40 large tumor antigen forms specific complexes with p53 and pRb in human brain tumors. Chin Med J (Engl). 2001;114(4):382–386. [PubMed] [Google Scholar]
- 15. Danks RA, Orian JM, Gonzales MF, et al. Transformation of astrocytes in transgenic mice expressing SV40 T antigen under the transcriptional control of the glial fibrillary acidic protein promoter. Cancer Res. 1995;55(19):4302–4310. [PubMed] [Google Scholar]
- 16. Xu H, Zong H, Ma C, et al. Epidermal growth factor receptor in glioblastoma. Oncol Lett. 2017;14(1):512–516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Brennan CW, Verhaak RG, McKenna A, et al. ; TCGA Research Network. The somatic genomic landscape of glioblastoma. Cell. 2013;155(2):462–477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Eskilsson E, Rosland GV, Talasila KM, et al. EGFRvIII mutations can emerge as late and heterogenous events in glioblastoma development and promote angiogenesis through Src activation. Neuro Oncol. 2016;18(12):1644–1655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Holland EC, Hively WP, DePinho RA, Varmus HE.. A constitutively active epidermal growth factor receptor cooperates with disruption of G1 cell-cycle arrest pathways to induce glioma-like lesions in mice. Genes Dev. 1998;12(23):3675–3685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Ahronian LG, Lewis BC.. Using the RCAS–TVA system to model human cancer in mice. Cold Spring Harb Protoc. 2014;2014(11):1128–1135. [DOI] [PubMed] [Google Scholar]
- 21. Ding H, Shannon P, Lau N, et al. Oligodendrogliomas result from the expression of an activated mutant epidermal growth factor receptor in a RAS transgenic mouse astrocytoma model. Cancer Res. 2003;63(5):1106–1113. [PubMed] [Google Scholar]
- 22. Noorani I, de la Rosa J, Choi YH, et al. PiggyBac mutagenesis and exome sequencing identify genetic driver landscapes and potential therapeutic targets of EGFR-mutant gliomas. Genome Biol. 2020;21(1):181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Wei Q, Clarke L, Scheidenhelm DK, et al. High-grade glioma formation results from postnatal pten loss or mutant epidermal growth factor receptor expression in a transgenic mouse glioma model. Cancer Res. 2006;66(15):7429–7437. [DOI] [PubMed] [Google Scholar]
- 24. Weiss WA, Burns MJ, Hackett C, et al. Genetic determinants of malignancy in a mouse model for oligodendroglioma. Cancer Res. 2003;63(7):1589–1595. [PubMed] [Google Scholar]
- 25. Chen PH, Chen X, He X.. Platelet-derived growth factors and their receptors: structural and functional perspectives. Biochim Biophys Acta. 2013;1834(10):2176–2186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Cancer Genome Atlas Research N. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature. 2008;455(7216):1061–1068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Jun HJ, Appleman VA, Wu HJ, et al. A PDGFRalpha-driven mouse model of glioblastoma reveals a stathmin1-mediated mechanism of sensitivity to vinblastine. Nat Commun. 2018;9(1):3116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Rahme GJ, Luikart BW, Cheng C, Israel MA.. A recombinant lentiviral PDGF-driven mouse model of proneural glioblastoma. Neuro Oncol. 2018;20(3):332–342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. England B, Huang T, Karsy M.. Current understanding of the role and targeting of tumor suppressor p53 in glioblastoma multiforme. Tumour Biol. 2013;34(4):2063–2074. [DOI] [PubMed] [Google Scholar]
- 30. Zhang Y, Dube C, GibertM, Jr, et al. The p53 pathway in glioblastoma. Cancers (Basel). 2018;10(9):297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Reilly KM, Loisel DA, Bronson RT, McLaughlin ME, Jacks T.. Nf1;Trp53 mutant mice develop glioblastoma with evidence of strain-specific effects. Nat Genet. 2000;26(1):109–113. [DOI] [PubMed] [Google Scholar]
- 32. Zhu Y, Guignard F, Zhao D, et al. Early inactivation of p53 tumor suppressor gene cooperating with NF1 loss induces malignant astrocytoma. Cancer Cell. 2005;8(2):119–130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Zheng H, Ying H, Yan H, et al. p53 and Pten control neural and glioma stem/progenitor cell renewal and differentiation. Nature. 2008;455(7216):1129–1133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Alcantara Llaguno S, Chen J, Kwon CH, et al. Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell. 2009;15(1):45–56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Kwon CH, Zhao D, Chen J, et al. Pten haploinsufficiency accelerates formation of high-grade astrocytomas. Cancer Res. 2008;68(9):3286–3294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Kim GB, Rincon Fernandez Pacheco D, Saxon D, et al. Rapid generation of somatic mouse mosaics with locus-specific, stably integrated transgenic elements. Cell. 2019;179(1):251–267.e24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Comba A, Faisal SM, Dunn PJ, et al. Spatiotemporal analysis of glioma heterogeneity reveals COL1A1 as an actionable target to disrupt tumor progression. Nat Commun. 2022;13(1):3606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Hu X, Pandolfi PP, Li Y, et al. mTOR promotes survival and astrocytic characteristics induced by Pten/AKT signaling in glioblastoma. Neoplasia. 2005;7(4):356–368. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Quelle DE, Zindy F, Ashmun RA, Sherr CJ.. Alternative reading frames of the INK4a tumor suppressor gene encode two unrelated proteins capable of inducing cell cycle arrest. Cell. 1995;83(6):993–1000. [DOI] [PubMed] [Google Scholar]
- 40. Gallagher SJ, Kefford RF, Rizos H.. The ARF tumour suppressor. Int J Biochem Cell Biol. 2006;38(10):1637–1641. [DOI] [PubMed] [Google Scholar]
- 41. Kim WY, Sharpless NE.. The regulation of INK4/ARF in cancer and aging. Cell. 2006;127(2):265–275. [DOI] [PubMed] [Google Scholar]
- 42. Uhrbom L, Dai C, Celestino JC, et al. Ink4a-Arf loss cooperates with KRas activation in astrocytes and neural progenitors to generate glioblastomas of various morphologies depending on activated Akt. Cancer Res. 2002;62(19):5551–5558. [PubMed] [Google Scholar]
- 43. Lai A, Kharbanda S, Pope WB, et al. Evidence for sequenced molecular evolution of IDH1 mutant glioblastoma from a distinct cell of origin. J Clin Oncol. 2011;29(34):4482–4490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Watanabe T, Nobusawa S, Kleihues P, Ohgaki H.. IDH1 mutations are early events in the development of astrocytomas and oligodendrogliomas. Am J Pathol. 2009;174(4):1149–1153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Parsons DW, Jones S, Zhang X, et al. An integrated genomic analysis of human glioblastoma multiforme. Science. 2008;321(5897):1807–1812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Nekrutenko A, Hillis DM, Patton JC, Bradley RD, Baker RJ.. Cytosolic isocitrate dehydrogenase in humans, mice, and voles and phylogenetic analysis of the enzyme family. Mol Biol Evol. 1998;15(12):1674–1684. [DOI] [PubMed] [Google Scholar]
- 47. Xu X, Zhao J, Xu Z, et al. Structures of human cytosolic NADP-dependent isocitrate dehydrogenase reveal a novel self-regulatory mechanism of activity. J Biol Chem. 2004;279(32):33946–33957. [DOI] [PubMed] [Google Scholar]
- 48. Dang L, White DW, Gross S, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2010;465(7300):966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Ye D, Xiong Y, Guan KL.. The mechanisms of IDH mutations in tumorigenesis. Cell Res. 2012;22(7):1102–1104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Jiang B, Zhang J, Xia J, et al. IDH1 mutation promotes tumorigenesis by inhibiting JNK activation and apoptosis induced by serum starvation. Cell Rep. 2017;19(2):389–400. [DOI] [PubMed] [Google Scholar]
- 51. Bardella C, Al-Dalahmah O, Krell D, et al. Expression of Idh1(R132H) in the murine subventricular zone stem cell niche recapitulates features of early gliomagenesis. Cancer Cell. 2016;30(4):578–594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Sasaki M, Knobbe CB, Itsumi M, et al. D-2-hydroxyglutarate produced by mutant IDH1 perturbs collagen maturation and basement membrane function. Genes Dev. 2012;26(18):2038–2049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Bunse L, Pusch S, Bunse T, et al. Suppression of antitumor T cell immunity by the oncometabolite (R)-2-hydroxyglutarate. Nat Med. 2018;24(8):1192–1203. [DOI] [PubMed] [Google Scholar]
- 54. Nunez FJ, Mendez FM, Kadiyala P, et al. IDH1-R132H acts as a tumor suppressor in glioma via epigenetic up-regulation of the DNA damage response. Sci Transl Med. 2019;11(479):eaaq1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Ding H, Roncari L, Shannon P, et al. Astrocyte-specific expression of activated p21-ras results in malignant astrocytoma formation in a transgenic mouse model of human gliomas. Cancer Res. 2001;61(9):3826–3836. [PubMed] [Google Scholar]
- 56. Shchors K, Persson AI, Rostker F, et al. Using a preclinical mouse model of high-grade astrocytoma to optimize p53 restoration therapy. Proc Natl Acad Sci U S A. 2013;110(16):E1480–E1489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Marumoto T, Tashiro A, Friedmann-Morvinski D, et al. Development of a novel mouse glioma model using lentiviral vectors. Nat Med. 2009;15(1):110–116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Seyfried TN, el-Abbadi M, Roy ML.. Ganglioside distribution in murine neural tumors. Mol Chem Neuropathol. 1992;17(2):147–167. [DOI] [PubMed] [Google Scholar]
- 59. Szatmari T, Lumniczky K, Desaknai S, et al. Detailed characterization of the mouse glioma 261 tumor model for experimental glioblastoma therapy. Cancer Sci. 2006;97(6):546–553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Chen X, Zhang L, Zhang IY, et al. RAGE expression in tumor-associated macrophages promotes angiogenesis in glioma. Cancer Res. 2014;74(24):7285–7297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Licon-Munoz Y, Avalos V, Subramanian S, et al. Single-nucleus and spatial landscape of the sub-ventricular zone in human glioblastoma. bioRxiv. 2024. [Google Scholar]
- 62. da Silva Siqueira L, Majolo F, da Silva APB, da Costa JC, Marinowic DR.. Neurospheres: a potential in vitro model for the study of central nervous system disorders. Mol Biol Rep. 2021;48(4):3649–3663. [DOI] [PubMed] [Google Scholar]
- 63. Binello E, Qadeer ZA, Kothari HP, Emdad L, Germano IM.. Stemness of the CT-2A immunocompetent mouse brain tumor model: characterization in vitro. J Cancer. 2012;3:166–174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Martinez-Murillo R, Martinez A.. Standardization of an orthotopic mouse brain tumor model following transplantation of CT-2A astrocytoma cells. Histol Histopathol. 2007;22(12):1309–1326. [DOI] [PubMed] [Google Scholar]
- 65. Ahmad M, Frei K, Willscher E, et al. How stemlike are sphere cultures from long-term cancer cell lines? Lessons from mouse glioma models. J Neuropathol Exp Neurol. 2014;73(11):1062–1077. [DOI] [PubMed] [Google Scholar]
- 66. Fraser H. Astrocytomas in an inbred mouse strain. J Pathol. 1971;103(4):266–270. [DOI] [PubMed] [Google Scholar]
- 67. Serano RD, Pegram CN, Bigner DD.. Tumorigenic cell culture lines from a spontaneous VM/Dk murine astrocytoma (SMA). Acta Neuropathol. 1980;51(1):53–64. [DOI] [PubMed] [Google Scholar]
- 68. Bradford R, Darling JL, Thomas DG.. The in-vitro chemosensitivity of three cell lines derived from the VM/DK spontaneous murine astrocytoma. J Neurol Neurosurg Psychiatry. 1986;49(12):1361–1366. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Chakraborty A, Yang C, Kresak JL, et al. KR158 spheres harboring slow-cycling cells recapitulate GBM features in an immunocompetent system. bioRxiv. 2024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Costa B, Fletcher MNC, Boskovic P, et al. A set of cell lines derived from a genetic murine glioblastoma model recapitulates molecular and morphological characteristics of human tumors. Cancers (Basel). 2021;13(2):230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Costa B, Eisemann T, Strelau J, et al. Intratumoral platelet aggregate formation in a murine preclinical glioma model depends on podoplanin expression on tumor cells. Blood Adv. 2019;3(7):1092–1102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Wiesner SM, Decker SA, Larson JD, et al. De novo induction of genetically engineered brain tumors in mice using plasmid DNA. Cancer Res. 2009;69(2):431–439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Kosaka A, Ohkuri T, Okada H.. Combination of an agonistic anti-CD40 monoclonal antibody and the COX-2 inhibitor celecoxib induces anti-glioma effects by promotion of type-1 immunity in myeloid cells and T-cells. Cancer Immunol Immunother. 2014;63(8):847–857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Knobbe CB, Reifenberger J, Reifenberger G.. Mutation analysis of the Ras pathway genes NRAS, HRAS, KRAS and BRAF in glioblastomas. Acta Neuropathol. 2004;108(6):467–470. [DOI] [PubMed] [Google Scholar]
- 75. Zamler DB, Shingu T, Kahn LM, et al. Immune landscape of a genetically engineered murine model of glioma compared with human glioma. JCI Insight. 2022;7(12):e148990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Cheema TA, Wakimoto H, Fecci PE, et al. Multifaceted oncolytic virus therapy for glioblastoma in an immunocompetent cancer stem cell model. Proc Natl Acad Sci U S A. 2013;110(29):12006–12011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Sampetrean O, Saga I, Nakanishi M, et al. Invasion precedes tumor mass formation in a malignant brain tumor model of genetically modified neural stem cells. Neoplasia. 2011;13(9):784–791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Hide T, Takezaki T, Nakatani Y, et al. Sox11 prevents tumorigenesis of glioma-initiating cells by inducing neuronal differentiation. Cancer Res. 2009;69(20):7953–7959. [DOI] [PubMed] [Google Scholar]
- 79. Glasgow SM, Zhu W, Stolt CC, et al. Mutual antagonism between Sox10 and NFIA regulates diversification of glial lineages and glioma subtypes. Nat Neurosci. 2014;17(10):1322–1329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Lee JH, Lee JE, Kahng JY, et al. Human glioblastoma arises from subventricular zone cells with low-level driver mutations. Nature. 2018;560(7717):243–247. [DOI] [PubMed] [Google Scholar]
- 81. Tritz ZP, Ayasoufi K, Johnson AJ.. Anti-PD-1 checkpoint blockade monotherapy in the orthotopic GL261 glioma model: the devil is in the detail. Neurooncol. Adv.. 2021;3(1):vdab066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Ladomersky E, Zhai L, Lenzen A, et al. IDO1 inhibition synergizes with radiation and PD-1 blockade to durably increase survival against advanced glioblastoma. Clin Cancer Res. 2018;24(11):2559–2573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Zhang P, Miska J, Lee-Chang C, et al. Therapeutic targeting of tumor-associated myeloid cells synergizes with radiation therapy for glioblastoma. Proc Natl Acad Sci U S A. 2019;116(47):23714–23723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Genoud V, Marinari E, Nikolaev SI, et al. Responsiveness to anti-PD-1 and anti-CTLA-4 immune checkpoint blockade in SB28 and GL261 mouse glioma models. Oncoimmunology. 2018;7(12):e1501137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Bayik D, Zhou Y, Park C, et al. Myeloid-derived suppressor cell subsets drive glioblastoma growth in a sex-specific manner. Cancer Discov. 2020;10(8):1210–1225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Hung AL, Maxwell R, Theodros D, et al. TIGIT and PD-1 dual checkpoint blockade enhances antitumor immunity and survival in GBM. Oncoimmunology. 2018;7(8):e1466769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Kim JE, Patel MA, Mangraviti A, et al. Combination therapy with anti-PD-1, Anti-TIM-3, and focal radiation results in regression of murine gliomas. Clin Cancer Res. 2017;23(1):124–136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Zeng J, See AP, Phallen J, et al. Anti-PD-1 blockade and stereotactic radiation produce long-term survival in mice with intracranial gliomas. Int J Radiat Oncol Biol Phys. 2013;86(2):343–349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Enriquez Perez J, Kopecky J, Visse E, Darabi A, Siesjo P.. Convection-enhanced delivery of temozolomide and whole cell tumor immunizations in GL261 and KR158 experimental mouse gliomas. BMC Cancer. 2020;20(1):7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Karachi A, Yang C, Dastmalchi F, et al. Modulation of temozolomide dose differentially affects T-cell response to immune checkpoint inhibition. Neuro Oncol. 2019;21(6):730–741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Vom Berg J, Vrohlings M, Haller S, et al. Intratumoral IL-12 combined with CTLA-4 blockade elicits T cell-mediated glioma rejection. J Exp Med. 2013;210(13):2803–2811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Nair S, Mazzoccoli L, Jash A, et al. Zika virus oncolytic activity requires CD8+ T cells and is boosted by immune checkpoint blockade. JCI Insight. 2021;6(1):e144619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Amoozgar Z, Kloepper J, Ren J, et al. Targeting Treg cells with GITR activation alleviates resistance to immunotherapy in murine glioblastomas. Nat Commun. 2021;12(1):2582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94. Berger G, Knelson EH, Jimenez-Macias JL, et al. STING activation promotes robust immune response and NK cell-mediated tumor regression in glioblastoma models. Proc Natl Acad Sci U S A. 2022;119(28):e2111003119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Simonds EF, Lu ED, Badillo O, et al. Deep immune profiling reveals targetable mechanisms of immune evasion in immune checkpoint inhibitor-refractory glioblastoma. J ImmunoTher Cancer. 2021;9(6):e002181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Chen D, Varanasi SK, Hara T, et al. CTLA-4 blockade induces a microglia-Th1 cell partnership that stimulates microglia phagocytosis and anti-tumor function in glioblastoma. Immunity. 2023;56(9):2086–2104.e8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Saha D, Martuza RL, Rabkin SD.. Macrophage polarization contributes to glioblastoma eradication by combination immunovirotherapy and immune checkpoint blockade. Cancer Cell. 2017;32(2):253–267.e5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Flores-Toro JA, Luo D, Gopinath A, et al. CCR2 inhibition reduces tumor myeloid cells and unmasks a checkpoint inhibitor effect to slow progression of resistant murine gliomas. Proc Natl Acad Sci U S A. 2020;117(2):1129–1138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Khalsa JK, Cheng N, Keegan J, et al. Immune phenotyping of diverse syngeneic murine brain tumors identifies immunologically distinct types. Nat Commun. 2020;11(1):3912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Iorgulescu JB, Gokhale PC, Speranza MC, et al. Concurrent dexamethasone limits the clinical benefit of immune checkpoint blockade in glioblastoma. Clin Cancer Res. 2021;27(1):276–287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Speranza MC, Passaro C, Ricklefs F, et al. Preclinical investigation of combined gene-mediated cytotoxic immunotherapy and immune checkpoint blockade in glioblastoma. Neuro Oncol. 2018;20(2):225–235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Chandramohan V, Bao X, Yu X, et al. Improved efficacy against malignant brain tumors with EGFRwt/EGFRvIII targeting immunotoxin and checkpoint inhibitor combinations. J ImmunoTher Cancer. 2019;7(1):142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Johanns TM, Ward JP, Miller CA, et al. Endogenous neoantigen-specific CD8 T cells identified in two glioblastoma models using a cancer immunogenomics approach. Cancer Immunol Res. 2016;4(12):1007–1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Fecci PE, Ochiai H, Mitchell DA, et al. Systemic CTLA-4 blockade ameliorates glioma-induced changes to the CD4+ T cell compartment without affecting regulatory T-cell function. Clin Cancer Res. 2007;13(7):2158–2167. [DOI] [PubMed] [Google Scholar]
- 105. Flores CT, Wildes TJ, Drake JA, et al. Lin(–)CCR2(+) hematopoietic stem and progenitor cells overcome resistance to PD-1 blockade. Nat Commun. 2018;9(1):4313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Flores C, Pham C, Snyder D, et al. Novel role of hematopoietic stem cells in immunologic rejection of malignant gliomas. Oncoimmunology. 2015;4(3):e994374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Wildes TJ, Grippin A, Dyson KA, et al. Cross-talk between T Cells and hematopoietic stem cells during adoptive cellular therapy for malignant glioma. Clin Cancer Res. 2018;24(16):3955–3966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108. Gangoso E, Southgate B, Bradley L, et al. Glioblastomas acquire myeloid-affiliated transcriptional programs via epigenetic immunoediting to elicit immune evasion. Cell. 2021;184(9):2454–2470.e26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109. Shingu T, Ho AL, Yuan L, et al. Qki deficiency maintains stemness of glioma stem cells in suboptimal environment by downregulating endolysosomal degradation. Nat Genet. 2017;49(1):75–86. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110. Chen CH, Chin RL, Hartley GP, et al. Novel murine glioblastoma models that reflect the immunotherapy resistance profile of a human disease. Neuro Oncol. 2023;25(8):1415–1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Johnson M, Bell A, Lauing KL, et al. Advanced age in humans and mouse models of glioblastoma show decreased survival from extratumoral influence. Clin Cancer Res. 2023;29(23):4973–4989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Choe JH, Watchmaker PB, Simic MS, et al. SynNotch-CAR T cells overcome challenges of specificity, heterogeneity, and persistence in treating glioblastoma. Sci Transl Med. 2021;13(591):eabe7378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Hegi ME, Diserens AC, Gorlia T, et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005;352(10):997–1003. [DOI] [PubMed] [Google Scholar]
- 114. Kushihara Y, Tanaka S, Kobayashi Y, et al. Glioblastoma with high O6-methyl-guanine DNA methyltransferase expression are more immunologically active than tumors with low MGMT expression. Front Immunol. 2024;15:1328375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Liu L, van Schaik TA, Chen KS, et al. Establishment and immune phenotyping of patient-derived glioblastoma models in humanized mice. Front Immunol. 2023;14:1324618. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116. Ashizawa T, Iizuka A, Nonomura C, et al. Antitumor effect of programmed death-1 (PD-1) blockade in humanized the NOG-MHC double knockout mouse. Clin Cancer Res. 2017;23(1):149–158. [DOI] [PubMed] [Google Scholar]
- 117. Srivastava R, Labani-Motlagh A, Chen A, et al. Development of a human glioblastoma model using humanized DRAG mice for immunotherapy. Antib Ther. 2023;6(4):253–264. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118. Klawitter M, El-Ayoubi A, Buch J, et al. The oncolytic adenovirus XVir-N-31, in combination with the blockade of the PD-1/PD-L1 axis, conveys abscopal effects in a humanized glioblastoma mouse model. Int J Mol Sci . 2022;23(17):9965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Lei L, Sonabend AM, Guarnieri P, et al. Glioblastoma models reveal the connection between adult glial progenitors and the proneural phenotype. PLoS One. 2011;6(5):e20041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120. Li Y, Li B, Li W, et al. Murine models of IDH-wild-type glioblastoma exhibit spatial segregation of tumor initiation and manifestation during evolution. Nat Commun. 2020;11(1):3669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Verma R, Chen X, Xin D, et al. Olig1/2-expressing intermediate lineage progenitors are predisposed to PTEN/p53-loss-induced gliomagenesis and harbor specific therapeutic vulnerabilities. Cancer Res. 2023;83(6):890–905. [DOI] [PubMed] [Google Scholar]
- 122. Zhu H, Acquaviva J, Ramachandran P, et al. Oncogenic EGFR signaling cooperates with loss of tumor suppressor gene functions in gliomagenesis. Proc Natl Acad Sci U S A. 2009;106(8):2712–2716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. Garg AD, Vandenberk L, Van Woensel M, et al. Preclinical efficacy of immune-checkpoint monotherapy does not recapitulate corresponding biomarkers-based clinical predictions in glioblastoma. Oncoimmunology. 2017;6(4):e1295903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Park J, Kim CG, Shim JK, et al. Effect of combined anti-PD-1 and temozolomide therapy in glioblastoma. Oncoimmunology. 2019;8(1):e1525243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125. Jahan N, Talat H, Alonso A, Saha D, Curry WT.. Triple combination immunotherapy with GVAX, anti-PD-1 monoclonal antibody, and agonist anti-OX40 monoclonal antibody is highly effective against murine intracranial glioma. Oncoimmunology. 2019;8(5):e1577108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126. Reardon DA, Gokhale PC, Klein SR, et al. Glioblastoma eradication following immune checkpoint blockade in an orthotopic, immunocompetent model. Cancer Immunol Res. 2016;4(2):124–135. [DOI] [PubMed] [Google Scholar]
- 127. Dixon KO, Schorer M, Nevin J, et al. Functional anti-TIGIT antibodies regulate development of autoimmunity and antitumor immunity. J Immunol. 2018;200(8):3000–3007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Kim SS, Harford JB, Moghe M, et al. A tumor-targeting nanomedicine carrying the p53 gene crosses the blood–brain barrier and enhances anti-PD-1 immunotherapy in mouse models of glioblastoma. Int J Cancer. 2019;145(9):2535–2546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129. Belcaid Z, Berrevoets C, Choi J, et al. Low-dose oncolytic adenovirus therapy overcomes tumor-induced immune suppression and sensitizes intracranial gliomas to anti-PD-1 therapy. Neurooncol Adv. 2020;2(1):vdaa011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130. Wainwright DA, Chang AL, Dey M, et al. Durable therapeutic efficacy utilizing combinatorial blockade against IDO, CTLA-4, and PD-L1 in mice with brain tumors. Clin Cancer Res. 2014;20(20):5290–5301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Yang F, He Z, Duan H, et al. Synergistic immunotherapy of glioblastoma by dual targeting of IL-6 and CD40. Nat Commun. 2021;12(1):3424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132. Liu CJ, Schaettler M, Blaha DT, et al. Treatment of an aggressive orthotopic murine glioblastoma model with combination checkpoint blockade and a multivalent neoantigen vaccine. Neuro Oncol. 2020;22(9):1276–1288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133. Weiss T, Weller M, Guckenberger M, Sentman CL, Roth P.. NKG2D-based CAR T cells and radiotherapy exert synergistic efficacy in glioblastoma. Cancer Res. 2018;78(4):1031–1043. [DOI] [PubMed] [Google Scholar]