Abstract

Throughout the family of coronaviruses, structured RNA elements within the 5′ region of the genome are highly conserved. The fifth stem-loop element from SARS-CoV-2 (5_SL5) represents an example of an RNA structural element, repeatedly occurring in coronaviruses. It contains a conserved, repetitive fold within its substructures SL5a and SL5b. We herein report the detailed characterization of the structure and dynamics of elements SL5a and SL5b that are located immediately upstream of the SARS-CoV-2 ORF1a/b start codon. Exploiting the unique ability of solution NMR methods, we show that the structures of both apical loops are modulated by structural differences in the remote parts located in their stem regions. We further integrated our high-resolution models of SL5a/b into the context of full-length 5_SL5 structures by combining different structural biology methods. Finally, we evaluated the impact of the two most common VoC mutations within 5_SL5 with respect to individual base-pair stability.
Introduction
In late 2019, the outbreak of COVID-19, caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), initiated research efforts worldwide. The majority of structural biology initiatives focused on the structure determination of the viral proteins and their complexes. Using nuclear magnetic resonance spectroscopy (NMR spectroscopy, short: NMR), we investigated the RNA genome of SARS-CoV-2, demonstrating that 15 phylogenetically conserved RNA elements located in the 5′-genomic end, the 3′-untranslated region, and the frame-shifting region constitute autonomously folded RNA elements.1,2 We determined their secondary structures by NMR experiments and DMS probing-based cross-validation, while other groups employed inline probing,3 SHAPE,4 and also DMS probing.1,4−6 Eight RNA stem-loop elements within the 5′-genomic end were shown to be involved in numerous virus–virus or virus–host RNA–RNA and RNA–protein interactions.7−9 In contrast to the vast number of structures, covering nearly the complete viral proteome, only a small number of 3D structures of the viral RNAs have been reported to date.6,10−15
The SL5 elements of alpha and beta coronaviruses conserve a similar structure.6,16,17 Sequence and structure comparisons of SL5 elements among different coronaviruses show that SL5 typically consists of a well-conserved subset of helical elements arranged in a four-way junction.18 In both alpha and beta genera, SL5 includes part of the coding sequence.19 Even prior to the emergence of SARS-CoV-2, the importance of the large SL5 element was illustrated by its high conservation. Especially the highly conserved hexaloop sequences, formed by UUYCGU repetitive structural motifs (RSMs), have been described for nearly all coronaviruses.20 To date, high-resolution structural characterizations of these RSMs are not available.
The 5_SL5 (stem-loop 5 within the 5′-genomic end) element of SARS-CoV-2 comprises nucleotides 149 to 297. It contains the AUG translation start codon of the nonstructural polypeptide chains (location: A266-U267-G268). Early on, its tertiary structure has been modeled by FARFAR2,21 and recently, cryogenic electron microscopy (cryo-EM) structures have been determined.6,11 Consistently, the overall architecture features a four-way junction with a T-shaped structure composed of three large stems entitled 5stem, 5a, and 5b, as well as an additional small 5c-stem (Supporting Information Figure S1A).
Here, we investigate the two individual stems 5_SL5a and 5_SL5b, capped by UUYCGU RSMs. The RSM of the two stem loops extends to a sequence identity of 10 nucleotides, comprising the six loop nucleotides and the two GC-closing base pairs: G198–C207 for SL5a and G236–C245 for SL5b, respectively.
During SARS-CoV-2 evolution, two major mutations have occurred: G210U in SL5a and C241U in SL5b (GISAID database, https://gisaid.org/hcov19-variants/). While the mutation in SL5a exclusively occurred in the delta variant of the virus, the mutation in SL5b is present in all lineages except the original one. The G210U mutation is located within a bulge region of SL5a, while the C241U mutation is located at position 4 within the hexaloop of SL5b. Interestingly, the mutations thus affected only one of the two RSMs, removing their perfect identity.
In this article, we report the solution structures of the two RNA subelements SL5a and SL5b derived from NMR experiments to collect extensive NMR chemical shifts, NOEs, RDC, and scalar coupling data. We complement those structures with insights in dynamics and individual base-pair stabilities employing hetNOE data and proton-filtered 13C-detected and hydrogen exchange experiments. Comparing the NMR signatures of isolated 5a and 5b elements to their counterparts within the full-length SL5,1 we find that both elements maintain their overall fold in isolation. We further show by small-angle X-ray scattering (SAXS) data that identical global folds are obtained for the entire 5_SL5 element compared to structures previously obtained by cryo-EM. Our solution NMR data augment these studies in atomic detail, revealing unexpected differences with regard to structure and dynamics in the sequentially identical RSM regions of 5a and 5b. These differences are also observed for sequences containing the two major mutations G210U and C241U.
Taken together, this work represents a valuable contribution to our understanding of the SARS-CoV-2 genome structure, manifested in the high-resolution structures and the dynamic nature of two crucial RNA elements.
Materials and Methods
RNA Synthesis
RNA samples were prepared as described in refs (1) and (15) (for sequences see Supporting Information Table S1). Additionally, an unlabeled SL5b RNA sample was purchased from Horizon Discovery and was used for SAXS and selected NMR measurements (as indicated). Briefly, if synthesized in-house, previously amplified and linearized double-stranded template DNA was incubated for 6 h at 37 °C together with the bacteriophage-derived RNA-polymerase T7 along with cofactors and substrates (nucleoside triphosphates, Mg2+, dithiothreitol, and spermidine) buffered in 0.2 M Tris–HCl pH 8.0. In vitro transcribed RNA was purified via polyacrylamide gel purification under denaturing conditions and with reverse-phase (RP) high-performance liquid chromatography (HPLC). Final RNA samples were buffer-exchanged to 25 mM potassium phosphate (KPi, pH 6.2) and 50 mM potassium chloride (KCl) (NMR buffer) using 2- to 3-kDa molecular weight cutoff (MWCO) VivaSpin filtration units (Sartorius).
NMR Experiments
Additional NMR chemical shift assignments augmenting previous data deposited in BMRB (see Supporting Information Table S2) and assignments for the mutation constructs were carried out as previously described.1,22,23 For structure calculations, additional nuclear Overhauser enhancement and exchange spectroscopy (NOESY) spectra with homogeneous excitation profiles24 for nuclear Overhauser effect (NOE)-derived distance restraints were measured each for the exchangeable protons (in 5% D2O; 298 K; 75, 100, or 150 ms NOESY mixing time) and for nonexchangeable protons (in 99.95% D2O; 298 K; 75, 100, or 150 ms NOESY mixing times), respectively. For the incorporation of residual dipolar couplings (RDCs), a NOESY spectrum of SL5b 13C,15N-labeled RNA with a GC-stabilized sequence was used with a mixing of 150 ms. 13C{F2}-filtered NOESY25 spectra were measured and used for final calculation of SL5b. NOESY peak picking and peak list generation were performed with NMRFAM-Sparky.26
RDC Determination
NH and CH RDC values were extracted from the difference in coupled in-phase/antiphase (IPAP) 1H, 15N/1H, and 13C correlation experiments in the direct dimension. One set of couplings was measured in Pf1 filamentous bacteriophage-aligned sample versus nonaligned RNA sample. Partial alignment of SL5a and SL5b was achieved by the addition of 10–20 mg/mL Pf1 phages, yielding deuterium splittings of 22 Hz for SL5a and 9 Hz for SL5b at 800 MHz. Peak picking and RDC determination were performed in NMRFAM-Sparky. RDC values were used to iteratively estimate the rhombicity (final value: SL5a = 0.06, SL5b = 0.45) and anisotropy coefficients (final value: SL5a = 12, SL5b = −10) of the alignment tensor by using preliminary 3D coordinates generated from structure calculations without the incorporation of RDCs in PALES.27 Additionally, we scaled down the size of the RDCs for SL5a by a factor of 4 due to the large concentration of phages and the resulting large size of the RDCs (60–80 Hz). Scaling down the RDCs allowed easier integration into ARIA. To confirm this approach, we back-calculated the scaled and unscaled RDCs using PALES from our ARIA calculations. There were no differences observed between the scaled and unscaled RDCs.
Mg2+ Titration and Chemical Shift Perturbation Analysis
For the Mg2+ titrations, 15N-labeled RNA was used for all of the experiments. Mg2+ solutions were added to yield final concentration between 0.5 and 4 mM (final concentration in sample volume) by addition of MgCl2 stock solution (1 M in NMR buffer). Chemical shift perturbations Δδ (CSPs) were calculated using referenced resonance assignment and the formula 1 stated in (28). 1H chemical shifts were referenced to DSS and 13C and 15N chemical shifts were indirectly referenced from the 1H chemical shift as previously described.29
| 1 |
Base-Pair Stabilities from Cross-Hydrogen Bond Scalar Couplings and Solvent Exchange Measurements
Experiments for measurement of 2hJ(N,N) couplings were conducted using the sequence according to ref (30). The transfer delay (Δ) was set to 30 ms, and experiments were performed in a temperature range between 278 and 313 K with 5 K steps. The couplings were calculated by formula 2, quantifying peak integral intensities of donor and acceptor (I(Nx)) nitrogen to the imino proton from the 1H,15N–HNN–COSY experiment.
| 2 |
To characterize base-pair stabilities, exchange rates of imino resonances with the solvent were determined. 15N- or 13C,15N-labeled samples were dissolved in an aqueous NMR buffer containing 5% D2O. 1H-detected inversion recovery experiments and their analysis were conducted according to ref (31). Measurements were performed at temperatures 278 to 313 K (5 K steps) with exchange delays 4 μs to 4 s. The temperature-dependent kex values for the H–N signals were estimated by fitting intensities to delay. Those were then used to calculate the ΔGdiss (Gibbs free enthalpy of dissociation) by ΔHdiss and ΔSdiss from fit of kex in formula 3, as described in ref (32).
 |
3 |
13C-detected 13C,15N-HSQC experiments33 were measured using a 13C,15N-labeled RNA sample in aqueous NMR buffer containing 5% D2O. Proton-filtered variants of these experiments were conducted at 298 K. Exchange delays were 0 ms (no exchange), 5.3 ms matching a 1J(N,H) coupling of 94 Hz (“proton filter”), and 10.6 ms with 47 Hz for intensity trace analysis.
Determination of Fast Internal Motions
{1H},13C hetNOEs were measured using constant-time chemical shift evolution in the 13C dimension, with and without NOE buildup in an interleaved manner and with temperature compensation. The interscan delay was set to 5 s and the number of scans was set to 64 to achieve a signal-to-noise ratio >10 for both NOE and noNOE spectra. hetNOE values were calculated by the formula hetNOE = (Isat/Iref) as described in ref (34). Errors were calculated based on the signal-to-noise ratios by error propagation.
Structure Calculations
Structure calculations were performed with ARIA 1.235 using CNS 1.136 and the adapted force field DNA-RNA-all-atom-hj.37 NOE restraints from NOESY spectra with mixing times of 75, 100, and 150 ms were initially incorporated (Figure 1), yielding a total number of 370 unambiguous NOE restraints for SL5a and 298 for SL5b, respectively. Spin diffusion was corrected within the ARIA routine. We further restrained H bonds for intra-base-pair 2hJ(N,N) couplings >4 Hz and hydrogen exchange rates >4 kJ/mol and applied base coplanarities for these base pairs. Initial dihedral angle restraints (dihedrals) were defined for the assumed A-form helical parts of the RNAs according to their secondary structure [SL5a: g-188:193; 196:199; 206:209; 213:218-c and SL5b: (g-227/)228:233; 236; 244:252(-cc)]. Nucleotides were restrained to ribose 3′-endo conformation with the exception of 2′-endo for U202 and C203 in SL5a and U240 and C241 in SL5b, in agreement with canonical coordinates derived from ribose 13C chemical shifts.22,23 The glycosidic torsion angles χ for nucleotides showing weak-to-moderate H1′-H8 (purine) or H1′-H6 (pyrimidine) NOE intensities were set to anticonformation. Calculations of stem-loop 5a were conducted for a GC-stabilized construct (g-188-218-c) with and without 13C,15N-labeling. NOE distance restraints for 5b were obtained for a construct representing the native stem (228–252) without GC modification. The final SL5b NOE distance list was incorporated to the GC-stabilized system (g-227-252-cc), as chemical shift assignments could successfully be transferred (see ref (23)). For the iterative improvement of calculated structures, RDCs for aromatic CH resonances were incorporated. NH RDCs were left out for cross-validation of the final structures.
Figure 1.
Schematic representation of the RNA structure calculation workflow. The iterative process was repeated until the structure bundles reached convergence. Convergence was defined as an average >0.98 for back-calculated CORR R values by PALES.
An iterative process was performed (Figure 1) in which dihedral angle restraints were repeatedly adapted based on geometric analysis of converged structures. H-bond restraints were iteratively adjusted according to convergence of the structures and based on chemical shift similarities and RDC analysis. Structure calculations were repeated until an agreement with the full set of RDCs of >0.98 was achieved. The complete list of final structure calculation restraints is included in the Supporting Information (Supporting Information Tables S3 and S4). A total of eight iterations were calculated for each individual structure calculation, with 100 structures per iteration. The 20 lowest-energy structures were used for each following iteration. The final iteration was performed with a total of 200 structures where the 20 lowest-energy structures were subjected to water refinement in ARIA. The final 20 lowest-energy structures were filtered according to their correlation of back-calculated RDCs with the total set of 64 experimental RDCs for SL5a and 52 for SL5b. The correlation score was determined with PALES using the bestFit option.27 Clash scores and structure geometries were analyzed with MolProbity and w3DNA.38,39 Finally, 3D coordinates of the bundle of the best 10 structures were converted to mmCIF format using the wwPDB validation system Web servers (https://validate.wwpdb.org).39 PDB entries are 9FO9 (SL5a) and 9FO8 (SL5b).
Rigid-Body Fitting
We employed a two-step protocol of rigid-body fitting using standard tools within the UCSF Chimera software package:40 (I) structural alignment of our entire NMR bundles to the deposited cryo-EM structures11 [EM density map EMD-42818, atomic structure bundle PDB 8UYS] based on the sequence using the MatchMaker routine, followed by (II) rigid-body fitting of the sequence-aligned structures into the cryo-EM 3D map using the Fit in Map routine. This process was conducted individually for each of the 20 atomic structures within each bundle.
SAXS Experiments
SAXS data for 5_SL5 and SL5a were collected on the EMBL P12 beamline at the PETRA III storage ring (DESY, Hamburg, Germany) by using a Pilatus 2 M detector at a sample–detector distance of 3 m and a wavelength of λ = 0.124 nm. Data were measured at 298 K at continuous flow with a total exposure time of 3.895 s (40 × 95 ms frames). SAXS data for SL5b_native were acquired at the BM29 BioSAXS beamline of the European Synchrotron Radiation Facility in Grenoble, France, in remote measurement mode. Sample buffers were identical to NMR conditions except for lacking D2O. Sample volumes were 60 μL at a concentration of 2.5 mg/mL. Samples were measured by exposure at 12.5 keV beam energy; scattering was acquired on a Pilatus3 2 M detector placed under vacuum and with local standard setups of distance and beam size. The herein used q range for SL5b was 0.0388–5.1388 nm–1 sampled by 1000 points. Acquired frames were automatically scanned for sufficient quality and to exclude radiation damage and summed up over a period of 5 s of measurement. For all RNAs, automated processing included normalization of scattering data to the intensity of the transmitted beam and radial averaging as well as subtraction of the buffer scattering data. SAXS scattering curves were plotted as I(s) vs s, where s = 4πsin θ/λ and 2θ is the scattering angle.
For SL5a, we recorded SAXS data from a sample of 1 mg/mL in NMR buffer without D2O at an angular space of 0.0256 < q < 7.3476 nm–1 covering a total of 2625 points. SAXS curve fits to experimental structures were calculated using CRYSOL via the ATSAS v 4.0.41,42 For full comparability of 5_SL5b NMR and SAXS (“native”) constructs, we used an in silico truncated version of our 5_SL5b ensemble lacking the two basal GC base pairs during the CRYSOL runs.
For the fl-SL5, we used SAXS data measured from a sample of 1 mg/mL (corresponding to 20 μM) in NMR buffer without D2O at an angular space of 0.0256 < q < 7.3476 nm–1 covering a total of 2625 points. SL5 SAXS curve fits to experimental structures were calculated using CRYSOL via the ATSAS online package.41,42 The particle distance distribution function P(r) plots were calculated using GNOM43 selecting the q range 0.1148 < s < 1.8504 nm–1 and further subjected to repetitive ab initio model building using DAMMIN (ATSAS online)44 setting P1 symmetry and no further anisometry. The model shown herein represents its underlying scattering curve through a χ2 value of 1.039. Theoretical Dmax and Rg values for PDB entry 8QO5 were obtained from CRYSOL and cross-checked via the HullRad v9 server.45
All SAXS data have been deposited in the SASBDB and entries are available through the IDs SASDVX2 (5_SL5), SASDVY2 (5_SL5a), and SASDVC3 (5_SL5b).
Results and Discussion
Definition of Autonomous Subelements by Detailed Analysis of the Chemical Shift Signature
We first confirmed that the subconstructs 5_SL5a and 5_SL5b+c (hereafter named SL5a and SL5b+c) retain their structure in the full-length 5_SL5 element (SL5) comprising nucleotides 149 to 297. A schematic and secondary structure representation can be found in the Supporting Information (Supporting Information Figure S1B) along with an assigned 1H,15N-TROSY overlay of 5_SL5 with the subelements 5a and 5b.
Titrations of full-length SL5 were performed to investigate whether SL5 changes conformation upon Mg2+ addition,46 possibly leading to involvement of 5a and/or 5b in tertiary interactions. Up to 4 mM Mg2+, very few changes in the SL5 reporter imino signal were observed. More importantly, for temperatures between 293 and 308 K, no additional signals were observed (Supporting Information Figure S2), confirming that the three elements retain their individual stable folds. The consistency of the imino signal pattern argues for a conserved local structural environment of SL5a and SL5b in the full SL5 context. Thus, it is unlikely that these stem loops are involved in significant tertiary architectures within SL5. We also performed Mg2+ titrations for isolated SL5a and SL5b+c (Supporting Information Figure S3), showing that while Mg2+ indeed has few pronounced local effects, global refolding or tertiary structure formation was also not observed for the isolated stem loops.
For SL5a, the largest observed CSPs were 0.12 ppm for G198 at 283 K and ∼0.1 ppm for G188 at 313 K. The latter is in line with Mg2+-induced stabilization of terminal base pairs at elevated temperatures (Supporting Information Figure S3).
For SL5b+c, the most significant differences are observed around the hinge (G252, N1H1 CSP4mM@283K = 0.09 ppm) between SL5b and SL5c (Supporting Information Figure S3, bottom, gray squares). The size of CSP is in the same range as those observed for SL5a. There are no significant Mg2+-induced changes in the stem part of SL5b (sequence 230–250). Given that the Mg2+-induced differences in SL5b+c localize to the hinge region between SL5b and SL5c and the observation that the structure in SL5c is not stable over a sufficiently large temperature range, we performed structure calculations for the isolated SL5b.
5a and 5b Resonance Assignment
We previously published resonance assignments of the subelements 5a,22 5b, and 5b+c.23 Here, we identified previously missing assignments for both stem loops and updated the corresponding BRMB IDs 50346 and 50339/51138 accordingly. Our previous assignments were derived from experiments conducted at 283 and 298 K. However, hydrogen atoms involved in more labile hydrogen bonds can often only be observed at lower temperatures due to reduced solvent exchange. To unambiguously identify all RSM residues and the individual residues of 5a′s bulge region, we thus also recorded spectra at 278 K (Supporting Information Figure S1C) and observed additional imino proton signals. Precisely, for 5a, we detected three signals in the spectral region characteristic for “wobble” base pairs (chemical shift of 11.37/157.80, 10.95/157.47, and 10.33/155.85 ppm (1H/15N)), indicating that three uridine residues form wobble-type interactions including a hydrogen-bonded imino proton. The assignment for the residues U194, U195, and U212 was performed by 1H,15N-TROSY combined with 1H,13C–H(N)CO47,48 and 13C,15N-HSQC,331H,13C-HSQC, and 1H,13C–H8(H6)–C8(C6)–N9(N1)–C4(C2)49 spectra. Further, ribose chemical shift assignments for SL5a as well as SL5b were obtained from the forward-directed 3D 1H,13C-HCC-TOCSY-CCH–COSY50,51 experiment and 3D 1H,13C-HC(C)H-TOCSY/3D 1H,13C-HCC(H)-TOCSY52 experiments. The RSM motifs of 5a and 5b were assigned through 1H,1H-NOESY sequential walks.
RSMs: Conformational Differences Revealed by Different Chemical Shifts
The sequences of SL5a and SL5b share an identity of 10 nucleotides containing two GC base pairs as loop closing and the characteristic core RSM, a 5′-UUUCGU-3′ hexaloop. The chemical shift assignment allowed us to compare similarities and differences in their overall architecture. We previously recorded 1H,13C-HSQC and 1H,1H-TOCSY spectra for nonexchangeable CH resonances, including the aromatic C5H5, C6H6, C8H8 and C2H2, and C1′H1′ regions. As shown in Figure 2, the chemical shifts of the 10 loop nucleotides differ for some of the aromatic CH (C6H6/C8H8 used here) and ribose C1′H1′ resonances despite the sequence identity. Interestingly, in this comparison, the largest chemical shift differences in the C1′H1′ resonance are observed for the two uridine nucleotides at the 3′ ends of the hexaloops, U205/U243 in SL5a/SL5b, respectively. Since the nucleotide sequences for the two RSMs are identical, the more remote regions (U194, U195, G210, U211, and U212 in SL5a and A235 in SL5b) that interrupt the regular A-form stems of 5a and 5b have to cause the subtle differences in RSM conformation detected via the differences in chemical shifts. Thus, structural differences in the remote bulges impact the structure and dynamics of the loop nucleotides even if these bulges are separated by 2–4 base pairs. Another striking observation is the difference of the C1′H1′ chemical shifts of nucleotides G204 and G242. While chemical shift reporters for U202/U240 and C203/C241 are rather similar, the NMR reporters of G204/G242 differ by a CS difference of 0.3 ppm. These trends are combined in Supporting Information Figure S4 which summarizes the CS differences of all aromatic C6H6/C8H8 and C1′H1′ resonances of the 5a and 5b RSMs. Further subtle differences between the RSM architecture of 5a and 5b are revealed when comparing the 1H,15N-TROSY of SL5b, which shows one weak additional and unusual chemical shift for a G nucleotide involved in a wobble hydrogen bond at 10.7 ppm/144 ppm (Supporting Information Figure S1C, also see ref (23)). This signal was assigned as loop G242 due to its characteristic chemical shift similar to the G of the well-characterized UUCG tetraloop.37 Detectability of this signal is temperature-dependent. The signal is no longer visible at 303 K and above. In contrast, the signal of the corresponding nucleotide G204 N1H1 is not observable in the 1H,15N-TROSY for SL5a, at neither 298 nor 278 K.
Figure 2.
Differences in 1H,13C-chemical shifts of aromatic C6H6/C8H8 (dark) and ribose C1′H1′ (light) in the sequence-identical loop regions of SL5a to SL5b. Secondary structure representations with the respective sequential numbering for comparison of SL5a (red) to SL5b (blue). Identical sequential loop region of 10 bases highlighted in the center. For an overlay of spectra, see Supporting Information Figure S4.
NMR Reveals Structural Dynamics within the Bulge and Loop Motifs
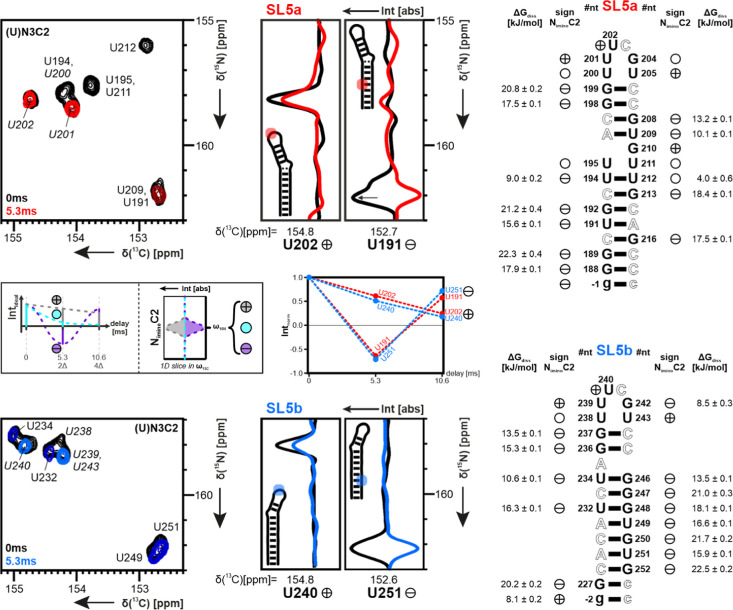
We comprehensively characterized the dynamics of the helical, bulge and loop regions of the two stem-loop RNAs recording atom-resolved heteronuclear {1H},13C-NOEs (hetNOE), 2hJ(N,N) (scalar coupling across hydrogen bonds),30 imino-solvent hydrogen exchange using 1H inversion recovery experiments,31,32,53 and low-γ-detected exchange-dependent 1J(HN)-evolution experiments33 (see underlying principles in Figure 4).
Figure 4.
Comparison of the base-pair properties for SL5a (top) and SL5b (bottom) via hydrogen-exchange-sensitive experiments. From left to right: 13C,15N-HSQC with exemplary 1D slices for U nucleotides without (black) and with a proton filter (colored), showing the impact on the phase.33 Scheme of analysis in small legend in-between. CN-HSQC measured at 298 K, with all corresponding peaks U N1C2 phased to a positive sign for comparison. Behavior of the signal sign in CN-HSQC with the proton filter present shown next to the sequence. See also Supporting Information Figures S6 and S7 for whole spectra. Also, next to the sequence, the stability of base pairs by ΔGdiss is calculated from kex of hydrogen exchange. Exchange rates measured in range of 278–313 K. Values of ΔGdiss highlighting the structure surfaces of SL5a (top) and SL5b (bottom). Watson–Crick (WC) base-pair partners A and C were colored the same as their U and G counterparts. The mapping shows the lower stabilities of the bonds around stem closing, bulge, and loop regions of the RNA. Color range from blue (4 kJ/mol) to red (23 kJ/mol), with transient bases colored in cyan. Structure models for the RNA are exemplary from the calculated 20 lowest-energy bundles.
hetNOE values report on motions faster than the rotational correlation time τC of the RNA and thus on locally flexible RNA residues. Our hetNOE data for 5a show consistently elevated values for the hexaloop C1′H1′ resonances, with U200 showing the greatest flexibility. Apart from the loop, G210 is by far the most flexible (C8H8 = 1.25, C1′H1’ = 1.35) nucleotide in the bulge region (colored red in Figure 3) of SL5a. The C1′,H1′-ribose moiety of U195 also shows an increased hetNOE in line with the nondetectability of the U195 imino proton at temperatures T > 278 K. In addition, the hetNOE of the aromatic C6H6 of U211 (hetNOE: 1.21) is slightly increased, indicating flexibility of the region comprising U195, G210, and U211. In contrast, hetNOE values for U194 and U212 are similar to the average value of stem residues, supporting the formation of a UU wobble base pair as indicated by the presence of their imino proton signals in the 1H,15N-TROSY at 278 K (Supporting Information Figure S1C).
Figure 3.
hetNOE values for aromatic and ribose-1′ CH bonds. (Left): SL5a (red); (right): SL5b (light blue). Measurements performed at 298 K. Sequence shown on top and bottom. Highlighted regions to identify the bulge region (color) and hexaloop (gray). A value of 1.2 was chosen as the threshold for dynamic evaluation and therefore visual orientation (dotted line). For the extended version with C2, see Supporting Information Figure S5.
Analysis of the hetNOE data of SL5b shows an overall rigid stem structure with most nucleotides having hetNOE values around 1.1 for the aromatic as well as the C1′H1′ resonances (Figure 3). The exception is A235, which shows slightly increased dynamics for the C1′H1′ resonance (hetNOE: 1.27) and significantly increased dynamics for the aromatic C8H8 resonance (hetNOE: 1.39), which is in line with its unpaired state. Significant dynamics is observed for the loop nucleotides. The largest hetNOE value of 1.35 is observed for U240. Consistent with the CS differences observed for the two RSMs of 5a and 5b, also the dynamics within the RSMs differ between both RNAs.
NMR Reveals Local Stability Differences for Base-Paired Regions
Not only does the detectability of imino proton signals in RNA indicate protection against solvent exchange, most often due to its involvement in stable hydrogen bonds their 1H,15N “fingerprint” chemical shift also identifies the type and even the sequence context of the precise WC base pair it is part of.54 Here, we define a canonical WC base pair as the detected formation of the hydrogen bond by the donor–acceptor pair G N1–H1···N3 C or A N1···H3–N3 U. To quantitatively analyze the (stable) WC base pairs, we determined 2hJ(N,N) constants (Supporting Information Figure S3, diamonds). The values for 2hJ(N,N) couplings agree with the literature in size and temperature dependence.30,55 For WC AU base pairs, we observe higher values between 6.5 and 7 Hz, while couplings for WC GC are in the range of 4.5–6 Hz. Values decrease in size with a rising temperature. The values are an indicator for distances between the donor and acceptor nitrogen atoms within the base pair and therefore report on the strength of the hydrogen bond. For the stem nucleotides involved in stable base pairs, we only see small changes across the tested temperature range. This is in contrast to the modulation of the cross peak intensities of the diagonal and cross peak intensities in the HNN–COSY experiment. Quantification of the peak intensities shows that the base pairs have different stabilities. Thus, while the size 2hJ(N,N) remains constant for the population of the closed base pair, the fraction of closed base pairs changes with temperature.
To determine individual stabilities of imino sites involved in hydrogen bonding, the exchange rates of individually resolved imino hydrogen with bulk water were measured.31 ΔHdiss and ΔSdiss (enthalpy and entropy of base-pair dissociation) were extracted by fitting of exchange rates kex of HSQC-imino signal intensities at temperatures between 278 and 313 K. Gibbs free enthalpies of dissociation ΔGdiss for individual base pairs were determined (Figure 4). Imino hydrogen exchange for SL5a shows stable AU base pairs with ΔGdiss of about 16 kJ/mol (U192–A215) to 10 kJ/mol (A196–U209) (Figure 4). Overall, the ΔGdiss for helical context GC base pairs was higher (∼20 kJ/mol) in comparison to the AU and GU (∼16 kJ/mol) ones. The dissociation energy was lowered for the base pairs at terminal, bulge, or loop locations. Note that ΔGdiss is higher for more stable base pairs.
The implemented inversion recovery experiment relies on the detectability of imino resonances due to their solvent protection. As a consequence, these experiments do not allow delineation of hydrogen bond stabilities for those residues which are only transiently involved in hydrogen bonds, exhibiting proton exchange rates faster than ∼100 s–1,33 since their solvent exchange is too rapid and imino signals become no longer detectable. We thus also recorded low-γ-detected 13C,15N-HSQCs (short: CN-HSQC) that do not require 1H excitation or detection which thus allowed us to detect transient hydrogen bonds with faster solvent exchange rates located within the RNA loop and bulge regions. The relative strengths of exchange protection can be qualitatively assessed by letting 15N in-phase coherence evolve under 1J(H,N) during an INEPT-based proton filter.33 In the absence of exchange, the sign of the cross peak evolves according to cos(π1J(H,N)Δ), while solvent exchange of the hydrogen imino protons covalently bound to the imino nitrogen leads to solvent-exchange-induced self-decoupling.56 The signal signature of the 1H-filtered CN-HSQC pending on the applied proton filter is exemplarily shown in Figure 4. A differentiation of the imino proton state can be made via the measurement of an echo delay corresponding to the 1J(H,N) of the imino bond (Δ = 5.3 ms for 1/1J(N,H) and Δ = 10.6 ms for 2/1J(N,H)). The signal is modulated in its sign according to its exchange protection: protected protons experience sign change, while no sign change or no corresponding signal is a result of an exchangeable state. The complete CN-HSQC for both constructs is shown in Supporting Information Figure S6. Additionally, the intensities are recovered in the experiment with Δ = 10.6 ms. The extent of recovered intensity is dependent on exchange protection and stability of the hydrogen bond. The readout of the experiment is shown in Figure 4, exemplified for the unpaired loop residue U202 which displays rapid imino proton exchange, in contrast to one of the canonical stem residues U191 which is involved in a stable WC base pair and can also be observed in the HNN–COSY (Supporting Information Figure S3). As expected, all canonical stem G and U residues of residues 5a and 5b with a detectable 2JNN coupling and measurable hydrogen exchange rates also show imino proton solvent protection in the CN-HSQC, notably including residue G242 in the loop of residue 5b. Residues with undetectable imino protons, however, do not show a uniform behavior: Residues U201, U202, and U205 of 5a, as well as their counterparts U239, U240, and U243 of 5b, carry rapidly exchanging imino protons. In 5a, U200 and G204, however, show transient protection, and the same is observed for U238 in 5b. The nature of the potential pairwise interaction of U200(/U238) with G204(/G242) remains speculative at this point.
The collective experimental data yield consistent hydrogen bonding for the bulge region of SL5a. With spectra obtained at 278 K (Supporting Information Figure S1C), and the information gained by the CN-HSQC and hydrogen exchange data, the assumed U194–U212 wobble base pair could be verified. U194 has a ΔGdiss of 9 kJ/mol, while its partner nucleotide U212 shows a more labile hydrogen bond, which is stabilized only by 4 kJ/mol and displays a transient character as evidenced by the CN-HSQC (Supporting Information Figure S7 top). U195 and U211 display transient protection of their imino protons but were not detectable in the hydrogen exchange experiment. Therefore, we conclude that their dissociation free energies are below 4 kJ/mol. G210 of the 5a bulge, in line with its elevated hetNOE values (Figure 3), shows rapid solvent exchange of its imino proton in the CN-HSQC.
NMR Solution Structures of SL5a and SL5b Contain Structurally Flexible Motifs
We describe the lowest-energy structured states in the dynamic ensemble of SL5a and SL5b. Several iterations of the structure calculations were performed. After each step, additional experimental data were integrated and energy-minimized. Structure models were used as starting points to build the final models, each consisting of 10 structures.
For the structure determination of SL5a, initial restraints for those nucleotides involved in hydrogen bonding were retrieved from 1H,15N-TROSY, 13C,15N-HSQC, and 1H,15N–HNN–COSY experiments (see the previous section). We loosely restrained U194 and U212 to form a hydrogen bond (Supporting Information) but left the nucleobases U195, G210, and U211 unrestrained due to faster exchange of these imino protons. Standard values for dihedrals for A-form helices were used for the upper and lower stems (nts g-188–193; 196–199; 206–209; 213-218-c). In addition, based on carbon chemical shift assignments, canonical coordinates were determined,22 and the ribose puckers for stem nucleotides U194, U195, U211, and U212 were set to adopt C3′-endo conformation. The characteristic chemical shift signature the UUCG tetraloop in aromatic, amino, and ribose signals provided preliminary loose restraints of the dihedral angles for the corresponding loop residues.57 All dihedral angles were continuously adapted based on w3DNA analysis, yielding the final restraints shown in Supporting Information for SL5a and Supporting Information for SL5b.
Initially, three 1H,1H-NOESYs, measured in 95% H2O, 5% D2O with 75 ms, 100 ms, and 150 ms mixing times, and two 1H,1H-NOESYs, measured in 100% D2O with 75 ms and 100 ms mixing times, were analyzed. In general, the bundle of 20 structures adopts a similar structure. Aligning the lower and the upper stem separately shows well-aligned stems, resulting in RMSDs of ∼1 Å. The bulge area comprising U195, U211, and G210, which steers the relative orientation of the two stems to be around 160°, shows the largest differences between the structures. Since the orientation cannot be deduced based on NOEs alone, RDCs that provide long-range orientation-dependent structural information were measured and included in subsequent structure calculations.
A total of 64 unique RDC values were measured in 1H,13C-IPAP-HSQCs. Initially, only a subset of the 42 unique RDCs were used to refine the calculated structure bundle. The initially implemented 42 RDCs stem from nucleotides exclusively in the base-paired regions and not from the bulge or loop regions. The rhombicity and axial components were determined using PALES and were interactively improved. During these iterations, the respective orientation of both stems was refined significantly. In the second step, the structure was further refined by including the RDCs of the bulge nucleotides. As a result, we were able to discard structure motifs featuring a strong S shape between U195 and A196. Furthermore, it was obvious from the back-calculated RDCs that implementing restraints for the nucleobase adopting an inward position brings U195, G210, and U211 in close proximity, which was required to adequately fulfill the RDCs of the bulge. Specifically, the flipped-out orientation of either of the three nucleobases was ruled out by our RDC data. Being aware of missing imino proton protection for G210 but based on chemical shift pattern of its exocyclic amino group, we restrained this residue by using its amino group as a hydrogen bond donor instead and introduced ambiguous H-bond restraints for U195 and U211, respectively (Supporting Information Table S3). The final correlation between the input and back-calculated RDCs by PALES is R2 = 0.99 for the final 10-structure ensemble (Table 1). In summary, we conclude that the predominant conformation in the structural ensemble of SL5a represents an overall continuous helical structure.
Table 1. NMR and Structural Statistics for Structure Determination of SL5a and SL5ba.
| construct | SL5a | SL5b |
|---|---|---|
| number of nucleotides (nt) | 33 nt | 29 nt |
| NMR experimental restraints | ||
| total number of experimental restraints | 816 | 698 |
| average no. of restraints per nucleotide | 24.7 | 24.1 |
| average no of NOE-derived restraints per nt | 11.75 | 10.38 |
| NOE-derived restraints | 388 | 301 |
| intraresidue | 223 | 173 |
| inter-residue | 166 | 127 |
| sequential |i–j| = 1 | 118 | 83 |
| nonsequential |i–j| > 1 | 48 | 45 |
| dihedral restraints | 284 | 261 |
| hydrogen-bonding restraints | 74 or 82 | 60 |
| planarity restraints | 14 | 24 |
| RDCs | 42 | 52 |
| structure analysis of 10 lowest-energy structure bundle | ||
| av R2 RDC back-calculated vs RDC observed | 0.99 | 0.98 |
| χ2to SAXS data | 4.66 | 7.49* |
| all atom r.m.s.d. from mean structure, Å | 3.9 | 3.4 |
| structure analysis best model | ||
| R2 RDC back-calculated vs RDC observed | 0.99 | 0.98 |
| χ2to SAXS data | 2.11 | 4.74* |
Dihedrals and hydrogen bond restraints for final calculation can be found in Supporting Information Table S3 (SL5a) and Table S4 (SL5b). (*SAXS was conducted on the native RNA sequence of SL5b (25 nt), therefore the ensemble was reduced by two stabilized GC pairs; for scattering curves, see Supporting Information Figure S8).
We used RNApdbee58−60 to describthe SL5a NMR ensemble based on the Leontis–Westhof classification61 (Figure 5). The final bundle of 10 structures displays three options that agree with our experimental results for nucleobase orientations for the nucleotides U195, G210, and U211. As shown in Figure 5, U195 and G210 can form either a trans WC/Sugar edge (option 1, occupied by 2 out of 10 states) or trans Hoogsteen/sugar edge (option 2) interaction, while either a trans WC/Hoogsteen (options 2 and 3) is detected between U195 and U211 or as displayed in option 1, U211 is stacked between G210 and U212 without identifiable hydrogen bonds. In line with our previous observations, the imino group of G210 is never involved in a direct hydrogen bond, but in 8 out of 10 states, it is its amino group instead. This is in line with the absence of any imino signal for G204. The structure evaluation by RNApdbee found involvement of the amino group instead. Additionally, our calculations showed that the imino proton of U211 was also not involved in any hydrogen bond. The cis WC/WC base pair for U194 and U212 is restrained in the final structure calculation, since we observed consistent convergence in the early ambiguously restrained calculations. All states are in agreement with the full set of back-calculated RDCs (R2 = 0.982 to 0.989). Comparison of the final bundle to a SAXS curve of SL5a showed a χ2 of 4.66, further validating this structural ensemble Table 1 and Supporting Information Figure S8).
Figure 5.
(left) Representation of the final 10 best structures for SL5a. Bulge highlighted in red; loop set to be transparent due to dynamic behavior. Structure calculations with ARIA used experimental data recorded at 298 K. Shown are structures with RDC incorporated. (right) Depiction of calculated possible hydrogen bond interaction patterns for the bulge region of SL5a. Analysis performed with the RNApdbee Web server. Schematic representation and structure details colored the same for each nucleobase. Symbols of base pairs according to the Leontis-Westhof classification.
For the structure determination of SL5b, we conducted NOE experiments using an unlabeled sample without terminal GC stabilization. Assignment transfer from BMRB 51138 yielded additional assignments summarized in the first paragraph of the methods section. For SL5b, initial NOE-based calculations were performed using five 1H,1H-NOESY spectra. Two spectra were measured in NMR buffer with 95% H2O, 5% D2O using mixing times of 100 and 150 ms, and three spectra were measured in NMR buffer with 100% D2O spectra with mixing times of 75, 100, and 150 ms.
With information gained from the herein extended chemical shift assignments, 13C,15N-HSQC, canonical coordinates (data see ref (23)), and hetNOE as well as NOE data, input for the experimental restraints was chosen. Initial restraints for this construct contained a UG base pair in the hexaloop as well as UUCG-like dihedral angles based on chemical shift similarity (Supporting Information Tables S4) for the loop. Those were refined in the iterative process where converged structures were analyzed by w3DNA. The backbone and ribose dihedral angles of the bulged nucleotide A235 and neighboring U234 and G236 were left unrestrained, since we observed flexibility of these residues in our hetNOE data. Distance restraints from 1H,1H-NOESY of 150 ms and a 1H,1H-xfilter-NOESY of 250 ms mixing times using a 13C,15N-labeled GC-stabilized sample and RDCs were incorporated for the structure calculation. 79 unique RDCs were measured in 1H,13C-IPAP-HSQCs for aromatic and C1′H1′ resonances and 1H,15N-IPAP-TROSY for imino NH resonances. For structure calculations of GC-stabilized SL5b, 52 CH-RDCs were finally used. The axial component as well as the rhombicity was determined using PALES in an iterative approach (see Materials and Methods).
The RDC incorporation resulted in convergence of the inward conformation for bulge A235 (Figure 6). Without the A235 C8H8 RDC, we observed a significantly higher number of different states for A235, which yielded an overall lower agreement with back-calculated RDCs (R2 < 0.90). Final structure bundles are in good agreement with back-calculated RDCs (R2 = 0.98) and show that, albeit inward pointing, bulge nucleotide A235 is still flexible compared to other stem nucleobases, in line with its elevated hetNOE value (Figure 3 and Supporting Information Figure S5). Its inward conformation is supported by ∼10 NOE contacts, including the A-form-typical A235-H2 to G246-H1′ sequential NOE. Comparison of the final bundle to SAXS curves showed a χ2 of 7.49 (Supporting Information Figure S8).
Figure 6.
(left) Representation of the final 10 best structures for SL5b. Bulge highlighted in blue; loop set to be transparent due to dynamic behavior. Structure calculations with ARIA used experimental data recorded at 298 K with ARIA. Shown are structures with RDC incorporated. (right) Depiction of the bulge A235 (blue) of SL5b in relation to surrounding stem nucleobases (gray). Analysis for base pairs performed with the RNApdbee Web server. Symbols according to the Leontis–Westhof classification.
Our results for SL5b indicate that the bulge region around A235 is structurally more defined compared with the bulge region of SL5a. The hexaloop for SL5b is likewise less dynamic than for SL5a, since the G242 is stabilized by a detectable hydrogen bond, suggesting the formation of a UG base pair (U238–G242). In line with its strong H1′-H8 NOE, structures consistently show G242 to adopt syn conformation, which is also observed for the corresponding loop residue in the 2KOC structure.37
SL5a and SL5b Solution Structures Add High-Resolution Information to the Cryo-EM Structure of SL5
Cryo-EM studies of SL56,11 show the stems of SL5a and SL5b to be horizontally stretched, only connected via their basal stems at the junction region. We here provide NMR evidence of the absence of any tertiary interaction between the subelements also under solution conditions. To compare our NMR structures with the cryo-EM structures available in the literature, we used the highest-resolution cryo-EM density map publicly available (EMD-42818) and its associated atomic structure bundle (PDB 8UYS) as in ref (11).
Employing a rigid-body fitting described in the methods section, we derived two values to assess the quality of the fitting of our NMR structures into the cryo-EM experimental data (overlay see Supporting Information Figure S9): (a) correlation of the number of atoms fitted into the cryo-EM 3D map relative to the total number of atoms in our bundles and (b) correlation of the goodness of the fitting of those atoms docked into the map. For SL5a, we obtained average values of 0.70 ± 0.04 and 0.38 ± 0.02 for (a) and (b), respectively. For SL5b, the average values were 0.67 ± 0.03 and 0.41 ± 0.01 for (a) and (b), respectively (see also Supporting Information Table S5). These analyses demonstrate that even with conservative fitting approaches, there is a clear correlation between our NMR structures and the available cryo-EM data.
Inspired by our NMR data on SL5a and SL5b in the full SL5 context, we also took a more comprehensive look at the full-length SL5 three-dimensional shape in solution. We used SAXS data recorded under given NMR conditions to compare them to the available cryo-EM structures of SL56,11 (Supporting Information Figures S8, S9 and S10). In accordance with the almost identical construct size, we find the expected geometric dimensions Dmax and Rg of PDB entry 8QO5 well-represented by our SAXS-derived solution data. A CRYSOL-based comparison of experimental SAXS data with theoretical scattering from 8QO5 yielded a χ2 value of 6.97 (Supporting Information Figure S8). In line with this, a denovo built dummy model shape (Figure S10) excellently represents the dimensions and fold of 8QO5 and, importantly, supports the independently oriented arrangement of elements 5a and 5b within the full SL5 element seen by NMR. Altogether, these data provide evidence for a full support of the suggested SL5 structures in solution and underscore the complementary potential of NMR spectroscopy, SAXS, and cryo-EM in the structural analysis of larger and complex folded RNAs.
Structural and dynamic characteristics of SL5a and SL5b are retained in full-length 5_SL5. To compare the structural dynamics of the SL5a and SL5b + c subelements with the full-length SL5 construct, only a subset of signals can be used. This restriction is particularly due to the size of the full-length SL5, leading to insufficient chemical shift resolution. The overlay and comparison of the 2hJ(N,N) by 1H,15N–HNN–COSY Figure 7 shows the comparison of the hydrogen bond coupling constant selected base pairs. Most of the values show significant similarity, demonstrating the power of the divide-and-conquer strategy.
Figure 7.
Comparison of 2hJ(N,N) values for the subconstructs (SL5a red, SL5b + c blue) with the full-length element (fl-SL5 gray) by assignment transfer via reporter signals (#1–10). 1H,15N–HNN–COSY experiments performed at 15N-labeled samples in a temperature range of 278–313 K. Exemplary overlay shown for 293 K. Note: Both positive and negative contour levels were colored the same for visual reasons.
Similarly, we observed remarkable agreement between the dissociation energies of evaluated base pairs in SL5 compared to the isolated RNAs (Figure 8). For example, U234, directly adjacent to the bulge in SL5b, has the values ΔGdiss(SL5b) = 10.6 kJ/mol, ΔGdiss(SL5b + c) = 10.2 kJ/mol, and ΔGdiss(SL5) = 10.6 kJ/mol (Supporting Information Figure S11).
Figure 8.
Comparison of ΔGdiss values for the subconstructs SL5a (red) and SL5b + c (blue) to the full-length element SL5 (gray) by assignment transfer via reporter signals. Hydrogen exchange measured at 15N- or 13C,15N-labeled samples in a T range of 278–313 K. For exemplary spectrum overlay for reporter selection, see Supporting Information Figure S12.
VoC Mutations in SL5a and SL5b Resulted in Subtle Changes on Transient Structure Motifs
Taking into account that a large number of mutations have evolved predominantly in the coding regions of the SARS-Cov-2 genome, we were intrigued by the appearance of two of the most abundant mutations within the 5′-untranslated region. The mutation G210U in SL5a occurred within the delta variant but reverted to G210 in the later variants, while the mutation of C241U in SL5b first occurred as early as January 2020 and manifested in all variants since (GISAID database, https://gisaid.org/hcov19-variants/).
Architecture of the Bulge Motif Is Significantly Altered in SL5a_G210U
The G210U mutant of SL5a (“SL5a_G210U”) reveals some striking differences in the pattern of protected hydrogen bonds in the 1H,15N-TROSY spectra. In detail, SL5a_G210U shows one additional N1H1 G-wobble signal at lower temperatures (Supporting Information Figure S1C) assigned to G204, a signal which was not observed in wild-type (wt) SL5a. Further, in SL5a_G210U, an additional wobble U residue is detected, while the wt signal from the A196–U209 base pair is no longer observable. The largest chemical shift changes between SL5a and SL5a_G210U are detected for the ribose C1′H1′ of the nucleotides in the bulge: U195, A196, U211, and U212 (Figure 9). This is also found for the aromatics of A196 as well as U209. In contrast, the aromatic signals of the SL5a_G210U hexaloop as well as the two adjacent GC base pairs show only minor changes between SL5a wt and SL5a_G210U. The only hexaloop residue showing significant chemical shift differences from wt is U200, indicating conformational changes in the backbone of remote loop nucleotides originating from the changes in the bulge.
Figure 9.
Comparison of CSP of the VoC to the Wuhan sequence constructs SL5a (red) and SL5b (blue). Atoms of nucleobases colored for C6/8-H6/8 values; ribose and backbone atoms colored for C1′-H1′ values. The CSPs are mapped on exemplary structures of SL5a′s and SL5b′s final bundles of 20 lowest-energy models. The mutated nucleotides are colored in black. Secondary structure representations to show the formed base pairs. For spectra and values of CSP, see Supporting Information Figures S13 and S14.
We further characterized the altered bulge motif of SL5a_G210U by hetNOE (Supporting Information Figure S5), 2hJNN (Supporting Information Figures S15 and S16), CN-HSQC (Supporting Information Figures 17 and 18), and hydrogen exchange (Supporting Information Figure S11) and compared the results to our wt data. From solvent exchange experiments, we observe that the U194–U212 base pair is further stabilized (ΔGdiss↑), which concomitantly results in an increased dissociation free energy of the adjacent U195. Both U194 and U195 show decreased flexibility as their hetNOEs are comparable to A-helix-like values (<1.2) (Supporting Information Figure S5). In line, the CN-HSQC shows both U194 and U195 to be involved in stable hydrogen bonds, since a sign change can be detected in the intensity trace (Supporting Information Figure S17). However, the 3′-nucleotides U209, U211, and U212 show transient involvement in hydrogen bonds, while U210 shows no imino proton protection.
The C241U mutation of SL5b destabilizes the hexaloop. Chemical shift differences in SL5b_C241U are most pronounced around the location of the mutation at position 241 in the hexaloop. G242 C1′H1′ also displays the highest chemical shift difference to wt with a value of 0.62 ppm (Figure 9). The C241U mutation also results in the observation loss of the C8H8 resonance of G242. Most importantly, in the 1H,15N-TROSY of the SL5b_C241U, the G242 N1H1 signal is no longer observable (Supporting Information Figure S1C), in line with a destabilization of the former U238–G242 base pair observed in the hexaloop of wt SL5b. The loss of this hydrogen bond is validated in the intensity trace analysis for CN-HSQC (Supporting Information Figures S17 and S18), where the behavior of G242 becomes more like that of wt SL5a_G204. Likewise, hetNOE values (Supporting Information Figure S5) report on increased flexibility for U240 and U243 in SL5b_C241U.
While CN experiments showed no base-pair stability changes within the stem, the comparison of hetNOE values shows a slight decrease for the bulge A235 C8H8, while C2H2 is line-broadened in the spectrum and no longer detectable. However, the U234 C1′H1′ hetNOE is increased, thus reflecting a difference in the nature of bulge dynamics inferred by the altered loop architecture in SL5b_C241U (SI Figures S5 and Figure 10).
Figure 10.
Proposed changes in loop architecture based on experimental findings gathered by 1H,15N-TROSY (278 K) and 13C,15N-HSQC (298 K) experiments. The mutations cause changes in the chemical shifts and stability of the loops. Base-pair formation indicated by dotted lines. The prevalent C241U mutation in SL5b appears SL5a-like. Mutations are indicated by colored boxes in schemes. [Note: chemical shift difference due to temperature difference.].
To conclude, a comparison of the structural and dynamical properties derived from our NMR experiments shows a similarity for SL5b_C241U to SL5a, while the RSM of SL5a_G210U more resembles the one of SL5b, with restricted dynamics and its protected loop hydrogen bond (Figure 10).
Conclusions
The genomic location of SL5, guarding the AUG start codon of the open reading frames, renders SL5 an important target for future pharmacological attempts, given its conservation of sequential composition among several genera of coronaviruses. In this work, we presented detailed insight in the subelements SL5a and SL5b, the main branches of the T-shaped SL5. Our NMR data provides a comprehensive understanding of the hydrogen bond pattern within elements as well as their individual stabilities. We show that under near-native conditions, different and partly transient interactions are present in the bulge of SL5a as compared to the structures solved by cryo-EM.6,11 Despite the sequence identity of the SL5a and SL5b RSM, their different stems lead to different loop dynamics. We attribute these effects mainly to the differences in the frequency of hydrogen bond formation present in SL5a and SL5b, respectively.
The formation of the hydrogen bond of U238 to G242 in the SL5b loop also occurs in the UUCG tetraloop of 2KOC.37 The initially assumed similarity of both the SL5a and SL5b hexaloops to 2KOC was based on the comparison of chemical shift characteristics and canonical coordinates22,23 and refined according to our experimental results. Interestingly, for SL5a and SL5b, the architecture of the loop is register-shifted, rather resembling a pentaloop with tetraloop character, as previously shown in refs (62) and (63). We also observed dynamic long-range effects in the VoCs with mutations in the bulge of SL5a and the loop of SL5b, respectively. The G210U mutation in SL5a results in stabilization of the formerly highly dynamic loop and bulge. In contrast, the original SL5b hexaloop contained a UG base pair, reducing the dynamics of the loop. This was altered by the early C241U mutation, which renders the hexaloop more dynamic and more similar to the SL5a hexaloop, as previous studies established that even some of the smallest (tetra)loops show significant dynamics.64
The spatial arrangement of SL5a and SL5b appears conserved and lacks intramolecular interactions of the loops within the full SL5 element. This was previously shown by two independent cryo-EM studies6,11 and further confirmed in this manuscript both by NMR and SAXS. Collectively, SAXS confirms the overall shape of SL5 observed by cryo-EM and thus manifests a justified link between global folds under cryogenic and native conditions. Our NMR-derived model of the 5a bulge architecture is slightly different from the cryo-EM study 8UYS that, by w3DNA analysis of all deposited structures, suggested that G210 is forming a WC-type base pair with U195, which would result in some sort of imino proton protection for G210, which we do not observe in any of our experiments.
Despite numerous previous works on establishing the secondary and tertiary structure of 5_SL5, the precise interactions within its highly conserved and structured subelements have remained poorly understood.7−9,65 With our work, we provide a complementary contribution to a detailed understanding of the SL5 system for future investigations and the fight against coronaviral diseases.
Acknowledgments
We thank Marie Hutchison for critical reading of the manuscript. We acknowledge the excellent technical support by Elke Stirnal and Kerstin Witt. We also thank Lucas Planer and Nils Kehrer for wet lab support, and Karthikeyan Dhamotharan for organizing SAXS measurements and remote data collection.
Glossary
Abbreviations
- SARS-CoV-2
severe acute respiratory syndrome coronavirus 2
- NMR
nuclear magnetic resonance (spectroscopy)
- SL
stem-loop
- ORF
open reading frame
- cryo-EM
cryogenic electron microscopy
- (het)NOE
(heteronuclear) nuclear Overhauser effect
- IPAP
in-phase antiphase
- RDC
residual dipolar coupling
- SAXS
small-angle X-ray scattering
- CSP
chemical shift perturbation
- 2hJ(N,N)
scalar coupling across hydrogen bond
Supporting Information Available
The Supporting Information is available free of charge at https://pubs.acs.org/doi/10.1021/jacs.4c08406.
Author Contributions
# K.R.M. and A.O. contributed equally to this work.
This work was supported by state of Hesse (BMRZ). Funding for the work comes from the IWB-EFRE-program 20007375 of the State of Hesse, the Deutsche Forschungsgemeinschaft (DFG) through CRC 902: “Molecular Principles of RNA-based regulation” (Project number 161793742) and through individual grant (project number 495006306 H.S.). SAXS measurements were facilitated by the BAGs with numbers SAXS990, 1102, and 1106 funded by DESY/EMBL Hamburg. The authors JCP and JMC acknowledge the financial support from the Ministry of Science, Innovation and Universities (BDNS n. 716450) to the Instruct Image Processing Center (I2PC) as part of the Spanish participation in Instruct-ERIC, the European strategic infrastructure project (ESFRI) in the area of structural biology. AS was funded by DFG through grant numbers SCHL2062/2-1 and 2-2 as well as by the Johanna Quandt Young Academy at Goethe University (grant number 2019/AS01).
The authors declare no competing financial interest.
Supplementary Material
References
- Wacker A.; Weigand J. E.; Akabayov S. R.; Altincekic N.; Bains J. K.; Banijamali E.; Binas O.; Castillo-Martinez J.; Cetiner E.; Ceylan B.; Chiu L.-Y.; Davila-Calderon J.; Dhamotharan K.; Duchardt-Ferner E.; Ferner J.; Frydman L.; Fürtig B.; Gallego J.; Grün J. T.; Hacker C.; Haddad C.; Hähnke M.; Hengesbach M.; Hiller F.; Hohmann K. F.; Hymon D.; de Jesus V.; Jonker H.; Keller H.; Knezic B.; Landgraf T.; Löhr F.; Luo L.; Mertinkus K. R.; Muhs C.; Novakovic M.; Oxenfarth A.; Palomino-Schätzlein M.; Petzold K.; Peter S. A.; Pyper D. J.; Qureshi N. S.; Riad M.; Richter C.; Saxena K.; Schamber T.; Scherf T.; Schlagnitweit J.; Schlundt A.; Schnieders R.; Schwalbe H.; Simba-Lahuasi A.; Sreeramulu S.; Stirnal E.; Sudakov A.; Tants J.-N.; Tolbert B. S.; Vögele J.; Weiß L.; Wirmer-Bartoschek J.; Wirtz Martin M. A.; Wöhnert J.; Zetzsche H. Secondary structure determination of conserved SARS-CoV-2 RNA elements by NMR spectroscopy. Nucleic Acids Res. 2020, 48 (22), 12415–12435. 10.1093/nar/gkaa1013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rangan R.; Zheludev I. N.; Hagey R. J.; Pham E. A.; Wayment-Steele H. K.; Glenn J. S.; Das R. RNA genome conservation and secondary structure in SARS-CoV-2 and SARS-related viruses: a first look. RNA (New York, N.Y.) 2020, 26 (8), 937–959. 10.1261/rna.076141.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miao Z.; Tidu A.; Eriani G.; Martin F. Secondary structure of the SARS-CoV-2 5′-UTR. RNA biology 2021, 18 (4), 447–456. 10.1080/15476286.2020.1814556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manfredonia I.; Nithin C.; Ponce-Salvatierra A.; Ghosh P.; Wirecki T. K.; Marinus T.; Ogando N. S.; Snijder E. J.; van Hemert M. J.; Bujnicki J. M.; Incarnato D. Genome-wide mapping of SARS-CoV-2 RNA structures identifies therapeutically-relevant elements. Nucleic Acids Res. 2020, 48 (22), 12436–12452. 10.1093/nar/gkaa1053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan T. C. T.; Allan M. F.; Malsick L. E.; Woo J. Z.; Zhu C.; Zhang F.; Khandwala S.; Nyeo S. S. Y.; Sun Y.; Guo J. U.; Bathe M.; Näär A.; Griffiths A.; Rouskin S. Secondary structural ensembles of the SARS-CoV-2 RNA genome in infected cells. Nat. Commun. 2022, 13 (1), 1128. 10.1038/s41467-022-28603-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Moura T. R.; Purta E.; Bernat A.; Martín-Cuevas E.; Kurkowska M.; Baulin E. F.; Mukherjee S.; Nowak J.; Biela A. P.; Rawski M.; Glatt S.; Moreno-Herrero F.; Bujnicki J. M. Conserved structures and dynamics in 5′-proximal regions of Betacoronavirus RNA genomes. Nucleic Acids Res. 2024, 52, 3419–3432. 10.1093/nar/gkae144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ziv O.; Price J.; Shalamova L.; Kamenova T.; Goodfellow I.; Weber F.; Miska E. A. The Short- and Long-Range RNA-RNA Interactome of SARS-CoV-2. Mol. Cell . 2020, 80 (6), 1067–1077.e5. 10.1016/j.molcel.2020.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt N.; Lareau C. A.; Keshishian H.; Ganskih S.; Schneider C.; Hennig T.; Melanson R.; Werner S.; Wei Y.; Zimmer M.; Ade J.; Kirschner L.; Zielinski S.; Dölken L.; Lander E. S.; Caliskan N.; Fischer U.; Vogel J.; Carr S. A.; Bodem J.; Munschauer M. The SARS-CoV-2 RNA-protein interactome in infected human cells. Nat. Microbiol. 2020, 6 (3), 339–353. 10.1038/s41564-020-00846-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flynn R. A.; Belk J. A.; Qi Y.; Yasumoto Y.; Wei J.; Alfajaro M. M.; Shi Q.; Mumbach M. R.; Limaye A.; DeWeirdt P. C.; Schmitz C. O.; Parker K. R.; Woo E.; Chang H. Y.; Horvath T. L.; Carette J. E.; Bertozzi C. R.; Wilen C. B.; Satpathy A. T. Discovery and functional interrogation of SARS-CoV-2 RNA-host protein interactions. Cell 2021, 184 (9), 2394–2411.e16. 10.1016/j.cell.2021.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vögele J.; Duchardt-Ferner E.; Bains J. K.; Knezic B.; Wacker A.; Sich C.; Weigand J. E.; Šponer J.; Schwalbe H.; Krepl M.; Wöhnert J. Structure of an internal loop motif with three consecutive U•U mismatches from stem-loop 1 in the 3′-UTR of the SARS-CoV-2 genomic RNA. Nucleic Acids Res. 2024, 52, 6687–6706. 10.1093/nar/gkae349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretsch R. C.; Xu L.; Zheludev I. N.; Zhou X.; Huang R.; Nye G.; Li S.; Zhang K.; Chiu W.; Das R. Tertiary folds of the SL5 RNA from the 5′ proximal region of SARS-CoV-2 and related coronaviruses. Proc. Natl. Acad. Sci. U.S.A. 2024, 121 (10), e2320493121 10.1073/pnas.2320493121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vögele J.; Hymon D.; Martins J.; Ferner J.; Jonker H. A.; Hargrove A. E.; Weigand J. E.; Wacker A.; Schwalbe H.; Wöhnert J.; Duchardt-Ferner E. High-resolution structure of stem-loop 4 from the 5′-UTR of SARS-CoV-2 solved by solution state NMR. Nucleic Acids Res. 2023, 51 (20), 11318–11331. 10.1093/nar/gkad762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones C. P.; Ferré-D’Amaré A. R. Crystal structure of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) frameshifting pseudoknot. RNA (New York, N.Y.) 2022, 28 (2), 239–249. 10.1261/rna.078825.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhatt P. R.; Scaiola A.; Loughran G.; Leibundgut M.; Kratzel A.; Meurs R.; Dreos R.; O’Connor K. M.; McMillan A.; Bode J. W.; Thiel V.; Gatfield D.; Atkins J. F.; Ban N. Structural basis of ribosomal frameshifting during translation of the SARS-CoV-2 RNA genome. Science (New York, N.Y.) 2021, 372 (6548), 1306–1313. 10.1126/science.abf3546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toews S.; Wacker A.; Faison E. M.; Duchardt-Ferner E.; Richter C.; Mathieu D.; Bottaro S.; Zhang Q.; Schwalbe H. The 5′-terminal stem-loop RNA element of SARS-CoV-2 features highly dynamic structural elements that are sensitive to differences in cellular pH. Nucleic Acids Res. 2024, 52, 7971–7986. 10.1093/nar/gkae477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang D.; Leibowitz J. L. The structure and functions of coronavirus genomic 3′ and 5′ ends. Virus Res. 2015, 206, 120–133. 10.1016/j.virusres.2015.02.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madhugiri R.; Karl N.; Petersen D.; Lamkiewicz K.; Fricke M.; Wend U.; Scheuer R.; Marz M.; Ziebuhr J. Structural and functional conservation of cis-acting RNA elements in coronavirus 5′-terminal genome regions. Virology 2018, 517, 44–55. 10.1016/j.virol.2017.11.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen S.-C.; Olsthoorn R. C. L. Group-specific structural features of the 5′-proximal sequences of coronavirus genomic RNAs. Virology 2010, 401 (1), 29–41. 10.1016/j.virol.2010.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madhugiri R.; Fricke M.; Marz M.; Ziebuhr J. RNA structure analysis of alphacoronavirus terminal genome regions. Virus Res. 2014, 194, 76–89. 10.1016/j.virusres.2014.10.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen S.-C.; Olsthoorn R. C. L.; Yu C.-H. Structural phylogenetic analysis reveals lineage-specific RNA repetitive structural motifs in all coronaviruses and associated variations in SARS-CoV-2. Virus Evol. 2021, 7 (1), veab021. 10.1093/ve/veab021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rangan R.; Watkins A. M.; Chacon J.; Kretsch R.; Kladwang W.; Zheludev I. N.; Townley J.; Rynge M.; Thain G.; Das R. De novo 3D models of SARS-CoV-2 RNA elements from consensus experimental secondary structures. Nucleic Acids Res. 2021, 49, 3092–3108. 10.1093/nar/gkab119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnieders R.; Peter S. A.; Banijamali E.; Riad M.; Altincekic N.; Bains J. K.; Ceylan B.; Fürtig B.; Grün J. T.; Hengesbach M.; Hohmann K. F.; Hymon D.; Knezic B.; Oxenfarth A.; Petzold K.; Qureshi N. S.; Richter C.; Schlagnitweit J.; Schlundt A.; Schwalbe H.; Stirnal E.; Sudakov A.; Vögele J.; Wacker A.; Weigand J. E.; Wirmer-Bartoschek J.; Wöhnert J. 1H, 13C and 15N chemical shift assignment of the stem-loop 5a from the 5′-UTR of SARS-CoV-2. Biomol. NMR Assignments 2021, 15 (1), 203–211. 10.1007/s12104-021-10007-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mertinkus K. R.; Grün J. T.; Altincekic N.; Bains J. K.; Ceylan B.; Ferner J.-P.; Frydman L.; Fürtig B.; Hengesbach M.; Hohmann K. F.; Hymon D.; Kim J.; Knezic B.; Novakovic M.; Oxenfarth A.; Peter S. A.; Qureshi N. S.; Richter C.; Scherf T.; Schlundt A.; Schnieders R.; Schwalbe H.; Stirnal E.; Sudakov A.; Vögele J.; Wacker A.; Weigand J. E.; Wirmer-Bartoschek J.; Martin M. A. W.; Wöhnert J. 1H, 13C and 15N chemical shift assignment of the stem-loops 5b + c from the 5′-UTR of SARS-CoV-2. Biomol. NMR Assignments 2022, 16 (1), 17–25. 10.1007/s12104-021-10053-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sklenar V. Suppression of Radiation Damping in Multidimensional NMR Experiments Using Magnetic Field Gradients. J. Magn. Reson., Ser. A 1995, 114 (1), 132–135. 10.1006/jmra.1995.1119. [DOI] [Google Scholar]
- Ikura M.; Bax A. Isotope-filtered 2D NMR of a protein-peptide complex: study of a skeletal muscle myosin light chain kinase fragment bound to calmodulin. J. Am. Chem. Soc. 1992, 114 (7), 2433–2440. 10.1021/ja00033a019. [DOI] [Google Scholar]
- Lee W.; Tonelli M.; Markley J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 2015, 31 (8), 1325–1327. 10.1093/bioinformatics/btu830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zweckstetter M. NMR: prediction of molecular alignment from structure using the PALES software. Nat. Protoc. 2008, 3 (4), 679–690. 10.1038/nprot.2008.36. [DOI] [PubMed] [Google Scholar]
- Getz M. M.; Andrews A. J.; Fierke C. A.; Al-Hashimi H. M. Structural plasticity and Mg2+ binding properties of RNase P P4 from combined analysis of NMR residual dipolar couplings and motionally decoupled spin relaxation. RNA 2007, 13 (2), 251–266. 10.1261/rna.264207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wishart D. S.; Bigam C. G.; Yao J.; Abildgaard F.; Dyson H. J.; Oldfield E.; Markley J. L.; Sykes B. D. 1H. 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J. Biomol. NMR 1995, 6 (2), 135–140. 10.1007/BF00211777. [DOI] [PubMed] [Google Scholar]
- Dingley A. J.; Nisius L.; Cordier F.; Grzesiek S. Direct detection of N-HN hydrogen bonds in biomolecules by NMR spectroscopy. Nat. Protoc. 2008, 3 (2), 242–248. 10.1038/nprot.2007.497. [DOI] [PubMed] [Google Scholar]
- Steinert H. S.; Rinnenthal J.; Schwalbe H. Individual basepair stability of DNA and RNA studied by NMR-detected solvent exchange. Biophys. J. 2012, 102 (11), 2564–2574. 10.1016/j.bpj.2012.03.074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinnenthal J.; Klinkert B.; Narberhaus F.; Schwalbe H. Direct observation of the temperature-induced melting process of the Salmonella fourU RNA thermometer at base-pair resolution. Nucleic Acids Res. 2010, 38 (11), 3834–3847. 10.1093/nar/gkq124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fürtig B.; Schnieders R.; Richter C.; Zetzsche H.; Keyhani S.; Helmling C.; Kovacs H.; Schwalbe H. Direct 13C-detected NMR experiments for mapping and characterization of hydrogen bonds in RNA. J. Biomol. NMR 2016, 64 (3), 207–221. 10.1007/s10858-016-0021-5. [DOI] [PubMed] [Google Scholar]
- Farrow N. A.; Muhandiram R.; Singer A. U.; Pascal S. M.; Kay C. M.; Gish G.; Shoelson S. E.; Pawson T.; Forman-Kay J. D.; Kay L. E. Backbone Dynamics of a Free and a Phosphopeptide-Complexed Src Homology 2 Domain Studied by 15N NMR Relaxation. Biochemistry 1994, 33 (19), 5984–6003. 10.1021/bi00185a040. [DOI] [PubMed] [Google Scholar]
- Linge J. P.; O’Donoghue S. I.; Nilges M. Automated assignment of ambiguous nuclear overhauser effects with ARIA. Methods Enzymol. 2001, 339, 71–90. 10.1016/S0076-6879(01)39310-2. [DOI] [PubMed] [Google Scholar]
- Brünger A. T.; Adams P. D.; Clore G. M.; DeLano W. L.; Gros P.; Grosse-Kunstleve R. W.; Jiang J. S.; Kuszewski J.; Nilges M.; Pannu N. S.; Read R. J.; Rice L. M.; Simonson T.; Warren G. L. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 1998, 54 (5), 905–921. 10.1107/S0907444998003254. [DOI] [PubMed] [Google Scholar]
- Nozinovic S.; Fürtig B.; Jonker H. R. A.; Richter C.; Schwalbe H. High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA. Nucleic Acids Res. 2010, 38 (2), 683–694. 10.1093/nar/gkp956. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams C. J.; Headd J. J.; Moriarty N. W.; Prisant M. G.; Videau L. L.; Deis L. N.; Verma V.; Keedy D. A.; Hintze B. J.; Chen V. B.; Jain S.; Lewis S. M.; Arendall W. B.; Snoeyink J.; Adams P. D.; Lovell S. C.; Richardson J. S.; Richardson D. C. MolProbity: More and better reference data for improved all-atom structure validation. Protein science: a publication of the Protein Society 2018, 27 (1), 293–315. 10.1002/pro.3330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li S.; Olson W. K.; Lu X.-J. Web 3DNA 2.0 for the analysis, visualization, and modeling of 3D nucleic acid structures. Nucleic Acids Res. 2019, 47 (W1), W26–W34. 10.1093/nar/gkz394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pettersen E. F.; Goddard T. D.; Huang C. C.; Couch G. S.; Greenblatt D. M.; Meng E. C.; Ferrin T. E. UCSF Chimera--a visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25 (13), 1605–1612. 10.1002/jcc.20084. [DOI] [PubMed] [Google Scholar]
- Svergun D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 1992, 25 (4), 495–503. 10.1107/S0021889892001663. [DOI] [Google Scholar]
- Manalastas-Cantos K.; Konarev P. V.; Hajizadeh N. R.; Kikhney A. G.; Petoukhov M. V.; Molodenskiy D. S.; Panjkovich A.; Mertens H. D. T.; Gruzinov A.; Borges C.; Jeffries C. M.; Svergun D. I.; Franke D. ATSAS 3.0: expanded functionality and new tools for small-angle scattering data analysis. J. Appl. Crystallogr. 2021, 54 (1), 343–355. 10.1107/S1600576720013412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franke D.; Petoukhov M. V.; Konarev P. V.; Panjkovich A.; Tuukkanen A.; Mertens H. D. T.; Kikhney A. G.; Hajizadeh N. R.; Franklin J. M.; Jeffries C. M.; Svergun D. I. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 2017, 50 (4), 1212–1225. 10.1107/S1600576717007786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Svergun D. I. Restoring low resolution structure of biological macromolecules from solution scattering using simulated annealing. Biophys. J. 1999, 76 (6), 2879–2886. 10.1016/S0006-3495(99)77443-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleming P. J.; Correia J. J.; Fleming K. G. Revisiting macromolecular hydration with HullRadSAS. Eur Biophys J.: EBJ. 2023, 52 (4–5), 215–224. 10.1007/s00249-022-01627-8. [DOI] [PubMed] [Google Scholar]
- Bowman J. C.; Lenz T. K.; Hud N. V.; Williams L. D. Cations in charge: magnesium ions in RNA folding and catalysis. Curr. Opin. Struct. Biol. 2012, 22 (3), 262–272. 10.1016/j.sbi.2012.04.006. [DOI] [PubMed] [Google Scholar]
- Favier A.; Brutscher B. Recovering lost magnetization: polarization enhancement in biomolecular NMR. J. Biomol. NMR 2011, 49, 9–15. 10.1007/s10858-010-9461-5. [DOI] [PubMed] [Google Scholar]
- Solyom Z.; Schwarten M.; Geist L.; Konrat R.; Willbold D.; Brutscher B. BEST-TROSY experiments for time-efficient sequential resonance assignment of large disordered proteins. J. Biomol. NMR 2013, 55, 311–321. 10.1007/s10858-013-9715-0. [DOI] [PubMed] [Google Scholar]
- Fürtig B.; Richter C.; Bermel W.; Schwalbe H. New NMR experiments for RNA nucleobase resonance assignment and chemical shift analysis of an RNA UUCG tetraloop. J. Biomol. NMR 2004, 28 (1), 1573–5001. 10.1023/B:JNMR.0000012863.63522.1f. [DOI] [PubMed] [Google Scholar]
- Schwalbe H.; Marino J. P.; Glaser S. J.; Griesinger C. Measurement of H,H-Coupling Constants Associated with.nu.1, nu. 2, and.nu.3 in Uniformly 13C-Labeled RNA by HCC-TOCSY-CCH-E.COSY. J. Am. Chem. Soc. 1995, 117 (27), 7251–7252. 10.1021/ja00132a028. [DOI] [Google Scholar]
- Glaser S. J.; Schwalbe H.; Marino J. P.; Griesinger C. Directed TOCSY, a Method for Selection of Directed Correlations by Optimal Combinations of Isotropic and Longitudinal Mixing. J. Magn. Reson., Ser. B 1996, 112 (2), 160–180. 10.1006/jmrb.1996.0126. [DOI] [PubMed] [Google Scholar]
- Kay L. E.; Xu G. Y.; Singer A. U.; Muhandiram D. R.; Formankay J. D. A Gradient-Enhanced HCCH-TOCSY Experiment for Recording Side-Chain 1H and 13C Correlations in H2O Samples of Proteins. J. Magn. Reson., Ser. B 1993, 101 (3), 333–337. 10.1006/jmrb.1993.1053. [DOI] [Google Scholar]
- Wagner D.; Rinnenthal J.; Narberhaus F.; Schwalbe H. Mechanistic insights into temperature-dependent regulation of the simple cyanobacterial hsp17 RNA thermometer at base-pair resolution. Nucleic Acids Res. 2015, 43 (11), 5572–5585. 10.1093/nar/gkv414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y.; Han G.; Jiang X.; Yuwen T.; Xue Y. Chemical shift prediction of RNA imino groups: application toward characterizing RNA excited states. Nat. Commun. 2021, 12, 1595. 10.1038/s41467-021-21840-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bytchenkoff D.; Chiarparin E.; Früh D.; Rüdisser S.; Bodenhausen G. Temperature dependence of internucleotide nitrogen–nitrogen scalar couplings in RNA. Magn. Reson. Chem. 2002, 40 (5), 377–379. 10.1002/mrc.1018. [DOI] [Google Scholar]
- Levitt M. H.Spin Dynamics:Basics of Nuclear Magnetic Resonance, 2.ed., reprint. With Corr; Wiley, 2009. [Google Scholar]
- Lawrence C.; Grishaev A. Chemical shifts-based similarity restraints improve accuracy of RNA structures determined via NMR. RNA 2020, 26, 2051–2061. 10.1261/rna.074617.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antczak M.; Popenda M.; Zok T.; Zurkowski M.; Adamiak R. W.; Szachniuk M. New algorithms to represent complex pseudoknotted RNA structures in dot-bracket notation. Bioinformatics 2018, 34 (8), 1304–1312. 10.1093/bioinformatics/btx783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zok T.; Antczak M.; Zurkowski M.; Popenda M.; Blazewicz J.; Adamiak R. W.; Szachniuk M. RNApdbee 2.0: multifunctional tool for RNA structure annotation. Nucleic Acids Res. 2018, 46 (W1), W30–W35. 10.1093/nar/gky314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Antczak M.; Zok T.; Popenda M.; Lukasiak P.; Adamiak R. W.; Blazewicz J.; Szachniuk M. RNApdbee--a webserver to derive secondary structures from pdb files of knotted and unknotted RNAs. Nucleic Acids Res. 2014, 42 (W1), W368–W372. 10.1093/nar/gku330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leontis N. B.; Westhof E. Geometric nomenclature and classification of RNA base pairs. RNA (New York, N.Y.) 2001, 7 (4), 499–512. 10.1017/S1355838201002515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thapar R.; Denmon A. P.; Nikonowicz E. P. Recognition modes of RNA tetraloops and tetraloop-like motifs by RNA-binding proteins. RNA 2014, 5 (1), 49–67. 10.1002/wrna.1196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee C. W.; Li L.; Giedroc D. P. The solution structure of coronaviral stem-loop 2 (SL2) reveals a canonical CUYG tetraloop fold. FEBS Lett. 2011, 585 (7), 1049–1053. 10.1016/j.febslet.2011.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oxenfarth A.; Kümmerer F.; Bottaro S.; Schnieders R.; Pinter G.; Jonker H. R. A.; Fürtig B.; Richter C.; Blackledge M.; Lindorff-Larsen K.; Schwalbe H. Integrated NMR/Molecular Dynamics Determination of the Ensemble Conformation of a Thermodynamically Stable CUUG RNA Tetraloop. J. Am. Chem. Soc. 2023, 145 (30), 16557–16572. 10.1021/jacs.3c03578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sola I.; Mateos-Gomez P. A.; Almazan F.; Zuñiga S.; Enjuanes L. RNA-RNA and RNA-protein interactions in coronavirus replication and transcription. RNA biology 2011, 8 (2), 237–248. 10.4161/rna.8.2.14991. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.